The Fox gene family in Xenopus laevis : FoxI2, FoxM1 and FoxP1 in early development
|
|
|
- Eleanore Griffith
- 6 years ago
- Views:
Transcription
1 Int. J. Dev. iol. 49: (2004) doi: /ijdb bp Developmental Expression Pattern The Fox gene family in Xenopus laevis : FoxI2, FoxM1 and FoxP1 in early development RR S. POHL, NTJE RÖSSNER and WLTER KNÖCHEL* bt. iochemie, Universität Ulm, Ulm, Germany STRCT We here describe the sequences and expression patterns of three new Fox (fork head box) transcription factors in Xenopus laevis embryos. xlfoxi2, another member of subclass I, is maternally transcribed. Zygotic transcripts are first detected during neurulation and become localised to the dorsal part of epibranchial placodes. xlfoxm1 like xlfoxp1 are the first members of subclasses M and P described in Xenopus. oth genes are maternally expressed and transcripts are found during early cleavage stages in the animal blastomeres. xlfoxm1 is strongly upregulated during neurula stages and transcripts are localised in the neuroectoderm. Later, expression is found in the spinal cord, the rhombencephalon, the retina and in the branchial arches. xlfoxp1 is activated during organogenesis and is mainly expressed in the brain, head mesenchyme and in the splanchnic layer of the lateral plate mesoderm. KEY WORDS: X. laevis, embryogenesis, fork head/winged helix factor, expression pattern Fox (fork head box) transcription factors are involved in various signalling pathways and cell fate decisions throughout development. ccording to conserved amino acid residues at distinct positions within the winged helix domain, these factors are categorised into the subclasses to Q (Kaestner et al., 2000). While for the mouse and human species, Fox family members from all subclasses have been reported, in the South frican clawed frog, Xenopus laevis, currently only the subclasses to L are known. Here we report the sequence and initial characterisation of two additional Xenopus laevis Fox gene family members, xlfoxm1 and xlfoxp1 and add xlfoxi2 as a new member to the already existing subclass I. We describe the full length cdns encoding these proteins and investigate the temporal and spatial expression patterns of corresponding genes by RT-PCR and by whole mount in situ hybridisations. Results and Discussion xlfoxi2 The FoxI subclass is already known in different organisms to contain several members and has recently been analysed regarding the phylogenetic relationship inside the subclass based on the zebrafish members (Solomon et al., 2003). y searching in EST data bases we have found an IMGE-clone ( ), which covers the complete translated region of a FoxI subclass member in Xenopus laevis. Referring to the suggested terminology regard- ing subclass I (Solomon et al., 2003), this clone is preliminarily designated as xlfoxi2, even if the true orthologue relationship inside this subclass remains to be elucidated. The complete sequence of the IMGE-clone has been determined and deposited under EML accession number J Starting 78 bp in front of the start codon, the complete cdn sequence comprises 1537 bp and a poly() tail. The translated region encompassing 1107 bp gives rise to a protein of 369 amino acids, with the fork head domain located between amino acids 115 and 224. The predicted amino acid sequence is aligned to the pseudo-allelelic variants xlfoxi1a and xlfoxi1b (Lef et al., 1994) as well as to the close relative xlfoxi1c (Pohl et al., 2002) (Fig. 1). While xlfoxi2 is 44% identical to xlfoxi1a and 46% to xlfoxi1b, it shows 48% identity to xlfoxi1c. However, conservation within the winged helix domain exhibits 84% and 86% identity between xlfoxi2 compared to xlfoxi1a and to xlfoxi1b, respectively and 88% to xlfox1c. Interestingly, all so far identified members of subclass I, xlfoxi1a/b, xlfoxi1c and xlfoxi2, show characteristic and different temporal expression patterns (Fig. 2) (Pohl et al., 2002). Starting with high amounts of maternal transcripts, xlfoxi2 expression decreases rapidly during early cleavage stages. Zygotic expression starts at neurulation and continues at low levels at stage 39. The xlfoxi1a/b genes, which exhibit identical patterns (Lef et al., 1994), are strongly upregulated bbreviations used in this paper: Fox, fork head box. *ddress correspondence to: Dr. W. Knöchel. bteilung iochemie, Universität Ulm, lbert-einstein-llee 11, D Ulm, Germany. Fax: walter.knoechel@medizin.uni-ulm.de /2005/$25.00 UC Press Printed in Spain
2 54.S. Pohl et al. during the early blastula stage reaching a maximum number of transcripts at the gastrula stage. Thereafter, the amount of transcripts declines during neurulation, with low levels even detected at stage 40. Finally, xlfoxi1c starts expression at the late gastrula stage and transcripts constantly accumulate during later embryogenesis (Pohl et al., 2002). Thus it is evident that the genes belonging to subclass I are subject to different regulatory mechanisms and that different developmental stages are associated with the expression of distinct xlfoxi proteins. Results obtained by whole mount in situ hybridisation confirm the temporal patterns as determined by RT-PCR. High amounts of xlfoxi2 transcripts are present at early cleavage stages in the animal half of the blastomeres (Fig. 3, ). Zygotic expression is exclusively found in the placodes of the head (Fig. 3C). In contrast to the placodal expression of xlfoxi1a (Fig. 3F) and xlfoxi1c (Fig. 3G), xlfoxi2 is only expressed in the dorsal part of the epibranchial placodes (for an overview see Schlosser and Northcutt, 2000). s shown by a transverse section (Fig. 3D), stained xlfoxi2 positive cells are located between the mesodermal foregut tissue and the more dorsal head mesenchyme. horizontal section demonstrates that xlfoxi2 is expressed within a restricted region located near the tip of the first, second and third visceral pouch (Fig. 3E). xlfoxm1 FoxM1 was originally isolated in a search for proteins that are phosphorylated during M-phase (Westendorf et al., 1994) without having been identified as a winged helix factor. fter the isolation of the complete cdn from mouse (Genbank accession number: NM_008021, Korver et al., 1997a) and the human orthologue (Genbank accession number: NM_202002, Ye et al., 1997; Korver et al., 1997 b; Yao et al., 1997) it became clear, that the expression of FoxM1 (also named MPP2, Trident, FKHL16, HFH-11 and WIN) is strictly correlated to proliferating cells. xlfoxm1 was originally identified in form of the incomplete ESTclones J and XL058d22. Screening of a stage 30 cdn library led to the isolation of an incomplete cdn overlapping with XL058d22. With 5 -primers derived from J and 3 -primers derived from our cdn, the complete coding sequence could finally be amplified from a reverse transcribed stage 30 RN. The cdn contains 2762 bp and a poly() tail (deposited under EML accession number J853462). The protein is encoded by 2277 bp, thus comprising 759 amino acids. The fork head domain is located at the N-terminal half in between amino acids 251 and 360. Figure 4 shows an alignment of the Xenopus sequence to the human and mouse orthologues. While the human and mouse proteins share 79% identity, the identities between xlfoxm1 and human Fig. 1. FoxI2 sequences. lignment of the predicted amino acid sequence of Xenopus laevis (x.l.) FoxI2 to FoxI1a, FoxI1b (Lef et al., 1994) and FoxI1c (Pohl et al., 2002). Identical residues are shown in blue. The fork head domain is shaded in gray. FOXM1 (36%) and mouse Foxm1 (37%) are rather low. However, this low degree is mainly due to a stretch of deviating amino acids following the fork head domain. In this context it should be noted that several mammalian splice variants are described (Yao et al., 1997). djacent to this region, the rate of identity is significantly higher and that of the winged helix domains is even 84% (see Fig. 4, calculated by DILIGN, Morgenstern et al., 1998). To determine the temporal expression pattern of xlfoxm1 in Xenopus embryos, RNs of different developmental stages were analysed by RT-PCR. s shown in Fig. 2, maternal gene transcription yields high amounts of RN in early cleavage stages, but transcripts are rapidly degraded until the blastula stage. Zygotic expression of xlfoxm1 starts during neurulation and transcripts persist and accumulate until the end of organogenesis. The spatial expression was determined by in situ hybridisation (Fig. 5). Transcripts are found in the animal half at early cleavage stages (Fig. 5), but are absent from gastrula stage embryos. During neurulation, expression is observed in the neural folds and, later, in the spinal cord as well as in the eye field (Fig. 5, C). This localisation becomes even more prominent at stage 22, when transcripts demarcate the region of the eye (Fig. 5D). Thus, it can
3 xlfoxi2, xlfoxm1 and xlfoxp1 genes 55 Fig. 2. Temporal expression of xlfoxi2, xlfoxm1 and xlfoxp1. RT-PCR for xlfoxi2, xlfoxm1 and xlfoxp1 was performed with RNs of different developmental stages. (-) indicates a negative control in the absence of RN. Histone H4 was used as internal control. be concluded that xlfoxm1 is involved in early stages of eye formation. During tailbud stages, xlfoxm1 expression is still restricted to the neuroectoderm, predominantly to the hindbrain, the eye and the spinal cord (Fig. 5E). With ongoing development, expression is also found at lower levels in the branchial arches (Fig. 5F). Sections of embryos at stage 35 reveal distinct expression of xlfoxm1 in the rhombencephalon, in the eye retina but not the in forming lens and in the branchial arches (Fig. 5 G-I). While in mouse development Foxm1 was found to be ubiquitously localised in all cell types during proliferation, its expression is downregulated when cells enter terminal differentiation (Korver et al., 1997a; Ye et al., 1997). In adult human and mouse tissues, FoxM1 is predominantly found within the thymus and testis and, at a lower extent, in intestine, liver and lung. Interestingly, a Foxm1 knock out mouse was found to be post-natally lethal due to a failure in development of the myocardium. oth for hepatocytes and myocytes, polyploidy was observed in combination with a dramatic increase in DN content (Korver et al., 1998). Human FOXM1 is localised on chromosome 12p13. Chromosomal abnormalities of this region are known from several tumors which were initially explained by the cyclin dependent kinase inhibitor p27 kip1 localised in close proximity (Korver et al., 1997b). However, it has recently been found that FoxM1 itself is an important part of the precise machinery ensuring the correct progression through the cell cycle. This is achieved via a direct regulation of cyclin 1 and D promoters by FoxM1, in association with a diminished expression of p27 Kip1 (reviewed in Costa et al., 2003). xlfoxp1 The subclass P of Fox transcription factors gained special interest, because FOXP2 was found to be associated with language disorders in humans (Lai et al., 2001). y sequence comparisons with great apes, FOXP2 was suggested to be an important target of selection during recent evolution in humans (Enard et al., 2002). FOXP1 was isolated in search for a protein known to be involved in several human tumors (anham et al., 2001). Members of the FoxP subclass are also of interest, because these proteins contain a zinc-finger and a leucine-zipper located N- terminal to their fork head domain. For the first time in case of Fox proteins, FoxP factors are not only able to bind as heterodimers to DN, but were shown to interact with each other due to these motifs (Wang et al., 2003; Li et al., 2004). xlfoxp1 was originally identified by the IMGE-clone However, this clone encodes only the C-terminal part of the protein. Further database searches for sequences encoding the N-terminal part led to additional EST clones (e.g. E679963) which were used to elongate the Xenopus FoxP1 sequence. Finally, the complete coding sequence could be amplified by RT-PCR from stage 30 RN. The xlfoxp1 cdn sequence encompasses 1968 bp starting 54 bp in front to the start codon. It contains 180 bp of the 3 -UTR and, additionally, a poly() tail (deposited under EML accession number J853463). Since the EST-clone contains a stop codon in front of the translation start site, the derived amino acid sequence is shorter than those listed for mammals (homo sapiens Genbank accession number: NM_032682; mus musculus Genbank accession number: NM_053202). However, it should be mentioned that different isoforms with varying start sites are also reported for mice (Wang et al., 2003). Moreover, translation start site of the zebrafish FoxP1 ( ENSDRT ) and, interestingly, also of the closely related FoxP2 gene in human (NM_014491) were uniformly determined to the respective start codon as we have found for xlfoxp1. The open reading frame of xlfoxp1 contains 1734 bp encoding 578 amino acids. The fork head domain is located within the C-terminal half, a rather unusual feature for a Fox protein. The alignment of FoxP1 protein sequences of X. laevis, human, mouse and zebrafish is shown in Fig. 4. oth on DN and amino acid level, FoxP1 shows the highest homology Fig. 3. Whole mount in situ hybridisation of xlfoxi2. () 2-cell stage, animal view; () 4-cell stage, lateral view; (C) stage 35, the red lines denote sections shown in (D,E) which are transverse and horizontal sections respectively, through the dorsal epibranchial placodes; (F) stage 35 stained for xlfoxi1a; (G) stage 34 stained for xlfoxi1c. he, heart anlage; rh, rhombencephalon; I., II., III., first, second and third visceral pouches. C D E F G
4 56.S. Pohl et al. Fig. 4. FoxM1 and FoxP1 proteins. () lignment of the predicted amino acid sequence of Xenopus laevis (x.l.) FoxM1 with human (h.s.) FOXM1 and mouse (m.m.) Foxm1. () lignment of the predicted amino acid sequence of xlfoxp1 with human (h.s.) FOXP1, mouse (m.m.) Foxp1 and zebrafish (d.r.) FoxP1. Identical residues are shown in blue. The fork head domain is shaded gray. The green box shows the position of the Gli-like zinc-finger with the relevant cysteine and histidine residues denoted by arrows, the red box marks the leucine-zipper, the magenta box represents the CtP-binding side. between different species, that to our knowledge was ever reported for Fox genes. xlfoxp1 protein is 86% identical to its mouse and human orthologues and 70% identical to zebrafish FoxP1. oth the fork head domain (grey) and the zinc-finger motif (green) are highly conserved. The latter was found to be closely related to the first zinc-finger found in Gli-proteins (Shu et al., 2001). Furthermore, the leucine-zipper that is localised directly adjacent to the zincfinger and a binding site for the transcriptional co-repressor CtP- 1 (Li et al., 2004) are also found to be conserved. Fig. 2 shows the temporal expression pattern of xlfoxp1. Low levels of maternal transcripts are present, but disappear before gastrulation. Zygotic transcription starts at stage 26 and continues throughout embryo-
5 xlfoxi2, xlfoxm1 and xlfoxp1 genes 57 C D 6C,F). In the brain the anterior most staining is restricted to the outer region of the mesencephalon (Fig. 6E). With ongoing development additional expression is found in the curling gut (Fig. 6D). This corresponds to the in situ analyses performed in mice, where Foxp1, besides its expression in the lung, is also described in the developing central nervous system and in the intestine (Shu et al., 2001; Tamura et al., 2003). Thus, the relationship between the mammalian and Xenopus FoxP1 genes is not only reflected by sequence homology but also by similar expression patterns. Experimental Procedures RT-PCR, in situ hybridisation and handling of Xenopus embryos was done according to standard procedures (for more details see: Pohl and Knöchel, 2001). Developmental stages were determined according to Nieuwkoop and Faber, The IMGE-clone was commercially obtained by RZPD (Deutsches Resourcenzentrum für Genomforschung GmbH, erlin). Primers used for amplification of complete coding regions for xlfoxp1 and xlfoxm1 are as follows: E F C D G H I Fig. 5. Whole mount in situ hybridisation of xlfoxm1. () 2-cell stage, lateral view; () stage 16, dorsal view, anterior to the bottom; (C) stage 19, anterior view; (D) stage 22, lateral view; (E) stage 26, dorsal view; (F) stage 35; in (D-F), anterior is to the right. Red lines in (F) denote sections shown in (G-I). (G) Transverse section revealing xlfoxm1 expression in the rhombencephalon; (H) transverse section demonstrating additional staining of the branchial arches; (I) horizontal section of the head with staining of the retina, anterior is to the bottom. ba, branchial arches; le, lens; he, heart; pn, pronephros; rh, rhombencephalon. genesis. In situ hybridisations reveal the presence of xlfoxp1 transcripts in the animal blastomeres of early cleavage stages (Fig. 6). t tailbud stages, xlfoxp1 expression is visible in regions of the brain, eyes and the splanchnic mesodermal layer of the lateral plate mesoderm surrounding the gut (Fig. 6,G). t stage 35, xlfoxp1 is expressed within the lens of the eye, in distinct regions of the head mesenchyme and the area anterior to the gut (Fig. E F G Fig. 6. Whole mount in situ hybridisation of xlfoxp1. () 8-cell stage, lateral view; () stage 31; (C) stage 35, red lines in (C) indicate the plane of sections shown in (E-G); (D) stage 41, (-D) are lateral views; (E) horizontal section, anterior is to the bottom; (F,G) transverse sections, dorsal is on top. ep, epidermis; he, heart; hm, head mesenchyme; le, lens; me, mesencephalon; pn, pronephros; ve, brain ventricle.
6 58.S. Pohl et al. xlfoxp1-for: 5 -CCC C G GG TG C C-3 ; xlfoxp1-rev: 5 -TT CTC CT GTC GTC TT TC-3 ; xlfoxm1-for: 5 -TG CT TTT GG CTC TC TG-3 ; xlfoxm1-rev: 5 -GT T G TCT C G CC TTG-3. Primers used in RT-PCR for temporal expression were xlfoxi2-rt-for: 5 -GC GC GT CC TGT TTC C-3 ; xlfoxi2-rt-rev: 5 -GGC CG GG GT TC TG CG-3 ; xlfoxp1-rt-for: 5 -CT GT TCC GC TG CT GC-3 ; xlfoxp1-rt-rev: 5 -GC CTC GT CT GG TTC G-3, xlfoxm1-rt-for: 5 -CCC G GTG TGC GT GG-3; xlfoxm1-rt-rev: 5 -TTC C CGC TTG CTG CTT T-3. cknowledgements We thank M. Köster and M.Schuff for screening of Xenopus cdn libraries and C. Donow for help with in situ hybridisation. This work was supported by grants from the Deutsche Forschungsgemeinschaft (SF 497/3) and by Fonds der Chemischen Industrie. References NHM,.H., ESLEY, N., CMPO, E., FERNNDEZ, P.L., FIDLER, C., GTTER, K., JONES, M., MSON, D.Y., PRIME, J.E., TROUGOUOFF, P., WOOD, K. and CORDELL, J.L. (2001). The FOXP1 winged helix transcription factor is a novel candidate tumor suppressor gene on chromosome 3p. Cancer Res. 61: COST, R.H., KLINICHENKO, V.V., HOLTERMN,.X. and WNG, X. (2003). Transcription factors in liver development, differentiation and regeneration. Hepatology 38: ENRD, W., PRZEWORSKI, M., FISHER, S.E., LI, C.S., WIEE, V., KITNO, T., MONCO,.P. and PO, S. (2002). Molecular evolution of FOXP2, a gene involved in speech and language. Nature 418: KESTNER, K.H., KNÖCHEL, W. and MRTINEZ, D.E. (2000). Unified nomenclature for the winged helix/forkhead transcription factors. Genes Dev. 14: KORVER, W., ROOSE, J. and CLEVERS, H. (1997a). The winged-helix transcription factor Trident is expressed in cycling cells. Nucleic cids Res. 25: KORVER, W., ROOSE, J., HEINEN, K., WEGHUIS, D.O., DE RUIJN, D., VN KESSEL,.G. and CLEVERS, H. (1997b). The human TRIDENT/HFH-11/ FKHL16 gene: structure, localization and promoter characterization. Genomics 46: KORVER, W., SCHILHM, M.W., MOERER, P., VN DEN HOFF, M.J., DM, K., LMERS, W.H., MEDEM, R.H. and CLEVERS, H. (1998). Uncoupling of S phase and mitosis in cardiomyocytes and hepatocytes lacking the winged-helix transcription factor Trident. Curr. iol. 8: LI, C.S., FISHER, S.E., HURST, J.., VRGH-KHDEM, F. and MONCO,.P. (2001). forkhead-domain gene is mutated in a severe speech and language disorder. Nature 413: LEF, J., CLEMENT, J.H., OSCHWLD, R., KÖSTER, M. and KNÖCHEL, W. (1994). Spatial and temporal transcription patterns of the forkhead related XFD-2/XFD-2' genes in Xenopus laevis embryos. Mech. Dev. 45: LI, S., WEIDENFELD, J. and MORRISEY, E.E. (2004). Transcriptional and DN binding activity of the Foxp1/2/4 family is modulated by heterotypic and homotypic protein interactions. Mol. Cell. iol. 24: MORGENSTERN,., FRECH, K., DRESS,. and WERNER, T. (1998). DILIGN: Finding local similarities by multiple sequence alignment. ioinformatics 14: NIEUWKOOP, P.D. and FER, J. (1967). Normal table of Xenopus laevis (Daudin), 2nd edn., Elsevier/North Holland, msterdam. POHL,.S. and KNÖCHEL, W. (2001). Overexpression of the transcriptional repressor FoxD3 prevents neural crest formation in Xenopus embryos. Mech. Dev. 103: POHL,.S., KNÖCHEL, S., DILLINGER, K. and KNÖCHEL, W. (2002). Sequence and expression of Fox2 (XFD-5) and FoxI1c (XFD-10) in Xenopus embryogenesis. Mech. Dev. 117: SCHLOSSER, G. and NORTHCUTT, R.G. (2000). Development of neurogenic placodes in Xenopus laevis. J. Comp. Neurol. 418: SHU, W., YNG, H., ZHNG, L., LU, M.M. and MORRISEY, E.E. (2001). Characterization of a new subfamily of winged-helix/forkhead (Fox) genes that are expressed in the lung and act as transcriptional repressors. J. iol. Chem. 276: SOLOMON, K.S., LOGSDON, J.M., JR. and FRITZ,. (2003). Expression and phylogenetic analyses of three zebrafish FoxI class genes. Dev. Dyn. 228: TMUR, S., MORIKW, Y., IWNISHI, H., HISOK, T. and SEN, E. (2003). Expression pattern of the winged-helix/forkhead transcription factor Foxp1 in the developing central nervous system. Gene Expr. Patterns 3: WNG,., LIN, D., LI, C. and TUCKER, P. (2003). Multiple Domains Define the Expression and Regulatory Properties of Foxp1 Forkhead Transcriptional Repressors. J. iol. Chem. 278: WESTENDORF, J.M., RO, P.N. and GERCE, L. (1994). Cloning of cdns for M- phase phosphoproteins recognized by the MPM2 monoclonal antibody and determination of the phosphorylated epitope. Proc. Natl. cad. Sci. US 91: YO, K.M., SH, M., LU, Z. and WONG, G.G. (1997). Molecular analysis of a novel winged helix protein, WIN. Expression pattern, DN binding property and alternative splicing within the DN binding domain. J. iol. Chem. 272: YE, H., KELLY, T.F., SMDNI, U., LIM, L., RUIO, S., OVERDIER, D.G., ROEUCK, K.. and COST, R.H. (1997). Hepatocyte nuclear factor 3/fork head homolog 11 is expressed in proliferating epithelial and mesenchymal cells of embryonic and adult tissues. Mol. Cell. iol. 17: Received: November 2004 Reviewed by Referees: November 2004 Modified by uthors and ccepted for Publication: January 2005 Edited by: Makoto sashima
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
Early Development in Invertebrates
 Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Zoology. Ectodermal derivatives (ZOO ) Developmental Stages. Developmental Stages
 Developmental Zoology (ZOO 228.1.0) Ectodermal derivatives 1 Developmental Stages Ø Early Development Fertilization Cleavage Gastrulation Neurulation Ø Later Development Organogenesis Larval molts Metamorphosis
Developmental Zoology (ZOO 228.1.0) Ectodermal derivatives 1 Developmental Stages Ø Early Development Fertilization Cleavage Gastrulation Neurulation Ø Later Development Organogenesis Larval molts Metamorphosis
Maternal Control of GermLayer Formation in Xenopus
 Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Supplemental Table 1. Primers used for cloning and PCR amplification in this study
 Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Supplemental Table 1. Primers used for cloning and PCR amplification in this study Target Gene Primer sequence NATA1 (At2g393) forward GGG GAC AAG TTT GTA CAA AAA AGC AGG CTT CAT GGC GCC TCC AAC CGC AGC
Role of Organizer Chages in Late Frog Embryos
 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
The Pax and large Maf families of genes in mammalian eye development
 The Pax and large Maf families of genes in mammalian eye development Vertebrate eye development is dependent on the coordinated action of thousands of genes. A specific group of over one hundred of regulatory
The Pax and large Maf families of genes in mammalian eye development Vertebrate eye development is dependent on the coordinated action of thousands of genes. A specific group of over one hundred of regulatory
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
Group activities: Making animal model of human behaviors e.g. Wine preference model in mice
 Lecture schedule 3/30 Natural selection of genes and behaviors 4/01 Mouse genetic approaches to behavior 4/06 Gene-knockout and Transgenic technology 4/08 Experimental methods for measuring behaviors 4/13
Lecture schedule 3/30 Natural selection of genes and behaviors 4/01 Mouse genetic approaches to behavior 4/06 Gene-knockout and Transgenic technology 4/08 Experimental methods for measuring behaviors 4/13
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
CHAPTER 3 DIFFERENTIAL EXPRESSION OF THE HMG BOX TRANSCRIPTION FACTORS XTCF-3 AND XLEF-1 DURING EARLY XENOPUS DEVELOPMENT.
 CHAPTER 3 DFFERENTAL EXPRESSON OF THE HMG BOX TRANSCRPTON FACTORS XTCF-3 AND XLEF-1 DURNG EARLY XENOPUS DEVELOPMENT. Miranda Molenaar, Jeroen Roose #, Josi Peterson, Serenella Venanzi*, Hans Clevers #
CHAPTER 3 DFFERENTAL EXPRESSON OF THE HMG BOX TRANSCRPTON FACTORS XTCF-3 AND XLEF-1 DURNG EARLY XENOPUS DEVELOPMENT. Miranda Molenaar, Jeroen Roose #, Josi Peterson, Serenella Venanzi*, Hans Clevers #
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Questions in developmental biology. Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration
 Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Computational Biology: Basics & Interesting Problems
 Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
Computational Biology: Basics & Interesting Problems Summary Sources of information Biological concepts: structure & terminology Sequencing Gene finding Protein structure prediction Sources of information
CHAPTER 1 THE STRUCTURAL BIOLOGY OF THE FGF19 SUBFAMILY
 CHAPTER 1 THE STRUCTURAL BIOLOGY OF THE FGF19 SUBFAMILY Andrew Beenken and Moosa Mohammadi* Department of Pharmacology, New York University School of Medicine, New York, New York, USA. *Corresponding Author:
CHAPTER 1 THE STRUCTURAL BIOLOGY OF THE FGF19 SUBFAMILY Andrew Beenken and Moosa Mohammadi* Department of Pharmacology, New York University School of Medicine, New York, New York, USA. *Corresponding Author:
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Pacific Symposium on Biocomputing 5: (2000)
 I N S I L I C O A N A L Y S I S O F G E N E E X P R E S S I O N P A T T E R N S D U R I N G E A R L Y D E V E L O P M E N T O F X E N O P U S L A E V I S N. P O L L E T, H. A. S C H M I D T, V. G A W A
I N S I L I C O A N A L Y S I S O F G E N E E X P R E S S I O N P A T T E R N S D U R I N G E A R L Y D E V E L O P M E N T O F X E N O P U S L A E V I S N. P O L L E T, H. A. S C H M I D T, V. G A W A
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam
 Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Eukaryotic vs. Prokaryotic genes
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 18: Eukaryotic genes http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Eukaryotic vs. Prokaryotic genes Like in prokaryotes,
Chapter 10 Development and Differentiation
 Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Zn 2+ -binding sites in USP18. (a) The two molecules of USP18 present in the asymmetric unit are shown. Chain A is shown in blue, chain B in green. Bound Zn 2+ ions are shown as
Supplementary Figure 1 Zn 2+ -binding sites in USP18. (a) The two molecules of USP18 present in the asymmetric unit are shown. Chain A is shown in blue, chain B in green. Bound Zn 2+ ions are shown as
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Caenorhabditis elegans
 Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Unit 5: Cell Division and Development Guided Reading Questions (45 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 12 The Cell Cycle Unit 5: Cell Division and Development Guided
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 12 The Cell Cycle Unit 5: Cell Division and Development Guided
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
RANK. Alternative names. Discovery. Structure. William J. Boyle* SUMMARY BACKGROUND
 RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Genomics and bioinformatics summary. Finding genes -- computer searches
 Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Genomics and bioinformatics summary 1. Gene finding: computer searches, cdnas, ESTs, 2. Microarrays 3. Use BLAST to find homologous sequences 4. Multiple sequence alignments (MSAs) 5. Trees quantify sequence
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
The Radiata-Bilateria split. Second branching in the evolutionary tree
 The Radiata-Bilateria split Second branching in the evolutionary tree Two very important characteristics are used to distinguish between the second bifurcation of metazoans Body symmetry Germinal layers
The Radiata-Bilateria split Second branching in the evolutionary tree Two very important characteristics are used to distinguish between the second bifurcation of metazoans Body symmetry Germinal layers
Gene Control Mechanisms at Transcription and Translation Levels
 Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
BIOLOGY - CLUTCH CH.32 - OVERVIEW OF ANIMALS.
 !! www.clutchprep.com Animals are multicellular, heterotrophic eukaryotes that feed by ingesting their food Most animals are diploid, and produce gametes produced directly by meiosis Animals lack cell
!! www.clutchprep.com Animals are multicellular, heterotrophic eukaryotes that feed by ingesting their food Most animals are diploid, and produce gametes produced directly by meiosis Animals lack cell
Cell Death & Trophic Factors II. Steven McLoon Department of Neuroscience University of Minnesota
 Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Supplemental data. Pommerrenig et al. (2011). Plant Cell /tpc
 Supplemental Figure 1. Prediction of phloem-specific MTK1 expression in Arabidopsis shoots and roots. The images and the corresponding numbers showing absolute (A) or relative expression levels (B) of
Supplemental Figure 1. Prediction of phloem-specific MTK1 expression in Arabidopsis shoots and roots. The images and the corresponding numbers showing absolute (A) or relative expression levels (B) of
Ch 10, 11 &14 Preview
 Ch 10, 11 &14 Preview Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme hypothesis have to be modified? a. Some
Ch 10, 11 &14 Preview Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme hypothesis have to be modified? a. Some
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development June 23, 2009 Concepts Cell-Cell Communication Cells develop in the context of their environment, including: - their immediate
Biology 4361 - Developmental Biology Cell-Cell Communication in Development June 23, 2009 Concepts Cell-Cell Communication Cells develop in the context of their environment, including: - their immediate
Exam 2 ID#: November 9, 2006
 Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Regulation of Transcription in Eukaryotes. Nelson Saibo
 Regulation of Transcription in Eukaryotes Nelson Saibo saibo@itqb.unl.pt In eukaryotes gene expression is regulated at different levels 1 - Transcription 2 Post-transcriptional modifications 3 RNA transport
Regulation of Transcription in Eukaryotes Nelson Saibo saibo@itqb.unl.pt In eukaryotes gene expression is regulated at different levels 1 - Transcription 2 Post-transcriptional modifications 3 RNA transport
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells
 Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
Molecular Evolution I. Molecular Evolution: pertains to evolution at the levels of DNA, RNA, and proteins. Outline. Mutations 10/25/18
 Molecular Evolution I Molecular Evolution: pertains to evolution at the levels of DNA, RNA, and proteins Carol Eunmi Lee University of Wisconsin Evolution 410 Outline (1) Types of Mutations that could
Molecular Evolution I Molecular Evolution: pertains to evolution at the levels of DNA, RNA, and proteins Carol Eunmi Lee University of Wisconsin Evolution 410 Outline (1) Types of Mutations that could
ASSESSING TRANSLATIONAL EFFICIACY THROUGH POLY(A)- TAIL PROFILING AND IN VIVO RNA SECONDARY STRUCTURE DETERMINATION
 ASSESSING TRANSLATIONAL EFFICIACY THROUGH POLY(A)- TAIL PROFILING AND IN VIVO RNA SECONDARY STRUCTURE DETERMINATION Journal Club, April 15th 2014 Karl Frontzek, Institute of Neuropathology POLY(A)-TAIL
ASSESSING TRANSLATIONAL EFFICIACY THROUGH POLY(A)- TAIL PROFILING AND IN VIVO RNA SECONDARY STRUCTURE DETERMINATION Journal Club, April 15th 2014 Karl Frontzek, Institute of Neuropathology POLY(A)-TAIL
Small RNA in rice genome
 Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Vol. 45 No. 5 SCIENCE IN CHINA (Series C) October 2002 Small RNA in rice genome WANG Kai ( 1, ZHU Xiaopeng ( 2, ZHONG Lan ( 1,3 & CHEN Runsheng ( 1,2 1. Beijing Genomics Institute/Center of Genomics and
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
TRANSCRIPTOMICS. (or the analysis of the transcriptome) Mario Cáceres. Main objectives of genomics. Determine the entire DNA sequence of an organism
 TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
Endoderm Specification and Differentiation in Xenopus Embryos
 Developmental Biology 236, 330 343 (2001) doi:10.1006/dbio.2001.0347, available online at http://www.idealibrary.com on Endoderm Specification and Differentiation in Xenopus Embryos Marko E. Horb 1 and
Developmental Biology 236, 330 343 (2001) doi:10.1006/dbio.2001.0347, available online at http://www.idealibrary.com on Endoderm Specification and Differentiation in Xenopus Embryos Marko E. Horb 1 and
Name. Biology Developmental Biology Winter Quarter 2013 KEY. Midterm 3
 Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
3/8/ Complex adaptations. 2. often a novel trait
 Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Markov Models & DNA Sequence Evolution
 7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
7.91 / 7.36 / BE.490 Lecture #5 Mar. 9, 2004 Markov Models & DNA Sequence Evolution Chris Burge Review of Markov & HMM Models for DNA Markov Models for splice sites Hidden Markov Models - looking under
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Regulation of Transcription in Eukaryotes
 Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
From Gene to Protein
 From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
Supplementary Information for
 Supplementary Information for Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding Sebastian Pechmann & Judith Frydman Department of Biology and BioX, Stanford
Supplementary Information for Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding Sebastian Pechmann & Judith Frydman Department of Biology and BioX, Stanford
Biology 218, practise Exam 2, 2011
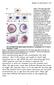 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
An essential role of a FoxD gene in notochord induction in Ciona embryos
 Development 129, 3441-3453 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV5028 3441 An essential role of a FoxD gene in notochord induction in Ciona embryos Kaoru S. Imai*, Nori
Development 129, 3441-3453 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV5028 3441 An essential role of a FoxD gene in notochord induction in Ciona embryos Kaoru S. Imai*, Nori
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Cardiac cell-cell Communication Part 1 Alonso P. Moreno D.Sc. CVRTI, Cardiology
 Bioengineering 6003 Cellular Electrophysiology and Biophysics Cardiac cell-cell Communication Part 1 Alonso P. Moreno D.Sc. CVRTI, Cardiology moreno@cvrti.utah.edu November 2010 poster Physiological Relevance
Bioengineering 6003 Cellular Electrophysiology and Biophysics Cardiac cell-cell Communication Part 1 Alonso P. Moreno D.Sc. CVRTI, Cardiology moreno@cvrti.utah.edu November 2010 poster Physiological Relevance
SUPPLEMENTARY DATA - 1 -
 - 1 - SUPPLEMENTARY DATA Construction of B. subtilis rnpb complementation plasmids For complementation, the B. subtilis rnpb wild-type gene (rnpbwt) under control of its native rnpb promoter and terminator
- 1 - SUPPLEMENTARY DATA Construction of B. subtilis rnpb complementation plasmids For complementation, the B. subtilis rnpb wild-type gene (rnpbwt) under control of its native rnpb promoter and terminator
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny
 Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Phylogenetics and chromosomal synteny of the GATAs 1273 GATA family of transcription factors of vertebrates: phylogenetics and chromosomal synteny CHUNJIANG HE, HANHUA CHENG* and RONGJIA ZHOU* Department
Maternal VegT and b-catenin: Patterning the Xenopus Blastula
 CHAPTER 1 Maternal VegT and b-catenin: Patterning the Xenopus Blastula Matthew Kofron, Jennifer Xanthos, and Janet Heasman 1 1.1 Introduction Loss of the maternal T-box transcription factor VegT has a
CHAPTER 1 Maternal VegT and b-catenin: Patterning the Xenopus Blastula Matthew Kofron, Jennifer Xanthos, and Janet Heasman 1 1.1 Introduction Loss of the maternal T-box transcription factor VegT has a
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p
 Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Organization of Genes Differs in Prokaryotic and Eukaryotic DNA Chapter 10 p.110-114 Arrangement of information in DNA----- requirements for RNA Common arrangement of protein-coding genes in prokaryotes=
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Mesoderm Development
 Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
DOWNLOAD PDF DETERMINATION OF PREPLACODAL ECTODERM AND SENSORY PLACODES SALLY A. MOODY
 Chapter 1 : Olfactory epithelium - Wikipedia The sensory organs of the vertebrate head derive from two embryological structures, the neural crest and the ectodermal placodes. Although quite a lot is known
Chapter 1 : Olfactory epithelium - Wikipedia The sensory organs of the vertebrate head derive from two embryological structures, the neural crest and the ectodermal placodes. Although quite a lot is known
Practical Bioinformatics
 5/2/2017 Dictionaries d i c t i o n a r y = { A : T, T : A, G : C, C : G } d i c t i o n a r y [ G ] d i c t i o n a r y [ N ] = N d i c t i o n a r y. h a s k e y ( C ) Dictionaries g e n e t i c C o
5/2/2017 Dictionaries d i c t i o n a r y = { A : T, T : A, G : C, C : G } d i c t i o n a r y [ G ] d i c t i o n a r y [ N ] = N d i c t i o n a r y. h a s k e y ( C ) Dictionaries g e n e t i c C o
