K ex. Conformational equilibrium. equilibrium K B
|
|
|
- Louise Powell
- 6 years ago
- Views:
Transcription
1
2 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar coupling, or relaxation) differ. DNMR deals with the effects in a broad sense of chemical exchange processes on NMR spectra; and conversely with the information about the changes in the environment of magnetic nuclei that can be derived from observation of NMR spectra. K ex Conformational equilibrium K B Chemical equilibrium
3 Types of Chemical Exchange Intramolecular exchange Motions of sidechains in proteins A B Helix-coil transitions of nucleic acids Unfolding of proteins Conformational equilibria Intermolecular exchange Binding of small molecules l to macromolecules Protonation/deprotonation equilibria Isotope exchange processes Enzyme catalyzed reactions M+L ML Because NMR detects the molecular motion itself, rather the numbers of molecules in different states, NMR is able to detect chemical exchange even when the system is in equilibrium
4 2-state First Order Exchange A B k1 k -1 Lifetime of state A: τ A = 1/k +1 Lifetime of state B: τ B = 1/k -1 Use a single lifetime 1/ τ =1/ τ A + 1/τ B = k +1 + k -1
5 Rationale for Chemical Exchange For slow exchange For fast exchange FT FT Bloch equation approach: dm AX /dt = -(Δω A )M AY -M AX /τ A + M BX /τ B dm BX /dt = -(Δω B )M BY -M BX /τ B + M AX /τ A
6 2-state 2nd Order Exchange M+L k +1 M+L ML K d = [M] [L]/[ML] = k -1 /k +1 k -1 K -3-9 d =10 10 M k on = k +1 ~ 10 8 M -1 s -1 (diffusion-limited) k ~ s Lifetime 1/ τ =1/ τ ML + 1/τ l = k -1 (1+f ML /f L ) f ML and f L are the mole fractions of bound and free ligand, ML L g respectively
7 Typical Motion Time Scale for Physical Processes SLOW very slow slow fast very fast ultrafast s ms μs ns ps fs FAST MACROSCOPIC DIFFUSION, FLOW CHEMICAL EXCHANGE MOLECULAR ROTATIONS RELAXATION SPECTRAL LARMOR TIMESCALE TIMESCALE TIMESCALE MOLECULAR VIBRATIONS
8 NMR Time Scale Time Scale Chem. Shift, δ Coupling Const., J T2 relaxation Slow k << δ Α δ Β k << J Α J Β k << 1/ T 2, Α 1/ T 2, B Intermediate k = δ Α δ Β k = J Α J Β k = 1/ T 2, Α 1/ T 2, B Fast k >> δ Α δ Β k>>j Α J Β k>>1/t 2, Α 1/ T 2, B Sec NMR time-scale refers to the chemical shift timescale. The range of the rate can be studied s -1 for H can be extended to faster rate using 19 F, 13 C and etc.
9 k1 Slow Exchange k << δ -δ A B A δ B k -1 Separate lines are observed for each state. The exchange rate can be readily measured from the line widths of the resonances Like the apparent spin-spin relaxation rates, 1/T 2i,obs 1/T 2A,obs = 1/T 2A +1/ 1/τ A = 1/T 2A +1/k 1 1/T 2B,obs = 1/T 2B + 1/τ Β = 1/T 2B + 1/k -1 line width Lw = 1/πT 2 = 1/πT 2 +k 1 /π Each resonance is broadened by Δ Lw = k/π Increasing temperature increases k, line width increases
10 Slow Exchange for M+L ML k 1 k- 1 Separate resonances potentially are observable for both the free and bound states M F,M B,L F, and L B The addition of a ligand to a solution of a protein can be used to determine the stoichiometry of the complex. Once a stoichiometric mole ratio is achieved, peaks from free ligand appear with increasing i intensity it as the excess of free ligand increases. Obtain spectra over a range of [L]/[M] ratios from 1 to 10
11 Slow Exchange for M+L ML k 1 k- 1 For free form 1/T 2L,obs = 1/T 2L + 1/τ L = 1/T 2L + k -1 f ML /f L 1/T 2M,obs = 1/T 2M + 1/τ Μ = 1/T 2M + k -1 f ML /f M For complex form 1/T 2ML,obs = 1/T 2ML + 1/τ ML = 1/T 2ML + k -1 Measurements of line width during a titration can be used to derive k -1 (k off ).
12 19 F spectra of the enzyme-inhibitor complex at various mole ratio of carbonic anhydrase:inhibitor Free inhibitor 1:4 1:3 1:2 1:1 1:0.5 Bound ligand At -6 ppm the broadened peak for the bound ligand is in slow exchange with the peak from free ligand at 0 ppm. The stoichiometry t of the complex is 2:1. No signal from the free ligand is visible until more than 2 moles of inhibitor are present.
13 Coalescence Rate For A B equal concentrations, there will be a rate of interchange where the separate lines for two species are no longer discernible The coalescence rate k c = π Δδ / 2 = 2.22 Δδ Δδ is the chemical shift difference between the two signals in the unit of Hz. Δδ depends on the magnetic field
14 Coalescence Temperature Since the rate depends on the ΔG of the inversion, and the ΔG is affected by T, higher temperature will make things go faster. Tc is the temperature t at which fast and slow exchange meet. T>Tc, fast exchange T<Tc, slow exchange T we can calculate the ΔG of the process ΔG = R * T C * [ ln ( T C / Δδ ) ] T C
15 Fast Exchange k >> δ A -δ B k1 A B k -1 A single resonance is observed, whose chemical shift is the weight average of the chemical shifts of the two individual states δ obs = f A δ A +f B δ B, f A + f B = 1 For very fast limit 1/T 2,obs = f A /T 2A + f B /T 2B For moderately fast 1/T 2,obs = f A /T 2A + f B /T 2B + f A f B2 4π (Δδ AB ) 2 / k -1 Maximal line broadening is observed when f A = f B = 0.5
16 Fast Exchange k >> δ A -δδ B k+1 M+L ML k-1 For M δ M,obs = f M δ M +f ML δ ML For L δ L.obs = f L δ L +f ML δ ML 1/T 2,obs = f ML /T 2ML 2,ML + f L /T 2L 2,L + f ML f L2 4π (δ ML -δ L ) 2 / k -1 A maximum in the line broadening of ligand or protein resonances occurs during the titration at a mole ratio of approx. ligand:protein 1:3 The dissociation i constant t for the complex can be obtained by measuring the chemical shift of the ligand resonance at a series of [L].
17
18
19
20
21
22
23
24 Diffusion by Pulse Field Gradient Price et al. Concepts Magn. Reson. (1997) 9,
25 Pulse Sequence Δ I = I o exp[-(γδg) 2 (Δ + 2τ 2δ/3)D] D = kt/f γ: gyromagnetic ratio f = 6πηr D: diffusion i constant t k: Boltzman constant t Δ: interval between gradient pulse f: friction coefficient δ: PFG duration time η: solvent viscosity G: gradient strength r: hydrodynamic radius
26 N15 Edited Chou et al. J. Biomol. NMR (2004) 29,
27 Calibration of Gradient Strength 500 MHz 600 MHz y = exp(-m1* M0^2) Value Error m e e-10 1 Chisq e-05 NA 1 R NA 0.8 y = exp(-m1* M0^2) Value Error 0.8 m e e-10 Chisq NA 0.6 R NA 0.6 m1 Chisq R y = exp(-m1 * M0^2) Value Error 1.465e e NA NA gradient strength gradient strength Gzlvl1=32768 = 53.2 G/cm = 64.6 G/cm
28 Diffusion Constant is Related to Molecular Weight diameter Lee et al. Biochimica et Biophysica Acta (2002) 1598,80 87
29 Diffusion Dependent on Conformation ApoCaM r = 22.4 ± 0.3 Å NMR: 22 Å 1.2 m1 Chisq R y = exp(-m1 * M0^2) Value Error e-06 NA NA m1 Chisq R y = exp(-m1 * M0^2) Value Error e-06 NA NA Ca 2+ -CaM r = 22.8 ± 0.5 Å apocam Ca-CaM X-ray: 23 Å gradient strength
30 Binding Constant Determination D obs = X L D L + X PL D PL X PL = (D L D obs )/(D L D PL ) K a = X PL /((1 X PL )([P] 0 X PL [L] 0 )) 0.5 mm cyclohexylacetic acid in D 2 O; D = 6.85 x10 6 cm 2 s mm cyclohexylacetic acid plus 0.5 mm β-cyclodextrin in D 2 O; D = 5.39 x10 6 cm 2 s -1 K a = 1800 ± 100 M -1 Cameron et al. J. Org. Chem. (2001) 66,
31
32 Limitations The transverse relaxation time constant T 2 limits the interval where diffusion i can be observed. Small diffusion i coefficients i (D < m 2 s -1 ), associated with macromolecules with masses larger than 50 kda are difficult to measure For weak binding, the change in diffusion constant is too small, and it is difficult to get the binding constant Singlet State Diffusion Spectroscopy stores the nuclear spin order as a singlet state. This state relaxes with a time constant T s that can be much longer than both T 1 and T 2 Further reading: Cavadini et al. Concepts Magn. Reson. (2008) 32A,
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Chemical Exchange and Ligand Binding
 Chemical Exchange and Ligand Binding NMR time scale Fast exchange for binding constants Slow exchange for tight binding Single vs. multiple binding mode Calcium binding process of calcium binding proteins
Chemical Exchange and Ligand Binding NMR time scale Fast exchange for binding constants Slow exchange for tight binding Single vs. multiple binding mode Calcium binding process of calcium binding proteins
Georgia State University
 Chemical Exchange and Calcium Signaling Jenny J. Yang Jenny@gsu.edu Chemistry Department Center for Diagnostics & Therapeutics Georgia State University http://www.gsuyanglab.com/research Ligand Protein
Chemical Exchange and Calcium Signaling Jenny J. Yang Jenny@gsu.edu Chemistry Department Center for Diagnostics & Therapeutics Georgia State University http://www.gsuyanglab.com/research Ligand Protein
Slow symmetric exchange
 Slow symmetric exchange ϕ A k k B t A B There are three things you should notice compared with the Figure on the previous slide: 1) The lines are broader, 2) the intensities are reduced and 3) the peaks
Slow symmetric exchange ϕ A k k B t A B There are three things you should notice compared with the Figure on the previous slide: 1) The lines are broader, 2) the intensities are reduced and 3) the peaks
Spin Relaxation and NOEs BCMB/CHEM 8190
 Spin Relaxation and NOEs BCMB/CHEM 8190 T 1, T 2 (reminder), NOE T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations
Spin Relaxation and NOEs BCMB/CHEM 8190 T 1, T 2 (reminder), NOE T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations
Introduction to Relaxation Theory James Keeler
 EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
Protein dynamics from NMR Relaxation data
 Protein dynamics from NMR Relaxation data Clubb 3/15/17 (S f2 ) ( e ) Nitrogen-15 relaxation ZZ-exchange R 1 = 1/T 1 Longitudinal relaxation (decay back to z-axis) R 2 = 1/T 2 Spin-spin relaxation (dephasing
Protein dynamics from NMR Relaxation data Clubb 3/15/17 (S f2 ) ( e ) Nitrogen-15 relaxation ZZ-exchange R 1 = 1/T 1 Longitudinal relaxation (decay back to z-axis) R 2 = 1/T 2 Spin-spin relaxation (dephasing
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Relaxation. Ravinder Reddy
 Relaxation Ravinder Reddy Relaxation What is nuclear spin relaxation? What causes it? Effect on spectral line width Field dependence Mechanisms Thermal equilibrium ~10-6 spins leads to NMR signal! T1 Spin-lattice
Relaxation Ravinder Reddy Relaxation What is nuclear spin relaxation? What causes it? Effect on spectral line width Field dependence Mechanisms Thermal equilibrium ~10-6 spins leads to NMR signal! T1 Spin-lattice
NMR in Structural Biology
 NMR in Structural Biology Exercise session 2 1. a. List 3 NMR observables that report on structure. b. Also indicate whether the information they give is short/medium or long-range, or perhaps all three?
NMR in Structural Biology Exercise session 2 1. a. List 3 NMR observables that report on structure. b. Also indicate whether the information they give is short/medium or long-range, or perhaps all three?
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Biophysical Chemistry: NMR Spectroscopy
 Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
Biochemistry 530 NMR Theory and Practice
 Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington Lecturer: Gabriele Varani Biochemistry and Chemistry Room J479 and
Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington Lecturer: Gabriele Varani Biochemistry and Chemistry Room J479 and
Chemistry 431. Lecture 23
 Chemistry 431 Lecture 23 Introduction The Larmor Frequency The Bloch Equations Measuring T 1 : Inversion Recovery Measuring T 2 : the Spin Echo NC State University NMR spectroscopy The Nuclear Magnetic
Chemistry 431 Lecture 23 Introduction The Larmor Frequency The Bloch Equations Measuring T 1 : Inversion Recovery Measuring T 2 : the Spin Echo NC State University NMR spectroscopy The Nuclear Magnetic
Chemical Exchange. Spin-interactions External interactions Magnetic field Bo, RF field B1
 Chemical Exchange Spin-interactions External interactions Magnetic field Bo, RF field B1 Internal Interactions Molecular motions Chemical shifts J-coupling Chemical Exchange 1 Outline Motional time scales
Chemical Exchange Spin-interactions External interactions Magnetic field Bo, RF field B1 Internal Interactions Molecular motions Chemical shifts J-coupling Chemical Exchange 1 Outline Motional time scales
Lecture #6 Chemical Exchange
 Lecture #6 Chemical Exchange Topics Introduction Effects on longitudinal magnetization Effects on transverse magnetization Examples Handouts and Reading assignments Kowalewski, Chapter 13 Levitt, sections
Lecture #6 Chemical Exchange Topics Introduction Effects on longitudinal magnetization Effects on transverse magnetization Examples Handouts and Reading assignments Kowalewski, Chapter 13 Levitt, sections
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
T 1, T 2, NOE (reminder)
 T 1, T 2, NOE (reminder) T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations of the system following perturbation
T 1, T 2, NOE (reminder) T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations of the system following perturbation
Biophysical Chemistry: NMR Spectroscopy
 Spin Dynamics & Vrije Universiteit Brussel 25th November 2011 Outline 1 Pulse/Fourier Transform NMR Thermal Equilibrium Effect of RF Pulses The Fourier Transform 2 Symmetric Exchange Between Two Sites
Spin Dynamics & Vrije Universiteit Brussel 25th November 2011 Outline 1 Pulse/Fourier Transform NMR Thermal Equilibrium Effect of RF Pulses The Fourier Transform 2 Symmetric Exchange Between Two Sites
Introduction solution NMR
 2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
Filtered/edited NOESY spectra
 Filtered/edited NOESY spectra NMR Seminar HS 207 Nina Ripin 22..7 Overview NMR of biomolecular complexes Problems and Solutions Filtered/edited nomenclature Experimental elements NOESY vs filtered pulse
Filtered/edited NOESY spectra NMR Seminar HS 207 Nina Ripin 22..7 Overview NMR of biomolecular complexes Problems and Solutions Filtered/edited nomenclature Experimental elements NOESY vs filtered pulse
Measuring Spin-Lattice Relaxation Time
 WJP, PHY381 (2009) Wabash Journal of Physics v4.0, p.1 Measuring Spin-Lattice Relaxation Time L.W. Lupinski, R. Paudel, and M.J. Madsen Department of Physics, Wabash College, Crawfordsville, IN 47933 (Dated:
WJP, PHY381 (2009) Wabash Journal of Physics v4.0, p.1 Measuring Spin-Lattice Relaxation Time L.W. Lupinski, R. Paudel, and M.J. Madsen Department of Physics, Wabash College, Crawfordsville, IN 47933 (Dated:
The Physical Basis of the NMR Experiment
 The Physical Basis of the NMR Experiment 1 Interaction of Materials with Magnetic Fields F F S N S N Paramagnetism Diamagnetism 2 Microscopic View: Single Spins an electron has mass and charge in addition
The Physical Basis of the NMR Experiment 1 Interaction of Materials with Magnetic Fields F F S N S N Paramagnetism Diamagnetism 2 Microscopic View: Single Spins an electron has mass and charge in addition
PRACTICAL ASPECTS OF NMR RELAXATION STUDIES OF BIOMOLECULAR DYNAMICS
 PRACTICAL ASPECTS OF MR RELAXATIO STUDIES OF BIOMOLECULAR DYAMICS Further reading: Can be downloaded from my web page Korzhnev D.E., Billeter M., Arseniev A.S., and Orekhov V. Y., MR Studies of Brownian
PRACTICAL ASPECTS OF MR RELAXATIO STUDIES OF BIOMOLECULAR DYAMICS Further reading: Can be downloaded from my web page Korzhnev D.E., Billeter M., Arseniev A.S., and Orekhov V. Y., MR Studies of Brownian
NMR Spectroscopy. Guangjin Hou
 NMR Spectroscopy Guangjin Hou 22-04-2009 NMR History 1 H NMR spectra of water H NMR spectra of water (First NMR Spectra on Water, 1946) 1 H NMR spectra ethanol (First bservation of the Chemical Shift,
NMR Spectroscopy Guangjin Hou 22-04-2009 NMR History 1 H NMR spectra of water H NMR spectra of water (First NMR Spectra on Water, 1946) 1 H NMR spectra ethanol (First bservation of the Chemical Shift,
SUPPLEMENTARY NOTE 1: ADDITIONAL CHARACTERIZATION OF NANODIAMOND SOLUTIONS AND THE OVERHAUSER EFFECT
 1 SUPPLEMENTARY NOTE 1: ADDITIONAL CHARACTERIZATION OF NANODIAMOND SOLUTIONS AND THE OVERHAUSER EFFECT Nanodiamond (ND) solutions were prepared using high power probe sonication and analyzed by dynamic
1 SUPPLEMENTARY NOTE 1: ADDITIONAL CHARACTERIZATION OF NANODIAMOND SOLUTIONS AND THE OVERHAUSER EFFECT Nanodiamond (ND) solutions were prepared using high power probe sonication and analyzed by dynamic
Chapter 7. Nuclear Magnetic Resonance Spectroscopy
 Chapter 7 Nuclear Magnetic Resonance Spectroscopy I. Introduction 1924, W. Pauli proposed that certain atomic nuclei have spin and magnetic moment and exposure to magnetic field would lead to energy level
Chapter 7 Nuclear Magnetic Resonance Spectroscopy I. Introduction 1924, W. Pauli proposed that certain atomic nuclei have spin and magnetic moment and exposure to magnetic field would lead to energy level
Protein-protein interactions (PPIs) via NMR. Paola Turano
 Protein-protein interactions (PPIs) via NMR Paola Turano turano@cerm.unifi.it The magnetic field at the The chemical shift nucleus (the effective field) is generally less than the applied field by a fraction
Protein-protein interactions (PPIs) via NMR Paola Turano turano@cerm.unifi.it The magnetic field at the The chemical shift nucleus (the effective field) is generally less than the applied field by a fraction
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
TITAN: Two-dimensional lineshape analysis
 TITAN: Two-dimensional lineshape analysis Chris Waudby Christodoulou Group c.waudby@ucl.ac.uk Andres Ramos Lisa Cabrita John Christodoulou Inhibition of fatty acid synthesis for treatment of tularemia
TITAN: Two-dimensional lineshape analysis Chris Waudby Christodoulou Group c.waudby@ucl.ac.uk Andres Ramos Lisa Cabrita John Christodoulou Inhibition of fatty acid synthesis for treatment of tularemia
Protein-protein interactions (PPIs) via NMR. Paola Turano
 Protein-protein interactions (PPIs) via NMR Paola Turano turano@cerm.unifi.it The magnetic field at the The chemical shift nucleus (the effective field) is generally less than the applied field by a fraction
Protein-protein interactions (PPIs) via NMR Paola Turano turano@cerm.unifi.it The magnetic field at the The chemical shift nucleus (the effective field) is generally less than the applied field by a fraction
The Basics of Magnetic Resonance Imaging
 The Basics of Magnetic Resonance Imaging Nathalie JUST, PhD nathalie.just@epfl.ch CIBM-AIT, EPFL Course 2013-2014-Chemistry 1 Course 2013-2014-Chemistry 2 MRI: Many different contrasts Proton density T1
The Basics of Magnetic Resonance Imaging Nathalie JUST, PhD nathalie.just@epfl.ch CIBM-AIT, EPFL Course 2013-2014-Chemistry 1 Course 2013-2014-Chemistry 2 MRI: Many different contrasts Proton density T1
Spin Dynamics Basics of Nuclear Magnetic Resonance. Malcolm H. Levitt
 Spin Dynamics Basics of Nuclear Magnetic Resonance Second edition Malcolm H. Levitt The University of Southampton, UK John Wiley &. Sons, Ltd Preface xxi Preface to the First Edition xxiii Introduction
Spin Dynamics Basics of Nuclear Magnetic Resonance Second edition Malcolm H. Levitt The University of Southampton, UK John Wiley &. Sons, Ltd Preface xxi Preface to the First Edition xxiii Introduction
Chem 325 NMR Intro. The Electromagnetic Spectrum. Physical properties, chemical properties, formulas Shedding real light on molecular structure:
 Physical properties, chemical properties, formulas Shedding real light on molecular structure: Wavelength Frequency ν Wavelength λ Frequency ν Velocity c = 2.998 10 8 m s -1 The Electromagnetic Spectrum
Physical properties, chemical properties, formulas Shedding real light on molecular structure: Wavelength Frequency ν Wavelength λ Frequency ν Velocity c = 2.998 10 8 m s -1 The Electromagnetic Spectrum
LineShapeKin NMR Line Shape Analysis Software for Studies of Protein-Ligand Interaction Kinetics
 LineShapeKin NMR Line Shape Analysis Software for Studies of Protein-Ligand Interaction Kinetics http://lineshapekin.net Spectral intensity Evgenii L. Kovrigin Department of Biochemistry, Medical College
LineShapeKin NMR Line Shape Analysis Software for Studies of Protein-Ligand Interaction Kinetics http://lineshapekin.net Spectral intensity Evgenii L. Kovrigin Department of Biochemistry, Medical College
Protein Dynamics, Allostery and Function
 Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
NMR Spectroscopy: A Quantum Phenomena
 NMR Spectroscopy: A Quantum Phenomena Pascale Legault Département de Biochimie Université de Montréal Outline 1) Energy Diagrams and Vector Diagrams 2) Simple 1D Spectra 3) Beyond Simple 1D Spectra 4)
NMR Spectroscopy: A Quantum Phenomena Pascale Legault Département de Biochimie Université de Montréal Outline 1) Energy Diagrams and Vector Diagrams 2) Simple 1D Spectra 3) Beyond Simple 1D Spectra 4)
Spectral Broadening Mechanisms
 Spectral Broadening Mechanisms Lorentzian broadening (Homogeneous) Gaussian broadening (Inhomogeneous, Inertial) Doppler broadening (special case for gas phase) The Fourier Transform NC State University
Spectral Broadening Mechanisms Lorentzian broadening (Homogeneous) Gaussian broadening (Inhomogeneous, Inertial) Doppler broadening (special case for gas phase) The Fourier Transform NC State University
Polarised Nucleon Targets for Europe, 2nd meeting, Bochum 2005
 Polarised Nucleon Targets for Europe, nd meeting, Bochum Temperature dependence of nuclear spin-lattice relaxations in liquid ethanol with dissolved TEMPO radicals H. Štěpánková, J. Englich, J. Kohout,
Polarised Nucleon Targets for Europe, nd meeting, Bochum Temperature dependence of nuclear spin-lattice relaxations in liquid ethanol with dissolved TEMPO radicals H. Štěpánková, J. Englich, J. Kohout,
Nuclear magnetic resonance spectroscopy II. 13 C NMR. Reading: Pavia Chapter , 6.7, 6.11, 6.13
 Nuclear magnetic resonance spectroscopy II. 13 NMR Reading: Pavia hapter 6.1-6.5, 6.7, 6.11, 6.13 1. General - more/better/additional structural information for larger compounds -problems: a) isotopes
Nuclear magnetic resonance spectroscopy II. 13 NMR Reading: Pavia hapter 6.1-6.5, 6.7, 6.11, 6.13 1. General - more/better/additional structural information for larger compounds -problems: a) isotopes
Supplementary Information. Profiling Formulated Monoclonal Antibodies by 1 H NMR Spectroscopy
 Supplementary Information Profiling Formulated Monoclonal Antibodies by 1 H NMR Spectroscopy Leszek Poppe 1 *, John B. Jordan 1, Ken Lawson 2, Matthew Jerums 2, Izydor Apostol 2 and Paul D. Schnier 1 1
Supplementary Information Profiling Formulated Monoclonal Antibodies by 1 H NMR Spectroscopy Leszek Poppe 1 *, John B. Jordan 1, Ken Lawson 2, Matthew Jerums 2, Izydor Apostol 2 and Paul D. Schnier 1 1
Magnetic Resonance Imaging. Pål Erik Goa Associate Professor in Medical Imaging Dept. of Physics
 Magnetic Resonance Imaging Pål Erik Goa Associate Professor in Medical Imaging Dept. of Physics pal.e.goa@ntnu.no 1 Why MRI? X-ray/CT: Great for bone structures and high spatial resolution Not so great
Magnetic Resonance Imaging Pål Erik Goa Associate Professor in Medical Imaging Dept. of Physics pal.e.goa@ntnu.no 1 Why MRI? X-ray/CT: Great for bone structures and high spatial resolution Not so great
NMR-spectroscopy of proteins in solution. Peter Schmieder
 NMR-spectroscopy of proteins in solution Basic aspects of NMR-Spektroskopie Basic aspects of NMR-spectroscopy 3/84 Prerequisite for NMR-spectroscopy is a nuclear spin that can be thought of as a mixture
NMR-spectroscopy of proteins in solution Basic aspects of NMR-Spektroskopie Basic aspects of NMR-spectroscopy 3/84 Prerequisite for NMR-spectroscopy is a nuclear spin that can be thought of as a mixture
Quantification of Dynamics in the Solid-State
 Bernd Reif Quantification of Dynamics in the Solid-State Technische Universität München Helmholtz-Zentrum München Biomolecular Solid-State NMR Winter School Stowe, VT January 0-5, 206 Motivation. Solid
Bernd Reif Quantification of Dynamics in the Solid-State Technische Universität München Helmholtz-Zentrum München Biomolecular Solid-State NMR Winter School Stowe, VT January 0-5, 206 Motivation. Solid
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Christopher Pavlik Bioanalytical Chemistry March 2, 2011
 Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
The NMR Inverse Imaging Problem
 The NMR Inverse Imaging Problem Nuclear Magnetic Resonance Protons and Neutrons have intrinsic angular momentum Atoms with an odd number of proton and/or odd number of neutrons have a net magnetic moment=>
The NMR Inverse Imaging Problem Nuclear Magnetic Resonance Protons and Neutrons have intrinsic angular momentum Atoms with an odd number of proton and/or odd number of neutrons have a net magnetic moment=>
Influence of Calcium-induced Aggregation on the Sensitivity of. Aminobis(methylenephosphonate)-Containing Potential MRI Contrast.
 Supporting Information for Influence of Calcium-induced Aggregation on the Sensitivity of Aminobis(methylenephosphonate)-Containing Potential MRI Contrast Agents Jörg Henig, Ilgar Mamedov, Petra Fouskova,
Supporting Information for Influence of Calcium-induced Aggregation on the Sensitivity of Aminobis(methylenephosphonate)-Containing Potential MRI Contrast Agents Jörg Henig, Ilgar Mamedov, Petra Fouskova,
NMR in Medicine and Biology
 NMR in Medicine and Biology http://en.wikipedia.org/wiki/nmr_spectroscopy MRI- Magnetic Resonance Imaging (water) In-vivo spectroscopy (metabolites) Solid-state t NMR (large structures) t Solution NMR
NMR in Medicine and Biology http://en.wikipedia.org/wiki/nmr_spectroscopy MRI- Magnetic Resonance Imaging (water) In-vivo spectroscopy (metabolites) Solid-state t NMR (large structures) t Solution NMR
Spectroscopy of Polymers
 Spectroscopy of Polymers Jack L. Koenig Case Western Reserve University WOMACS Professional Reference Book American Chemical Society, Washington, DC 1992 Contents Preface m xiii Theory of Polymer Characterization
Spectroscopy of Polymers Jack L. Koenig Case Western Reserve University WOMACS Professional Reference Book American Chemical Society, Washington, DC 1992 Contents Preface m xiii Theory of Polymer Characterization
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
CHEM / BCMB 4190/6190/8189. Introductory NMR. Lecture 10
 CHEM / BCMB 490/690/889 Introductory NMR Lecture 0 - - CHEM 490/690 Spin-Echo The spin-echo pulse sequence: 90 - τ - 80 - τ(echo) Spins echoes are widely used as part of larger pulse sequence to refocus
CHEM / BCMB 490/690/889 Introductory NMR Lecture 0 - - CHEM 490/690 Spin-Echo The spin-echo pulse sequence: 90 - τ - 80 - τ(echo) Spins echoes are widely used as part of larger pulse sequence to refocus
General NMR basics. Solid State NMR workshop 2011: An introduction to Solid State NMR spectroscopy. # nuclei
 : An introduction to Solid State NMR spectroscopy Dr. Susanne Causemann (Solid State NMR specialist/ researcher) Interaction between nuclear spins and applied magnetic fields B 0 application of a static
: An introduction to Solid State NMR spectroscopy Dr. Susanne Causemann (Solid State NMR specialist/ researcher) Interaction between nuclear spins and applied magnetic fields B 0 application of a static
Longitudinal-relaxation enhanced fast-pulsing techniques: New tools for biomolecular NMR spectroscopy
 Longitudinal-relaxation enhanced fast-pulsing techniques: New tools for biomolecular NMR spectroscopy Bernhard Brutscher Laboratoire de Résonance Magnétique Nucléaire Institut de Biologie Structurale -
Longitudinal-relaxation enhanced fast-pulsing techniques: New tools for biomolecular NMR spectroscopy Bernhard Brutscher Laboratoire de Résonance Magnétique Nucléaire Institut de Biologie Structurale -
NMR, the vector model and the relaxation
 NMR, the vector model and the relaxation Reading/Books: One and two dimensional NMR spectroscopy, VCH, Friebolin Spin Dynamics, Basics of NMR, Wiley, Levitt Molecular Quantum Mechanics, Oxford Univ. Press,
NMR, the vector model and the relaxation Reading/Books: One and two dimensional NMR spectroscopy, VCH, Friebolin Spin Dynamics, Basics of NMR, Wiley, Levitt Molecular Quantum Mechanics, Oxford Univ. Press,
Supporting Information Elucidating Lithium-Ion and Proton Dynamics in Anti- Perovskite Solid Electrolytes
 Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2018 Supporting Information Elucidating Lithium-Ion and Proton Dynamics in Anti-
Electronic Supplementary Material (ESI) for Energy & Environmental Science. This journal is The Royal Society of Chemistry 2018 Supporting Information Elucidating Lithium-Ion and Proton Dynamics in Anti-
Magnetic Resonance Spectroscopy
 INTRODUCTION TO Magnetic Resonance Spectroscopy ESR, NMR, NQR D. N. SATHYANARAYANA Formerly, Chairman Department of Inorganic and Physical Chemistry Indian Institute of Science, Bangalore % I.K. International
INTRODUCTION TO Magnetic Resonance Spectroscopy ESR, NMR, NQR D. N. SATHYANARAYANA Formerly, Chairman Department of Inorganic and Physical Chemistry Indian Institute of Science, Bangalore % I.K. International
Magnetic Nuclei other than 1 H
 Magnetic Nuclei other than 1 H 2 H (Deuterium): I = 1 H,D-Exchange might be used to simplify 1 H-NMR spectra since H-D couplings are generally small; - - - -O- - - -D 2 -O- triplet of triplets slightly
Magnetic Nuclei other than 1 H 2 H (Deuterium): I = 1 H,D-Exchange might be used to simplify 1 H-NMR spectra since H-D couplings are generally small; - - - -O- - - -D 2 -O- triplet of triplets slightly
Introduction to solution NMR. Alexandre Bonvin. The NMR research group. Bijvoet Center for Biomolecular Research
 Introduction to solution NMR 1 Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben Bente%Vestergaard% The NMR research group Prof. Marc Baldus Prof. Rolf Boelens
Introduction to solution NMR 1 Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben Bente%Vestergaard% The NMR research group Prof. Marc Baldus Prof. Rolf Boelens
Magnetisation Transfer Schemes
 Magnetisation Transfer Schemes P. K. Madhu Department of Chemical Sciences Tata Institute of Fundamental Research Homi Bhabha Road Colaba Mumbai 400 005, India Sensitivity of NMR Spectroscopy S/N Nγ exc
Magnetisation Transfer Schemes P. K. Madhu Department of Chemical Sciences Tata Institute of Fundamental Research Homi Bhabha Road Colaba Mumbai 400 005, India Sensitivity of NMR Spectroscopy S/N Nγ exc
Topics. The concept of spin Precession of magnetic spin Relaxation Bloch Equation. Bioengineering 280A Principles of Biomedical Imaging
 Bioengineering 280A Principles of Biomedical Imaging Fall Quarter 2006 MRI Lecture 1 Topics The concept of spin Precession of magnetic spin Relaxation Bloch Equation 1 Spin Intrinsic angular momentum of
Bioengineering 280A Principles of Biomedical Imaging Fall Quarter 2006 MRI Lecture 1 Topics The concept of spin Precession of magnetic spin Relaxation Bloch Equation 1 Spin Intrinsic angular momentum of
NMR journey. Introduction to solution NMR. Alexandre Bonvin. Topics. Why use NMR...? Bijvoet Center for Biomolecular Research
 2 NMR journey Introduction to solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, CCMB, Hyderabad, India November 29th
2 NMR journey Introduction to solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, CCMB, Hyderabad, India November 29th
MOLECULAR SPECTROSCOPY AND PHOTOCHEMISTRY
 20 CHAPTER MOLECULAR SPECTROSCOPY AND PHOTOCHEMISTRY 20.1 Introduction to Molecular Spectroscopy 20.2 Experimental Methods in Molecular Spectroscopy 20.3 Rotational and Vibrational Spectroscopy 20.4 Nuclear
20 CHAPTER MOLECULAR SPECTROSCOPY AND PHOTOCHEMISTRY 20.1 Introduction to Molecular Spectroscopy 20.2 Experimental Methods in Molecular Spectroscopy 20.3 Rotational and Vibrational Spectroscopy 20.4 Nuclear
Experiment 7: Dynamic NMR spectroscopy (Dated: April 19, 2010)
 Experiment 7: Dynamic NMR spectroscopy (Dated: April 19, 2010) I. INTRODUCTION In general spectroscopic experiments are divided into two categories: optical spectroscopy and magnetic spectroscopy. In previous
Experiment 7: Dynamic NMR spectroscopy (Dated: April 19, 2010) I. INTRODUCTION In general spectroscopic experiments are divided into two categories: optical spectroscopy and magnetic spectroscopy. In previous
PROTEIN NMR SPECTROSCOPY
 List of Figures List of Tables xvii xxvi 1. NMR SPECTROSCOPY 1 1.1 Introduction to NMR Spectroscopy 2 1.2 One Dimensional NMR Spectroscopy 3 1.2.1 Classical Description of NMR Spectroscopy 3 1.2.2 Nuclear
List of Figures List of Tables xvii xxvi 1. NMR SPECTROSCOPY 1 1.1 Introduction to NMR Spectroscopy 2 1.2 One Dimensional NMR Spectroscopy 3 1.2.1 Classical Description of NMR Spectroscopy 3 1.2.2 Nuclear
10.4 Continuous Wave NMR Instrumentation
 10.4 Continuous Wave NMR Instrumentation coherent detection bulk magnetization the rotating frame, and effective magnetic field generating a rotating frame, and precession in the laboratory frame spin-lattice
10.4 Continuous Wave NMR Instrumentation coherent detection bulk magnetization the rotating frame, and effective magnetic field generating a rotating frame, and precession in the laboratory frame spin-lattice
V27: RF Spectroscopy
 Martin-Luther-Universität Halle-Wittenberg FB Physik Advanced Lab Course V27: RF Spectroscopy ) Electron spin resonance (ESR) Investigate the resonance behaviour of two coupled LC circuits (an active rf
Martin-Luther-Universität Halle-Wittenberg FB Physik Advanced Lab Course V27: RF Spectroscopy ) Electron spin resonance (ESR) Investigate the resonance behaviour of two coupled LC circuits (an active rf
Scalar (contact) vs dipolar (pseudocontact) contributions to isotropic shifts.
 Scalar (contact) vs dipolar (pseudocontact) contributions to isotropic shifts. Types of paramagnetic species: organic radicals, and complexes of transition metals, lanthanides, and actinides. Simplest
Scalar (contact) vs dipolar (pseudocontact) contributions to isotropic shifts. Types of paramagnetic species: organic radicals, and complexes of transition metals, lanthanides, and actinides. Simplest
Structurele Biologie NMR
 MR journey Structurele Biologie MR 5 /3C 3 /65 MR & Structural biology course setup lectures - Sprangers R & Kay LE ature (27) basics of MR (Klaartje ouben: k.houben@uu.nl; 4/2) from peaks to data (ans
MR journey Structurele Biologie MR 5 /3C 3 /65 MR & Structural biology course setup lectures - Sprangers R & Kay LE ature (27) basics of MR (Klaartje ouben: k.houben@uu.nl; 4/2) from peaks to data (ans
- Basic understandings: - Mapping interactions:
 NMR-lecture April 6th, 2009, FMP Berlin Outline: Christian Freund - Basic understandings: Relaxation Chemical exchange - Mapping interactions: -Chemical shift mapping (fast exchange) Linewidth analysis
NMR-lecture April 6th, 2009, FMP Berlin Outline: Christian Freund - Basic understandings: Relaxation Chemical exchange - Mapping interactions: -Chemical shift mapping (fast exchange) Linewidth analysis
Basic One- and Two-Dimensional NMR Spectroscopy
 Horst Friebolin Basic One- and Two-Dimensional NMR Spectroscopy Third Revised Edition Translated by Jack K. Becconsall WILEY-VCH Weinheim New York Chichester Brisbane Singapore Toronto Contents XV 1 The
Horst Friebolin Basic One- and Two-Dimensional NMR Spectroscopy Third Revised Edition Translated by Jack K. Becconsall WILEY-VCH Weinheim New York Chichester Brisbane Singapore Toronto Contents XV 1 The
Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10
 Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10 Instructions: This is an open book, limited time, exam. You may use notes you have from class and any text book you find useful. You may also use
Name: BCMB/CHEM 8190, BIOMOLECULAR NMR FINAL EXAM-5/5/10 Instructions: This is an open book, limited time, exam. You may use notes you have from class and any text book you find useful. You may also use
Fluorescence Spectroscopy
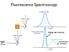 Fluorescence Spectroscopy Raleigh light scattering light all freqs Fluorescence emission Raleigh Scattering 10 nm Raleigh light scattering Fluorescence emission 400 nm Scattering - TWO particle 10 nm Particle
Fluorescence Spectroscopy Raleigh light scattering light all freqs Fluorescence emission Raleigh Scattering 10 nm Raleigh light scattering Fluorescence emission 400 nm Scattering - TWO particle 10 nm Particle
Nuclear Magnetic Resonance Spectroscopy Chem 4010/5326: Organic Spectroscopic Analysis Andrew Harned
 Nuclear Magnetic Resonance Spectroscopy Chem 4010/5326: Organic Spectroscopic Analysis 2015 Andrew Harned NMR Spectroscopy NMR Spectroscopy All nuclei have a nuclear spin quantum number (I) I = 0, 1/2,
Nuclear Magnetic Resonance Spectroscopy Chem 4010/5326: Organic Spectroscopic Analysis 2015 Andrew Harned NMR Spectroscopy NMR Spectroscopy All nuclei have a nuclear spin quantum number (I) I = 0, 1/2,
Fundamental MRI Principles Module 2 N. Nuclear Magnetic Resonance. X-ray. MRI Hydrogen Protons. Page 1. Electrons
 Fundamental MRI Principles Module 2 N S 1 Nuclear Magnetic Resonance There are three main subatomic particles: protons positively charged neutrons no significant charge electrons negatively charged Protons
Fundamental MRI Principles Module 2 N S 1 Nuclear Magnetic Resonance There are three main subatomic particles: protons positively charged neutrons no significant charge electrons negatively charged Protons
Manual to NMR2SCO-scripts
 Manual to NMR2SCO-scripts By Holm Petzold Introduction NMR 2 SCO is a collection of 3 Visual Basic scripts to be used for elucidation of solution H-NMR data of Fe 2+ -SCO complexes. They run on every MS
Manual to NMR2SCO-scripts By Holm Petzold Introduction NMR 2 SCO is a collection of 3 Visual Basic scripts to be used for elucidation of solution H-NMR data of Fe 2+ -SCO complexes. They run on every MS
NMR-spectroscopy in solution - an introduction. Peter Schmieder
 NMR-spectroscopy in solution - an introduction 2/92 Advanced Bioanalytics NMR-Spectroscopy Introductory session (11:00 12:30) Basic aspects of NMR-spectroscopy NMR parameter Multidimensional NMR-spectroscopy
NMR-spectroscopy in solution - an introduction 2/92 Advanced Bioanalytics NMR-Spectroscopy Introductory session (11:00 12:30) Basic aspects of NMR-spectroscopy NMR parameter Multidimensional NMR-spectroscopy
Suppression of Static Magnetic Field in Diffusion Measurements of Heterogeneous Materials
 PIERS ONLINE, VOL. 5, NO. 1, 2009 81 Suppression of Static Magnetic Field in Diffusion Measurements of Heterogeneous Materials Eva Gescheidtova 1 and Karel Bartusek 2 1 Faculty of Electrical Engineering
PIERS ONLINE, VOL. 5, NO. 1, 2009 81 Suppression of Static Magnetic Field in Diffusion Measurements of Heterogeneous Materials Eva Gescheidtova 1 and Karel Bartusek 2 1 Faculty of Electrical Engineering
A Hands on Introduction to NMR Lecture #1 Nuclear Spin and Magnetic Resonance
 A Hands on Introduction to NMR 22.920 Lecture #1 Nuclear Spin and Magnetic Resonance Introduction - The aim of this short course is to present a physical picture of the basic principles of Nuclear Magnetic
A Hands on Introduction to NMR 22.920 Lecture #1 Nuclear Spin and Magnetic Resonance Introduction - The aim of this short course is to present a physical picture of the basic principles of Nuclear Magnetic
Solid-state NMR and proteins : basic concepts (a pictorial introduction) Barth van Rossum,
 Solid-state NMR and proteins : basic concepts (a pictorial introduction) Barth van Rossum, 16.02.2009 Solid-state and solution NMR spectroscopy have many things in common Several concepts have been/will
Solid-state NMR and proteins : basic concepts (a pictorial introduction) Barth van Rossum, 16.02.2009 Solid-state and solution NMR spectroscopy have many things in common Several concepts have been/will
Introduction of Key Concepts of Nuclear Magnetic Resonance
 I have not yet lost that sense of wonder, and delight, that this delicate motion should reside in all ordinary things around us, revealing itself only to those who looks for it. E. M. Purcell, Nobel Lecture.
I have not yet lost that sense of wonder, and delight, that this delicate motion should reside in all ordinary things around us, revealing itself only to those who looks for it. E. M. Purcell, Nobel Lecture.
PRACTICAL ASPECTS OF NMR RELAXATION STUDIES OF BIOMOLECULAR DYNAMICS
 PRACTICAL ASPECTS OF MR RELAXATIO STUDIES OF BIOMOLECULAR DYAMICS Further reading: (Can be downloaded from my web page Korzhnev D.E., Billeter M., Arseniev A.S., and Orekhov V. Y., MR Studies of Brownian
PRACTICAL ASPECTS OF MR RELAXATIO STUDIES OF BIOMOLECULAR DYAMICS Further reading: (Can be downloaded from my web page Korzhnev D.E., Billeter M., Arseniev A.S., and Orekhov V. Y., MR Studies of Brownian
Spin-spin coupling I Ravinder Reddy
 Spin-spin coupling I Ravinder Reddy Spin-interactions External interactions Magnetic field Bo, RF field B1 Internal Interactions Molecular motions Exchange Chemical shifts J-coupling Spin Diffusion Dipolar
Spin-spin coupling I Ravinder Reddy Spin-interactions External interactions Magnetic field Bo, RF field B1 Internal Interactions Molecular motions Exchange Chemical shifts J-coupling Spin Diffusion Dipolar
NMR Spectroscopy of Polymers
 UNESCO/IUPAC Course 2005/2006 Jiri Brus NMR Spectroscopy of Polymers Brus J 1. part At the very beginning the phenomenon of nuclear spin resonance was studied predominantly by physicists and the application
UNESCO/IUPAC Course 2005/2006 Jiri Brus NMR Spectroscopy of Polymers Brus J 1. part At the very beginning the phenomenon of nuclear spin resonance was studied predominantly by physicists and the application
Physikalische Chemie IV (Magnetische Resonanz) HS Solution Set 2. Hand out: Hand in:
 Solution Set Hand out:.. Hand in:.. Repetition. The magnetization moves adiabatically during the application of an r.f. pulse if it is always aligned along the effective field axis. This behaviour is observed
Solution Set Hand out:.. Hand in:.. Repetition. The magnetization moves adiabatically during the application of an r.f. pulse if it is always aligned along the effective field axis. This behaviour is observed
Control of Spin Systems
 Control of Spin Systems The Nuclear Spin Sensor Many Atomic Nuclei have intrinsic angular momentum called spin. The spin gives the nucleus a magnetic moment (like a small bar magnet). Magnetic moments
Control of Spin Systems The Nuclear Spin Sensor Many Atomic Nuclei have intrinsic angular momentum called spin. The spin gives the nucleus a magnetic moment (like a small bar magnet). Magnetic moments
An introduction to Solid State NMR and its Interactions
 An introduction to Solid State NMR and its Interactions From tensor to NMR spectra CECAM Tutorial September 9 Calculation of Solid-State NMR Parameters Using the GIPAW Method Thibault Charpentier - CEA
An introduction to Solid State NMR and its Interactions From tensor to NMR spectra CECAM Tutorial September 9 Calculation of Solid-State NMR Parameters Using the GIPAW Method Thibault Charpentier - CEA
Fundamental MRI Principles Module Two
 Fundamental MRI Principles Module Two 1 Nuclear Magnetic Resonance There are three main subatomic particles: protons neutrons electrons positively charged no significant charge negatively charged Protons
Fundamental MRI Principles Module Two 1 Nuclear Magnetic Resonance There are three main subatomic particles: protons neutrons electrons positively charged no significant charge negatively charged Protons
NMR Dynamics and Relaxation
 NMR Dynamics and Relaxation Günter Hempel MLU Halle, Institut für Physik, FG Festkörper-NMR 1 Introduction: Relaxation Two basic magnetic relaxation processes: Longitudinal relaxation: T 1 Relaxation Return
NMR Dynamics and Relaxation Günter Hempel MLU Halle, Institut für Physik, FG Festkörper-NMR 1 Introduction: Relaxation Two basic magnetic relaxation processes: Longitudinal relaxation: T 1 Relaxation Return
Spectral Broadening Mechanisms. Broadening mechanisms. Lineshape functions. Spectral lifetime broadening
 Spectral Broadening echanisms Lorentzian broadening (Homogeneous) Gaussian broadening (Inhomogeneous, Inertial) Doppler broadening (special case for gas phase) The Fourier Transform NC State University
Spectral Broadening echanisms Lorentzian broadening (Homogeneous) Gaussian broadening (Inhomogeneous, Inertial) Doppler broadening (special case for gas phase) The Fourier Transform NC State University
Introduction to MRI. Spin & Magnetic Moments. Relaxation (T1, T2) Spin Echoes. 2DFT Imaging. K-space & Spatial Resolution.
 Introduction to MRI Spin & Magnetic Moments Relaxation (T1, T2) Spin Echoes 2DFT Imaging Selective excitation, phase & frequency encoding K-space & Spatial Resolution Contrast (T1, T2) Acknowledgement:
Introduction to MRI Spin & Magnetic Moments Relaxation (T1, T2) Spin Echoes 2DFT Imaging Selective excitation, phase & frequency encoding K-space & Spatial Resolution Contrast (T1, T2) Acknowledgement:
NMR Relaxation and Molecular Dynamics
 Ecole RMN Cargese Mars 2008 NMR Relaxation and Molecular Dynamics Martin Blackledge IBS Grenoble Carine van Heijenoort ICSN, CNRS Gif-sur-Yvette Solution NMR Timescales for Biomolecular Motion ps ns µs
Ecole RMN Cargese Mars 2008 NMR Relaxation and Molecular Dynamics Martin Blackledge IBS Grenoble Carine van Heijenoort ICSN, CNRS Gif-sur-Yvette Solution NMR Timescales for Biomolecular Motion ps ns µs
Measuring S using an analytical ultracentrifuge. Moving boundary
 Measuring S using an analytical ultracentrifuge Moving boundary [C] t = 0 t 1 t 2 0 top r bottom 1 dr b r b (t) r b ω 2 = S ln = ω 2 S (t-t dt r b (t o ) o ) r b = boundary position velocity = dr b dt
Measuring S using an analytical ultracentrifuge Moving boundary [C] t = 0 t 1 t 2 0 top r bottom 1 dr b r b (t) r b ω 2 = S ln = ω 2 S (t-t dt r b (t o ) o ) r b = boundary position velocity = dr b dt
Polarized solid deuteron targets EU-SpinMap Dubrovnik
 Experimentalphysik I Arbeitsgruppe Physik der Hadronen und Kerne Prof. Dr. W. Meyer G. Reicherz, Chr. Heß, A. Berlin, J. Herick Polarized solid deuteron targets EU-SpinMap 11.10.2010 Dubrovnik Polarized
Experimentalphysik I Arbeitsgruppe Physik der Hadronen und Kerne Prof. Dr. W. Meyer G. Reicherz, Chr. Heß, A. Berlin, J. Herick Polarized solid deuteron targets EU-SpinMap 11.10.2010 Dubrovnik Polarized
NMR BMB 173 Lecture 16, February
 NMR The Structural Biology Continuum Today s lecture: NMR Lots of slides adapted from Levitt, Spin Dynamics; Creighton, Proteins; And Andy Rawlinson There are three types of particles in the universe Quarks
NMR The Structural Biology Continuum Today s lecture: NMR Lots of slides adapted from Levitt, Spin Dynamics; Creighton, Proteins; And Andy Rawlinson There are three types of particles in the universe Quarks
EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December Suggested resolution
 page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
Protein Dynamics, Allostery and Function
 Protein Dynamics, Allostery and Function Lecture 2. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Functional Protein Dynamics Proteins are dynamic and
Protein Dynamics, Allostery and Function Lecture 2. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Functional Protein Dynamics Proteins are dynamic and
Magnetic Resonance Spectroscopy EPR and NMR
 Magnetic Resonance Spectroscopy EPR and NMR A brief review of the relevant bits of quantum mechanics 1. Electrons have spin, - rotation of the charge about its axis generates a magnetic field at each electron.
Magnetic Resonance Spectroscopy EPR and NMR A brief review of the relevant bits of quantum mechanics 1. Electrons have spin, - rotation of the charge about its axis generates a magnetic field at each electron.
