Supporting Text - Cubic Ternary Complex Activation Model (ctcam)
|
|
|
- Meagan Campbell
- 6 years ago
- Views:
Transcription
1 Both Ligand- and Cell-Specific Parameters Control Ligand Agonism in a Kinetic Model of G protein Coupled Receptor Signaling by Tamara L. Kinzer-Ursem and Jennifer J. Linderman Supporting Text - Cubic Ternary Complex Activation Model (ctcam) The reactions and the accompanying rate constants necessary to describe the ctcam are shown in Fig. S1. Model detail and assumptions are explained in this supporting text. Differential equations describing the ctcam were written according to mass action kinetics and are also outlined below (Eqns. S.19 S.30). G-protein activation/inactivation steps Under physiological conditions, activation of G protein by an active receptor (R*) initiates a multi-step process of GDP dissociation, GTP association, and dissociation of the G protein heterotrimer into Gα and Gβγ subunits. These elementary processes are represented in the ctcam (Fig. 1 and A1) as a single step with active receptor-g protein complexes (R*G and LR*G) activating G protein with the rate constant k Gact in accordance with findings that show the dissociation of GDP from the Gα protein subunit to be the rate limiting step [1,2]. It is assumed that rate constant of G protein activation (k Gact ) is the same or similar for both constitutively active receptor/g-protein complexes (R*G) and ligand-associated active receptor/g protein complexes (LR*G). Gα GTP is considered to be the response signal although it is known that Gβγ also couples to and activates signal transduction proteins [3,4]. The active Gα GTP subunit is deactivated when the nucleotide is hydrolyzed to form Gα GDP with a rate constant k GTP. This rate constant can vary over several 100 fold depending on the intrinsic GTPase activity of the Gα subunit or the presence of RGS proteins in the system [5,6]. Recombination of the Gα GDP and Gβγ subunits to form free G protein occurs with rate constant k G. Assumptions regarding rate constants describing ligand, receptor, and G-protein interactions 1
2 When the system is at equilibrium (i.e. k Gact =0) the principle of microscopic balance dictates that the forward and backward velocities of each elementary step must be equal [7,8]. This principle implies that the free energy change clockwise and counterclockwise around a path must be equal. For example, for the front face of the cube (Fig. S.1) the rate constants must be set so that k 1 k 5 k 8 k 4 =k 3 k 7 k 6 k 2. Due to this thermodynamic constraint, only 19 of the 24 rate constants describing the ligandreceptor-g protein interactions are independent. Therefore, it is assumed that the ratio of the forward and backward rate constants in the model should reduce to the seven equilibrium constants of the ctcm ([9] and Fig. 1). These consist of three cellular dependent parameters (K act, K g, and β) that dictate receptor and G protein interaction in absence of ligand, three ligand dependent parameters (K a, α, and γ), and one parameter (δ) that is both ligand- and cell-specific. These relationships are (S.1) k 1 /k 2 = K act (S.7) k 13 /k 14 = βk act (S.2) k 3 /k 4 = K a (S.8) k 15 /k 16 = γk g (S.3) k 5 /k 6 = αk a (S.9) k 17 /k 18 = βγδkg (S.4) k 7 /k 8 = αk act (S.10) k 19 /k 20 = αβγkact (S.5) k 9 / k 10 = K g (S.11) k 21 /k 22 = αγδk a (S.6) k 11 /k 12 = βk g (S.12) k 23 /k 24 = γk a By specifying these equilibrium parameters (7 in total), then, if the forward rate constant is known, the reverse rate constant can be calculated, leaving 12 parameters to specify. The three forward rate constants k 1, k 3, and k 11 will be specified. Assumptions for the remaining 9 rate constants can be grouped into and are presented below in three categories. The independent ligand- and cell- specific parameters that make up the ctcam are summarized in Table 1. Rate constants describing the reversible association of ligands and receptors It is assumed that G protein binding does not affect the ligand association rate constant such that the rate constants for L+R LR and L+RG LRG are equal (k 3 =k 23 ), and L+R* LR*, L+R*G LR*G are equal (k 5 =k 21 ). In at least one particular system it has been shown that the association rate constants for ligand binding to R and RG are the same [10]. Ligand association with the active receptor state R* is assumed to be influenced by the ligand-specific parameter α such that k 5 = αk 3. Ligand-receptor 2
3 dissociation rate constants are assumed not to be affected by receptor conformation such that the reactions LR R and LR* R* have equal rate constants (k 4 =k 6 ). These assumptions are summarized in the following equations: (S.13) k 3 = k 23 (S.14) k 5 = k 21 = αk 3 (S.15) k 4 = k 6 Rate constants describing the reversible association of receptors and G proteins It is assumed that the inactive receptor conformation is unable to couple with full efficiency to G protein. The collision efficiency (η), an adjustable parameter ranging 0 to 1, is used to implement this assumption [11] such that k 9 =ηk 11. Ligand biases receptor conformation based on the parameter α; however it is assumed here that ligand does not have an effect on the association of either the active or inactive receptor conformation with G protein such that the rate constants for the reactions R RG and LR LRG are equal (k 9 =k 15 ) and R* R*G and LR* LR*G are equal (k 11 =k 17 ). Thus: (S.16) k 9 = k 15 = ηk 11 (S.17) k 11 = k 17 Rate constants for the interconversion of receptor conformations The receptor inactivation is assumed not to be affected by ligand or G protein binding such that the rate constants describing the R* R, LR* LR, R*G RG, and LR*G LRG are equal. Thus: (S.18) k 2 = k 8 = k 14 = k 20 Model equations (S.19) dr[t]/dt = -k 1 R[t] + (k 1 /K act )R*[t] - (ηk 11 )R[t]G[t] + ((ηk 11 )/K g )RG[t] k 3 R[t]L + (k 3 /K a )*LR[t]; (S.20) dlr[t]/dt = -(k 3 /K a )LR[t] + k 3 L*R[t] - αk 1 LR[t] + k 1 /K act LR*[t] ηk 11 LR[t] G[t] + (ηk 11 /γk g )LRG[t] (S.21) drg[t]/dt = ηk 11 R[t]G[t] - (ηk 11 /K g )RG[t] + (k 3 /γk a )LRG[t] - k f RG[t]L[t] - βk 1 RG[t] + (k 1 /K act )RG[t] 3
4 (S.22) dlrg[t]/dt = k 3 RG[t]L - (k 3 /γk a )LRG[t] + ηk 11 LR[t]G[t] - (ηk 11 /γk g ))LRG[t] + (k 1 /K act )LR*G[t] - k 1 αβδlrg[t] (S.23) dr*[t]/dt = k 1 R[t] - (k 1 /K act )R*[t] + (k 3 /K a )LR*[t] - (αk 3 )R*[t]L - βk 11 R*[t] G[t] + (k 11 /K g )R*G[t] + k Gact R*G[t] (S.24) dlr*[t]/dt = αk 1 LR[t] - (k 1 /K act )LR*[t] - (k 3 /K a )LR*[t] + αk 3 R*[t]L - βk 11 LR*[t]G[t] + (k 11 /δγk g )LR*G[t] + k Gact LR*G[t] (S.25) dr*g[t]/dt = βk 11 G[t]R*[t] - (k 11 /K g )R*G[t] + βk 1 RG[t] - (k 1 /K act )R*G[t] - αk 3 R*G[t]L + (k 3 /δγk a )LR*G[t] - k Gact R*G[t] (S.26) dlr*g[t]/dt = αk 3 R*G[t]L - (k 3 /δγk a )LR*G[t] + βk 11 LR*[t]G[t] - (k 11 /δγk g )LR*G[t] + αβδk 1 LRG[t] - (k 1 /K act )LR*G[t] - k Gact LR*G[t] (S.27) dgα GTP [t]/dt = k Gact (R*G[t] + LR*G[t]) - k GTP Gα GTP [t] (S.28) dgα GDP [t]/dt = k GTP Gα GTP [t] - k G Gα GDP [t]gβγ[t] (S.29) dgβγ[t]/dt = k Gact (R*G[t] + LR*G[t]) - k G Gα GDP [t]gβγ[t] (S.30) dg[t]/dt = -ηk 11 R[t]G[t] + (ηk 11 /K g )RG[t] - ηk 11 LR[t]G[t] + (ηk 11 /γk g )LRG[t] - βk 11 LR*[t]G[t] + (k 11 /δγk g )LR*G[t] - βk 11 G[t]R*[t] + (k 11 /K g )R*G[t] + k G Gα GDP [t]gβγ[t] Equations were solved numerically using the NDSolve function in Mathematica (Wolfram Research Inc., Champaign, IL). Total receptor and G protein (R total and G total ) were set as the initial values of R and G while all other species were set to zero. The system was allowed to come to steady state (typically less that 100 seconds) at which point ligand (L) was added (denoted as time = 0) and each species tracked over time. References 1. Remmers AE, Engel C, Liu M, Neubig RR (1999) Interdomain interactions regulate GDP release from heterotrimeric G proteins. Biochemistry 38: Traynor JR, Clark MJ, Remmers AE (2002) Relationship between rate and extent of G protein activation: Comparison between full and partial opioid agonists. J Pharmacol Exp Ther 300: Robishaw JD, Berlot CH (2004) Translating G protein subunit diversity into functional specificity. Curr Opin Cell Biol 16: Cabrera-Vera TM, Vanhauwe J, Thomas TO, Medkova M, Preininger A, et al. (2003) Insights into G protein structure, function, and regulation. Endocr Rev 24:
5 5. Lan KL, Zhong H, Nanamori M, Neubig RR (2000) Rapid kinetics of regulator of G- protein signaling (RGS)-mediated Galphai and Galphao deactivation. Galpha specificity of RGS4 and RGS7. J Biol Chem 275: Zhong H, Wade SM, Woolf PJ, Linderman JJ, Traynor JR, et al. (2002) A spatial focusing model for G protein signals: RGS protein-mediated kinetic scaffolding. J Biol Chem 278: Wyman J (1975) The turning wheel: A study in steady states. Proc Natl Acad Sci U S A 72: Lauffenburger DA, Linderman JJ (1993) Receptors: Models for binding, trafficking, and signaling. New York: Oxford University Press. 365 p. 9. Shea LD, Neubig RR, Linderman JJ (2000) Timing is everything - the role of kinetics in G protein activation. Life Sci 68: Posner RG, Fay SP, Domalewski MD, Sklar LA (1994) Continuous spectrofluorometric analysis of formyl peptide receptor ternary complex interactions. Mol Pharmacol 45: Shea LD, Omann GM, Linderman JJ (1997) Calculation of diffusion-limited kinetics for the reactions in collision coupling and receptor cross-linking. Biophys J 73: Figure Legend Figure S.1. The cubic ternary complex activation model (ctcam). The numbered (k 1 -k 24 ) rate constants describe the forward and reverse reactions for ligand, receptor and G protein interactions, and the lettered rate constants (k Gact, k GTP, and k G ) describe G protein activation and recycling [9]. 5
6 Figure S.1 6
COMPUTATIONAL ANALYSIS OF G-PROTEIN COUPLED RECEPTOR SCREENING, DIMERIZATION, AND DESENSITIZATION. Peter J. Woolf
 COMPUTATIONAL ANALYSIS OF G-PROTEIN COUPLED RECEPTOR SCREENING, DIMERIZATION, AND DESENSITIZATION by Peter J. Woolf A dissertation submitted in partial fulfillment of the requirements for the degree of
COMPUTATIONAL ANALYSIS OF G-PROTEIN COUPLED RECEPTOR SCREENING, DIMERIZATION, AND DESENSITIZATION by Peter J. Woolf A dissertation submitted in partial fulfillment of the requirements for the degree of
Heterotrimeric G proteins and the role of lipids in signaling. John Sondek, Ph.D. Depts. of Pharmacology and Biochemistry & Biophyscis
 Heterotrimeric G proteins and the role of lipids in signaling John Sondek, Ph.D. Depts. of Pharmacology and Biochemistry & Biophyscis The GTPase cycle molecular switch A GTPases is NOT a kinase Two major
Heterotrimeric G proteins and the role of lipids in signaling John Sondek, Ph.D. Depts. of Pharmacology and Biochemistry & Biophyscis The GTPase cycle molecular switch A GTPases is NOT a kinase Two major
arxiv:physics/ v3 [physics.bio-ph] 16 Sep 2003
![arxiv:physics/ v3 [physics.bio-ph] 16 Sep 2003 arxiv:physics/ v3 [physics.bio-ph] 16 Sep 2003](/thumbs/92/110902603.jpg) Accepted for publication in Biophysical Chemistry Special Issue for Walter Kauzmann arxiv:physics/0207049v3 [physics.bio-ph] 16 Sep 2003 Thermodynamic and Kinetic Analysis of Sensitivity Amplification
Accepted for publication in Biophysical Chemistry Special Issue for Walter Kauzmann arxiv:physics/0207049v3 [physics.bio-ph] 16 Sep 2003 Thermodynamic and Kinetic Analysis of Sensitivity Amplification
References on Kinetics and Mechanisms
 References on Kinetics and Mechanisms Excellent reference for all aspects of enzyme kinetics including important elements of Metabolic Control Analysis of relevance to systems analysis of enzyme function
References on Kinetics and Mechanisms Excellent reference for all aspects of enzyme kinetics including important elements of Metabolic Control Analysis of relevance to systems analysis of enzyme function
Signal Transduction Phosphorylation Protein kinases. Misfolding diseases. Protein Engineering Lysozyme variants
 Signal Transduction Phosphorylation Protein kinases Misfolding diseases Protein Engineering Lysozyme variants Cells and Signals Regulation The cell must be able to respond to stimuli Cellular activities
Signal Transduction Phosphorylation Protein kinases Misfolding diseases Protein Engineering Lysozyme variants Cells and Signals Regulation The cell must be able to respond to stimuli Cellular activities
Visual pigments. Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2019
 Visual pigments Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2019 References Webvision: The Organization of the Retina and Visual System (http://www.ncbi.nlm.nih.gov/books/nbk11522/#a 127) The
Visual pigments Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2019 References Webvision: The Organization of the Retina and Visual System (http://www.ncbi.nlm.nih.gov/books/nbk11522/#a 127) The
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Application of the Markov State Model to Molecular Dynamics of Biological Molecules. Speaker: Xun Sang-Ni Supervisor: Prof. Wu Dr.
 Application of the Markov State Model to Molecular Dynamics of Biological Molecules Speaker: Xun Sang-Ni Supervisor: Prof. Wu Dr. Jiang Introduction Conformational changes of proteins : essential part
Application of the Markov State Model to Molecular Dynamics of Biological Molecules Speaker: Xun Sang-Ni Supervisor: Prof. Wu Dr. Jiang Introduction Conformational changes of proteins : essential part
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
R7.3 Receptor Kinetics
 Chapter 7 9/30/04 R7.3 Receptor Kinetics Professional Reference Shelf Just as enzymes are fundamental to life, so is the living cell s ability to receive and process signals from beyond the cell membrane.
Chapter 7 9/30/04 R7.3 Receptor Kinetics Professional Reference Shelf Just as enzymes are fundamental to life, so is the living cell s ability to receive and process signals from beyond the cell membrane.
PARAMETER UNCERTAINTY QUANTIFICATION USING SURROGATE MODELS APPLIED TO A SPATIAL MODEL OF YEAST MATING POLARIZATION
 Ching-Shan Chou Department of Mathematics Ohio State University PARAMETER UNCERTAINTY QUANTIFICATION USING SURROGATE MODELS APPLIED TO A SPATIAL MODEL OF YEAST MATING POLARIZATION In systems biology, we
Ching-Shan Chou Department of Mathematics Ohio State University PARAMETER UNCERTAINTY QUANTIFICATION USING SURROGATE MODELS APPLIED TO A SPATIAL MODEL OF YEAST MATING POLARIZATION In systems biology, we
Physical Models of Allostery: Allosteric Regulation in Capsid Assembly
 Physical Models of Allostery: Allosteric Regulation in Capsid Assembly QCB Journal Club Prof. Sima Setayeshgar JB Holmes Nov. 2, 2017 Mechanisms of Allosteric Regulation From R.A. Laskowski, FEBS Letters,
Physical Models of Allostery: Allosteric Regulation in Capsid Assembly QCB Journal Club Prof. Sima Setayeshgar JB Holmes Nov. 2, 2017 Mechanisms of Allosteric Regulation From R.A. Laskowski, FEBS Letters,
Modeling Yeast Cell Polarization Induced by Pheromone Gradients
 Journal of Statistical Physics ( C 27 ) DOI: 1.17/s1955-7-9285-1 Modeling Yeast Cell Polarization Induced by Pheromone Gradients Tau-Mu Yi, 1 Shanqin Chen, 2 Ching-Shan Chou 2 and Qing Nie 2 Received March
Journal of Statistical Physics ( C 27 ) DOI: 1.17/s1955-7-9285-1 Modeling Yeast Cell Polarization Induced by Pheromone Gradients Tau-Mu Yi, 1 Shanqin Chen, 2 Ching-Shan Chou 2 and Qing Nie 2 Received March
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM
 The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
Computational modeling of G-Protein Coupled Receptors (GPCRs) has recently become
 Homology Modeling and Docking of Melatonin Receptors Andrew Kohlway, UMBC Jeffry D. Madura, Duquesne University 6/18/04 INTRODUCTION Computational modeling of G-Protein Coupled Receptors (GPCRs) has recently
Homology Modeling and Docking of Melatonin Receptors Andrew Kohlway, UMBC Jeffry D. Madura, Duquesne University 6/18/04 INTRODUCTION Computational modeling of G-Protein Coupled Receptors (GPCRs) has recently
Lecture 2: Receptor-ligand binding and cooperativity
 Lecture 2: Receptor-ligand binding and cooperativity Paul C Bressloff (Spring 209) A biochemical receptor is a protein molecule that receives a chemical signal in the form of ligand molecules. The ligands
Lecture 2: Receptor-ligand binding and cooperativity Paul C Bressloff (Spring 209) A biochemical receptor is a protein molecule that receives a chemical signal in the form of ligand molecules. The ligands
Supporting Text Z = 2Γ 2+ + Γ + Γ [1]
![Supporting Text Z = 2Γ 2+ + Γ + Γ [1] Supporting Text Z = 2Γ 2+ + Γ + Γ [1]](/thumbs/91/105230015.jpg) Supporting Text RNA folding experiments are typically carried out in a solution containing a mixture of monovalent and divalent ions, usually MgCl 2 and NaCl or KCl. All three species of ions, Mg, M +
Supporting Text RNA folding experiments are typically carried out in a solution containing a mixture of monovalent and divalent ions, usually MgCl 2 and NaCl or KCl. All three species of ions, Mg, M +
Receptors and Ion Channels
 Receptors and Ion Channels Laurie Kellaway Senior Lecturer Department of Human Biology Laurie@curie.uct.ac.za Tel. +27 +21 4066 271 What are the two types of Neurotransmitter receptors Ionotropic receptors
Receptors and Ion Channels Laurie Kellaway Senior Lecturer Department of Human Biology Laurie@curie.uct.ac.za Tel. +27 +21 4066 271 What are the two types of Neurotransmitter receptors Ionotropic receptors
Analysis of correlated mutations in Ras G-domain
 www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
www.bioinformation.net Volume 13(6) Hypothesis Analysis of correlated mutations in Ras G-domain Ekta Pathak * Bioinformatics Department, MMV, Banaras Hindu University. Ekta Pathak - E-mail: ektavpathak@gmail.com;
Why study protein dynamics?
 Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
Why study protein dynamics? Protein flexibility is crucial for function. One average structure is not enough. Proteins constantly sample configurational space. Transport - binding and moving molecules
SINGLE-MOLECULE PHYSIOLOGY
 SINGLE-MOLECULE PHYSIOLOGY Kazuhiko Kinosita, Jr. Center for Integrative Bioscience, Okazaki National Research Institutes Higashiyama 5-1, Myodaiji, Okazaki 444-8585, Japan Single-Molecule Physiology under
SINGLE-MOLECULE PHYSIOLOGY Kazuhiko Kinosita, Jr. Center for Integrative Bioscience, Okazaki National Research Institutes Higashiyama 5-1, Myodaiji, Okazaki 444-8585, Japan Single-Molecule Physiology under
An Allosteric Theory for Hemoglobin Incorporating Asymmetric States to Test the Putative Molecular Code for Cooperativity
 J. Mol. Biol. (1996) 257, 737 744 COMMUNICATION An Allosteric Theory for Hemoglobin Incorporating Asymmetric States to Test the Putative Molecular Code for Cooperativity Stuart J. Edelstein Département
J. Mol. Biol. (1996) 257, 737 744 COMMUNICATION An Allosteric Theory for Hemoglobin Incorporating Asymmetric States to Test the Putative Molecular Code for Cooperativity Stuart J. Edelstein Département
MOLECULAR DRUG TARGETS
 MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Visual pigments. Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2015
 Visual pigments Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2015 References Photoreceptors and visual pigments Webvision: The Organization of the Retina and Visual System (http://www.ncbi.nlm.nih.gov/books/nbk11522/#a127)
Visual pigments Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2015 References Photoreceptors and visual pigments Webvision: The Organization of the Retina and Visual System (http://www.ncbi.nlm.nih.gov/books/nbk11522/#a127)
six lectures on systems biology
 six lectures on systems biology jeremy gunawardena department of systems biology harvard medical school lecture 3 5 april 2011 part 2 seminar room, department of genetics a rather provisional syllabus
six lectures on systems biology jeremy gunawardena department of systems biology harvard medical school lecture 3 5 april 2011 part 2 seminar room, department of genetics a rather provisional syllabus
Ligand screening system using fusion proteins of G protein coupled receptors with G protein α subunits
 2 Ligand screening system using fusion proteins of G protein coupled receptors with G protein α subunits G protein coupled receptors A key player of signaling transduction. Cell membranes are packed with
2 Ligand screening system using fusion proteins of G protein coupled receptors with G protein α subunits G protein coupled receptors A key player of signaling transduction. Cell membranes are packed with
arxiv: v1 [q-bio.cb] 31 Dec 2015
![arxiv: v1 [q-bio.cb] 31 Dec 2015 arxiv: v1 [q-bio.cb] 31 Dec 2015](/thumbs/77/76028420.jpg) A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network arxiv:52.09233v [q-bio.cb] 3
A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network arxiv:52.09233v [q-bio.cb] 3
Chemical kinetics and catalysis
 Chemical kinetics and catalysis Outline Classification of chemical reactions Definition of chemical kinetics Rate of chemical reaction The law of chemical raction rate Collision theory of reactions, transition
Chemical kinetics and catalysis Outline Classification of chemical reactions Definition of chemical kinetics Rate of chemical reaction The law of chemical raction rate Collision theory of reactions, transition
William D. Singer, Ph.D.
 William D. Singer, Ph.D. billsinger@dcccd.edu EDUCATION AND TRAINING University of Texas Southwestern Medical Center; Dallas, Texas 1991-1995 ~ Postdoctoral Studies, Cellular Signal Transduction University
William D. Singer, Ph.D. billsinger@dcccd.edu EDUCATION AND TRAINING University of Texas Southwestern Medical Center; Dallas, Texas 1991-1995 ~ Postdoctoral Studies, Cellular Signal Transduction University
Replica exchange methodology. John Karanicolas June 2003
 Replica exchange methodology John Karanicolas MMTSB@PSC, June 2003 Outline o Motivation o Theory o Practical considerations o MMTSB Tool Set Why? o Sampling on rugged potential energy surfaces is difficult
Replica exchange methodology John Karanicolas MMTSB@PSC, June 2003 Outline o Motivation o Theory o Practical considerations o MMTSB Tool Set Why? o Sampling on rugged potential energy surfaces is difficult
Advanced Topics in RNA and DNA. DNA Microarrays Aptamers
 Quiz 1 Advanced Topics in RNA and DNA DNA Microarrays Aptamers 2 Quantifying mrna levels to asses protein expression 3 The DNA Microarray Experiment 4 Application of DNA Microarrays 5 Some applications
Quiz 1 Advanced Topics in RNA and DNA DNA Microarrays Aptamers 2 Quantifying mrna levels to asses protein expression 3 The DNA Microarray Experiment 4 Application of DNA Microarrays 5 Some applications
Alliance for Cellular Signaling
 Alliance for Cellular Signaling 40 laboratories investigating basic questions in cell signaling How complex is signal processing in cells? What is the structure and dynamics of the network? Can functional
Alliance for Cellular Signaling 40 laboratories investigating basic questions in cell signaling How complex is signal processing in cells? What is the structure and dynamics of the network? Can functional
Abstract. Author Summary. Yougan Cheng 1, Hans Othmer 1,* 1 School of Mathematics, University of Minnesota, Minneapolis, MN, USA. *
 A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network - Submission to PLOS Computational
A Model for Direction Sensing in Dictyostelium Discoideum: Ras Activity and Symmetry Breaking Driven by a G βγ - Mediated, G α2 -Ric8 Dependent Signal Transduction Network - Submission to PLOS Computational
The EGF Signaling Pathway! Introduction! Introduction! Chem Lecture 10 Signal Transduction & Sensory Systems Part 3. EGF promotes cell growth
 Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 3 Question of the Day: Who is the son of Sevenless? Introduction! Signal transduction involves the changing of a cell s metabolism or gene
Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 3 Question of the Day: Who is the son of Sevenless? Introduction! Signal transduction involves the changing of a cell s metabolism or gene
Identification of Functional Amino Acids in the G Protein Alpha-Subunit
 The University of Maine DigitalCommons@UMaine University of Maine Office of Research and Sponsored Programs: Grant Reports Special Collections 7-31-2001 Identification of Functional Amino Acids in the
The University of Maine DigitalCommons@UMaine University of Maine Office of Research and Sponsored Programs: Grant Reports Special Collections 7-31-2001 Identification of Functional Amino Acids in the
An allosteric model (chemoreception/gustation/membrane receptors/membrane transition)
 Proc. Nati. Acad. Sci. USA Vol. 77, No. 3, pp. 1686-1690, NMarch 1980 Neurobiology Sodium and potassium salt stimulation of taste receptor cells: An allosteric model (chemoreception/gustation/membrane
Proc. Nati. Acad. Sci. USA Vol. 77, No. 3, pp. 1686-1690, NMarch 1980 Neurobiology Sodium and potassium salt stimulation of taste receptor cells: An allosteric model (chemoreception/gustation/membrane
26 Robustness in a model for calcium signal transduction dynamics
 26 Robustness in a model for calcium signal transduction dynamics U. Kummer 1,G.Baier 2 and L.F. Olsen 3 1 European Media Laboratory, Villa Bosch, Schloss-Wolfsbrunnenweg 33, 69118 Heidelberg, Germany
26 Robustness in a model for calcium signal transduction dynamics U. Kummer 1,G.Baier 2 and L.F. Olsen 3 1 European Media Laboratory, Villa Bosch, Schloss-Wolfsbrunnenweg 33, 69118 Heidelberg, Germany
Lecture 27. Transition States and Enzyme Catalysis
 Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
A NOVEL ROLE FOR THE RGS PROTEIN SST2 IN YEAST SIGNALING REVEALED BY MATHEMATICAL MODELING AND LIVE CELL IMAGING. Sai Phanindra Venkatapurapu
 A NOVEL ROLE FOR THE RGS PROTEIN SST2 IN YEAST SIGNALING REVEALED BY MATHEMATICAL MODELING AND LIVE CELL IMAGING Sai Phanindra Venkatapurapu A dissertation submitted to the faculty of the University of
A NOVEL ROLE FOR THE RGS PROTEIN SST2 IN YEAST SIGNALING REVEALED BY MATHEMATICAL MODELING AND LIVE CELL IMAGING Sai Phanindra Venkatapurapu A dissertation submitted to the faculty of the University of
Chem Lecture 10 Signal Transduction
 Chem 452 - Lecture 10 Signal Transduction 111202 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Chem 452 - Lecture 10 Signal Transduction 111202 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
SUPPLEMENTARY INFORMATION
 Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
Figure S1. Secondary structure of CAP (in the camp 2 -bound state) 10. α-helices are shown as cylinders and β- strands as arrows. Labeling of secondary structure is indicated. CDB, DBD and the hinge are
BINDING AND THE CONFORMATIONAL CHANGE OF HEMOGLOBIN
 THE RELATION BETWEEN CARBON MONOXIDE BINDING AND THE CONFORMATIONAL CHANGE OF HEMOGLOBIN CHARLES A. SAWICKI AND QUENTIN H. GIBSON, Section of Biochemistry, Molecular and Cell Biology, Cornell University,
THE RELATION BETWEEN CARBON MONOXIDE BINDING AND THE CONFORMATIONAL CHANGE OF HEMOGLOBIN CHARLES A. SAWICKI AND QUENTIN H. GIBSON, Section of Biochemistry, Molecular and Cell Biology, Cornell University,
Reception The target cell s detection of a signal coming from outside the cell May Occur by: Direct connect Through signal molecules
 Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Lecture 3 Regulation of Initiation: Met-tRNA-binding
 Institut für Biochemie und Molekulare Medizin Lecture 3 Regulation of Initiation: Met-tRNA-binding Michael Altmann FS 2010 Model of initiation eif4g eif4e AAA AAA PABP cap AAA AUG mrna eif4a eif4b ATP
Institut für Biochemie und Molekulare Medizin Lecture 3 Regulation of Initiation: Met-tRNA-binding Michael Altmann FS 2010 Model of initiation eif4g eif4e AAA AAA PABP cap AAA AUG mrna eif4a eif4b ATP
Lecture 7: Simple genetic circuits I
 Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer
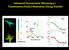 Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Switch Gears to Enzymes
 Switch Gears to Enzymes Their energetic profiles What we measure and the information content (macro vs microscopic constants Structural contributions to catalysis References Excellent reference for all
Switch Gears to Enzymes Their energetic profiles What we measure and the information content (macro vs microscopic constants Structural contributions to catalysis References Excellent reference for all
Resonance Assignment of the RGS Domain of Human RGS10
 Resonance Assignment of the RGS Domain of Human RGS10 Oleg Y. Fedorov 2, Victoria A. Higman 1, Peter Schmieder 1, Martina Leidert 1, Annette Diehl 1, Jonathan Elkins 2, Meera Soundararajan 2, Hartmut Oschkinat
Resonance Assignment of the RGS Domain of Human RGS10 Oleg Y. Fedorov 2, Victoria A. Higman 1, Peter Schmieder 1, Martina Leidert 1, Annette Diehl 1, Jonathan Elkins 2, Meera Soundararajan 2, Hartmut Oschkinat
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
5A Effects of receptor clustering and rebinding on intracellular signaling
 5A Effects of receptor clustering and rebinding on intracellular signaling In the analysis of bacterial chemotaxis, we showed how receptor clustering can amplify a biochemical signal through cooperative
5A Effects of receptor clustering and rebinding on intracellular signaling In the analysis of bacterial chemotaxis, we showed how receptor clustering can amplify a biochemical signal through cooperative
Many proteins spontaneously refold into native form in vitro with high fidelity and high speed.
 Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Medicinal Chemistry/ CHEM 458/658 Chapter 8- Receptors and Messengers
 Medicinal Chemistry/ CHEM 458/658 Chapter 8- Receptors and Messengers Bela Torok Department of Chemistry University of Massachusetts Boston Boston, MA 1 Introduction Receptor specific areas of proteins
Medicinal Chemistry/ CHEM 458/658 Chapter 8- Receptors and Messengers Bela Torok Department of Chemistry University of Massachusetts Boston Boston, MA 1 Introduction Receptor specific areas of proteins
The circle and the basics of signal transduction. Course Outline. Topic #! Topic lecture! Silverthorn! Membranes (pre-requisite material)" "
 Homeostasis 03 The goal of this lecture is to discuss the concept of homeostasis and to introduce basic signal transduction mechanisms involved in homeostatic regulation The sections for this lecture are:
Homeostasis 03 The goal of this lecture is to discuss the concept of homeostasis and to introduce basic signal transduction mechanisms involved in homeostatic regulation The sections for this lecture are:
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
GTPASE-ACTIVATING PROTEINS FOR HETEROTRIMERIC GPROTEINS: Regulators of G Protein Signaling (RGS) and RGS-Like Proteins
 Annu. Rev. Biochem. 2000. 69:795 827 Copyright c 2000 by Annual Reviews. All rights reserved GTPASE-ACTIVATING PROTEINS FOR HETEROTRIMERIC GPROTEINS: Regulators of G Protein Signaling (RGS) and RGS-Like
Annu. Rev. Biochem. 2000. 69:795 827 Copyright c 2000 by Annual Reviews. All rights reserved GTPASE-ACTIVATING PROTEINS FOR HETEROTRIMERIC GPROTEINS: Regulators of G Protein Signaling (RGS) and RGS-Like
Acetylcholine Receptor Channels Activated by a Single Agonist Molecule
 1840 Biophysical Journal Volume 98 May 2010 1840 1846 Acetylcholine Receptor Channels Activated by a Single Agonist Molecule Archana Jha and Anthony Auerbach Department of Physiology and Biophysics, State
1840 Biophysical Journal Volume 98 May 2010 1840 1846 Acetylcholine Receptor Channels Activated by a Single Agonist Molecule Archana Jha and Anthony Auerbach Department of Physiology and Biophysics, State
1. Supplemental figures and legends
 1. Supplemental figures and legends Figure S1. Phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands. AD, phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands at 72h. A, sea urchin
1. Supplemental figures and legends Figure S1. Phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands. AD, phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands at 72h. A, sea urchin
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
BCH 4054 Spring 2001 Chapter 33 Lecture Notes
 BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Invariant Manifold Reductions for Markovian Ion Channel Dynamics
 Invariant Manifold Reductions for Markovian Ion Channel Dynamics James P. Keener Department of Mathematics, University of Utah, Salt Lake City, UT 84112 June 4, 2008 Abstract We show that many Markov models
Invariant Manifold Reductions for Markovian Ion Channel Dynamics James P. Keener Department of Mathematics, University of Utah, Salt Lake City, UT 84112 June 4, 2008 Abstract We show that many Markov models
Signaling Proteins: Mechanical Force Generation by G-proteins G
 Signaling Proteins: Mechanical Force Generation by G-proteins G Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Collaborators: Robijn Bruinsma (Leiden & UCLA) Paul O Lague (UCLA)
Signaling Proteins: Mechanical Force Generation by G-proteins G Ioan Kosztin Beckman Institute University of Illinois at Urbana-Champaign Collaborators: Robijn Bruinsma (Leiden & UCLA) Paul O Lague (UCLA)
Protein synthesis II Biochemistry 302. Bob Kelm February 25, 2004
 Protein synthesis II Biochemistry 302 Bob Kelm February 25, 2004 Two idealized views of the 70S ribosomal complex during translation 70S cavity Fig. 27.25 50S tunnel View with 30S subunit in front, 50S
Protein synthesis II Biochemistry 302 Bob Kelm February 25, 2004 Two idealized views of the 70S ribosomal complex during translation 70S cavity Fig. 27.25 50S tunnel View with 30S subunit in front, 50S
Bacterial Chemotaxis
 Bacterial Chemotaxis Bacteria can be attracted/repelled by chemicals Mechanism? Chemoreceptors in bacteria. attractant Adler, 1969 Science READ! This is sensing, not metabolism Based on genetic approach!!!
Bacterial Chemotaxis Bacteria can be attracted/repelled by chemicals Mechanism? Chemoreceptors in bacteria. attractant Adler, 1969 Science READ! This is sensing, not metabolism Based on genetic approach!!!
Protein synthesis I Biochemistry 302. Bob Kelm February 23, 2004
 Protein synthesis I Biochemistry 302 Bob Kelm February 23, 2004 Key features of protein synthesis Energy glutton Essential metabolic activity of the cell. Consumes 90% of the chemical energy (ATP,GTP).
Protein synthesis I Biochemistry 302 Bob Kelm February 23, 2004 Key features of protein synthesis Energy glutton Essential metabolic activity of the cell. Consumes 90% of the chemical energy (ATP,GTP).
Inferring the in vivo looping properties of DNA
 1 Inferring the in vivo looping properties of DNA Leonor Saiz, J. Miguel Rubi *, and Jose M. G. Vilar Integrative Biological Modeling Laboratory, Computational Biology Program, Memorial Sloan-Kettering
1 Inferring the in vivo looping properties of DNA Leonor Saiz, J. Miguel Rubi *, and Jose M. G. Vilar Integrative Biological Modeling Laboratory, Computational Biology Program, Memorial Sloan-Kettering
Evolution of functionality in lattice proteins
 Evolution of functionality in lattice proteins Paul D. Williams,* David D. Pollock, and Richard A. Goldstein* *Department of Chemistry, University of Michigan, Ann Arbor, MI, USA Department of Biological
Evolution of functionality in lattice proteins Paul D. Williams,* David D. Pollock, and Richard A. Goldstein* *Department of Chemistry, University of Michigan, Ann Arbor, MI, USA Department of Biological
Analysis of Ligand Bias in Functional Studies Involving the Allosteric Modulation of G Protein- Coupled Receptors
 Chapman University Chapman University Digital Commons Biology, Chemistry, and Environmental Sciences Faculty Articles and Research Biology, Chemistry, and Environmental Sciences 5-2014 Analysis of Ligand
Chapman University Chapman University Digital Commons Biology, Chemistry, and Environmental Sciences Faculty Articles and Research Biology, Chemistry, and Environmental Sciences 5-2014 Analysis of Ligand
Chemical Reactions and the enzimes
 Chemical Reactions and the enzimes LESSON N. 6 - PSYCHOBIOLOGY Chemical reactions consist of interatomic interactions that take place at the level of their orbital, and therefore different from nuclear
Chemical Reactions and the enzimes LESSON N. 6 - PSYCHOBIOLOGY Chemical reactions consist of interatomic interactions that take place at the level of their orbital, and therefore different from nuclear
Biol2174 Cell Physiology in Health & Disease
 Biol2174 Cell Physiology in Health & Disease Lecture 4: Membrane Transport Proteins Kiaran Kirk Research School of Biology Learning objectives To understand: The need for membrane transport proteins in
Biol2174 Cell Physiology in Health & Disease Lecture 4: Membrane Transport Proteins Kiaran Kirk Research School of Biology Learning objectives To understand: The need for membrane transport proteins in
a systems approach to biology
 a systems approach to biology jeremy gunawardena department of systems biology harvard medical school lecture 8 27 september 2011 4. metabolism, continued recap warning: a single number, like the CI or
a systems approach to biology jeremy gunawardena department of systems biology harvard medical school lecture 8 27 september 2011 4. metabolism, continued recap warning: a single number, like the CI or
Lecture 19 (10/30/17) Enzyme Regulation
 Reading: Ch5; 164, 166-169 Problems: none Remember Today at 6:30 in PHO-206 is the first MB lecture & quiz NEXT Reading: Ch5; 158-169, 162-166, 169-174 Lecture 19 (10/30/17) Problems: Ch5 (text); 3,7,8,10
Reading: Ch5; 164, 166-169 Problems: none Remember Today at 6:30 in PHO-206 is the first MB lecture & quiz NEXT Reading: Ch5; 158-169, 162-166, 169-174 Lecture 19 (10/30/17) Problems: Ch5 (text); 3,7,8,10
Epibatidine Binds with Unique Site and State Selectivity to Muscle Nicotinic Acetylcholine Receptors*
 THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 14, Issue of April 3, pp. 7843 7849, 1998 1998 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Epibatidine Binds
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 273, No. 14, Issue of April 3, pp. 7843 7849, 1998 1998 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. Epibatidine Binds
Principal Component Analysis (PCA)
 Principal Component Analysis (PCA) Projection onto 2-dim subspace: v 2 v 1 Trajectory x(t) in 3N-dim configuration space A 100-dimensional data set viewed at random orientation Example: Conformational
Principal Component Analysis (PCA) Projection onto 2-dim subspace: v 2 v 1 Trajectory x(t) in 3N-dim configuration space A 100-dimensional data set viewed at random orientation Example: Conformational
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
BME Engineering Molecular Cell Biology. Basics of the Diffusion Theory. The Cytoskeleton (I)
 BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
BME 42-620 Engineering Molecular Cell Biology Lecture 07: Basics of the Diffusion Theory The Cytoskeleton (I) BME42-620 Lecture 07, September 20, 2011 1 Outline Diffusion: microscopic theory Diffusion:
The Photon Counting Histogram (PCH)
 The Photon Counting Histogram (PCH) While Fluorescence Correlation Spectroscopy (FCS) can measure the diffusion coefficient and concentration of fluorophores, a complementary method, a photon counting
The Photon Counting Histogram (PCH) While Fluorescence Correlation Spectroscopy (FCS) can measure the diffusion coefficient and concentration of fluorophores, a complementary method, a photon counting
Oscillations and waves in the concentration of free intracellular
 A dynamic model of the type-2 inositol trisphosphate receptor James Sneyd* and Jean-François Dufour *Institute of Information and Mathematical Sciences Massey University Albany Campus Private Bag 102-904
A dynamic model of the type-2 inositol trisphosphate receptor James Sneyd* and Jean-François Dufour *Institute of Information and Mathematical Sciences Massey University Albany Campus Private Bag 102-904
Topic 4: Equilibrium binding and chemical kinetics
 Topic 4: Equilibrium binding and chemical kinetics Outline: Applications, applications, applications use Boltzmann to look at receptor-ligand binding use Boltzmann to look at PolII-DNA binding and gene
Topic 4: Equilibrium binding and chemical kinetics Outline: Applications, applications, applications use Boltzmann to look at receptor-ligand binding use Boltzmann to look at PolII-DNA binding and gene
Molecular Mechanism for Conformational Dynamics of Ras GTP Elucidated from In-Situ Structural Transition in Crystal
 Molecular Mechanism for Conformational Dynamics of Ras GTP Elucidated from In-Situ Structural Transition in Crystal Shigeyuki Matsumoto, Nao Miyano, Seiki Baba, Jingling Liao, Takashi Kawamura, Chiemi
Molecular Mechanism for Conformational Dynamics of Ras GTP Elucidated from In-Situ Structural Transition in Crystal Shigeyuki Matsumoto, Nao Miyano, Seiki Baba, Jingling Liao, Takashi Kawamura, Chiemi
Supporting Information
 Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins
 13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
Irreversible Inhibition Kinetics
 1 Irreversible Inhibition Kinetics Automation and Simulation Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Automate the determination of biochemical parameters 2. PK/PD simulations with multiple injections Irreversible
1 Irreversible Inhibition Kinetics Automation and Simulation Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Automate the determination of biochemical parameters 2. PK/PD simulations with multiple injections Irreversible
Energetics and Thermodynamics
 DNA/Protein structure function analysis and prediction Protein Folding and energetics: Introduction to folding Folding and flexibility (Ch. 6) Energetics and Thermodynamics 1 Active protein conformation
DNA/Protein structure function analysis and prediction Protein Folding and energetics: Introduction to folding Folding and flexibility (Ch. 6) Energetics and Thermodynamics 1 Active protein conformation
Chemistry 123: Physical and Organic Chemistry Topic 4: Gaseous Equilibrium
 Topic 4: Introduction, Topic 4: Gaseous Equilibrium Text: Chapter 6 & 15 4.0 Brief review of Kinetic theory of gasses (Chapter 6) 4.1 Concept of dynamic equilibrium 4.2 General form & properties of equilbrium
Topic 4: Introduction, Topic 4: Gaseous Equilibrium Text: Chapter 6 & 15 4.0 Brief review of Kinetic theory of gasses (Chapter 6) 4.1 Concept of dynamic equilibrium 4.2 General form & properties of equilbrium
Activation Free Energies in Partial Steps of the. Sarcoplasmic Reticulum Ca-ATPase Cycle during. the Transient States of the Reaction
 Advanced Studies in Biology, Vol. 6, 2014, no. 1, 35-46 HIKARI Ltd, www.m-hikari.com http://dx.doi.org/10.12988/asb.2014.4414 Activation Free Energies in Partial Steps of the Sarcoplasmic Reticulum Ca-ATPase
Advanced Studies in Biology, Vol. 6, 2014, no. 1, 35-46 HIKARI Ltd, www.m-hikari.com http://dx.doi.org/10.12988/asb.2014.4414 Activation Free Energies in Partial Steps of the Sarcoplasmic Reticulum Ca-ATPase
PHARMACOLOGY G PROTEIN COUPLED RECEPTORS
 PHARMACOLOGY G PROTEIN COUPLED RECEPTORS Edited by Richard R. Neubig Department of of Michigan Medical School Ann Arbor, Michigan, USA Serial Editor S. J. Enna Department of Molecular and Integrative Physiology
PHARMACOLOGY G PROTEIN COUPLED RECEPTORS Edited by Richard R. Neubig Department of of Michigan Medical School Ann Arbor, Michigan, USA Serial Editor S. J. Enna Department of Molecular and Integrative Physiology
COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION
 COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION I. Aksan 1, M. Sen 2, M. K. Araz 3, and M. L. Kurnaz 3 1 School of Biological Sciences, University of Manchester,
COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION I. Aksan 1, M. Sen 2, M. K. Araz 3, and M. L. Kurnaz 3 1 School of Biological Sciences, University of Manchester,
5. Kinetics of Allosteric Enzymes. Sigmoidal Kinetics. Cooperativity Binding Constant
 5. Kinetics of Allosteric Enzymes Sigmoidal Kinetics Cooperativity Binding Constant Kinetics of Allosteric Enzymes Contents Definitions Allosteric enzymes Cooperativity Homoallostery Heteroallostery Biphasic
5. Kinetics of Allosteric Enzymes Sigmoidal Kinetics Cooperativity Binding Constant Kinetics of Allosteric Enzymes Contents Definitions Allosteric enzymes Cooperativity Homoallostery Heteroallostery Biphasic
Macromolecular Crowding
 Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp
![Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp](/thumbs/74/70848503.jpg) Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp. 216-225 Updated on: 2/4/07 at 9:00 pm Key Concepts Kinetics is the study of reaction rates. Study of enzyme kinetics
Lecture 11: Enzymes: Kinetics [PDF] Reading: Berg, Tymoczko & Stryer, Chapter 8, pp. 216-225 Updated on: 2/4/07 at 9:00 pm Key Concepts Kinetics is the study of reaction rates. Study of enzyme kinetics
Information Content in Genetics:
 Information Content in Genetics: DNA, RNA and protein mrna translation into protein (protein synthesis) Francis Crick, 1958 [Crick, F. H. C. in Symp. Soc. Exp. Biol., The Biological Replication of Macromolecules,
Information Content in Genetics: DNA, RNA and protein mrna translation into protein (protein synthesis) Francis Crick, 1958 [Crick, F. H. C. in Symp. Soc. Exp. Biol., The Biological Replication of Macromolecules,
arxiv:cond-mat/ v1 [cond-mat.soft] 19 Mar 2001
![arxiv:cond-mat/ v1 [cond-mat.soft] 19 Mar 2001 arxiv:cond-mat/ v1 [cond-mat.soft] 19 Mar 2001](/thumbs/88/114574654.jpg) Modeling two-state cooperativity in protein folding Ke Fan, Jun Wang, and Wei Wang arxiv:cond-mat/0103385v1 [cond-mat.soft] 19 Mar 2001 National Laboratory of Solid State Microstructure and Department
Modeling two-state cooperativity in protein folding Ke Fan, Jun Wang, and Wei Wang arxiv:cond-mat/0103385v1 [cond-mat.soft] 19 Mar 2001 National Laboratory of Solid State Microstructure and Department
KINETICS PROBLEMS. (Alberty R A, Physical Chemistry, 7 th ed., Wiley, NY, 1987) (Answers on the last page)
 KINETICS PROBLEMS (Alberty R A, Physical Chemistry, 7 th ed., Wiley, NY, 1987) (Answers on the last page) 20.1 The half-life of a first-order chemical reaction A --> B is 10 min. What percent of A remains
KINETICS PROBLEMS (Alberty R A, Physical Chemistry, 7 th ed., Wiley, NY, 1987) (Answers on the last page) 20.1 The half-life of a first-order chemical reaction A --> B is 10 min. What percent of A remains
MATHEMATICAL MODELING OF SPECIES-SPECIFIC DIACYLGLYCEROL DYNAMICS IN THE RAW MACROPHAGE FOLLOWING P2Y 6
 MATHEMATICAL MODELING OF SPECIES-SPECIFIC DIACYLGLYCEROL DYNAMICS IN THE RAW 264.7 MACROPHAGE FOLLOWING P2Y 6 RECEPTOR ACTIVATION BY URIDINE 5 -DIPHOSPHATE. By Hannah L. Callender Dissertation Submitted
MATHEMATICAL MODELING OF SPECIES-SPECIFIC DIACYLGLYCEROL DYNAMICS IN THE RAW 264.7 MACROPHAGE FOLLOWING P2Y 6 RECEPTOR ACTIVATION BY URIDINE 5 -DIPHOSPHATE. By Hannah L. Callender Dissertation Submitted
2013 W. H. Freeman and Company. 6 Enzymes
 2013 W. H. Freeman and Company 6 Enzymes CHAPTER 6 Enzymes Key topics about enzyme function: Physiological significance of enzymes Origin of catalytic power of enzymes Chemical mechanisms of catalysis
2013 W. H. Freeman and Company 6 Enzymes CHAPTER 6 Enzymes Key topics about enzyme function: Physiological significance of enzymes Origin of catalytic power of enzymes Chemical mechanisms of catalysis
Enzyme Enzymes are proteins that act as biological catalysts. Enzymes accelerate, or catalyze, chemical reactions. The molecules at the beginning of
 Enzyme Enzyme Enzymes are proteins that act as biological catalysts. Enzymes accelerate, or catalyze, chemical reactions. The molecules at the beginning of the process are called substrates and the enzyme
Enzyme Enzyme Enzymes are proteins that act as biological catalysts. Enzymes accelerate, or catalyze, chemical reactions. The molecules at the beginning of the process are called substrates and the enzyme
Supplementary information
 Supplementary information Figure S1 Mass spectrum of a 3 µm solution of Hsp90 2 incubated with FKBP51 and FKBP52 at 6 µm concentrations. This spectrum illustrates the mass spectral assignment protocol
Supplementary information Figure S1 Mass spectrum of a 3 µm solution of Hsp90 2 incubated with FKBP51 and FKBP52 at 6 µm concentrations. This spectrum illustrates the mass spectral assignment protocol
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
