Supplementary Information Mechanism of influenza A M2 transmembrane domain assembly in lipid membranes
|
|
|
- Julianna Pierce
- 5 years ago
- Views:
Transcription
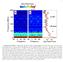
1 Supplementary Information Mechanism of influenza A M2 transmembrane domain assembly in lipid membranes Elka R. Georgieva 1,2*, Peter P. Borbat 1,2, Haley D. Norman 3 and Jack H. Freed 1,2* 1 Department of Chemistry and Chemical Biology, 2 National Biomedical Center for Advanced ESR Technology, and 3 School of Engineering, Cornell University, Ithaca, NY Correspondence should be addressed to erg54@cornell.edu and jhf3@cornell.edu Supplementary Figure 1. DEER data preprocessing and presentation as used in this work. Supplementary Fig. 1 illustrates DEER data preprocessing and presentation used in this work, given that other procedures are common in utilizing this method. The raw time-domain DEER signal (in black) and the background signal commonly called baseline (in green) are shown on 1
2 a semi-logarithmic scale in the upper panel. All DEER data in this study appear as nonoscillating decays, each reaches the asymptotic value, V at t=, V( ). It is needed to determine the modulation depth Δ(p, N) 1 DEER(t= )/DEER(t=0) defined in the main text and designated as Δ(p, N) 1 DEER( )/DEER(0) in the figure. After fitting the baseline to a straight line or a 2 nd degree polynomial, the background-subtracted DEER signal is plotted on a linear scale in the middle panel. Now it becomes V(0) = 1 + Δ and V( ) = 1. After subtracting unity and normalization by dividing by V(0) it finally appears in the lowest panel such that V(0) is Δ(p, N) and V( ) = 0. Thus, the modulation depth, Δ, can be read directly from the plot and is used in this work to estimate the number of interacting spins in a nano-object based on known approaches 1-4. Supplementary Figure 2. Mass spectrum shows close to 100% (>95%) spin-labeling efficiency of M2TMD construct (M2TMD), based on the experimentally obtained molecular weight, which for M2TMD singly spin-labeled with MTSL is Da 2
3 Supplementary Figure 3. Room temperature (23 C) cw-esr spectra for spin-labeled L46C residue in M2TMD residing in DOPC/POPS membranes. a Low ph data, ph 5.5, for P/L of 1:235, 1:500 and 1:4,100 are shown. b Data taken at ph 5.5 and ph 8 are compared for P/L of 1:235. The cw-esr spectra are all similar indicating very similar dynamic properties with rotational correlation time, c, in the nanosecond range. No contribution from free spin-label to the recorded cw-esr spectra could be noticed. 3
4 Supplementary Figure 4. a A ph-induced effect on M2TMD assembly is demonstrated for M2TMD reconstituted in DOPC/POPS membranes at peptide-to-lipid ratios (P/L) of 1:235 and 1:1,300 and at ph s 5.5 (red) and 8 (blue). b The dependence on P/L of DEER time-domain signals (left) and corresponding distances (right) at both ph 5.5 and ph 8 is demonstrated by comparing the data for P/L s of 1:235 and 1:4,100. In all cases the DEER data and distance distributions are normalized to have the same intensity. In both a and b, the experimental time-domain DEER data are shown on the left and respective distance distributions are on the right. The distance distributions in these plots were reconstructed using Tikhonov regularization 5 and refined by the Maximum Entropy Method 6. c The distance distributions generated using Molecular Modeling Software (MMM) 7 based on high ph (ph 7.5) NMR structures in DOPC/DOPE (pdb ID 2L0J) 8 and in DMPC with amantadine drug bound in M2 channel (pdb ID 2KQT) 9 are shown in the upper panel. The 4
5 calculated distance distribution based on the x-ray crystal structure of detergent-solubilized M2TMD at ph 5.3 (pdb ID 3C9J) 10 is shown in the lower panel. Overall, we find good agreement between calculated and experimental distances. However, in most cases our distance distributions are produced by co-existing M2TMD dimers and tetramers with their contributions depending on P/L ratio. In general, dimers contribute just one distance, whereas for a tetramer, both laterally and proximally distant spins define the DEER signal and distances. In the case of a homotetramer with 4-fold symmetry two distances r 2 and r 1 = 2 1/2 r 2 with populations in ratio 2:1 could be reconstructed from the time-domain DEER signal, as previously discussed 11. (r 1 and r 2 here denote proximal and lateral distances, respectively). In the relevant case of a lower symmetry or for an asymmetric tetramer more distances (up to 6) may be present, resulting in a less tractable case. It is noteworthy that we observed an increased population of longer distances in the range of nm when lowering the P/L, i.e. increasing the population of M2TMD dimers. Another trend is a shift to longer distances upon decreasing ph from 8 to 5.5, that was observed for all P/L conditions, including those where dimer is prevalent (P/L s higher than 1:4,100), thus indicating that the dimer also experiences ph sensitive conformational changes. In summary, we did observe a conspicuous ph-induced shift to longer distances upon decreasing the ph from 8 to 5.5, which might be reasonably attributed to M2TMD channel opening. However, investigating the details of this structural transition is outside the focus of the present study. 5
6 Supplementary Figure 5. High degree of DEER data reproducibility for independent sample preparations is illustrated for selected typical cases, P/L 1:160 and 1:2,300 at ph 5.5, and 1:4,100 at ph 8. Data for duplicate samples from different preparations are superimposed in each panel. In all cases the extent of data variations is negligible. Supplementary Figure 6. The lack of effect of sample freezing rate on the DEER modulation depth for M2TMD spin-labeled at position L46C reconstituted into DOPC/POPS membranes at ph 8 and P/L 1:3,170. Data for samples plunge-frozen in liquid nitrogen or much faster by quenching in n-pentane kept just above its melting point of -130 ºC, are shown in shades of blue. The raw DEER data rendered on semi-logarithmic scale together with the corresponding second-order polynomial baselines and the background-subtracted DEER data are plotted in the left and right panels, respectively. For these two freezing methods, we found only very minor difference ~0.01 in the DEER modulation depths, which is almost within the level of uncertainty encountered in fitting 6
7 the baselines. Moreover, no change of the signal shape exceeding the noise level was detected. This indicates that it is unlikely that in our procedures there could be significant effects of lipid freezing on the M2TMD state and its assembly in particular. This is in line with observing just minor effects in freezing of a globular soluble protein 12. Supplementary Figure 7. DEER modulation depth variations that are too small to be significant are produced due to uncertainties in background fitting. The data shown in Supplementary Fig. 7 were taken for M2TMD in DOPC/POPS at 85:15 molar ratio. In the left panels, the raw DEER data (in black) are plotted on a semi-logarithmic scale together with manually varied baselines (BL1-BL3) rendered in shades of blue and cyan for ph 8 and in shades of red for ph 5.5. The background-subtracted and normalized DEER data presented in the right panels are rendered in colors corresponding to those of the baselines used in the subtractions. The data for sample with P/L 1:18,800 at ph 8 are shown in row a, whereas the data for sample with P/L 1:2,300 at ph 5.5 are shown in row b. The baselines for ph 8 were fit to a second order polynomial on semi-logarithmic scale. In the case of ph 5.5, linear baseline (plotted in violet) was also adequate. Using the approach 7
8 outlined, we estimated an error due to uncertainty in background subtraction not to exceed ±2.5% for all processed DEER data. Supplementary Figure 8. Low-temperature (163 K) cw-esr spectra for spin-labeled L46C residue in M2TMD in DOPC/POPS membranes. Data for ph 5.5 and ph 8 at P/Ls of 1:160 and 1:500 are shown. To examine the possible presence of short distances (shorter than is in the sensitivity range of DEER, i.e. less than 1.7 nm), we recorded the cw-esr spectra of M2TMD frozen samples. Finding the d 1 /d parameter from the cw-esr spectra of frozen samples, as illustrated by Supplementary Fig. 8, is a commonly used empirical method for estimating strong dipole-dipole interactions arising at shorter than 2.0 nm distances between paramagnetic centers 13. According to this method an inter-spin distance of 1.89 nm corresponds to d 1 /d = 0.54; further increase in the inter-spin distances leads to smaller values for the d 1 /d parameter. In our study, we measured a d 1 /d parameter of less than 0.54, indicating the presence of distances, which are well within the sensitivity range of DEER experiment. Furthermore, the largest d 1 /d values were obtained for P/L of 1:160 for both ph 5.5 and ph 8. These values gradually decrease with the increase of P/L (P/L 1:160 vs. P/L 1:500 in the figure), i.e. when reducing the population of M2TMD
9 tetramer. Hence, most likely, only the M2TMD tetramer has a distance in the range of 2.0 nm or slightly less, which also agrees with our data in Supplementary Fig. 4. Supplementary Figure 9. a Primary spin-echo echo decays for samples spin-labeled at position L46C in M2TMD are shown. Data plotted on the left were taken at ph 5.5 for P/L of 1:1,300 and for P/L of 1:160, but for peptide magnetically diluted with non-labeled peptide to about the same number of spins. Data on the right were taken at ph 8 for P/L of 1:500 and for P/L of 1:160, but with peptide magnetically diluted with non-labeled peptide to about the same number of spins. b Time-domain double-quantum coherence (DQC) signal (left) and reconstructed distances (right) for the magnetically diluted sample at ph 8. The time-domain DQC was normalized to unity at zero evolution time. 9
10 To collect the primary spin-echo decay data, a standard π/2-π pulse sequence with 250 ns initial inter-pulse separation was used. Within the accuracy of relaxation experiment, our data for non-magnetically diluted and magnetically diluted samples at ph 5.5 are nearly identical. Only a minor difference is noticed for ph 8. Thus, the primary spin-echo decays due to phase memory relaxation, at P/L ratio of 1:160, where tetramers are dominant species, are very close to those where dimers are also significantly present, i.e. P/L 1:1,300 and 1:500. The DQC data indicate that at most there is only a very small population of distances shorter than 1.7 nm (green dashed line). Given that there is usually a broadening ( wings ) of distance distribution due to noise-dependent stabilization properties of Tikhonov regularization, the fraction of spin-pairs at distances less than 2.0 nm is likely even less than what is seen in panel c. 10
11 Supplementary References 1. Milov, A.D., Ponomarev, A.B. & Tsvetkov, Y.D. Electron-electron double resonance in electron spin echo: Model biradical systems and the sensitized photolysis of decalin. Chemical Physics Letters 110, (1984). 2. Bode, B.E. et al. Counting the monomers in nanometer-sized oligomers by pulsed electronelectron double resonance. J Am Chem Soc 129, (2007). 3. Borbat, P.P. & Freed, J.H. Pulse Dipolar ESR: Distance Measurements. in Structure and Bonding (Springer, Heidelberg 2014). 4. Jeschke, G., Sajid, M., Schulte, M. & Godt, A. Three-spin correlations in double electronelectron resonance. Phys Chem Chem Phys 11, (2009). 5. Chiang, Y.W., Borbat, P.P. & Freed, J.H. The determination of pair distance distributions by pulsed ESR using Tikhonov regularization. J Magn Reson 172, (2005). 6. Chiang, Y.W., Borbat, P.P. & Freed, J.H. Maximum entropy: a complement to Tikhonov regularization for determination of pair distance distributions by pulsed ESR. J Magn Reson 177, (2005). 7. Polyhach, Y., Bordignon, E. & Jeschke, G. Rotamer libraries of spin labelled cysteines for protein studies. Phys Chem Chem Phys 13, (2011). 8. Sharma, M. et al. Insight into the mechanism of the influenza A proton channel from a structure in a lipid bilayer. Science 330, (2010). 9. Cady, S.D. et al. Structure of the amantadine binding site of influenza M2 proton channels in lipid bilayers. Nature 463, (2010). 10. Stouffer, A.L. et al. Structural basis for the function and inhibition of an influenza virus proton channel. Nature 451, (2008). 11. Dalmas, O., Hyde, H.C., Hulse, R.E. & Perozo, E. Symmetry-constrained analysis of pulsed double electron-electron resonance (DEER) spectroscopy reveals the dynamic nature of the KcsA activation gate. J Am Chem Soc 134, (2012). 12. Georgieva, E.R. et al. Effect of freezing conditions on distances and their distributions derived from Double Electron Electron Resonance (DEER): a study of doubly-spin-labeled T4 lysozyme. J Magn Reson 216, (2012). 13. Kokorin, A.I. Forty years of the d1/d parameter. (ed. Kokorin, A.I.) (InTech, 2012, p ). 11
SUPPLEMENTARY MATERIAL FOR
 SUPPLEMENTARY MATERIAL FOR THE LIPID-BINDING DOMAIN OF WILD TYPE AND MUTANT ALPHA- SYNUCLEIN: COMPACTNESS AND INTERCONVERSION BETWEEN THE BROKEN- AND EXTENDED-HELIX FORMS. Elka R. Georgieva 1, Trudy F.
SUPPLEMENTARY MATERIAL FOR THE LIPID-BINDING DOMAIN OF WILD TYPE AND MUTANT ALPHA- SYNUCLEIN: COMPACTNESS AND INTERCONVERSION BETWEEN THE BROKEN- AND EXTENDED-HELIX FORMS. Elka R. Georgieva 1, Trudy F.
SUPPLEMENTARY MATERIAL
 SUPPLEMENTARY MATERIAL rientation selection in distance measurements between nitroxide spin labels at 94 GHz EPR with variable frequency separation Igor Tkach 1, Soraya Pornsuwan 1, Claudia Höbartner 2,
SUPPLEMENTARY MATERIAL rientation selection in distance measurements between nitroxide spin labels at 94 GHz EPR with variable frequency separation Igor Tkach 1, Soraya Pornsuwan 1, Claudia Höbartner 2,
Maximum entropy: A complement to Tikhonov regularization for determination of pair distance distributions by pulsed ESR
 Journal of Magnetic Resonance 177 (2005) 184 196 www.elsevier.com/locate/jmr Maximum entropy: A complement to Tikhonov regularization for determination of pair distance distributions by pulsed ESR Yun-Wei
Journal of Magnetic Resonance 177 (2005) 184 196 www.elsevier.com/locate/jmr Maximum entropy: A complement to Tikhonov regularization for determination of pair distance distributions by pulsed ESR Yun-Wei
Pulsed Electron Electron Double Resonance
 Pulsed Electron Electron Double Resonance (PELDOR oder DEER) T. F. Prisner Institute of Phys. & Theor. Chemistry Center of Biological Magnetic Resonance Goethe University Frankfurt www.prisner.de Teaching
Pulsed Electron Electron Double Resonance (PELDOR oder DEER) T. F. Prisner Institute of Phys. & Theor. Chemistry Center of Biological Magnetic Resonance Goethe University Frankfurt www.prisner.de Teaching
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
Prediction of favourable sites for spin labelling of proteins
 Spectroscopy 24 (2010) 651 659 651 DOI 10.3233/SPE-2010-0490 IOS Press Prediction of favourable sites for spin labelling of proteins Yevhen Polyhach and Gunnar Jeschke Laboratory of Physical Chemistry,
Spectroscopy 24 (2010) 651 659 651 DOI 10.3233/SPE-2010-0490 IOS Press Prediction of favourable sites for spin labelling of proteins Yevhen Polyhach and Gunnar Jeschke Laboratory of Physical Chemistry,
Protein Structure Determination Using Long-Distance Constraints from Double-Quantum Coherence ESR: Study of T4 Lysozyme
 Published on Web 04/19/2002 Protein Structure Determination Using Long-Distance Constraints from Double-Quantum Coherence ESR: Study of T4 Lysozyme Petr P. Borbat, Hassane S. Mchaourab, and Jack H. Freed*,
Published on Web 04/19/2002 Protein Structure Determination Using Long-Distance Constraints from Double-Quantum Coherence ESR: Study of T4 Lysozyme Petr P. Borbat, Hassane S. Mchaourab, and Jack H. Freed*,
Site directed spin labelling and pulsed dipolar electron paramagnetic resonance (double electron electron resonance) of force activation in muscle
 INTITUTE OF PHYI PUBLIHING JOURNAL OF PHYI: ONDENED MATTER J. Phys.: ondens. Matter 17 (2005) 1459 1469 doi:10.1088/0953-8984/17/18/004 ite directed spin labelling and pulsed dipolar electron paramagnetic
INTITUTE OF PHYI PUBLIHING JOURNAL OF PHYI: ONDENED MATTER J. Phys.: ondens. Matter 17 (2005) 1459 1469 doi:10.1088/0953-8984/17/18/004 ite directed spin labelling and pulsed dipolar electron paramagnetic
Archived Information. Comment on Distinct Populations in Spin-Label EPR Spectra from Nitroxides [J. Phys. Chem. B 2018, 122, ]
![Archived Information. Comment on Distinct Populations in Spin-Label EPR Spectra from Nitroxides [J. Phys. Chem. B 2018, 122, ] Archived Information. Comment on Distinct Populations in Spin-Label EPR Spectra from Nitroxides [J. Phys. Chem. B 2018, 122, ]](/thumbs/93/112745958.jpg) Archived Information Comment on Distinct Populations in Spin-Label EPR Spectra from Nitroxides [J. Phys. Chem. B 2018, 122, 6129-6133] Eva Meirovitch, 1,* Boris Dzikovski, 2 and Jack H. Freed 2,* 1 The
Archived Information Comment on Distinct Populations in Spin-Label EPR Spectra from Nitroxides [J. Phys. Chem. B 2018, 122, 6129-6133] Eva Meirovitch, 1,* Boris Dzikovski, 2 and Jack H. Freed 2,* 1 The
e 2m e c I, (7.1) = g e β B I(I +1), (7.2) = erg/gauss. (7.3)
 Chemistry 126 Molecular Spectra & Molecular Structure Week # 7 Electron Spin Resonance Spectroscopy, Supplement Like the hydrogen nucleus, an unpaired electron in a sample has a spin of I=1/2. The magnetic
Chemistry 126 Molecular Spectra & Molecular Structure Week # 7 Electron Spin Resonance Spectroscopy, Supplement Like the hydrogen nucleus, an unpaired electron in a sample has a spin of I=1/2. The magnetic
ELECTRON PARAMAGNETIC RESONANCE
 ELECTRON PARAMAGNETIC RESONANCE = MAGNETIC RESONANCE TECHNIQUE FOR STUDYING PARAMAGNETIC SYSTEMS i.e. SYSTEMS WITH AT LEAST ONE UNPAIRED ELECTRON Examples of paramagnetic systems Transition-metal complexes
ELECTRON PARAMAGNETIC RESONANCE = MAGNETIC RESONANCE TECHNIQUE FOR STUDYING PARAMAGNETIC SYSTEMS i.e. SYSTEMS WITH AT LEAST ONE UNPAIRED ELECTRON Examples of paramagnetic systems Transition-metal complexes
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Diphthamide biosynthesis requires a radical iron-sulfur enzyme. Pennsylvania State University, University Park, Pennsylvania 16802, USA
 Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Diphthamide biosynthesis requires a radical iron-sulfur enzyme Yang Zhang, 1,4 Xuling Zhu, 1,4 Andrew T. Torelli, 1 Michael Lee, 2 Boris Dzikovski, 1 Rachel Koralewski, 1 Eileen Wang, 1 Jack Freed, 1 Carsten
Biophysical Chemistry: NMR Spectroscopy
 Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
Originally published in: Annual Review of Physical Chemistry 63,
 Research Collection Review Article DEER Distance Measurements on Proteins Author(s): Jeschke, Gunnar Publication Date: 2012-05 Permanent Link: https://doi.org/10.3929/ethz-a-010784314 Originally published
Research Collection Review Article DEER Distance Measurements on Proteins Author(s): Jeschke, Gunnar Publication Date: 2012-05 Permanent Link: https://doi.org/10.3929/ethz-a-010784314 Originally published
Matthias Lütgens, Frank Friedriszik, and Stefan Lochbrunner* 1 Concentration dependent CARS and Raman spectra of acetic acid in carbon tetrachloride
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 SUPPORTING INFORMATION Direct observation of the cyclic dimer in liquid acetic
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2014 SUPPORTING INFORMATION Direct observation of the cyclic dimer in liquid acetic
Ifs and Buts of DEER. Likai Song, Ilker Sen, Marco Bonora, Florida State University, NHMFL
 Ifs and Buts of DEER Likai Song, Ilker Sen, Marco Bonora, P. Fajer Florida State University, NHMFL 1. X-band v. W-band: is bigger better? DEER STEPR 2. DEER Analysis: how not to over interpret the data?
Ifs and Buts of DEER Likai Song, Ilker Sen, Marco Bonora, P. Fajer Florida State University, NHMFL 1. X-band v. W-band: is bigger better? DEER STEPR 2. DEER Analysis: how not to over interpret the data?
Supporting Information
 Supporting Information Oxaliplatin binding to human copper chaperone Atox1 and protein dimerization Benny D. Belviso, 1 Angela Galliani, 2 Alessia Lasorsa, 2 Valentina Mirabelli, 1,3 Rocco Caliandro, 1
Supporting Information Oxaliplatin binding to human copper chaperone Atox1 and protein dimerization Benny D. Belviso, 1 Angela Galliani, 2 Alessia Lasorsa, 2 Valentina Mirabelli, 1,3 Rocco Caliandro, 1
T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y. jgp
 S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4-5pm and SFL 229, Monday 3/5 4-5:30pm. 1. NMR
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4-5pm and SFL 229, Monday 3/5 4-5:30pm. 1. NMR
PLEASE SCROLL DOWN FOR ARTICLE
 This article was downloaded by:[prisner, T. F.] On: 18 December 2007 Access Details: [subscription number 788607874] Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number:
This article was downloaded by:[prisner, T. F.] On: 18 December 2007 Access Details: [subscription number 788607874] Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number:
A Combined Optical and EPR Spectroscopy Study: Azobenzene-Based Biradicals as Reversible Molecular Photoswitches
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 A Combined Optical and EPR Spectroscopy Study: Azobenzene-Based Biradicals as Reversible
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2017 A Combined Optical and EPR Spectroscopy Study: Azobenzene-Based Biradicals as Reversible
pyridoxal phosphate synthase
 Supplementary Information 13 C-NMR snapshots of the complex reaction coordinate of pyridoxal phosphate synthase Jeremiah W. Hanes, Ivan Keresztes, and Tadhg P. Begley * Department of Chemistry and Chemical
Supplementary Information 13 C-NMR snapshots of the complex reaction coordinate of pyridoxal phosphate synthase Jeremiah W. Hanes, Ivan Keresztes, and Tadhg P. Begley * Department of Chemistry and Chemical
METHODOLOGIES AND APPLICATION DEVELOPMENT OF HIGH FIELD PELDOR FOR SPIN LABELLED PROTEINS. Johannes Erik McKay
 METHODOLOGIES AND APPLICATION DEVELOPMENT OF HIGH FIELD PELDOR FOR SPIN LABELLED PROTEINS Johannes Erik McKay A Thesis Submitted for the Degree of PhD at the University of St Andrews 2016 Full metadata
METHODOLOGIES AND APPLICATION DEVELOPMENT OF HIGH FIELD PELDOR FOR SPIN LABELLED PROTEINS Johannes Erik McKay A Thesis Submitted for the Degree of PhD at the University of St Andrews 2016 Full metadata
Methods EPR sample preparation: EPR Spectroscopy:
 Methods EPR sample preparation: For CW dipolar EPR measurements (Fig. 2 A), 32-TOAC-DR5 was reconstituted at 0.4 mm in DPC buffer (15 mm dodecyl phosphocholine, 50 mm phosphate buffer, ph 8.1), with 1
Methods EPR sample preparation: For CW dipolar EPR measurements (Fig. 2 A), 32-TOAC-DR5 was reconstituted at 0.4 mm in DPC buffer (15 mm dodecyl phosphocholine, 50 mm phosphate buffer, ph 8.1), with 1
Theory of double quantum two-dimensional electron spin resonance with application to distance measurements
 Theory of double quantum two-dimensional electron spin resonance with application to distance measurements Sunil Saxena and Jack H. Freed Baker Laboratory of Chemistry, Cornell University, Ithaca, New
Theory of double quantum two-dimensional electron spin resonance with application to distance measurements Sunil Saxena and Jack H. Freed Baker Laboratory of Chemistry, Cornell University, Ithaca, New
Pulsed Dipolar EPR Spectroscopy
 Pulsed Dipolar EPR Spectroscopy Maxim Yulikov (Zurich, Switzerland) V International School for Young Scientists Magnetic Resonance and Magnetic Phenomena in Chemical and Biological Physics September 5-2,
Pulsed Dipolar EPR Spectroscopy Maxim Yulikov (Zurich, Switzerland) V International School for Young Scientists Magnetic Resonance and Magnetic Phenomena in Chemical and Biological Physics September 5-2,
Supplementary Figure 1 Interlayer exciton PL peak position and heterostructure twisting angle. a, Photoluminescence from the interlayer exciton for
 Supplementary Figure 1 Interlayer exciton PL peak position and heterostructure twisting angle. a, Photoluminescence from the interlayer exciton for six WSe 2 -MoSe 2 heterostructures under cw laser excitation
Supplementary Figure 1 Interlayer exciton PL peak position and heterostructure twisting angle. a, Photoluminescence from the interlayer exciton for six WSe 2 -MoSe 2 heterostructures under cw laser excitation
BCMB / CHEM 8190 Biomolecular NMR GRADUATE COURSE OFFERING IN NUCLEAR MAGNETIC RESONANCE
 BCMB / CHEM 8190 Biomolecular NMR GRADUATE COURSE OFFERING IN NUCLEAR MAGNETIC RESONANCE "Biomolecular Nuclear Magnetic Resonance" is a course intended for all graduate students with an interest in applications
BCMB / CHEM 8190 Biomolecular NMR GRADUATE COURSE OFFERING IN NUCLEAR MAGNETIC RESONANCE "Biomolecular Nuclear Magnetic Resonance" is a course intended for all graduate students with an interest in applications
Supplementary Materials belonging to
 Supplementary Materials belonging to Solution conformation of the 70 kda E.coli Hsp70 complexed with ADP and substrate. Eric B. Bertelsen 1, Lyra Chang 2, Jason E. Gestwicki 2 and Erik R. P. Zuiderweg
Supplementary Materials belonging to Solution conformation of the 70 kda E.coli Hsp70 complexed with ADP and substrate. Eric B. Bertelsen 1, Lyra Chang 2, Jason E. Gestwicki 2 and Erik R. P. Zuiderweg
Solid-State NMR Distance & Dynamics Techniques for Structure Determination of the Influenza M2 Protein
 Solid-State NMR Distance & Dynamics Techniques for Structure Determination of the Influenza M2 Protein Mei Hong Department of Chemistry and Ames Laboratory, Iowa State University 2nd CCPN / Extend-NMR
Solid-State NMR Distance & Dynamics Techniques for Structure Determination of the Influenza M2 Protein Mei Hong Department of Chemistry and Ames Laboratory, Iowa State University 2nd CCPN / Extend-NMR
Determining the Oligomeric Structure of Proteorhodopsin by Gd 3+ -Based Pulsed Dipolar Spectroscopy of Multiple Distances
 Resource Determining the Oligomeric Structure of Proteorhodopsin by Gd 3+ -Based Pulsed Dipolar Spectroscopy of Multiple Distances Devin T. Edwards, 1 Thomas Huber, 2 Sunyia Hussain, 3 Katherine M. Stone,
Resource Determining the Oligomeric Structure of Proteorhodopsin by Gd 3+ -Based Pulsed Dipolar Spectroscopy of Multiple Distances Devin T. Edwards, 1 Thomas Huber, 2 Sunyia Hussain, 3 Katherine M. Stone,
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS
 Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Biological Opportunities with Solution Scattering. Brian R. Crane
 Biological Opportunities with Solution Scattering XDL 2011 Brian R. Crane Cornell University, Ithaca NY bc69@cornell.edu Bacterial Transmembrane Receptors Histidine kinases, adenylyl cyclases, methyl accepting
Biological Opportunities with Solution Scattering XDL 2011 Brian R. Crane Cornell University, Ithaca NY bc69@cornell.edu Bacterial Transmembrane Receptors Histidine kinases, adenylyl cyclases, methyl accepting
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Quantification of Dynamics in the Solid-State
 Bernd Reif Quantification of Dynamics in the Solid-State Technische Universität München Helmholtz-Zentrum München Biomolecular Solid-State NMR Winter School Stowe, VT January 0-5, 206 Motivation. Solid
Bernd Reif Quantification of Dynamics in the Solid-State Technische Universität München Helmholtz-Zentrum München Biomolecular Solid-State NMR Winter School Stowe, VT January 0-5, 206 Motivation. Solid
Electron Spin Resonance Spectroscopy: A Renaissance
 Electron Spin Resonance Spectroscopy: A Renaissance Jack H. Freed Department of Chemistry and Chemical Biology & ACERT Cornell University Ithaca, NY, USA ACS 235 th National Meeting Physical Chemistry
Electron Spin Resonance Spectroscopy: A Renaissance Jack H. Freed Department of Chemistry and Chemical Biology & ACERT Cornell University Ithaca, NY, USA ACS 235 th National Meeting Physical Chemistry
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Spectroscopy of Polymers
 Spectroscopy of Polymers Jack L. Koenig Case Western Reserve University WOMACS Professional Reference Book American Chemical Society, Washington, DC 1992 Contents Preface m xiii Theory of Polymer Characterization
Spectroscopy of Polymers Jack L. Koenig Case Western Reserve University WOMACS Professional Reference Book American Chemical Society, Washington, DC 1992 Contents Preface m xiii Theory of Polymer Characterization
Christopher Pavlik Bioanalytical Chemistry March 2, 2011
 Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Raman Optical Activity Comes of Age
 Raman Optical Activity Comes of Age Laurence D. Barron Department of Chemistry, University of Glasgow, Glasgow G12 8QQ, UK E-mail: laurence.barron@glasgow.ac.uk Raman optical activity (ROA) provides vibrational
Raman Optical Activity Comes of Age Laurence D. Barron Department of Chemistry, University of Glasgow, Glasgow G12 8QQ, UK E-mail: laurence.barron@glasgow.ac.uk Raman optical activity (ROA) provides vibrational
XV 74. Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go?
 XV 74 Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go? 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet S = 0 could be any of them
XV 74 Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go? 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet S = 0 could be any of them
Structure of p53 binding to the BAX response element reveals DNA. unwinding and compression to accommodate base-pair insertion
 Structure of p53 binding to the BAX response element reveals DNA unwinding and compression to accommodate base-pair insertion Yongheng Chen 1,2,*, Xiaojun Zhang 3, Ana Carolina Dantas Machado 1, Yuan Ding
Structure of p53 binding to the BAX response element reveals DNA unwinding and compression to accommodate base-pair insertion Yongheng Chen 1,2,*, Xiaojun Zhang 3, Ana Carolina Dantas Machado 1, Yuan Ding
Protein dynamics from NMR Relaxation data
 Protein dynamics from NMR Relaxation data Clubb 3/15/17 (S f2 ) ( e ) Nitrogen-15 relaxation ZZ-exchange R 1 = 1/T 1 Longitudinal relaxation (decay back to z-axis) R 2 = 1/T 2 Spin-spin relaxation (dephasing
Protein dynamics from NMR Relaxation data Clubb 3/15/17 (S f2 ) ( e ) Nitrogen-15 relaxation ZZ-exchange R 1 = 1/T 1 Longitudinal relaxation (decay back to z-axis) R 2 = 1/T 2 Spin-spin relaxation (dephasing
Quantum technologies based on nitrogen-vacancy centers in diamond: towards applications in (quantum) biology
 Quantum technologies based on nitrogen-vacancy centers in diamond: towards applications in (quantum) biology 3 E 532 nm 1 2δω 1 Δ ESR 0 1 A 1 3 A 2 Microwaves 532 nm polarization Pulse sequence detection
Quantum technologies based on nitrogen-vacancy centers in diamond: towards applications in (quantum) biology 3 E 532 nm 1 2δω 1 Δ ESR 0 1 A 1 3 A 2 Microwaves 532 nm polarization Pulse sequence detection
Superoperators for NMR Quantum Information Processing. Osama Usman June 15, 2012
 Superoperators for NMR Quantum Information Processing Osama Usman June 15, 2012 Outline 1 Prerequisites 2 Relaxation and spin Echo 3 Spherical Tensor Operators 4 Superoperators 5 My research work 6 References.
Superoperators for NMR Quantum Information Processing Osama Usman June 15, 2012 Outline 1 Prerequisites 2 Relaxation and spin Echo 3 Spherical Tensor Operators 4 Superoperators 5 My research work 6 References.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR
 Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR Residual Dipolar Couplings (RDC s) Relatively new technique ~ 1996 Nico Tjandra, Ad Bax- NIH, Jim Prestegard, UGA Combination of
Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR Residual Dipolar Couplings (RDC s) Relatively new technique ~ 1996 Nico Tjandra, Ad Bax- NIH, Jim Prestegard, UGA Combination of
Ferdowsi University of Mashhad
 Spectroscopy in Inorganic Chemistry Nuclear Magnetic Resonance Spectroscopy spin deuterium 2 helium 3 The neutron has 2 quarks with a -e/3 charge and one quark with a +2e/3 charge resulting in a total
Spectroscopy in Inorganic Chemistry Nuclear Magnetic Resonance Spectroscopy spin deuterium 2 helium 3 The neutron has 2 quarks with a -e/3 charge and one quark with a +2e/3 charge resulting in a total
NMR, X-ray Diffraction, Protein Structure, and RasMol
 NMR, X-ray Diffraction, Protein Structure, and RasMol Introduction So far we have been mostly concerned with the proteins themselves. The techniques (NMR or X-ray diffraction) used to determine a structure
NMR, X-ray Diffraction, Protein Structure, and RasMol Introduction So far we have been mostly concerned with the proteins themselves. The techniques (NMR or X-ray diffraction) used to determine a structure
Nature Structural & Molecular Biology: doi: /nsmb.3194
 Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Supplementary Figures:
 Supplementary Figures: dcdtbt vibration spectrum: Ground state blue vs Cation state red Intensity a.u. 1000 1100 1200 1300 1400 1500 1600 1700 Frequency cm^1 dcdtbt vibration spectrum: Ground state blue
Supplementary Figures: dcdtbt vibration spectrum: Ground state blue vs Cation state red Intensity a.u. 1000 1100 1200 1300 1400 1500 1600 1700 Frequency cm^1 dcdtbt vibration spectrum: Ground state blue
PROTEIN NMR SPECTROSCOPY
 List of Figures List of Tables xvii xxvi 1. NMR SPECTROSCOPY 1 1.1 Introduction to NMR Spectroscopy 2 1.2 One Dimensional NMR Spectroscopy 3 1.2.1 Classical Description of NMR Spectroscopy 3 1.2.2 Nuclear
List of Figures List of Tables xvii xxvi 1. NMR SPECTROSCOPY 1 1.1 Introduction to NMR Spectroscopy 2 1.2 One Dimensional NMR Spectroscopy 3 1.2.1 Classical Description of NMR Spectroscopy 3 1.2.2 Nuclear
Appendix A Anti-parallel aggregation of WALP peptides in DOPC
 Appendix A Appendix A Anti-parallel aggregation of WALP peptides in DOPC The study of WALP spin labeled at the central position had suggested linear aggregates in DOPC and cluster aggregates in DPPC, both
Appendix A Appendix A Anti-parallel aggregation of WALP peptides in DOPC The study of WALP spin labeled at the central position had suggested linear aggregates in DOPC and cluster aggregates in DPPC, both
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
NMR Spectroscopy of Polymers
 UNESCO/IUPAC Course 2005/2006 Jiri Brus NMR Spectroscopy of Polymers Brus J 1. part At the very beginning the phenomenon of nuclear spin resonance was studied predominantly by physicists and the application
UNESCO/IUPAC Course 2005/2006 Jiri Brus NMR Spectroscopy of Polymers Brus J 1. part At the very beginning the phenomenon of nuclear spin resonance was studied predominantly by physicists and the application
Supplementary Information. Overlap between folding and functional energy landscapes for. adenylate kinase conformational change
 Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
Supplementary Information Overlap between folding and functional energy landscapes for adenylate kinase conformational change by Ulrika Olsson & Magnus Wolf-Watz Contents: 1. Supplementary Note 2. Supplementary
ESR Spectroscopy in Membrane Biophysics. Volume 27
 ESR Spectroscopy in Membrane Biophysics Volume 27 A Continuation Order Plan is available for this series. A continuation order will bring delivery of each new volume immediately upon publication. Volumes
ESR Spectroscopy in Membrane Biophysics Volume 27 A Continuation Order Plan is available for this series. A continuation order will bring delivery of each new volume immediately upon publication. Volumes
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1. Comparison of apoa-i with and without helix 5.
 Supplementary Figure 1 Comparison of apoa-i with and without helix 5. (a) Overlay of two TROSY-HNCO spectra of ApoA-I with helix 5 (red spectrum) and without helix 5 (black spectrum). Virtually identical
Supplementary Figure 1 Comparison of apoa-i with and without helix 5. (a) Overlay of two TROSY-HNCO spectra of ApoA-I with helix 5 (red spectrum) and without helix 5 (black spectrum). Virtually identical
Chapter 6. The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR
 The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
Light irradiation experiments with coumarin [1]
![Light irradiation experiments with coumarin [1] Light irradiation experiments with coumarin [1]](/thumbs/88/114731970.jpg) Materials and instruments All the chemicals were purchased from commercial suppliers and used as received. Thin-layer chromatography (TLC) analysis was carried out on pre-coated silica plates. Column chromatography
Materials and instruments All the chemicals were purchased from commercial suppliers and used as received. Thin-layer chromatography (TLC) analysis was carried out on pre-coated silica plates. Column chromatography
Intensity (a.u.) 2Theta (degrees) Supplementary data O 3 BO 3. Sup-Figure 1: Room temperature XRD patterns of KCaBO 3 :Eu 3+ sample treated at 500 ºC.
 Supplementary data The following results and discussions are for the low-temperature (5 C) processed phosphors. 5 ºC Intensity (a.u.) KCaBO 3 Eu 2 O 3 H 3 BO 3 2 3 4 5 2Theta (degrees) 6 7 Sup-Figure 1:
Supplementary data The following results and discussions are for the low-temperature (5 C) processed phosphors. 5 ºC Intensity (a.u.) KCaBO 3 Eu 2 O 3 H 3 BO 3 2 3 4 5 2Theta (degrees) 6 7 Sup-Figure 1:
NMR Spectroscopy Laboratory Experiment Introduction. 2. Theory
 1. Introduction 64-311 Laboratory Experiment 11 NMR Spectroscopy Nuclear Magnetic Resonance (NMR) spectroscopy is a powerful and theoretically complex analytical tool. This experiment will introduce to
1. Introduction 64-311 Laboratory Experiment 11 NMR Spectroscopy Nuclear Magnetic Resonance (NMR) spectroscopy is a powerful and theoretically complex analytical tool. This experiment will introduce to
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray
 CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CONTENTS. 2 CLASSICAL DESCRIPTION 2.1 The resonance phenomenon 2.2 The vector picture for pulse EPR experiments 2.3 Relaxation and the Bloch equations
 CONTENTS Preface Acknowledgements Symbols Abbreviations 1 INTRODUCTION 1.1 Scope of pulse EPR 1.2 A short history of pulse EPR 1.3 Examples of Applications 2 CLASSICAL DESCRIPTION 2.1 The resonance phenomenon
CONTENTS Preface Acknowledgements Symbols Abbreviations 1 INTRODUCTION 1.1 Scope of pulse EPR 1.2 A short history of pulse EPR 1.3 Examples of Applications 2 CLASSICAL DESCRIPTION 2.1 The resonance phenomenon
NMR in Structural Biology
 NMR in Structural Biology Exercise session 2 1. a. List 3 NMR observables that report on structure. b. Also indicate whether the information they give is short/medium or long-range, or perhaps all three?
NMR in Structural Biology Exercise session 2 1. a. List 3 NMR observables that report on structure. b. Also indicate whether the information they give is short/medium or long-range, or perhaps all three?
Supplemental data for
 Supplemental data for A Real-Time Guanine Nucleotide Exchange Assay using NMR: Activation of RhoA by PDZ- RhoGEF. Geneviève M.C. Gasmi-Seabrook 1,3, Christopher B. Marshall 1,3, Melissa Cheung 1,3, Bryan
Supplemental data for A Real-Time Guanine Nucleotide Exchange Assay using NMR: Activation of RhoA by PDZ- RhoGEF. Geneviève M.C. Gasmi-Seabrook 1,3, Christopher B. Marshall 1,3, Melissa Cheung 1,3, Bryan
Structure and RNA-binding properties. of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex
 Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
Structure and RNA-binding properties of the Not1 Not2 Not5 module of the yeast Ccr4 Not complex Varun Bhaskar 1, Vladimir Roudko 2,3, Jerome Basquin 1, Kundan Sharma 4, Henning Urlaub 4, Bertrand Seraphin
DNP enhanced frequency-selective TEDOR experiments in bacteriorhodopsin
 DNP enhanced frequency-selective TEDOR experiments in bacteriorhodopsin Journal of Magnetic Resonance 202 (2010) 9-13 Bajaj S. V., Mak-Jurkauskus, A. L., Belenky, M., Herzfeld, J. and Griffin, R. MR Seminar
DNP enhanced frequency-selective TEDOR experiments in bacteriorhodopsin Journal of Magnetic Resonance 202 (2010) 9-13 Bajaj S. V., Mak-Jurkauskus, A. L., Belenky, M., Herzfeld, J. and Griffin, R. MR Seminar
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Supporting Information Elucidating Lithium-Ion and Proton Dynamics in Anti- Perovskite Solid Electrolytes
 Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2018 Supporting Information Elucidating Lithium-Ion and Proton Dynamics in Anti-
Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2018 Supporting Information Elucidating Lithium-Ion and Proton Dynamics in Anti-
Supplementary Materials
 Supplementary Materials Sample characterization The presence of Si-QDs is established by Transmission Electron Microscopy (TEM), by which the average QD diameter of d QD 2.2 ± 0.5 nm has been determined
Supplementary Materials Sample characterization The presence of Si-QDs is established by Transmission Electron Microscopy (TEM), by which the average QD diameter of d QD 2.2 ± 0.5 nm has been determined
Principles of Nuclear Magnetic Resonance in One and Two Dimensions
 Principles of Nuclear Magnetic Resonance in One and Two Dimensions Richard R. Ernst, Geoffrey Bodenhausen, and Alexander Wokaun Laboratorium für Physikalische Chemie Eidgenössische Technische Hochschule
Principles of Nuclear Magnetic Resonance in One and Two Dimensions Richard R. Ernst, Geoffrey Bodenhausen, and Alexander Wokaun Laboratorium für Physikalische Chemie Eidgenössische Technische Hochschule
Coupling of Functional Hydrogen Bonds in Pyridoxal-5 -phosphate- Enzyme Model Systems Observed by Solid State NMR
 Supporting Information for Coupling of Functional ydrogen Bonds in Pyridoxal-5 -phosphate- Enzyme Model Systems bserved by Solid State NMR Shasad Sharif, David Schagen, Michael D. Toney, and ans-einrich
Supporting Information for Coupling of Functional ydrogen Bonds in Pyridoxal-5 -phosphate- Enzyme Model Systems bserved by Solid State NMR Shasad Sharif, David Schagen, Michael D. Toney, and ans-einrich
NMR SATELLITES AS A PROBE FOR CHEMICAL
 NMR SATELLITES AS A PROBE FOR CHEMICAL INVESTIGATIONS SHIzuo FUJIWARA, Yogi ARATA, HIR0sHI OZAWA and MASAYUKI KuruGI Department of Chemistry, The University of Tokyo, Japan ABSTRACT Satellite lines in
NMR SATELLITES AS A PROBE FOR CHEMICAL INVESTIGATIONS SHIzuo FUJIWARA, Yogi ARATA, HIR0sHI OZAWA and MASAYUKI KuruGI Department of Chemistry, The University of Tokyo, Japan ABSTRACT Satellite lines in
Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10
 Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10 Instructions: This is an open book, limited time, exam. You may use notes you have from class and any text book you find useful. You may also use
Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10 Instructions: This is an open book, limited time, exam. You may use notes you have from class and any text book you find useful. You may also use
Supplementary Figure 1: Spin noise spectra of 55 Mn in bulk sample at BL =10.5 mt, before subtraction of the zero-frequency line. a, Contour plot of
 1 Supplementary Figure 1: Spin noise spectra of 55 Mn in bulk sample at BL =10.5 mt, before subtraction of the zero-frequency line. a, Contour plot of the spin noise spectra calculated with Eq. (2) for
1 Supplementary Figure 1: Spin noise spectra of 55 Mn in bulk sample at BL =10.5 mt, before subtraction of the zero-frequency line. a, Contour plot of the spin noise spectra calculated with Eq. (2) for
Structural characterization of NiV N 0 P in solution and in crystal.
 Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
Supplementary Figure 1 Structural characterization of NiV N 0 P in solution and in crystal. (a) SAXS analysis of the N 32-383 0 -P 50 complex. The Guinier plot for complex concentrations of 0.55, 1.1,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/NCHEM.1299 Protein fold determined by paramagnetic magic-angle spinning solid-state NMR spectroscopy Ishita Sengupta 1, Philippe S. Nadaud 1, Jonathan J. Helmus 1, Charles D. Schwieters 2
DOI: 10.1038/NCHEM.1299 Protein fold determined by paramagnetic magic-angle spinning solid-state NMR spectroscopy Ishita Sengupta 1, Philippe S. Nadaud 1, Jonathan J. Helmus 1, Charles D. Schwieters 2
Room Temperature Quantum Coherence and Rabi Oscillations in Vanadyl Phthalocyanine: Toward Multifunctional Molecular Spin Qubits
 Room Temperature Quantum Coherence and Rabi Oscillations in Vanadyl Phthalocyanine: Toward Multifunctional Molecular Spin Qubits Matteo Atzori, Lorenzo Tesi, Elena Morra, Mario Chiesa, Lorenzo Sorace,
Room Temperature Quantum Coherence and Rabi Oscillations in Vanadyl Phthalocyanine: Toward Multifunctional Molecular Spin Qubits Matteo Atzori, Lorenzo Tesi, Elena Morra, Mario Chiesa, Lorenzo Sorace,
Asian Journal of Chemistry; Vol. 25, No. 4 (2013),
 Asian Journal of Chemistry; Vol. 25, No. 4 (213), 214-218 http://dx.doi.org/1.14233/ajchem.213.13346 Observation of Triplet Traces Obtained with Inversion Recovery Method in Both Residual Water- and H
Asian Journal of Chemistry; Vol. 25, No. 4 (213), 214-218 http://dx.doi.org/1.14233/ajchem.213.13346 Observation of Triplet Traces Obtained with Inversion Recovery Method in Both Residual Water- and H
NMR of large protein systems: Solid state and dynamic nuclear polarization. Sascha Lange, Leibniz-Institut für Molekulare Pharmakologie (FMP)
 NMR of large protein systems: Solid state and dynamic nuclear polarization Sascha Lange, Leibniz-Institut für Molekulare Pharmakologie (FMP) The Aim of the Game solution NMR other methods solid state NMR
NMR of large protein systems: Solid state and dynamic nuclear polarization Sascha Lange, Leibniz-Institut für Molekulare Pharmakologie (FMP) The Aim of the Game solution NMR other methods solid state NMR
DETECTION OF UNPAIRED ELECTRONS
 DETECTION OF UNPAIRED ELECTRONS There are experimental methods for the detection of unpaired electrons. One of the hallmarks of unpaired electrons in materials is interaction with a magnetic field. That
DETECTION OF UNPAIRED ELECTRONS There are experimental methods for the detection of unpaired electrons. One of the hallmarks of unpaired electrons in materials is interaction with a magnetic field. That
Spatially resolved two-dimensional Fourier transform electron spin resonance
 Spatially resolved two-dimensional Fourier transform electron spin resonance Uwe Ewert I, Richard H. Crepeau, Sanghyuk Lee, Curt R. Dunnam *, Dajiang Xu and Jack H. Freed Bakerkboratory of Chemistry, Cornell
Spatially resolved two-dimensional Fourier transform electron spin resonance Uwe Ewert I, Richard H. Crepeau, Sanghyuk Lee, Curt R. Dunnam *, Dajiang Xu and Jack H. Freed Bakerkboratory of Chemistry, Cornell
Long-lived spin echoes in magnetically diluted system: an NMR study of the Ge single crystals Alexander M. Panich,
 Long-lived spin echoes in magnetically diluted system: an NMR study of the Ge single crystals Alexander M. Panich, Department of Physics, Ben-Gurion University of the Negev, Beer Sheva, Israel N. A. Sergeev,
Long-lived spin echoes in magnetically diluted system: an NMR study of the Ge single crystals Alexander M. Panich, Department of Physics, Ben-Gurion University of the Negev, Beer Sheva, Israel N. A. Sergeev,
Circular Dichroism & Optical Rotatory Dispersion. Proteins (KCsa) Polysaccharides (agarose) DNA CHEM 305. Many biomolecules are α-helical!
 Circular Dichroism & Optical Rotatory Dispersion Polysaccharides (agarose) DNA Proteins (KCsa) Many biomolecules are α-helical! How can we measure the amount and changes in amount of helical structure
Circular Dichroism & Optical Rotatory Dispersion Polysaccharides (agarose) DNA Proteins (KCsa) Many biomolecules are α-helical! How can we measure the amount and changes in amount of helical structure
Observation of quadrupole helix chirality and its domain structure in DyFe 3 (BO 3 ) 4
 Observation of quadrupole helix chirality and its domain structure in DyFe 3 (BO 3 ) 4 T. Usui, Y. Tanaka, H. Nakajima, M. Taguchi, A. Chainani, M. Oura, S. Shin, N. Katayama, H. Sawa, Y. Wakabayashi,
Observation of quadrupole helix chirality and its domain structure in DyFe 3 (BO 3 ) 4 T. Usui, Y. Tanaka, H. Nakajima, M. Taguchi, A. Chainani, M. Oura, S. Shin, N. Katayama, H. Sawa, Y. Wakabayashi,
Fluorescence 2009 update
 XV 74 Fluorescence 2009 update Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
XV 74 Fluorescence 2009 update Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
Linear and nonlinear spectroscopy
 Linear and nonlinear spectroscopy We ve seen that we can determine molecular frequencies and dephasing rates (for electronic, vibrational, or spin degrees of freedom) from frequency-domain or timedomain
Linear and nonlinear spectroscopy We ve seen that we can determine molecular frequencies and dephasing rates (for electronic, vibrational, or spin degrees of freedom) from frequency-domain or timedomain
Protein Structure Determination from Pseudocontact Shifts Using ROSETTA
 Supporting Information Protein Structure Determination from Pseudocontact Shifts Using ROSETTA Christophe Schmitz, Robert Vernon, Gottfried Otting, David Baker and Thomas Huber Table S0. Biological Magnetic
Supporting Information Protein Structure Determination from Pseudocontact Shifts Using ROSETTA Christophe Schmitz, Robert Vernon, Gottfried Otting, David Baker and Thomas Huber Table S0. Biological Magnetic
Supplemental Information for. Quaternary dynamics of B crystallin as a direct consequence of localised tertiary fluctuations in the C terminus
 Supplemental Information for Quaternary dynamics of B crystallin as a direct consequence of localised tertiary fluctuations in the C terminus Andrew J. Baldwin 1, Gillian R. Hilton 2, Hadi Lioe 2, Claire
Supplemental Information for Quaternary dynamics of B crystallin as a direct consequence of localised tertiary fluctuations in the C terminus Andrew J. Baldwin 1, Gillian R. Hilton 2, Hadi Lioe 2, Claire
Determining Chemical Structures with NMR Spectroscopy the ADEQUATEAD Experiment
 Determining Chemical Structures with NMR Spectroscopy the ADEQUATEAD Experiment Application Note Author Paul A. Keifer, Ph.D. NMR Applications Scientist Agilent Technologies, Inc. Santa Clara, CA USA Abstract
Determining Chemical Structures with NMR Spectroscopy the ADEQUATEAD Experiment Application Note Author Paul A. Keifer, Ph.D. NMR Applications Scientist Agilent Technologies, Inc. Santa Clara, CA USA Abstract
SUPPLEMENTARY INFORMATION
 Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
Dph2 SeMet (iron-free) # Dph2 (iron-free) Dph2-[4Fe-4S] Data collection Space group P2 1 2 1 2 1 P2 1 2 1 2 1 P2 1 2 1 2 1 Cell dimensions a, b, c (Å) 58.26, 82.08, 160.42 58.74, 81.87, 160.01 55.70, 80.53,
dots) and max max without energies
 Supplementary Figure 1 Light-polarization-dependent the crystal b-axis. Scale bar, 25 m. (b) Polarization-dependent absorption spectra of bilayer ReS 2. (c) Corresponding spectral weights of Lorentzian
Supplementary Figure 1 Light-polarization-dependent the crystal b-axis. Scale bar, 25 m. (b) Polarization-dependent absorption spectra of bilayer ReS 2. (c) Corresponding spectral weights of Lorentzian
Molecular Modeling lecture 2
 Molecular Modeling 2018 -- lecture 2 Topics 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction 4. Where do protein structures come from? X-ray crystallography
Molecular Modeling 2018 -- lecture 2 Topics 1. Secondary structure 3. Sequence similarity and homology 2. Secondary structure prediction 4. Where do protein structures come from? X-ray crystallography
Introduction solution NMR
 2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
Chemical Science EDGE ARTICLE
 Chemical Science EDGE ARTICLE Cite this: Chem. Sci., 2013, 4, 2776 Ab initio calculations and validation of the ph-dependent structures of the His37-Trp41 quartet, the heart of acid activation and proton
Chemical Science EDGE ARTICLE Cite this: Chem. Sci., 2013, 4, 2776 Ab initio calculations and validation of the ph-dependent structures of the His37-Trp41 quartet, the heart of acid activation and proton
Structurele Biologie NMR
 MR journey Structurele Biologie MR 5 /3C 3 /65 MR & Structural biology course setup lectures - Sprangers R & Kay LE ature (27) basics of MR (Klaartje ouben: k.houben@uu.nl; 4/2) from peaks to data (ans
MR journey Structurele Biologie MR 5 /3C 3 /65 MR & Structural biology course setup lectures - Sprangers R & Kay LE ature (27) basics of MR (Klaartje ouben: k.houben@uu.nl; 4/2) from peaks to data (ans
You are advised to spend an equal amount of time on each question.
 UNIVERSITY OF EAST ANGLIA School of Chemistry Main Series UG Examination 2015-16 BIOPHYSICAL CHEMISTRY CHE-5601Y Time allowed: 2 hours Answer THREE questions. You are advised to spend an equal amount of
UNIVERSITY OF EAST ANGLIA School of Chemistry Main Series UG Examination 2015-16 BIOPHYSICAL CHEMISTRY CHE-5601Y Time allowed: 2 hours Answer THREE questions. You are advised to spend an equal amount of
