Molecular Mechanics. Yohann Moreau. November 26, 2015
|
|
|
- Bruno Hunt
- 5 years ago
- Views:
Transcription
1 Molecular Mechanics Yohann Moreau November 26, 2015 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
2 Introduction A so-called Force-Field is based on an empirical approach No explicit description of electrons (no QM) - Electrons are implicitely included in atomic parameters Bonds are not a result of the calculation (as in QM) but are parameters - Balls & Springs model - Energy is a function of geometry AND topology! Different types of atoms that interact with defined potentials - Parameters depend on the nature of interacting atoms A force field is made of a huge number of parameters A force field must be limited to a defined kind of system - DNA, Solids, fatty acids, proteins, etc... Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
3 Generality The total energy of a system is the sum of different terms They account for interactions between all the atoms of the system: Interactions between bound atoms - Bonding term between 2 atoms a & b: E r ab =f(r ab) - Angle (3 atoms): E=f(θ abc ) - Torsional (4 atoms): E fct of the dihedral between 4 atoms - Improper Torsional (4 atoms): for planar env. (e.g. sp2 carbon) Atoms separated by 4 bonds or more are unbound - Their interaction is made of Van der Waals and electrostatic forces (charge/charge) Some other specific terms can also be added (after all it s empirical) The total energy can thus be written: E tot = E bond + E angle + E dihedral + E VdW + E Elst Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
4 Illustration of the different terms E tot = E bond + E angle + E dihedrah + E VdW + E Elst Termes liés E a =f(θ) E l =f(r) E d =f(φ) E d =f(χ) Termes non liés E VdW =f(r) δ+ δ- E Elec =f(r) Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
5 Determination of the energy of a system To know the energy as a function of the geometry, one has to: know the topology of the molecule - how atoms are bound together? know the type of each atom and their position Count all the interaction terms associated Once all of this is gathered, energy can be computed If one parameter is missing, energy can t be determined Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
6 Atom types The atomic type depends on - its nature - its hybridization - its neighbours Hence a FF is generally restrained to a given type of system such as proteins Parameters are considered transferable (similar system) See charmm22.prm e.g. from Introduction to Computational Chemistry, p23, F. Jensen Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
7 Exercise: propanal in interaction with H 2 O Using the following representation of the system: - determine the number of degrees of freedom - identify all the terms needed to calculate the total potential energy of the system Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
8 Application: propanal in interaction with H 2 O Count of the different terms - 8 bonds: 3 C-H, 1 C-C, 1 C-O, 1 C-H ; 2 O-H - 10 angles : 3 H-C-H, 3 H-C-C, 1 C-C-H, 1 C-C-O, 1 H-O-H - 6 dihedral: 3 H-C-C-H, 3 H-C-C-O - 1 improper dihedral: C-C-O-H = 21 VdW interactions and 21 electrostatic interactions - 1 possible H-bond (not in all force fields) Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
9 Potentials of interaction Once terms identified, energy associated to each can be computed Each potential accounts for the energy modification in fonction of the modification of the degree of freedom associated This must allow to reproduce the PES around the equilibrium geometry (region for wich E is 40 kj/mol or less above minimum) Potentials have a simple analytic expression so as to be easily derived for dynamics and optimisation Remarks: - The equilibrium point of an interaction is a force-field parameter! - A given degree of freedom will remain close to this value (vide infra) - Exemple: r CC 0 varies close to 1,523Å in MM2 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
10 math point: the Taylor development near a given point x 0, a function can be expressed as a sum of the derivatives (polynomial approx.): f (x) = f (k) (x 0 ) (x x 0 ) k + O(k + 1) n! k=0,n At the second order, one gets a parabola: f (x) = f (x 0 ) + (x x 0 ).f (x 0 ) + 1/2.(x x 0 ) 2.f (x 0 ) f (x 0 ): value of the function at x 0 - if x 0 is the equilibrium parameter, this is the lowest point (equilibrium energy) f (x 0 ) = 0 if x 0 is a minimum: the curve is flat f (x 0 ) = force constant af an harmonic potential The goal is to mimic the unknown fonction f(x) around x 0, the equilibrium position by a parabola Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
11 Application: Bonding potential between two atoms (stretching) The energy variation in function of the interatomic distance is complex At the vicinity of the equilibrium distance, the curve is flat (f (x)=0) At the actual equilibrium distance r 0, energy is zero relative to the equilibrium energy The stretch energy between atoms i and j is expressed by a harmonic potential E lij = 1 2 k ij(r ij rij 0)2 Where: - rij 0 is the equilibrium distance between i & j - k ij is the force constant of the spring, usually in kcal/mol/å2 Stretching terms can also be more complex (including exponentials, e.g.) but the harmonic form is the most common Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
12 Beyond the harmonic approximation Anharmonic potential -Taylor development can be made at a greater order (4, e.g.) E l,anh (r r 0 ) = k ij 2 (r r 0) 2 + k ij 3 (r r 0) 3 + k ij 4 (r r 0) 4 More parameters to define and higher computational cost Morse potential: - E l,morse (r r 0 ) = D(1 exp [ α(r r 0)] 2 ), α = k 2D - D: dissociation energy, α = f (k) - good reproduciton of the dissociation curve but less usable Common force fields use quadratic potentials Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
13 Illustration: Different C-H stretching potentials from: Introduction to Computational Chemistry, p28, F. Jensen Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
14 How is the parameter k ij determined? The fonction to fit has an unknown analytical expression The value of k depends on the type of atoms bound Different ways to get the force constant and equilibrium distance: - From vibrational spectra (IR) - From a dissociation profile obtained by quantum chemistry the creation of a force field is a huge work of parametrization Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
15 Bending terms Increase or decrease the angle makes the energy increase One uses a harmonic potential in function of the angle θ E a = 1 2 k ijk.(θ θ 0 ) 2 k ijk is expressed in kcal/mol/degrees 2 and depends on the nature of atoms i, j and k The harmonic approximation is good enough at ±30 around θ 0 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
16 Torsion: I- out of plane bending This term accounts for the cost of puting an atom out of plane - Example: pyramidalization of an sp 2 atom (planar environment at equilibrium) E oop (φ) = k.φ 2 ; at equilibrium: φ 0 = 0 Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
17 Torsion: II- dihedral For 4 bound atoms, the term represents the angle between tyhe two external bonds - Torsions have a periodic character: E(360 ) = E(0 ) - several minima and maxima (weak rotational barriers) - A quadratic potential (harmonic) is not suitable cosine-type function is often used: x E diedre (φ) = V n cos(nφ) n=1 - x: nb. of minima (max: 5), V n fixes the barrier height φ [0 360 ] Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
18 Example: torsional potential with 3 minima A term (1+) is added to shift position of zero, as well as a phase factor τ E = 1 2 [0.5(1+cos(1φ τ) 0.2(1 cos(2φ τ))+0.5(1+cos(3φ τ))] Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
19 Non bonded terms: electrostatic interactions This strong interactions takes place between charges atoms - Charge accounts for the difference between the amount of negative (e ) and positive (protons) charges carried by an atom. - It depends on the nature of each type of atom (electronegativity and neighbours) The pair interaction writes: E elec,ij = 1 4πɛ 0 q i.q j r ij The electrostatic energy is thus: - positive (repulsive) for two atoms with charge of the same sign - negative (attractive) between a negative and a positive atom - proportionnal to the product of the two charges - inverse to the interatomic distance This interaction is counted only for atoms with more than 3 bonds between Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
20 Non bonded terms: electrostatic interactions Determination of charges Generally, each type of atom has a specific charge Charges are determined by mean of quantum calculation - always the same level of theory (MP2/6-31G* e.g.) for consistency Charges are fitted so as to reproduce the QM electrostatic potential - In protein force fields, amino-acids are split into groups: side chain and backbone - Each group has a whole charge (zero for backbone) See charmm topology file eg Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
21 Higher order electrostatic terms The electronic structure around an atom can be developped as a series of multipoles of different orders - Charge is a multipole of zeroth order - An atomic dipole can be added, rarely higher order terms - It can account for the local dissymmetry of the electronic structure This adds supplementary terms: charge/dipole et dipole/dipole General force fields don t make use of such terms There are also polarizable force fields in which environment modifies the atomic multipolar structure - totally beyond our scope Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
22 Non bonded terms: Van der Waals interactions The Van der Waals interaction is the sum of non-polar interactions between atoms These interactions originate in quantum phenomena - Pauli repulsion: at short distance, two electronic clouds can t interpenetrate - Dispersion interactions (London, Keesom, Debye forces): slightly attractive ( behave as 1/r 6 ) A Van der Waals potential is written as the sum of the two phenomena and vanishes quickly: E VdW (r ij ) = E repulsion (r ij ) C ij r 6 ij The expression of E repulsion can be written in different ways Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
23 Van der Waals potentials Lennard-Jones-type potential is the most common - repulsion varies as r 12 : E LJ (r ij ) = 4ε ij [( σ ij r ij ) 12 ( σ ij r ij ) 6 ] - σ ij, VdW radius, distance for which E=0: σ ij = σ i + σ j - ε is the well-depth (energy) ε ij = ε i ε j There are other expressions for the repuslive part: - Example: Buckingham/Hill potential: E Hill = A exp BR C R 6 Other specific terms, such as H-bonds: - Very rare: E LH = ε ij [5( σ ij r ij ) 12 6( σ ij r ij ) 10 ] At long distance, such interactions can be neglected beyond a cut-off value (often around 12 to 14 Å) Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
24 Lennard-Jones potential between two non polar hydrogens Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
25 Common ForceFields A force field is the whole set of parameters and equations that allows the energetic description of a given system force fields are specific to defined kind of systems: - CHARMM, Gromos & Amber for proteins - MM2/3/4: general application ( - UFF (Universal Force Field): parameters are calcultaed on-the-fly Whatever the force field, the molecular topology as well as all parameters MUST be known and remain unchanged - Applied to a given geometry, one can compute the associated energy Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
26 Example: the CHARMM Force Field Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
27 A specific force field for water: TIP3P Rather old: 1983 for the first version but still in use -only O has L-J params: σ O = 1, 7683 Å, ε O = 0, 1520 kcal.mol 1 - geometry is fixed: 104,52 ; r OH = 0,9572 Å - charges: q H = +0,417 et q O = -0,834 It was built to reproduce properties of liquid water: -density, strucutre, diffusion Exercise: calculate the dipole moment of TIP3P water with: 1D = 2,54 u.a. ; µ exp = 1, 86D ; discuss your result Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
28 Application force fields Strength: very fast (energy and derivatives) - allows the study of very large systems and/or carry out a large number of calculation Drawbacks: very specific to the kind of system studied - impossible to modify the topology and atomic types Common applications: - Molecular dynamics: evolution along time of a system - Statistical sampling in general - Conformational analysis (see labs) - Structure refinement for X-ray or NMR structure determination - Molecular docking between a protein and a ligand - Qantitative Structure-Activity Relation Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
29 Other force field-like methods Force fields have been defined at the atomic level but other cases exist: Coarse grained FFs: -A particle represents several atoms such as the side chain of an amino-acid Allows to study way larger systems Allows to study slow phenomena such as protein folding/assembly Reactive force fields: parameters vary in function of the geometry - Narrow domain of application, quantum-based methods are more suited Yohann Moreau (UJF) Molecular Mechanics, Label RFCT 2015 November 26, / 29
Lecture 11: Potential Energy Functions
 Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Lecture 11: Potential Energy Functions Dr. Ronald M. Levy ronlevy@temple.edu Originally contributed by Lauren Wickstrom (2011) Microscopic/Macroscopic Connection The connection between microscopic interactions
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland
 Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
Example questions for Molecular modelling (Level 4) Dr. Adrian Mulholland 1) Question. Two methods which are widely used for the optimization of molecular geometies are the Steepest descents and Newton-Raphson
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
T6.2 Molecular Mechanics
 T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
T6.2 Molecular Mechanics We have seen that Benson group additivities are capable of giving heats of formation of molecules with accuracies comparable to those of the best ab initio procedures. However,
Dihedral Angles. Homayoun Valafar. Department of Computer Science and Engineering, USC 02/03/10 CSCE 769
 Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Molecular mechanics. classical description of molecules. Marcus Elstner and Tomáš Kubař. April 29, 2016
 classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
classical description of molecules April 29, 2016 Chemical bond Conceptual and chemical basis quantum effect solution of the SR numerically expensive (only small molecules can be treated) approximations
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D.
 3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
3rd Advanced in silico Drug Design KFC/ADD Molecular mechanics intro Karel Berka, Ph.D. Martin Lepšík, Ph.D. Pavel Polishchuk, Ph.D. Thierry Langer, Ph.D. Jana Vrbková, Ph.D. UP Olomouc, 23.1.-26.1. 2018
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii
 The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
The Potential Energy Surface (PES) And the Basic Force Field Chem 4021/8021 Video II.iii Fundamental Points About Which to Be Thinking It s clear the PES is useful, so how can I construct it for an arbitrary
k θ (θ θ 0 ) 2 angles r i j r i j
 1 Force fields 1.1 Introduction The term force field is slightly misleading, since it refers to the parameters of the potential used to calculate the forces (via gradient) in molecular dynamics simulations.
1 Force fields 1.1 Introduction The term force field is slightly misleading, since it refers to the parameters of the potential used to calculate the forces (via gradient) in molecular dynamics simulations.
Force Fields for Classical Molecular Dynamics simulations of Biomolecules. Emad Tajkhorshid
 Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Theoretical and Computational Biophysics Group, Beckman Institute Departments of Biochemistry and Pharmacology,
Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Theoretical and Computational Biophysics Group, Beckman Institute Departments of Biochemistry and Pharmacology,
Force fields in computer simulation of soft nanomaterials
 Lecture 2 Part B Force fields in computer simulation of soft nanomaterials Recommended reading: Leach Chapter 4 1 Force Field Methods Force field methods are simulation methods that use classical force
Lecture 2 Part B Force fields in computer simulation of soft nanomaterials Recommended reading: Leach Chapter 4 1 Force Field Methods Force field methods are simulation methods that use classical force
Semi Empirical Force Fields and Their Limitations. Potential Energy Surface (PES)
 Semi Empirical Force Fields and Their Limitations Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Potential Energy Surface (PES) Schrödinger equation: H T Ψ( r, = E Ψ( r, H =
Semi Empirical Force Fields and Their Limitations Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Potential Energy Surface (PES) Schrödinger equation: H T Ψ( r, = E Ψ( r, H =
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro. Karel Berka, Ph.D.
 4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
Subject of the Lecture:
 Subject of the Lecture: Conceptual basis for the development of force fields. Implementation/validation Water - a worked example Extensions - combining molecular mechanics and quantum mechanics (QM/MM)
Subject of the Lecture: Conceptual basis for the development of force fields. Implementation/validation Water - a worked example Extensions - combining molecular mechanics and quantum mechanics (QM/MM)
Why study protein dynamics?
 Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
Scuola di Chimica Computazionale
 Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica Organica e Biologica Modellistica
Societa Chimica Italiana Gruppo Interdivisionale di Chimica Computazionale Scuola di Chimica Computazionale Introduzione, per Esercizi, all Uso del Calcolatore in Chimica Organica e Biologica Modellistica
Molecular Mechanics, Dynamics & Docking
 Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics. C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology. January 2001
 Molecular Mechanics C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology January 2001 Introduction Molecular Mechanics uses classical type models to predict the energy
Molecular Mechanics C. David Sherrill School of Chemistry and Biochemistry Georgia Institute of Technology January 2001 Introduction Molecular Mechanics uses classical type models to predict the energy
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
3. An Introduction to Molecular Mechanics
 3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
This semester. Books
 Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
Models mostly proteins from detailed to more abstract models Some simulation methods This semester Books None necessary for my group and Prof Rarey Molecular Modelling: Principles and Applications Leach,
Why Is Molecular Interaction Important in Our Life
 Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
An Introduction to Quantum Chemistry and Potential Energy Surfaces. Benjamin G. Levine
 An Introduction to Quantum Chemistry and Potential Energy Surfaces Benjamin G. Levine This Week s Lecture Potential energy surfaces What are they? What are they good for? How do we use them to solve chemical
An Introduction to Quantum Chemistry and Potential Energy Surfaces Benjamin G. Levine This Week s Lecture Potential energy surfaces What are they? What are they good for? How do we use them to solve chemical
3. An Introduction to Molecular Mechanics
 3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
3. An Introduction to Molecular Mechanics Introduction When you use Chem3D to draw molecules, the program assigns bond lengths and bond angles based on experimental data. The program does not contain real
Biomolecular modeling I
 2016, December 6 Biomolecular structure Structural elements of life Biomolecules proteins, nucleic acids, lipids, carbohydrates... Biomolecular structure Biomolecules biomolecular complexes aggregates...
2016, December 6 Biomolecular structure Structural elements of life Biomolecules proteins, nucleic acids, lipids, carbohydrates... Biomolecular structure Biomolecules biomolecular complexes aggregates...
An introduction to Molecular Dynamics. EMBO, June 2016
 An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
Hyeyoung Shin a, Tod A. Pascal ab, William A. Goddard III abc*, and Hyungjun Kim a* Korea
 The Scaled Effective Solvent Method for Predicting the Equilibrium Ensemble of Structures with Analysis of Thermodynamic Properties of Amorphous Polyethylene Glycol-Water Mixtures Hyeyoung Shin a, Tod
The Scaled Effective Solvent Method for Predicting the Equilibrium Ensemble of Structures with Analysis of Thermodynamic Properties of Amorphous Polyethylene Glycol-Water Mixtures Hyeyoung Shin a, Tod
Colloid Chemistry. La chimica moderna e la sua comunicazione Silvia Gross.
 Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Molecular Mechanics / ReaxFF
 Molecular Dynamics simulations Lecture 09: Molecular Mechanics / ReaxFF Dr. Olli Pakarinen University of Helsinki Fall 2012 Lecture notes based on notes by Dr. Jani Kotakoski, 2010 CONTENTS Molecular mechanics
Molecular Dynamics simulations Lecture 09: Molecular Mechanics / ReaxFF Dr. Olli Pakarinen University of Helsinki Fall 2012 Lecture notes based on notes by Dr. Jani Kotakoski, 2010 CONTENTS Molecular mechanics
Solutions to Assignment #4 Getting Started with HyperChem
 Solutions to Assignment #4 Getting Started with HyperChem 1. This first exercise is meant to familiarize you with the different methods for visualizing molecules available in HyperChem. (a) Create a molecule
Solutions to Assignment #4 Getting Started with HyperChem 1. This first exercise is meant to familiarize you with the different methods for visualizing molecules available in HyperChem. (a) Create a molecule
Water models in classical simulations
 Water models in classical simulations Maria Fyta Institut für Computerphysik, Universität Stuttgart Stuttgart, Germany Water transparent, odorless, tasteless and ubiquitous really simple: two H atoms attached
Water models in classical simulations Maria Fyta Institut für Computerphysik, Universität Stuttgart Stuttgart, Germany Water transparent, odorless, tasteless and ubiquitous really simple: two H atoms attached
Biomolecular modeling I
 2015, December 15 Biomolecular simulation Elementary body atom Each atom x, y, z coordinates A protein is a set of coordinates. (Gromacs, A. P. Heiner) Usually one molecule/complex of interest (e.g. protein,
2015, December 15 Biomolecular simulation Elementary body atom Each atom x, y, z coordinates A protein is a set of coordinates. (Gromacs, A. P. Heiner) Usually one molecule/complex of interest (e.g. protein,
A Molecular Dynamics Simulation of a Homogeneous Organic-Inorganic Hybrid Silica Membrane
 A Molecular Dynamics Simulation of a Homogeneous Organic-Inorganic Hybrid Silica Membrane Supplementary Information: Simulation Procedure and Physical Property Analysis Simulation Procedure The molecular
A Molecular Dynamics Simulation of a Homogeneous Organic-Inorganic Hybrid Silica Membrane Supplementary Information: Simulation Procedure and Physical Property Analysis Simulation Procedure The molecular
TOPOLOGIES AND FORCE FIELD PARAMETERS FOR NITROXIDE SPIN LABELS
 TOPOLOGIES AND FORCE FIELD PARAMETERS FOR NITROXIDE SPIN LABELS 1. Introduction The force field used in these studies is the CHARMM19 (Chemistry at HARvard Molecular Mechanics) extended atom force field
TOPOLOGIES AND FORCE FIELD PARAMETERS FOR NITROXIDE SPIN LABELS 1. Introduction The force field used in these studies is the CHARMM19 (Chemistry at HARvard Molecular Mechanics) extended atom force field
Potential Energy (hyper)surface
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
Supporting Information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 207 Supporting Information Carbon Nanoscroll- Silk Crystallite Hybrid Structure with Controllable Hydration
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 207 Supporting Information Carbon Nanoscroll- Silk Crystallite Hybrid Structure with Controllable Hydration
Chimica Farmaceutica
 Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Molecular Simulation II. Classical Mechanical Treatment
 Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Molecular Simulation II Quantum Chemistry Classical Mechanics E = Ψ H Ψ ΨΨ U = E bond +E angle +E torsion +E non-bond Jeffry D. Madura Department of Chemistry & Biochemistry Center for Computational Sciences
Chapter 11 Molecular Mechanics
 Chapter 11 Molecular Mechanics Molecular Mechanics uses an analytical, differentiable, and relatively simple potential energy function, (R), for describing the interactions between a set of atoms specified
Chapter 11 Molecular Mechanics Molecular Mechanics uses an analytical, differentiable, and relatively simple potential energy function, (R), for describing the interactions between a set of atoms specified
Session 1. Introduction to Computational Chemistry. Computational (chemistry education) and/or (Computational chemistry) education
 Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Session 1 Introduction to Computational Chemistry 1 Introduction to Computational Chemistry Computational (chemistry education) and/or (Computational chemistry) education First one: Use computational tools
Force Fields for MD simulations
 Force Fields for MD simulations Topology/parameter files Where do the numbers an MD code uses come from? ow to make topology files for ligands, cofactors, special amino acids, ow to obtain/develop missing
Force Fields for MD simulations Topology/parameter files Where do the numbers an MD code uses come from? ow to make topology files for ligands, cofactors, special amino acids, ow to obtain/develop missing
Molecular Dynamics Simulations. Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia
 Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Force fields, thermo- and barostats. Berk Hess
 Force fields, thermo- and barostats Berk Hess What is a force field? A force field usually consists of three parts: a set of functional forms parameters for the functional forms that, usually, depend on
Force fields, thermo- and barostats Berk Hess What is a force field? A force field usually consists of three parts: a set of functional forms parameters for the functional forms that, usually, depend on
CHEMISTRY 4021/8021 MIDTERM EXAM 1 SPRING 2014
 CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
CHEMISTRY 4021/8021 Q1) Propose a simple, united-atom molecular mechanics force-field needed to generate a potential energy surface for an isolated molecule of acetone (Me 2 CO). I.e., provide an energy
Biomolecules are dynamic no single structure is a perfect model
 Molecular Dynamics Simulations of Biomolecules References: A. R. Leach Molecular Modeling Principles and Applications Prentice Hall, 2001. M. P. Allen and D. J. Tildesley "Computer Simulation of Liquids",
Molecular Dynamics Simulations of Biomolecules References: A. R. Leach Molecular Modeling Principles and Applications Prentice Hall, 2001. M. P. Allen and D. J. Tildesley "Computer Simulation of Liquids",
Energy functions and their relationship to molecular conformation. CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror
 Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Outline Energy functions for proteins (or biomolecular systems more generally)
Energy functions and their relationship to molecular conformation CS/CME/BioE/Biophys/BMI 279 Oct. 3 and 5, 2017 Ron Dror Outline Energy functions for proteins (or biomolecular systems more generally)
Soot - Developing anisotropic potentials from first principles for PAH molecules. Tim Totton, Alston Misquitta and Markus Kraft 12/11/2009
 Soot - Developing anisotropic potentials from first principles for PAH molecules. Tim Totton, Alston Misquitta and 12/11/2009 HRTEM images of soot Some evidence for different soot structures based on different
Soot - Developing anisotropic potentials from first principles for PAH molecules. Tim Totton, Alston Misquitta and 12/11/2009 HRTEM images of soot Some evidence for different soot structures based on different
Computational Methods. Chem 561
 Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Computational Methods Chem 561 Lecture Outline 1. Ab initio methods a) HF SCF b) Post-HF methods 2. Density Functional Theory 3. Semiempirical methods 4. Molecular Mechanics Computational Chemistry " Computational
Molecular Mechanics. I. Quantum mechanical treatment of molecular systems
 Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Adsorption of gases on solids (focus on physisorption)
 Adsorption of gases on solids (focus on physisorption) Adsorption Solid surfaces show strong affinity towards gas molecules that it comes in contact with and some amt of them are trapped on the surface
Adsorption of gases on solids (focus on physisorption) Adsorption Solid surfaces show strong affinity towards gas molecules that it comes in contact with and some amt of them are trapped on the surface
Molecular Modelling for Medicinal Chemistry (F13MMM) Room A36
 Molecular Modelling for Medicinal Chemistry (F13MMM) jonathan.hirst@nottingham.ac.uk Room A36 http://comp.chem.nottingham.ac.uk Assisted reading Molecular Modelling: Principles and Applications. Andrew
Molecular Modelling for Medicinal Chemistry (F13MMM) jonathan.hirst@nottingham.ac.uk Room A36 http://comp.chem.nottingham.ac.uk Assisted reading Molecular Modelling: Principles and Applications. Andrew
Computational Molecular Biophysics. Computational Biophysics, GRS Jülich SS 2013
 Computational Molecular Biophysics Computational Biophysics, GRS Jülich SS 2013 Computational Considerations Number of terms for different energy contributions / E el ~N ~N ~N ~N 2 Computational cost for
Computational Molecular Biophysics Computational Biophysics, GRS Jülich SS 2013 Computational Considerations Number of terms for different energy contributions / E el ~N ~N ~N ~N 2 Computational cost for
Molecular Dynamics. A very brief introduction
 Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Molecular Dynamics A very brief introduction Sander Pronk Dept. of Theoretical Physics KTH Royal Institute of Technology & Science For Life Laboratory Stockholm, Sweden Why computer simulations? Two primary
Advanced Molecular Molecular Dynamics
 Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Advanced Molecular Molecular Dynamics Technical details May 12, 2014 Integration of harmonic oscillator r m period = 2 k k and the temperature T determine the sampling of x (here T is related with v 0
Homework Problem Set 1 Solutions
 Chemistry 380.37 Dr. Jean M. Standard omework Problem Set 1 Solutions 1. A student investigates a bond between atoms A and B in a molecule using a software package for molecular mechanics. The student
Chemistry 380.37 Dr. Jean M. Standard omework Problem Set 1 Solutions 1. A student investigates a bond between atoms A and B in a molecule using a software package for molecular mechanics. The student
Computer simulation of Biological Systems
 UNIVERSITY OF TRENTO Faculty of Mathematical, Physical and Natural Sciences Physics Department Ph.D. Thesis in Physics Computer simulation of Biological Systems Supervisors: Dr. Maurizio Dapor Dr. Giovanni
UNIVERSITY OF TRENTO Faculty of Mathematical, Physical and Natural Sciences Physics Department Ph.D. Thesis in Physics Computer simulation of Biological Systems Supervisors: Dr. Maurizio Dapor Dr. Giovanni
Atoms can form stable units called molecules by sharing electrons.
 Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule) e.g. ionic, covalent. Forces that cause the aggregation
Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule) e.g. ionic, covalent. Forces that cause the aggregation
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
An Informal AMBER Small Molecule Force Field :
 An Informal AMBER Small Molecule Force Field : parm@frosst Credit Christopher Bayly (1992-2010) initiated, contributed and lead the efforts Daniel McKay (1997-2010) and Jean-François Truchon (2002-2010)
An Informal AMBER Small Molecule Force Field : parm@frosst Credit Christopher Bayly (1992-2010) initiated, contributed and lead the efforts Daniel McKay (1997-2010) and Jean-François Truchon (2002-2010)
Chapter 3. Crystal Binding
 Chapter 3. Crystal Binding Energy of a crystal and crystal binding Cohesive energy of Molecular crystals Ionic crystals Metallic crystals Elasticity What causes matter to exist in three different forms?
Chapter 3. Crystal Binding Energy of a crystal and crystal binding Cohesive energy of Molecular crystals Ionic crystals Metallic crystals Elasticity What causes matter to exist in three different forms?
Homology modeling. Dinesh Gupta ICGEB, New Delhi 1/27/2010 5:59 PM
 Homology modeling Dinesh Gupta ICGEB, New Delhi Protein structure prediction Methods: Homology (comparative) modelling Threading Ab-initio Protein Homology modeling Homology modeling is an extrapolation
Homology modeling Dinesh Gupta ICGEB, New Delhi Protein structure prediction Methods: Homology (comparative) modelling Threading Ab-initio Protein Homology modeling Homology modeling is an extrapolation
THEORY OF MOLECULE. A molecule consists of two or more atoms with certain distances between them
 THEORY OF MOLECULE A molecule consists of two or more atoms with certain distances between them through interaction of outer electrons. Distances are determined by sum of all forces between the atoms.
THEORY OF MOLECULE A molecule consists of two or more atoms with certain distances between them through interaction of outer electrons. Distances are determined by sum of all forces between the atoms.
Intermolecular and Intramolecular Forces. Introduction
 Intermolecular and Intramolecular Forces Introduction Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule)
Intermolecular and Intramolecular Forces Introduction Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule)
Specific Ion Solvtion in Ethylene Carbonate and Propylene Carbonate
 Specific Ion Solvtion in Ethylene Carbonate and Propylene Carbonate A. Arslanargin, A. Powers, S. Rick, T. Pollard, T. Beck Univ Cincinnati Chemistry Support: NSF, OSC TSRC 2016 November 2, 2016 A. Arslanargin,
Specific Ion Solvtion in Ethylene Carbonate and Propylene Carbonate A. Arslanargin, A. Powers, S. Rick, T. Pollard, T. Beck Univ Cincinnati Chemistry Support: NSF, OSC TSRC 2016 November 2, 2016 A. Arslanargin,
Molecular Modelling. part of Bioinformatik von RNA- und Proteinstrukturen. Sonja Prohaska. Leipzig, SS Computational EvoDevo University Leipzig
 part of Bioinformatik von RNA- und Proteinstrukturen Computational EvoDevo University Leipzig Leipzig, SS 2011 Protein Structure levels or organization Primary structure: sequence of amino acids (from
part of Bioinformatik von RNA- und Proteinstrukturen Computational EvoDevo University Leipzig Leipzig, SS 2011 Protein Structure levels or organization Primary structure: sequence of amino acids (from
5 Classical Molecular Dynamics (MD) Simulations with Tinker
 5 Classical Molecular Dynamics (MD) Simulations with Tinker In this set of exercises, you will be briey introduced to the concept of classical force elds and molecular dynamics via propagation of atomic
5 Classical Molecular Dynamics (MD) Simulations with Tinker In this set of exercises, you will be briey introduced to the concept of classical force elds and molecular dynamics via propagation of atomic
Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations
 Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations Alexandre V. Morozov, Tanja Kortemme, Kiril Tsemekhman, David Baker
Close agreement between the orientation dependence of hydrogen bonds observed in protein structures and quantum mechanical calculations Alexandre V. Morozov, Tanja Kortemme, Kiril Tsemekhman, David Baker
Molecular Dynamics, Monte Carlo and Docking. Lecture 21. Introduction to Bioinformatics MNW2
 Molecular Dynamics, Monte Carlo and Docking Lecture 21 Introduction to Bioinformatics MNW2 If you throw up a stone, it is Physics. If you throw up a stone, it is Physics. If it lands on your head, it is
Molecular Dynamics, Monte Carlo and Docking Lecture 21 Introduction to Bioinformatics MNW2 If you throw up a stone, it is Physics. If you throw up a stone, it is Physics. If it lands on your head, it is
Force Fields for Classical Molecular Dynamics simulations of Biomolecules. Emad Tajkhorshid
 Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Beckman Institute Departments of Biochemistry Center for Biophysics and Computational Biology University of Illinois
Force Fields for Classical Molecular Dynamics simulations of Biomolecules Emad Tajkhorshid Beckman Institute Departments of Biochemistry Center for Biophysics and Computational Biology University of Illinois
74 these states cannot be reliably obtained from experiments. In addition, the barriers between the local minima can also not be obtained reliably fro
 73 Chapter 5 Development of Adiabatic Force Field for Polyvinyl Chloride (PVC) and Chlorinated PVC (CPVC) 5.1 Introduction Chlorinated polyvinyl chloride has become an important specialty polymer due to
73 Chapter 5 Development of Adiabatic Force Field for Polyvinyl Chloride (PVC) and Chlorinated PVC (CPVC) 5.1 Introduction Chlorinated polyvinyl chloride has become an important specialty polymer due to
Computer simulation methods (2) Dr. Vania Calandrini
 Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Computer simulation methods (2) Dr. Vania Calandrini in the previous lecture: time average versus ensemble average MC versus MD simulations equipartition theorem (=> computing T) virial theorem (=> computing
Computational Modeling of Protein-Ligand Interactions
 Computational Modeling of Protein-Ligand Interactions Steven R. Gwaltney Department of Chemistry Mississippi State University Mississippi State, MS 39762 Auguste Comte, 1830 Every attempt to refer chemical
Computational Modeling of Protein-Ligand Interactions Steven R. Gwaltney Department of Chemistry Mississippi State University Mississippi State, MS 39762 Auguste Comte, 1830 Every attempt to refer chemical
Quantum mechanics can be used to calculate any property of a molecule. The energy E of a wavefunction Ψ evaluated for the Hamiltonian H is,
 Chapter : Molecules Quantum mechanics can be used to calculate any property of a molecule The energy E of a wavefunction Ψ evaluated for the Hamiltonian H is, E = Ψ H Ψ Ψ Ψ 1) At first this seems like
Chapter : Molecules Quantum mechanics can be used to calculate any property of a molecule The energy E of a wavefunction Ψ evaluated for the Hamiltonian H is, E = Ψ H Ψ Ψ Ψ 1) At first this seems like
Lecture 1. Conformational Analysis in Acyclic Systems
 Lecture 1 Conformational Analysis in Acyclic Systems Learning Outcomes: by the end of this lecture and after answering the associated problems, you will be able to: 1. use Newman and saw-horse projections
Lecture 1 Conformational Analysis in Acyclic Systems Learning Outcomes: by the end of this lecture and after answering the associated problems, you will be able to: 1. use Newman and saw-horse projections
Imperfect Gases. NC State University
 Chemistry 431 Lecture 3 Imperfect Gases NC State University The Compression Factor One way to represent the relationship between ideal and real gases is to plot the deviation from ideality as the gas is
Chemistry 431 Lecture 3 Imperfect Gases NC State University The Compression Factor One way to represent the relationship between ideal and real gases is to plot the deviation from ideality as the gas is
Gromacs Workshop Spring CSC
 Gromacs Workshop Spring 2007 @ CSC Erik Lindahl Center for Biomembrane Research Stockholm University, Sweden David van der Spoel Dept. Cell & Molecular Biology Uppsala University, Sweden Berk Hess Max-Planck-Institut
Gromacs Workshop Spring 2007 @ CSC Erik Lindahl Center for Biomembrane Research Stockholm University, Sweden David van der Spoel Dept. Cell & Molecular Biology Uppsala University, Sweden Berk Hess Max-Planck-Institut
Atomic Structure and Bonding. Chapter 1 Organic Chemistry, 8 th Edition John McMurry
 Atomic Structure and Bonding Chapter 1 Organic Chemistry, 8 th Edition John McMurry 1 Common Elements Groups First row Second row In most organic molecules carbon is combined with relatively few elements
Atomic Structure and Bonding Chapter 1 Organic Chemistry, 8 th Edition John McMurry 1 Common Elements Groups First row Second row In most organic molecules carbon is combined with relatively few elements
Atomic and Molecular Dimensions
 1 Atomic and Molecular Dimensions Equilibrium Interatomic Distances When two atoms approach each other, their positively charged nuclei and negatively charged electronic clouds interact. The total interaction
1 Atomic and Molecular Dimensions Equilibrium Interatomic Distances When two atoms approach each other, their positively charged nuclei and negatively charged electronic clouds interact. The total interaction
PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS
 TASKQUARTERLYvol.20,No4,2016,pp.353 360 PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS MARTIN ZACHARIAS Physics Department T38, Technical University of Munich James-Franck-Str.
TASKQUARTERLYvol.20,No4,2016,pp.353 360 PROTEIN-PROTEIN DOCKING REFINEMENT USING RESTRAINT MOLECULAR DYNAMICS SIMULATIONS MARTIN ZACHARIAS Physics Department T38, Technical University of Munich James-Franck-Str.
Analysis of the simulation
 Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Analysis of the simulation Marcus Elstner and Tomáš Kubař January 7, 2014 Thermodynamic properties time averages of thermodynamic quantites correspond to ensemble averages (ergodic theorem) some quantities
Computational Biology & Computational Medicine
 Computational Biology & Computational Medicine Homayoun Valafar Outline Why proteins? What are proteins? How do we compute them? How do we use computational approaches? Why Proteins? Molecular basis of
Computational Biology & Computational Medicine Homayoun Valafar Outline Why proteins? What are proteins? How do we compute them? How do we use computational approaches? Why Proteins? Molecular basis of
Fondamenti di Chimica Farmaceutica. Computer Chemistry in Drug Research: Introduction
 Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Fondamenti di Chimica Farmaceutica Computer Chemistry in Drug Research: Introduction Introduction Introduction Introduction Computer Chemistry in Drug Design Drug Discovery: Target identification Lead
Chem 1140; Molecular Modeling
 P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
P. Wipf 1 Chem 1140 $E = -! (q a + q b )" ab S ab + ab!k
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Aqueous solutions. Solubility of different compounds in water
 Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Applications of Molecular Dynamics
 June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
June 4, 0 Molecular Modeling and Simulation Applications of Molecular Dynamics Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, The University of Tokyo Tohru
Scientific Computing II
 Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
Scientific Computing II Molecular Dynamics Simulation Michael Bader SCCS Summer Term 2015 Molecular Dynamics Simulation, Summer Term 2015 1 Continuum Mechanics for Fluid Mechanics? Molecular Dynamics the
2 Structure. 2.1 Coulomb interactions
 2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
General Physical Chemistry II
 General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model. John Dood Hope College
 Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
Developing Monovalent Ion Parameters for the Optimal Point Charge (OPC) Water Model John Dood Hope College What are MD simulations? Model and predict the structure and dynamics of large macromolecules.
CHEM-UA 127: Advanced General Chemistry I
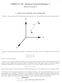 1 CHEM-UA 127: Advanced General Chemistry I Notes for Lecture 6 I MOLECULAR GEOMETRY AND COORDINATES Consider a diatomic molecule AB Imagine fixing this molecule at a very specific spatial location, as
1 CHEM-UA 127: Advanced General Chemistry I Notes for Lecture 6 I MOLECULAR GEOMETRY AND COORDINATES Consider a diatomic molecule AB Imagine fixing this molecule at a very specific spatial location, as
Interatomic Potentials. The electronic-structure problem
 Interatomic Potentials Before we can start a simulation, we need the model! Interaction between atoms and molecules is determined by quantum mechanics: Schrödinger Equation + Born-Oppenheimer approximation
Interatomic Potentials Before we can start a simulation, we need the model! Interaction between atoms and molecules is determined by quantum mechanics: Schrödinger Equation + Born-Oppenheimer approximation
Lecture 2: Series Expansion
 Lecture 2: Series Expansion Key points Maclaurin series : For, Taylor expansion: Maple commands series convert taylor, Student[NumericalAnalysis][Taylor] 1 Maclaurin series of elementary functions for
Lecture 2: Series Expansion Key points Maclaurin series : For, Taylor expansion: Maple commands series convert taylor, Student[NumericalAnalysis][Taylor] 1 Maclaurin series of elementary functions for
Supplementary Information
 Supplementary Information Ballistic Thermal Transport in Carbyne and Cumulene with Micron-Scale Spectral Acoustic Phonon Mean Free Path Mingchao Wang and Shangchao Lin * Department of Mechanical Engineering,
Supplementary Information Ballistic Thermal Transport in Carbyne and Cumulene with Micron-Scale Spectral Acoustic Phonon Mean Free Path Mingchao Wang and Shangchao Lin * Department of Mechanical Engineering,
