Testing the independence hypothesis of accepted mutations for pairs of adjacent amino acids in protein sequences
|
|
|
- Alban McKenzie
- 5 years ago
- Views:
Transcription
1 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln CSE Journal Articles Computer Science and Engineering, Department of Summer Testing the independence hypothesis of accepted mutations for pairs of adjacent amino acids in protein sequences Jyotsna Ramanan University of Nebraska-Lincoln, jor.compscie@gmail.com Peter Revesz University of Nebraska - Lincoln, prevesz1@unl.edu Follow this and additional works at: Part of the Bioinformatics Commons, Biostatistics Commons, Computational Biology Commons, Computer Sciences Commons, Molecular Biology Commons, Molecular Genetics Commons, Statistical Methodology Commons, and the Statistical Models Commons Ramanan, Jyotsna and Revesz, Peter, "Testing the independence hypothesis of accepted mutations for pairs of adjacent amino acids in protein sequences" (2017). CSE Journal Articles This Article is brought to you for free and open access by the Computer Science and Engineering, Department of at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in CSE Journal Articles by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln.
2 Testing the Independence Hypothesis of Accepted Mutations for Pairs of Adjacent Amino Acids in Protein Sequences Jyotsna Ramanan and Peter Z. Revesz Abstract Evolutionary studies usually assume that the genetic mutations are independent of each other. However, that does not imply that the observed mutations are independent of each other because it is possible that when a nucleotide is mutated, then it may be biologically beneficial if an adjacent nucleotide mutates too. With a number of decoded genes currently available in various genome libraries and online databases, it is now possible to have a large-scale computer-based study to test whether the independence assumption holds for pairs of adjacent amino acids. Hence the independence question also arises for pairs of adjacent amino acids within proteins. The independence question can be tested by considering the evolution of proteins within a closely related sets of proteins, which are called protein families. In this thesis, we test the independence hypothesis for three protein families from the PFAM library, which is a publicly available online database that records a growing number of protein families. For each protein family, we construct a hypothetical common ancestor, or consensus sequence. We compare the hypothetical common ancestor of a protein family with each of the descendant protein sequences in the family to test where the mutations occurred during evolution. The comparison yields actual probabilities for each pair of amino acids changing into another pair of amino acids. By comparing the actual probabilities with the theoretical probabilities under the independence assumption, we identify anomalies that indicate that the independence assumption does not hold for many pairs of amino acids. Keywords amino acid, independent probabilities, nucleotide, genetic mutation, protein. I. INTRODUCTION IOLOGICAL evolution depends on random mutations B accompanied by natural selection for the more fit genes. That simple statement does not imply that the observed mutations are independent from each other. It is possible that if a nucleotide changes, then it is biologically beneficial to have some of the adjacent or nearby nucleotides change as The work of the second author was supported in part by a Fulbright Scholarship sponsored by the U.S. Department of State while he was on leave from the University of Nebraska-Lincoln. Jytosna Ramanan was with the Department of Computer Science and Engineering of the University of Nebraska-Lincoln, while she was an M.S. student. Now she is working as a software engineer in Chicago, IL, USA ( jor.compscie@gmail.com). Peter Z. Revesz is a professor in the Department of Computer Science and Engineering, University of Nebraska-Lincoln, Lincoln, NE USA (phone: ; fax: ; revesz@cse.unl.edu). For part of this research he was on a Fulbright Scholarship leave at the Aquincum Institute of Technology in Budapest, Hungary. well. For example, if in some protein-coding region within some triplet that encodes a hydrophilic amino acid a nucleotide changes such that the triplet would encode a hydrophobic amino acid, then a mutation of another nucleotide in the same triplet may be advantageous if with that mutation the triplet would again encode a hydrophilic amino acid (or preserve another key property of amino acids). In other words, some mutations within a triplet slightly increase the probability that some accompanying mutation with a readjusting effect would survive in the offspring. With the greatly increasing number of decoded genes currently available in a number of genome libraries and online databases, it is now possible to have a large-scale computerbased study to test whether the independence assumption holds. One difficulty, however, is to find the coding regions and coding triplets. Hence it seems more convenient to investigate proteins derived from the coding regions. The mutations in the coding regions of the DNA are usually reflected in the mutations of amino acids. Therefore, instead of the evolution of genes, one may talk about the evolution of proteins within a closely related set of proteins, which is called a protein family. The PFAM library [4] records a growing number of protein families. Each protein in a protein family can be assumed to be genetically related to the other proteins in that family and to have evolved from a single ancestor protein. For any set of DNA strings and any set of proteins, there are several algorithms that can be used to find a hypothetical evolutionary tree (see the textbooks by Baum and Smith [1], Hall [2], and Lerney et al. [3] for an overview of these algorithms.) Revesz [5] has recently proposed a new phylogenetic tree-building algorithm called the Common Mutation Similarity Matrixes (CMSM) algorithm. This algorithm finds a hypothetical evolutionary tree. The first step of the CMSM algorithm is to find a hypothetical common ancestor, which is denoted by µ. In this paper, we will use the idea of a hypothetical common ancestor. We can compare the hypothetical common ancestor of a family of proteins with each of the proteins in the family to test where the mutations occur. We can also test for each adjacent pair of amino acids how many times that pair changed into another pair of amino acids. The resulting experimental statistics can be compared with the theoretical probability under the independence assumption. If the ISSN:
3 deviation from the theoretical probability is significant, then the independence assumption fails to provide a satisfying explanation for the experimental results. Evolutionary studies usually assume that the genetic mutations are independent of each other. This paper tests the independence hypothesis for genetic mutations with regard to protein coding regions. As discussed in Section III, according to our experimental results the independence assumption generally holds, but there seem to be certain exceptions. We give examples in Section III of some particular adjacent amino acid pairs that seem to change in ways that deviate significantly from the expected theoretical probability under the independence assumption. This paper is organized as follows. Section II describes our method with an extended example. Section III describes the protein families that were used in the experiments. Section IV presents our experimental results. Finally, Section V gives some conclusions and directions for further research. II. THE INDEPENDENCE TESTING METHOD In this section, we describe the step-by-step procedure that we used to test whether among the surviving descendants of the hypothetical common ancestor µ the adjacent pairs of amino acids are mutated independently of each other. As an artificial and simplified example, suppose that there exists an ancestor protein µ that is made up of only the amino acids A, D, N and R as shown in Fig. 1. Further assume during evolution each of these four amino acids either remains unchanged or is mutated into only one of the other three amino acids within this group of four amino acids. Suppose that the seven descendants S 1 S 7 are as shown in Fig. 1. S 1 S 2 S 3 RNARDANDRADNRDANRARA NRARDANRADADNANARNAD RADNRANDANDRANDRDRAN proteins in the given set of protein family using the method that is also used by the Common Mutation Similarity Matrix. In the case of amino acid sequences, the hypothetical common ancestor, µ, is constructed by taking an alignment of the amino acid sequences, and in each column of the alignment finding the amino acid (out of the twenty possible amino acids that are used in almost every protein in all organisms) that is overall closest to the all the amino acids in that column. The overall closest amino acid is by definition the one for which the sum of the PAM250 matrix distance values between it and the amino acids in the column considered is minimal. If there are two or more values that are minimal, then we make a random selection. For the example in Fig. 1, consisting of seven artificial sequences from S 1, S 2,... S 7, each with a length of twenty nucleotides, the consensus sequence is as shown in Fig. 2. µ RNARDANDRADNRDANRNAA Fig. 2 The consensus sequence for the artificial protein family in Fig. 1 B. Calculate Mutation Matrix Next, we calculate a mutation probability matrix. The mutation probability matrix contains the probabilities of any amino acid changing into another amino acid. For the running example with the data shown in Fig. 1, the mutation probability matrix is shown in Table 1. The mutation Matrix in Table 1 shows the frequencies of the each of the four amino acid changes into one of the other three amino acids or remains the same. The column Total shows the total number of the possibility of one amino acid can mutate into another amino acid, or remain the same throughout the entire sequence (S 1 to S 7 ). S 4 S 5 S 6 S 7 DNARDNARDRNARDANRANR RNDRANRDRDANDNANDRAN RNARDANDRADNRDANRARA RNARDADDRADNRDANDADA Table 1 The mutation probability matrix for the data in Fig.1 A R N D Total A R N D Fig. 1 A set of seven artificial sequences Our testing method consists of five steps that are explained in Sections (A-E). A. Find Consensus Sequence Construct the hypothetical common ancestor for the C. Find Theoretical Probabilities Based on the mutation probability matrix values, we estimate the probability of the changes of any adjacent pair of amino acids into another pair of amino acids assuming that the mutations are independent of each other. For example, the probability of AN changing into DR can be computed as follows: ISSN:
4 Prob (AN, DR) = Prob (A, D) * Prob (N, R) =!! =!!"!"!"# Hence the theoretical probability corresponding to the amino acid pair AN changing to DR is approximately The theoretical probabilities for all possible combinations of amino acid pairs of the artificial sequence in Fig. 1 mutating into another possible pair of the same set are shown in Table 2. Note that the table values are in decimal format for the purpose of calculation. D. Find Actual Probabilities Now, we calculate the actual probabilities of changes for each pair of amino acids in the consensus sequence. Starting from the first pair to the end of the consensus string, we first calculate the number of times and the index, each pair in the consensus string occurs. We then calculate the frequencies of that specific pair in the consensus string mutating into another pair among the rest of the descendent sequences in that column. If the current adjacent amino acid pair of the consensus string happens to appear in another index of the same consensus string, then we repeat the step to check for frequencies of that pair mutating into other possible pairs in that column, for the rest of the descendant sequences. We then slide the window of the current pair in the consensus string to the adjacent consecutive pair of the same consensus string, to calculate their respective frequencies of mutations among the descendent pairs of that column. The steps mentioned in the above paragraph are repeated until we encounter the last possible pair of the consensus sequence. The results for the example in Fig. 1 with the seven artificial sequences are shown in Table 3. Note that in Table 3, the column Total refers to the total number of ways in which a pair of the consensus sequence can mutate into another possible pair in its descendant sequence, whose value is the product of the number of times a single pair appears in the consensus string and the total number of sequences in the protein family. For example, in consensus string µ for the artificial sequence in Fig. 1, NR appears in two indices as underlined in Fig. 3 below. µ RNARDANDRADNRDANRNAA Fig. 2 The consensus sequence for the artificial protein family in Fig. 1 In this case, the total number of possibilities of NR changing into another pair is 2 * 7 = 14, where seven is the total number of sequences of the protein family. Algorithm ACTUAL-PROBABILITY (S, n, m) INPUT: The set S of aligned sequences of a protein family represented as a matrix. The sequences are S[1][1 m], S[2][1 m],, S[n][1 m] where n denotes the total number of sequences and m denotes their lengths. //TOT is the overall total number of possible ways a //particular pair can mutate to another pair. //The auxiliary function Find_Consensus_Sequence(S) finds //the consensus sequence of S. 1 C := Find_Consensus_Sequence(S) 2 for i 1 to m-1 do 3 Calculate the count and index of all the adjacent pairs in the consensus sequence 4 TOT := count * n 5 end for 6 for i 1 to m-1 do 7 for j 2 to n do 8 Calculate the occurrences of possible pairs in the descendent sequences corresponding to the column S[i][i+1] which is the consensus sequence. 14 end for 15 end for The algorithm ACTUAL-PROBABILITY is a dynamic programming computer algorithm. We iterate through the consensus sequence m number of times for each adjacent pair in the consensus sequence. During each of those iterations we count the frequencies of the pair in that window which may or may not mutate into another pair in their descendant sequences of the corresponding window, which takes about n number of comparisons. This operation can be seen under the nested loops that start on lines 6 and 7 in the algorithm. Line 8 calculates the occurrences of the pair in the consensus mutating into one of the possible 400 pairs in the descendent sequences. That takes about n number of comparisons in the worst case. Hence we can prove the following theorem. Theorem: The running time of the algorithm ACTUAL- PROBABILITY is O(n 2 m) where m n, and m is the size of the consensus sequence and n is the number of the sequences of the protein family. ISSN:
5 Table 2 The theoretical probabilities of changes for each pair of amino acids for the artificial example protein family in Fig. 3. AA AR AN AD RA RR RN RD NA NR NN ND DA DR DN DD AA AR AN AD RA RR RN RD NA NR NN ND DA DR DN DD Table 3 The actual probabilities of changes for each pair of amino acids for the artificial example protein family in Fig. 3. AA AR AN AD RA RR RN RD NA NR NN ND DA DR DN DD TOTAL AA AR AN AD RA RR RN RD NA NR NN ND DA DR DN DD ISSN:
6 E. Find Discrepancies between the Actual and Theoretical Probabilities We compare the theoretical and the actual probabilities and note the most important discrepancies. The percentage probability difference in the theoretical and actual probabilities of the mutations of amino acid pairs is the absolute value of the difference between the two types of probabilities divided by the maximum of the two probabilities. Let T (p1, p2) and E (p1, p2) be the theoretical and the experimental probabilities, respectively, that the amino acid pair p1 changes into the amino acid pair p2. Let PD (p1, p2) be the percent probability difference defined as follows: PD(p1, p2) = T p1, p2 E(p1, p2) Max(T p1, p2, E p1, p2 ) The percentage Difference (PD) for the top eight pairs of the consensus sequence mutating into other pairs in the descendant sequences of the artificial protein family is shown in Table 4 below. Table 4 Differences for the protein sequences in Fig. 1 Pair of Amino Acids From à To Theoretical (T) III. THE PROTEIN FAMILIES USED IN THE EXPERIMENTS Our testing methods were also conducted on three other protein families for obtaining robust outcomes. We present the experimental values of the following protein families below. The PF series number indicates the serial number of the protein in the PFAM library (pfam.xfam.org) [4]. The PFAM library can be also browsed using the PROFESS protein database system [12] and references in the InterPro protein classification system [9]. DAGK_cat (PF00781) IL17 (PF06083) KA1 (PF02149) Actual (A) Freq Out of Difference PD (T, A) AD à DA % AR à DN % RN à DR % AA à AN % AN à NR % NA à RA % NR à ND % RN à RA % Note that the theoretical probabilities and actual probabilities are presented in Tables 4-6. In that, the theoretical probabilities of certain amino acids that had a negligible probability value (<0.05) were rounded to zeros as they seemed to be meager for large sets of protein sequences. The actual mutation probability matrices for the respective protein families are presented in Tables 7 9 at the end of the article. are using Word, use either the Microsoft Equation Editor or the MathType add-on ( for equations in your paper (Insert Object Create New Microsoft Equation or MathType Equation). Float over text should not be selected. In the next three sections we describe the protein families that were used in the experiments. A. The DAGK_cat Protein Family (PF00781) The protein family used here to test the method on large data set is the Diacylglycerol kinase catalytic domain (DAGK_cat) whose sequences can be referred from the PFAM Library. This domain consists of sequences, out of which 110 seed sequences were used for the experiment in this paper. The common mutation ancestor µ was calculated to be: KALVIVNPKSGTARGGKGKKLLERKVRPLLEEAGVS DDELDLRLTENPGPGDVLRRGYGNLEKLKSNALELL AGAAREAAEANEQSDGDTLLPWSENLAYGYCPDLIV AAGGDGTVNEVLNGLAGNARRDDLELATRNHPRAV LVPSSPPLGIIPLGRTGNDFARALNAHGGFEEGIPLGY DPEEAARAALELIKKIKGQTRPVDVGKV B. The KA Protein Family (PF02149) This family consists of 1349 sequences in total, where around 105 sequences were used for the experiment discussed in this paper. The common mutation ancestor µ was calculated to be: LVVKFEIEVCKVPLLSGNSNSQEHLYGVQFKRINSGD TWQYKNLASKILSELKL C. The IL17 Protein Family (PF06083) This family consists of 531 sequences in total, where around 102 sequences were used for the experiment discussed in this paper. The common mutation ancestor µ was calculated to be: RSLSPWDYREIDPHDPNRYPRVIAEARCLLCSGGSRCI GDLNPATGQGEDDIAELQGLRRSLNSVPIYQEILVAF LDGGGKLRRLCDKPCSRPKTHEPCAGCRYSYRLEPV KETVTVGCTV IV. EXPERIMENTAL RESULTS Table 5 shows the experimental results for the DAGK_cat protein family (PF00781). ISSN:
7 Table 5 Anomalous probability differences for the DAGK_cat protein family for pairs of amino acids with or without mutations Pair of Amino Acids From à To Theoretical (T) Actual (A) Out Freq of Difference (PD) FA à LA % VI à VF % SG à AG % PK à PT % EV à EV % SG à SG % FA à FA % NP à NP % VA à IA % IP à LP % AR à AR % NG à NG % VD à ID % DG à DG % IP à IP % TV à TL % LN à VN % LE à LN % IV à VI % VG à LG % GD à GD % TV à TV % IP à VP % GT à GT % LG à AG % GN à GN % GG à GG % PL à PL % VL à VV % Table 6 Anomalous probability differences for the KA protein family for pairs of amino acids with or without mutations Pair of Amino Acids From à To Theoretical (T) Actual (A) Out Freq of Difference (PD) PL à PR % LS à LS % VC à IV % LY à LH % YG à HG % KL à RL % YK à YK % EI à EI % GV à GI % CK à VK % FE à FE % QF à QF % KF à RF % KV à KV % VC à VC % EV à EV % KR à QR % VP à LP % EL à EL % KV à KL % KR à RR % IL à IL % GD à GN % RI à RV % GV à GV % FK à FK % RI à RL % KR à KR % VP à VP % KF à KF % Next, Table 6 shows the experimental results for the KA protein family (PF02149). Next, Table 7 shows the experimental results for the IL17 protein family (PF06083). ISSN:
8 Table 7 Anomalous probability differences for the IL17 protein family for pairs of amino acids with or without mutations Pair of Amino Acids From à To LV à PV VT à VP TV à PV LN à MN VG à VA TV à AV YQ à QQ GC à AC VG à VG LR à LK SP à CP LS à IS Theoretical (T) Actual (A) Out Freq of Difference (PD) % % % % % % % % % % % % AR à AK % IY à IQ % AR à AQ PR à PS EA à EA PR à PQ RC à KC YP à FP YP à IP GQ à GK RC à QC DP à DE SV à SV SL à SI LS à LS SY à SF SL à SL AE à PE VP à LP RY à RI ED à ED RY à RF CI à CL CI à CV % % % % % % % % % % % % % % % % % % % % % % SP à SP PI à PV YR à FR DY à TY NR à NR YP à YP RS à RS HD à ID NS à NS YR à YR LV à VV % % % % % % % % % % % HD à ED % PW à PW % PR à PR DP à DP WD à WT % % % Table 8 shows five pairwise mutations that are common in at least two of the three protein families that we studied. The first three mutations occur exactly the same in the corresponding protein families. In the fourth and the fifth mutations, the pairs are interchanged. For example, when we take the IP à VP mutation, which occurs in the DAGK protein, and interchange the pairs on both the left and the right hand sides, then we get the symmetric mutation PI à PV, which occurs in the IL17 protein. These two mutations are very similar to each other because proteins are amino acid chains, and the two mutations simple read these amino acid chains from different directions. Table 8 Common or similar mutations in the three protein families Mutation DAGK_cat IL17 KA1 1 EV à EV EV à EV 2 LS àls LS à LS 3 VP à LP VP à LP 4 IP à VP PI à PV 5 VL à VV LV à VV There are a total of 400 x 400 = 160,000 possible pairwise mutations. The probability of finding a common pairwise mutation out of the top 31 of IL17 mutations and the top 18 KA1 mutations, is: Prob(out of the 18 newly picked from 160,000 at least one will match one of the 31 picked before) = 1 Prob(none of 18 newly picked matches the 31 picked before) ISSN:
9 In terms of number of permutations, this problem could be solved as: n!! 1 P! n r! = 1! P! m! m r! where m = , n = and r =18. After the substitution of the respective values, we get: 1!"####!!" P!"!"#### P!" Let us set the above-calculated value to be our P- value. As can be seen from Table 8, the probability of finding five common mutations in at least two of the protein families was calculated to be about which is significantly lesser than the P-value. Figs. 4 and 5 show the calculations and the statistical results generated using SAS for our proof. Fig. 5 Finding 5 common pairwise mutations out of the top 31 IL17 mutations, the top 18 KA1 mutations and the top 31 DAGK_cat mutation Fig. 4 SAS results showing the probability of finding at least one common pairwise mutation out of the top 31 of IL17 mutations and the top 18 KA1 mutations Since the above probability of overlap is so small, our finding cannot be explained as a random event. This shows that the anomalies we found are not accidental but are some consequence of the chemical nature of these particular amino acid pairs and evolutionary forces acting on those pairs. Moreover, the above low probability is just for finding at least one common pairwise mutation whereas we have found three of them plus two other pairs that are complements of each other. ISSN:
10 Fig. 6 Histograms showing the number of times each amino acid appears in µ for the protein family DAGK_cat (blue), IL17 (red) and KA (orange), and the histogram showing the percent probability differences (PDs) for the mutations that are commonly anomalous in at least two of the three protein families. A. Charts Fig. 6 shows the histograms of the probability of each amino acid in the sample protein families. The amino acids are along the x-axis and the total possible outcomes (in numbers) are along the y-axis. V. CONCLUSION AND FUTURE WORK Large The experimental results suggest that adjacent pairs of amino acids in the surviving descendants are sometimes mutated in a dependent instead of an independent way. Since the probability of overlap seems to be small about and evidently lesser than out P-value which about implies that we have a concrete proof that our findings cannot be explained as a random event. This shows that the anomalies we found are not accidental but are some consequence of the chemical nature of these particular amino acid pairs and evolutionary forces acting on those pairs. Moreover, the above low probability is just for finding at least one common pairwise mutation whereas we have found three of them plus two other pairs that are complements of each other. From the overall set of experiments, we can infer that the pairwise mutations of a protein sequence in a protein family does not have to be independent all the time. However, the experimental data is based only on three protein families. In the future we plan to use our independence testing method for many other protein families. We also plan to experiment with using other amino acid substitution matrixes beside the PAM250 matrix [6]. We also plan to look at longer sequences, that is, consider adjacent N-mers of amino acids for N > 2. Finally, it is an intriguing question what the changed view of the amino acid similarity table implies about evolution [17]. ACKNOWLEDGMENT We thank Dr. Stephen D. Kachman of the Department of Statistics at University of Nebraska- Lincoln for his pertinent ISSN:
11 help in this research. One of us (J.R) would like to thank Dr. Juan Cui and Dr. Stephen Scott for investing their time to serve in the M.S. thesis committee. REFERENCES [1] D. Baum and S. Smith, Tree Thinking: An Introduction to Phylogenetic Biology, Roberts and Company Publishers [2] B. G. Hall, Phylogenetic Trees Made Easy: A How to Manual, 4th edition, Sinauer Associates, [3] P. Lerney, M. Salemi, and A.-M Vandamme, editors. The Phylogenetic Handbook: A Practical Approach to Phylogenetic Analysis and Hypothesis Testing, 2nd edition, Cambridge University Press, [4] The PFAM Protein Library. Available: [5] P. Z. Revesz, An algorithm for constructing hypothetical evolutionary trees using common mutations similarity matrices, Proc. 4th ACM International Conference on Bioinformatics and Computational Biology, ACM Press, Bethesda, MD, USA, September 2013, pp [6] P. Z. Revesz, Introduction to Databases: From Biological to Spacio- Temporal, Springer, [7] H. M. Bakali, M. D. Herman, K. A. Johnson, A. A. Kelly, A. Wieslander, B. M. Hallberg, and P. Nordlund; Crystal structure of YegS, a homologue to the mammalian diacylglycerol kinases, reveals a novel regulatory metal binding site, J. Biol Chem 282: , [8] I. Merida, A. Avila-Flores, and E. Merino, "Diacylglycerol kinases: at the hub of cell signaling, Biochem. J. 409 (1): doi: /bj , [9] InterPro Protein sequence analysis & classification. Available: [10] C. Brocker, D. Thompson, A. Matsumoto, D. W. Nebert, V. Vasiliou "Evolutionary divergence and functions of the human interleukin (IL) gene family. Human Genomics 5 (1), [11] S. Aggarwal and A. L. Gurney "IL-17: prototype member of an emerging cytokine family," J. Leukoc. Biol. 71(1), [12] T. Triplet, M. D. Shortridge, M. A. Griep, R. Powers and P. Revesz PROFESS: A PROtein Function, Evolution, Structure and Sequence database. Database, 2010:baq011. PMC , [13] J. P. Tassan and X. Le Goff "An overview of the KIN1/PAR-1/MARK kinase family," Biol. Cell 96 (3): 193 9, [14] J. Biernat, Y. Z. Wu, T. Timm, Q. Zheng-Fischhofer, E. Mandelkow, L. Meijer, E. M. Mandelkor (November 2002). "Protein kinase MARK/PAR-1 is required for neurite outgrowth and establishment of neuronal polarity". Mol. Biol. Cell 13 (11): [15] S. Guo and K. J. Kemphues (May 1995). "par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Ser/Thr kinase that is asymmetrically distributed". Cell 81 (4): [16] M. Elbert, G. Rossi and P. Brennwald "The yeast par-1 homologs kin1 and kin2 show genetic and physical interactions with components of the exocytic machinery," Mol. Biol. Cell 16 (2): , [17] M. D. Shortridge, T. Triplet, P. Revesz, M. Griep and R. Powers "Bacterial Protein Structures Reveal Phylum Dependent Divergence," Comput. Biol. Chem., 35: PMC , Jyotsna Ramanan (MS 16) was a graduate student in the Department of Computer Science and Engineering at the University of Nebraska- Lincoln and studied previously at Pondicherry University, Pondicherry, India. Her research interests are in bioinformatics, big data and database system concepts. She is currently working as a software engineer in Chicago. Peter Z. Revesz (Ph.D 91) holds a Ph.D. degree in Computer Science from Brown University and was a postdoctoral fellow at the University of Toronto. He is an expert in databases, data mining, big data analytics and bioinformatics. He is the author of Introduction to Databases: From Biological to Spatio-Temporal (Springer, 2010) and Introduction to Constraint Databases (Springer, 2002). He is currently a professor in the Department of Computer Science and Engineering at the University of Nebraska- Lincoln, Lincoln, NE 6815, USA. Dr. Revesz also held visiting appointments at the Aquincum Institute of Technology, the IBM T. J. Watson Research Center, INRIA, the Max Planck Institute for Computer Science, the University of Athens, the University of Hasselt, the U.S. Air Force Office of Scientific Research and the U.S. Department of State. He is a recipient of an AAAS Science & Technology Policy Fellowship, a J. William Fulbright Scholarship, an Alexander von Humboldt Research Fellowship, a Jefferson Science Fellowship, a National Science Foundation CAREER award, and a Faculty International Scholar of the Year award by Phi Beta Delta, the Honor Society for International Scholars. ISSN:
TESTING THE INDEPENDENCE HYPOTHESIS OF ACCEPTED MUTATIONS FOR PAIRS OF ADJACENT AMINO ACIDS IN PROTEIN SEQUENCES
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Computer Science and Engineering: Theses, Dissertations, and Student Research Computer Science and Engineering, Department
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Computer Science and Engineering: Theses, Dissertations, and Student Research Computer Science and Engineering, Department
Incremental Phylogenetics by Repeated Insertions: An Evolutionary Tree Algorithm
 Incremental Phylogenetics by Repeated Insertions: An Evolutionary Tree Algorithm Peter Z. Revesz, Zhiqiang Li Abstract We introduce the idea of constructing hypothetical evolutionary trees using an incremental
Incremental Phylogenetics by Repeated Insertions: An Evolutionary Tree Algorithm Peter Z. Revesz, Zhiqiang Li Abstract We introduce the idea of constructing hypothetical evolutionary trees using an incremental
A Computational Model of the Spread of Ancient Human Populations Based on Mitochondrial DNA Samples
 A Computational Model of the Spread of Ancient Human Populations Based on Mitochondrial DNA Samples Peter Z. Revesz Abstract The extraction of mitochondrial DNA (mtdna) from ancient human population samples
A Computational Model of the Spread of Ancient Human Populations Based on Mitochondrial DNA Samples Peter Z. Revesz Abstract The extraction of mitochondrial DNA (mtdna) from ancient human population samples
A mitochondrial DNA-based computational model of the spread of human populations
 University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln CSE Journal Articles Computer Science and Engineering, Department of Spring 3-15-2016 A mitochondrial DNA-based computational
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln CSE Journal Articles Computer Science and Engineering, Department of Spring 3-15-2016 A mitochondrial DNA-based computational
A L A BA M A L A W R E V IE W
 A L A BA M A L A W R E V IE W Volume 52 Fall 2000 Number 1 B E F O R E D I S A B I L I T Y C I V I L R I G HT S : C I V I L W A R P E N S I O N S A N D TH E P O L I T I C S O F D I S A B I L I T Y I N
A L A BA M A L A W R E V IE W Volume 52 Fall 2000 Number 1 B E F O R E D I S A B I L I T Y C I V I L R I G HT S : C I V I L W A R P E N S I O N S A N D TH E P O L I T I C S O F D I S A B I L I T Y I N
Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species
 Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species Zhiqiang Li 1 and Peter Z. Revesz 1,a 1 Department of Computer Science, University of Nebraska-Lincoln, Lincoln, NE,
Cubic Spline Interpolation Reveals Different Evolutionary Trends of Various Species Zhiqiang Li 1 and Peter Z. Revesz 1,a 1 Department of Computer Science, University of Nebraska-Lincoln, Lincoln, NE,
Local Alignment Statistics
 Local Alignment Statistics Stephen Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health Bethesda, MD Central Issues in Biological Sequence Comparison
Local Alignment Statistics Stephen Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health Bethesda, MD Central Issues in Biological Sequence Comparison
Sequence analysis and comparison
 The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
Bioinformatics. Scoring Matrices. David Gilbert Bioinformatics Research Centre
 Bioinformatics Scoring Matrices David Gilbert Bioinformatics Research Centre www.brc.dcs.gla.ac.uk Department of Computing Science, University of Glasgow Learning Objectives To explain the requirement
Bioinformatics Scoring Matrices David Gilbert Bioinformatics Research Centre www.brc.dcs.gla.ac.uk Department of Computing Science, University of Glasgow Learning Objectives To explain the requirement
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Solutions and Ions. Pure Substances
 Class #4 Solutions and Ions CHEM 107 L.S. Brown Texas A&M University Pure Substances Pure substance: described completely by a single chemical formula Fixed composition 1 Mixtures Combination of 2 or more
Class #4 Solutions and Ions CHEM 107 L.S. Brown Texas A&M University Pure Substances Pure substance: described completely by a single chemical formula Fixed composition 1 Mixtures Combination of 2 or more
Sara C. Madeira. Universidade da Beira Interior. (Thanks to Ana Teresa Freitas, IST for useful resources on this subject)
 Bioinformática Sequence Alignment Pairwise Sequence Alignment Universidade da Beira Interior (Thanks to Ana Teresa Freitas, IST for useful resources on this subject) 1 16/3/29 & 23/3/29 27/4/29 Outline
Bioinformática Sequence Alignment Pairwise Sequence Alignment Universidade da Beira Interior (Thanks to Ana Teresa Freitas, IST for useful resources on this subject) 1 16/3/29 & 23/3/29 27/4/29 Outline
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Tools and Algorithms in Bioinformatics
 Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Tools and Algorithms in Bioinformatics GCBA815, Fall 2015 Week-4 BLAST Algorithm Continued Multiple Sequence Alignment Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and
Quantifying sequence similarity
 Quantifying sequence similarity Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 16 th 2016 After this lecture, you can define homology, similarity, and identity
Quantifying sequence similarity Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 16 th 2016 After this lecture, you can define homology, similarity, and identity
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Sequence Analysis 17: lecture 5. Substitution matrices Multiple sequence alignment
 Sequence Analysis 17: lecture 5 Substitution matrices Multiple sequence alignment Substitution matrices Used to score aligned positions, usually of amino acids. Expressed as the log-likelihood ratio of
Sequence Analysis 17: lecture 5 Substitution matrices Multiple sequence alignment Substitution matrices Used to score aligned positions, usually of amino acids. Expressed as the log-likelihood ratio of
Lecture Notes: Markov chains
 Computational Genomics and Molecular Biology, Fall 5 Lecture Notes: Markov chains Dannie Durand At the beginning of the semester, we introduced two simple scoring functions for pairwise alignments: a similarity
Computational Genomics and Molecular Biology, Fall 5 Lecture Notes: Markov chains Dannie Durand At the beginning of the semester, we introduced two simple scoring functions for pairwise alignments: a similarity
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Introduction to Evolutionary Concepts
 Introduction to Evolutionary Concepts and VMD/MultiSeq - Part I Zaida (Zan) Luthey-Schulten Dept. Chemistry, Beckman Institute, Biophysics, Institute of Genomics Biology, & Physics NIH Workshop 2009 VMD/MultiSeq
Introduction to Evolutionary Concepts and VMD/MultiSeq - Part I Zaida (Zan) Luthey-Schulten Dept. Chemistry, Beckman Institute, Biophysics, Institute of Genomics Biology, & Physics NIH Workshop 2009 VMD/MultiSeq
An Evolutionary Trend Discovery Algorithm Based on Cubic Spline Interpolation
 An Evolutionary Trend Discovery Algorithm Based on Cubic Spline Interpolation ZHIQIANG LI and PETER Z. REVESZ Department of Computer Science and Engineering University of Nebraska-Lincoln Lincoln, NE 68588-0115
An Evolutionary Trend Discovery Algorithm Based on Cubic Spline Interpolation ZHIQIANG LI and PETER Z. REVESZ Department of Computer Science and Engineering University of Nebraska-Lincoln Lincoln, NE 68588-0115
MATHEMATICAL MODELS - Vol. III - Mathematical Modeling and the Human Genome - Hilary S. Booth MATHEMATICAL MODELING AND THE HUMAN GENOME
 MATHEMATICAL MODELING AND THE HUMAN GENOME Hilary S. Booth Australian National University, Australia Keywords: Human genome, DNA, bioinformatics, sequence analysis, evolution. Contents 1. Introduction:
MATHEMATICAL MODELING AND THE HUMAN GENOME Hilary S. Booth Australian National University, Australia Keywords: Human genome, DNA, bioinformatics, sequence analysis, evolution. Contents 1. Introduction:
EVOLUTIONARY DISTANCES
 EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
EVOLUTIONARY DISTANCES FROM STRINGS TO TREES Luca Bortolussi 1 1 Dipartimento di Matematica ed Informatica Università degli studi di Trieste luca@dmi.units.it Trieste, 14 th November 2007 OUTLINE 1 STRINGS:
INFORMATION-THEORETIC BOUNDS OF EVOLUTIONARY PROCESSES MODELED AS A PROTEIN COMMUNICATION SYSTEM. Liuling Gong, Nidhal Bouaynaya and Dan Schonfeld
 INFORMATION-THEORETIC BOUNDS OF EVOLUTIONARY PROCESSES MODELED AS A PROTEIN COMMUNICATION SYSTEM Liuling Gong, Nidhal Bouaynaya and Dan Schonfeld University of Illinois at Chicago, Dept. of Electrical
INFORMATION-THEORETIC BOUNDS OF EVOLUTIONARY PROCESSES MODELED AS A PROTEIN COMMUNICATION SYSTEM Liuling Gong, Nidhal Bouaynaya and Dan Schonfeld University of Illinois at Chicago, Dept. of Electrical
Single alignment: Substitution Matrix. 16 march 2017
 Single alignment: Substitution Matrix 16 march 2017 BLOSUM Matrix BLOSUM Matrix [2] (Blocks Amino Acid Substitution Matrices ) It is based on the amino acids substitutions observed in ~2000 conserved block
Single alignment: Substitution Matrix 16 march 2017 BLOSUM Matrix BLOSUM Matrix [2] (Blocks Amino Acid Substitution Matrices ) It is based on the amino acids substitutions observed in ~2000 conserved block
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT
 5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT
 3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT.03.239 25.09.2012 SEQUENCE ANALYSIS IS IMPORTANT FOR... Prediction of function Gene finding the process of identifying the regions of genomic DNA that encode
3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT.03.239 25.09.2012 SEQUENCE ANALYSIS IS IMPORTANT FOR... Prediction of function Gene finding the process of identifying the regions of genomic DNA that encode
Introduction to population genetics & evolution
 Introduction to population genetics & evolution Course Organization Exam dates: Feb 19 March 1st Has everybody registered? Did you get the email with the exam schedule Summer seminar: Hot topics in Bioinformatics
Introduction to population genetics & evolution Course Organization Exam dates: Feb 19 March 1st Has everybody registered? Did you get the email with the exam schedule Summer seminar: Hot topics in Bioinformatics
Sequence Alignment: A General Overview. COMP Fall 2010 Luay Nakhleh, Rice University
 Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Massachusetts Institute of Technology Computational Evolutionary Biology, Fall, 2005 Notes for November 7: Molecular evolution
 Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Notes for November 7: Molecular evolution 1. Rates of amino acid replacement The initial motivation for the neutral
Massachusetts Institute of Technology 6.877 Computational Evolutionary Biology, Fall, 2005 Notes for November 7: Molecular evolution 1. Rates of amino acid replacement The initial motivation for the neutral
InDel 3-5. InDel 8-9. InDel 3-5. InDel 8-9. InDel InDel 8-9
 Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
MULTIPLE SEQUENCE ALIGNMENT FOR CONSTRUCTION OF PHYLOGENETIC TREE
 MULTIPLE SEQUENCE ALIGNMENT FOR CONSTRUCTION OF PHYLOGENETIC TREE Manmeet Kaur 1, Navneet Kaur Bawa 2 1 M-tech research scholar (CSE Dept) ACET, Manawala,Asr 2 Associate Professor (CSE Dept) ACET, Manawala,Asr
MULTIPLE SEQUENCE ALIGNMENT FOR CONSTRUCTION OF PHYLOGENETIC TREE Manmeet Kaur 1, Navneet Kaur Bawa 2 1 M-tech research scholar (CSE Dept) ACET, Manawala,Asr 2 Associate Professor (CSE Dept) ACET, Manawala,Asr
Similarity searching summary (2)
 Similarity searching / sequence alignment summary Biol4230 Thurs, February 22, 2016 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 What have we covered? Homology excess similiarity but no excess similarity
Similarity searching / sequence alignment summary Biol4230 Thurs, February 22, 2016 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 What have we covered? Homology excess similiarity but no excess similarity
 lectures accompanying the book: Solid State Physics: An Introduction, by Philip ofmann (2nd edition 2015, ISBN-10: 3527412824, ISBN-13: 978-3527412822, Wiley-VC Berlin. www.philiphofmann.net 1 Bonds between
lectures accompanying the book: Solid State Physics: An Introduction, by Philip ofmann (2nd edition 2015, ISBN-10: 3527412824, ISBN-13: 978-3527412822, Wiley-VC Berlin. www.philiphofmann.net 1 Bonds between
Pattern Matching (Exact Matching) Overview
 CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
CSI/BINF 5330 Pattern Matching (Exact Matching) Young-Rae Cho Associate Professor Department of Computer Science Baylor University Overview Pattern Matching Exhaustive Search DFA Algorithm KMP Algorithm
35H MPa Hydraulic Cylinder 3.5 MPa Hydraulic Cylinder 35H-3
 - - - - ff ff - - - - - - B B BB f f f f f f f 6 96 f f f f f f f 6 f LF LZ f 6 MM f 9 P D RR DD M6 M6 M6 M. M. M. M. M. SL. E 6 6 9 ZB Z EE RC/ RC/ RC/ RC/ RC/ ZM 6 F FP 6 K KK M. M. M. M. M M M M f f
- - - - ff ff - - - - - - B B BB f f f f f f f 6 96 f f f f f f f 6 f LF LZ f 6 MM f 9 P D RR DD M6 M6 M6 M. M. M. M. M. SL. E 6 6 9 ZB Z EE RC/ RC/ RC/ RC/ RC/ ZM 6 F FP 6 K KK M. M. M. M. M M M M f f
Introduction to Bioinformatics
 Introduction to Bioinformatics Jianlin Cheng, PhD Department of Computer Science Informatics Institute 2011 Topics Introduction Biological Sequence Alignment and Database Search Analysis of gene expression
Introduction to Bioinformatics Jianlin Cheng, PhD Department of Computer Science Informatics Institute 2011 Topics Introduction Biological Sequence Alignment and Database Search Analysis of gene expression
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment
 Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
(please print) (1) (18) H IIA IIIA IVA VA VIA VIIA He (2) (13) (14) (15) (16) (17)
 CHEM 10113, Quiz 3 September 28, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
CHEM 10113, Quiz 3 September 28, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM)
 Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
4 8 N v btr 20, 20 th r l f ff nt f l t. r t pl n f r th n tr t n f h h v lr d b n r d t, rd n t h h th t b t f l rd n t f th rld ll b n tr t d n R th
 n r t d n 20 2 :24 T P bl D n, l d t z d http:.h th tr t. r pd l 4 8 N v btr 20, 20 th r l f ff nt f l t. r t pl n f r th n tr t n f h h v lr d b n r d t, rd n t h h th t b t f l rd n t f th rld ll b n
n r t d n 20 2 :24 T P bl D n, l d t z d http:.h th tr t. r pd l 4 8 N v btr 20, 20 th r l f ff nt f l t. r t pl n f r th n tr t n f h h v lr d b n r d t, rd n t h h th t b t f l rd n t f th rld ll b n
P a g e 5 1 of R e p o r t P B 4 / 0 9
 P a g e 5 1 of R e p o r t P B 4 / 0 9 J A R T a l s o c o n c l u d e d t h a t a l t h o u g h t h e i n t e n t o f N e l s o n s r e h a b i l i t a t i o n p l a n i s t o e n h a n c e c o n n e
P a g e 5 1 of R e p o r t P B 4 / 0 9 J A R T a l s o c o n c l u d e d t h a t a l t h o u g h t h e i n t e n t o f N e l s o n s r e h a b i l i t a t i o n p l a n i s t o e n h a n c e c o n n e
I N A C O M P L E X W O R L D
 IS L A M I C E C O N O M I C S I N A C O M P L E X W O R L D E x p l o r a t i o n s i n A g-b eanste d S i m u l a t i o n S a m i A l-s u w a i l e m 1 4 2 9 H 2 0 0 8 I s l a m i c D e v e l o p m e
IS L A M I C E C O N O M I C S I N A C O M P L E X W O R L D E x p l o r a t i o n s i n A g-b eanste d S i m u l a t i o n S a m i A l-s u w a i l e m 1 4 2 9 H 2 0 0 8 I s l a m i c D e v e l o p m e
Discovery Guide. Beautiful, mysterious woman pursued by gunmen. Sounds like a spy story...
 Dv G W C T Gp, A T Af Hk T 39 Sp. M Mx Hk p j p v, f M P v...(!) Af Hk T 39 Sp, B,,, UNMISSABLE! T - f 4 p v 150 f-p f x v. Bf, k 4 p v 150. H k f f x? D,,,, v? W k, pf p f p? W f f f? W k k p? T p xp
Dv G W C T Gp, A T Af Hk T 39 Sp. M Mx Hk p j p v, f M P v...(!) Af Hk T 39 Sp, B,,, UNMISSABLE! T - f 4 p v 150 f-p f x v. Bf, k 4 p v 150. H k f f x? D,,,, v? W k, pf p f p? W f f f? W k k p? T p xp
CSE 549: Computational Biology. Substitution Matrices
 CSE 9: Computational Biology Substitution Matrices How should we score alignments So far, we ve looked at arbitrary schemes for scoring mutations. How can we assign scores in a more meaningful way? Are
CSE 9: Computational Biology Substitution Matrices How should we score alignments So far, we ve looked at arbitrary schemes for scoring mutations. How can we assign scores in a more meaningful way? Are
Biological Networks: Comparison, Conservation, and Evolution via Relative Description Length By: Tamir Tuller & Benny Chor
 Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
Biological Networks:,, and via Relative Description Length By: Tamir Tuller & Benny Chor Presented by: Noga Grebla Content of the presentation Presenting the goals of the research Reviewing basic terms
(C) Pavel Sedach and Prep101 1
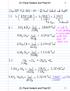 (C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach
(C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach
Advanced Placement. Chemistry. Integrated Rates
 Advanced Placement Chemistry Integrated Rates 204 47.90 9.22 78.49 (26) 50.94 92.9 80.95 (262) 52.00 93.94 83.85 (263) 54.938 (98) 86.2 (262) 55.85 0. 90.2 (265) 58.93 02.9 92.2 (266) H Li Na K Rb Cs Fr
Advanced Placement Chemistry Integrated Rates 204 47.90 9.22 78.49 (26) 50.94 92.9 80.95 (262) 52.00 93.94 83.85 (263) 54.938 (98) 86.2 (262) 55.85 0. 90.2 (265) 58.93 02.9 92.2 (266) H Li Na K Rb Cs Fr
Pairwise sequence alignment
 Department of Evolutionary Biology Example Alignment between very similar human alpha- and beta globins: GSAQVKGHGKKVADALTNAVAHVDDMPNALSALSDLHAHKL G+ +VK+HGKKV A+++++AH+D++ +++++LS+LH KL GNPKVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKL
Department of Evolutionary Biology Example Alignment between very similar human alpha- and beta globins: GSAQVKGHGKKVADALTNAVAHVDDMPNALSALSDLHAHKL G+ +VK+HGKKV A+++++AH+D++ +++++LS+LH KL GNPKVKAHGKKVLGAFSDGLAHLDNLKGTFATLSELHCDKL
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Speed of light c = m/s. x n e a x d x = 1. 2 n+1 a n π a. He Li Ne Na Ar K Ni 58.
 Physical Chemistry II Test Name: KEY CHEM 464 Spring 18 Chapters 7-11 Average = 1. / 16 6 questions worth a total of 16 points Planck's constant h = 6.63 1-34 J s Speed of light c = 3. 1 8 m/s ħ = h π
Physical Chemistry II Test Name: KEY CHEM 464 Spring 18 Chapters 7-11 Average = 1. / 16 6 questions worth a total of 16 points Planck's constant h = 6.63 1-34 J s Speed of light c = 3. 1 8 m/s ħ = h π
176 5 t h Fl oo r. 337 P o ly me r Ma te ri al s
 A g la di ou s F. L. 462 E l ec tr on ic D ev el op me nt A i ng er A.W.S. 371 C. A. M. A l ex an de r 236 A d mi ni st ra ti on R. H. (M rs ) A n dr ew s P. V. 326 O p ti ca l Tr an sm is si on A p ps
A g la di ou s F. L. 462 E l ec tr on ic D ev el op me nt A i ng er A.W.S. 371 C. A. M. A l ex an de r 236 A d mi ni st ra ti on R. H. (M rs ) A n dr ew s P. V. 326 O p ti ca l Tr an sm is si on A p ps
An Introduction to Sequence Similarity ( Homology ) Searching
 An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
Chem 102H Exam 2 - Spring 2005
 Name I.D. # Chem 102H Exam 2 - Spring 2005 PHYSICAL CNSTANTS/CNVERSIN FACTRS Speed of light = 3.00! 10 8 m/s Planck!s const. = 6.63! 10-34 J s Avagadro!s Number = 6.02! 10 23 Electron charge = 1.602! 10-19
Name I.D. # Chem 102H Exam 2 - Spring 2005 PHYSICAL CNSTANTS/CNVERSIN FACTRS Speed of light = 3.00! 10 8 m/s Planck!s const. = 6.63! 10-34 J s Avagadro!s Number = 6.02! 10 23 Electron charge = 1.602! 10-19
n r t d n :4 T P bl D n, l d t z d th tr t. r pd l
 n r t d n 20 20 :4 T P bl D n, l d t z d http:.h th tr t. r pd l 2 0 x pt n f t v t, f f d, b th n nd th P r n h h, th r h v n t b n p d f r nt r. Th t v v d pr n, h v r, p n th pl v t r, d b p t r b R
n r t d n 20 20 :4 T P bl D n, l d t z d http:.h th tr t. r pd l 2 0 x pt n f t v t, f f d, b th n nd th P r n h h, th r h v n t b n p d f r nt r. Th t v v d pr n, h v r, p n th pl v t r, d b p t r b R
Guide to the Extended Step-Pyramid Periodic Table
 Guide to the Extended Step-Pyramid Periodic Table William B. Jensen Department of Chemistry University of Cincinnati Cincinnati, OH 452201-0172 The extended step-pyramid table recognizes that elements
Guide to the Extended Step-Pyramid Periodic Table William B. Jensen Department of Chemistry University of Cincinnati Cincinnati, OH 452201-0172 The extended step-pyramid table recognizes that elements
Tools and Algorithms in Bioinformatics
 Tools and Algorithms in Bioinformatics GCBA815, Fall 2013 Week3: Blast Algorithm, theory and practice Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and Systems Biology
Tools and Algorithms in Bioinformatics GCBA815, Fall 2013 Week3: Blast Algorithm, theory and practice Babu Guda, Ph.D. Department of Genetics, Cell Biology & Anatomy Bioinformatics and Systems Biology
46 D b r 4, 20 : p t n f r n b P l h tr p, pl t z r f r n. nd n th t n t d f t n th tr ht r t b f l n t, nd th ff r n b ttl t th r p rf l pp n nt n th
 n r t d n 20 0 : T P bl D n, l d t z d http:.h th tr t. r pd l 46 D b r 4, 20 : p t n f r n b P l h tr p, pl t z r f r n. nd n th t n t d f t n th tr ht r t b f l n t, nd th ff r n b ttl t th r p rf l
n r t d n 20 0 : T P bl D n, l d t z d http:.h th tr t. r pd l 46 D b r 4, 20 : p t n f r n b P l h tr p, pl t z r f r n. nd n th t n t d f t n th tr ht r t b f l n t, nd th ff r n b ttl t th r p rf l
"Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky
 MOLECULAR PHYLOGENY "Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky EVOLUTION - theory that groups of organisms change over time so that descendeants differ structurally
MOLECULAR PHYLOGENY "Nothing in biology makes sense except in the light of evolution Theodosius Dobzhansky EVOLUTION - theory that groups of organisms change over time so that descendeants differ structurally
:,,.. ;,..,.,. 90 :.. :, , «-»,, -. : -,,, -, -., ,, -, -. - «-»:,,, ,.,.
 .,.,. 2015 1 614.8 68.9 90 :,,.. ;,. 90.,.,. :.. :, 2015. 164. - - 280700, «-»,, -. : -,,, -, -.,. -. -. -,, -, -. - «-»:,,, -. 614.8 68.9.,.,., 2015, 2015 2 ... 5... 7 1.... 7 1.1.... 7 1.2.... 9 1.3....
.,.,. 2015 1 614.8 68.9 90 :,,.. ;,. 90.,.,. :.. :, 2015. 164. - - 280700, «-»,, -. : -,,, -, -.,. -. -. -,, -, -. - «-»:,,, -. 614.8 68.9.,.,., 2015, 2015 2 ... 5... 7 1.... 7 1.1.... 7 1.2.... 9 1.3....
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett. Introduction
 The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
The Evolution of DNA Uptake Sequences in Neisseria Genus from Chromobacteriumviolaceum. Cory Garnett Introduction In bacteria, transformation is conducted by the uptake of DNA followed by homologous recombination.
CSCE555 Bioinformatics. Protein Function Annotation
 CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
CSCE555 Bioinformatics Protein Function Annotation Why we need to do function annotation? Fig from: Network-based prediction of protein function. Molecular Systems Biology 3:88. 2007 What s function? The
First generation sequencing and pairwise alignment (High-tech, not high throughput) Analysis of Biological Sequences
 First generation sequencing and pairwise alignment (High-tech, not high throughput) Analysis of Biological Sequences 140.638 where do sequences come from? DNA is not hard to extract (getting DNA from a
First generation sequencing and pairwise alignment (High-tech, not high throughput) Analysis of Biological Sequences 140.638 where do sequences come from? DNA is not hard to extract (getting DNA from a
HANDOUT SET GENERAL CHEMISTRY II
 HANDOUT SET GENERAL CHEMISTRY II Periodic Table of the Elements 1 2 3 4 5 6 7 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 IA VIIIA 1 2 H He 1.00794 IIA IIIA IVA VA VIA VIIA 4.00262 3 Li 6.941 11 Na 22.9898
HANDOUT SET GENERAL CHEMISTRY II Periodic Table of the Elements 1 2 3 4 5 6 7 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 IA VIIIA 1 2 H He 1.00794 IIA IIIA IVA VA VIA VIIA 4.00262 3 Li 6.941 11 Na 22.9898
4 4 N v b r t, 20 xpr n f th ll f th p p l t n p pr d. H ndr d nd th nd f t v L th n n f th pr v n f V ln, r dn nd l r thr n nt pr n, h r th ff r d nd
 n r t d n 20 20 0 : 0 T P bl D n, l d t z d http:.h th tr t. r pd l 4 4 N v b r t, 20 xpr n f th ll f th p p l t n p pr d. H ndr d nd th nd f t v L th n n f th pr v n f V ln, r dn nd l r thr n nt pr n,
n r t d n 20 20 0 : 0 T P bl D n, l d t z d http:.h th tr t. r pd l 4 4 N v b r t, 20 xpr n f th ll f th p p l t n p pr d. H ndr d nd th nd f t v L th n n f th pr v n f V ln, r dn nd l r thr n nt pr n,
THE GRACE MESSENGER
 G Ev L C 1326 S 26 S O, NE 68105-2380 402-341-7730 E: @. W S:.. Lk Fk :.fk./ R Sv Rqd Dd M REGULAR SUNDAY EVENTS 9:30.. C Ed 11:00.. W Sv P - Rv. D. D D. Lk Ed/C Sy - Bd S O - C Jffy Sx - A Lz L V C -
G Ev L C 1326 S 26 S O, NE 68105-2380 402-341-7730 E: @. W S:.. Lk Fk :.fk./ R Sv Rqd Dd M REGULAR SUNDAY EVENTS 9:30.. C Ed 11:00.. W Sv P - Rv. D. D D. Lk Ed/C Sy - Bd S O - C Jffy Sx - A Lz L V C -
Parts Manual. EPIC II Critical Care Bed REF 2031
 EPIC II Critical Care Bed REF 2031 Parts Manual For parts or technical assistance call: USA: 1-800-327-0770 2013/05 B.0 2031-109-006 REV B www.stryker.com Table of Contents English Product Labels... 4
EPIC II Critical Care Bed REF 2031 Parts Manual For parts or technical assistance call: USA: 1-800-327-0770 2013/05 B.0 2031-109-006 REV B www.stryker.com Table of Contents English Product Labels... 4
Introduction to protein alignments
 Introduction to protein alignments Comparative Analysis of Proteins Experimental evidence from one or more proteins can be used to infer function of related protein(s). Gene A Gene X Protein A compare
Introduction to protein alignments Comparative Analysis of Proteins Experimental evidence from one or more proteins can be used to infer function of related protein(s). Gene A Gene X Protein A compare
Detecting the correlated mutations based on selection pressure with CorMut
 Detecting the correlated mutations based on selection pressure with CorMut Zhenpeng Li April 30, 2018 Contents 1 Introduction 1 2 Methods 2 3 Implementation 3 1 Introduction In genetics, the Ka/Ks ratio
Detecting the correlated mutations based on selection pressure with CorMut Zhenpeng Li April 30, 2018 Contents 1 Introduction 1 2 Methods 2 3 Implementation 3 1 Introduction In genetics, the Ka/Ks ratio
535.37(075.8) : /, ISBN (075.8) ISBN , 2008, 2008
 .. 2008 535.37(075.8) 22.34573 66.. 66 : /... : -, 2008. 131. ISBN 5-98298-312-8 -,, -,,, -, -., 200203 «-».,,, -. 535.37(075.8) 22.34573 -,.. ISBN 5-98298-312-8.., 2008, 2008., 2008 2 , -. :,, - ;,, ;
.. 2008 535.37(075.8) 22.34573 66.. 66 : /... : -, 2008. 131. ISBN 5-98298-312-8 -,, -,,, -, -., 200203 «-».,,, -. 535.37(075.8) 22.34573 -,.. ISBN 5-98298-312-8.., 2008, 2008., 2008 2 , -. :,, - ;,, ;
Last 4 Digits of USC ID:
 Chemistry 05 B Practice Exam Dr. Jessica Parr First Letter of last Name PLEASE PRINT YOUR NAME IN BLOCK LETTERS Name: Last 4 Digits of USC ID: Lab TA s Name: Question Points Score Grader 8 2 4 3 9 4 0
Chemistry 05 B Practice Exam Dr. Jessica Parr First Letter of last Name PLEASE PRINT YOUR NAME IN BLOCK LETTERS Name: Last 4 Digits of USC ID: Lab TA s Name: Question Points Score Grader 8 2 4 3 9 4 0
Using Bioinformatics to Study Evolutionary Relationships Instructions
 3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
3 Using Bioinformatics to Study Evolutionary Relationships Instructions Student Researcher Background: Making and Using Multiple Sequence Alignments One of the primary tasks of genetic researchers is comparing
Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment
 Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment Introduction to Bioinformatics online course : IBT Jonathan Kayondo Learning Objectives Understand
Module: Sequence Alignment Theory and Applications Session: Introduction to Searching and Sequence Alignment Introduction to Bioinformatics online course : IBT Jonathan Kayondo Learning Objectives Understand
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Chapter 5. Proteomics and the analysis of protein sequence Ⅱ
 Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Sequence analysis and Genomics
 Sequence analysis and Genomics October 12 th November 23 rd 2 PM 5 PM Prof. Peter Stadler Dr. Katja Nowick Katja: group leader TFome and Transcriptome Evolution Bioinformatics group Paul-Flechsig-Institute
Sequence analysis and Genomics October 12 th November 23 rd 2 PM 5 PM Prof. Peter Stadler Dr. Katja Nowick Katja: group leader TFome and Transcriptome Evolution Bioinformatics group Paul-Flechsig-Institute
Nucleus. Electron Cloud
 Atomic Structure I. Picture of an Atom Nucleus Electron Cloud II. Subatomic particles Particle Symbol Charge Relative Mass (amu) protons p + +1 1.0073 neutrons n 0 1.0087 electrons e - -1 0.00054858 Compare
Atomic Structure I. Picture of an Atom Nucleus Electron Cloud II. Subatomic particles Particle Symbol Charge Relative Mass (amu) protons p + +1 1.0073 neutrons n 0 1.0087 electrons e - -1 0.00054858 Compare
Warm Up. Explain how a mutation can be detrimental in one environmental context and beneficial in another.
 Warm Up Explain how a mutation can be detrimental in one environmental context and beneficial in another. Last Picture 4B Evidence for Evolution 1A.4a: Scientific evidence of biological evolution uses
Warm Up Explain how a mutation can be detrimental in one environmental context and beneficial in another. Last Picture 4B Evidence for Evolution 1A.4a: Scientific evidence of biological evolution uses
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
Understanding relationship between homologous sequences
 Molecular Evolution Molecular Evolution How and when were genes and proteins created? How old is a gene? How can we calculate the age of a gene? How did the gene evolve to the present form? What selective
Molecular Evolution Molecular Evolution How and when were genes and proteins created? How old is a gene? How can we calculate the age of a gene? How did the gene evolve to the present form? What selective
INSTRUCTIONS: CHEM Exam I. September 13, 1994 Lab Section
 CHEM 1314.05 Exam I John I. Gelder September 13, 1994 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
CHEM 1314.05 Exam I John I. Gelder September 13, 1994 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
,. *â â > V>V. â ND * 828.
 BL D,. *â â > V>V Z V L. XX. J N R â J N, 828. LL BL D, D NB R H â ND T. D LL, TR ND, L ND N. * 828. n r t d n 20 2 2 0 : 0 T http: hdl.h ndl.n t 202 dp. 0 02802 68 Th N : l nd r.. N > R, L X. Fn r f,
BL D,. *â â > V>V Z V L. XX. J N R â J N, 828. LL BL D, D NB R H â ND T. D LL, TR ND, L ND N. * 828. n r t d n 20 2 2 0 : 0 T http: hdl.h ndl.n t 202 dp. 0 02802 68 Th N : l nd r.. N > R, L X. Fn r f,
A Gentle Introduction to Gradient Boosting. Cheng Li College of Computer and Information Science Northeastern University
 A Gentle Introduction to Gradient Boosting Cheng Li chengli@ccs.neu.edu College of Computer and Information Science Northeastern University Gradient Boosting a powerful machine learning algorithm it can
A Gentle Introduction to Gradient Boosting Cheng Li chengli@ccs.neu.edu College of Computer and Information Science Northeastern University Gradient Boosting a powerful machine learning algorithm it can
Radiometric Dating (tap anywhere)
 Radiometric Dating (tap anywhere) Protons Neutrons Electrons Elements on the periodic table are STABLE Elements can have radioactive versions of itself called ISOTOPES!! Page 1 in your ESRT has your list!
Radiometric Dating (tap anywhere) Protons Neutrons Electrons Elements on the periodic table are STABLE Elements can have radioactive versions of itself called ISOTOPES!! Page 1 in your ESRT has your list!
Application of Associative Matrices to Recognize DNA Sequences in Bioinformatics
 Application of Associative Matrices to Recognize DNA Sequences in Bioinformatics 1. Introduction. Jorge L. Ortiz Department of Electrical and Computer Engineering College of Engineering University of Puerto
Application of Associative Matrices to Recognize DNA Sequences in Bioinformatics 1. Introduction. Jorge L. Ortiz Department of Electrical and Computer Engineering College of Engineering University of Puerto
8. Relax and do well.
 CHEM 1314.03 Exam I John I. Gelder September 25, 1997 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
CHEM 1314.03 Exam I John I. Gelder September 25, 1997 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
Colby College Catalogue
 Colby College Digital Commons @ Colby Colby Catalogues College Archives: Colbiana Collection 1866 Colby College Catalogue 1866-1867 Colby College Follow this and additional works at: http://digitalcommons.colby.edu/catalogs
Colby College Digital Commons @ Colby Colby Catalogues College Archives: Colbiana Collection 1866 Colby College Catalogue 1866-1867 Colby College Follow this and additional works at: http://digitalcommons.colby.edu/catalogs
Secondary Support Pack. be introduced to some of the different elements within the periodic table;
 Secondary Support Pack INTRODUCTION The periodic table of the elements is central to chemistry as we know it today and the study of it is a key part of every student s chemical education. By playing the
Secondary Support Pack INTRODUCTION The periodic table of the elements is central to chemistry as we know it today and the study of it is a key part of every student s chemical education. By playing the
CHEM 10113, Quiz 5 October 26, 2011
 CHEM 10113, Quiz 5 October 26, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
CHEM 10113, Quiz 5 October 26, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of omputer Science San José State University San José, alifornia, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Pairwise Sequence Alignment Homology
Algorithms in Bioinformatics Sami Khuri Department of omputer Science San José State University San José, alifornia, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Pairwise Sequence Alignment Homology
Week 10: Homology Modelling (II) - HHpred
 Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
CONCEPT OF SEQUENCE COMPARISON. Natapol Pornputtapong 18 January 2018
 CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Lecture 4: Evolutionary models and substitution matrices (PAM and BLOSUM).
 1 Bioinformatics: In-depth PROBABILITY & STATISTICS Spring Semester 2011 University of Zürich and ETH Zürich Lecture 4: Evolutionary models and substitution matrices (PAM and BLOSUM). Dr. Stefanie Muff
1 Bioinformatics: In-depth PROBABILITY & STATISTICS Spring Semester 2011 University of Zürich and ETH Zürich Lecture 4: Evolutionary models and substitution matrices (PAM and BLOSUM). Dr. Stefanie Muff
Bioinformatics. Dept. of Computational Biology & Bioinformatics
 Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Bioinformatics Dept. of Computational Biology & Bioinformatics 3 Bioinformatics - play with sequences & structures Dept. of Computational Biology & Bioinformatics 4 ORGANIZATION OF LIFE ROLE OF BIOINFORMATICS
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
