Evolutionary analysis of the genomic region of FOXP3 in vertebrates
|
|
|
- Conrad West
- 5 years ago
- Views:
Transcription
1 University of Arkansas, Fayetteville Biological Sciences Undergraduate Honors Theses Biological Sciences Evolutionary analysis of the genomic region of FOXP3 in vertebrates Ethan G. Guffey University of Arkansas, Fayetteville Follow this and additional works at: Recommended Citation Guffey, Ethan G., "Evolutionary analysis of the genomic region of FOXP3 in vertebrates" (2015). Biological Sciences Undergraduate Honors Theses This Thesis is brought to you for free and open access by the Biological Sciences at It has been accepted for inclusion in Biological Sciences Undergraduate Honors Theses by an authorized administrator of For more information, please contact
2 Evolutionary Analysis of the Genomic Region of FOXP3 in Vertebrates An Honors Thesis submitted in partial fulfillment of the requirements of Honors Studies in Biology By Ethan Guffey Spring 2015 Biology J. William Fulbright College of Arts and Sciences The University of Arkansas
3 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 2 Table of Contents Section Page Number Abstract 3 Introduction 4 Results 6 Discussion 10 References 14 Appendix 16
4 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 3 Abstract The Human FOXP3 gene plays a role in immunosuppression, which includes the suppression of the maternal immune system to allow the growth of a semi- foreign fetus inside her uterus. This has lead scientists to hypothesize that FOXP3 gene played a crucial role in the development of viviparous species. FOXP3 is highly conserved across Eutherian mammals, but the degree of conservation in other viviparous species has not been investigated. This study found that FOXP3 orthologs are much less highly conserved in aplacental species and may be more closely related to other members of the FOXP gene family than to the FOXP3 found in mammals. Through the creation of phylogenetic trees, I found that all of the members of the Forkhead Box Protein Family were named correctly, but FOXP3 had the highest degree of substitutions, which may explain its relatively novel function in immunosuppression. Keywords: FOXP3, forkhead box protein, viviparity, Eutheria, immunosuppression, orthologs
5 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 4 The Forkhead Box P3 gene, or FOXP3, belongs to a larger group called the forkhead box winged helix and more specifically the FOXP subfamily including FOXP1, FOXP2, FOXP3, and FOXP54, which are a group of transcription factors grouped together based DNA binding domain structure. The FOXP3 gene produces a protein by the same name that works as a transcription factor and serves as a master regulator in the immune system most importantly for CD4+ CD25+ T regulatory cell (T- reg) development and function (Andersen, Nissen, & Betz, 2012). It s role as master regulator means that it works on multiple genes that activate or inhibit immune function given certain conditions. For example, it is important in preventing autoimmunity, or self- tolerance, by deleting self- reactive T cells in the thymus, which is the site of T- reg development (Samstein, Josefewicz, Arvey, Treuting, & Rudensky, 2012). An additional function of the FOXP3 protein as the master regulator is the response to non- self antigens. During pregnancy, the mother constantly comes into contact with paternal antigens expressed by the fetus through the maternal- fetal barrier, known as the placenta. These non- self antigens trigger a response in the immune system of the mother that may result in a termination of the pregnancy. Obviously, termination of pregnancy would not allow for propagation of the species and the proliferation of the parental gene pool. For placental pregnancy to be viable, a mechanism had to evolve to allow maternal- fetal interaction and suppress these immune responses, and FOXP3 mediates this interaction (Andersen et al., 2012). Samstein et al. (2012) found that when female mice deficient in CNS1, an enhancer of FOXP3, were mated with males that were antigenically different from the female, they showed increased rates of spontaneous fetal absorption while the CNS1 positive individuals did not. This shows that sufficient expression of the FOXP3 gene plays a role in reducing the immune response of the mother to the growing fetus in the uterus.
6 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 5 FOXP3 is just one among many genes that have been recruited over the course of evolutionary time to slowly give rise to placentals. In mammalia major transitions exist in the form evolutionary novel phenotypes that are observable examples of the development of placentation (Lynch et al., 2015). For example, the monotremes are mammals that grow embryos in utero for 10 days transferring nutrients via a yolk- like placenta before laying thin, poorly mineralized eggs that hatch within 10 to 20 days. The next group of mammals appears to have embellished the yolk- sac into a structure more recognizable as the placenta. This evolutionary advancement marks the rise of viviparity, or live- birth, in a group called Theria, which includes both marsupials and Eutherians or placental mammals. While viviparity evolved only once in Eutherians, interestingly evidence suggests that it evolve over 100 times in squamous reptile species including lizards and snakes (Dykes & Beaupre, 2012). Marsupials have relatively short pregnancies of around 25 days, which is shorter than one estrous cycle. Placentals have evolved much longer pregnancies lasting on average 131 days but as much as 670 days. These longer pregnancies led to differentiation of cells in the uterus and induced large- scale gene regulation, which includes maternal immunotolerance of the semi- foreign fetus (Lynch et al., 2015; Murphy, Thompson, & Belov, 2009). Due to the unique function of the FOXP3 gene in viviparity, studies have suggested that FOXP3 is only found in placentals. According to Andersen et al. (2012), the forkhead box subfamily can only be found in jawed vertebrates. They went further to say that FOXP3- like genes could be found in bony fish, amphibians, and reptiles, but not in bird species. Samstein et al. (2012) found that the FOXP3 enhancer, CNS1, may have played a role in the development of viviparity in mammals through regulation of the FOXP3 gene. Andersen et al. (2012) specifically looked at the domains that make up FOXP3. These regions evolved in a step- wise fashion that allowed for the rise of placentals. They found that all
7 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 6 mammals contained the proline rich (ProR) region and the forkhead domain that functions in nuclear localization and DNA binding. They hypothesized that the very N- terminal tail of the ProR region, which was only found in placentals, may have increased the evolutionary differentiation from other FOXPs because it may aid in binding to factors specifically associated with T cell activation. The goal of this study was to analyze the genomic region surrounding FOXP3 in mammals and use that information to explore the genomes of mammals and other animals to find FOXP3 or FOXP3- like genes to determine its evolutionary origins, but the goal evolved into a search for orthologs of FOXP3 in other species in which it has not previously been found. Results FOXP3 In order to discover the origins of the FOXP3 gene and its role in placentation, sequences across all species included on NCBI blast were analyzed to determine which were most related to human FOXP3. A preliminary blastn of the NCBI NR nucleotide database with human FOXP3 DNA sequence (Accession: EF ) returned different isoforms of human FOXP3. A follow- up blastn search was conducted excluding the human taxid to remove other isoforms of the same gene. The results were all FOXP3 isoforms of mammalian species. To increase the likelihood of more comprehensive findings, blastp was used for the remainder of this study rather than blastn because amino acid sequences are more likely to be highly conserved across species than nucleotide sequences due to the degenerate nature of the genetic code. Additionally, substitution of an amino acid with similar properties would lead to
8 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 7 similar protein structures. For example, a non- polar amino acid such as alanine would likely offer very little change if substituted for another non- polar amino acid such as leucine. The same can be said for other groups of amino acids with acidic and basic sidechains. A blastp of the NCBI protein database using the human FOXP3 protein (Accession: ABQ ) showed high percent identity and high query coverage among mammals with the most highly conserved being among primate species including chimpanzee (Pan troglodytes), bonobo (Pan paniscus), gorilla (Gorilla gorilla), and macaque (Macaca fascicularis, Figure 1). All four of these species have a 99% identity and 100% query coverage, which suggests very similar sequences between these species and human FOXP3. Since the overall genomes of chimpanzee and these other species are the most closely related genomes to humans, these results seem reasonable. Cow (Bos taurus), cat (Felis catus), camel (Camelus ferus), armadillo (Dasypus novemcinctus), rhinoceros (Ceratotherium simum), and horse (Equus caballus) are all non- primates but have 100% query coverage and at least 90% identity. However, all of these species are mammals. It is likely that mammals would share more closely related sequences to each other than to non- mammalian vertebrates. Figure 2 shows a visual representation of the blastp results with some additional information about taxonomy. All of these species mentioned above belong to a larger group called Eutheria, more commonly known as placentals. Considering the function of the products of the FOXP3 gene in live- bearing mothers, it stands to reason that this gene would be highly conserved across placentals. It is probable that the advantageous mutation occurred in a common ancestor of placentals that allowed for the successful development of viviparous species, because of the function of the FOXP3 gene in immunosuppression of the mother. The highly conserved nature of the FOXP3 gene across mammals leads to looking outside this group into other species including amphibians, fish, and birds to determine whether these contain
9 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 8 FOXP3. If a FOXP3 appears in these species but in a much less highly conserved form, it may suggest that the mutation entered the genome in a more distant common ancestor than previously thought. A blastp of the NCBI protein database search was completed using human FOXP3 as the query but excluding the entire taxid of mammals. This allowed a closer look at species believed by previous studies not to contain FOXP3. The results showed much lower query coverage and percent identity than the search including mammals (Figure 3). These results include genes named for other members of the forkhead box family including FOXP1 and FOXP2 in addition to a select few named FOXP3. In contrast with Figure 1, the results including non- mammalian vertebrates (notably they are aplacental) show query coverage from 50-60% and percent identity from 40-50%. Since the gene products of FOXP3 do not have a known equivalent in aplacental vertebrates, there appears to be a lack of fitness advantage and therefore no selection in these species, which may account for the varying degree of amino acid substitutions in non- Eutherian species. This suggests that the fitness advantage of FOXP3 in placentals serves to support the high similarity of the gene across the majority of mammals. Forkhead Box Protein Family Aside from the low coverage and identity, FOXP3 appears to be largely non- existent outside of mammalian species with a few exceptions. When FOXP3 is conserved in non- mammalian vertebrates, as in the case of some frogs (Xenopus tropicalis) and fish (Dano rerio), it is much less highly conserved than in mammals. Other than those few species where there is a counterpart, FOXP3 in fish, reptiles, and birds appears to be much more similar to other members of the forkhead box protein family, FOXP1, FOXP2, and FOXP4, than to forms of FOXP3
10 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 9 in other species as shown in Figure 3. We then considered whether some of the genes in these other lineages might be misnamed, which might account for absence of the gene named, FOXP3 in other species. To determine whether the members of the forkhead box family are named properly, each member of the forkhead box family was analyzed, and a chromosomal region was obtained including each gene and their adjacent genes. It is likely that adjacent genes either upstream or downstream of the gene in question are likely to be linked, meaning that these genes are very unlikely to separate over the course of time in a species genome. Throughout the course of evolutionary time, these genes are likely to remain in proximity even if that region is translocated to another chromosome. I therefore considered whether I could identify forkhead protein orthologs by their gene neighborhood. The hypothesis was that the true ortholog of FOXP3 would have the some of the same proximal genes in different lineages. The region of interest, or the FOXP3 gene region, is located on the X chromosome in humans. According to figure 4, genes within close proximity to FOXP3 are called PPP1R3F, which is a member of the protein phosphatase 1 family, and CCDC22, which is part of the coiled- coil domain family. In addition to FOXP3, the gene neighborhoods of FOXP1 (Figure 5), FOXP2 (Figure 6), and FOXP4 (Figure 7) were analyzed as well in case they are misnamed in other species. EIF4E3 is in close proximity to FOXP1. FOXP2 has two genes nearby called PPP1R3A and MDFIC. MDFI is in close proximity to FOXP4. I used the MultiAlign program in the LaserGene software package (DNAStar, Madison, WI) to generate sequence alignments and distance trees for human (Homo sapien), mouse (Mus musculus), Chicken (Gallus gallus), American alligator (Alligator mississipiensis), western painted turtle (Chrysemys picta bellii), cobra (Ophiophagus hannah), python (Python bivittatus), and
11 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 10 western clawed frog (Xenopus tropicalis) to analyze the gene families of Forkhead Box Protein (FOXP_, Figure 8), Protein Phosphatase 1 (PPP1R3_, Figure 9), Coiled- Coil Domain Containing (CCDC_, Figure 10), and MyoD Family Inhibitor (MDFI, Figure 11). From Figure 8 the forkhead box protein family members are different from one another meaning they are evolutionarily distinct. A closer look at the forkhead box protein family tree reveals that clades of FOXP1, FOXP2, and FOXP4 differ by fewer than 30 amino acid substitutions. However, FOXP3 has many more substitutions with the fewest being 60 substitutions in both cobra (Ophiophagus hannah) and python (Pyton bivittatus). According to these trees, each separate gene forms its own clade, which implies that all of the genes are named correctly in the NCBI blast database. The inconsistent nature of the amino acid sequence for FOXP3, as evidenced by the high degree of substitutions (Figure 8) and the low percent identity and query coverage (Figure 3), shows that this gene is the least highly conserved of the forkhead box family across species and possibly the newest paralog of this family that arose from another member of this family. In terms of evolution, FOXP3 appears to be the most diverse gene within the forkhead box protein family, which might suggest the reason for the relatively novel function in the gene of this family that may have aided in the rise of placentals and viviparity. Discussion The FOXP3 protein is highly conserved across mammalian species with highest identity and coverage among the various primate species. The FOXP3 gene is the most diverse paralog of the forkhead box protein family and, from these results, appears to be highly conserved in placentals with few clear orthologs in non- placental species. Representative orthologs could be
12 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 11 identified in some species of frogs and snakes. FOXP3 has more substitutions than any other member of this family, which may explain the reason for the relatively diverse gene products. Interestingly there is a commonality between genes proximal to FOXP2 and those of FOXP3 and FOXP4. Both FOXP2 and FOXP3 regions have neighboring genes from the protein phosphatase 1 family (PPP1R3A and PPP1R3F, respectively). Also the FOXP2 region shares a gene neighbor with the FOXP4 family MyoD Family Inhibitor (MDFIC and MDFI, respectively). Because of the gene maps of the forkhead box family (Figures 4-7), FOXP2 appears to have the most similarity to the other family members. For example, the FOXP2 gene region includes genes from the Protein Phosphatase 1 family, which is also in the FOXP3 region, and the MyoD Inhibitor family, which is also in the FOXP4 region. It is possible that the forkhead box family arose in a paralogous nature in which a whole or partial genome duplication gave rise to duplicate copies of the original ancestor of this gene family. The commonly shared genes in the region around FOXP2 with other forkhead box members may be evidence that FOXP2 is closest to the ancestor of this family, but it has since undergone its own evolutionary divergence. Of course more research needs to be conducted on all members of the forkhead box protein family with specific focus on the regions around the genes and how they relate to each other to determine the validity of this conjecture of the paralogous nature of the rise members of this family. Due to the interests of my research laboratory, I specifically searched the chicken (Gallus gallus) genome for sequences similar to human FOXP3 but no such gene was found. The result of all searches of the chicken genome using the human FOXP3 as query resulted in different isoforms of FOXP2. After the majority of research was completed for this project, it was brought to my attention that the chicken genome is not complete. This genome is missing
13 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 12 sequences for chromosome 16 and many of the microchromosomes. It is possible that a FOXP3 ortholog does exist in chicken but that the region containing it has not yet been sequenced. This revealing gap in the knowledge about this genome brings concern to other genomes as well. During my research, I was unaware of these gaps, so future research should be done to determine which genomes used in these findings contain sequenced chromosomes and regions. Additionally, new methods of sequencing should also be researched allow the possibility to fill in these gaps. Due to the function of the FOXP3 protein in suppressing the immune system, additional research may have practical applications in medicine. The autoimmune disorder IPEX syndrome, or Immunodysregulation Polyendocrinopathy Enteropathy X- linked syndrome, results from mutations in FOXP3 that result in non- conservative substitutions rendering the protein ineffective by changing the shape of the protein s structure or prematurely stopping translation creating a shorter, incomplete protein (Passerini, Santoni, Roncarolo, & Bacchetta, 2014). This ineffective or incomplete protein will not be able to bind correctly to DNA and serve as an enhancer for genes that lead to CD4+ CD25+ T regulator cell production. As a result there will be little or no regulatory response due to lower serum levels of T- regulator cells in the blood and no way to suppress immune responses including those against the individual s own body. This is known to cause diseases such as insulin dependent diabetes, eczema, and food allergies. This is a recessive, X- linked trait, therefore males are more commonly affected than females, which are more commonly carriers. Research into the origin of this gene may offer insight into the structure that may lead to potential gene therapies that bolster the levels of T- reg cells when needed to reduce the immune responses and promote self- tolerance. Similarly women who have difficulty carrying their pregnancy to term as a result of spontaneous abortions may benefit from this research. Pregnant women generally have
14 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 13 elevated levels of T- reg cells in their blood than women who are not pregnant. However, this elevated level of T- reg cells was not found in women experiencing unexplained recurrent spontaneous abortions (URSA), which is defined as two or more consecutive losses of pregnancy before twenty weeks (Zaigui et al., 2012). With reduced T- reg counts, these women are unable to suppress the immune responses triggered by the introduction of the paternal antigens by the fetus. The lower level of T- reg in the blood of URSA patients is likely caused by reduced expression or functionality of FOXP3. Again it appears that mutations in FOXP3 are the cause of reduced functionality. Research needs to be done to determine more specifically which regions of this gene are prone to mutations that results in undesired loss of pregnancy in these women and to determine if any form of therapy can be implemented to raise the T- reg cell counts of these women during pregnancy to prevent spontaneous abortions. With continued research into the origins and sequence variations of FOXP3, it may be possible to increase the quality of life in people affected by mutations that result in deficient forms of this immunologically significant protein.
15 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 14 References Andersen, K. G., Nissen, J. K., & Betz, A. G. (2012). Comparative Genomics Reveals Key Gain- of- Function Events in Foxp3 during Regulatory T Cell Evolution. Frontiers In Immunology, doi: /fimmu Lynch, V. J., Nnamani, M. C., Kapusta, A., Brayer, K., Plaza, S. L., Mazur, E. C., &... Wagner, G. P. (2015). Ancient Transposable Elements Transformed the Uterine Regulatory Landscape and Transcriptome during the Evolution of Mammalian Pregnancy. Cell Reports, doi: /j.celrep Murphy, B. F., Belov, K., & Thompson, M. B. (2010). Evolution of viviparity and uterine angiogenesis: vascular endothelial growth factor (VEGF) in oviparous and viviparous skinks. Journal Of Experimental Zoology. Part B, Molecular And Developmental Evolution, 314(2), doi: /jez.b O'Leary, M. On the Trail of the First Placental Mammals. American Scientist [serial online]. May 2014; 102(3): Available from: Academic Search Complete, Ipswich, MA. Accessed March 18, Passerini, L., Santoni de Sio, F. R., Roncarolo, M. G., & Bacchetta, R. (2014). Forkhead box P3: The Peacekeeper of the Immune System. International Reviews Of Immunology, 33(2), doi: / Samstein, R. M., Josefowicz, S. Z., Arvey, A., Treuting, P. M., & Rudensky, A. Y. (2012). Extrathymic generation of regulatory T cells in placental mammals mitigates maternal- fetal conflict. Cell, 150(1), doi: /j.cell
16 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 15 Van Dyke, J. U., & Beaupre, S. J. (2012). Stable isotope tracer reveals that viviparous snakes transport amino acids to offspring during gestation.the Journal Of Experimental Biology, 215(Pt 5), doi: /jeb Zaigui, W., Zeshan, Y., Cai, Z., Zhuyu, L., Xiumei, S., Xiuming, Z., & Yinguang, L. (2012). Association between Functional Polymorphisms of Foxp3 Gene and the Occurrence of Unexplained Recurrent Spontaneous Abortion in a Chinese Han Population. Clinical & Developmental Immunology, 1-7. doi: /2012/896458
17 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 16 Appendix Figure 1 Results of Blastp of human FOXP3 (ABQ15210) on NR protein database
18 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 17 Figure 2Taxonomy report of blastp of human FOXP3.
19 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 18 Figure 3 Blastp of human FOXP3 (ABQ15210) excluding mammals Figure 4 Map of human FOXP3 gene region
20 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 19 Figure 5 Map of human FOXP1 gene region Figure 6 Map of human FOXP2 gene region Figure 7 Map of human FOXP4 gene region
21 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 20 Figure 8 Distance tree of vertebrate Forkhead Box family. [Key: human (Homo sapien, Hsa ), mouse (Mus musculus, Mmu ), Chicken (Gallus gallus, Gga ), American alligator (Alligator mississipiensis, Ami ), western painted turtle (Chrysemys picta bellii, Cpb ), cobra (Ophiophagus hannah, Oha ), python (Python bivittatus, Pbi ), and western clawed frog (Xenopus tropicalis, Xtr )]
22 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 21 Figure 9 Distance Tree of Protein Phosphatase 1 family. [Key: same as Figure 8]
23 EVOLUTIONARY ANALYSIS OF THE GENOMIC REGION OF FOXP3 IN VERTEBRATES 22 Figure 10 Distance tree of Coiled- Coil Domain family. [Key: same as Figure 8] Figure 11 Distance Tree of MyoD Family Inhibitor [Key: same as Figure 8]
Supplemental Figure 1.
 Supplemental Material: Annu. Rev. Genet. 2015. 49:213 42 doi: 10.1146/annurev-genet-120213-092023 A Uniform System for the Annotation of Vertebrate microrna Genes and the Evolution of the Human micrornaome
Supplemental Material: Annu. Rev. Genet. 2015. 49:213 42 doi: 10.1146/annurev-genet-120213-092023 A Uniform System for the Annotation of Vertebrate microrna Genes and the Evolution of the Human micrornaome
Piecing It Together. 1) The envelope contains puzzle pieces for 5 vertebrate embryos in 3 different stages of
 Piecing It Together 1) The envelope contains puzzle pieces for 5 vertebrate embryos in 3 different stages of development. Lay out the pieces so that you have matched up each animal name card with its 3
Piecing It Together 1) The envelope contains puzzle pieces for 5 vertebrate embryos in 3 different stages of development. Lay out the pieces so that you have matched up each animal name card with its 3
Genomes and Their Evolution
 Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 21 Genomes and Their Evolution PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Biased amino acid composition in warm-blooded animals
 Biased amino acid composition in warm-blooded animals Guang-Zhong Wang and Martin J. Lercher Bioinformatics group, Heinrich-Heine-University, Düsseldorf, Germany Among eubacteria and archeabacteria, amino
Biased amino acid composition in warm-blooded animals Guang-Zhong Wang and Martin J. Lercher Bioinformatics group, Heinrich-Heine-University, Düsseldorf, Germany Among eubacteria and archeabacteria, amino
Comparing Genomes! Homologies and Families! Sequence Alignments!
 Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Comparing Genomes! Homologies and Families! Sequence Alignments! Allows us to achieve a greater understanding of vertebrate evolution! Tells us what is common and what is unique between different species
Chapter 16: Reconstructing and Using Phylogenies
 Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES
 Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Molecular Biology-2018 1 Definitions: RELATIONSHIPS BETWEEN GENES/PROTEINS HOMOLOGUES Heterologues: Genes or proteins that possess different sequences and activities. Homologues: Genes or proteins that
Science in Motion Ursinus College
 Science in Motion Ursinus College NAME EVIDENCE FOR EVOLUTION LAB INTRODUCTION: Evolution is not just a historical process; it is occurring at this moment. Populations constantly adapt in response to changes
Science in Motion Ursinus College NAME EVIDENCE FOR EVOLUTION LAB INTRODUCTION: Evolution is not just a historical process; it is occurring at this moment. Populations constantly adapt in response to changes
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Lecture 11 Friday, October 21, 2011
 Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Visit to BPRC. Data is crucial! Case study: Evolution of AIRE protein 6/7/13
 Visit to BPRC Adres: Lange Kleiweg 161, 2288 GJ Rijswijk Utrecht CS à Den Haag CS 9:44 Spoor 9a, arrival 10:22 Den Haag CS à Delft 10:28 Spoor 1, arrival 10:44 10:48 Delft Voorzijde à Bushalte TNO/Lange
Visit to BPRC Adres: Lange Kleiweg 161, 2288 GJ Rijswijk Utrecht CS à Den Haag CS 9:44 Spoor 9a, arrival 10:22 Den Haag CS à Delft 10:28 Spoor 1, arrival 10:44 10:48 Delft Voorzijde à Bushalte TNO/Lange
1 Introduction. Abstract
 CBS 530 Assignment No 2 SHUBHRA GUPTA shubhg@asu.edu 993755974 Review of the papers: Construction and Analysis of a Human-Chimpanzee Comparative Clone Map and Intra- and Interspecific Variation in Primate
CBS 530 Assignment No 2 SHUBHRA GUPTA shubhg@asu.edu 993755974 Review of the papers: Construction and Analysis of a Human-Chimpanzee Comparative Clone Map and Intra- and Interspecific Variation in Primate
DO NOT WRITE ON THIS. Evidence from Evolution Activity. The Fossilization Process. Types of Fossils
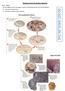 Evidence from Evolution Activity Part 1 - Fossils Use the diagrams on the next page to answer the following questions IN YOUR NOTEBOOK. 1. Describe how fossils form. 2. Describe the different types of
Evidence from Evolution Activity Part 1 - Fossils Use the diagrams on the next page to answer the following questions IN YOUR NOTEBOOK. 1. Describe how fossils form. 2. Describe the different types of
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
Evolutionary change. Evolution and Diversity. Two British naturalists, one revolutionary idea. Darwin observed organisms in many environments
 Evolutionary change Evolution and Diversity Ch 13 How populations evolve Organisms change over time In baby steps Species (including humans) are descended from other species Two British naturalists, one
Evolutionary change Evolution and Diversity Ch 13 How populations evolve Organisms change over time In baby steps Species (including humans) are descended from other species Two British naturalists, one
Evolution. Changes over Time
 Evolution Changes over Time TEKS Students will analyze and evaluate B. 7 C how natural selection produces change in populations, not individuals B. 7 E/F effects of genetic mechanisms and their relationship
Evolution Changes over Time TEKS Students will analyze and evaluate B. 7 C how natural selection produces change in populations, not individuals B. 7 E/F effects of genetic mechanisms and their relationship
Chapter 26: Phylogeny and the Tree of Life
 Chapter 26: Phylogeny and the Tree of Life 1. Key Concepts Pertaining to Phylogeny 2. Determining Phylogenies 3. Evolutionary History Revealed in Genomes 1. Key Concepts Pertaining to Phylogeny PHYLOGENY
Chapter 26: Phylogeny and the Tree of Life 1. Key Concepts Pertaining to Phylogeny 2. Determining Phylogenies 3. Evolutionary History Revealed in Genomes 1. Key Concepts Pertaining to Phylogeny PHYLOGENY
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
USING BLAST TO IDENTIFY PROTEINS THAT ARE EVOLUTIONARILY RELATED ACROSS SPECIES
 USING BLAST TO IDENTIFY PROTEINS THAT ARE EVOLUTIONARILY RELATED ACROSS SPECIES HOW CAN BIOINFORMATICS BE USED AS A TOOL TO DETERMINE EVOLUTIONARY RELATIONSHPS AND TO BETTER UNDERSTAND PROTEIN HERITAGE?
USING BLAST TO IDENTIFY PROTEINS THAT ARE EVOLUTIONARILY RELATED ACROSS SPECIES HOW CAN BIOINFORMATICS BE USED AS A TOOL TO DETERMINE EVOLUTIONARY RELATIONSHPS AND TO BETTER UNDERSTAND PROTEIN HERITAGE?
Many of the slides that I ll use have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Many of the slides that I ll use have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Many of the slides that I ll use have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Evidence of Evolution
 Evidence of Evolution Biogeography The Age of Earth and Fossils Ancient artiodactyl Modern whale Ancestors of Whales Ambulocetus could both swim in shallow water and walk on land. Rodhocetus probably spent
Evidence of Evolution Biogeography The Age of Earth and Fossils Ancient artiodactyl Modern whale Ancestors of Whales Ambulocetus could both swim in shallow water and walk on land. Rodhocetus probably spent
Evidence for Evolution
 Evidence for Evolution 1. 2. 3. 4. 5. Paleontology Comparative Anatomy Embryology Comparative Biochemistry Geographical Distribution How old is everything? The History of Earth as a Clock Station 1: Paleontology
Evidence for Evolution 1. 2. 3. 4. 5. Paleontology Comparative Anatomy Embryology Comparative Biochemistry Geographical Distribution How old is everything? The History of Earth as a Clock Station 1: Paleontology
Biology EOC Review Study Questions
 Biology EOC Review Study Questions Microscopes and Characteristics of Life 1. How do you calculate total magnification on a compound light microscope? 2. What is the basic building block of all living
Biology EOC Review Study Questions Microscopes and Characteristics of Life 1. How do you calculate total magnification on a compound light microscope? 2. What is the basic building block of all living
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Emily Blanton Phylogeny Lab Report May 2009
 Introduction It is suggested through scientific research that all living organisms are connected- that we all share a common ancestor and that, through time, we have all evolved from the same starting
Introduction It is suggested through scientific research that all living organisms are connected- that we all share a common ancestor and that, through time, we have all evolved from the same starting
08/21/2017 BLAST. Multiple Sequence Alignments: Clustal Omega
 BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
BLAST Multiple Sequence Alignments: Clustal Omega What does basic BLAST do (e.g. what is input sequence and how does BLAST look for matches?) Susan Parrish McDaniel College Multiple Sequence Alignments
Full file at CHAPTER 2 Genetics
 CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
Reproduction and Evolution Practice Exam
 Reproduction and Evolution Practice Exam Topics: Genetic concepts from the lecture notes including; o Mitosis and Meiosis, Homologous Chromosomes, Haploid vs Diploid cells Reproductive Strategies Heaviest
Reproduction and Evolution Practice Exam Topics: Genetic concepts from the lecture notes including; o Mitosis and Meiosis, Homologous Chromosomes, Haploid vs Diploid cells Reproductive Strategies Heaviest
thebiotutor.com AS Biology Unit 2 Classification, Adaptation & Biodiversity
 thebiotutor.com AS Biology Unit 2 Classification, Adaptation & Biodiversity 1 Classification and taxonomy Classification Phylogeny Taxonomy The process of sorting living things into groups. The study of
thebiotutor.com AS Biology Unit 2 Classification, Adaptation & Biodiversity 1 Classification and taxonomy Classification Phylogeny Taxonomy The process of sorting living things into groups. The study of
BIOLOGY FINAL EXAM REVIEW SHEET Chapters 10-15, 17-30
 Name Hour Due Date: BIOLOGY FINAL EXAM REVIEW SHEET Chapters 10-15, 17-30 The exam was prepared by the Biology teachers in the science departments of CVHS and DHS. 1. What is a Punnett Square? 2. Cross
Name Hour Due Date: BIOLOGY FINAL EXAM REVIEW SHEET Chapters 10-15, 17-30 The exam was prepared by the Biology teachers in the science departments of CVHS and DHS. 1. What is a Punnett Square? 2. Cross
How Biological Diversity Evolves
 CHAPTER 14 How Biological Diversity Evolves PowerPoint Lectures for Essential Biology, Third Edition Neil Campbell, Jane Reece, and Eric Simon Essential Biology with Physiology, Second Edition Neil Campbell,
CHAPTER 14 How Biological Diversity Evolves PowerPoint Lectures for Essential Biology, Third Edition Neil Campbell, Jane Reece, and Eric Simon Essential Biology with Physiology, Second Edition Neil Campbell,
Review sheet for Mendelian genetics through human evolution. What organism did Mendel study? What characteristics of this organism did he examine?
 Review sheet for Mendelian genetics through human evolution WARNING: I have tried to be complete, but I may have missed something. You are responsible for all the material discussed in class. This is only
Review sheet for Mendelian genetics through human evolution WARNING: I have tried to be complete, but I may have missed something. You are responsible for all the material discussed in class. This is only
Constructing a Pedigree
 Constructing a Pedigree Use the appropriate symbols: Unaffected Male Unaffected Female Affected Male Affected Female Male carrier of trait Mating of Offspring 2. Label each generation down the left hand
Constructing a Pedigree Use the appropriate symbols: Unaffected Male Unaffected Female Affected Male Affected Female Male carrier of trait Mating of Offspring 2. Label each generation down the left hand
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Biodiversity. The Road to the Six Kingdoms of Life
 Biodiversity The Road to the Six Kingdoms of Life How the 6 kingdoms came about At first, only two kingdoms were recognized Then Haeckel proposed a third kingdom Protista (where protists had both plant
Biodiversity The Road to the Six Kingdoms of Life How the 6 kingdoms came about At first, only two kingdoms were recognized Then Haeckel proposed a third kingdom Protista (where protists had both plant
3/8/ Complex adaptations. 2. often a novel trait
 Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Biodiversity. The Road to the Six Kingdoms of Life
 Biodiversity The Road to the Six Kingdoms of Life How the 6 kingdoms came about At first, only two kingdoms were recognized Then Haeckel proposed a third kingdom Protista (where protists had both plant
Biodiversity The Road to the Six Kingdoms of Life How the 6 kingdoms came about At first, only two kingdoms were recognized Then Haeckel proposed a third kingdom Protista (where protists had both plant
Review sheet for the material covered by exam III
 Review sheet for the material covered by exam III WARNING: Like last time, I have tried to be complete, but I may have missed something. You are responsible for all the material discussed in class. This
Review sheet for the material covered by exam III WARNING: Like last time, I have tried to be complete, but I may have missed something. You are responsible for all the material discussed in class. This
Bioinformatics Exercises
 Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Bioinformatics Exercises AP Biology Teachers Workshop Susan Cates, Ph.D. Evolution of Species Phylogenetic Trees show the relatedness of organisms Common Ancestor (Root of the tree) 1 Rooted vs. Unrooted
Biology Chapter 15 Evolution Notes
 Biology Chapter 15 Evolution Notes Section 1: Darwin's Theory of Evolution by Natural Selection Charles Darwin- English naturalist that studied animals over a number of years before developing the theory
Biology Chapter 15 Evolution Notes Section 1: Darwin's Theory of Evolution by Natural Selection Charles Darwin- English naturalist that studied animals over a number of years before developing the theory
Mechanisms of Evolution Darwinian Evolution
 Mechanisms of Evolution Darwinian Evolution Descent with modification by means of natural selection All life has descended from a common ancestor The mechanism of modification is natural selection Concept
Mechanisms of Evolution Darwinian Evolution Descent with modification by means of natural selection All life has descended from a common ancestor The mechanism of modification is natural selection Concept
Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution
 Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution Background How does an evolutionary biologist decide how closely related two different species are? The simplest way is to compare
Bioinformatics: Investigating Molecular/Biochemical Evidence for Evolution Background How does an evolutionary biologist decide how closely related two different species are? The simplest way is to compare
Reading for Lecture 13 Release v10
 Reading for Lecture 13 Release v10 Christopher Lee November 15, 2011 Contents 1 Evolutionary Trees i 1.1 Evolution as a Markov Process...................................... ii 1.2 Rooted vs. Unrooted Trees........................................
Reading for Lecture 13 Release v10 Christopher Lee November 15, 2011 Contents 1 Evolutionary Trees i 1.1 Evolution as a Markov Process...................................... ii 1.2 Rooted vs. Unrooted Trees........................................
Understanding relationship between homologous sequences
 Molecular Evolution Molecular Evolution How and when were genes and proteins created? How old is a gene? How can we calculate the age of a gene? How did the gene evolve to the present form? What selective
Molecular Evolution Molecular Evolution How and when were genes and proteins created? How old is a gene? How can we calculate the age of a gene? How did the gene evolve to the present form? What selective
HEREDITY AND EVOLUTION
 HEREDITY AND EVOLUTION 1. What is a gene? Answer. Gene is the unit of inheritance. Gene is the part of a chromosome which controls the appearance of a set of hereditary characteristics. 2. What is meant
HEREDITY AND EVOLUTION 1. What is a gene? Answer. Gene is the unit of inheritance. Gene is the part of a chromosome which controls the appearance of a set of hereditary characteristics. 2. What is meant
Hands-On Nine The PAX6 Gene and Protein
 Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Hands-On Nine The PAX6 Gene and Protein Main Purpose of Hands-On Activity: Using bioinformatics tools to examine the sequences, homology, and disease relevance of the Pax6: a master gene of eye formation.
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2017-2018 GENETICS BIO-5009A Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2017-2018 GENETICS BIO-5009A Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section
Anatomy. Species may share similar physical features because the feature was present in a common ancestor (homologous and analogous structures).
 Evidence for Evolution Evidence for evolution comes from many different areas of biology: Anatomy. Species may share similar physical features because the feature was present in a common ancestor (homologous
Evidence for Evolution Evidence for evolution comes from many different areas of biology: Anatomy. Species may share similar physical features because the feature was present in a common ancestor (homologous
Homework Assignment, Evolutionary Systems Biology, Spring Homework Part I: Phylogenetics:
 Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST
 Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Investigation 3: Comparing DNA Sequences to Understand Evolutionary Relationships with BLAST Introduction Bioinformatics is a powerful tool which can be used to determine evolutionary relationships and
Evidence of Evolution by Natural Selection. Dodo bird
 Evidence of Evolution by Natural Selection Dodo bird 2007-2008 Evidence supporting evolution Fossil record transition species Anatomical record homologous & vestigial structures embryology & development
Evidence of Evolution by Natural Selection Dodo bird 2007-2008 Evidence supporting evolution Fossil record transition species Anatomical record homologous & vestigial structures embryology & development
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Phylogenetic inference
 Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Group activities: Making animal model of human behaviors e.g. Wine preference model in mice
 Lecture schedule 3/30 Natural selection of genes and behaviors 4/01 Mouse genetic approaches to behavior 4/06 Gene-knockout and Transgenic technology 4/08 Experimental methods for measuring behaviors 4/13
Lecture schedule 3/30 Natural selection of genes and behaviors 4/01 Mouse genetic approaches to behavior 4/06 Gene-knockout and Transgenic technology 4/08 Experimental methods for measuring behaviors 4/13
BIOINFORMATICS LAB AP BIOLOGY
 BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
BIOINFORMATICS LAB AP BIOLOGY Bioinformatics is the science of collecting and analyzing complex biological data. Bioinformatics combines computer science, statistics and biology to allow scientists to
Phylogenetic analysis. Characters
 Typical steps: Phylogenetic analysis Selection of taxa. Selection of characters. Construction of data matrix: character coding. Estimating the best-fitting tree (model) from the data matrix: phylogenetic
Typical steps: Phylogenetic analysis Selection of taxa. Selection of characters. Construction of data matrix: character coding. Estimating the best-fitting tree (model) from the data matrix: phylogenetic
Name: Hour: Teacher: ROZEMA. Inheritance & Mutations Connected to Speciation
 Name: Hour: Teacher: ROZEMA Inheritance & Mutations Connected to Speciation Let s Review What We Already Know: What Have We Learned? Lesson 26: PI 1 (Projected Image) - Human Karyotype (image from https://en.wikipedia.org/wiki/karyotype#/media/file:nhgri_human_male_karyotype.png)
Name: Hour: Teacher: ROZEMA Inheritance & Mutations Connected to Speciation Let s Review What We Already Know: What Have We Learned? Lesson 26: PI 1 (Projected Image) - Human Karyotype (image from https://en.wikipedia.org/wiki/karyotype#/media/file:nhgri_human_male_karyotype.png)
Roadmap. Sexual Selection. Evolution of Multi-Gene Families Gene Duplication Divergence Concerted Evolution Survey of Gene Families
 1 Roadmap Sexual Selection Evolution of Multi-Gene Families Gene Duplication Divergence Concerted Evolution Survey of Gene Families 2 One minute responses Q: How do aphids start producing males in the
1 Roadmap Sexual Selection Evolution of Multi-Gene Families Gene Duplication Divergence Concerted Evolution Survey of Gene Families 2 One minute responses Q: How do aphids start producing males in the
THE EVIDENCE FOR EVOLUTION
 Unit 37 THE EVIDENCE FOR EVOLUTION LEARNING OBJECTIVES 1. Understand the meaning of the term evolution. 2. Learn about fossil evidence including how fossils are formed. 3. Learn how comparative anatomy
Unit 37 THE EVIDENCE FOR EVOLUTION LEARNING OBJECTIVES 1. Understand the meaning of the term evolution. 2. Learn about fossil evidence including how fossils are formed. 3. Learn how comparative anatomy
(Again) Midterm and Essay 1 = April 12th, Thursday the week after Spring Break
 Announcements (Again) Midterm and Essay 1 = April 12th, Thursday the week after Spring Break This week: More chapter 5 - classification practice, new species concepts, fossils 1 On the midterm 882-E scantron
Announcements (Again) Midterm and Essay 1 = April 12th, Thursday the week after Spring Break This week: More chapter 5 - classification practice, new species concepts, fossils 1 On the midterm 882-E scantron
15.3 Darwin Presents his Case. Biology Mr. Hines
 15.3 Darwin Presents his Case Biology Mr. Hines Darwin returned to England with a wealth of new data. He brought many specimens from the Galapagos to further his studies and to present his data to others.
15.3 Darwin Presents his Case Biology Mr. Hines Darwin returned to England with a wealth of new data. He brought many specimens from the Galapagos to further his studies and to present his data to others.
BLAST Database Searching. BME 110: CompBio Tools Todd Lowe April 8, 2010
 BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
Sperm & Eggs & Variation..OH MY!
 Sperm & Eggs & Variation..OH MY! 1 What if a new individual was formed through mitosis? 2 allele amniocentesis asexual reproduction autosome binary fission chorionic villi sampling crossing over diploid
Sperm & Eggs & Variation..OH MY! 1 What if a new individual was formed through mitosis? 2 allele amniocentesis asexual reproduction autosome binary fission chorionic villi sampling crossing over diploid
1 low Humans Evolved
 1 low Humans Evolved Robert Howl IOIIIB Silk UNIVERS1. i 1 \..UK I..1 I \ Nv I Technische Unive-^itdt Darmstadt FACHDCRLICH 10 BIOLOGIE B i!. I i o t h p k -_ ScLninspilinstiafiG 10 D-6 4287 Darmstadt
1 low Humans Evolved Robert Howl IOIIIB Silk UNIVERS1. i 1 \..UK I..1 I \ Nv I Technische Unive-^itdt Darmstadt FACHDCRLICH 10 BIOLOGIE B i!. I i o t h p k -_ ScLninspilinstiafiG 10 D-6 4287 Darmstadt
Name: Class: Date: ID: A
 Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species
 Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species Zhiqiang Li 1 and Peter Z. Revesz 1,a 1 Department of Computer Science, University of Nebraska-Lincoln, Lincoln, NE,
Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species Zhiqiang Li 1 and Peter Z. Revesz 1,a 1 Department of Computer Science, University of Nebraska-Lincoln, Lincoln, NE,
Face area (cm 2 ) Brain surface area (cm 2 ) Cranial capacity (cm 3 ) 1, Jaw Angle ( º )
 Honors Biology Test : Evolution GOOD LUCK! You ve learned so much! Multiple Choice: Identify the choice that best completes the statement or answers the question. (2 pts each) 1. As we move through the
Honors Biology Test : Evolution GOOD LUCK! You ve learned so much! Multiple Choice: Identify the choice that best completes the statement or answers the question. (2 pts each) 1. As we move through the
GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny
 Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Gene Families part 2. Review: Gene Families /727 Lecture 8. Protein family. (Multi)gene family
 Review: Gene Families Gene Families part 2 03 327/727 Lecture 8 What is a Case study: ian globin genes Gene trees and how they differ from species trees Homology, orthology, and paralogy Last tuesday 1
Review: Gene Families Gene Families part 2 03 327/727 Lecture 8 What is a Case study: ian globin genes Gene trees and how they differ from species trees Homology, orthology, and paralogy Last tuesday 1
The Theory of Evolution
 Name Date Class CHAPTER 13 DIRECTED READING The Theory of Evolution Section 13-1: The Theory of Evolution by Natural Selection Darwin Proposed a Mechanism for Evolution Mark each statement below T if it
Name Date Class CHAPTER 13 DIRECTED READING The Theory of Evolution Section 13-1: The Theory of Evolution by Natural Selection Darwin Proposed a Mechanism for Evolution Mark each statement below T if it
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Unit 3 - Molecular Biology & Genetics - Review Packet
 Name Date Hour Unit 3 - Molecular Biology & Genetics - Review Packet True / False Questions - Indicate True or False for the following statements. 1. Eye color, hair color and the shape of your ears can
Name Date Hour Unit 3 - Molecular Biology & Genetics - Review Packet True / False Questions - Indicate True or False for the following statements. 1. Eye color, hair color and the shape of your ears can
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Basic Local Alignment Search Tool
 Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny
 CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
GENETICS - CLUTCH CH.1 INTRODUCTION TO GENETICS.
 !! www.clutchprep.com CONCEPT: HISTORY OF GENETICS The earliest use of genetics was through of plants and animals (8000-1000 B.C.) Selective breeding (artificial selection) is the process of breeding organisms
!! www.clutchprep.com CONCEPT: HISTORY OF GENETICS The earliest use of genetics was through of plants and animals (8000-1000 B.C.) Selective breeding (artificial selection) is the process of breeding organisms
7. Where do most crustaceans live? A. in the air B. in water C. on the land D. underground. 10. Which of the following is true about all mammals?
 1 A flounder is a type of fish The flounder can change its color to match the surroundings If a shark approaches, the flounder lays still, blending into the sandy ocean bottom This is known as 2 Which
1 A flounder is a type of fish The flounder can change its color to match the surroundings If a shark approaches, the flounder lays still, blending into the sandy ocean bottom This is known as 2 Which
Topic 7: Evolution. 1. The graph below represents the populations of two different species in an ecosystem over a period of several years.
 1. The graph below represents the populations of two different species in an ecosystem over a period of several years. Which statement is a possible explanation for the changes shown? (1) Species A is
1. The graph below represents the populations of two different species in an ecosystem over a period of several years. Which statement is a possible explanation for the changes shown? (1) Species A is
#Evolution. Nothing in Biology makes sense except in the light of evolution.
 #Evolution Nothing in Biology makes sense except in the light of evolution. The Theory of Evolution Change over time. People used to think that species did not change. DARWIN WAS NOT THE PERSON TO COME
#Evolution Nothing in Biology makes sense except in the light of evolution. The Theory of Evolution Change over time. People used to think that species did not change. DARWIN WAS NOT THE PERSON TO COME
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are:
 Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Comparative genomics and proteomics Species available Ensembl focuses on metazoan (animal) genomes. The genomes currently available at the Ensembl site are: Vertebrates: human, chimpanzee, mouse, rat,
Biology Keystone (PA Core) Quiz Theory of Evolution - (BIO.B ) Theory Of Evolution, (BIO.B ) Scientific Terms
 Biology Keystone (PA Core) Quiz Theory of Evolution - (BIO.B.3.2.1 ) Theory Of Evolution, (BIO.B.3.3.1 ) Scientific Terms Student Name: Teacher Name: Jared George Date: Score: 1) Evidence for evolution
Biology Keystone (PA Core) Quiz Theory of Evolution - (BIO.B.3.2.1 ) Theory Of Evolution, (BIO.B.3.3.1 ) Scientific Terms Student Name: Teacher Name: Jared George Date: Score: 1) Evidence for evolution
Organizing Life s Diversity
 17 Organizing Life s Diversity section 2 Modern Classification Classification systems have changed over time as information has increased. What You ll Learn species concepts methods to reveal phylogeny
17 Organizing Life s Diversity section 2 Modern Classification Classification systems have changed over time as information has increased. What You ll Learn species concepts methods to reveal phylogeny
full file at
 Chapter 1 1. Genetics contribute to advances in: Answer: E A. agriculture. B. pharmaceuticals. C. medicine. D. modern biology. E. All of the above. 2. Genetic information can be carried in which of the
Chapter 1 1. Genetics contribute to advances in: Answer: E A. agriculture. B. pharmaceuticals. C. medicine. D. modern biology. E. All of the above. 2. Genetic information can be carried in which of the
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Objective 3.01 (DNA, RNA and Protein Synthesis)
 Objective 3.01 (DNA, RNA and Protein Synthesis) DNA Structure o Discovered by Watson and Crick o Double-stranded o Shape is a double helix (twisted ladder) o Made of chains of nucleotides: o Has four types
Objective 3.01 (DNA, RNA and Protein Synthesis) DNA Structure o Discovered by Watson and Crick o Double-stranded o Shape is a double helix (twisted ladder) o Made of chains of nucleotides: o Has four types
The formation of new species from existing species by the accumulation of variation is called as speciation.
 HEREDITY AND EVOLUTION Speciation :- The formation of new species from existing species by the accumulation of variation is called as speciation. It is mainly due to one or more of the following factors.
HEREDITY AND EVOLUTION Speciation :- The formation of new species from existing species by the accumulation of variation is called as speciation. It is mainly due to one or more of the following factors.
Variation of Traits. genetic variation: the measure of the differences among individuals within a population
 Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Lecture Outline Overview: Investigating the Tree of Life Evolutionary biology is about both process and pattern. o The processes of evolution are natural selection
Chapter 26 Phylogeny and the Tree of Life Lecture Outline Overview: Investigating the Tree of Life Evolutionary biology is about both process and pattern. o The processes of evolution are natural selection
CHAPTERS 24-25: Evidence for Evolution and Phylogeny
 CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
 www.lessonplansinc.com Topic: Dinosaur Evolution Project Summary: Students pretend to evolve two dinosaurs using genetics and watch how the dinosaurs adapt to an environmental change. This is a very comprehensive
www.lessonplansinc.com Topic: Dinosaur Evolution Project Summary: Students pretend to evolve two dinosaurs using genetics and watch how the dinosaurs adapt to an environmental change. This is a very comprehensive
Homeotic Genes and Body Patterns
 Homeotic Genes and Body Patterns Every organism has a unique body pattern. Although specialized body structures, such as arms and legs, may be similar in makeup (both are made of muscle and bone), their
Homeotic Genes and Body Patterns Every organism has a unique body pattern. Although specialized body structures, such as arms and legs, may be similar in makeup (both are made of muscle and bone), their
Mitosis and Meiosis. 2. The distribution of chromosomes in one type of cell division is shown in the diagram below.
 Name: Date: 1. Jack bought a small turtle. Three months later, the turtle had grown to twice its original size. Which of the following statements best describes why Jack s turtle got bigger? A. Parts of
Name: Date: 1. Jack bought a small turtle. Three months later, the turtle had grown to twice its original size. Which of the following statements best describes why Jack s turtle got bigger? A. Parts of
Name Period. 3. How many rounds of DNA replication and cell division occur during meiosis?
 Name Period GENERAL BIOLOGY Second Semester Study Guide Chapters 3, 4, 5, 6, 11, 14, 16, 17, 18 and 19. SEXUAL REPRODUCTION AND MEIOSIS 1. What is the purpose of meiosis? 2. Distinguish between diploid
Name Period GENERAL BIOLOGY Second Semester Study Guide Chapters 3, 4, 5, 6, 11, 14, 16, 17, 18 and 19. SEXUAL REPRODUCTION AND MEIOSIS 1. What is the purpose of meiosis? 2. Distinguish between diploid
