Circular Dichroism. For students of HI Computational Structural Biology
|
|
|
- Bruno Golden
- 6 years ago
- Views:
Transcription
1 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Circular Dichroism For students of HI Computational Structural Biology Willy Wriggers, Ph.D. Adopted from material by Mathew Baker, Univ. of Bath
2 What is Circular Dichroism? Circular Dichroism (CD) is a type of absorption spectroscopy that can provide information on the structures of many types of biological macromolecules It measures the difference between the absorption of left and right handed circularly-polarized light by proteins. CD is used for; Protein structure determination. Induced structural changes, i.e. ph, heat & solvent. Protein folding/unfolding. Ligand binding Structural aspects of nucleic acids, polysaccharides, peptides, hormones & other small molecules.
3 Plane Polarized Light E M Direction of propagation E vectors Polarizer Direction of propagation
4 Plane & Circularly Polarized Light A light source usually consists of a collection of randomly orientated emitters, the emitted light is a collection of waves with all possible orientations of the E vectors. Plane polarized light is is obtained by passing light through a polarizer that transmits light with only a single plane of polarization. i.e. it passes only those components of the E vector that are parallel to the axis of the polarizer. Circular polarized light; The E vectors of two electromagnetic waves are ¼ wavelength out of phase & are perpendicular. The vector that is the sum of the E vectors of the two components rotates so that its tip follows a helical path (dotted line).
5 Circularly Polarized Light Linearly polarized light: Electric vector direction constant - magnitude varies. Circularly polarized light: Electric vector direction varies - magnitude constant
6 Circularly Polarized Light Circular polarized light: Electric vector direction varies - magnitude constant So its in two forms: left and right handed E E Righthanded Lefthanded
7 Circular Dichroism CD measures the difference between the absorption of left and right handed circularly-polarized light. polarized light: This is measured as a function of wavelength, & the difference is always very small (<<1/10000 of total). After passing through the sample, the L & R beams have different amplitudes & the combination of the two unequal beams gives elliptically polarized light.hence, CD measures the ellipicity of the transmitted light (the light that remains that is not absorbed):
8 Absorption Spectroscopy Shine light through a sample and measure the proportion absorbed as a function of wavelength. Absorbance A = log(i 0 /I) Beer-Lambert law: A(λ) = ε(λ)lc ε: extinction coefficient wavelength λ l I 0 Sample conc. C I The longer the path or the more concentrated the sample, the higher the absorbance
9 Absorption Spectroscopy CD measures the difference between the absorption of left and right handed circularly-polarized light: A(λ) = A R (λ)-a L (λ) = [ε R (λ) - ε L (λ)]lc or A(λ) = ε (λ)lc ε is the difference in the extinction coefficients typically < 10 M -1 cm -1 typical ε around M -1 cm -1 So the CD signal is a very small difference between two large originals.
10 Absorption Spectroscopy CD is only observed at wavelengths where absorption occurs, in absorption bands. CD arises because of the interaction between different transition dipoles doing the absorption. As this depends on the relative orientation of different groups in space, the signal is very sensitive to conformation. So in general ε is much more conformation dependent that ε. We will concentrate on the electronic CD of peptides and proteins below 240nm. This region is dominated by the absorption of peptide bond and is sensitive to changes in secondary structure. Can also do CD in near UV (look at trp side chains), visible (cofactors etc.) and IR regions.
11 The peptide bond is inherently asymmetric & is always optically active. Any optical activity from side-chain chromophores is induced & results from interactions with asymmetrical neighbouring groups.
12 CD Signal is a Small Difference Between Two Large Originals CD of E.Coli DNA Native Denatured For instance at 260 nm ε = ~3 M -1 cm -1 ε = ~6000 M -1 cm -1 i.e. CD signal 0.05% of original need to measure signals ~1/100 of this!
13 Mean residue ellipicity in deg cm 2 dmol CD Signals for Different Secondary Structures wavelength in nm α-helix β-sheet random coil These are Fasman standard curves for polylysine in different environments E L E R > 0 E L E R < 0 (data from ftp://jgiqc.llnl.gov)
14 CD Spectra of Protein 2 ndary Structures α-helix β-sheet -ve band (nm) ve band (nm) β-turn L.H polypro II helix (weak) (strong) weak Random coil 200
15 CD Signals are Sensitive to 2 ndary Structure CD signals for GCN4-p1 O'Shea et al. Science (1989) 243:538 figure 3: 34µM GCN4-p1 in 0.15M NaCl, 10mM phosphate ph 7.0 GCN4-p1 is a coiled coil: [θ] in 1000 deg cm 2 dmol O C 50 O C 75 O C wavelength in nm 100% helical at 0 o C It melts to a random coil at high temperature
16 Applications of CD Determination of secondary structure of proteins that cannot be crystallised Investigation of the effect of e.g. drug binding on protein secondary structure Dynamic processes, e.g. protein folding Studies of the effects of environment on protein structure Secondary structure and super-secondary structure of membrane proteins Study of ligand-induced conformational changes Carbohydrate conformation Investigations of protein-protein and protein-nucleic acid interactions Fold recognition
17 Advantages Simple and quick experiments No extensive preparation Measurements on solution phase Relatively low concentrations/amounts of sample Microsecond time resolution Any size of macromolecule
18 Total Signal for a Protein Depends on its 2 ndary Structure chymotrypsin (~all β) lysozyme (mixed α & β) triosephosphate isomerase (mostly α some β) myoglobin (all α) Notice the progressive change in θ 222 as the amount of helix increases from chymotrypsin to myoglobin
19 Finding Proportion of 2 ndary Structures Fit the unknown curve θ u to a combination of standard curves. In the simplest case use the Fasman standards θ t = x α θ α + x β θ β + x c θ c Vary x α, x β and x c to give the best fit of θ t to θ u while x α + x β + x c = 1.0 Do this by least squares minimization Mean residue ellipicity in deg cm 2 dmol wavelength in nm α-helix β-sheet random coil
20 Example Fit: Myoglobin In this case: θ t = x α θ α + x β θ β + x c θ c fits best with x α = 80%, x β = 0% x c = 20% Mean residue ellipicity in deg cm 2 dmol α-helix β-sheet random coil agrees well with structure 78% helix, 22% coil wavelength in nm
21 Example Fit (2): GCN4-p1 [θ] in 1000 deg cm 2 dmol -1 MRE in 1000 degs cm 2 mol CD signals for GCN4-p1 O'Shea et al. Science (1989) 243:538 figure 3: 34µM GCN4-p1 in 0.15M NaCl, 10mM phosphate ph O C 50 O C 75 O C wavelength in nm fit to GCN4-p1 50 O C data original data best fit with mix of 0 O C and 75 O C spectra wavelength in nm At 0 o C 100% helix 75 o C 0% helix Q: what about 50 o C? θ t = x o θ o + x 75 θ 75 fits best with x o = 50%, x 75 = 50% Shows that at 50 o C 1/2 of peptide α-helix dimer 1/2 of peptide random coil monomer
22 However, CD Signal Depends Somewhat on Environment Can see this by looking at the effect of trifluoroethanol (TFE) on a coiled-coil similar to GCN4-p1 TFE induces helicity in all peptides MRE Effect of 50% TFE on a monomeric peptide peptide in water peptide in 50% TFE TFE wavelength in nm But on a coiled-coil breaks down helical dimer to single helices MRE Effect of 50% TFE on a coiled-coil TM-36 aqueous TM-36 + TFE TFE wavelength in nm Although 2ndry structure same CD changes Lau, Taneja and Hodges (1984) J.Biol.Chem. 259:
23 Best Fitting Procedures Use Many Different Proteins For Standard Spectra There are many different algorithms. All rely on using up to 20 CD spectra of proteins of known structure. By mixing these together a fit spectra is obtained for an unknown. For full details see Dichroweb: the online CD analysis tool Can generally get accuracies of 0.97 for helices, 0.75 for beta sheet, 0.50 for turns, and 0.89 for other structure types (Manavalan & Johnson, 1987, Anal. Biochem. 167, 76-85).
24 Limitations of Secondary Structure Analysis The simple deconvolution of a CD spectrum into 4 or 5 components which do not vary from one protein to another is a gross oversimplification. The reference CD spectra corresponding to 100% helix, sheet, turn etc are not directly applicable to proteins which contain short sections of the various structures e.g. The CD of an α-helix is known to increase with increasing helix length, CD of β-sheets are very sensitive to environment & geometry. Far UV curves (>275nm) can contain contributions from aromatic amino-acids, in practice CD is measured at wavelengths below this. The shapes of far UV CD curves depend on tertiary as well as secondary structure.
25 CD is Very Useful for Looking at Membrane Proteins Membrane proteins are difficult to study. Crystallography difficult - need to use detergents Even when structure obtained: Q- is it the same as in lipid membrane? CD ideal can do spectra of protein in lipid vesicles. We look at Staphylococcal α-hemolysin as an example
26 α-hemolysin Channel Formation Soluble monomer Lipid-bound monomer pore prepore From Walker et al. (1995) Chemistry & Biology 2: This model was built by combining CD results with mutagenesis cross-linking, channel measurements
27 α-hemolysin CD Results Tobkes et al. (1985) Biochemistry 24:1915 soluble monomer ---- lipid monomer assembled hexamer or heptamer Mean residue ellipicity in deg cm 2 dmol wavelength in nm The 3 spectra are similar So the 3 conformations have similar 2ndry structure What is the 2ndry structure? Is this normal for a membrane protein? α-helix β-sheet random coil
28 α-hemolysin Structures Crystal structure of pore Song et al. Science :1859 CD used to show that the structure in detergent was the same as that of the pore in lipid. Just about all β Recently have structure of related soluble monomer
29 Using CD to Test a Peptide Designed to be Controlled by Light Uses a bifunctional iodoacetamide derivative of azobenzene that cross links a pair of cys residues. The azobenzene group adopts a trans conformation in the dark but can be forced to adopt a cis conformation by exposure to visible light of the appropriate wavelength: dark trans unstable light cis stable N N hv' or time Designed peptide to be helical in the cis (light) but helix to be unstable in the dark hv N N
30 Using CD to Test a Peptide Designed to be Controlled by Light Dark adapted peptide After light exposure (380nm 7mW 5 mins dark trans unstable light cis stable Can roughly gauge helicity helicity = [θ] 222 /32000 In this case dark 11% helix, light 48% Kumita, Smart & Woolley PNAS (2000) 97:
31 Practicalities CD is based on measuring a very small difference between two large signals must be done carefully the Abs must be reasonable max between ~0.5 and ~1.5. Quartz cells path lengths between cm and 10 cm. 1cm and 0.1 cm common have to be careful with buffers TRIS bad - high UV abs Measure cell base line with solvent Then sample with same cell inserted same way around Turbidity kills - filter solutions Everything has to be clean For accurate 2ndary structure estimation must know concentration of sample
32 Typical Conditions for CD Protein Concentration: 0.25 mg/ml Cell Path Length: 1 mm Volume 400 µl Need very little sample 0.1 mg Concentration reasonable Stabilizers (Metal ions, etc.): minimum Buffer Concentration : 5 mm or as low as possible while maintaining protein stability A structural biology method that can give real answers in a day.
33 Instrumentation - Lab-Based Spectropolarimeter $120k+ automatic vs λ, time, temperature, stopped flow down to 190nm (if you are lucky) 450W Xe bulb - produces ozone: hazard for health and silver coated optics So flush with large amounts of N 2 use boil off from liquid N 2
34 mean residue ellipticity in 1000 deg cm 2 dmol -1 cd signal in millidegrees raw CD spectra wavelength in nm buffer baseline 7.5uM GramPM in buffer (1) 7.5uM GramPM in buffer(2) TFE baseline 7.5uM GramPM in TFE(1) 7.5uM GramPM in TFE(2) 7.5uM GramPM in TFE(3) Average of 5 runs - converted to MRE wavelength in nm CD Analysis GramPM in buffer GramPM in TFE Looking at the CD of a gramicidin suspension in water Raw spectrum - data every 0.2nm from 205nm to 250nm, each data point measured 5 times 1sec avg. Note the baselines, these vary from cell to cell or if instrument moved, new bulb, recalibrated. Note noise - that s why we measure so many points. Final spectra the average of 5 runs (with about 3 baselines). Result - gramicidin suspension in buffer has a novel CD spectrum But what is the structure??
35 Instrumentation: Synchrotron-Based Synchrotron - whiz electrons around a ring. Can be used to produce very intense radiation by wiggling beam. Commonly used to produce X rays (λ around 0.1nm) But can be used to push signals down to 160nm and below Great for fast stopped-flow to see rapid changes
36 Amyloid Diseases
37 Summary CD is a useful method for looking at secondary structures of proteins and peptides. It is an adaptation of standard absorption spectroscopy in which the difference in the abs between left and right hand circularly polarized light is measured. CD can be measured under a wide range of conditions - e.g., good for membrane proteins. CD can be used to measure change. CD compliments other more detailed techniques such as crystallography.
38 Resources Books: van Holde KE, Johnson W & Ho P, Principles of Physical Biochemistry Prentice Hall 1998 or Campbell, ID and Dwek, RA Biological Spectroscopy Benjamin/Cummings Publishing, MIT CD Links Birkbeck College CD Tutorial CD links page at Daresbury Synchrotron Dichroweb: online CD analysis tool
39 Figure and Text Credits Text and figures for this lecture were adapted from the following source: Mathew Baker, University of Bath
Circular Dichroism & Optical Rotatory Dispersion. Proteins (KCsa) Polysaccharides (agarose) DNA CHEM 305. Many biomolecules are α-helical!
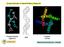 Circular Dichroism & Optical Rotatory Dispersion Polysaccharides (agarose) DNA Proteins (KCsa) Many biomolecules are α-helical! How can we measure the amount and changes in amount of helical structure
Circular Dichroism & Optical Rotatory Dispersion Polysaccharides (agarose) DNA Proteins (KCsa) Many biomolecules are α-helical! How can we measure the amount and changes in amount of helical structure
4. Circular Dichroism - Spectroscopy
 4. Circular Dichroism - Spectroscopy The optical rotatory dispersion (ORD) and the circular dichroism (CD) are special variations of absorption spectroscopy in the UV and VIS region of the spectrum. The
4. Circular Dichroism - Spectroscopy The optical rotatory dispersion (ORD) and the circular dichroism (CD) are special variations of absorption spectroscopy in the UV and VIS region of the spectrum. The
Protein folding. Today s Outline
 Protein folding Today s Outline Review of previous sessions Thermodynamics of folding and unfolding Determinants of folding Techniques for measuring folding The folding process The folding problem: Prediction
Protein folding Today s Outline Review of previous sessions Thermodynamics of folding and unfolding Determinants of folding Techniques for measuring folding The folding process The folding problem: Prediction
CONFOCHECK. Innovation with Integrity. Infrared Protein Analysis FT-IR
 CONFOCHECK Infrared Protein Analysis Innovation with Integrity FT-IR CONFOCHECK: FT-IR System for Protein Analytics FT-IR Protein Analysis Infrared spectroscopy measures molecular vibrations due to the
CONFOCHECK Infrared Protein Analysis Innovation with Integrity FT-IR CONFOCHECK: FT-IR System for Protein Analytics FT-IR Protein Analysis Infrared spectroscopy measures molecular vibrations due to the
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray
 CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
CD Basis Set of Spectra that is used is that derived from comparing the spectra of globular proteins whose secondary structures are known from X-ray crystallography An example of the use of CD Modeling
THE UNIVERSITY OF MANITOBA. PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS
 PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS EXAMINATION: Biochemistry of Proteins EXAMINER: J. O'Neil Section 1: You must answer all of
PAPER NO: 409 LOCATION: Fr. Kennedy Gold Gym PAGE NO: 1 of 6 DEPARTMENT & COURSE NO: CHEM 4630 TIME: 3 HOURS EXAMINATION: Biochemistry of Proteins EXAMINER: J. O'Neil Section 1: You must answer all of
BIRKBECK COLLEGE (University of London)
 BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
BIRKBECK COLLEGE (University of London) SCHOOL OF BIOLOGICAL SCIENCES M.Sc. EXAMINATION FOR INTERNAL STUDENTS ON: Postgraduate Certificate in Principles of Protein Structure MSc Structural Molecular Biology
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Visible and IR Absorption Spectroscopy. Andrew Rouff and Kyle Chau
 Visible and IR Absorption Spectroscopy Andrew Rouff and Kyle Chau The Basics wavelength= (λ) original intensity= Ι o sample slab thickness= dl Final intensity= I f ε = molar extinction coefficient -di=
Visible and IR Absorption Spectroscopy Andrew Rouff and Kyle Chau The Basics wavelength= (λ) original intensity= Ι o sample slab thickness= dl Final intensity= I f ε = molar extinction coefficient -di=
Human Biology. The Chemistry of Living Things. Concepts and Current Issues. All Matter Consists of Elements Made of Atoms
 2 The Chemistry of Living Things PowerPoint Lecture Slide Presentation Robert J. Sullivan, Marist College Michael D. Johnson Human Biology Concepts and Current Issues THIRD EDITION Copyright 2006 Pearson
2 The Chemistry of Living Things PowerPoint Lecture Slide Presentation Robert J. Sullivan, Marist College Michael D. Johnson Human Biology Concepts and Current Issues THIRD EDITION Copyright 2006 Pearson
Spectroscopy Chapter 13
 Spectroscopy Chapter 13 Electromagnetic Spectrum Electromagnetic spectrum in terms of wavelength, frequency and Energy c=λν c= speed of light in a vacuum 3x108 m/s v= frequency in Hertz (Hz s-1 ) λ= wavelength
Spectroscopy Chapter 13 Electromagnetic Spectrum Electromagnetic spectrum in terms of wavelength, frequency and Energy c=λν c= speed of light in a vacuum 3x108 m/s v= frequency in Hertz (Hz s-1 ) λ= wavelength
Model Worksheet Student Handout
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
ion mobility spectrometry IR spectroscopy
 Debasmita Gho 29.10.2016 Introducti on Owing to its accuracy, sensitivity, and speed, mass spectrometry (MS) coupled to fragmentation techniques is the method of choice for determining the primary structure
Debasmita Gho 29.10.2016 Introducti on Owing to its accuracy, sensitivity, and speed, mass spectrometry (MS) coupled to fragmentation techniques is the method of choice for determining the primary structure
Presenter: She Zhang
 Presenter: She Zhang Introduction Dr. David Baker Introduction Why design proteins de novo? It is not clear how non-covalent interactions favor one specific native structure over many other non-native
Presenter: She Zhang Introduction Dr. David Baker Introduction Why design proteins de novo? It is not clear how non-covalent interactions favor one specific native structure over many other non-native
NH 2. Biochemistry I, Fall Term Sept 9, Lecture 5: Amino Acids & Peptides Assigned reading in Campbell: Chapter
 Biochemistry I, Fall Term Sept 9, 2005 Lecture 5: Amino Acids & Peptides Assigned reading in Campbell: Chapter 3.1-3.4. Key Terms: ptical Activity, Chirality Peptide bond Condensation reaction ydrolysis
Biochemistry I, Fall Term Sept 9, 2005 Lecture 5: Amino Acids & Peptides Assigned reading in Campbell: Chapter 3.1-3.4. Key Terms: ptical Activity, Chirality Peptide bond Condensation reaction ydrolysis
Code Course name CFU Year G6403B Info not available 3 1
 Basic aims The aim of the course is an in-depth discussion of the structureactivity relationships of the main classes of biological molecules. Strategies for synthesis, isolation and structural characterization
Basic aims The aim of the course is an in-depth discussion of the structureactivity relationships of the main classes of biological molecules. Strategies for synthesis, isolation and structural characterization
Supporting Information
 Supporting Information Figure S1. 2D ( 1 H- 1 H) COSY90 NMR (300 MHz, 3:1 TFA:TFA-d) spectrum of oligo(l-glu-co- 25%L-Cys) synthesized using from 7:3 L-Glu-(Et) 2 :L-Cys-Et, 0.5 M total substrate concentration,
Supporting Information Figure S1. 2D ( 1 H- 1 H) COSY90 NMR (300 MHz, 3:1 TFA:TFA-d) spectrum of oligo(l-glu-co- 25%L-Cys) synthesized using from 7:3 L-Glu-(Et) 2 :L-Cys-Et, 0.5 M total substrate concentration,
To be covered (and why) Spectroscopy of Proteins. UV-Vis Absorption. UV-Vis Absorption. Spectra
 To be covered (and why) Spectroscopy of Proteins General considerations UV-Vis Absorption quantitation Fluorescence hydrophobicity Foldedness FT-Infrared Foldedness ircular Dichroism Foldedness NMR (a
To be covered (and why) Spectroscopy of Proteins General considerations UV-Vis Absorption quantitation Fluorescence hydrophobicity Foldedness FT-Infrared Foldedness ircular Dichroism Foldedness NMR (a
Unit 1: Chemistry - Guided Notes
 Scientific Method Notes: Unit 1: Chemistry - Guided Notes 1 Common Elements in Biology: Atoms are made up of: 1. 2. 3. In order to be stable, an atom of an element needs a full valence shell of electrons.
Scientific Method Notes: Unit 1: Chemistry - Guided Notes 1 Common Elements in Biology: Atoms are made up of: 1. 2. 3. In order to be stable, an atom of an element needs a full valence shell of electrons.
The Chemistry of Life
 The Chemistry of Life Things you should be able to do 1. Describe how the unique properties of water support life on Earth. 2. Explain how carbon is uniquely suited to form biological macromolecules. 3.
The Chemistry of Life Things you should be able to do 1. Describe how the unique properties of water support life on Earth. 2. Explain how carbon is uniquely suited to form biological macromolecules. 3.
March CUME Organic Chemistry
 March CUME Organic Chemistry Department of Chemistry, University of Missouri Columbia Saturday, March 21, 1998, @Nine O Clock, 221 Chemistry Dr. Rainer Glaser Announced Reading Circular Dichroism and Exciton-Coupled
March CUME Organic Chemistry Department of Chemistry, University of Missouri Columbia Saturday, March 21, 1998, @Nine O Clock, 221 Chemistry Dr. Rainer Glaser Announced Reading Circular Dichroism and Exciton-Coupled
Supplementary Figures
 Supplementary Figures Supplementary Figure 1. The asymmetric unit in para-iodio-phenylalanine crystal. The 50% probability ellipsoid representation was prepared using the Mercury Software. Colors are as
Supplementary Figures Supplementary Figure 1. The asymmetric unit in para-iodio-phenylalanine crystal. The 50% probability ellipsoid representation was prepared using the Mercury Software. Colors are as
E The oscillating E-field defines the polarization of the wave. B
 This sheet is the lab document your TA will use to score your lab. It is to be turned in at the end of lab. To receive full credit you must use complete sentences and explain your reasoning. A. Describing
This sheet is the lab document your TA will use to score your lab. It is to be turned in at the end of lab. To receive full credit you must use complete sentences and explain your reasoning. A. Describing
Chem 524 Lecture Notes CD (Section 18) update 2011
 Chem 5 Lecture Notes CD (Section 8) update For HTML of 5 notes, click here XV. Circular Dichroism A. Differential absorption of left and right circular polarized light by molecular transition. Measure
Chem 5 Lecture Notes CD (Section 8) update For HTML of 5 notes, click here XV. Circular Dichroism A. Differential absorption of left and right circular polarized light by molecular transition. Measure
4.3A: Electronic transitions
 Ashley Robison My Preferences Site Tools Popular pages MindTouch User Guide FAQ Sign Out If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it
Ashley Robison My Preferences Site Tools Popular pages MindTouch User Guide FAQ Sign Out If you like us, please share us on social media. The latest UCD Hyperlibrary newsletter is now complete, check it
Chemistry in Biology. Section 1. Atoms, Elements, and Compounds
 Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Section 1 Atoms, Elements, and Compounds Atoms! Chemistry is the study of matter.! Atoms are the building blocks of matter.! Neutrons and protons are located at the center of the atom.! Protons are positively
Supplemental Material
 Supplemental Material Plasmonic Circular Dichroism of Peptide Functionalized Gold Nanoparticles Joseph M. Slocik, Alexander O. Govorov, and Rajesh R. Naik * Methods Nanoparticle functionalization. A peptide
Supplemental Material Plasmonic Circular Dichroism of Peptide Functionalized Gold Nanoparticles Joseph M. Slocik, Alexander O. Govorov, and Rajesh R. Naik * Methods Nanoparticle functionalization. A peptide
Exploration of Protein Folding
 Exploration of Protein Folding Question: What conditions affect the folding of a protein? Pre-lab reading Atkins & Jones (5 th ed.): Sections 5.1 5.5; 9.8 9.9; and Section 19.13 Safety and Waste Disposal
Exploration of Protein Folding Question: What conditions affect the folding of a protein? Pre-lab reading Atkins & Jones (5 th ed.): Sections 5.1 5.5; 9.8 9.9; and Section 19.13 Safety and Waste Disposal
It s really this simple.
 Background Light harvesting complexes exist to facilitate and maximize the absorption capacity of the reaction centers (RC) as well as PSI and PSII Purple bacteria utilize these functions by having an
Background Light harvesting complexes exist to facilitate and maximize the absorption capacity of the reaction centers (RC) as well as PSI and PSII Purple bacteria utilize these functions by having an
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Chapter 2 The Chemistry of Biology. Dr. Ramos BIO 370
 Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Chapter 2 The Chemistry of Biology Dr. Ramos BIO 370 2 Atoms, Bonds, and Molecules Matter - all materials that occupy space and have mass Matter is composed of atoms. Atom simplest form of matter not divisible
Introduction to Electromagnetic Radiation and Radiative Transfer
 Introduction to Electromagnetic Radiation and Radiative Transfer Temperature Dice Results Visible light, infrared (IR), ultraviolet (UV), X-rays, γ-rays, microwaves, and radio are all forms of electromagnetic
Introduction to Electromagnetic Radiation and Radiative Transfer Temperature Dice Results Visible light, infrared (IR), ultraviolet (UV), X-rays, γ-rays, microwaves, and radio are all forms of electromagnetic
Chapter 2: Chemistry. What does chemistry have to do with biology? Vocabulary BIO 105
 Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Experimental Methods in Kinetics (2014)
 IIIa- 35 Experimental Methods in Kinetics (2014) Initial examples assume start R = A 0 + B 0 etc., P = 0 follow reaction forward initial rate, etc. measure [A] vs. t Chemical methods: Could mix and take
IIIa- 35 Experimental Methods in Kinetics (2014) Initial examples assume start R = A 0 + B 0 etc., P = 0 follow reaction forward initial rate, etc. measure [A] vs. t Chemical methods: Could mix and take
Elements and Isotopes
 Section 2-1 Notes Atoms Life depends on chemistry. The basic unit of matter is the atom. Atoms are incredibly small The subatomic particles that make up atoms are protons, neutrons, and electrons. Parts
Section 2-1 Notes Atoms Life depends on chemistry. The basic unit of matter is the atom. Atoms are incredibly small The subatomic particles that make up atoms are protons, neutrons, and electrons. Parts
Protocol for Use of BCMP Jasco J-815 Circular Dichroism Spectropolarimeter
 Protocol for Use of BCMP Jasco J-815 Circular Dichroism Spectropolarimeter Getting Started 1. Turn on the nitrogen: Open the valve on the nitrogen tank (max 10 psi) so that the pressure gauge on the left
Protocol for Use of BCMP Jasco J-815 Circular Dichroism Spectropolarimeter Getting Started 1. Turn on the nitrogen: Open the valve on the nitrogen tank (max 10 psi) so that the pressure gauge on the left
A very brief history of the study of light
 1. Sir Isaac Newton 1672: A very brief history of the study of light Showed that the component colors of the visible portion of white light can be separated through a prism, which acts to bend the light
1. Sir Isaac Newton 1672: A very brief history of the study of light Showed that the component colors of the visible portion of white light can be separated through a prism, which acts to bend the light
Supporting Information
 Supporting Information Wiley-VCH 27 6941 Weinheim, Germany Homogeneous Biocatalysis in both Fluorous Biphasic and Supercritical Carbon Dioxide Systems H. R. Hobbs, H. M. Kirke, M. Poliakoff, N.R. T ho
Supporting Information Wiley-VCH 27 6941 Weinheim, Germany Homogeneous Biocatalysis in both Fluorous Biphasic and Supercritical Carbon Dioxide Systems H. R. Hobbs, H. M. Kirke, M. Poliakoff, N.R. T ho
Chapter 6 Chemistry in Biology
 Section 1: Atoms, Elements, and Compounds Section 2: Chemical Reactions Section 3: Water and Solutions Section 4: The Building Blocks of Life Click on a lesson name to select. 6.1 Atoms, Elements, and
Section 1: Atoms, Elements, and Compounds Section 2: Chemical Reactions Section 3: Water and Solutions Section 4: The Building Blocks of Life Click on a lesson name to select. 6.1 Atoms, Elements, and
Fluorescence 2009 update
 XV 74 Fluorescence 2009 update Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
XV 74 Fluorescence 2009 update Jablonski diagram Where does the energy go? Can be viewed like multistep kinetic pathway 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Course Syllabus. Department: Science & Technology. Date: April I. Course Prefix and Number: CHM 212. Course Name: Organic Chemistry II
 Department: Science & Technology Date: April 2012 I. Course Prefix and Number: CHM 212 Course Name: Organic Chemistry II Course Syllabus Credit Hours and Contact Hours: 5 credit hours and 7 (3:3:1) contact
Department: Science & Technology Date: April 2012 I. Course Prefix and Number: CHM 212 Course Name: Organic Chemistry II Course Syllabus Credit Hours and Contact Hours: 5 credit hours and 7 (3:3:1) contact
Ch 3: Chemistry of Life. Chemistry Water Macromolecules Enzymes
 Ch 3: Chemistry of Life Chemistry Water Macromolecules Enzymes Chemistry Atom = smallest unit of matter that cannot be broken down by chemical means Element = substances that have similar properties and
Ch 3: Chemistry of Life Chemistry Water Macromolecules Enzymes Chemistry Atom = smallest unit of matter that cannot be broken down by chemical means Element = substances that have similar properties and
CHEMISTRY. 2 Types of Properties Associated with Matter. Composition of Matter. Physical: properties that do not change the identity of the substance
 CHEMISTRY Composition of Matter Matter Mass Anything that occupies space and has mass Quantity of matter an object has Weight Pull of gravity on an object 2 Types of Properties Associated with Matter Physical:
CHEMISTRY Composition of Matter Matter Mass Anything that occupies space and has mass Quantity of matter an object has Weight Pull of gravity on an object 2 Types of Properties Associated with Matter Physical:
Foundations in Microbiology Seventh Edition
 Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 2 The Chemistry of Biology Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Lecture PowerPoint to accompany Foundations in Microbiology Seventh Edition Talaro Chapter 2 The Chemistry of Biology Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display.
Model Worksheet Teacher Key
 Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Introduction Despite the complexity of life on Earth, the most important large molecules found in all living things (biomolecules) can be classified into only four main categories: carbohydrates, lipids,
Experiment#1 Beer s Law: Absorption Spectroscopy of Cobalt(II)
 : Absorption Spectroscopy of Cobalt(II) OBJECTIVES In successfully completing this lab you will: prepare a stock solution using a volumetric flask; use a UV/Visible spectrometer to measure an absorption
: Absorption Spectroscopy of Cobalt(II) OBJECTIVES In successfully completing this lab you will: prepare a stock solution using a volumetric flask; use a UV/Visible spectrometer to measure an absorption
Analytical Technologies in Biotechnology Prof. Dr. Ashwani K Sharma Department of Biotechnology Indian Institute of Technology, Roorkee
 Analytical Technologies in Biotechnology Prof. Dr. Ashwani K Sharma Department of Biotechnology Indian Institute of Technology, Roorkee Module - 6 Spectroscopic Techniques Lecture - 2 UV-Visible Spectroscopy
Analytical Technologies in Biotechnology Prof. Dr. Ashwani K Sharma Department of Biotechnology Indian Institute of Technology, Roorkee Module - 6 Spectroscopic Techniques Lecture - 2 UV-Visible Spectroscopy
Chemistry Comes to Life
 BIOLOGY OF HUMANS Concepts, Applications, and Issues Fifth Edition Judith Goodenough Betty McGuire 2 Chemistry Comes to Life Lecture Presentation Anne Gasc Hawaii Pacific University and University of Hawaii
BIOLOGY OF HUMANS Concepts, Applications, and Issues Fifth Edition Judith Goodenough Betty McGuire 2 Chemistry Comes to Life Lecture Presentation Anne Gasc Hawaii Pacific University and University of Hawaii
Advanced Certificate in Principles in Protein Structure. You will be given a start time with your exam instructions
 BIRKBECK COLLEGE (University of London) Advanced Certificate in Principles in Protein Structure MSc Structural Molecular Biology Date: Thursday, 1st September 2011 Time: 3 hours You will be given a start
BIRKBECK COLLEGE (University of London) Advanced Certificate in Principles in Protein Structure MSc Structural Molecular Biology Date: Thursday, 1st September 2011 Time: 3 hours You will be given a start
Nanobiotechnology. Place: IOP 1 st Meeting Room Time: 9:30-12:00. Reference: Review Papers. Grade: 40% midterm, 60% final report (oral + written)
 Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 40% midterm, 60% final report (oral + written) Midterm: 5/18 Oral Presentation 1. 20 minutes each person
Nanobiotechnology Place: IOP 1 st Meeting Room Time: 9:30-12:00 Reference: Review Papers Grade: 40% midterm, 60% final report (oral + written) Midterm: 5/18 Oral Presentation 1. 20 minutes each person
Background BIOCHEMISTRY LAB CHE-554. Experiment #1 Spectrophotometry
 BIOCHEMISTRY LAB Experiment #1 Spectrophotometry CHE-554 In day 1 we will use spectrophotometry as an analytical technique using a known extinction coefficient to assess the precision and accuracy of common
BIOCHEMISTRY LAB Experiment #1 Spectrophotometry CHE-554 In day 1 we will use spectrophotometry as an analytical technique using a known extinction coefficient to assess the precision and accuracy of common
UV-Vis optical fiber assisted spectroscopy in thin films and solutions
 UV-Vis optical fiber assisted spectroscopy in thin films and solutions Description UV-Visible absorption and transmission spectra provide fundamental information for all experiments related to the attenuation
UV-Vis optical fiber assisted spectroscopy in thin films and solutions Description UV-Visible absorption and transmission spectra provide fundamental information for all experiments related to the attenuation
B DAYS BIOCHEMISTRY UNIT GUIDE Due 9/13/16 Monday Tuesday Wednesday Thursday Friday 8/22 - A 8/23 - B
 B DAYS BIOCHEMISTRY UNIT GUIDE Due 9/13/16 Monday Tuesday Wednesday Thursday Friday 8/22 - A 8/23 - B 8/24 - A 8/25 - B 8/26 - A 8/29 - B *Hydrolysis and dehydration synthesis *Macromolecule active reading
B DAYS BIOCHEMISTRY UNIT GUIDE Due 9/13/16 Monday Tuesday Wednesday Thursday Friday 8/22 - A 8/23 - B 8/24 - A 8/25 - B 8/26 - A 8/29 - B *Hydrolysis and dehydration synthesis *Macromolecule active reading
Denaturation and renaturation of proteins
 Denaturation and renaturation of proteins Higher levels of protein structure are formed without covalent bonds. Therefore, they are not as stable as peptide covalent bonds which make protein primary structure
Denaturation and renaturation of proteins Higher levels of protein structure are formed without covalent bonds. Therefore, they are not as stable as peptide covalent bonds which make protein primary structure
MODULE 2: BIOLOGICAL MOLECULES
 PEER-LED TEAM LEARNING INTRDUCTRY BILGY MDULE 2: BILGICAL MLECULES JSEP GRISWLD, + DEAN STETLER,* AND MICAEL GAINES, ( + City College of New York, *University of Kansas, Univ. of Miami;) I. Introduction
PEER-LED TEAM LEARNING INTRDUCTRY BILGY MDULE 2: BILGICAL MLECULES JSEP GRISWLD, + DEAN STETLER,* AND MICAEL GAINES, ( + City College of New York, *University of Kansas, Univ. of Miami;) I. Introduction
( 1 vρ) = ( sec)( N sec m 1 )
 Page 1 of 5 + formula sheet 1. You have isolated a novel protein that has been implicated in a disease and you would like to use what you have learned in Chem/Biochem 471 to deduce some of its properties.
Page 1 of 5 + formula sheet 1. You have isolated a novel protein that has been implicated in a disease and you would like to use what you have learned in Chem/Biochem 471 to deduce some of its properties.
Bio110 Lab 3: Basic Chemistry A. Carranza
 NAME Basic Chemistry The following chart lists the important elements found in cytoplasm by weight. On the chart, fill in the symbol and the number of electrons found in each element Use the periodic table
NAME Basic Chemistry The following chart lists the important elements found in cytoplasm by weight. On the chart, fill in the symbol and the number of electrons found in each element Use the periodic table
UV-Vis Absorption Experiment 5: Beer- Lambert Law and the Temperature Dependence of the Crystal Violet- Sodium Hydroxide Reaction
 1 UV-Vis Absorption Experiment 5: Beer- Lambert Law and the Temperature Dependence of the Crystal Violet- Sodium Hydroxide Reaction Overview In Part A of this experiment, the absorption behaviour of crystal
1 UV-Vis Absorption Experiment 5: Beer- Lambert Law and the Temperature Dependence of the Crystal Violet- Sodium Hydroxide Reaction Overview In Part A of this experiment, the absorption behaviour of crystal
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
Reflection = EM strikes a boundary between two media differing in η and bounces back
 Reflection = EM strikes a boundary between two media differing in η and bounces back Incident ray θ 1 θ 2 Reflected ray Medium 1 (air) η = 1.00 Medium 2 (glass) η = 1.50 Specular reflection = situation
Reflection = EM strikes a boundary between two media differing in η and bounces back Incident ray θ 1 θ 2 Reflected ray Medium 1 (air) η = 1.00 Medium 2 (glass) η = 1.50 Specular reflection = situation
Hole s Human Anatomy and Physiology Eleventh Edition. Chapter 2
 Hole s Human Anatomy and Physiology Eleventh Edition Shier Butler Lewis Chapter 2 1 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. CHAPTER 2 CHEMICAL BASIS OF
Hole s Human Anatomy and Physiology Eleventh Edition Shier Butler Lewis Chapter 2 1 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. CHAPTER 2 CHEMICAL BASIS OF
Raman Optical Activity Comes of Age
 Raman Optical Activity Comes of Age Laurence D. Barron Department of Chemistry, University of Glasgow, Glasgow G12 8QQ, UK E-mail: laurence.barron@glasgow.ac.uk Raman optical activity (ROA) provides vibrational
Raman Optical Activity Comes of Age Laurence D. Barron Department of Chemistry, University of Glasgow, Glasgow G12 8QQ, UK E-mail: laurence.barron@glasgow.ac.uk Raman optical activity (ROA) provides vibrational
2/25/2013. Electronic Configurations
 1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
1 2 3 4 5 Chapter 2 Chemical Principles The Structure of Atoms Chemistry is the study of interactions between atoms and molecules The atom is the smallest unit of matter that enters into chemical reactions
4 Proteins: Structure, Function, Folding W. H. Freeman and Company
 4 Proteins: Structure, Function, Folding 2013 W. H. Freeman and Company CHAPTER 4 Proteins: Structure, Function, Folding Learning goals: Structure and properties of the peptide bond Structural hierarchy
4 Proteins: Structure, Function, Folding 2013 W. H. Freeman and Company CHAPTER 4 Proteins: Structure, Function, Folding Learning goals: Structure and properties of the peptide bond Structural hierarchy
J- vs. H-type assembly: pentamethine cyanine (Cy5) as near IR chiroptical. reporter
 S1 Supporting Information: J- vs. H-type assembly: pentamethine cyanine (Cy5) as near IR chiroptical reporter Larysa I. Markova, a,b Vladimir L. Malinovskii, a Leonid D. Patsenker b and Robert Häner* a
S1 Supporting Information: J- vs. H-type assembly: pentamethine cyanine (Cy5) as near IR chiroptical reporter Larysa I. Markova, a,b Vladimir L. Malinovskii, a Leonid D. Patsenker b and Robert Häner* a
XV 74. Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go?
 XV 74 Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go? 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet S = 0 could be any of them
XV 74 Flouorescence-Polarization-Circular-Dichroism- Jablonski diagram Where does the energy go? 1) Excite system through A Absorbance S 0 S n Excite from ground excited singlet S = 0 could be any of them
Skoog Chapter 6 Introduction to Spectrometric Methods
 Skoog Chapter 6 Introduction to Spectrometric Methods General Properties of Electromagnetic Radiation (EM) Wave Properties of EM Quantum Mechanical Properties of EM Quantitative Aspects of Spectrochemical
Skoog Chapter 6 Introduction to Spectrometric Methods General Properties of Electromagnetic Radiation (EM) Wave Properties of EM Quantum Mechanical Properties of EM Quantitative Aspects of Spectrochemical
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
UNIT 3 CHEMISTRY. Fundamental Principles in Chemistry
 UNIT 3 CHEMISTRY NOTE: This list has been compiled based on the topics covered in the 2016 Master Class program. Once all of the 2017 Chemistry program materials have been finalised, this summary will
UNIT 3 CHEMISTRY NOTE: This list has been compiled based on the topics covered in the 2016 Master Class program. Once all of the 2017 Chemistry program materials have been finalised, this summary will
BIOCHEMISTRY Course Outline (Fall, 2011)
 BIOCHEMISTRY 402 - Course Outline (Fall, 2011) Number OVERVIEW OF LECTURE TOPICS: of Lectures INSTRUCTOR 1. Structural Components of Proteins G. Brayer (a) Amino Acids and the Polypeptide Chain Backbone...2
BIOCHEMISTRY 402 - Course Outline (Fall, 2011) Number OVERVIEW OF LECTURE TOPICS: of Lectures INSTRUCTOR 1. Structural Components of Proteins G. Brayer (a) Amino Acids and the Polypeptide Chain Backbone...2
 UV-Visible, Fluorescence and CD Spectroscopy Of Proteins Sepideh hkhorasanizadeh h Department of Biochemistry and Molecular Genetics Jordan Hall 6042 http://en.wikipedia.org/wiki/uv-vis_spectroscopy http://en.wikipedia.org/wiki/fluorescence_spectroscopy
UV-Visible, Fluorescence and CD Spectroscopy Of Proteins Sepideh hkhorasanizadeh h Department of Biochemistry and Molecular Genetics Jordan Hall 6042 http://en.wikipedia.org/wiki/uv-vis_spectroscopy http://en.wikipedia.org/wiki/fluorescence_spectroscopy
Lecture 5. More on UV-visible Spectrophotometry: Beer s Law and Measuring Protein Concentration
 Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 5 More on UV-visible Spectrophotometry: Beer s Law and Measuring Protein Concentration 23 January 2018 c David P. Goldenberg
Biological Chemistry Laboratory Biology 3515/Chemistry 3515 Spring 2018 Lecture 5 More on UV-visible Spectrophotometry: Beer s Law and Measuring Protein Concentration 23 January 2018 c David P. Goldenberg
Central Dogma. modifications genome transcriptome proteome
 entral Dogma DA ma protein post-translational modifications genome transcriptome proteome 83 ierarchy of Protein Structure 20 Amino Acids There are 20 n possible sequences for a protein of n residues!
entral Dogma DA ma protein post-translational modifications genome transcriptome proteome 83 ierarchy of Protein Structure 20 Amino Acids There are 20 n possible sequences for a protein of n residues!
Protein Structure. W. M. Grogan, Ph.D. OBJECTIVES
 Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Protein Structure W. M. Grogan, Ph.D. OBJECTIVES 1. Describe the structure and characteristic properties of typical proteins. 2. List and describe the four levels of structure found in proteins. 3. Relate
Kang, Lin-Woo, Ph.D. Professor Department of Biological Sciences Konkuk University Seoul, Korea nd Semester
 Kang, Lin-Woo, Ph.D. Professor Department of Biological Sciences Konkuk University Seoul, Korea 2018. 2 nd Semester Absorbance Assay (280 nm) Considerations for use Absorbance assays are fast and
Kang, Lin-Woo, Ph.D. Professor Department of Biological Sciences Konkuk University Seoul, Korea 2018. 2 nd Semester Absorbance Assay (280 nm) Considerations for use Absorbance assays are fast and
1 Introduction FOURIER TRANSFORM INFRARED STUDIES IN SOLID EGG WHITE LYSOZYME. 1C/94/403 INTERNAL REPORT (Limited Distribution)
 1C/94/403 INTERNAL REPORT (Limited Distribution) International Atomic Energy Agency and Nations Educational Scientific and Cultural Organization INTERNATIONAL CENTRE FOR THEORETICAL PHYSICS 1 Introduction
1C/94/403 INTERNAL REPORT (Limited Distribution) International Atomic Energy Agency and Nations Educational Scientific and Cultural Organization INTERNATIONAL CENTRE FOR THEORETICAL PHYSICS 1 Introduction
Optical Spectroscopy 1 1. Absorption spectroscopy (UV/vis)
 Optical Spectroscopy 1 1. Absorption spectroscopy (UV/vis) 2 2. Circular dichroism (optical activity) CD / ORD 3 3. Fluorescence spectroscopy and energy transfer Electromagnetic Spectrum Electronic Molecular
Optical Spectroscopy 1 1. Absorption spectroscopy (UV/vis) 2 2. Circular dichroism (optical activity) CD / ORD 3 3. Fluorescence spectroscopy and energy transfer Electromagnetic Spectrum Electronic Molecular
Figure ) Letter E represents a nucleic acid building block known as a. Answer: nucleotide Diff: 3 Page Ref: 54
 Essentials of Human Anatomy and Physiology, 10e (Marieb) Chapter 2 Basic Chemistry 2.1 Short Answer Figure 2.1 Using Figure 2.1, identify the following: 1) Which letter represents a carbohydrate polymer?
Essentials of Human Anatomy and Physiology, 10e (Marieb) Chapter 2 Basic Chemistry 2.1 Short Answer Figure 2.1 Using Figure 2.1, identify the following: 1) Which letter represents a carbohydrate polymer?
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Protein Structure Prediction and Display
 Protein Structure Prediction and Display Goal Take primary structure (sequence) and, using rules derived from known structures, predict the secondary structure that is most likely to be adopted by each
Protein Structure Prediction and Display Goal Take primary structure (sequence) and, using rules derived from known structures, predict the secondary structure that is most likely to be adopted by each
Photochemical principles
 Chapter 1 Photochemical principles Dr. Suzan A. Khayyat 1 Photochemistry Photochemistry is concerned with the absorption, excitation and emission of photons by atoms, atomic ions, molecules, molecular
Chapter 1 Photochemical principles Dr. Suzan A. Khayyat 1 Photochemistry Photochemistry is concerned with the absorption, excitation and emission of photons by atoms, atomic ions, molecules, molecular
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Paper: 12, Organic Spectroscopy Module: 5, Applications of UV spectroscopy
 Subject Chemistry Paper No and Title Module No and Title Module Tag Paper 12: Organic Spectroscopy Applications of UV-visible Spectroscopy CHE_P12_M5 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction
Subject Chemistry Paper No and Title Module No and Title Module Tag Paper 12: Organic Spectroscopy Applications of UV-visible Spectroscopy CHE_P12_M5 TABLE OF CONTENTS 1. Learning Outcomes 2. Introduction
Electromagnetic wave energy & polarization
 Phys 0 Lecture 6 Electromagnetic wave energy & polarization Today we will... Learn about properties p of electromagnetic waves Energy density & intensity Polarization linear, circular, unpolarized Apply
Phys 0 Lecture 6 Electromagnetic wave energy & polarization Today we will... Learn about properties p of electromagnetic waves Energy density & intensity Polarization linear, circular, unpolarized Apply
Circular Dichroism Spectroscopy of Membrane Proteins: A tutorial review
 Circular Dichroism Spectroscopy of Membrane Proteins: A tutorial review Journal: Manuscript ID CS-SYN-01-2015-000084.R3 Article Type: Tutorial Review Date Submitted by the Author: 16-Jun-2016 Complete
Circular Dichroism Spectroscopy of Membrane Proteins: A tutorial review Journal: Manuscript ID CS-SYN-01-2015-000084.R3 Article Type: Tutorial Review Date Submitted by the Author: 16-Jun-2016 Complete
DNA Condensation Using Self-Assembled Peptides
 DNA Condensation Using Self-Assembled Peptides Angela m. Jimenez, Class of 2008, Major: Chemical Engineering Mentor: Raymond S. Tu, Department of Chemical Engineering ABSTRACT Counterion-mediated DNA-condensation
DNA Condensation Using Self-Assembled Peptides Angela m. Jimenez, Class of 2008, Major: Chemical Engineering Mentor: Raymond S. Tu, Department of Chemical Engineering ABSTRACT Counterion-mediated DNA-condensation
BIOCHEMISTRY NOTES - UNIT 2-
 BIOCHEMISTRY NOTES - UNIT 2- ATOMS - the basic unit of matter. Contains subatomic particles o (+ charge) o (no charge/neutral) o (- charge) Protons and neutrons have about the same mass. Electrons are
BIOCHEMISTRY NOTES - UNIT 2- ATOMS - the basic unit of matter. Contains subatomic particles o (+ charge) o (no charge/neutral) o (- charge) Protons and neutrons have about the same mass. Electrons are
THz spectroscopy in biology
 Preparation of the concerned sectors for educational and R&D activities related to the Hungarian ELI project THz spectroscopy in biology Andrea Buzády 6. lecture Spectroscopy TÁMOP-4.1.1.C-12/1/KONV-2012-0005
Preparation of the concerned sectors for educational and R&D activities related to the Hungarian ELI project THz spectroscopy in biology Andrea Buzády 6. lecture Spectroscopy TÁMOP-4.1.1.C-12/1/KONV-2012-0005
Optical Activity as a Biosignature in the Search for Extraterrestrial Life
 ptical Activity as a Biosignature in the Search for Extraterrestrial Life Andrew K. Boal AST740 21 April 2006 utline of this talk Science ow will we detect life on other planets? Some proposed biosignatures
ptical Activity as a Biosignature in the Search for Extraterrestrial Life Andrew K. Boal AST740 21 April 2006 utline of this talk Science ow will we detect life on other planets? Some proposed biosignatures
A Single Outer Sphere Mutation Stabilizes apo- Mn Superoxide Dismutase by 35 C and. Disfavors Mn Binding.
 Supporting information for A Single Outer Sphere Mutation Stabilizes apo- Mn Superoxide Dismutase by 35 C and Disfavors Mn Binding. Anne-Frances Miller* and Ting Wang Department of Chemistry, University
Supporting information for A Single Outer Sphere Mutation Stabilizes apo- Mn Superoxide Dismutase by 35 C and Disfavors Mn Binding. Anne-Frances Miller* and Ting Wang Department of Chemistry, University
Optical spectroscopies. Rita P.Y. Chen. Electromagnetic Spectrum
 Optical spectroscopies Rita P.Y. Chen Electromagnetic Spectrum The longer the wavelength the lower the energy!!!! h = 6.63 x 10-34 J s 1 ev = 1.6 x 10-19 J, c= 3 x 10 8 m Transmittance, T = I f / I
Optical spectroscopies Rita P.Y. Chen Electromagnetic Spectrum The longer the wavelength the lower the energy!!!! h = 6.63 x 10-34 J s 1 ev = 1.6 x 10-19 J, c= 3 x 10 8 m Transmittance, T = I f / I
2015 AP Biology Unit 2 PRETEST- Introduction to the Cell and Biochemistry
 Name: Class: _ Date: _ 2015 AP Biology Unit 2 PRETEST- Introduction to the Cell and Biochemistry Multiple Choice Identify the choice that best completes the statement or answers the question. 1) In what
Name: Class: _ Date: _ 2015 AP Biology Unit 2 PRETEST- Introduction to the Cell and Biochemistry Multiple Choice Identify the choice that best completes the statement or answers the question. 1) In what
UNIT 1: BIOCHEMISTRY
 UNIT 1: BIOCHEMISTRY UNIT 1: Biochemistry Chapter 6.1: Chemistry of Life I. Atoms, Ions, and Molecules A. Living things consist of atoms of different elements 1. An atom is the smallest basic unit of matter
UNIT 1: BIOCHEMISTRY UNIT 1: Biochemistry Chapter 6.1: Chemistry of Life I. Atoms, Ions, and Molecules A. Living things consist of atoms of different elements 1. An atom is the smallest basic unit of matter
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Lecture 4: Polarisation of light, introduction
 Lecture 4: Polarisation of light, introduction Lecture aims to explain: 1. Light as a transverse electro-magnetic wave 2. Importance of polarisation of light 3. Linearly polarised light 4. Natural light
Lecture 4: Polarisation of light, introduction Lecture aims to explain: 1. Light as a transverse electro-magnetic wave 2. Importance of polarisation of light 3. Linearly polarised light 4. Natural light
Chapter 02 Chemistry of Life
 Chapter 02 Chemistry of Life Multiple Choice Questions 1. The smallest unit of matter is the A. molecule. B. atom. C. compound. D. isotope. HAPS Objective: C.01.03 Compare and contrast the terms atoms,
Chapter 02 Chemistry of Life Multiple Choice Questions 1. The smallest unit of matter is the A. molecule. B. atom. C. compound. D. isotope. HAPS Objective: C.01.03 Compare and contrast the terms atoms,
BIOCHEMISTRY Unit 2 Part 4 ACTIVITY #6 (Chapter 5) PROTEINS
 BIOLOGY BIOCHEMISTRY Unit 2 Part 4 ACTIVITY #6 (Chapter 5) NAME NAME PERIOD PROTEINS GENERAL CHARACTERISTICS AND IMPORTANCES: Polymers of amino acids Each has unique 3-D shape Vary in sequence of amino
BIOLOGY BIOCHEMISTRY Unit 2 Part 4 ACTIVITY #6 (Chapter 5) NAME NAME PERIOD PROTEINS GENERAL CHARACTERISTICS AND IMPORTANCES: Polymers of amino acids Each has unique 3-D shape Vary in sequence of amino
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Effect of the Single and Double Chain Surfactant Cobalt(III) Complexes on Their Hydrophobicity, Micelle Formation,
 Electronic Supplementary Material (ESI) for Inorganic Chemistry Frontiers. This journal is the Partner Organisations 2014 Supplementary Information Effect of the Single and Double Chain Surfactant Cobalt(III)
Electronic Supplementary Material (ESI) for Inorganic Chemistry Frontiers. This journal is the Partner Organisations 2014 Supplementary Information Effect of the Single and Double Chain Surfactant Cobalt(III)
