SNPs versus sequences for phylogeography an explora:on using simula:ons and massively parallel sequencing in a non- model bird
|
|
|
- Katherine French
- 5 years ago
- Views:
Transcription
1 SNPs versus sequences for phylogeography an explora:on using simula:ons and massively parallel sequencing in a non- model bird Michael G. Harvey, Brian T. Smith, Brant C. Faircloth, Travis C. Glenn, and Robb T. Brumfield 100 CH CH CA NA CA SA SA 0 0 SA 100 NA NA
2
3 Which method of genera:ng massively parallel sequence data should I use for phylogeography?
4 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
5 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
6 What are some of the op:ons? Whole genome sequencing Reduced representa:on library sequencing: - PCR amplicon sequencing - RNA sequencing - Sequence capture - Restric:on digest- based methods (RAD tags, GBS, etc.)
7 What are some of the op:ons? Whole genome sequencing Reduced representa:on library sequencing: - PCR amplicon sequencing - RNA sequencing - Sequence capture - Restric:on digest- based methods (RAD tags, GBS, etc.)
8 How do they work?
9 1. Random fragmenta:on 2. Addi:on of barcodes and adapters, PCR Sequence Capture 3. Hybridiza:on to bio:nylated probes 4. Enrichment with streptavidin beads 5. Amplifica:on, Illumina sequencing
10 Sequence Capture probe 1. Random fragmenta:on 2. Addi:on of barcodes and adapters, PCR 3. Hybridiza:on to bio:nylated probes Enrichment Amplifica:on, with Illumina streptavidin sequencing beads
11 Sequence Capture probe 1. Random fragmenta:on 2. Addi:on of barcodes and adapters, PCR Sequences up to 1000 bp 3. Hybridiza:on to bio:nylated probes Enrichment Amplifica:on, with Illumina streptavidin sequencing beads
12 Ultraconserved Elements (UCEs) are useful for sequence capture Frequency of variable positions Core UCE (probe) UCE alignments across all birds (from McCormack et al. 2013) Distance from center of alignment (bp)
13 UCEs are useful for deep phylogeny
14 But do UCEs work at phylogeographic :mescales?
15 Restric:on Digest- based Methods 1. RE diges:on 2. Addi:on of barcodes and adapters 3. (Size selec:on), amplifica:on, Illumina sequencing
16 Restric:on Digest- based Methods 1. RE diges:on 2. Addi:on of barcodes and adapters 3. (Size selec:on), amplifica:on, Illumina sequencing
17 Restric:on Digest- based Methods 1. RE diges:on 2. Addi:on of barcodes and adapters 3. (Size selec:on), amplifica:on, Illumina sequencing Read- length sequences 100 bp
18 Restric:on Digest- based Methods Methods: CRoPs (Van Orsouw et al. 2007) RAD- seq (Baird et al. 2008) Modified CRoPs (Gompert et al. 2010) MSG (Andolfaao et al. 2011) GBS (Elshire et al. 2011) Modified GBS (Parchman et al. 2012) ddrad- seq (Hohenlohe et al. 2012, Peterson et al. 2012) Modified GBS (Poland et al. 2012) SBG (Truong et al. 2012) 2b- RAD- seq (Wang et al. 2012)
19 Restric:on Digest- based Methods Methods: CRoPs (Van Orsouw et al. 2007) RAD- seq (Baird et al. 2008) Modified CRoPs (Gompert et al. 2010) MSG (Andolfaao et al. 2011) GBS (Elshire et al. 2011) Modified GBS (Parchman et al. 2012) ddrad- seq (Hohenlohe et al. 2012, Peterson et al. 2012) Modified GBS (Poland et al. 2012) SBG (Truong et al. 2012) 2b- RAD- seq (Wang et al. 2012)
20 Sequence Capture RD- based Methods Fewer, longer sequences up to 1000 bp Many, read- length sequences 100 bp (SNPs)
21 What have they been used for? Sequence Capture Re- sequencing in model species (e.g. exome) Phylogene:cs RD- based methods GWAS QTL- mapping Popula:on Gene:cs
22 What have they been used for? Sequence Capture RD- based methods Phylogene:cs Phylogeography? Popula:on Gene:cs
23 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
24 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
25 Let s try them out! Xenops minutus (Aves; Furnariidae) Smith et al. (in prep.) See talk this agernoon! Peruvian B, 3:30pm Photo: L. C. Ribenboim
26 Study Design UCEs 2 individuals / popula:on 2,560 probes targe:ng 2,386 UCEs ~1/5 lane of Illumina HiSeq (PE) Bioinforma:cs (illumiprocessor, Velvet, phyluce, BWA, MAFFT) ultraconserved.org SNPs 8 individuals / popula:on (+ outgroup) Ins:tute for Genomic Diversity (Cornell Univ.) ~2/3 lane of Illumina HiSeq (SE) Bioinforma:cs (UNEAK)
27 Challenges read quality manual edi:ng sample size thresholds mt excess paralogy keeping indexes straight hybridiza:on contamina:on CPU :me run:mes failed runs qpcr sucks enrichment? file size computa:on memory number of popula:ons
28 UCE Sequence Capture Results
29 1,368 UCE Alignments 834 bp = snps dark gray = missing data
30 UCE Intraspecific Varia:on
31 UCE Tree Reconstruc:on 1.0 *BEAST (Heled and Drummond 2010) 137 loci 9.0E NA SA CA CH
32 UCE Tree Reconstruc:on 1.0 *BEAST (Heled and Drummond 2010) 137 loci 9.0E NA SA CA CH
33 UCE Tree Reconstruc:on 1.0 *BEAST (Heled and Drummond 2010) 137 loci 9.0E NA SA CA CH
34 UCE Demographic Modeling 1.0 G- PhoCS (Gronau et al. 2011) 1,368 loci 9.0E SA NA CA CH
35 UCE Demographic Modeling Time before present (My) SA NA CA CH 0.0
36 Genotyping by Sequencing Results
37 GBS Dataset Summary 106,784 SNPs pre- filtering 2,355 SNPs with data for all individuals
38 SNP Tree Reconstruc:on Methods Concatena:on SNAPP (Bryant et al. 2012) TreeMix (Pickrell and Pritchard 2012)
39 SNP Tree Reconstruc:on Methods Concatena:on SNAPP (Bryant et al. 2012) TreeMix (Pickrell and Pritchard 2012)
40 SNP Tree 100 TreeMix (Pickrell and Pritchard 2012) NA SA CA CH
41 SNP vs. UCE Trees E NA SA CA CH NA SA CA CH
42 SNP Demography Allele Frequency Spectra 16 a i (Gutenkunst Recent High Unequal migra:on divergence et al. N 2009) e Popula:on Popula:on 2 16
43 SNP Demography Allele Frequency Spectra 16 Popula:on Popula:on 2 16
44 SNP Demography Allele Frequency Spectra Popula:on Popula:on 2 16
45 SNP Demography Allele Frequency Spectra Unequal N e Popula:on Popula:on 2 16
46 SNP Demography Allele Frequency Spectra Recent divergence Popula:on Popula:on 2 16
47 SNP Demography Allele Frequency Spectra High migra:on Popula:on Popula:on 2 16
48 SNP Demography Allele Frequency Spectra a i (Gutenkunst et al. 2009) Popula:on Popula:on 2 16
49 SNP Demographic Modeling data model
50 SNP Demographic Modeling NA SA CA CH
51 SNP Demographic Modeling NA NASA SA CA CACH CH
52 SNP Demographic Modeling data model ((NASA) (CACH)) NA NASA SA CA CACH CH
53 SNP Demographic Modeling data model ((NASA) (CACH)) ((NA)(SA)) NA SA CA CH
54 SNP Demographic Modeling data model ((NASA) (CACH)) ((NA)(SA)) NA SA CA CH ((CA)(CH))
55 SNP Demographic Modeling Time before present (My) SA NA CA CH 0.0
56 SNP vs UCE Demography Time before present (My) SA NA CA CH SA NA CA CH 0.0
57 Summary of Empirical Results Species trees are similar between datasets Demographic models differ somewhat in parameter es:mates between datasets
58 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
59 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
60 Simula:on Treatments Tau (species divergence =me) 0.1, 0.2, 0.4, 0.8, 1.6 θ :me Theta (4N e μ) 0.2, 0.4, 0.8, 1.6, 3.2 θ τ Migra=on rate (N e m) 0, 0.1, 0.3, 0.7, 1.5 github.com/mgharvey/mps- sim θ m m θ m m m m θ τ / 2
61 Divergence Time Es:mate Accuracy a I and 50,000 GBS SNPs G- PhoCS and 1,000 UCEs 3e+06 2 Divergence :me 1 2e+06 1e+06 P 0e+00 Tau treatment Tau Treatment Tau treatment
62 Effec:ve Popula:on Size Es:mate Accuracy a I and 50,000 GBS SNPs G- PhoCS and 1,000 UCEs 5 5e+06 Migrants per genera:on e+06 3e+06 2e+06 1e+06 0e+00 Theta treatment Theta treatment Theta Treatment
63 Migra:on Es:mate Accuracy a I and 50,000 GBS SNPs G- PhoCS and 1,000 UCEs Migrants per genera:on e+00 Migra:on treatment Migration Treatment Migra:on treatment
64 Summary of Simula:on Results SNPs+ a i and sequences+g- PhoCS es:mate divergence :mes with similar accuracy SNPs+ a i es:mate N e and migra:on with higher accuracy
65 Why the Varia:on in Accuracy? 1. Varia:on across datasets 2. Number of loci 3. Number of individuals 4. Analy:cal method
66 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
67 Outline 1. Overview of methods for genera:ng massively parallel sequencing data 2. Phylogeography of a non- model bird a. Sequence capture of ultraconserved elements (UCEs) b. SNPs from Genotyping by Sequencing 3. Simula=ons: UCEs vs. SNPs for phylogeography 4. Conclusions
68 Xenops minutus Popula:on History 9.0E Time before present (My) NA SA CA CH SA NA CA CH 0.0
69 Sequences vs. SNPs Sequences from Sequence Capture SNPs from Genotyping by Sequencing CH CH 100 CA CA NA SA SA NA NA 0 0 SA θ 9.0E-5 θ θ τ NA SA CA CH θ m θ m τ m / 2
70 Which method of genera:ng massively parallel sequence data should I use for phylogeography? Sequence Capture RD- based methods Phylogene:cs Popula:on Gene:cs
71 Which method of genera:ng massively parallel sequence data should I use for phylogeography? Sequence Capture RD- based methods Phylogene:cs Phylogeography Popula:on Gene:cs
72 Harvey Commiaee Members Mike Hellberg Jeremy Brown James V. Remsen Brumfield Lab John McCormack Liz Derryberry Glenn Seeholzer Caroline Duffie James Maley Sarah Hird Andrés Cuervo Gustavo Bravo Discussion Bryan Carstens John Andersen Eric Riameyer Clare Brown Noah Reid LSU Bird Lunch LSU Museum of Natural Science LSU Biological Sciences Acknowledgements LSU High Performance Compu:ng Le Yan Bhupender Thakur Support Society of Systema:c Biologists Na:onal Science Founda:on For UCE probe sets, protocols, and code: ultraconserved.org For simula:on code: github.com/mgharvey/mps- sim
Fei Lu. Post doctoral Associate Cornell University
 Fei Lu Post doctoral Associate Cornell University http://www.maizegenetics.net Genotyping by sequencing (GBS) is simple and cost effective 1. Digest DNA 2. Ligate adapters with barcodes 3. Pool DNAs 4.
Fei Lu Post doctoral Associate Cornell University http://www.maizegenetics.net Genotyping by sequencing (GBS) is simple and cost effective 1. Digest DNA 2. Ligate adapters with barcodes 3. Pool DNAs 4.
Genotyping By Sequencing (GBS) Method Overview
 enotyping By Sequencing (BS) Method Overview Sharon E Mitchell Institute for enomic Diversity Cornell University http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina sequencing
enotyping By Sequencing (BS) Method Overview Sharon E Mitchell Institute for enomic Diversity Cornell University http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina sequencing
Genotyping By Sequencing (GBS) Method Overview
 enotyping By Sequencing (BS) Method Overview RJ Elshire, JC laubitz, Q Sun, JV Harriman ES Buckler, and SE Mitchell http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina
enotyping By Sequencing (BS) Method Overview RJ Elshire, JC laubitz, Q Sun, JV Harriman ES Buckler, and SE Mitchell http://wwwmaizegeneticsnet/ Topics Presented Background/oals BS lab protocol Illumina
From Gene Trees to Species Trees. Tandy Warnow The University of Texas at Aus<n
 From Gene Trees to Species Trees Tandy Warnow The University of Texas at Aus
From Gene Trees to Species Trees Tandy Warnow The University of Texas at Aus
Evolu&on, Popula&on Gene&cs, and Natural Selec&on Computa.onal Genomics Seyoung Kim
 Evolu&on, Popula&on Gene&cs, and Natural Selec&on 02-710 Computa.onal Genomics Seyoung Kim Phylogeny of Mammals Phylogene&cs vs. Popula&on Gene&cs Phylogene.cs Assumes a single correct species phylogeny
Evolu&on, Popula&on Gene&cs, and Natural Selec&on 02-710 Computa.onal Genomics Seyoung Kim Phylogeny of Mammals Phylogene&cs vs. Popula&on Gene&cs Phylogene.cs Assumes a single correct species phylogeny
Prac%cal Bioinforma%cs for Life Scien%sts. Week 14, Lecture 28. István Albert Bioinforma%cs Consul%ng Center Penn State
 Prac%cal Bioinforma%cs for Life Scien%sts Week 14, Lecture 28 István Albert Bioinforma%cs Consul%ng Center Penn State Final project A group of researchers are interested in studying protein binding loca%ons
Prac%cal Bioinforma%cs for Life Scien%sts Week 14, Lecture 28 István Albert Bioinforma%cs Consul%ng Center Penn State Final project A group of researchers are interested in studying protein binding loca%ons
Target Capture and Massively Parallel Sequencing of Ultraconserved Elements (UCEs) for. Comparative Studies at Shallow Evolutionary Time Scales
 1 Target Capture and Massively Parallel Sequencing of Ultraconserved Elements (UCEs) for Comparative Studies at Shallow Evolutionary Time Scales Brian Tilston Smith 1*, Michael G. Harvey 1,2*, Brant C.
1 Target Capture and Massively Parallel Sequencing of Ultraconserved Elements (UCEs) for Comparative Studies at Shallow Evolutionary Time Scales Brian Tilston Smith 1*, Michael G. Harvey 1,2*, Brant C.
Demographic Inference
 Demographic Inference Mike Hickerson City University of New York outline 1. Historical Demographic inference with genetics 2. Hierarchical Bayesian inference & Bayesian model averaging (w/abc) 3. software
Demographic Inference Mike Hickerson City University of New York outline 1. Historical Demographic inference with genetics 2. Hierarchical Bayesian inference & Bayesian model averaging (w/abc) 3. software
GBS Bioinformatics Pipeline(s) Overview
 GBS Bioinformatics Pipeline(s) Overview Getting from sequence files to genotypes. Pipeline Coding: Ed Buckler Jeff Glaubitz James Harriman Presentation: Terry Casstevens With supporting information from
GBS Bioinformatics Pipeline(s) Overview Getting from sequence files to genotypes. Pipeline Coding: Ed Buckler Jeff Glaubitz James Harriman Presentation: Terry Casstevens With supporting information from
GBS Bioinformatics Pipeline(s) Overview
 GBS Bioinformatics Pipeline(s) Overview Getting from sequence files to genotypes. Pipeline Coding: Ed Buckler Jeff Glaubitz James Harriman Presentation: Rob Elshire With supporting information from the
GBS Bioinformatics Pipeline(s) Overview Getting from sequence files to genotypes. Pipeline Coding: Ed Buckler Jeff Glaubitz James Harriman Presentation: Rob Elshire With supporting information from the
Analysis of Y-STR Profiles in Mixed DNA using Next Generation Sequencing
 Analysis of Y-STR Profiles in Mixed DNA using Next Generation Sequencing So Yeun Kwon, Hwan Young Lee, and Kyoung-Jin Shin Department of Forensic Medicine, Yonsei University College of Medicine, Seoul,
Analysis of Y-STR Profiles in Mixed DNA using Next Generation Sequencing So Yeun Kwon, Hwan Young Lee, and Kyoung-Jin Shin Department of Forensic Medicine, Yonsei University College of Medicine, Seoul,
Introduction to Statistical Genetics (BST227) Lecture 6: Population Substructure in Association Studies
 Introduction to Statistical Genetics (BST227) Lecture 6: Population Substructure in Association Studies Confounding in gene+c associa+on studies q What is it? q What is the effect? q How to detect it?
Introduction to Statistical Genetics (BST227) Lecture 6: Population Substructure in Association Studies Confounding in gene+c associa+on studies q What is it? q What is the effect? q How to detect it?
Comparison of Target-Capture and Restriction-Site Associated DNA Sequencing for Phylogenomics: A Test in Cardinalid Tanagers (Aves, Genus: Piranga)
 Syst. Biol. 65(4):640 650, 2016 The Author(s) 2016. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 65(4):640 650, 2016 The Author(s) 2016. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Phylogene)cs. IMBB 2016 BecA- ILRI Hub, Nairobi May 9 20, Joyce Nzioki
 Phylogene)cs IMBB 2016 BecA- ILRI Hub, Nairobi May 9 20, 2016 Joyce Nzioki Phylogenetics The study of evolutionary relatedness of organisms. Derived from two Greek words:» Phle/Phylon: Tribe/Race» Genetikos:
Phylogene)cs IMBB 2016 BecA- ILRI Hub, Nairobi May 9 20, 2016 Joyce Nzioki Phylogenetics The study of evolutionary relatedness of organisms. Derived from two Greek words:» Phle/Phylon: Tribe/Race» Genetikos:
Measurement of the meridional flow from eigenfunc5on perturba5ons
 Measurement of the meridional flow from eigenfunc5on perturba5ons Ariane Schad, Markus Roth Kiepenheuer- Ins5tut für Sonnenphysik Solar Subsurface Flows from Helioseismology: Problems and Prospects Helioseismology
Measurement of the meridional flow from eigenfunc5on perturba5ons Ariane Schad, Markus Roth Kiepenheuer- Ins5tut für Sonnenphysik Solar Subsurface Flows from Helioseismology: Problems and Prospects Helioseismology
Introduc)on to RNA- Seq Data Analysis. Dr. Benilton S Carvalho Department of Medical Gene)cs Faculty of Medical Sciences State University of Campinas
 Introduc)on to RNA- Seq Data Analysis Dr. Benilton S Carvalho Department of Medical Gene)cs Faculty of Medical Sciences State University of Campinas Material: hep://)ny.cc/rnaseq Slides: hep://)ny.cc/slidesrnaseq
Introduc)on to RNA- Seq Data Analysis Dr. Benilton S Carvalho Department of Medical Gene)cs Faculty of Medical Sciences State University of Campinas Material: hep://)ny.cc/rnaseq Slides: hep://)ny.cc/slidesrnaseq
Creative Commons Attribution 2.0 (
 v1.5 Target Enrichment of Illumina Libraries B. Faircloth August 21, 2013 Ecol. and Evol. Biology Univ. of California Los Angeles The goal here is to adapt the SureSelect/Mycroarray enrichment kits for
v1.5 Target Enrichment of Illumina Libraries B. Faircloth August 21, 2013 Ecol. and Evol. Biology Univ. of California Los Angeles The goal here is to adapt the SureSelect/Mycroarray enrichment kits for
Test of neutral theory predic3ons for the BCI tree community informed by regional abundance data
 Test of neutral theory predic3ons for the BCI tree community informed by regional abundance data Anne%e Ostling Cody Weinberger Devin Riley Ecology and Evolu:onary Biology University of Michigan 1 Outline
Test of neutral theory predic3ons for the BCI tree community informed by regional abundance data Anne%e Ostling Cody Weinberger Devin Riley Ecology and Evolu:onary Biology University of Michigan 1 Outline
Bias in RNA sequencing and what to do about it
 Bias in RNA sequencing and what to do about it Walter L. (Larry) Ruzzo Computer Science and Engineering Genome Sciences University of Washington Fred Hutchinson Cancer Research Center Seattle, WA, USA
Bias in RNA sequencing and what to do about it Walter L. (Larry) Ruzzo Computer Science and Engineering Genome Sciences University of Washington Fred Hutchinson Cancer Research Center Seattle, WA, USA
CSCI1950 Z Computa3onal Methods for Biology* (*Working Title) Lecture 1. Ben Raphael January 21, Course Par3culars
 CSCI1950 Z Computa3onal Methods for Biology* (*Working Title) Lecture 1 Ben Raphael January 21, 2009 Course Par3culars Three major topics 1. Phylogeny: ~50% lectures 2. Func3onal Genomics: ~25% lectures
CSCI1950 Z Computa3onal Methods for Biology* (*Working Title) Lecture 1 Ben Raphael January 21, 2009 Course Par3culars Three major topics 1. Phylogeny: ~50% lectures 2. Func3onal Genomics: ~25% lectures
Accounting for read depth in the analysis of genotyping-by-sequencing data
 Accounting for read depth in the analysis of genotyping-by-sequencing data Ken Dodds, John McEwan, Timothy Bilton, Rudi Brauning, Rayna Anderson, Tracey Van Stijn, Theodor Kristjánsson, Shannon Clarke
Accounting for read depth in the analysis of genotyping-by-sequencing data Ken Dodds, John McEwan, Timothy Bilton, Rudi Brauning, Rayna Anderson, Tracey Van Stijn, Theodor Kristjánsson, Shannon Clarke
Whole-genome amplification in doubledigest RADseq results in adequate libraries but fewer sequenced loci
 Whole-genome amplification in doubledigest RADseq results in adequate libraries but fewer sequenced loci Bruno A. S. de Medeiros and Brian D. Farrell Department of Organismic and Evolutionary Biology and
Whole-genome amplification in doubledigest RADseq results in adequate libraries but fewer sequenced loci Bruno A. S. de Medeiros and Brian D. Farrell Department of Organismic and Evolutionary Biology and
Heritability and the response to selec2on
 Heritability and the response to selec2on Resemblance between rela2ves in Quan2ta2ve traits A trait with L loci Each segregating an allele A 1 at freq. p l Each copy of the A 1 allele at a locus increasing
Heritability and the response to selec2on Resemblance between rela2ves in Quan2ta2ve traits A trait with L loci Each segregating an allele A 1 at freq. p l Each copy of the A 1 allele at a locus increasing
Phylogenomics of Phrynosomatid Lizards: Conflicting Signals from Sequence Capture. Versus Restriction Site Associated DNA Sequencing
 Genome Biology and Evolution Advance Access published February 7, 2015 doi:10.1093/gbe/evv026 Phylogenomics of Phrynosomatid Lizards: Conflicting Signals from Sequence Capture Versus Restriction Site Associated
Genome Biology and Evolution Advance Access published February 7, 2015 doi:10.1093/gbe/evv026 Phylogenomics of Phrynosomatid Lizards: Conflicting Signals from Sequence Capture Versus Restriction Site Associated
Construc)ng the Tree of Life: Divide-and-Conquer! Tandy Warnow University of Illinois at Urbana-Champaign
 Construc)ng the Tree of Life: Divide-and-Conquer! Tandy Warnow University of Illinois at Urbana-Champaign Phylogeny (evolutionary tree) Orangutan Gorilla Chimpanzee Human From the Tree of the Life Website,
Construc)ng the Tree of Life: Divide-and-Conquer! Tandy Warnow University of Illinois at Urbana-Champaign Phylogeny (evolutionary tree) Orangutan Gorilla Chimpanzee Human From the Tree of the Life Website,
Explore SNP polymorphism data. A. Dereeper, Y. Hueber
 Explore SNP polymorphism data A. Dereeper, Y. Hueber Bioinformatics trainings, Supagro, February, 2016 Tablet Graphical tool to visualize assemblies Accept many formats ACE, SAM, BAM GATK (Genome Analysis
Explore SNP polymorphism data A. Dereeper, Y. Hueber Bioinformatics trainings, Supagro, February, 2016 Tablet Graphical tool to visualize assemblies Accept many formats ACE, SAM, BAM GATK (Genome Analysis
Historical biogeography models with dispersal probability as a func7on of distance(s)
 Historical biogeography models with dispersal probability as a func7on of distance(s) Nicholas J. Matzke, Postdoctoral Fellow, NIMBioS (Na6onal Ins6tute of Mathema6cal, www.nimbios.org) SSE symposium:
Historical biogeography models with dispersal probability as a func7on of distance(s) Nicholas J. Matzke, Postdoctoral Fellow, NIMBioS (Na6onal Ins6tute of Mathema6cal, www.nimbios.org) SSE symposium:
Detecting selection from differentiation between populations: the FLK and hapflk approach.
 Detecting selection from differentiation between populations: the FLK and hapflk approach. Bertrand Servin bservin@toulouse.inra.fr Maria-Ines Fariello, Simon Boitard, Claude Chevalet, Magali SanCristobal,
Detecting selection from differentiation between populations: the FLK and hapflk approach. Bertrand Servin bservin@toulouse.inra.fr Maria-Ines Fariello, Simon Boitard, Claude Chevalet, Magali SanCristobal,
Ben Hecht Columbia River Inter-Tribal Fish Commission March 19, 2013
 The Genetic Basis for the Propensity to Migrate in a Sequestered Population of Rainbow and Steelhead Trout Ben Hecht Columbia River Inter-Tribal Fish Commission March 19, 2013 Migration Cultural, ecological,
The Genetic Basis for the Propensity to Migrate in a Sequestered Population of Rainbow and Steelhead Trout Ben Hecht Columbia River Inter-Tribal Fish Commission March 19, 2013 Migration Cultural, ecological,
Statistics for Differential Expression in Sequencing Studies. Naomi Altman
 Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Statistics for Differential Expression in Sequencing Studies Naomi Altman naomi@stat.psu.edu Outline Preliminaries what you need to do before the DE analysis Stat Background what you need to know to understand
Phylogenomics of Phrynosomatid Lizards: Conflicting Signals from Sequence Capture versus Restriction Site Associated DNA Sequencing
 Phylogenomics of Phrynosomatid Lizards: Conflicting Signals from Sequence Capture versus Restriction Site Associated DNA Sequencing Adam D. Leaché 1,2, *, Andreas S. Chavez 1,2,5,LeonardN.Jones 1,2,JaredA.Grummer
Phylogenomics of Phrynosomatid Lizards: Conflicting Signals from Sequence Capture versus Restriction Site Associated DNA Sequencing Adam D. Leaché 1,2, *, Andreas S. Chavez 1,2,5,LeonardN.Jones 1,2,JaredA.Grummer
Recombina*on and Linkage Disequilibrium (LD)
 Recombina*on and Linkage Disequilibrium (LD) A B a b r = recombina*on frac*on probability of an odd Number of crossovers occur Between our markers 0
Recombina*on and Linkage Disequilibrium (LD) A B a b r = recombina*on frac*on probability of an odd Number of crossovers occur Between our markers 0
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Our typical RNA quantification pipeline
 RNA-Seq primer Our typical RNA quantification pipeline Upload your sequence data (fastq) Align to the ribosome (Bow>e) Align remaining reads to genome (TopHat) or transcriptome (RSEM) Make report of quality
RNA-Seq primer Our typical RNA quantification pipeline Upload your sequence data (fastq) Align to the ribosome (Bow>e) Align remaining reads to genome (TopHat) or transcriptome (RSEM) Make report of quality
Inferring phylogenetic history from restriction site associated DNA (RADseq)
 Chapter 6 Inferring phylogenetic history from restriction site associated DNA (RADseq) Richard H. Ree 1 & Andrew L. Hipp 1,2 1 The Field Museum, 1400 South Lake Shore Drive, Chicago, Illinois 60605, U.S.A.
Chapter 6 Inferring phylogenetic history from restriction site associated DNA (RADseq) Richard H. Ree 1 & Andrew L. Hipp 1,2 1 The Field Museum, 1400 South Lake Shore Drive, Chicago, Illinois 60605, U.S.A.
Methods to reconstruct phylogene1c networks accoun1ng for ILS
 Methods to reconstruct phylogene1c networks accoun1ng for ILS Céline Scornavacca some slides have been kindly provided by Fabio Pardi ISE-M, Equipe Phylogénie & Evolu1on Moléculaires Montpellier, France
Methods to reconstruct phylogene1c networks accoun1ng for ILS Céline Scornavacca some slides have been kindly provided by Fabio Pardi ISE-M, Equipe Phylogénie & Evolu1on Moléculaires Montpellier, France
Comparative Genomics of Fagaceae
 Fagaceae Images.google.com Linkage Map www.quia.com TM www.clipartlord.com Selection of mapping parents SM2 SM1 Predominant pollinator? Progeny Exclusion for Full Sib Linkage Mapping Year Acorns genotyped
Fagaceae Images.google.com Linkage Map www.quia.com TM www.clipartlord.com Selection of mapping parents SM2 SM1 Predominant pollinator? Progeny Exclusion for Full Sib Linkage Mapping Year Acorns genotyped
Introduction to Phylogenetic Analysis
 Introduction to Phylogenetic Analysis Tuesday 24 Wednesday 25 July, 2012 School of Biological Sciences Overview This free workshop provides an introduction to phylogenetic analysis, with a focus on the
Introduction to Phylogenetic Analysis Tuesday 24 Wednesday 25 July, 2012 School of Biological Sciences Overview This free workshop provides an introduction to phylogenetic analysis, with a focus on the
Next-generation sequencing and the expanding domain of phylogeography
 Next-generation sequencing and the expanding domain of phylogeography The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation
Next-generation sequencing and the expanding domain of phylogeography The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters. Citation
, Helen K. Pigage 1, Peter J. Wettstein 2, Stephanie A. Prosser 1 and Jon C. Pigage 1ˆ. Jeremy M. Bono 1*
 Bono et al. BMC Evolutionary Biology (2018) 18:139 https://doi.org/10.1186/s12862-018-1248-4 RESEARCH ARTICLE Genome-wide markers reveal a complex evolutionary history involving divergence and introgression
Bono et al. BMC Evolutionary Biology (2018) 18:139 https://doi.org/10.1186/s12862-018-1248-4 RESEARCH ARTICLE Genome-wide markers reveal a complex evolutionary history involving divergence and introgression
Genetic Drift in Human Evolution
 Genetic Drift in Human Evolution (Part 2 of 2) 1 Ecology and Evolutionary Biology Center for Computational Molecular Biology Brown University Outline Introduction to genetic drift Modeling genetic drift
Genetic Drift in Human Evolution (Part 2 of 2) 1 Ecology and Evolutionary Biology Center for Computational Molecular Biology Brown University Outline Introduction to genetic drift Modeling genetic drift
Visualizing Population Genetics
 Visualizing Population Genetics Supervisors: Rachel Fewster, James Russell, Paul Murrell Louise McMillan 1 December 2015 Louise McMillan Visualizing Population Genetics 1 December 2015 1 / 29 Outline 1
Visualizing Population Genetics Supervisors: Rachel Fewster, James Russell, Paul Murrell Louise McMillan 1 December 2015 Louise McMillan Visualizing Population Genetics 1 December 2015 1 / 29 Outline 1
Supplementary Information for Discovery and characterization of indel and point mutations
 Supplementary Information for Discovery and characterization of indel and point mutations using DeNovoGear Avinash Ramu 1 Michiel J. Noordam 1 Rachel S. Schwartz 2 Arthur Wuster 3 Matthew E. Hurles 3 Reed
Supplementary Information for Discovery and characterization of indel and point mutations using DeNovoGear Avinash Ramu 1 Michiel J. Noordam 1 Rachel S. Schwartz 2 Arthur Wuster 3 Matthew E. Hurles 3 Reed
Exponen'al growth Limi'ng factors Environmental resistance Carrying capacity logis'c growth curve
 Exponen'al growth Popula)on increases by a fixed percent Fixed percent of a large number produces a large increase Graphed as a J- shaped curve Cannot be sustained indefinitely It occurs in nature With
Exponen'al growth Popula)on increases by a fixed percent Fixed percent of a large number produces a large increase Graphed as a J- shaped curve Cannot be sustained indefinitely It occurs in nature With
New imputation strategies optimized for crop plants: FILLIN (Fast, Inbred Line Library ImputatioN) FSFHap (Full Sib Family Haplotype)
 New imputation strategies optimized for crop plants: FILLIN (Fast, Inbred Line Library ImputatioN) FSFHap (Full Sib Family Haplotype) Kelly Swarts PAG Allele Mining 1/11/2014 Imputation is the projection
New imputation strategies optimized for crop plants: FILLIN (Fast, Inbred Line Library ImputatioN) FSFHap (Full Sib Family Haplotype) Kelly Swarts PAG Allele Mining 1/11/2014 Imputation is the projection
Lecture WS Evolutionary Genetics Part I 1
 Quantitative genetics Quantitative genetics is the study of the inheritance of quantitative/continuous phenotypic traits, like human height and body size, grain colour in winter wheat or beak depth in
Quantitative genetics Quantitative genetics is the study of the inheritance of quantitative/continuous phenotypic traits, like human height and body size, grain colour in winter wheat or beak depth in
Differen'al Privacy with Bounded Priors: Reconciling U+lity and Privacy in Genome- Wide Associa+on Studies
 Differen'al Privacy with Bounded Priors: Reconciling U+lity and Privacy in Genome- Wide Associa+on Studies Florian Tramèr, Zhicong Huang, Erman Ayday, Jean- Pierre Hubaux ACM CCS 205 Denver, Colorado,
Differen'al Privacy with Bounded Priors: Reconciling U+lity and Privacy in Genome- Wide Associa+on Studies Florian Tramèr, Zhicong Huang, Erman Ayday, Jean- Pierre Hubaux ACM CCS 205 Denver, Colorado,
Unlocking RNA-seq tools for zero inflation and single cell applications using observation weights
 Unlocking RNA-seq tools for zero inflation and single cell applications using observation weights Koen Van den Berge, Ghent University Statistical Genomics, 2018-2019 1 The team Koen Van den Berge Fanny
Unlocking RNA-seq tools for zero inflation and single cell applications using observation weights Koen Van den Berge, Ghent University Statistical Genomics, 2018-2019 1 The team Koen Van den Berge Fanny
Specia'on. The linkage between geographical circumstance and evolu5onary development is embodied in the very word biogeography."
 Specia'on 1 Specia'on "Wallace and Darwin s great insight [natural selec5on as a factor in specia5on] only began the era of asking. The mystery of mysteries had been solved, at least in rough outline;
Specia'on 1 Specia'on "Wallace and Darwin s great insight [natural selec5on as a factor in specia5on] only began the era of asking. The mystery of mysteries had been solved, at least in rough outline;
DART Tutorial Sec'on 1: Filtering For a One Variable System
 DART Tutorial Sec'on 1: Filtering For a One Variable System UCAR The Na'onal Center for Atmospheric Research is sponsored by the Na'onal Science Founda'on. Any opinions, findings and conclusions or recommenda'ons
DART Tutorial Sec'on 1: Filtering For a One Variable System UCAR The Na'onal Center for Atmospheric Research is sponsored by the Na'onal Science Founda'on. Any opinions, findings and conclusions or recommenda'ons
Characterizing transi.ng planets with JWST spectra: Simula.ons and Retrievals
 Characterizing transi.ng planets with JWST spectra: Simula.ons and Retrievals Tom Greene (NASA ARC) Michael Line (UCSC / Hubble Fellow / ARC), Jonathan Fortney (UCSC) October 15, 2015 Planet transmission
Characterizing transi.ng planets with JWST spectra: Simula.ons and Retrievals Tom Greene (NASA ARC) Michael Line (UCSC / Hubble Fellow / ARC), Jonathan Fortney (UCSC) October 15, 2015 Planet transmission
Microbiota: Its Evolution and Essence. Hsin-Jung Joyce Wu "Microbiota and man: the story about us
 Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
Microbiota: Its Evolution and Essence Overview q Define microbiota q Learn the tool q Ecological and evolutionary forces in shaping gut microbiota q Gut microbiota versus free-living microbe communities
CS 581 / BIOE 540: Algorithmic Computa<onal Genomics. Tandy Warnow Departments of Bioengineering and Computer Science hhp://tandy.cs.illinois.
 CS 581 / BIOE 540: Algorithmic Computa
CS 581 / BIOE 540: Algorithmic Computa
Introduction to Advanced Population Genetics
 Introduction to Advanced Population Genetics Learning Objectives Describe the basic model of human evolutionary history Describe the key evolutionary forces How demography can influence the site frequency
Introduction to Advanced Population Genetics Learning Objectives Describe the basic model of human evolutionary history Describe the key evolutionary forces How demography can influence the site frequency
Microsatellite evolution in Adélie penguins
 Microsatellite evolution in Adélie penguins Bennet McComish School of Mathematics and Physics Microsatellites Tandem repeats of motifs up to 6bp, e.g. (AC) 6 = ACACACACACAC Length is highly polymorphic.
Microsatellite evolution in Adélie penguins Bennet McComish School of Mathematics and Physics Microsatellites Tandem repeats of motifs up to 6bp, e.g. (AC) 6 = ACACACACACAC Length is highly polymorphic.
What to do with a billion years of evolution
 What to do with a billion years of evolution Melissa DeBiasse University of Florida Whitney Laboratory for Marine Bioscience @MelissaDeBiasse melissa.debiasse@gmail.com melissadebiasse.weebly.com Acknowledgements
What to do with a billion years of evolution Melissa DeBiasse University of Florida Whitney Laboratory for Marine Bioscience @MelissaDeBiasse melissa.debiasse@gmail.com melissadebiasse.weebly.com Acknowledgements
Gene Regulatory Networks II Computa.onal Genomics Seyoung Kim
 Gene Regulatory Networks II 02-710 Computa.onal Genomics Seyoung Kim Goal: Discover Structure and Func;on of Complex systems in the Cell Identify the different regulators and their target genes that are
Gene Regulatory Networks II 02-710 Computa.onal Genomics Seyoung Kim Goal: Discover Structure and Func;on of Complex systems in the Cell Identify the different regulators and their target genes that are
opulation genetics undamentals for SNP datasets
 opulation genetics undamentals for SNP datasets with crocodiles) Sam Banks Charles Darwin University sam.banks@cdu.edu.au I ve got a SNP genotype dataset, now what? Do my data meet the requirements of
opulation genetics undamentals for SNP datasets with crocodiles) Sam Banks Charles Darwin University sam.banks@cdu.edu.au I ve got a SNP genotype dataset, now what? Do my data meet the requirements of
Genotype Imputation. Biostatistics 666
 Genotype Imputation Biostatistics 666 Previously Hidden Markov Models for Relative Pairs Linkage analysis using affected sibling pairs Estimation of pairwise relationships Identity-by-Descent Relatives
Genotype Imputation Biostatistics 666 Previously Hidden Markov Models for Relative Pairs Linkage analysis using affected sibling pairs Estimation of pairwise relationships Identity-by-Descent Relatives
Mass of the electron m 0
 Mass of the electron m 0 1 Objective To determine the rest mass of the electron, m e, via γ-ray interactions (mainly Compton scattering and photoeffect) in a NaI scintillation detector. Based on the enclosed
Mass of the electron m 0 1 Objective To determine the rest mass of the electron, m e, via γ-ray interactions (mainly Compton scattering and photoeffect) in a NaI scintillation detector. Based on the enclosed
Introduction to Phylogenetic Analysis
 Introduction to Phylogenetic Analysis Tuesday 23 Wednesday 24 July, 2013 School of Biological Sciences Overview This workshop will provide an introduction to phylogenetic analysis, leading to a focus on
Introduction to Phylogenetic Analysis Tuesday 23 Wednesday 24 July, 2013 School of Biological Sciences Overview This workshop will provide an introduction to phylogenetic analysis, leading to a focus on
New method developments in pollen DNA barcoding for forensic applica8ons
 New method developments in pollen DNA barcoding for forensic applica8ons Karen Leanne Bell, Emory University Homeland Security Symposium Series, University of Texas El Paso, June 2016 1 Overview Background
New method developments in pollen DNA barcoding for forensic applica8ons Karen Leanne Bell, Emory University Homeland Security Symposium Series, University of Texas El Paso, June 2016 1 Overview Background
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Statistical Models for sequencing data: from Experimental Design to Generalized Linear Models
 Best practices in the analysis of RNA-Seq and CHiP-Seq data 4 th -5 th May 2017 University of Cambridge, Cambridge, UK Statistical Models for sequencing data: from Experimental Design to Generalized Linear
Best practices in the analysis of RNA-Seq and CHiP-Seq data 4 th -5 th May 2017 University of Cambridge, Cambridge, UK Statistical Models for sequencing data: from Experimental Design to Generalized Linear
Parallelizing Gaussian Process Calcula1ons in R
 Parallelizing Gaussian Process Calcula1ons in R Christopher Paciorek UC Berkeley Sta1s1cs Joint work with: Benjamin Lipshitz Wei Zhuo Prabhat Cari Kaufman Rollin Thomas UC Berkeley EECS (formerly) IBM
Parallelizing Gaussian Process Calcula1ons in R Christopher Paciorek UC Berkeley Sta1s1cs Joint work with: Benjamin Lipshitz Wei Zhuo Prabhat Cari Kaufman Rollin Thomas UC Berkeley EECS (formerly) IBM
Automated Illumina TruSight HLA v2 Sequencing Panel Library Preparation with the epmotion 5075t
 SHORT PROTOCOL No. 41 Automated Illumina TruSight HLA v2 Sequencing Panel Library Preparation with the epmotion 5075t Introduction This protocol describes the workstation configuration and pre-programmed
SHORT PROTOCOL No. 41 Automated Illumina TruSight HLA v2 Sequencing Panel Library Preparation with the epmotion 5075t Introduction This protocol describes the workstation configuration and pre-programmed
The applicability of next-generation sequencing to native plant materials development
 The applicability of next-generation sequencing to native plant materials development Rob Massatti, USGS-Southwest Biological Science Center Flagstaff, AZ Michael Luth www.blackfootnativeplants.com Next-generation
The applicability of next-generation sequencing to native plant materials development Rob Massatti, USGS-Southwest Biological Science Center Flagstaff, AZ Michael Luth www.blackfootnativeplants.com Next-generation
394C, October 2, Topics: Mul9ple Sequence Alignment Es9ma9ng Species Trees from Gene Trees
 394C, October 2, 2013 Topics: Mul9ple Sequence Alignment Es9ma9ng Species Trees from Gene Trees Mul9ple Sequence Alignment Mul9ple Sequence Alignments and Evolu9onary Histories (the meaning of homologous
394C, October 2, 2013 Topics: Mul9ple Sequence Alignment Es9ma9ng Species Trees from Gene Trees Mul9ple Sequence Alignment Mul9ple Sequence Alignments and Evolu9onary Histories (the meaning of homologous
CS 6140: Machine Learning Spring What We Learned Last Week. Survey 2/26/16. VS. Model
 Logis@cs CS 6140: Machine Learning Spring 2016 Instructor: Lu Wang College of Computer and Informa@on Science Northeastern University Webpage: www.ccs.neu.edu/home/luwang Email: luwang@ccs.neu.edu Assignment
Logis@cs CS 6140: Machine Learning Spring 2016 Instructor: Lu Wang College of Computer and Informa@on Science Northeastern University Webpage: www.ccs.neu.edu/home/luwang Email: luwang@ccs.neu.edu Assignment
Tools and Algorithms in Bioinformatics
 Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
CS 6140: Machine Learning Spring 2016
 CS 6140: Machine Learning Spring 2016 Instructor: Lu Wang College of Computer and Informa?on Science Northeastern University Webpage: www.ccs.neu.edu/home/luwang Email: luwang@ccs.neu.edu Logis?cs Assignment
CS 6140: Machine Learning Spring 2016 Instructor: Lu Wang College of Computer and Informa?on Science Northeastern University Webpage: www.ccs.neu.edu/home/luwang Email: luwang@ccs.neu.edu Logis?cs Assignment
Association Testing with Quantitative Traits: Common and Rare Variants. Summer Institute in Statistical Genetics 2014 Module 10 Lecture 5
 Association Testing with Quantitative Traits: Common and Rare Variants Timothy Thornton and Katie Kerr Summer Institute in Statistical Genetics 2014 Module 10 Lecture 5 1 / 41 Introduction to Quantitative
Association Testing with Quantitative Traits: Common and Rare Variants Timothy Thornton and Katie Kerr Summer Institute in Statistical Genetics 2014 Module 10 Lecture 5 1 / 41 Introduction to Quantitative
Misconceptions on Missing Data in RAD-seq Phylogenetics with a Deep-scale Example from Flowering Plants
 Syst. Biol. 66(3):399 412, 2017 The Author(s) 2016. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Syst. Biol. 66(3):399 412, 2017 The Author(s) 2016. Published by Oxford University Press, on behalf of the Society of Systematic Biologists. All rights reserved. For Permissions, please email: journals.permissions@oup.com
Evidence of adaptation from ancestral variation in young populations of beach mice
 Evidence of adaptation from ancestral variation in young populations of beach mice Journal: Evolution Manuscript ID: -0.R Manuscript Type: Original Article Date Submitted by the Author: -Jan- Complete
Evidence of adaptation from ancestral variation in young populations of beach mice Journal: Evolution Manuscript ID: -0.R Manuscript Type: Original Article Date Submitted by the Author: -Jan- Complete
Purging of inbreeding load in par;ally selfing popula;ons
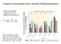 Purging of inbreeding load in par;ally selfing popula;ons Given an inbreeding coefficient of f, the equilibrium frequency is q e µ ( h(1 f ) + f )s Data from various Schiedea species with different extent
Purging of inbreeding load in par;ally selfing popula;ons Given an inbreeding coefficient of f, the equilibrium frequency is q e µ ( h(1 f ) + f )s Data from various Schiedea species with different extent
Invasion genomics and adaptation in Australian fireweed. Andrew Lowe Peter Prentis, Elly Dormontt University of Adelaide, Australia
 Invasion genomics and adaptation in Australian fireweed Andrew Lowe Peter Prentis, Elly Dormontt University of Adelaide, Australia S. vulgaris var. vulgaris S. squalidus S. eboracensis Oliver Payne & Nick
Invasion genomics and adaptation in Australian fireweed Andrew Lowe Peter Prentis, Elly Dormontt University of Adelaide, Australia S. vulgaris var. vulgaris S. squalidus S. eboracensis Oliver Payne & Nick
Learning Your Identity and Disease from Research Papers: Information Leaks in Genome-Wide Association Study
 Learning Your Identity and Disease from Research Papers: Information Leaks in Genome-Wide Association Study Rui Wang, Yong Li, XiaoFeng Wang, Haixu Tang and Xiaoyong Zhou Indiana University at Bloomington
Learning Your Identity and Disease from Research Papers: Information Leaks in Genome-Wide Association Study Rui Wang, Yong Li, XiaoFeng Wang, Haixu Tang and Xiaoyong Zhou Indiana University at Bloomington
Eppendorf twin.tec PCR Plates 96 LoBind Increase Yield of Transcript Species and Number of Reads of NGS Libraries
 APPLICATION NOTE No. 375 I December 2016 Eppendorf twin.tec PCR Plates 96 LoBind Increase Yield of Transcript Species and Number of Reads of NGS Libraries Hanae A. Henke¹, Björn Rotter² ¹Eppendorf AG,
APPLICATION NOTE No. 375 I December 2016 Eppendorf twin.tec PCR Plates 96 LoBind Increase Yield of Transcript Species and Number of Reads of NGS Libraries Hanae A. Henke¹, Björn Rotter² ¹Eppendorf AG,
Introduction to PLINK H3ABionet Course Covenant University, Nigeria
 UNIVERSITY OF THE WITWATERSRAND, JOHANNESBURG Introduction to PLINK H3ABionet Course Covenant University, Nigeria Scott Hazelhurst H3ABioNet funded by NHGRI grant number U41HG006941 Wits Bioinformatics
UNIVERSITY OF THE WITWATERSRAND, JOHANNESBURG Introduction to PLINK H3ABionet Course Covenant University, Nigeria Scott Hazelhurst H3ABioNet funded by NHGRI grant number U41HG006941 Wits Bioinformatics
EvoluDonary lineages and phylogenedcs. (from: Darwin, C The Origin of Species. John Murray, Albermarle St., London.)
 EvoluDonary lineages and phylogenedcs (from: Darwin, C. 1859. The Origin of Species. John Murray, Albermarle St., London.) 1 EvoluDonary lineages and phylogenedcs Overview of this sec.on: 1) Basic PhylogeneDcs
EvoluDonary lineages and phylogenedcs (from: Darwin, C. 1859. The Origin of Species. John Murray, Albermarle St., London.) 1 EvoluDonary lineages and phylogenedcs Overview of this sec.on: 1) Basic PhylogeneDcs
Going Beyond SNPs with Next Genera5on Sequencing Technology Personalized Medicine: Understanding Your Own Genome Fall 2014
 Going Beyond SNPs with Next Genera5on Sequencing Technology 02-223 Personalized Medicine: Understanding Your Own Genome Fall 2014 Next Genera5on Sequencing Technology (NGS) NGS technology Discover more
Going Beyond SNPs with Next Genera5on Sequencing Technology 02-223 Personalized Medicine: Understanding Your Own Genome Fall 2014 Next Genera5on Sequencing Technology (NGS) NGS technology Discover more
Systems Biology and Neuroengineering
 Systems Biology and Neuroengineering Dion Khodagholy Columbia University Electrical Engineering Department TRANSCENDING DISCIPLINES, TRANSFORMING LIVES What is Systems Biology and Neuroengineering? Development
Systems Biology and Neuroengineering Dion Khodagholy Columbia University Electrical Engineering Department TRANSCENDING DISCIPLINES, TRANSFORMING LIVES What is Systems Biology and Neuroengineering? Development
Probing the Dynamical History of Exoplanets: Spectroscopic Observa<ons of Transi<ng Systems
 June. 4, 215 @ NAOJ Probing the Dynamical History of Exoplanets: Spectroscopic Observa
June. 4, 215 @ NAOJ Probing the Dynamical History of Exoplanets: Spectroscopic Observa
Crust and Lithosphere
 Crust and Lithosphere Our Charge descrip5on of scien5fic problems importance for broader society importance of the topics within Earth and other sciences exis5ng and required resources for fundamental
Crust and Lithosphere Our Charge descrip5on of scien5fic problems importance for broader society importance of the topics within Earth and other sciences exis5ng and required resources for fundamental
Genomic data detect corresponding signatures of population size change on an ecological time scale in two salamander species
 Molecular Ecology (2017) 26, 1060 1074 doi: 10.1111/mec.13988 Genomic data detect corresponding signatures of population size change on an ecological time scale in two salamander species SCHYLER O. NUNZIATA,*
Molecular Ecology (2017) 26, 1060 1074 doi: 10.1111/mec.13988 Genomic data detect corresponding signatures of population size change on an ecological time scale in two salamander species SCHYLER O. NUNZIATA,*
High-throughput sequence alignment. November 9, 2017
 High-throughput sequence alignment November 9, 2017 a little history human genome project #1 (many U.S. government agencies and large institute) started October 1, 1990. Goal: 10x coverage of human genome,
High-throughput sequence alignment November 9, 2017 a little history human genome project #1 (many U.S. government agencies and large institute) started October 1, 1990. Goal: 10x coverage of human genome,
Dr. Jennifer Weller WORKFLOW DURING THE B3 CAMP MAKING SOLUTIONS FROM STOCKS. B3 Summer Science Camp at Olympic High School
 Dr. Jennifer Weller WORKFLOW DURING THE B3 CAMP MAKING SOLUTIONS FROM STOCKS B3 Summer Science Camp at Olympic High School LAB WORKFLOW OVERVIEW Collect Samples (June 12 th ) Extract DNA from the samples
Dr. Jennifer Weller WORKFLOW DURING THE B3 CAMP MAKING SOLUTIONS FROM STOCKS B3 Summer Science Camp at Olympic High School LAB WORKFLOW OVERVIEW Collect Samples (June 12 th ) Extract DNA from the samples
Machine Learning & Data Mining CS/CNS/EE 155. Lecture 11: Hidden Markov Models
 Machine Learning & Data Mining CS/CNS/EE 155 Lecture 11: Hidden Markov Models 1 Kaggle Compe==on Part 1 2 Kaggle Compe==on Part 2 3 Announcements Updated Kaggle Report Due Date: 9pm on Monday Feb 13 th
Machine Learning & Data Mining CS/CNS/EE 155 Lecture 11: Hidden Markov Models 1 Kaggle Compe==on Part 1 2 Kaggle Compe==on Part 2 3 Announcements Updated Kaggle Report Due Date: 9pm on Monday Feb 13 th
Hidden histories of gene flow in highland birds revealed with genomic markers
 Molecular Ecology (2016) 25, 5144 5157 doi: 10.1111/mec.13813 Hidden histories of gene flow in highland birds revealed with genomic markers EUGENIA ZARZA,* BRANT C. FAIRCLOTH, WHITNEY L.E. TSAI,* ROBERT
Molecular Ecology (2016) 25, 5144 5157 doi: 10.1111/mec.13813 Hidden histories of gene flow in highland birds revealed with genomic markers EUGENIA ZARZA,* BRANT C. FAIRCLOTH, WHITNEY L.E. TSAI,* ROBERT
Amy Driskell. Laboratories of Analytical Biology National Museum of Natural History Smithsonian Institution, Wash. DC
 DNA Barcoding Amy Driskell Laboratories of Analytical Biology National Museum of Natural History Smithsonian Institution, Wash. DC 1 Outline 1. Barcoding in general 2. Uses & Examples 3. Barcoding Bocas
DNA Barcoding Amy Driskell Laboratories of Analytical Biology National Museum of Natural History Smithsonian Institution, Wash. DC 1 Outline 1. Barcoding in general 2. Uses & Examples 3. Barcoding Bocas
Evolution before Darwin
 http://www.beatricebiologist.com Charles Lyell and Geology Geological forces could account for the differences in the world Con7nental Dri9 and Plate Tectonics Only book that Darwin took with him on his
http://www.beatricebiologist.com Charles Lyell and Geology Geological forces could account for the differences in the world Con7nental Dri9 and Plate Tectonics Only book that Darwin took with him on his
Single Cell Sequencing
 Single Cell Sequencing Fundamental unit of life Autonomous and unique Interactive Dynamic - change over time Evolution occurs on the cellular level Robert Hooke s drawing of cork cells, 1665 Type Prokaryotes
Single Cell Sequencing Fundamental unit of life Autonomous and unique Interactive Dynamic - change over time Evolution occurs on the cellular level Robert Hooke s drawing of cork cells, 1665 Type Prokaryotes
Ion Torrent. The chip is the machine
 Ion Torrent Introduction The Ion Personal Genome Machine [PGM] is simple, more costeffective, and more scalable than any other sequencing technology. Founded in 2007 by Jonathan Rothberg. Part of Life
Ion Torrent Introduction The Ion Personal Genome Machine [PGM] is simple, more costeffective, and more scalable than any other sequencing technology. Founded in 2007 by Jonathan Rothberg. Part of Life
Evolutionary factors and synthetic biology
 Evolutionary factors and synthetic biology NAS Joint Session on Climate Change and Ecology Owain Edwards Group Leader, Environmental Synthetic Genomics, CSIRO, Perth, Australia Domain Leader, Biocontrol
Evolutionary factors and synthetic biology NAS Joint Session on Climate Change and Ecology Owain Edwards Group Leader, Environmental Synthetic Genomics, CSIRO, Perth, Australia Domain Leader, Biocontrol
Levels of genetic variation for a single gene, multiple genes or an entire genome
 From previous lectures: binomial and multinomial probabilities Hardy-Weinberg equilibrium and testing HW proportions (statistical tests) estimation of genotype & allele frequencies within population maximum
From previous lectures: binomial and multinomial probabilities Hardy-Weinberg equilibrium and testing HW proportions (statistical tests) estimation of genotype & allele frequencies within population maximum
Integer Programming in Computational Biology. D. Gusfield University of California, Davis Presented December 12, 2016.!
 Integer Programming in Computational Biology D. Gusfield University of California, Davis Presented December 12, 2016. There are many important phylogeny problems that depart from simple tree models: Missing
Integer Programming in Computational Biology D. Gusfield University of California, Davis Presented December 12, 2016. There are many important phylogeny problems that depart from simple tree models: Missing
Gene Genealogies Coalescence Theory. Annabelle Haudry Glasgow, July 2009
 Gene Genealogies Coalescence Theory Annabelle Haudry Glasgow, July 2009 What could tell a gene genealogy? How much diversity in the population? Has the demographic size of the population changed? How?
Gene Genealogies Coalescence Theory Annabelle Haudry Glasgow, July 2009 What could tell a gene genealogy? How much diversity in the population? Has the demographic size of the population changed? How?
Collaborative topic models: motivations cont
 Collaborative topic models: motivations cont Two topics: machine learning social network analysis Two people: " boy Two articles: article A! girl article B Preferences: The boy likes A and B --- no problem.
Collaborative topic models: motivations cont Two topics: machine learning social network analysis Two people: " boy Two articles: article A! girl article B Preferences: The boy likes A and B --- no problem.
Mathematical models in population genetics II
 Mathematical models in population genetics II Anand Bhaskar Evolutionary Biology and Theory of Computing Bootcamp January 1, 014 Quick recap Large discrete-time randomly mating Wright-Fisher population
Mathematical models in population genetics II Anand Bhaskar Evolutionary Biology and Theory of Computing Bootcamp January 1, 014 Quick recap Large discrete-time randomly mating Wright-Fisher population
Inferring Species Trees Directly from Biallelic Genetic Markers: Bypassing Gene Trees in a Full Coalescent Analysis. Research article.
 Inferring Species Trees Directly from Biallelic Genetic Markers: Bypassing Gene Trees in a Full Coalescent Analysis David Bryant,*,1 Remco Bouckaert, 2 Joseph Felsenstein, 3 Noah A. Rosenberg, 4 and Arindam
Inferring Species Trees Directly from Biallelic Genetic Markers: Bypassing Gene Trees in a Full Coalescent Analysis David Bryant,*,1 Remco Bouckaert, 2 Joseph Felsenstein, 3 Noah A. Rosenberg, 4 and Arindam
