Gene expression microarray technology measures the expression levels of thousands of genes. Research Article
|
|
|
- Timothy Page
- 5 years ago
- Views:
Transcription
1 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 7, Number 2, 2 # Mary Ann Liebert, Inc. Pp. 8 DOI:.89/cmb Research Article Reducing the Computational Complexity of Information Theoretic Approaches for Reconstructing Gene Regulatory Networks PENG QIU, ANDREW J. GENTLES, and SYLVIA K. PLEVRITIS ABSTRACT Information theoretic approaches are increasingly being used for reconstructing regulatory networks from microarray data. These approaches start by computing the pairwise mutual information (MI) between all gene pairs. The resulting MI matrix is then manipulated to identify regulatory relationships. A barrier to these approaches is the time-consuming step of computing the MI matrix. We present a method to reduce this computation time. We apply spectral analysis to re-order the genes, so that genes that share regulatory relationships are more likely to be placed close to each other. Then, using a sliding window approach with appropriate window size and step size, we compute the MI for the genes within the sliding window, and the remainder is assumed to be zero. Using both simulated data and microarray data, we demonstrate that our method does not incur performance loss in regions of high-precision and low-recall, while the computational time is significantly lowered. The proposed method can be used with any method that relies on the mutual information to reconstruct networks. Key words: algorithms, computational molecular biology, machine learning.. INTRODUCTION Gene expression microarray technology measures the expression levels of thousands of genes simultaneously and provides data for reconstructing large-scale gene regulatory networks. Recently, information theoretic approaches have been used increasingly for this purpose. Examples include the relevance network (RelNet) (Butte and Kohane, 2; Butte et al., 2) ARACNE ( Margolin et al., 26a), and the maximum relevance minimum redundance network (MRNet) ( Meyer et al., 27). These approaches and others start by computing the pairwise mutual information ( MI) between all possible pairs of genes, resulting in a MI matrix. The MI matrix is then manipulated to identify regulatory relationships. For example, in relevance networks, an edge exists between a pair of genes if their MI exceeds a given specific threshold. ARACNE improves on the relevance networks by applying the data processing inequality (Cover and Thomas, 26) (DPI) to each connected gene triple, removing potentially false positive edges. In MRNet, the maximum relevance minimum redundance (MRMR) criterion is applied, where the maximum relevance Department of Radiology, Stanford University, Stanford, California.
2 2 QIU ET AL. criterion assigns edges to gene pairs that share large MI, while the minimum redundance criterion controls false positives (Meyer et al., 27; Ding and Peng, 25). The MRMR criterion is essentially a pairwise approximation of the conditional MI. The conditional MI is explicitly used for inference in Zhao et al. (28). If two genes that share large MI but are conditionally independent give a third gene, there is no edge between them. These approaches have been successfully applied to simulated data and real microarray data on B-Cell, melanoma, and have identified interesting regulatory targets and pathways. The most time-consuming step in all of the above approaches (Butte and Kohane, 2; Margolin et al., 26a; Meyer et al., 27) is computing the entire MI matrix, which requires all possible pairs of genes to be examined. The computational complexity of Zhao et al., (28) is higher because the conditional MI has to be computed for each gene triple. To reduce the computational complexity, we only compute the MI for gene pairs with expected significant values. We identify these gene pairs by applying spectral analysis (Chung, 997) to re-order the genes, so that genes that share regulatory relationships are more likely to be placed close to each other. To determine the new gene ordering, a Laplacian matrix is derived from the correlation matrix of the gene expression data, assuming the correlation matrix provides an adequate approximation to the adjacency matrix for our purpose. Motivated by the spectral decomposition in Rapaport et al. (27), we then compute the Fiedler vector, which is the eigenvector associated with the second smallest eigenvalue of the Laplacian matrix. Since the Fiedler vector is smooth with respect to the connectivity described by the Laplacian matrix (Rapaport et al., 27), we sort the elements of the Fiedler vector to obtain the desired gene ordering. After re-ordering the genes, we apply a sliding window approach with respect to the new gene ordering and compute the MI among the genes within the sliding window. Since we only compute the MI among genes within the sliding window, only part of the MI matrix is computed and the remainder is assumed to be zero. The resulting MI matrix can then be used with a variety of information theoretic approaches for reconstructing the regulatory network, including, for example, RelNet, ARACNE, and MRNet. Depending on the window size, we demonstrate that the computational complexity of computing the MI matrix can be significantly reduced, with minor loss in the accuracy of the reconstructed regulatory network. 2.. Re-ordering the genes 2. METHODS A gene regulatory network reconstructed by the information theoretic approaches (Butte and Kohane, 2; Margolin et al., 26a; Meyer et al., 27) is undirected and can be represented by an adjacency matrix. The adjacency matrix is usually sparse, and a value of at the (i,j) element represents an edge connecting gene i and gene j. Since the adjacency matrix is sparse, its rows and columns can be reshuffled (resulting in a re-ordering of the genes), so that most s are concentrated along the diagonal of the reshuffled adjacency matrix. We apply spectral analysis to re-order the genes to obtain an adjacency matrix with the s concentrated along the diagonal. For example, consider Figure a, which shows the adjacency matrix of a simulated scale-free network, containing genes and 35 edges. For the moment, assume that we know the adjacency matrix A. We define the Laplacian matrix as L ¼ D A, where D is a diagonal matrix whose diagonal elements are the degrees of the genes. In spectral graph theory (Chung, 997), the Fiedler vector, v ¼ [v, v 2,..., v n ], is defined as the eigenvector associated with the second smallest eigenvalue of the Laplacian matrix (the smallest eigenvalue is trivial). As presented in the spectral decomposition (Rapaport et al., 27), the Fiedler vector is smooth with respect to the network connectivity described by the Laplacian matrix. By smooth with respect to the network, we mean that if two nodes i and j are connected by an edge, the difference between their corresponding entries in the Fiedler vector, jv i v j j,is small. The mathematics behind the above argument is as follows. Since the Fiedler vector is the first nontrivial eigenvector associated with the smallest non-trivial eigenvalue of L, the Fiedler vector solves the following minimization problem, where the objective function can be written as follows, min v Lv () s:t: jvj¼
3 REDUCING COMPUTATIONAL COMPLEXITY 3 a Adjacency Matrix b FIG.. (a) The adjacency matrix of a simulated scale-free network, consisting of genes and 35 edges. (b) The reshuffled adjacency matrix using spectral analysis. With a sliding window of size 2 and step size equal to half of the window size, the sliding window approach covers 28% of the adjacency matrix (gray area), which contains 85% of the edges v Lv ¼ v Dv v Av ¼ X d i v 2 i ¼ ¼ i2f,..., ng X fi, j:i 4 j, A ij ¼ g X fi, j:i 4 j, A ij ¼ g X i, j2f,..., ng v i A ij v j (v 2 i þ v2 j ) X fi, j:i 4 j, A ij ¼ g 2v i v j (v i v j ) 2 (2) Therefore, the Fielder vector is also minimizes (2), which means if two nodes i and j are connected, the difference between their corresponding entries in the Fiedler vector, jv i v j j, tends to be small. We compute the Fiedler vector of the Laplacian matrix L and sort its elements in either ascending or descending order. In the gene ordering obtained from the Fiedler vector, connected genes are more likely to be placed close to each other. Figure b shows the reshuffled adjacency matrix, with most edges ( s) now close to the diagonal. In this particular example, if we apply a window of size 2 and slide the window along the new gene ordering with a step size of half of the window size, the sliding window approach covers 28% of the adjacency matrix (the gray area), but reconstructs 85% of the edges. Although we miss 5% of the edges by applying the sliding window, to ensure high-precision and low-recall, most of these missing edges would not be identified as significant even when the entire MI matrix is computed. Simulation results in Section 3 show that applying the sliding window approach does not incur performance loss in operating regions of high-precision and low-recall. In practice, the adjacency matrix is an unknown, so we need to obtain the gene re-ordering from the expression data. We proceed by assuming that the correlation matrix provides an approximation of the adjacency matrix that is adequate for our purpose, because the correlation matrix reflects the linear relationship between gene pairs and is fast to compute. We normalize each gene s expression data to zero mean and unit variance, and compute the correlation matrix by multiplying the expression data and its transpose. The power adjacency function (Zhang and Horvath, 25) is applied to each entry of the correlation matrix (y ¼jxj K ). The resulting matrix, denoted by W, is an approximation to the adjacency matrix. We define the Laplacian as L ¼ D W, where D is a diagonal matrix, chosen such that the column sums of L are zeros. Eigendecomposition of the Laplacian is performed to find the Fiedler vector, which is sorted to re-order the genes. The computational complexity of obtaining the gene ordering is negligible compared to the computation of the MI matrix. For example, for a B-Cell gene expression dataset (Margolin et al., 26a), which contains 336 samples and 9563 genes per sample, computing the entire MI matrix takes about 42 hours, using the Java package developed in Margolin et al. (26b), whereas obtaining the gene ordering using the above procedure takes less than minutes in Matlab Reducing computation time by the sliding window approach After re-ordering the genes, connected genes are more likely to be placed close to each other. We then apply a sliding window along the new ordering, and compute MI within the sliding window. This concept
4 4 QIU ET AL. can be visualized in Figure b. If we reshuffled the genes according to the ordering, the gene pairs within the sliding window correspond to the gray area, along the diagonal line of the reshuffled MI matrix. The blocks are determined by the window size and the step size of the sliding window. With an appropriate window size and step size, the computational complexity can be significantly reduced with minimal performance loss. The reduction in computational complexity can be quantified as follows. Assume that the total number of genes is n, the size of the sliding window is w, and the step size is chosen to be half of the window size. To compute the entire MI matrix, n 2 pairs of genes need to be examined. The number of gene pairs covered by 4 þ 3w2 (2n ) 4 w the sliding window is w2 3w. The ratio is approximately 2n. In the example in Figure, there are n ¼ genes, and the window size w ¼ 2. Using the sliding window, the computational complexity is reduced to 3% of that needed to obtain the entire MI matrix. The reduction in computational complexity is the result of computing only the diagonal part of the reshuffled MI matrix. Because the remaining entries of the MI matrix are set to be zeros, there is potential loss of reconstruction accuracy. In the following section, we examine a simulated dataset and a B-Cell gene expression dataset to evaluate the performance loss. 3. RESULTS 3.. Simulated data We use the data generator in Rogers and Girolami (25) to simulate artificial gene regulatory networks and expression data. The generator produces topologically scale-free networks (Barabási and Bonabeau, 23), where the number of regulatory connections for each gene is generated according to an approximate power-law distribution. A symmetric adjacency matrix is derived from the simulated topology, serving as the ground truth in the simulation. The expression data is drawn from the steady state of the system, which is evaluated by integrating a system of differential equations. The generator is able to simulate gene knockout experiments by holding the expression of the knocked-out gene at zero during the integration (Rogers and Girolami, 25). In our simulation, we use the data generator to produce gene regulatory networks and simulate the expression data of knock-out experiments for each gene. The number of genes n is chosen to be either or 4. Similar to Margolin et al. (26a) and Meyer et al. (27), we use precision and recall as performance metrics. Denote the true positives N TP, the false positives N FP, and the false negatives N FN. The precision is defined as p ¼ N TP =(N TP þ N FP ), which is the fraction of the true edges among the edges identified by the algorithm. The recall is defined as r ¼ N TP =(N TP þ N FN ), which is the ratio between the correctly identified true edges over the total number of true edges. The precision-recall curve is generated by adjusting the MI threshold in the regulatory network reconstruction algorithms. Figure 2 shows an example of a simulated dataset containing genes and 35 edges. The expression data contains samples, corresponding to the gene knock-out experiment for each gene. Figure 2a shows the simulated adjacency matrix, which is the same as Figure a. We compute the correlation matrix of the simulated expression data, and obtain the gene ordering using spectral analysis as described in Section 2. Figure 2b shows the reshuffled adjacency matrix according to the ordering, which is inferior compared to that in Figure b. This is because the ordering in Figure b is based on the knowledge of the true adjacency matrix, while the ordering in Figure 2b is based on the correlation matrix of the expression data. In Figure 2c, the performance of RelNet, ARACNE, and MRNet are compared using precision-recall curves. Both ARACNE and MRNet show superior performance to RelNet, which is consistent with the results in Margolin et al. (26a) and Meyer et al. (27). In Figure 2d f, the three methods RelNet, ARACNE, and MRNet are compared with their windowed versions win-relnet, win-aracne, and win-mrnet, respectively. We apply a sliding window along the gene ordering in Figure 2b, with a window size of and a step size of 5. Such choice of the sliding window reduces the computational complexity to 5% of that when the entire MI matrix is computed. We observe that in the high-precision low-recall regime, applying the sliding window does not incur performance loss. In some cases, applying the sliding window yields slightly better performance. In the low-precision regime, the windowed version has lower recall. However, in practice, this regime is of little value, because it is not possible to distinguish biologically meaningful edges from false positives.
5 REDUCING COMPUTATIONAL COMPLEXITY 5 Precision a Adjacency Matrix b Reshuffled Adjacency Matrix c RelNet win RelNet.2.4 Recall Precision d e f ARACNE win ARACNE.2.4 Recall Precision Precision RelNet. ARACNE MRNet.2.4 Recall MRNet win MRNet.2.4 Recall FIG. 2. Performance comparison of RelNet, ARACNE, MRNet, and their windowed version, using a simulated dataset (GData). The number of genes is. The sliding window has window size and step size 5. (a) Adjacency matrix. (b) Reshuffled adjacency matrix. (c) Comparison of RelNet, ARACNE, MRNet. (d) RelNet versus its windowed version. (e) ARACNE versus its windowed version. (f) MRNet versus its windowed version. To further demonstrate that applying the sliding window approach does not incur performance loss in the high-precision low-recall regime, we summarize the results from simulated datasets in Table. We integrate the precision-recall curve from % to 3% recall, and use the area under curve as the metric of accuracy in the high-precision low-recall regime. We normalize the metric by its maximum possible value. The first 5 simulated datasets contain n ¼ genes and an average of 5 edges. The other 5 datasets contain n ¼ 4 genes and around 55 edges. In all cases, the size of the sliding window is chosen to be w ¼ n=, and the step size is half of the window size. Therefore, applying the sliding window reduces the computational complexity to 5% of that when the entire MI matrix is computed. From Table, we observe Table. Performance of RelNet, ARACNE, MRNet, and Their Windowed Versions, When Target Is a Recall of 3% or Less, Based on Simulated Data Simulated dataset RelNet win-relnet ARACNE win-aracne MRNet win-mrnet GData GData GData GData GData Average G4Data G4Data G4Data G4Data G4Data Average
6 6 QIU ET AL. that the windowed version of RelNet, ARACNE, and MRNet have similar performance compared with their original implementation B-Cell data We tested the gene ordering and sliding window approach on a publicly available B-Cell gene expression dataset. The dataset contains 336 samples and 9563 genes per sample (Margolin et al., 26a). Since the true B-Cell gene regulatory network is unknown, precision and recall cannot be used as metric to evaluate the reconstruction performance. Therefore, we only compare the reconstructed networks by the sliding windowed approach to the original implementations of ARACNE. The gene ordering is obtained as described in Section 2. The window size is chosen to be, and the step size is half of the window size. In Table 2, we show the edges and hub nodes (genes with degree higher than 3) identified by the original implementation of ARACNE and our windowed approach. It can be observed that, when the MI threshold is high, meaning when the algorithm operates at high-precision lowrecall regime, there is little difference between the results from ARACNE and our sliding window approach. On average, win-aracne recovers 94% of the edges that ARACNE identifies, and at the same time generates <% new edges that ARACNE does not identify. win-aracne recovers 96% of the hub genes that ARACNE identifies, and does not generate new hub genes that ARACNE does not identify. The computation time for obtaining the gene ordering is less than minutes, which is negligible compared to the 42 hours needed to compute the entire MI matrix. Using the sliding window approach, we reduce the computation time by 84%, that is from 42 to 23 hours. In order to examine the effect of the window size on the reconstructed regulatory network, we fix the MI threshold to be, and vary window size from 4 to 6. For each choice of the window size, we compute what percentage of the edges and hub genes identified by ARACNE are also identified by win- ARACNE. As the window size increases, the regulatory network constructed by win-aracne first quickly approaches the network constructed by ARACNE, and then saturates. When the window size increases to 9563, the total number of genes, win-aracne and ARACNE should produce identical results. In Figure 3, we also see that the computational complexity is 3w 2n, linear in the window size w. In summary, we can see that, if we are willing to sacrifice a certain amount of the edges identified by ARACNE, we can apply a sliding window along the proposed gene ordering, and significantly reduce the computational complexity. 4. CONCLUSION When information theoretic approaches are applied to reconstruct large scale gene regulatory networks, the mutual information between all possible gene pairs needs to be computed, which is a time-consuming Table 2. Comparison of ARACNE and win-aracne on a B-Cell Dataset, which Contains 9563 Genes MI threshold Edges ARACNE win-aracne Overlaps Hub genes ARACNE win-aracne Overlaps Window size is chosen to be and the step size is half of window size. On average, win-aracne recovers 94% of the edges that ARACNE identifies, and generates <% new edges that ARACNE does not identify. win-aracne recovers 96% of the hub genes that ARACNE identifies, and does not generate new hub genes that ARACNE does not identify. Computation time is reduced to 6% of the original implementation.
7 REDUCING COMPUTATIONAL COMPLEXITY overlap edges overlap of hub genes computational complexity window size FIG. 3. Comparison of ARACNE and win-aracne, in terms of the percentage of overlapping edges and overlapping high degree nodes, and the computational complexity as a function of window size. task. In this work, we present a method to reduce this computation time. We apply spectral analysis to reorder the genes, so that genes that share regulatory relationships are more likely to be placed close to each other. Using a sliding window with appropriate window size and step size, we can apply the information theoretic approaches to examine the genes subset by subset, along the new gene ordering. Through analysis of simulation and real gene expression data, we demonstrated that, in the high-precision low-recall regime, applying the sliding window approach yields similar performance compared with the case where the entire mutual information matrix is computed, while the computational complexity is significantly reduced. ACKNOWLEDGMENTS We gratefully acknowledge funding from the NCI Integrative Cancer Biology Program (ICBP; grant U56 CA2973). No competing financial interests exist. DISCLOSURE STATEMENT REFERENCES Barabási, A.L., and Bonabeau, E. 23. Scale-free networks. Sci. Am. 288, Butte, A.J., and Kohane, L.S. 2. Mutual information relevance networks: functional genomic clustering using pairwise entropy measurements. Paci. Symp. Biocomput. 4, Butte, A.J., Tamayo, P., Slonim, D., et al. 2. Discovering functional relationships between RNA expression and chemotherapeutic susceptibility using relevance networks. Proc. Natl. Acad. Sci. USA 97, Chung, F.R.K Spectral Graph Theory (CBMS Regional Conference Series in Mathematics, No. 92). American Mathematical Society, New York. Cover, T., and Thomas, J. 26. Elements of Information Theory, 2nd ed. Wiley Series in Telecommunications and Signal Processing. Wiley-Interscience, New York. Ding, C., and Peng, H. 25. Minimum redundancy feature selection from microarray gene expression data. J Bioinform. Comput. Biol. 3, Margolin, A.A., Nemenman, I., Basso, K., et al. 26a. ARACNE: an algorithm for the reconstruction of gene regulatory networks in a mammalian cellular context. BMC Bioinform. 7. Margolin, A.A., Wang, K., Lim, W.K., et al. 26b. Reverse engineering cellular networks. Nat. Protoc., Meyer, P.E., Kontos, K., Lafitte, F., et al. 27. Information-theoretic inference of large transcriptional regulatory networks. EURASIP J. Bioinform. Syst. Biol. 27, 9. Rapaport, F., Zinovyev, A., Dutreix, M., et al. 27. Classification of microarray data using gene networks. BMC Bioinform. 8, 35þ.
8 8 QIU ET AL. Rogers, S., and Girolami, M., 25. A Bayesian regression approach to the inference of regulatory networks from gene expression data. Bioinformatics 2, Zhang, B., and Horvath, S. 25. A general framework for weighted gene co-expression network analysis. Statist. Appl. Genet. Mol. Biol. 4. Zhao, W., Serpedin, E., and Dougherty, E.R. 28. Inferring connectivity of genetic regulatory networks using information-theoretic criteria. IEEE=ACM Trans. Comput. Biol. Bioinform. 5, Address correspondence to: Dr. Peng Qiu 2 Welch Road, Lucas Center Department of Radiology Stanford University Stanford, CA qiupeng@stanford.edu
Hub Gene Selection Methods for the Reconstruction of Transcription Networks
 for the Reconstruction of Transcription Networks José Miguel Hernández-Lobato (1) and Tjeerd. M. H. Dijkstra (2) (1) Computer Science Department, Universidad Autónoma de Madrid, Spain (2) Institute for
for the Reconstruction of Transcription Networks José Miguel Hernández-Lobato (1) and Tjeerd. M. H. Dijkstra (2) (1) Computer Science Department, Universidad Autónoma de Madrid, Spain (2) Institute for
A New Method to Build Gene Regulation Network Based on Fuzzy Hierarchical Clustering Methods
 International Academic Institute for Science and Technology International Academic Journal of Science and Engineering Vol. 3, No. 6, 2016, pp. 169-176. ISSN 2454-3896 International Academic Journal of
International Academic Institute for Science and Technology International Academic Journal of Science and Engineering Vol. 3, No. 6, 2016, pp. 169-176. ISSN 2454-3896 International Academic Journal of
HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH
 HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH Hoang Trang 1, Tran Hoang Loc 1 1 Ho Chi Minh City University of Technology-VNU HCM, Ho Chi
HYPERGRAPH BASED SEMI-SUPERVISED LEARNING ALGORITHMS APPLIED TO SPEECH RECOGNITION PROBLEM: A NOVEL APPROACH Hoang Trang 1, Tran Hoang Loc 1 1 Ho Chi Minh City University of Technology-VNU HCM, Ho Chi
Graph Wavelets to Analyze Genomic Data with Biological Networks
 Graph Wavelets to Analyze Genomic Data with Biological Networks Yunlong Jiao and Jean-Philippe Vert "Emerging Topics in Biological Networks and Systems Biology" symposium, Swedish Collegium for Advanced
Graph Wavelets to Analyze Genomic Data with Biological Networks Yunlong Jiao and Jean-Philippe Vert "Emerging Topics in Biological Networks and Systems Biology" symposium, Swedish Collegium for Advanced
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
Information-Theoretic Inference of Gene Networks Using Backward Elimination
 Information-Theoretic Inference of Gene Networks Using Backward Elimination Patrick E. Meyer 1,2,3, Daniel Marbach 1,2, Sushmita Roy 1,2 and Manolis Kellis 1,2 1 Computer Science and Artificial Intelligence
Information-Theoretic Inference of Gene Networks Using Backward Elimination Patrick E. Meyer 1,2,3, Daniel Marbach 1,2, Sushmita Roy 1,2 and Manolis Kellis 1,2 1 Computer Science and Artificial Intelligence
Markov Chains and Spectral Clustering
 Markov Chains and Spectral Clustering Ning Liu 1,2 and William J. Stewart 1,3 1 Department of Computer Science North Carolina State University, Raleigh, NC 27695-8206, USA. 2 nliu@ncsu.edu, 3 billy@ncsu.edu
Markov Chains and Spectral Clustering Ning Liu 1,2 and William J. Stewart 1,3 1 Department of Computer Science North Carolina State University, Raleigh, NC 27695-8206, USA. 2 nliu@ncsu.edu, 3 billy@ncsu.edu
Spectral Clustering. Guokun Lai 2016/10
 Spectral Clustering Guokun Lai 2016/10 1 / 37 Organization Graph Cut Fundamental Limitations of Spectral Clustering Ng 2002 paper (if we have time) 2 / 37 Notation We define a undirected weighted graph
Spectral Clustering Guokun Lai 2016/10 1 / 37 Organization Graph Cut Fundamental Limitations of Spectral Clustering Ng 2002 paper (if we have time) 2 / 37 Notation We define a undirected weighted graph
A Geometric Interpretation of Gene Co-Expression Network Analysis. Steve Horvath, Jun Dong
 A Geometric Interpretation of Gene Co-Expression Network Analysis Steve Horvath, Jun Dong Outline Network and network concepts Approximately factorizable networks Gene Co-expression Network Eigengene Factorizability,
A Geometric Interpretation of Gene Co-Expression Network Analysis Steve Horvath, Jun Dong Outline Network and network concepts Approximately factorizable networks Gene Co-expression Network Eigengene Factorizability,
Introduction to Machine Learning
 10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
Iterative Laplacian Score for Feature Selection
 Iterative Laplacian Score for Feature Selection Linling Zhu, Linsong Miao, and Daoqiang Zhang College of Computer Science and echnology, Nanjing University of Aeronautics and Astronautics, Nanjing 2006,
Iterative Laplacian Score for Feature Selection Linling Zhu, Linsong Miao, and Daoqiang Zhang College of Computer Science and echnology, Nanjing University of Aeronautics and Astronautics, Nanjing 2006,
Spectral Methods for Subgraph Detection
 Spectral Methods for Subgraph Detection Nadya T. Bliss & Benjamin A. Miller Embedded and High Performance Computing Patrick J. Wolfe Statistics and Information Laboratory Harvard University 12 July 2010
Spectral Methods for Subgraph Detection Nadya T. Bliss & Benjamin A. Miller Embedded and High Performance Computing Patrick J. Wolfe Statistics and Information Laboratory Harvard University 12 July 2010
Singular Value Decomposition and Principal Component Analysis (PCA) I
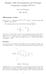 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Machine Learning for Data Science (CS4786) Lecture 11
 Machine Learning for Data Science (CS4786) Lecture 11 Spectral clustering Course Webpage : http://www.cs.cornell.edu/courses/cs4786/2016sp/ ANNOUNCEMENT 1 Assignment P1 the Diagnostic assignment 1 will
Machine Learning for Data Science (CS4786) Lecture 11 Spectral clustering Course Webpage : http://www.cs.cornell.edu/courses/cs4786/2016sp/ ANNOUNCEMENT 1 Assignment P1 the Diagnostic assignment 1 will
Spectral Generative Models for Graphs
 Spectral Generative Models for Graphs David White and Richard C. Wilson Department of Computer Science University of York Heslington, York, UK wilson@cs.york.ac.uk Abstract Generative models are well known
Spectral Generative Models for Graphs David White and Richard C. Wilson Department of Computer Science University of York Heslington, York, UK wilson@cs.york.ac.uk Abstract Generative models are well known
Inferring Models of cis-regulatory Modules using Information Theory
 Inferring Models of cis-regulatory Modules using Information Theory BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 28 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material,
Inferring Models of cis-regulatory Modules using Information Theory BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 28 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material,
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Diffusion and random walks on graphs
 Diffusion and random walks on graphs Leonid E. Zhukov School of Data Analysis and Artificial Intelligence Department of Computer Science National Research University Higher School of Economics Structural
Diffusion and random walks on graphs Leonid E. Zhukov School of Data Analysis and Artificial Intelligence Department of Computer Science National Research University Higher School of Economics Structural
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Learning from Sensor Data: Set II. Behnaam Aazhang J.S. Abercombie Professor Electrical and Computer Engineering Rice University
 Learning from Sensor Data: Set II Behnaam Aazhang J.S. Abercombie Professor Electrical and Computer Engineering Rice University 1 6. Data Representation The approach for learning from data Probabilistic
Learning from Sensor Data: Set II Behnaam Aazhang J.S. Abercombie Professor Electrical and Computer Engineering Rice University 1 6. Data Representation The approach for learning from data Probabilistic
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Genetic Networks. Korbinian Strimmer. Seminar: Statistical Analysis of RNA-Seq Data 19 June IMISE, Universität Leipzig
 Genetic Networks Korbinian Strimmer IMISE, Universität Leipzig Seminar: Statistical Analysis of RNA-Seq Data 19 June 2012 Korbinian Strimmer, RNA-Seq Networks, 19/6/2012 1 Paper G. I. Allen and Z. Liu.
Genetic Networks Korbinian Strimmer IMISE, Universität Leipzig Seminar: Statistical Analysis of RNA-Seq Data 19 June 2012 Korbinian Strimmer, RNA-Seq Networks, 19/6/2012 1 Paper G. I. Allen and Z. Liu.
Research Statement on Statistics Jun Zhang
 Research Statement on Statistics Jun Zhang (junzhang@galton.uchicago.edu) My interest on statistics generally includes machine learning and statistical genetics. My recent work focus on detection and interpretation
Research Statement on Statistics Jun Zhang (junzhang@galton.uchicago.edu) My interest on statistics generally includes machine learning and statistical genetics. My recent work focus on detection and interpretation
Weighted gene co-expression analysis. Yuehua Cui June 7, 2013
 Weighted gene co-expression analysis Yuehua Cui June 7, 2013 Weighted gene co-expression network (WGCNA) A type of scale-free network: A scale-free network is a network whose degree distribution follows
Weighted gene co-expression analysis Yuehua Cui June 7, 2013 Weighted gene co-expression network (WGCNA) A type of scale-free network: A scale-free network is a network whose degree distribution follows
Machine Learning (BSMC-GA 4439) Wenke Liu
 Machine Learning (BSMC-GA 4439) Wenke Liu 02-01-2018 Biomedical data are usually high-dimensional Number of samples (n) is relatively small whereas number of features (p) can be large Sometimes p>>n Problems
Machine Learning (BSMC-GA 4439) Wenke Liu 02-01-2018 Biomedical data are usually high-dimensional Number of samples (n) is relatively small whereas number of features (p) can be large Sometimes p>>n Problems
Class 4: Classification. Quaid Morris February 11 th, 2011 ML4Bio
 Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Spectral Graph Theory and You: Matrix Tree Theorem and Centrality Metrics
 Spectral Graph Theory and You: and Centrality Metrics Jonathan Gootenberg March 11, 2013 1 / 19 Outline of Topics 1 Motivation Basics of Spectral Graph Theory Understanding the characteristic polynomial
Spectral Graph Theory and You: and Centrality Metrics Jonathan Gootenberg March 11, 2013 1 / 19 Outline of Topics 1 Motivation Basics of Spectral Graph Theory Understanding the characteristic polynomial
CS224W: Social and Information Network Analysis Jure Leskovec, Stanford University
 CS224W: Social and Information Network Analysis Jure Leskovec Stanford University Jure Leskovec, Stanford University http://cs224w.stanford.edu Task: Find coalitions in signed networks Incentives: European
CS224W: Social and Information Network Analysis Jure Leskovec Stanford University Jure Leskovec, Stanford University http://cs224w.stanford.edu Task: Find coalitions in signed networks Incentives: European
Differential Modeling for Cancer Microarray Data
 Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
Differential Modeling for Cancer Microarray Data Omar Odibat Department of Computer Science Feb, 01, 2011 1 Outline Introduction Cancer Microarray data Problem Definition Differential analysis Existing
Spectral Clustering. Spectral Clustering? Two Moons Data. Spectral Clustering Algorithm: Bipartioning. Spectral methods
 Spectral Clustering Seungjin Choi Department of Computer Science POSTECH, Korea seungjin@postech.ac.kr 1 Spectral methods Spectral Clustering? Methods using eigenvectors of some matrices Involve eigen-decomposition
Spectral Clustering Seungjin Choi Department of Computer Science POSTECH, Korea seungjin@postech.ac.kr 1 Spectral methods Spectral Clustering? Methods using eigenvectors of some matrices Involve eigen-decomposition
Research Article On the Impact of Entropy Estimation on Transcriptional Regulatory Network Inference Based on Mutual Information
 Hindawi Publishing Corporation EURASIP Journal on Bioinformatics and Systems Biology Volume 29, Article ID 38959, 9 pages doi:1.1155/29/38959 Research Article On the Impact of Entropy Estimation on Transcriptional
Hindawi Publishing Corporation EURASIP Journal on Bioinformatics and Systems Biology Volume 29, Article ID 38959, 9 pages doi:1.1155/29/38959 Research Article On the Impact of Entropy Estimation on Transcriptional
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
ARACNE: An Algorithm for the Reconstruction of Gene Regulatory Networks in a Mammalian Cellular Context
 ARACNE: An Algorithm for the Reconstruction of Gene Regulatory Networks in a Mammalian Cellular Context Adam A. Margolin 1,2, Ilya Nemenman 2, Katia Basso 3, Chris Wiggins 2,4, Gustavo Stolovitzky 5, Riccardo
ARACNE: An Algorithm for the Reconstruction of Gene Regulatory Networks in a Mammalian Cellular Context Adam A. Margolin 1,2, Ilya Nemenman 2, Katia Basso 3, Chris Wiggins 2,4, Gustavo Stolovitzky 5, Riccardo
Data Analysis and Manifold Learning Lecture 3: Graphs, Graph Matrices, and Graph Embeddings
 Data Analysis and Manifold Learning Lecture 3: Graphs, Graph Matrices, and Graph Embeddings Radu Horaud INRIA Grenoble Rhone-Alpes, France Radu.Horaud@inrialpes.fr http://perception.inrialpes.fr/ Outline
Data Analysis and Manifold Learning Lecture 3: Graphs, Graph Matrices, and Graph Embeddings Radu Horaud INRIA Grenoble Rhone-Alpes, France Radu.Horaud@inrialpes.fr http://perception.inrialpes.fr/ Outline
1 Matrix notation and preliminaries from spectral graph theory
 Graph clustering (or community detection or graph partitioning) is one of the most studied problems in network analysis. One reason for this is that there are a variety of ways to define a cluster or community.
Graph clustering (or community detection or graph partitioning) is one of the most studied problems in network analysis. One reason for this is that there are a variety of ways to define a cluster or community.
Functional Analysis Review
 Outline 9.520: Statistical Learning Theory and Applications February 8, 2010 Outline 1 2 3 4 Vector Space Outline A vector space is a set V with binary operations +: V V V and : R V V such that for all
Outline 9.520: Statistical Learning Theory and Applications February 8, 2010 Outline 1 2 3 4 Vector Space Outline A vector space is a set V with binary operations +: V V V and : R V V such that for all
Model Accuracy Measures
 Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Singular value decomposition for genome-wide expression data processing and modeling. Presented by Jing Qiu
 Singular value decomposition for genome-wide expression data processing and modeling Presented by Jing Qiu April 23, 2002 Outline Biological Background Mathematical Framework:Singular Value Decomposition
Singular value decomposition for genome-wide expression data processing and modeling Presented by Jing Qiu April 23, 2002 Outline Biological Background Mathematical Framework:Singular Value Decomposition
11 : Gaussian Graphic Models and Ising Models
 10-708: Probabilistic Graphical Models 10-708, Spring 2017 11 : Gaussian Graphic Models and Ising Models Lecturer: Bryon Aragam Scribes: Chao-Ming Yen 1 Introduction Different from previous maximum likelihood
10-708: Probabilistic Graphical Models 10-708, Spring 2017 11 : Gaussian Graphic Models and Ising Models Lecturer: Bryon Aragam Scribes: Chao-Ming Yen 1 Introduction Different from previous maximum likelihood
8.1 Concentration inequality for Gaussian random matrix (cont d)
 MGMT 69: Topics in High-dimensional Data Analysis Falll 26 Lecture 8: Spectral clustering and Laplacian matrices Lecturer: Jiaming Xu Scribe: Hyun-Ju Oh and Taotao He, October 4, 26 Outline Concentration
MGMT 69: Topics in High-dimensional Data Analysis Falll 26 Lecture 8: Spectral clustering and Laplacian matrices Lecturer: Jiaming Xu Scribe: Hyun-Ju Oh and Taotao He, October 4, 26 Outline Concentration
Ensemble Non-negative Matrix Factorization Methods for Clustering Protein-Protein Interactions
 Belfield Campus Map Ensemble Non-negative Matrix Factorization Methods for Clustering Protein-Protein Interactions
Belfield Campus Map Ensemble Non-negative Matrix Factorization Methods for Clustering Protein-Protein Interactions
Clustering of Pathogenic Genes in Human Co-regulatory Network. Michael Colavita Mentor: Soheil Feizi Fifth Annual MIT PRIMES Conference May 17, 2015
 Clustering of Pathogenic Genes in Human Co-regulatory Network Michael Colavita Mentor: Soheil Feizi Fifth Annual MIT PRIMES Conference May 17, 2015 Topics Background Genetic Background Regulatory Networks
Clustering of Pathogenic Genes in Human Co-regulatory Network Michael Colavita Mentor: Soheil Feizi Fifth Annual MIT PRIMES Conference May 17, 2015 Topics Background Genetic Background Regulatory Networks
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data
 DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
Introduction to Bioinformatics
 Systems biology Introduction to Bioinformatics Systems biology: modeling biological p Study of whole biological systems p Wholeness : Organization of dynamic interactions Different behaviour of the individual
Systems biology Introduction to Bioinformatics Systems biology: modeling biological p Study of whole biological systems p Wholeness : Organization of dynamic interactions Different behaviour of the individual
Inverse Power Method for Non-linear Eigenproblems
 Inverse Power Method for Non-linear Eigenproblems Matthias Hein and Thomas Bühler Anubhav Dwivedi Department of Aerospace Engineering & Mechanics 7th March, 2017 1 / 30 OUTLINE Motivation Non-Linear Eigenproblems
Inverse Power Method for Non-linear Eigenproblems Matthias Hein and Thomas Bühler Anubhav Dwivedi Department of Aerospace Engineering & Mechanics 7th March, 2017 1 / 30 OUTLINE Motivation Non-Linear Eigenproblems
Graph Partitioning Using Random Walks
 Graph Partitioning Using Random Walks A Convex Optimization Perspective Lorenzo Orecchia Computer Science Why Spectral Algorithms for Graph Problems in practice? Simple to implement Can exploit very efficient
Graph Partitioning Using Random Walks A Convex Optimization Perspective Lorenzo Orecchia Computer Science Why Spectral Algorithms for Graph Problems in practice? Simple to implement Can exploit very efficient
Spectral Theory of Unsigned and Signed Graphs Applications to Graph Clustering. Some Slides
 Spectral Theory of Unsigned and Signed Graphs Applications to Graph Clustering Some Slides Jean Gallier Department of Computer and Information Science University of Pennsylvania Philadelphia, PA 19104,
Spectral Theory of Unsigned and Signed Graphs Applications to Graph Clustering Some Slides Jean Gallier Department of Computer and Information Science University of Pennsylvania Philadelphia, PA 19104,
PCA and admixture models
 PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
PCA and admixture models CM226: Machine Learning for Bioinformatics. Fall 2016 Sriram Sankararaman Acknowledgments: Fei Sha, Ameet Talwalkar, Alkes Price PCA and admixture models 1 / 57 Announcements HW1
Lecture 1: Graphs, Adjacency Matrices, Graph Laplacian
 Lecture 1: Graphs, Adjacency Matrices, Graph Laplacian Radu Balan January 31, 2017 G = (V, E) An undirected graph G is given by two pieces of information: a set of vertices V and a set of edges E, G =
Lecture 1: Graphs, Adjacency Matrices, Graph Laplacian Radu Balan January 31, 2017 G = (V, E) An undirected graph G is given by two pieces of information: a set of vertices V and a set of edges E, G =
Unsupervised Clustering of Human Pose Using Spectral Embedding
 Unsupervised Clustering of Human Pose Using Spectral Embedding Muhammad Haseeb and Edwin R Hancock Department of Computer Science, The University of York, UK Abstract In this paper we use the spectra of
Unsupervised Clustering of Human Pose Using Spectral Embedding Muhammad Haseeb and Edwin R Hancock Department of Computer Science, The University of York, UK Abstract In this paper we use the spectra of
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Predicting causal effects in large-scale systems from observational data
 nature methods Predicting causal effects in large-scale systems from observational data Marloes H Maathuis 1, Diego Colombo 1, Markus Kalisch 1 & Peter Bühlmann 1,2 Supplementary figures and text: Supplementary
nature methods Predicting causal effects in large-scale systems from observational data Marloes H Maathuis 1, Diego Colombo 1, Markus Kalisch 1 & Peter Bühlmann 1,2 Supplementary figures and text: Supplementary
Machine Learning. Principal Components Analysis. Le Song. CSE6740/CS7641/ISYE6740, Fall 2012
 Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Introduction to Spectral Graph Theory and Graph Clustering
 Introduction to Spectral Graph Theory and Graph Clustering Chengming Jiang ECS 231 Spring 2016 University of California, Davis 1 / 40 Motivation Image partitioning in computer vision 2 / 40 Motivation
Introduction to Spectral Graph Theory and Graph Clustering Chengming Jiang ECS 231 Spring 2016 University of California, Davis 1 / 40 Motivation Image partitioning in computer vision 2 / 40 Motivation
Spectral Clustering on Handwritten Digits Database
 University of Maryland-College Park Advance Scientific Computing I,II Spectral Clustering on Handwritten Digits Database Author: Danielle Middlebrooks Dmiddle1@math.umd.edu Second year AMSC Student Advisor:
University of Maryland-College Park Advance Scientific Computing I,II Spectral Clustering on Handwritten Digits Database Author: Danielle Middlebrooks Dmiddle1@math.umd.edu Second year AMSC Student Advisor:
Theorems. Least squares regression
 Theorems In this assignment we are trying to classify AML and ALL samples by use of penalized logistic regression. Before we indulge on the adventure of classification we should first explain the most
Theorems In this assignment we are trying to classify AML and ALL samples by use of penalized logistic regression. Before we indulge on the adventure of classification we should first explain the most
17.1 Directed Graphs, Undirected Graphs, Incidence Matrices, Adjacency Matrices, Weighted Graphs
 Chapter 17 Graphs and Graph Laplacians 17.1 Directed Graphs, Undirected Graphs, Incidence Matrices, Adjacency Matrices, Weighted Graphs Definition 17.1. A directed graph isa pairg =(V,E), where V = {v
Chapter 17 Graphs and Graph Laplacians 17.1 Directed Graphs, Undirected Graphs, Incidence Matrices, Adjacency Matrices, Weighted Graphs Definition 17.1. A directed graph isa pairg =(V,E), where V = {v
Prediction of double gene knockout measurements
 Prediction of double gene knockout measurements Sofia Kyriazopoulou-Panagiotopoulou sofiakp@stanford.edu December 12, 2008 Abstract One way to get an insight into the potential interaction between a pair
Prediction of double gene knockout measurements Sofia Kyriazopoulou-Panagiotopoulou sofiakp@stanford.edu December 12, 2008 Abstract One way to get an insight into the potential interaction between a pair
Basic Calculus Review
 Basic Calculus Review Lorenzo Rosasco ISML Mod. 2 - Machine Learning Vector Spaces Functionals and Operators (Matrices) Vector Space A vector space is a set V with binary operations +: V V V and : R V
Basic Calculus Review Lorenzo Rosasco ISML Mod. 2 - Machine Learning Vector Spaces Functionals and Operators (Matrices) Vector Space A vector space is a set V with binary operations +: V V V and : R V
Modeling Gene Expression from Microarray Expression Data with State-Space Equations. F.X. Wu, W.J. Zhang, and A.J. Kusalik
 Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Modeling Gene Expression from Microarray Expression Data with State-Space Equations FX Wu, WJ Zhang, and AJ Kusalik Pacific Symposium on Biocomputing 9:581-592(2004) MODELING GENE EXPRESSION FROM MICROARRAY
Regression Model In The Analysis Of Micro Array Data-Gene Expression Detection
 Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Jamal Fathima.J.I 1 and P.Venkatesan 1. Research Scholar -Department of statistics National Institute For Research In Tuberculosis, Indian Council For Medical Research,Chennai,India,.Department of statistics
Networks and Their Spectra
 Networks and Their Spectra Victor Amelkin University of California, Santa Barbara Department of Computer Science victor@cs.ucsb.edu December 4, 2017 1 / 18 Introduction Networks (= graphs) are everywhere.
Networks and Their Spectra Victor Amelkin University of California, Santa Barbara Department of Computer Science victor@cs.ucsb.edu December 4, 2017 1 / 18 Introduction Networks (= graphs) are everywhere.
Stat 315c: Introduction
 Stat 315c: Introduction Art B. Owen Stanford Statistics Art B. Owen (Stanford Statistics) Stat 315c: Introduction 1 / 14 Stat 315c Analysis of Transposable Data Usual Statistics Setup there s Y (we ll
Stat 315c: Introduction Art B. Owen Stanford Statistics Art B. Owen (Stanford Statistics) Stat 315c: Introduction 1 / 14 Stat 315c Analysis of Transposable Data Usual Statistics Setup there s Y (we ll
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Analyzing Microarray Time course Genome wide Data
 OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
OR 779 Functional Data Analysis Course Project Analyzing Microarray Time course Genome wide Data Presented by Xin Zhao April 29, 2002 Cornell University Overview 1. Introduction Biological Background Biological
This appendix provides a very basic introduction to linear algebra concepts.
 APPENDIX Basic Linear Algebra Concepts This appendix provides a very basic introduction to linear algebra concepts. Some of these concepts are intentionally presented here in a somewhat simplified (not
APPENDIX Basic Linear Algebra Concepts This appendix provides a very basic introduction to linear algebra concepts. Some of these concepts are intentionally presented here in a somewhat simplified (not
Learning undirected graphical models from multiple datasets with the generalized non-rejection rate
 Learning undirected graphical models from multiple datasets with the generalized non-rejection rate Alberto Roverato Università di Bologna, Italy alberto.roverato@unibo.it Robert Castelo Universitat Pompeu
Learning undirected graphical models from multiple datasets with the generalized non-rejection rate Alberto Roverato Università di Bologna, Italy alberto.roverato@unibo.it Robert Castelo Universitat Pompeu
Reducing storage requirements for biological sequence comparison
 Bioinformatics Advance Access published July 15, 2004 Bioinfor matics Oxford University Press 2004; all rights reserved. Reducing storage requirements for biological sequence comparison Michael Roberts,
Bioinformatics Advance Access published July 15, 2004 Bioinfor matics Oxford University Press 2004; all rights reserved. Reducing storage requirements for biological sequence comparison Michael Roberts,
CS60021: Scalable Data Mining. Dimensionality Reduction
 J. Leskovec, A. Rajaraman, J. Ullman: Mining of Massive Datasets, http://www.mmds.org 1 CS60021: Scalable Data Mining Dimensionality Reduction Sourangshu Bhattacharya Assumption: Data lies on or near a
J. Leskovec, A. Rajaraman, J. Ullman: Mining of Massive Datasets, http://www.mmds.org 1 CS60021: Scalable Data Mining Dimensionality Reduction Sourangshu Bhattacharya Assumption: Data lies on or near a
The Singular Value Decomposition (SVD) and Principal Component Analysis (PCA)
 Chapter 5 The Singular Value Decomposition (SVD) and Principal Component Analysis (PCA) 5.1 Basics of SVD 5.1.1 Review of Key Concepts We review some key definitions and results about matrices that will
Chapter 5 The Singular Value Decomposition (SVD) and Principal Component Analysis (PCA) 5.1 Basics of SVD 5.1.1 Review of Key Concepts We review some key definitions and results about matrices that will
1 Matrix notation and preliminaries from spectral graph theory
 Graph clustering (or community detection or graph partitioning) is one of the most studied problems in network analysis. One reason for this is that there are a variety of ways to define a cluster or community.
Graph clustering (or community detection or graph partitioning) is one of the most studied problems in network analysis. One reason for this is that there are a variety of ways to define a cluster or community.
Bioinformatics 2. Yeast two hybrid. Proteomics. Proteomics
 GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
GENOME Bioinformatics 2 Proteomics protein-gene PROTEOME protein-protein METABOLISM Slide from http://www.nd.edu/~networks/ Citrate Cycle Bio-chemical reactions What is it? Proteomics Reveal protein Protein
ECS 289 / MAE 298, Network Theory Spring 2014 Problem Set # 1, Solutions. Problem 1: Power Law Degree Distributions (Continuum approach)
 ECS 289 / MAE 298, Network Theory Spring 204 Problem Set #, Solutions Problem : Power Law Degree Distributions (Continuum approach) Consider the power law distribution p(k) = Ak γ, with support (i.e.,
ECS 289 / MAE 298, Network Theory Spring 204 Problem Set #, Solutions Problem : Power Law Degree Distributions (Continuum approach) Consider the power law distribution p(k) = Ak γ, with support (i.e.,
Information Propagation Analysis of Social Network Using the Universality of Random Matrix
 Information Propagation Analysis of Social Network Using the Universality of Random Matrix Yusuke Sakumoto, Tsukasa Kameyama, Chisa Takano and Masaki Aida Tokyo Metropolitan University, 6-6 Asahigaoka,
Information Propagation Analysis of Social Network Using the Universality of Random Matrix Yusuke Sakumoto, Tsukasa Kameyama, Chisa Takano and Masaki Aida Tokyo Metropolitan University, 6-6 Asahigaoka,
Data Mining and Analysis: Fundamental Concepts and Algorithms
 Data Mining and Analysis: Fundamental Concepts and Algorithms dataminingbook.info Mohammed J. Zaki 1 Wagner Meira Jr. 2 1 Department of Computer Science Rensselaer Polytechnic Institute, Troy, NY, USA
Data Mining and Analysis: Fundamental Concepts and Algorithms dataminingbook.info Mohammed J. Zaki 1 Wagner Meira Jr. 2 1 Department of Computer Science Rensselaer Polytechnic Institute, Troy, NY, USA
Spectra of Adjacency and Laplacian Matrices
 Spectra of Adjacency and Laplacian Matrices Definition: University of Alicante (Spain) Matrix Computing (subject 3168 Degree in Maths) 30 hours (theory)) + 15 hours (practical assignment) Contents 1. Spectra
Spectra of Adjacency and Laplacian Matrices Definition: University of Alicante (Spain) Matrix Computing (subject 3168 Degree in Maths) 30 hours (theory)) + 15 hours (practical assignment) Contents 1. Spectra
Bayesian Data Fusion with Gaussian Process Priors : An Application to Protein Fold Recognition
 Bayesian Data Fusion with Gaussian Process Priors : An Application to Protein Fold Recognition Mar Girolami 1 Department of Computing Science University of Glasgow girolami@dcs.gla.ac.u 1 Introduction
Bayesian Data Fusion with Gaussian Process Priors : An Application to Protein Fold Recognition Mar Girolami 1 Department of Computing Science University of Glasgow girolami@dcs.gla.ac.u 1 Introduction
Causal Graphical Models in Systems Genetics
 1 Causal Graphical Models in Systems Genetics 2013 Network Analysis Short Course - UCLA Human Genetics Elias Chaibub Neto and Brian S Yandell July 17, 2013 Motivation and basic concepts 2 3 Motivation
1 Causal Graphical Models in Systems Genetics 2013 Network Analysis Short Course - UCLA Human Genetics Elias Chaibub Neto and Brian S Yandell July 17, 2013 Motivation and basic concepts 2 3 Motivation
Supplemental Material
 Supplemental Material Article title: Construction and comparison of gene co expression networks shows complex plant immune responses Author names and affiliation: Luis Guillermo Leal 1*, Email: lgleala@unal.edu.co
Supplemental Material Article title: Construction and comparison of gene co expression networks shows complex plant immune responses Author names and affiliation: Luis Guillermo Leal 1*, Email: lgleala@unal.edu.co
Classification Semi-supervised learning based on network. Speakers: Hanwen Wang, Xinxin Huang, and Zeyu Li CS Winter
 Classification Semi-supervised learning based on network Speakers: Hanwen Wang, Xinxin Huang, and Zeyu Li CS 249-2 2017 Winter Semi-Supervised Learning Using Gaussian Fields and Harmonic Functions Xiaojin
Classification Semi-supervised learning based on network Speakers: Hanwen Wang, Xinxin Huang, and Zeyu Li CS 249-2 2017 Winter Semi-Supervised Learning Using Gaussian Fields and Harmonic Functions Xiaojin
Dimensionality Reduction Techniques (DRT)
 Dimensionality Reduction Techniques (DRT) Introduction: Sometimes we have lot of variables in the data for analysis which create multidimensional matrix. To simplify calculation and to get appropriate,
Dimensionality Reduction Techniques (DRT) Introduction: Sometimes we have lot of variables in the data for analysis which create multidimensional matrix. To simplify calculation and to get appropriate,
Data Mining: Concepts and Techniques. (3 rd ed.) Chapter 8. Chapter 8. Classification: Basic Concepts
 Data Mining: Concepts and Techniques (3 rd ed.) Chapter 8 1 Chapter 8. Classification: Basic Concepts Classification: Basic Concepts Decision Tree Induction Bayes Classification Methods Rule-Based Classification
Data Mining: Concepts and Techniques (3 rd ed.) Chapter 8 1 Chapter 8. Classification: Basic Concepts Classification: Basic Concepts Decision Tree Induction Bayes Classification Methods Rule-Based Classification
A NEW METHOD OF DETERMINING THRESHOLD OF GENE NETWORK BASED ON PHENOTYPES
 Journal of Biological Systems, Vol. 19, No. 4 (2011) 607 616 c World Scientific Publishing Company DOI: 10.1142/S0218339011004068 A NEW METHOD OF DETERMINING THRESHOLD OF GENE NETWORK BASED ON PHENOTYPES
Journal of Biological Systems, Vol. 19, No. 4 (2011) 607 616 c World Scientific Publishing Company DOI: 10.1142/S0218339011004068 A NEW METHOD OF DETERMINING THRESHOLD OF GENE NETWORK BASED ON PHENOTYPES
Certifying the Global Optimality of Graph Cuts via Semidefinite Programming: A Theoretic Guarantee for Spectral Clustering
 Certifying the Global Optimality of Graph Cuts via Semidefinite Programming: A Theoretic Guarantee for Spectral Clustering Shuyang Ling Courant Institute of Mathematical Sciences, NYU Aug 13, 2018 Joint
Certifying the Global Optimality of Graph Cuts via Semidefinite Programming: A Theoretic Guarantee for Spectral Clustering Shuyang Ling Courant Institute of Mathematical Sciences, NYU Aug 13, 2018 Joint
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 3 Linear
Dimensionality Reduction
 Dimensionality Reduction Le Song Machine Learning I CSE 674, Fall 23 Unsupervised learning Learning from raw (unlabeled, unannotated, etc) data, as opposed to supervised data where a classification of
Dimensionality Reduction Le Song Machine Learning I CSE 674, Fall 23 Unsupervised learning Learning from raw (unlabeled, unannotated, etc) data, as opposed to supervised data where a classification of
Machine learning for pervasive systems Classification in high-dimensional spaces
 Machine learning for pervasive systems Classification in high-dimensional spaces Department of Communications and Networking Aalto University, School of Electrical Engineering stephan.sigg@aalto.fi Version
Machine learning for pervasive systems Classification in high-dimensional spaces Department of Communications and Networking Aalto University, School of Electrical Engineering stephan.sigg@aalto.fi Version
Non-Negative Factorization for Clustering of Microarray Data
 INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
INT J COMPUT COMMUN, ISSN 1841-9836 9(1):16-23, February, 2014. Non-Negative Factorization for Clustering of Microarray Data L. Morgos Lucian Morgos Dept. of Electronics and Telecommunications Faculty
Power Grid Partitioning: Static and Dynamic Approaches
 Power Grid Partitioning: Static and Dynamic Approaches Miao Zhang, Zhixin Miao, Lingling Fan Department of Electrical Engineering University of South Florida Tampa FL 3320 miaozhang@mail.usf.edu zmiao,
Power Grid Partitioning: Static and Dynamic Approaches Miao Zhang, Zhixin Miao, Lingling Fan Department of Electrical Engineering University of South Florida Tampa FL 3320 miaozhang@mail.usf.edu zmiao,
Expectation Maximization
 Expectation Maximization Machine Learning CSE546 Carlos Guestrin University of Washington November 13, 2014 1 E.M.: The General Case E.M. widely used beyond mixtures of Gaussians The recipe is the same
Expectation Maximization Machine Learning CSE546 Carlos Guestrin University of Washington November 13, 2014 1 E.M.: The General Case E.M. widely used beyond mixtures of Gaussians The recipe is the same
B553 Lecture 5: Matrix Algebra Review
 B553 Lecture 5: Matrix Algebra Review Kris Hauser January 19, 2012 We have seen in prior lectures how vectors represent points in R n and gradients of functions. Matrices represent linear transformations
B553 Lecture 5: Matrix Algebra Review Kris Hauser January 19, 2012 We have seen in prior lectures how vectors represent points in R n and gradients of functions. Matrices represent linear transformations
CSE 291. Assignment Spectral clustering versus k-means. Out: Wed May 23 Due: Wed Jun 13
 CSE 291. Assignment 3 Out: Wed May 23 Due: Wed Jun 13 3.1 Spectral clustering versus k-means Download the rings data set for this problem from the course web site. The data is stored in MATLAB format as
CSE 291. Assignment 3 Out: Wed May 23 Due: Wed Jun 13 3.1 Spectral clustering versus k-means Download the rings data set for this problem from the course web site. The data is stored in MATLAB format as
An indicator for the number of clusters using a linear map to simplex structure
 An indicator for the number of clusters using a linear map to simplex structure Marcus Weber, Wasinee Rungsarityotin, and Alexander Schliep Zuse Institute Berlin ZIB Takustraße 7, D-495 Berlin, Germany
An indicator for the number of clusters using a linear map to simplex structure Marcus Weber, Wasinee Rungsarityotin, and Alexander Schliep Zuse Institute Berlin ZIB Takustraße 7, D-495 Berlin, Germany
Interaction Network Topologies
 Proceedings of International Joint Conference on Neural Networks, Montreal, Canada, July 31 - August 4, 2005 Inferrng Protein-Protein Interactions Using Interaction Network Topologies Alberto Paccanarot*,
Proceedings of International Joint Conference on Neural Networks, Montreal, Canada, July 31 - August 4, 2005 Inferrng Protein-Protein Interactions Using Interaction Network Topologies Alberto Paccanarot*,
Towards Detecting Protein Complexes from Protein Interaction Data
 Towards Detecting Protein Complexes from Protein Interaction Data Pengjun Pei 1 and Aidong Zhang 1 Department of Computer Science and Engineering State University of New York at Buffalo Buffalo NY 14260,
Towards Detecting Protein Complexes from Protein Interaction Data Pengjun Pei 1 and Aidong Zhang 1 Department of Computer Science and Engineering State University of New York at Buffalo Buffalo NY 14260,
Dimensionality reduction
 Dimensionality Reduction PCA continued Machine Learning CSE446 Carlos Guestrin University of Washington May 22, 2013 Carlos Guestrin 2005-2013 1 Dimensionality reduction n Input data may have thousands
Dimensionality Reduction PCA continued Machine Learning CSE446 Carlos Guestrin University of Washington May 22, 2013 Carlos Guestrin 2005-2013 1 Dimensionality reduction n Input data may have thousands
Computational Systems Biology
 Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm
 Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm Zhenqiu Liu, Dechang Chen 2 Department of Computer Science Wayne State University, Market Street, Frederick, MD 273,
Gene Expression Data Classification with Revised Kernel Partial Least Squares Algorithm Zhenqiu Liu, Dechang Chen 2 Department of Computer Science Wayne State University, Market Street, Frederick, MD 273,
