6.#Molecular#Mechanics#and# Molecular#Dynamics#Simula5on# MOLECULAR)MECHANICS) Determinant#of#Structure#(or#Lack#of#It)# Water#
|
|
|
- Cleopatra Green
- 6 years ago
- Views:
Transcription
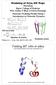
1 BIOCH 755: Biochemistry I Fall #Molecular#Mechanics#and# Molecular#Dynamics#Simula5on# Jianhan#Chen# Office#Hour:#M#1:30?2:30PM,#Chalmers#034# #jianhanc@ksu.edu# Office:#785?2518# Determinant#of#Structure#(or#Lack#of#It)# Probability#of#observing#a#par5cular#structure#(conforma5on)# is#determined#by#its#stability#(as#defined#by#the#free#energy)# Thermodynamics#and#sta5s5cal#mechanics!# No#single#structure#is#the$structure# It#is#all#about#probability#(sta5s5cal#mechanics!)# Mo5ons#and#flexibility#are#important#too# The#stability#depends#on#a#range#of#factors# Intramolecular#interac5ons# Bonded:#chemical#bonds,#angles,#dihedrals#etc# Nonbonded:# weak #interac5ons# Charged?charged,#van#der#Waals#(dispersion#and#repulsion)# Intermolecular#interac5ons:#nonbonded/weak#interac5ons# Cellular#environment:#solvent#(water),#membrane,#salt,#pH#etc# Associa5on#with#other#biomolecules,#small#molecules,#ions,#etc# (c)#jianhan#chen# 2# Water# Solvent#of#life# Many#unique#proper5es# Maximum#density#at#4# o C# Ice#is#lighter#than#liquid#water# Polar#molecule# hydrogen#bonding#network# High#specific#heat#capacity# Hydrophobicity#and#hydrophilicity# Solute#polarity#(carry#par5al#charges#or#not)# Salts#(e.g.#NaCl)#dissolve#in#water#readily# Hydrocarbons#(oil)#do#not#mix#with#water# Amphipathic#molecules# Self?assembly#to#micelles,#biological#membranes# MOLECULAR)MECHANICS) (c)#jianhan#chen# 3# (c)#jianhan#chen# 4#
2 Quantum#Mechanics#vs.#Molecular#Mechanics# Quantum#mechanics:# exact #and#most#applicable#to# understand#chemical#reac5ons# Separate#nuclei#and#electrons# Too#expensive,#and#not#sufficiently#accurate# Not#relevant#as#many#biological#processes# Molecular#mechanics:#classical#mechanics#at#molecular#level# Classical#treatment#of#all#atoms# No#electron,#no#chemistry# Allows#descrip5on#of#large#molecules# Experimental#methods#available#to#determine#the#key#parameters#in#a# molecular#mechanical#treatment# Hybrid#QM/MM# QM#for#the#ac5ve#site#(where#reac5on#occurs)#and#MM#for#the#rest# Accurate#treatment#of#MM/QM#Boundary#is#a#problem# (c)#jianhan#chen# 5# Classical#Mechanics# Total#energy:#E$=$K$+$V$ Kine5c#energy#(K#=#mv 2 /2),#poten5al#energy#V#(i.e.,#force#field)$ Newton s#second#law#of#mo5on:#f$=$m$a# Rela5on#of#force#and#poten5al#energy:#F$=#?$δV/δr$ (c)#jianhan#chen# 6# Molecular#Poten5als# Bonds#and#Angles# Basic#form:#V#=#V bonding #+#V nonbonding# # # # # ######=#(#ΣV bond #+#ΣV angle #+#ΣV dihe )#+#Σ(V elec #+#V vdw )# # # The#poten5al#energy#is#a#func5on#of#all#coordinates.# # Addi5vity,#empirical,#transferability# # V bond #=#k bond #(#r# #r o ) 2# Harmonic#approxima5on# OK#for#biomolecules # # V angle #=#k angle #(#θ# #θ o ) 2# (c)#jianhan#chen# 7# (c)#jianhan#chen# 8#
3 V dihe #=#k dihe $1$[1#+#cos(n $ φ $ 3$δ)]## Dihedral#Poten5als# Realis5c#Dihedral#Poten5als# Actual#dihedral#poten5als#omen#have#contribu5ons#with#mul5ple#periodici5es# V= 3 ( 1 + cos ( 3 φ ) ) trans (c)#jianhan#chen# 9# (c)#jianhan#chen# 10# Electrosta5c#Interac5ons# V elec #=#q 1 q 2 /4πε 0 r####coulomb s#law# ε 0 : # permipvity#constant#of#vacuum# A#simplified#form:#V elec# =#332#q 1 q 2 /r# Where#q#is#unit#of#electron#charge,##r#is#in#Å,#and#V#in#kcal/mol.# Dielectric#medium:##V elec# =#332#q 1 q 2 /εr# # ε#is#dielectric#constant#(rela5ve#permipvity).## ε=78#for#water#under#lab#condi5ons#(300k,#1atm)# V or δv/δr (kcazl/mol) 1/r 2 1/r r q 1 q 2 van#der#waals#interac5ons# London#dispersion:#arrac5ve#forces#that#arise#from# temporary#dipoles#(induced#dipole?induced#dipole# interac5ons)# van#der#waals#repulsion:#all#atoms#repel#at#short#distances# A#common#func5on#form:#V vdw #=# A/r 6 #+#B/r 12# Lennard?Jones#poten5al#func5on#(12?6)# # r (Å) 11# (c)#jianhan#chen# 12#
4 Lennard?Jones#Poten5al# ε = 1, r min = 1 Sub?Summary# Molecular)mechanics)as)an)effective)approach)for) calculating)the)energies)of)large)biomolecules) Common)functional)form)with)five)terms) Covalent)bonded)terms) Non;bonded)terms:)electrostatics,)vdW) V (kcazl/mol) Coming)up:)application)of)molecular)mechanics)in) biomolecular)modeling)and)simulation)(i.e.,)molecular) dynamics)) r (Å) (c)#jianhan#chen# 13# 14 Why#Modeling?# Visualiza5on# Interpreta5on#of#experimental#data# X?Ray#Crystallography,#NMR#Structures,#EM#models# Low#resolu5on#experimental#data:#mutagenesis,#FRET,#and#many#more# Novel#insights#not#accessible#to#experiments# Experiment Theory MOLECULAR)DYNAMICS) X-ray, NMR, Cyro-EM (structure & dynamics) Scattering, Calorimetry, pka, physical measurements Protein models (QM, MM, Coarse-grained ) Simulation techniques (MD, MC, REX ) (c)#jianhan#chen# 15# To#ra5onalize#and#predict#experimental#observa5ons.# To#provide#new#insights,#hypothesis,#and#direc5ons.## (c)#jianhan#chen# 16#
5 Molecular#Poten5als# Basic#form:#V#=#V bonding #+#V nonbonding# # # # # ######=#(#ΣV bond #+#ΣV angle #+#ΣV dihe )#+#Σ(V elec #+#V vdw )# # # The#poten5al#energy#is#a#func5on#of#all#coordinates.# # Addi5vity,#empirical,#transferability# # 1 2 bonds,i V MM = Σ k b 0 i (b i - b i ) angles,i Molecular#Mechanics# Classical Energy Functions 0 + Σ k θ i (θ i - θ i ) 2 + Σ k φ torsions,i i [1 + cos(n i φ i - δ i )] ij + Σ Σ ε min atoms,i i < j ij r min r ij 12 ij 6 r min r ij q i q j εr ij θ φ δ i Force Field CHARMM Amber OPLS δ j Molecular Dynamics (MD).. m i r i = F i = - V i Monte Carlo (MC) P(δr) = exp(-δv/kt) (c)#jianhan#chen# 17# 18# CHARMM#param22#Force#Field# Topology#file:#define#the#building#blocks#(atoms,#connec5vi5es)## atom types residue blocks atom compositions connectivity MASS 1 H ! polar H MASS 2 HC ! N-ter H MASS 3 HA ! nonpolar H name type charge RESI ALA 0.00 GROUP ATOM N NH1-0.47! ATOM HN H 0.31! HN-N ATOM CA CT1 0.07! HB1 ATOM HA HB 0.09! / GROUP! HA-CA--CB-HB2 ATOM CB CT3-0.27! \ ATOM HB1 HA 0.09! HB3 ATOM HB2 HA 0.09! O=C ATOM HB3 HA 0.09! GROUP! ATOM C C 0.51 ATOM O O BOND CB CA N HN N CA O C BOND C CA C +N CA HA CB HB1 CB HB2 CB HB3 IMPR N -C CA HN C CA +N O DONOR HN N excerpted from: top_all22_prot.inp (c)#jianhan#chen# 19# CHARMM#param22#Force#Field# Parameter#file:#define#the#parameters#of#interac5ons## BONDS C C ! ALLOW ARO HEM! Heme vinyl substituent (KK, from propene (JCS)) CA CA ! ALLOW ARO! benzene, JES 8/25/89 ANGLES CA CA CA ! ALLOW ARO! JES 8/25/89 CE1 CE1 CT !! for 2-butene, yin/adm jr., 12/95 DIHEDRALS C CT1 NH1 C ! ALLOW PEP! ala dipeptide update for new C VDW Rmin, adm jr., 3/3/93c C CT2 NH1 C ! ALLOW PEP! ala dipeptide update for new C VDW Rmin, adm jr., 3/3/93c NONBONDED nbxmod 5 atom cdiel shift vatom vdistance vswitch - cutnb 13.0 ctofnb 12.0 ctonnb 10.0 eps 1.0 e14fac 1.0 wmin 1.5!adm jr., 5/08/91, suggested cutoff scheme C ! ALLOW PEP POL ARO! NMA pure solvent, adm jr., 3/3/93 CA ! ALLOW ARO! benzene (JES) excerpted from: par_all22_prot.inp (c)#jianhan#chen# 20#
6 Parameteriza5on#of#Force#Fields# Bonded#terms:#spectroscopy#or#quantum#mechanics# Lennard?Jones:#Small#molecular#crystals# Electrosta5c:#quantum#mechanics#(fit#monopoles#to# electrosta5c#poten5al)# Many#challenges#in#prac5ce# which#(model)#molecules:#availability,#representa5ve#or#not### how#many#atomic#classes:#transferability#and#tractability# Which#proper5es#to#parameterize#for?# Correla5on#of#parameters# Higher#order/new#terms#or#not?# Electron#polariza5on,#non?addi5vity,### water,#water,#and#water# At#the#end,#do#they#add#up?#(cancella5on#of#errors)# (c)#jianhan#chen# 21# Energy#Minimiza5on# Minimiza5on#follows#gradient#of#poten5al#to#iden5fy#stable# points#on#energy#surface## Let#V(x)#=#k#(x?x 0 ) 2 ## Begin#at#x,#how#do#we#find#x 0 #if#we#don t#know#v(x)#in#detail?## #How#can#we#move#from#x #to#x 0?## #Steepest#descent#(SD):## x! x #=#x+δ### δ#=#?dx# V(x)/# x#=#?dx#k(x?#x 0 )## This#moves#us,#depending#on## #####the#step#size#dx,#toward#x 0.## On#a#simple#harmonic#surface,#we## #####will#reach#the#minimum,#x 0,#i.e.## #####converge,#in#a#certain#number#of## #####steps#related#to#dx.## x x 0 (c)#jianhan#chen# 22# Molecular#Dynamics# Objec5ve:#{r 1 (t),#,#r N (t)}#!#{r 1 (t+δt),#,#r N (t#+δt)}# f#=#ma# Basic#idea:#solve#Newton s#equa5on#of#mo5on#numerically# Given#current#coordinates#(x),#veloci5es#(v)# Forces#can#be#calculated#based#on#coordinates#(from#f#=#? V/ x)# x(t+δt)#=#x(t)#+#v(t)#δt# v(t+δt)#=#v(t)#+#f(t)/m#δt# Repeat#above#opera5ons# More#accurate#integrators#(berer#energy#conserva5on)# Verlet#Algorithm#(Verlet#J.#Chem.#Phys.#1967)# ###consider#taylor s#expansions:# x(t±δt)#=#x(t)#±#v(t) Δt#+#1/2m#f(t)#Δt 2# ±#1/6#d 3 x/dt 3 #Δt 3 +O(Δt 4 )## ###Adding#expansion#x(t+Δt)#and#x(t?Δt)#and#rearrange:### x(t+δt)#=#2x(t)#?#x(t?δt)#+#f(t)/m#δt 2 +O(Δt 4 )## ###Subtrac5ng#expansion#x(t+Δt)#and#x(t?Δt)#and#rearrange:# #########v(t)#=#[x(t+δt)?x(t?δt)/(2δt)+o(δt 3 )### velocities lag (c)#jianhan#chen# 23# Solvent#and#Periodic#Boundary#Condi5ons# van Gunsteren Angew Chem Int Ed (2006) (c)#jianhan#chen# 24#
7 Controlling#Thermodynamic#Variables# Basic#Flow#of#a#MD#Simula5on# MD#generate#sta5s5cal#ensembles#that#connect#microscopic# details#to#macroscopic/thermodynamic#proper5es# NVE#(microcanonical#?#Entropy#rules!)## Read#in#the#force# field#(topology#and# parameter#files)# Read#in#the#protein# sequence#and# generate#protein# structure#file#(psf)# Read#in#the#ini5al# coordinates# NVT#(Canonical#?#Helmholtz#free#energy#is#relevant,#A)## temperature#t#=# m<v2>/(3k B )## NPT#(Isothermal?isobaric#?#Gibbs#free#energy#is#relevant,#G)# #P=kine5c#+#virial#contribu5ons## Structural#and# trajectory#analysis# Nonbonded#op5ons,# PBC,#restraint# poten5als,#etc# Op5onal#Energy# minimiza5on# Op5onal# Equilibra5on#MD# Thermostats,#barostats,#etc.,#allow#one#to#choose#appropriate# ensembles## Following#Nose,#Hoover,#Evans#and#others# See#Brooks,#Curr.#Opin.#Struct.#Biol.,#5,#211(1995)]# PSF: a computer model or representation of the protein given the force field, including all atoms, their connectivities and how they interact, etc. Produc5on#MD:#type#of#integrator,# thermodynamic#variables#(t,#p,#etc),# simula5on#length,#output#op5ons#(file# names,#write#frequencies#etc),## (c)#jianhan#chen# 25# (c)#jianhan#chen# 26# Biomolecular#Simula5ons#are#computa5onally# very#expensive# Channel-forming peptides in a fully solvated membrane bilayer; Channel: 1795 atoms; All: atoms Simulated Time 1 ns (10-9 s) (500,000 MD steps) CPU Time ~200 hours (10 6 s) Wall Time ~1 days (10 5 s) / 8 CPUs very#small#5me#step#required## δt$~#fs#(10?15 #s)# interac5ons#between# thousands#of#atoms#need#to# be#computed# (c)#jianhan#chen# 27# Biological#Time#Scale# Bond#vibra5ons# # #1#fs#(10?15# s)# Sugar#repuckering## #1#ps#(10?12# s)## DNA#bending# # # #1#ns#(10?9 #s)## Domain#movement# #1#ms#(10?6 #s)## Base#pair#opening# # #1#ms#(10?3 #s)## Transcrip5on# # # #2.5#ms#/#nucleo5de## Protein#synthesis# # #6.5#ms#/#amino#acid## Protein#folding## #~#10#s#(speed#limit:#µs)## RNA#life5me# # # #~#300#s# Simulation time should exceed the time scale of interest by ~10-fold! (c)#jianhan#chen# 28#
8 Gap#in#Timescales# Prac5cal#Considera5ons# Simulation Timescales Peptides (~10 residues) Small Proteins (<50 residues) Large Proteins (>200 residues) s ms Biological Timescales 10 0 s 10-3 s µs 10-6 s ns ps fs 10-9 s s s Protein Folding, Conformational Transitions Internal Dynamics Bond Vibrations Long?range#forces# Using#cut?off#to#reduce#the#number#of# nonbonded#atom#pairs#(~#12?15å)# Electrosta5c#decays#slowing#(1/r)#and#cut# off#does#not#work#well;#par5cle#mesh# Eward#(PME)#is#needed.# Parallel#execu5on## Par55on#various#regions#of#the#system#to# different#cpus# Need#to#communicate#informa5on# between#nodes;#this#is#a#borleneck# Simplifica5ons#of#the#model# Enhanced#sampling#techniques# # van Gunsteren Angew Chem Int Ed (2006) Shaw JCC (2005) 29# (c)#jianhan#chen# 30# Anton#Computer# Specialized#computer#for#molecular#dynamics#simula5ons# Highly#parallel#with#extensive#specialized#hardware#for#MD# related#computa5ons# Rumored#to#cost#>$100M#to#design#and#build# 17#μs/day#for#small#proteins#(~25K#atoms)## ~5#μs/day#for#large#proteins#(500#aa,#>132K#atoms)# A#512?node#Anton#donated#by#DESRES#is#available#for#public# use#at#pirsburg#supercomputer#center# Implicit#Solvent# Solvent#increases#the#system#size#about#10?fold# It#is#possible#to#describe#the#mean#influence#of#water#w/o# explicitly#including#water## Explicit solvent Protein: 56 residues (855 atoms) Solvent: 5411 waters (16233 atoms) Implicit solvent Hybrid macroscopic (solvent) / microscopic (solute) (c)#jianhan#chen# 32#
9 Coarse?Grained#Models# Rely#on#reduced#representa5on#and/or#simplified#interac5on# schemes#to#access#larger#length#and#5me#scales# Klein and Shinoda, Science (2008) Biomembrane sculpting by protein-bar domains The simulation shown in the figure was carried out using a box with dimensions 100 x 16 x 50 nm and would correspond to a system of 10 million atoms. Using a shape-based CG model reduces the size to 3265 CG beads. The simulation showed that a concerted action of BAR domains arranged in a lattice results in the development of a global membrane curvature on a time scale of several µs, with the resulting curvature radius of ~30 nm that was observed experimentally. (c)#jianhan#chen# 33# Energy Surface Barriers,#Temperature#and#Timescales# * ΔG Reaction Coordinate τ = τ 0 exp(δg /kt) τ 0 ~ s ~ ps T = 300 K ΔG : 1 kcal/mol, τ ~ ps 5 kcal/mol, τ ~ ns 10 kcal/mol, τ ~ µs Protein energy landscape is highly complex and rugged with numerous local minima. 34# Enhanced#Sampling#Techniques# Applica5ons#of#Modeling# 300 K 355 K 420 K 500 K P i j = REP1 REP2 REP3 REP4 MD/MC Exchange criteria?? Δ = ( E i - E j ) ( 1/kT j - 1/kT i ) Replica Exchange (REX) 1 Δ 0 exp(-δ) Δ > 0 REP2 REP1 REP3 REP4 MD/MC? REP2 REP3 REP1 REP4 MD/MC Protein Energy Surface Sugita and Okamoto, CPL (1999); MMTSB Tool Set: * 35# Main#advantages# Offer#atomis5c#spa5al#resolu5on#and#femtosecond#5me#resolu5on# Allow#probing#the#system#in#many#nontrivial#ways#that#are#not# possible#or#too#dangerous#experimentally# Omen#much#cheaper#than#doing#the#experiment#itself# Can#be#applied#at#very#large#scales#(computers#are#cheap)# Can#provide#theore5cal#frameworks#for#experimental#studies# A#few#prototypical#areas# Protein#structure#predic5on#and#calcula5on# Virtual#screening#and#ra5onal#drug#design# Simula5on#of#important#systems:#mechanisms# Interpreta5on#of#(sta5c)#experimental#data# Protein#misfolding#and#aggrega5on# Biomolecular#engineering:#design#of#new#enzymes#etc# # (c)#jianhan#chen# 36#
10 NMR#Structure#Refinement# Protein#Folding#Problem# REX/GB Refinement PDB: 1XJH Refined Model Initial Model from CNS 1 day later 1 month later Chen et. al., JACS (2004); Chen et al., J. Biomol NMR (2004). (c)#jianhan#chen# 37# Inconclusive#Experiments# 500 Å (c)#jianhan#chen# 38# Homology#Modeling# Extreme simplification Limited force field accuracy Large gaps in timescales see also, Bionanotechnology D.S. Goodsell 2004 Wiley 39# (c)#jianhan#chen# 40#
11 Basic#Principles# Proteins#with#similar# sequences#should# have#similar# structure.## One#can#model#a# sequence#of# unknown#structure# (target)#based#on#a# homolog#of#known# structure#(template).# Structure#?>#func5on# Sali, A. & Kuriyan, J. Trends Biochem. Sci. 22, M20 M24 (1999) (c)#jianhan#chen# 41# (c)#jianhan#chen# 42# PDB#is# complete #(i.e.,#novel#fold#is#rare)# Homology#Modeling#Basic#Flow#Chart# " Backbone# genera5on# " Loop#modeling# " Side#chains# " Refinement# " Valida5on##) (c)#jianhan#chen# 43# (c)#jianhan#chen# 44#
12 Structure#Valida5on# Covalent#geometry:#typically#OK# Ramanchandron#plot:#usually#OK# Inside/outside#distribu5ons#of#polar#and#apolar#residues#can# be#useful.# Biological/biochemical#data# Ac5ve#site#residues# Modifica5on#sites# Interac5on#sites# Valida5on#servers/tools:## ProQ# WhatIF# Procheck# Popular#Structure#Predic5on#Servers# Modern#predic5on#tools/servers#employ#sophis5cated# integra5on#of#homology#modeling,#de#novo#modeling,# structure#refinement#and#many#other#empirical# tricks #to#get# the#most#out#of#exis5ng#sta5s5cal#and#physical#knowledge# Rosera/Robera#(David#Baker)# I?TASSER#(Yang#Zhang)# MULTICOM#(Jianlin#Cheng)# (c)#jianhan#chen# 45# (c)#jianhan#chen# 46#
!"#$%&#'()*+,$('%-"+.)'+/)01"2"#$%&#'() 3,(&%,&($.4))5"26&,$()!"/$#1+7)
 BIOCH 590: Biomacromolecules Spring 2010 81$('(%91%'#):(7'+1;'-"+)"
BIOCH 590: Biomacromolecules Spring 2010 81$('(%91%'#):(7'+1;'-"+)"
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Semi Empirical Force Fields and Their Limitations. Potential Energy Surface (PES)
 Semi Empirical Force Fields and Their Limitations Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Potential Energy Surface (PES) Schrödinger equation: H T Ψ( r, = E Ψ( r, H =
Semi Empirical Force Fields and Their Limitations Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Potential Energy Surface (PES) Schrödinger equation: H T Ψ( r, = E Ψ( r, H =
Understanding the determinants of selectivity in drug metabolism through modeling of dextromethorphan oxidation by cytochrome P450
 Understanding the determinants of selectivity in drug metabolism through modeling of dextromethorphan oxidation by cytochrome P450 Julianna Oláh, 1 Adrian J. Mulholland* and Jeremy N. Harvey* School of
Understanding the determinants of selectivity in drug metabolism through modeling of dextromethorphan oxidation by cytochrome P450 Julianna Oláh, 1 Adrian J. Mulholland* and Jeremy N. Harvey* School of
Potential Energy (hyper)surface
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
Journal Name ARTICLE. ESI for: CHARMM force field parameterization protocol for selfassembling peptide amphiphiles: The Fmoc moiety
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016Please do not adjust margins ESI for: Received 00th January 20xx, Accepted 00th
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2016Please do not adjust margins ESI for: Received 00th January 20xx, Accepted 00th
The Molecular Dynamics Simulation Process
 The Molecular Dynamics Simulation Process For textbooks see: M.P. Allen and D.J. Tildesley. Computer Simulation of Liquids.Oxford University Press, New York, 1987. D. Frenkel and B. Smit. Understanding
The Molecular Dynamics Simulation Process For textbooks see: M.P. Allen and D.J. Tildesley. Computer Simulation of Liquids.Oxford University Press, New York, 1987. D. Frenkel and B. Smit. Understanding
The Molecular Dynamics Method
 H-bond energy (kcal/mol) - 4.0 The Molecular Dynamics Method Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis 0 ATPase, a molecular
H-bond energy (kcal/mol) - 4.0 The Molecular Dynamics Method Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis 0 ATPase, a molecular
The Molecular Dynamics Method
 H-bond energy (kcal/mol) - 4.0 The Molecular Dynamics Method Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis 0 ATPase, a molecular
H-bond energy (kcal/mol) - 4.0 The Molecular Dynamics Method Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis 0 ATPase, a molecular
Molecular Mechanics. I. Quantum mechanical treatment of molecular systems
 Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Dihedral Angles. Homayoun Valafar. Department of Computer Science and Engineering, USC 02/03/10 CSCE 769
 Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
MD EXPERIMENT. Initialization : Relaxation : Equilibration : Productive run: Analysis : building molecular system; initial coordinates
 MD EXPERIMENT Initialization : Relaxation : building molecular system; initial coordinates the system is brought to a state where actual Equilibration : Productive run: Analysis : dynamics can be performed
MD EXPERIMENT Initialization : Relaxation : building molecular system; initial coordinates the system is brought to a state where actual Equilibration : Productive run: Analysis : dynamics can be performed
Force Fields for Classical Molecular Dynamics simulations of Biomolecules. Emad Tajkhorshid
 Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Theoretical and Computational Biophysics Group, Beckman Institute Departments of Biochemistry and Pharmacology,
Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Theoretical and Computational Biophysics Group, Beckman Institute Departments of Biochemistry and Pharmacology,
The Computational Microscope
 The Computational Microscope Computational microscope views at atomic resolution... Rs SER RER C E M RER N GA L... how living cells maintain health and battle disease John Stone Our Microscope is Made
The Computational Microscope Computational microscope views at atomic resolution... Rs SER RER C E M RER N GA L... how living cells maintain health and battle disease John Stone Our Microscope is Made
Lecture 11: Potential Energy Functions
 Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Is quantum mechanics necessary for predicting binding free energy?
 Supporting Information Is quantum mechanics necessary for predicting binding free energy? Ting Zhou, Danzhi Huang and Amedeo Caflisch* Department of Biochemistry, University of Zürich, Winterthurerstrasse
Supporting Information Is quantum mechanics necessary for predicting binding free energy? Ting Zhou, Danzhi Huang and Amedeo Caflisch* Department of Biochemistry, University of Zürich, Winterthurerstrasse
Figure 1. Molecules geometries of 5021 and Each neutral group in CHARMM topology was grouped in dash circle.
 Project I Chemistry 8021, Spring 2005/2/23 This document was turned in by a student as a homework paper. 1. Methods First, the cartesian coordinates of 5021 and 8021 molecules (Fig. 1) are generated, in
Project I Chemistry 8021, Spring 2005/2/23 This document was turned in by a student as a homework paper. 1. Methods First, the cartesian coordinates of 5021 and 8021 molecules (Fig. 1) are generated, in
Force Fields for Classical Molecular Dynamics simulations of Biomolecules. Emad Tajkhorshid
 Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Beckman Institute Departments of Biochemistry Center for Biophysics and Computational Biology University of Illinois
Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Beckman Institute Departments of Biochemistry Center for Biophysics and Computational Biology University of Illinois
Computational Studies of the Photoreceptor Rhodopsin. Scott E. Feller Wabash College
 Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
Methods of Computer Simulation. Molecular Dynamics and Monte Carlo
 Molecular Dynamics Time is of the essence in biological processes therefore how do we understand time-dependent processes at the molecular level? How do we do this experimentally? How do we do this computationally?
Molecular Dynamics Time is of the essence in biological processes therefore how do we understand time-dependent processes at the molecular level? How do we do this experimentally? How do we do this computationally?
Force Fields for MD simulations
 Force Fields for MD simulations Topology/parameter files Where do the numbers an MD code uses come from? ow to make topology files for ligands, cofactors, special amino acids, ow to obtain/develop missing
Force Fields for MD simulations Topology/parameter files Where do the numbers an MD code uses come from? ow to make topology files for ligands, cofactors, special amino acids, ow to obtain/develop missing
Environment Research Institute, Shandong University, Jinan , P. R. China. Keywords
 Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supplementary data for Dehydrochlorination mechanism of γ- hexachlorocyclohexane degraded by
Electronic Supplementary Material (ESI) for RSC Advances. This journal is The Royal Society of Chemistry 2015 Supplementary data for Dehydrochlorination mechanism of γ- hexachlorocyclohexane degraded by
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation
 Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Polypeptide Folding Using Monte Carlo Sampling, Concerted Rotation, and Continuum Solvation Jakob P. Ulmschneider and William L. Jorgensen J.A.C.S. 2004, 126, 1849-1857 Presented by Laura L. Thomas and
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Computational Biology & Computational Medicine
 Computational Biology & Computational Medicine Homayoun Valafar Outline Why proteins? What are proteins? How do we compute them? How do we use computational approaches? Why Proteins? Molecular basis of
Computational Biology & Computational Medicine Homayoun Valafar Outline Why proteins? What are proteins? How do we compute them? How do we use computational approaches? Why Proteins? Molecular basis of
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS
 Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Supporting Protocol This protocol describes the construction and the force-field parameters of the non-standard residue for the Ag + -site using CNS CNS input file generatemetal.inp: remarks file generate/generatemetal.inp
Protein Structure Prediction
 Protein Structure Prediction Michael Feig MMTSB/CTBP 2006 Summer Workshop From Sequence to Structure SEALGDTIVKNA Ab initio Structure Prediction Protocol Amino Acid Sequence Conformational Sampling to
Protein Structure Prediction Michael Feig MMTSB/CTBP 2006 Summer Workshop From Sequence to Structure SEALGDTIVKNA Ab initio Structure Prediction Protocol Amino Acid Sequence Conformational Sampling to
DISCRETE TUTORIAL. Agustí Emperador. Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING:
 DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
DISCRETE TUTORIAL Agustí Emperador Institute for Research in Biomedicine, Barcelona APPLICATION OF DISCRETE TO FLEXIBLE PROTEIN-PROTEIN DOCKING: STRUCTURAL REFINEMENT OF DOCKING CONFORMATIONS Emperador
Limitations of temperature replica exchange (T-REMD) for protein folding simulations
 Limitations of temperature replica exchange (T-REMD) for protein folding simulations Jed W. Pitera, William C. Swope IBM Research pitera@us.ibm.com Anomalies in protein folding kinetic thermodynamic 322K
Limitations of temperature replica exchange (T-REMD) for protein folding simulations Jed W. Pitera, William C. Swope IBM Research pitera@us.ibm.com Anomalies in protein folding kinetic thermodynamic 322K
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Free Energy Landscape of Protein Folding in Water: Explicit vs. Implicit Solvent
 PROTEINS: Structure, Function, and Genetics 53:148 161 (2003) Free Energy Landscape of Protein Folding in Water: Explicit vs. Implicit Solvent Ruhong Zhou* IBM T.J. Watson Research Center, Yorktown Heights,
PROTEINS: Structure, Function, and Genetics 53:148 161 (2003) Free Energy Landscape of Protein Folding in Water: Explicit vs. Implicit Solvent Ruhong Zhou* IBM T.J. Watson Research Center, Yorktown Heights,
Advanced Molecular Dynamics
 Advanced Molecular Dynamics Introduction May 2, 2017 Who am I? I am an associate professor at Theoretical Physics Topics I work on: Algorithms for (parallel) molecular simulations including GPU acceleration
Advanced Molecular Dynamics Introduction May 2, 2017 Who am I? I am an associate professor at Theoretical Physics Topics I work on: Algorithms for (parallel) molecular simulations including GPU acceleration
Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia
 University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
Protein Folding experiments and theory
 Protein Folding experiments and theory 1, 2,and 3 Protein Structure Fig. 3-16 from Lehninger Biochemistry, 4 th ed. The 3D structure is not encoded at the single aa level Hydrogen Bonding Shared H atom
Protein Folding experiments and theory 1, 2,and 3 Protein Structure Fig. 3-16 from Lehninger Biochemistry, 4 th ed. The 3D structure is not encoded at the single aa level Hydrogen Bonding Shared H atom
An introduction to Molecular Dynamics. EMBO, June 2016
 An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
Follow this and additional works at: Part of the Biomedical Engineering and Bioengineering Commons
 Clemson University TigerPrints All Dissertations Dissertations 5-2011 Methods Development and Force Field Evaluation for Molecular Simulations of Iinteractions Between Structured Peptides and Functionalized
Clemson University TigerPrints All Dissertations Dissertations 5-2011 Methods Development and Force Field Evaluation for Molecular Simulations of Iinteractions Between Structured Peptides and Functionalized
Multi-Scale Hierarchical Structure Prediction of Helical Transmembrane Proteins
 Multi-Scale Hierarchical Structure Prediction of Helical Transmembrane Proteins Zhong Chen Dept. of Biochemistry and Molecular Biology University of Georgia, Athens, GA 30602 Email: zc@csbl.bmb.uga.edu
Multi-Scale Hierarchical Structure Prediction of Helical Transmembrane Proteins Zhong Chen Dept. of Biochemistry and Molecular Biology University of Georgia, Athens, GA 30602 Email: zc@csbl.bmb.uga.edu
Principles and Applications of Molecular Dynamics Simulations with NAMD
 Principles and Applications of Molecular Dynamics Simulations with NAMD Nov. 14, 2016 Computational Microscope NCSA supercomputer JC Gumbart Assistant Professor of Physics Georgia Institute of Technology
Principles and Applications of Molecular Dynamics Simulations with NAMD Nov. 14, 2016 Computational Microscope NCSA supercomputer JC Gumbart Assistant Professor of Physics Georgia Institute of Technology
Modeling Biological Systems Opportunities for Computer Scientists
 Modeling Biological Systems Opportunities for Computer Scientists Filip Jagodzinski RBO Tutorial Series 25 June 2007 Computer Science Robotics & Biology Laboratory Protein: πρώτα, "prota, of Primary Importance
Modeling Biological Systems Opportunities for Computer Scientists Filip Jagodzinski RBO Tutorial Series 25 June 2007 Computer Science Robotics & Biology Laboratory Protein: πρώτα, "prota, of Primary Importance
Molecular dynamics simulation. CS/CME/BioE/Biophys/BMI 279 Oct. 5 and 10, 2017 Ron Dror
 Molecular dynamics simulation CS/CME/BioE/Biophys/BMI 279 Oct. 5 and 10, 2017 Ron Dror 1 Outline Molecular dynamics (MD): The basic idea Equations of motion Key properties of MD simulations Sample applications
Molecular dynamics simulation CS/CME/BioE/Biophys/BMI 279 Oct. 5 and 10, 2017 Ron Dror 1 Outline Molecular dynamics (MD): The basic idea Equations of motion Key properties of MD simulations Sample applications
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro. Karel Berka, Ph.D.
 4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
Supporting Information
 Supporting Information Constant ph molecular dynamics reveals ph-modulated binding of two small-molecule BACE1 inhibitors Christopher R. Ellis 1,, Cheng-Chieh Tsai 1,, Xinjun Hou 2, and Jana Shen 1, 1
Supporting Information Constant ph molecular dynamics reveals ph-modulated binding of two small-molecule BACE1 inhibitors Christopher R. Ellis 1,, Cheng-Chieh Tsai 1,, Xinjun Hou 2, and Jana Shen 1, 1
Gromacs Workshop Spring CSC
 Gromacs Workshop Spring 2007 @ CSC Erik Lindahl Center for Biomembrane Research Stockholm University, Sweden David van der Spoel Dept. Cell & Molecular Biology Uppsala University, Sweden Berk Hess Max-Planck-Institut
Gromacs Workshop Spring 2007 @ CSC Erik Lindahl Center for Biomembrane Research Stockholm University, Sweden David van der Spoel Dept. Cell & Molecular Biology Uppsala University, Sweden Berk Hess Max-Planck-Institut
LAMMPS Performance Benchmark on VSC-1 and VSC-2
 LAMMPS Performance Benchmark on VSC-1 and VSC-2 Daniel Tunega and Roland Šolc Institute of Soil Research, University of Natural Resources and Life Sciences VSC meeting, Neusiedl am See, February 27-28,
LAMMPS Performance Benchmark on VSC-1 and VSC-2 Daniel Tunega and Roland Šolc Institute of Soil Research, University of Natural Resources and Life Sciences VSC meeting, Neusiedl am See, February 27-28,
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Energy Landscapes and Accelerated Molecular- Dynamical Techniques for the Study of Protein Folding
 Energy Landscapes and Accelerated Molecular- Dynamical Techniques for the Study of Protein Folding John K. Prentice Boulder, CO BioMed Seminar University of New Mexico Physics and Astronomy Department
Energy Landscapes and Accelerated Molecular- Dynamical Techniques for the Study of Protein Folding John K. Prentice Boulder, CO BioMed Seminar University of New Mexico Physics and Astronomy Department
Procheck output. Bond angles (Procheck) Structure verification and validation Bond lengths (Procheck) Introduction to Bioinformatics.
 Structure verification and validation Bond lengths (Procheck) Introduction to Bioinformatics Iosif Vaisman Email: ivaisman@gmu.edu ----------------------------------------------------------------- Bond
Structure verification and validation Bond lengths (Procheck) Introduction to Bioinformatics Iosif Vaisman Email: ivaisman@gmu.edu ----------------------------------------------------------------- Bond
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Van Gunsteren et al. Angew. Chem. Int. Ed. 45, 4064 (2006)
 Van Gunsteren et al. Angew. Chem. Int. Ed. 45, 4064 (2006) Martini Workshop 2015 Coarse Graining Basics Alex de Vries Every word or concept, clear as it may seem to be, has only a limited range of applicability
Van Gunsteren et al. Angew. Chem. Int. Ed. 45, 4064 (2006) Martini Workshop 2015 Coarse Graining Basics Alex de Vries Every word or concept, clear as it may seem to be, has only a limited range of applicability
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Molecular Dynamics Simulations. Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia
 Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Protein Dynamics, Allostery and Function
 Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields
 Department of Chemistry and Biochemistry, Concordia University! page 1 of 6 CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields INTRODUCTION The goal of this tutorial
Department of Chemistry and Biochemistry, Concordia University! page 1 of 6 CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields INTRODUCTION The goal of this tutorial
Biomolecules are dynamic no single structure is a perfect model
 Molecular Dynamics Simulations of Biomolecules References: A. R. Leach Molecular Modeling Principles and Applications Prentice Hall, 2001. M. P. Allen and D. J. Tildesley "Computer Simulation of Liquids",
Molecular Dynamics Simulations of Biomolecules References: A. R. Leach Molecular Modeling Principles and Applications Prentice Hall, 2001. M. P. Allen and D. J. Tildesley "Computer Simulation of Liquids",
Folding WT villin in silico Three folding simulations reach native state within 5-8 µs
 Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign Molecular Modeling Flexible Fitting 1: Introduction to Molecular Dynamics MD force field Equation
Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign Molecular Modeling Flexible Fitting 1: Introduction to Molecular Dynamics MD force field Equation
This semester. Books
 Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
Applications of Molecular Dynamics
 June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
Molecular Simulation III
 Molecular Simulation III Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University Time and Length Scales Tamar Schlick s Biomolecular Structure and
Molecular Simulation III Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences Duquesne University Time and Length Scales Tamar Schlick s Biomolecular Structure and
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D.
 3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
3. Solutions W = N!/(N A!N B!) (3.1) Using Stirling s approximation ln(n!) = NlnN N: ΔS mix = k (N A lnn + N B lnn N A lnn A N B lnn B ) (3.
 3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
Why study protein dynamics?
 Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
Computational Modeling of Protein Dynamics. Yinghao Wu Department of Systems and Computational Biology Albert Einstein College of Medicine Fall 2014
 Computational Modeling of Protein Dynamics Yinghao Wu Department of Systems and Computational Biology Albert Einstein College of Medicine Fall 2014 Outline Background of protein dynamics Basic computational
Computational Modeling of Protein Dynamics Yinghao Wu Department of Systems and Computational Biology Albert Einstein College of Medicine Fall 2014 Outline Background of protein dynamics Basic computational
Monte Carlo simulations of polyalanine using a reduced model and statistics-based interaction potentials
 THE JOURNAL OF CHEMICAL PHYSICS 122, 024904 2005 Monte Carlo simulations of polyalanine using a reduced model and statistics-based interaction potentials Alan E. van Giessen and John E. Straub Department
THE JOURNAL OF CHEMICAL PHYSICS 122, 024904 2005 Monte Carlo simulations of polyalanine using a reduced model and statistics-based interaction potentials Alan E. van Giessen and John E. Straub Department
Molecular Mechanics. Yohann Moreau. November 26, 2015
 Molecular Mechanics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, 2015 1 / 29 Introduction A so-called Force-Field
Molecular Mechanics Yohann Moreau yohann.moreau@ujf-grenoble.fr November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, 2015 1 / 29 Introduction A so-called Force-Field
Molecular Simulation II. Classical Mechanical Treatment
 Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields
 Department of Chemistry and Biochemistry, Concordia University page 1 of 6 CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields INTRODUCTION The goal of this tutorial
Department of Chemistry and Biochemistry, Concordia University page 1 of 6 CHEM 498Q / CHEM 630Q: Molecular Modeling of Proteins TUTORIAL #3a: Empirical force fields INTRODUCTION The goal of this tutorial
Fondamenti di Chimica Farmaceutica. Computer Chemistry in Drug Research: Introduction
 Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
ONETEP PB/SA: Application to G-Quadruplex DNA Stability. Danny Cole
 ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
Electronic excitations in conjugated. Many-Body Green's Functions. Behnaz Bagheri Varnousfaderani. Behnaz Bagheri Varnousfaderani
 Technische Universiteit Eindhoven University of Technology Electronic excitations in conjugated polymers Electronic from excitations QM/MM simulations in conjugated pol based from on Many-Body QM/MM simulations
Technische Universiteit Eindhoven University of Technology Electronic excitations in conjugated polymers Electronic from excitations QM/MM simulations in conjugated pol based from on Many-Body QM/MM simulations
Protein Modeling Methods. Knowledge. Protein Modeling Methods. Fold Recognition. Knowledge-based methods. Introduction to Bioinformatics
 Protein Modeling Methods Introduction to Bioinformatics Iosif Vaisman Ab initio methods Energy-based methods Knowledge-based methods Email: ivaisman@gmu.edu Protein Modeling Methods Ab initio methods:
Protein Modeling Methods Introduction to Bioinformatics Iosif Vaisman Ab initio methods Energy-based methods Knowledge-based methods Email: ivaisman@gmu.edu Protein Modeling Methods Ab initio methods:
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model. John Dood Hope College
 Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
Some Accelerated/Enhanced Sampling Techniques
 Some Accelerated/Enhanced Sampling Techniques The Sampling Problem Accelerated sampling can be achieved by addressing one or more of the MD ingredients Back to free energy differences... Examples of accelerated
Some Accelerated/Enhanced Sampling Techniques The Sampling Problem Accelerated sampling can be achieved by addressing one or more of the MD ingredients Back to free energy differences... Examples of accelerated
Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached
 Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached to HpUreI. Urea hydrolysis products 2NH 3 and 1CO 2
Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached to HpUreI. Urea hydrolysis products 2NH 3 and 1CO 2
Coupling the Level-Set Method with Variational Implicit Solvent Modeling of Molecular Solvation
 Coupling the Level-Set Method with Variational Implicit Solvent Modeling of Molecular Solvation Bo Li Math Dept & CTBP, UCSD Li-Tien Cheng (Math, UCSD) Zhongming Wang (Math & Biochem, UCSD) Yang Xie (MAE,
Coupling the Level-Set Method with Variational Implicit Solvent Modeling of Molecular Solvation Bo Li Math Dept & CTBP, UCSD Li-Tien Cheng (Math, UCSD) Zhongming Wang (Math & Biochem, UCSD) Yang Xie (MAE,
TOPOLOGIES AND FORCE FIELD PARAMETERS FOR NITROXIDE SPIN LABELS
 TOPOLOGIES AND FORCE FIELD PARAMETERS FOR NITROXIDE SPIN LABELS 1. Introduction The force field used in these studies is the CHARMM19 (Chemistry at HARvard Molecular Mechanics) extended atom force field
TOPOLOGIES AND FORCE FIELD PARAMETERS FOR NITROXIDE SPIN LABELS 1. Introduction The force field used in these studies is the CHARMM19 (Chemistry at HARvard Molecular Mechanics) extended atom force field
Challenges in generation of conformational ensembles for peptides and small proteins
 Challenges in generation of conformational ensembles for peptides and small proteins Carlos Simmerling Stony Brook University What could (and does) go wrong? 1. Sampling: difficult to obtain converged
Challenges in generation of conformational ensembles for peptides and small proteins Carlos Simmerling Stony Brook University What could (and does) go wrong? 1. Sampling: difficult to obtain converged
Introduction The gramicidin A (ga) channel forms by head-to-head association of two monomers at their amino termini, one from each bilayer leaflet. Th
 Abstract When conductive, gramicidin monomers are linked by six hydrogen bonds. To understand the details of dissociation and how the channel transits from a state with 6H bonds to ones with 4H bonds or
Abstract When conductive, gramicidin monomers are linked by six hydrogen bonds. To understand the details of dissociation and how the channel transits from a state with 6H bonds to ones with 4H bonds or
Conformational sampling of macrocycles in solution and in the solid state
 Conformational sampling of macrocycles in solution and in the solid state Paul Hawkins, Ph.D. Head of Scientific Solutions Stanislaw Wlodek, Ph.D. Senior Scientific Developer 6/6/2018 https://berkonomics.com/?p=2437
Conformational sampling of macrocycles in solution and in the solid state Paul Hawkins, Ph.D. Head of Scientific Solutions Stanislaw Wlodek, Ph.D. Senior Scientific Developer 6/6/2018 https://berkonomics.com/?p=2437
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
Supplementary Figures:
 Supplementary Figures: Supplementary Figure 1: The two strings converge to two qualitatively different pathways. A) Models of active (red) and inactive (blue) states used as end points for the string calculations
Supplementary Figures: Supplementary Figure 1: The two strings converge to two qualitatively different pathways. A) Models of active (red) and inactive (blue) states used as end points for the string calculations
Exercise 2: Solvating the Structure Before you continue, follow these steps: Setting up Periodic Boundary Conditions
 Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Exercise 2: Solvating the Structure HyperChem lets you place a molecular system in a periodic box of water molecules to simulate behavior in aqueous solution, as in a biological system. In this exercise,
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
The Force Field Toolkit (fftk)
 Parameterizing Small Molecules Using: The Force Field Toolkit (fftk) Christopher G. Mayne, Emad Tajkhorshid Beckman Institute for Advanced Science and Technology University of Illinois, Urbana-Champaign
Parameterizing Small Molecules Using: The Force Field Toolkit (fftk) Christopher G. Mayne, Emad Tajkhorshid Beckman Institute for Advanced Science and Technology University of Illinois, Urbana-Champaign
Molecular simulation and structure prediction using CHARMM and the MMTSB Tool Set Free Energy Methods
 Molecular simulation and structure prediction using CHARMM and the MMTSB Tool Set Free Energy Methods Charles L. Brooks III MMTSB/CTBP 2006 Summer Workshop CHARMM Simulations The flow of data and information
Molecular simulation and structure prediction using CHARMM and the MMTSB Tool Set Free Energy Methods Charles L. Brooks III MMTSB/CTBP 2006 Summer Workshop CHARMM Simulations The flow of data and information
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data
 Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION
 THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
Computational protein design
 Computational protein design There are astronomically large number of amino acid sequences that needs to be considered for a protein of moderate size e.g. if mutating 10 residues, 20^10 = 10 trillion sequences
Computational protein design There are astronomically large number of amino acid sequences that needs to be considered for a protein of moderate size e.g. if mutating 10 residues, 20^10 = 10 trillion sequences
K ex. Conformational equilibrium. equilibrium K B
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Improving Protein Function Prediction with Molecular Dynamics Simulations. Dariya Glazer Russ Altman
 Improving Protein Function Prediction with Molecular Dynamics Simulations Dariya Glazer Russ Altman Motivation Sometimes the 3D structure doesn t score well for a known function. The experimental structure
Improving Protein Function Prediction with Molecular Dynamics Simulations Dariya Glazer Russ Altman Motivation Sometimes the 3D structure doesn t score well for a known function. The experimental structure
Interpreting and evaluating biological NMR in the literature. Worksheet 1
 Interpreting and evaluating biological NMR in the literature Worksheet 1 1D NMR spectra Application of RF pulses of specified lengths and frequencies can make certain nuclei detectable We can selectively
Interpreting and evaluating biological NMR in the literature Worksheet 1 1D NMR spectra Application of RF pulses of specified lengths and frequencies can make certain nuclei detectable We can selectively
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
2008 Biowerkzeug Ltd.
 2008 Biowerkzeug Ltd. 1 Contents Summary...3 1 Simulation...4 1.1 Setup...4 1.2 Output...4 2 Settings...5 3 Analysis...9 3.1 Setup...9 3.2 Input options...9 3.3 Descriptions...10 Please note that we cannot
2008 Biowerkzeug Ltd. 1 Contents Summary...3 1 Simulation...4 1.1 Setup...4 1.2 Output...4 2 Settings...5 3 Analysis...9 3.1 Setup...9 3.2 Input options...9 3.3 Descriptions...10 Please note that we cannot
COSMO-RS Theory. The Basics
 Theory The Basics From µ to properties Property µ 1 µ 2 activity coefficient vapor pressure Infinite dilution Gas phase Pure compound Pure bulk compound Partition coefficient Phase 1 Phase 2 Liquid-liquid
Theory The Basics From µ to properties Property µ 1 µ 2 activity coefficient vapor pressure Infinite dilution Gas phase Pure compound Pure bulk compound Partition coefficient Phase 1 Phase 2 Liquid-liquid
Protein Modeling. Generating, Evaluating and Refining Protein Homology Models
 Protein Modeling Generating, Evaluating and Refining Protein Homology Models Troy Wymore and Kristen Messinger Biomedical Initiatives Group Pittsburgh Supercomputing Center Homology Modeling of Proteins
Protein Modeling Generating, Evaluating and Refining Protein Homology Models Troy Wymore and Kristen Messinger Biomedical Initiatives Group Pittsburgh Supercomputing Center Homology Modeling of Proteins
Simulating biomolecular function from motions across multiple scales (I) Peter J. Bond (BII)
 Simulating biomolecular function from motions across multiple scales (I) Peter J. Bond (BII) peterjb@bii.a-star.edu.sg Structural Biology: Why the Need for Simulation? 2017 year 1972 0 RCSB PDB: RCSB Protein
Simulating biomolecular function from motions across multiple scales (I) Peter J. Bond (BII) peterjb@bii.a-star.edu.sg Structural Biology: Why the Need for Simulation? 2017 year 1972 0 RCSB PDB: RCSB Protein
Choosing weights for simulated tempering
 PHYSICAL REVIEW E 7, 173 7 Choosing weights for simulated tempering Sanghyun Park 1 and Vijay S. Pande 1, 1 Department of Chemistry, Stanford University, Stanford, California 935, USA Department of Structural
PHYSICAL REVIEW E 7, 173 7 Choosing weights for simulated tempering Sanghyun Park 1 and Vijay S. Pande 1, 1 Department of Chemistry, Stanford University, Stanford, California 935, USA Department of Structural
Protein Structure Prediction
 Protein Structure Prediction Michael Feig MMTSB/CTBP 2009 Summer Workshop From Sequence to Structure SEALGDTIVKNA Folding with All-Atom Models AAQAAAAQAAAAQAA All-atom MD in general not succesful for real
Protein Structure Prediction Michael Feig MMTSB/CTBP 2009 Summer Workshop From Sequence to Structure SEALGDTIVKNA Folding with All-Atom Models AAQAAAAQAAAAQAA All-atom MD in general not succesful for real
Methods for searching and sampling configurational space
 International Spring School Statistical Thermodynamics, Santiago de Chile Tuesday, ovember 28, 2017 Lecture 18 Methods for searching and sampling configurational space Prof. Dr. Wilfred F. van Gunsteren
International Spring School Statistical Thermodynamics, Santiago de Chile Tuesday, ovember 28, 2017 Lecture 18 Methods for searching and sampling configurational space Prof. Dr. Wilfred F. van Gunsteren
