5A Effects of receptor clustering and rebinding on intracellular signaling
|
|
|
- Kristopher Leonard
- 5 years ago
- Views:
Transcription
1 5A Effects of receptor clustering and rebinding on intracellular signaling In the analysis of bacterial chemotaxis, we showed how receptor clustering can amplify a biochemical signal through cooperative effects (see Sect ). Here we explore another consequence of membrane protein clustering, namely, it can increase the likelihood of rapid rebinding of a ligand. In mammalian cells, clustering of receptors may by facilitated by lipid rafts, which are membrane microdomains rich in cholesterol and having diameters in the range nm [7, 9]. One suggested role of lipid rafts is that they serve as mediators for several growth factors by localizing clusters of their corresponding receptors. (Growth factors acts as triggers for many cellular processes by binding to the extracellular domain of the receptors.) For example, in vitro experiments have shown that disruption of lipid rafts significantly effects the dissociation of fibroblast growth factor-2 (FGF-2) from heparin sulfate proteoglycans (HSPG), which is a co-receptor [5, 2]. Stochastic model of ligand rebinding We begin by describing a self-consistent stochastic mean-field model of ligand rebinding due to Gopalakrishnan et al. [3]. Consider a homogeneous distribution of receptors on a two-dimensional planar surface with mean surface density R 0 per unit area. Let R(t) denote the density of receptors bound to a ligand at time t, with dr dt = k R(t) + k + ρ(t)[r 0 R(t)], (5A.1) where k ± are the intrinsic association and dissociation rates, and ρ(t) is the ligand density in a neighborhood of the surface (a boundary layer of width that is comparable to the size of a ligand). Suppose that the initial density of ligand bound receptors is R(0) = R R 0 and the initial density of ligand in the bulk is zero. It follows that the only contribution to ρ(t) 0 when t > 0 is from ligands released from bound receptors at earlier times 0 < τ < t. Let G(r,,t) denote the probability density (per unit volume) that a ligand, which started in the boundary layer at time t = 0, is in contact with the surface at time t having shifted by r R 2 in the plane, with possible multiple visits to the semi-absorbing surface at intermediate times but without binding to a receptor. Since ρ(t) is independent of position in the plane, we have ] ρ(t) = k R(τ) G(r,,t τ)dr dτ. (5A.2) 0 R2 with k dτ the probability that a ligand is released from a bound receptor in the interval [τ,τ + dτ]. The next step is to calculate the Green s function G(r,,t). Let q(r,,t) denote the probability density (per unit volume) that a ligand released at time t = 0 first 2
2 returns to the surface at time t after shifting by r R 2 in the plane. The Green s function satisfies the integral equation ] dτ G(r,,t) = q(r,,t) + γ q(r,,τ)g(r r,,t τ)dr 0 R 2 δ. (5A.3) Here δ is the time interval for which a ligand resides in a volume element 3, before diffusing way into the bulk. The first term on the right-hand side is the probability that a ligand arrives back at the surface for the first time at t, whereas iterating the second term yields all contributions from multiple visits to the surcae (with binding to a receptor). The factor γ gives the probability of non-binding upon contact with the surface and takes the form γ = 1 k + R 0 δ/. (If R were comparable to R 0 then we would have to replace R 0 by the density of unbound receptors.) It turns out that one does not have to calculate q explicitly [4]. Let G 0 (r,,r) denote the Green s function in the case γ = 1, for which the planar surface is purely reflecting, and ] dτ G 0 (r,,t) = q(r,,t) + q(r,,τ)g 0 (r r,,t τ)dr 0 R 2 δ. (5A.4) We now observe that G 0 is the fundamental solution of the 3D diffusion equation in the domain (r,z),r R 2,z > 0 with a reflecting boundary at z = 0, which is wellknown. We can then express G in terms of G 0 by Fourier transforming equations (5A.3) and (5A.4) with respect to r and Laplace transforming with respect to t: Ĝ(k,,s) = q(k,,s) + γλ δ Ĝ(k,,s) q(k,,s) and Ĝ 0 (k,,s) = q(k,,s) + δ Ĝ0(k,,s) q(k,,s). It follows that Ĝ 0 (k,,s) Ĝ(k,,s) = 1 + ( /δ)ĝ 0 (k,,s) which can be rearranged to yield Ĝ(k,,s) = [1 + γ δ Ĝ(k,,s) ], Ĝ 0 (k,,s) 1 + k + R 0 Ĝ 0 (k,,s), (5A.5) where we have used (1 γ)( /δ) = k + R 0. Next, let p(t) = R(t)/R 0 denote the fraction of bound receptors. Laplace transforming equations (5A.1) and (5A.2), we find that and (k + s) p(s) p(0) = k + ρ(s) 3
3 Hence p(s) = ρ(s) = k R 0 p(s)ĝ(0,,s). p(0) s + k (1 Σ(s)), Σ(s) = k +R 0 Ĝ(0,,s). (5A.6) It can be shown that Ĝ 0 (0,,s) = 1/ Ds so that Σ(s) = k + R 0 Ds + k+ R 0. (5A.7) and, consequently, p(0) p(s) =. s + k Ds Ds + k+ R 0 (5A.8) Equation (5A.8) may be used to extract the short-time and long-time behavior of the bound receptor fraction p(t). At short times (large s), we have p(s) p(0)/(s+k ), which corresponds to exponential decay at the intrinsic rate k, that is, rebinding has no effect. On the other hand, in the late time regime (small s), we have p(s) p(0)/[s + k Ds/(k+ R 0 ]) which exhibits non-exponential behavior in time. More explicitly where c = D(k /k + R 0 ) 2. p(t) p(0)e k t, for t D (k + R 0 ) 2, (5A.9a) p(t) p(0)e ct erfc( ct), for t D (k + R 0 ) 2, (5A.9b) Extension to receptor clusters Gopalakrishnan et al. [3] used the above mean-field formalism to study the effects of receptor clustering on ligand rebinding. Suppose that we focus at a point on the surface within a cluster, which has increased receptor density R 1 compared to the background. The density of free ligand close to the surface at the given point is now given by ] ρ(t) = k p(τ) R 0( r )G(r,,t τ)dr dτ. (5A.10) 0 R 2 Here R 0 (r) = R 1 + (R 0 R 1 )Θ(r ξ ). That is, if the distance between the initial and final positions of a ligand in the plane is smaller than the size of a cluster ξ, then the receptor density is R 1, whereas if the ligand travels lateral distance greater than ξ then the receptor density is R 0, R 0 < R 1. 4
4 For the sake of illustration, suppose that the receptor distribution consists of dense isolated clusters so that R 0 0 and R 1 is large. Then ρ(t) k R 1 0 [ p(τ) G ] 0 (0,,t τ) 1 e ξ 2 /4D(t τ) dτ, (5A.11) where G(0,,t) is the inverse Laplace transform of Ĝ(0,,s) with R 0 R 1 : Ĝ(0,,t) = 1 πdt k +R 1 D e(k +R 1 ) 2 t/d erfc(k + R 1 t/d). (5A.12) The factor 1 e ξ 2 /4D(t τ) arises from integrating the 3D Green s function over a disc of radius ξ in the plane. The Laplace transform of the bound receptor fraction still has the form (5A.6) with [ ] Σ(s) = k + R 1 e st Ĝ(0,,t) 1 e ξ 2 /4Dt dt. (5A.13) 0 Performing an asymptotic analysis of p(s) in the limit s 0 shows that at large times t ξ 2 /D and densely packed clusters (large R 1 ) [3] p(t) p(0)e k ξ 0 t/2ξ, ξ ξ 0 2D k + R 1. (5A.14) Thus, in the case of dense isolated clusters of receptors, the effective dissociation rate of ligands at sufficiently long time-scales is reduced by a factor that is inversely proportional to the size of the cluster. Moreover, ξ 0 sets the critical cluster size below which clustering has negligible influence. It is interesting to relate the above result to a very different approach to determining an effective dissociation, which is rate based on the theory of Berg and Purcell [1], see Sect Recall that for a spherical cell of radius a, with N receptors on its surface and intrinsic ligand binding rate k +, the net ligand flux (in the absence of dissociation) is J = k + ρ 0, where ρ 0 is the far-field concentration of ligand and k+ = 4πaDNk + Nk + + 4πaD. (5A.15) Comparison with the diffusion-limited result in the limit k +, suggests that the effective absorption probability of a ligand in contact with the cell surface is γ = Nk + Nk + + 4πaD. (5A.16) It follows that the corresponding probability of non-absorption is 1 γ, indicating that the possible effect of rebinding can be accounted for by taking the effective dissociation rate to be [8] 5
5 b k = k (1 γ) = k 4πaD. Nk+ + 4πaD (5A.17) Now imagine that a receptor cluster of size ξ is sufficiently dense that it effectively acts like an absorbing disc of radius r. Assuming Nk+ 4πrD, we have 4k D 4πrD b, = k k Nk+ k+ rr1 after relating the receptor density within the cluster to the total number of receptors according to R1 = N/(πr2 ). This is consistent with the effective decay rate in equation (5A.14) under the identification ξ = r/4. The issue of protein clustering not only applies to receptors detecting extracellular signals (Sec. 5A), but also membrane-associated enzymes that play an important role in activating intracellular signaling pathways. A canonical motif is a signaling molecule in the cytoplasm being chemically modified by two antagonistic enzymes, with an activating enzyme such as a kinase located in the cell membrane and a deactivating enzyme such as a phosphatase distributed in the cytoplasm (see Sec. 9.1). The effectiveness of enzyme clustering on the substrate response depends on the number of substrate sites that have to be activated. If only one site of the cytoplasmic molecule has to be activated, then enzyme clustering reduces the response, since the effective target size presented to the substrate molecules is smaller. On the hand, if more than one site has to be activated, then clustering can enhance the response due to the rapid rebinding property of protein clusters [6]. This is shown schematically in Fig. 5A.1. Fig. 5A.1: (A) Clustering of enzyme molecules reduces their effective target size for substrate molecules in the bulk, which decreases substrate activation. (B) On the other hand, clustering also enhances rapid substrate rebinding, which increases substrate activation if multisite modification is required. Supplementary references 1. Berg, H. C., Purcell, E. M.: Physics of chemoreception. Biophys. J. 20, (1977). 6
6 2. Chu, C. L., J. A. Buczek-Thomas, and M. A. Nugent. Heparan sulfate proteoglycans modulate fibroblast growth factor-2 binding through a lipid-raft-mediated mechanism. Biochem. J. 379: (2004). 3. Gopalakrishnan, M., Forsten-Williams, K., Nugent, M. A., Taubery, U. C. Effects of receptor clustering on ligand dissociation kinetics: theory and simulations. Biophys. J. 89, (2005). 4. Ghosh, S., Gopalakrishnan, M., Forsten-Williams, K.: Self-consistent theory of reversible ligand binding to a spherical cell. Phys. Biol (2007). 5. Kramer, K. L., and H. J. Yost Heparan sulfate core proteins in cell-cell signaling. Annu. Rev. Genet. 37: Mugler, A., Bailey, A.G., Takahashi, K., ten Wolde, P.R., Membrane clustering and the role of rebinding in biochemical signaling. Biophys. J. 102, Munro, S Lipid rafts: elusive or illusive? Cell. 115: Shoup, D., Szabo, A.: Role of diffusion in ligand binding to macromolecules and cell-bound receptors Biophys. J. 40, (1982) 9. Simons, K., and W. L. C. Vaz Model systems, lipid rafts and cell membranes. Annu. Rev. Biophys. Biomol. Struct. 33:
Problems in diffusion and absorption: How fast can you hit a target with a random walk?
 Problems in diffusion and absorption: How fast can you hit a target with a random walk? Andrew J. Bernoff Harvey Mudd College In collaboration with Alan Lindsay (Notre Dame) Thanks to Alan Lindsay, Michael
Problems in diffusion and absorption: How fast can you hit a target with a random walk? Andrew J. Bernoff Harvey Mudd College In collaboration with Alan Lindsay (Notre Dame) Thanks to Alan Lindsay, Michael
Advanced Higher Biology. Unit 1- Cells and Proteins 2c) Membrane Proteins
 Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
When do diffusion-limited trajectories become memoryless?
 When do diffusion-limited trajectories become memoryless? Maciej Dobrzyński CWI (Center for Mathematics and Computer Science) Kruislaan 413, 1098 SJ Amsterdam, The Netherlands Abstract Stochastic description
When do diffusion-limited trajectories become memoryless? Maciej Dobrzyński CWI (Center for Mathematics and Computer Science) Kruislaan 413, 1098 SJ Amsterdam, The Netherlands Abstract Stochastic description
Lecture 1: Random walk
 Lecture : Random walk Paul C Bressloff (Spring 209). D random walk q p r- r r+ Figure 2: A random walk on a D lattice. Consider a particle that hops at discrete times between neighboring sites on a one
Lecture : Random walk Paul C Bressloff (Spring 209). D random walk q p r- r r+ Figure 2: A random walk on a D lattice. Consider a particle that hops at discrete times between neighboring sites on a one
Lecture 2: Receptor-ligand binding and cooperativity
 Lecture 2: Receptor-ligand binding and cooperativity Paul C Bressloff (Spring 209) A biochemical receptor is a protein molecule that receives a chemical signal in the form of ligand molecules. The ligands
Lecture 2: Receptor-ligand binding and cooperativity Paul C Bressloff (Spring 209) A biochemical receptor is a protein molecule that receives a chemical signal in the form of ligand molecules. The ligands
56:198:582 Biological Networks Lecture 11
 56:198:582 Biological Networks Lecture 11 Network Motifs in Signal Transduction Networks Signal transduction networks Signal transduction networks are composed of interactions between signaling proteins.
56:198:582 Biological Networks Lecture 11 Network Motifs in Signal Transduction Networks Signal transduction networks Signal transduction networks are composed of interactions between signaling proteins.
Macromolecular Crowding
 Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
The following question(s) were incorrectly answered.
 Name: Marcie Joseph Module: Cells & chemistry Test topic/animation: My animations/all animations Test type: Multiple choice Score: 48/50 Percent correct: 96% The following question(s) were incorrectly
Name: Marcie Joseph Module: Cells & chemistry Test topic/animation: My animations/all animations Test type: Multiple choice Score: 48/50 Percent correct: 96% The following question(s) were incorrectly
COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION
 COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION I. Aksan 1, M. Sen 2, M. K. Araz 3, and M. L. Kurnaz 3 1 School of Biological Sciences, University of Manchester,
COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION I. Aksan 1, M. Sen 2, M. K. Araz 3, and M. L. Kurnaz 3 1 School of Biological Sciences, University of Manchester,
School of Computer Science and Communication, Royal Institute of Technology, Stockholm, Sweden, 3
 Presented at the COMSOL Conference 2009 Milan Qasim Ali Chaudhry 1, Michael Hanke 2, Ralf Morgenstern 3 1,2 School of Computer Science and Communication, Royal Institute of Technology, Stockholm, Sweden,
Presented at the COMSOL Conference 2009 Milan Qasim Ali Chaudhry 1, Michael Hanke 2, Ralf Morgenstern 3 1,2 School of Computer Science and Communication, Royal Institute of Technology, Stockholm, Sweden,
Bacterial Chemotaxis
 Bacterial Chemotaxis Bacteria can be attracted/repelled by chemicals Mechanism? Chemoreceptors in bacteria. attractant Adler, 1969 Science READ! This is sensing, not metabolism Based on genetic approach!!!
Bacterial Chemotaxis Bacteria can be attracted/repelled by chemicals Mechanism? Chemoreceptors in bacteria. attractant Adler, 1969 Science READ! This is sensing, not metabolism Based on genetic approach!!!
Visual pigments. Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2019
 Visual pigments Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2019 References Webvision: The Organization of the Retina and Visual System (http://www.ncbi.nlm.nih.gov/books/nbk11522/#a 127) The
Visual pigments Neuroscience, Biochemistry Dr. Mamoun Ahram Third year, 2019 References Webvision: The Organization of the Retina and Visual System (http://www.ncbi.nlm.nih.gov/books/nbk11522/#a 127) The
Kinetics of protein binding in solid-phase immunoassays: Theory
 THE JOURNAL OF CHEMICAL PHYSICS 22, 2475 2005 Kinetics of protein binding in solid-phase immunoassays: Theory Konstantin V. Klenin Division of Biophysics of Macromolecules, German Cancer Research Center,
THE JOURNAL OF CHEMICAL PHYSICS 22, 2475 2005 Kinetics of protein binding in solid-phase immunoassays: Theory Konstantin V. Klenin Division of Biophysics of Macromolecules, German Cancer Research Center,
Reception The target cell s detection of a signal coming from outside the cell May Occur by: Direct connect Through signal molecules
 Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
From cell biology to Petri nets. Rainer Breitling, Groningen, NL David Gilbert, London, UK Monika Heiner, Cottbus, DE
 From cell biology to Petri nets Rainer Breitling, Groningen, NL David Gilbert, London, UK Monika Heiner, Cottbus, DE Biology = Concentrations Breitling / 2 The simplest chemical reaction A B irreversible,
From cell biology to Petri nets Rainer Breitling, Groningen, NL David Gilbert, London, UK Monika Heiner, Cottbus, DE Biology = Concentrations Breitling / 2 The simplest chemical reaction A B irreversible,
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
return in class, or Rm B
 Last lectures: Genetic Switches and Oscillators PS #2 due today bf before 3PM return in class, or Rm. 68 371B Naturally occurring: lambda lysis-lysogeny decision lactose operon in E. coli Engineered: genetic
Last lectures: Genetic Switches and Oscillators PS #2 due today bf before 3PM return in class, or Rm. 68 371B Naturally occurring: lambda lysis-lysogeny decision lactose operon in E. coli Engineered: genetic
Quantifying Intermittent Transport in Cell Cytoplasm
 Quantifying Intermittent Transport in Cell Cytoplasm Ecole Normale Supérieure, Mathematics and Biology Department. Paris, France. May 19 th 2009 Cellular Transport Introduction Cellular Transport Intermittent
Quantifying Intermittent Transport in Cell Cytoplasm Ecole Normale Supérieure, Mathematics and Biology Department. Paris, France. May 19 th 2009 Cellular Transport Introduction Cellular Transport Intermittent
NANO 243/CENG 207 Course Use Only
 L8. Drug Dispersion and Diffusion in Biological Systems (2) April 26, 2018 2. Diffusion in Water (3) Diffusion of small molecule in water drug molecules/solvents are identical to solvent Assume molecules
L8. Drug Dispersion and Diffusion in Biological Systems (2) April 26, 2018 2. Diffusion in Water (3) Diffusion of small molecule in water drug molecules/solvents are identical to solvent Assume molecules
BE 159: Signal Transduction and Mechanics in Morphogenesis
 BE 159: Signal Transduction and Mechanics in Morphogenesis Justin Bois Caltech Winter, 2018 2018 Justin Bois. This work is licensed under a Creative Commons Attribution License CC-BY 4.0. 5 Delta-Notch
BE 159: Signal Transduction and Mechanics in Morphogenesis Justin Bois Caltech Winter, 2018 2018 Justin Bois. This work is licensed under a Creative Commons Attribution License CC-BY 4.0. 5 Delta-Notch
Diffusion influence on Michaelis Menten kinetics
 JOURNAL OF CHEMICAL PHYSICS VOLUME 5, NUMBER 3 5 JULY 200 Diffusion influence on Michaelis Menten kinetics Hyojoon Kim, Mino Yang, Myung-Un Choi, and Kook Joe Shin a) School of Chemistry, Seoul National
JOURNAL OF CHEMICAL PHYSICS VOLUME 5, NUMBER 3 5 JULY 200 Diffusion influence on Michaelis Menten kinetics Hyojoon Kim, Mino Yang, Myung-Un Choi, and Kook Joe Shin a) School of Chemistry, Seoul National
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
2. At quasi-steady state or equilibrium, the net in-flux of the carrier-substrate complex CS is balanced by the net out-flux of the free carrier C.
 Facilitated Transport Instructor: Nam un Wang facilitmcd Process escription In facilitated transport, a carrier molecule C binds to the substrate to form a carrier-substrate complex C at the outer side
Facilitated Transport Instructor: Nam un Wang facilitmcd Process escription In facilitated transport, a carrier molecule C binds to the substrate to form a carrier-substrate complex C at the outer side
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biophysics I. DIFFUSION
 Biophysics I. DIFFUSION Experiment add a droplet of ink to a glass of water Observation: the stain spreads and eventually colours the entire fluid add a droplet of ink to HOT and COLD water Observation:
Biophysics I. DIFFUSION Experiment add a droplet of ink to a glass of water Observation: the stain spreads and eventually colours the entire fluid add a droplet of ink to HOT and COLD water Observation:
Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013
 Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013 Lecture 10. Biochemical Transformations II. Phosphoryl transfer and the kinetics and thermodynamics of energy currency in the cell: ATP and GTP.
Chemistry 5.07SC Biological Chemistry I Fall Semester, 2013 Lecture 10. Biochemical Transformations II. Phosphoryl transfer and the kinetics and thermodynamics of energy currency in the cell: ATP and GTP.
The energy speed accuracy trade-off in sensory adaptation
 SUPPLEMENTARY INFORMATION DOI: 1.138/NPHYS2276 The energy speed accuracy trade-off in sensory adaptation 1 The effective potential H(m) In the main text, we introduced a set of F a and F m functions to
SUPPLEMENTARY INFORMATION DOI: 1.138/NPHYS2276 The energy speed accuracy trade-off in sensory adaptation 1 The effective potential H(m) In the main text, we introduced a set of F a and F m functions to
Richik N. Ghosh, Linnette Grove, and Oleg Lapets ASSAY and Drug Development Technologies 2004, 2:
 1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
C C C C 2 C 2 C 2 C + u + v + (w + w P ) = D t x y z X. (1a) y 2 + D Z. z 2
 This chapter provides an introduction to the transport of particles that are either more dense (e.g. mineral sediment) or less dense (e.g. bubbles) than the fluid. A method of estimating the settling velocity
This chapter provides an introduction to the transport of particles that are either more dense (e.g. mineral sediment) or less dense (e.g. bubbles) than the fluid. A method of estimating the settling velocity
Intercellular communication
 Intercellular communication Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada Outline General principle of intercellular communicabon Signal molecules
Intercellular communication Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada Outline General principle of intercellular communicabon Signal molecules
Probabilistic Model Checking for Biochemical Reaction Systems
 Probabilistic Model Checing for Biochemical Reaction Systems Ratana Ty Ken-etsu Fujita Ken ichi Kawanishi Graduated School of Science and Technology Gunma University Abstract Probabilistic model checing
Probabilistic Model Checing for Biochemical Reaction Systems Ratana Ty Ken-etsu Fujita Ken ichi Kawanishi Graduated School of Science and Technology Gunma University Abstract Probabilistic model checing
Testing the transition state theory in stochastic dynamics of a. genetic switch
 Testing the transition state theory in stochastic dynamics of a genetic switch Tomohiro Ushikubo 1, Wataru Inoue, Mitsumasa Yoda 1 1,, 3, and Masaki Sasai 1 Department of Computational Science and Engineering,
Testing the transition state theory in stochastic dynamics of a genetic switch Tomohiro Ushikubo 1, Wataru Inoue, Mitsumasa Yoda 1 1,, 3, and Masaki Sasai 1 Department of Computational Science and Engineering,
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2011 Monday, 27 June, 9.30 am 12.30 pm Answer
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
Chemistry 25 (Spring term 2016) Final Examination for NON-SENIORS GOOD LUCK AND HAVE A GREAT SUMMER!!!
 Name ANSWER KEY Distributed Thursday, June 2 Due Chemistry 25 (Spring term 2016) Final Examination for NON-SENIORS GOOD LUCK AND HAVE A GREAT SUMMER!!! by Thursday, June 2 at 1 pm to 362 Broad a drop box
Name ANSWER KEY Distributed Thursday, June 2 Due Chemistry 25 (Spring term 2016) Final Examination for NON-SENIORS GOOD LUCK AND HAVE A GREAT SUMMER!!! by Thursday, June 2 at 1 pm to 362 Broad a drop box
An introduction to mathematical modeling. of signal transduction and gene control networks
 An introduction to mathematical modeling of signal transduction and gene control networks Hans G. Othmer Department of Mathematics University of Minnesota Minneapolis, MN Overview First Lecture: An introduction
An introduction to mathematical modeling of signal transduction and gene control networks Hans G. Othmer Department of Mathematics University of Minnesota Minneapolis, MN Overview First Lecture: An introduction
arxiv: v1 [q-bio.bm] 20 Dec 2007
![arxiv: v1 [q-bio.bm] 20 Dec 2007 arxiv: v1 [q-bio.bm] 20 Dec 2007](/thumbs/93/111959027.jpg) Abstract arxiv:0712.3467v1 [q-bio.bm] 20 Dec 2007 The mean time required by a transcription factor (TF) or an enzyme to find a target in the nucleus is of prime importance for the initialization of transcription,
Abstract arxiv:0712.3467v1 [q-bio.bm] 20 Dec 2007 The mean time required by a transcription factor (TF) or an enzyme to find a target in the nucleus is of prime importance for the initialization of transcription,
Cellular Automata Approaches to Enzymatic Reaction Networks
 Cellular Automata Approaches to Enzymatic Reaction Networks Jörg R. Weimar Institute of Scientific Computing, Technical University Braunschweig, D-38092 Braunschweig, Germany J.Weimar@tu-bs.de, http://www.jweimar.de
Cellular Automata Approaches to Enzymatic Reaction Networks Jörg R. Weimar Institute of Scientific Computing, Technical University Braunschweig, D-38092 Braunschweig, Germany J.Weimar@tu-bs.de, http://www.jweimar.de
System Biology - Deterministic & Stochastic Dynamical Systems
 System Biology - Deterministic & Stochastic Dynamical Systems System Biology - Deterministic & Stochastic Dynamical Systems 1 The Cell System Biology - Deterministic & Stochastic Dynamical Systems 2 The
System Biology - Deterministic & Stochastic Dynamical Systems System Biology - Deterministic & Stochastic Dynamical Systems 1 The Cell System Biology - Deterministic & Stochastic Dynamical Systems 2 The
Unit 2: Cells Guided Reading Questions (55 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 6 A Tour of the Cell Unit 2: Cells Guided Reading Questions (55
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 6 A Tour of the Cell Unit 2: Cells Guided Reading Questions (55
2 Dilution of Proteins Due to Cell Growth
 Problem Set 1 1 Transcription and Translation Consider the following set of reactions describing the process of maing a protein out of a gene: G β g G + M M α m M β m M + X X + S 1+ 1 X S 2+ X S X S 2
Problem Set 1 1 Transcription and Translation Consider the following set of reactions describing the process of maing a protein out of a gene: G β g G + M M α m M β m M + X X + S 1+ 1 X S 2+ X S X S 2
Unit 2: Cells Guided Reading Questions (60 pts total)
 Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 6 A Tour of the Cell Unit 2: Cells Guided Reading Questions (60
Name: AP Biology Biology, Campbell and Reece, 7th Edition Adapted from chapter reading guides originally created by Lynn Miriello Chapter 6 A Tour of the Cell Unit 2: Cells Guided Reading Questions (60
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
On the status of the Michaelis-Menten equation and its implications for enzymology
 1 On the status of the Michaelis-Menten equation and its implications for enzymology Sosale Chandrasekhar 1 Department of Organic Chemistry, Indian Institute of Science, Bangalore 560 012, India 1 E-mail:
1 On the status of the Michaelis-Menten equation and its implications for enzymology Sosale Chandrasekhar 1 Department of Organic Chemistry, Indian Institute of Science, Bangalore 560 012, India 1 E-mail:
R7.3 Receptor Kinetics
 Chapter 7 9/30/04 R7.3 Receptor Kinetics Professional Reference Shelf Just as enzymes are fundamental to life, so is the living cell s ability to receive and process signals from beyond the cell membrane.
Chapter 7 9/30/04 R7.3 Receptor Kinetics Professional Reference Shelf Just as enzymes are fundamental to life, so is the living cell s ability to receive and process signals from beyond the cell membrane.
Random Walks and Diffusion. APC 514 December 5, 2002 Cox&Shvartsman
 Random Walks and Diffusion APC 514 December 5, 2002 Cox&Shvartsman Cell motility over adhesive substrates: a periodic phenomenon Simple model for linear motion: DIMILLA PA, BARBEE K, LAUFFENBURGER DA MATHEMATICAL-MODEL
Random Walks and Diffusion APC 514 December 5, 2002 Cox&Shvartsman Cell motility over adhesive substrates: a periodic phenomenon Simple model for linear motion: DIMILLA PA, BARBEE K, LAUFFENBURGER DA MATHEMATICAL-MODEL
Quantum stochasticity and neuronal computations
 Institute for Clinical Neuroanatomy Dr. Senckenbergische Anatomie J.-W. Goethe Universität, Frankfurt am Main Quantum stochasticity and neuronal computations Peter Jedlička, MD Definition A stochastic
Institute for Clinical Neuroanatomy Dr. Senckenbergische Anatomie J.-W. Goethe Universität, Frankfurt am Main Quantum stochasticity and neuronal computations Peter Jedlička, MD Definition A stochastic
CELL BIOLOGY - CLUTCH CH. 9 - TRANSPORT ACROSS MEMBRANES.
 !! www.clutchprep.com K + K + K + K + CELL BIOLOGY - CLUTCH CONCEPT: PRINCIPLES OF TRANSMEMBRANE TRANSPORT Membranes and Gradients Cells must be able to communicate across their membrane barriers to materials
!! www.clutchprep.com K + K + K + K + CELL BIOLOGY - CLUTCH CONCEPT: PRINCIPLES OF TRANSMEMBRANE TRANSPORT Membranes and Gradients Cells must be able to communicate across their membrane barriers to materials
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer
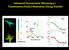 Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Where do differential equations come from?
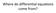 Where do differential equations come from? Example: Keeping track of cell growth (a balance equation for cell mass) Rate of change = rate in rate out Old example: Cell size The cell is spherical. Nutrient
Where do differential equations come from? Example: Keeping track of cell growth (a balance equation for cell mass) Rate of change = rate in rate out Old example: Cell size The cell is spherical. Nutrient
Lecture 3 13/11/2018
 Lecture 3 13/11/2018 1 Plasma membrane ALL cells have a cell membrane made of proteins and lipids. protein channel Cell Membrane Layer 1 Layer 2 lipid bilayer protein pump Lipid bilayer allows water, carbon
Lecture 3 13/11/2018 1 Plasma membrane ALL cells have a cell membrane made of proteins and lipids. protein channel Cell Membrane Layer 1 Layer 2 lipid bilayer protein pump Lipid bilayer allows water, carbon
Systems Biology Across Scales: A Personal View XIV. Intra-cellular systems IV: Signal-transduction and networks. Sitabhra Sinha IMSc Chennai
 Systems Biology Across Scales: A Personal View XIV. Intra-cellular systems IV: Signal-transduction and networks Sitabhra Sinha IMSc Chennai Intra-cellular biochemical networks Metabolic networks Nodes:
Systems Biology Across Scales: A Personal View XIV. Intra-cellular systems IV: Signal-transduction and networks Sitabhra Sinha IMSc Chennai Intra-cellular biochemical networks Metabolic networks Nodes:
Principles of Synthetic Biology: Midterm Exam
 Principles of Synthetic Biology: Midterm Exam October 28, 2010 1 Conceptual Simple Circuits 1.1 Consider the plots in figure 1. Identify all critical points with an x. Put a circle around the x for each
Principles of Synthetic Biology: Midterm Exam October 28, 2010 1 Conceptual Simple Circuits 1.1 Consider the plots in figure 1. Identify all critical points with an x. Put a circle around the x for each
GFD 2006 Lecture 2: Diffusion-controlled solidification
 GFD 2006 Lecture 2: Diffusion-controlled solidification Grae Worster; notes by Victor Tsai and Dan Goldberg March 15, 2007 1 Finishing off Lecture 1 As shown in Lecture 1, an approximation for the diffusion
GFD 2006 Lecture 2: Diffusion-controlled solidification Grae Worster; notes by Victor Tsai and Dan Goldberg March 15, 2007 1 Finishing off Lecture 1 As shown in Lecture 1, an approximation for the diffusion
Kinetic modelling of coupled transport across biological membranes
 Indian Journal of Biochemistry & Biophysics Vol. 51, April 14, pp. 93-99 Kinetic modelling of coupled transport across biological membranes Kalyani Korla and Chanchal K Mitra* School of Life Sciences,
Indian Journal of Biochemistry & Biophysics Vol. 51, April 14, pp. 93-99 Kinetic modelling of coupled transport across biological membranes Kalyani Korla and Chanchal K Mitra* School of Life Sciences,
Time allowed: 2 hours Answer ALL questions in Section A, ALL PARTS of the question in Section B and ONE question from Section C.
 UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2015-2016 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO-4004B Time allowed: 2 hours Answer ALL questions in Section
UNIVERSITY OF EAST ANGLIA School of Biological Sciences Main Series UG Examination 2015-2016 FUNDAMENTALS OF CELL BIOLOGY AND BIOCHEMISTRY BIO-4004B Time allowed: 2 hours Answer ALL questions in Section
Theory for Ligand Rebinding at Cell Membrane Surfaces
 Biophysical Journal Volume 74 March 998 25 228 25 Theory for Ligand Rebinding at Cell Membrane Surfaces B. Christoffer Lagerholm and Nancy L. Thompson Department of Chemistry, University of North Carolina,
Biophysical Journal Volume 74 March 998 25 228 25 Theory for Ligand Rebinding at Cell Membrane Surfaces B. Christoffer Lagerholm and Nancy L. Thompson Department of Chemistry, University of North Carolina,
Brownian Motion: Fokker-Planck Equation
 Chapter 7 Brownian Motion: Fokker-Planck Equation The Fokker-Planck equation is the equation governing the time evolution of the probability density of the Brownian particla. It is a second order differential
Chapter 7 Brownian Motion: Fokker-Planck Equation The Fokker-Planck equation is the equation governing the time evolution of the probability density of the Brownian particla. It is a second order differential
Problem Set No. 4 Due: Monday, 11/18/10 at the start of class
 Department of Chemical Engineering ChE 170 University of California, Santa Barbara Fall 2010 Problem Set No. 4 Due: Monday, 11/18/10 at the start of class Objective: To understand the thermodynamic and
Department of Chemical Engineering ChE 170 University of California, Santa Barbara Fall 2010 Problem Set No. 4 Due: Monday, 11/18/10 at the start of class Objective: To understand the thermodynamic and
Are DNA Transcription Factor Proteins Maxwellian Demons?
 Are DNA Transcription Factor Proteins Maxwellian Demons? * physics as a design constraint on molecular evolution. Alexander Grosberg, Longhua Hu, U. Minnesota, NYU U. Minnesota Biophysical Journal 2008
Are DNA Transcription Factor Proteins Maxwellian Demons? * physics as a design constraint on molecular evolution. Alexander Grosberg, Longhua Hu, U. Minnesota, NYU U. Minnesota Biophysical Journal 2008
AUTOMOTIVE EXHAUST AFTERTREATMENT
 AUTOMOTIVE EXHAUST AFTERTREATMENT CATALYST FUNDAMENTLS Catalyst in its simplest term is a material that increase the rate (molecules converted by unit time) of a chemical reaction while itself not undergoing
AUTOMOTIVE EXHAUST AFTERTREATMENT CATALYST FUNDAMENTLS Catalyst in its simplest term is a material that increase the rate (molecules converted by unit time) of a chemical reaction while itself not undergoing
One dimensional steady state diffusion, with and without source. Effective transfer coefficients
 One dimensional steady state diffusion, with and without source. Effective transfer coefficients 2 mars 207 For steady state situations t = 0) and if convection is not present or negligible the transport
One dimensional steady state diffusion, with and without source. Effective transfer coefficients 2 mars 207 For steady state situations t = 0) and if convection is not present or negligible the transport
Minimal encounter time and separation determine ligand-receptor binding in cell adhesion SUPPLEMENTARY INFORMATION
 Minimal encounter time and separation determine ligand-receptor binding in cell adhesion SUPPLEMENTARY INFORMATION Philippe Robert Alice Nicolas Said Aranda-Espinoza Pierre Bongrand Laurent Limozin Molecular
Minimal encounter time and separation determine ligand-receptor binding in cell adhesion SUPPLEMENTARY INFORMATION Philippe Robert Alice Nicolas Said Aranda-Espinoza Pierre Bongrand Laurent Limozin Molecular
Physics 4488/6562: Statistical Mechanics Material for Week 2 Exercises due Monday Feb 5 Last
 Physics 4488/6562: Statistical Mechanics http://www.physics.cornell.edu/sethna/teaching/562/ Material for Week 2 Exercises due Monday Feb 5 Last correction at January 26, 218, 11:5 am c 217, James Sethna,
Physics 4488/6562: Statistical Mechanics http://www.physics.cornell.edu/sethna/teaching/562/ Material for Week 2 Exercises due Monday Feb 5 Last correction at January 26, 218, 11:5 am c 217, James Sethna,
Macromolecular Interactions the equilibrium element
 Macromolecular Interactions the equilibrium element Physical Reality Quantitative P + L PL K d,overall K d,overall = [P][L] [PL] Driving force is difference in ground state free energies ΔG f o ΔG f o
Macromolecular Interactions the equilibrium element Physical Reality Quantitative P + L PL K d,overall K d,overall = [P][L] [PL] Driving force is difference in ground state free energies ΔG f o ΔG f o
Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme
 Supplementary Information: Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme Lee Freiburger, 1 Teresa Miletti, 1 Siqi Zhu, 1 Oliver Baettig, Albert Berghuis, Karine
Supplementary Information: Substrate-dependent switching of the allosteric binding mechanism of a dimeric enzyme Lee Freiburger, 1 Teresa Miletti, 1 Siqi Zhu, 1 Oliver Baettig, Albert Berghuis, Karine
Math 345 Intro to Math Biology Lecture 19: Models of Molecular Events and Biochemistry
 Math 345 Intro to Math Biology Lecture 19: Models of Molecular Events and Biochemistry Junping Shi College of William and Mary, USA Molecular biology and Biochemical kinetics Molecular biology is one of
Math 345 Intro to Math Biology Lecture 19: Models of Molecular Events and Biochemistry Junping Shi College of William and Mary, USA Molecular biology and Biochemical kinetics Molecular biology is one of
Ch 4: Cellular Metabolism, Part 1
 Developed by John Gallagher, MS, DVM Ch 4: Cellular Metabolism, Part 1 Energy as it relates to Biology Energy for synthesis and movement Energy transformation Enzymes and how they speed reactions Metabolism
Developed by John Gallagher, MS, DVM Ch 4: Cellular Metabolism, Part 1 Energy as it relates to Biology Energy for synthesis and movement Energy transformation Enzymes and how they speed reactions Metabolism
Concept review: Binding equilibria
 Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Lecture 7: Simple genetic circuits I
 Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
REVIEW 2: CELLS & CELL COMMUNICATION. A. Top 10 If you learned anything from this unit, you should have learned:
 Name AP Biology REVIEW 2: CELLS & CELL COMMUNICATION A. Top 10 If you learned anything from this unit, you should have learned: 1. Prokaryotes vs. eukaryotes No internal membranes vs. membrane-bound organelles
Name AP Biology REVIEW 2: CELLS & CELL COMMUNICATION A. Top 10 If you learned anything from this unit, you should have learned: 1. Prokaryotes vs. eukaryotes No internal membranes vs. membrane-bound organelles
Supporting Information
 Supporting Information Computation of a Theoretical Membrane Phase Diagram, and the Role of Phase in Lipid Raft-Mediated Protein Organization Eshan D. Mitra 1, Samuel C. Whitehead, David Holowka, Barbara
Supporting Information Computation of a Theoretical Membrane Phase Diagram, and the Role of Phase in Lipid Raft-Mediated Protein Organization Eshan D. Mitra 1, Samuel C. Whitehead, David Holowka, Barbara
Challenges of Single Ion-Channel Biosensors
 Vanderbilt Institute for Integrative Biosystems Research and Education Instrumenting and Control the Single Cell Challenges of Single Ion-Channel Biosensors Fredrick Sachs (SUNY Buffalo) John P. Wikswo
Vanderbilt Institute for Integrative Biosystems Research and Education Instrumenting and Control the Single Cell Challenges of Single Ion-Channel Biosensors Fredrick Sachs (SUNY Buffalo) John P. Wikswo
A A + B. ra + A + 1. We now want to solve the Einstein equations in the following cases:
 Lecture 29: Cosmology Cosmology Reading: Weinberg, Ch A metric tensor appropriate to infalling matter In general (see, eg, Weinberg, Ch ) we may write a spherically symmetric, time-dependent metric in
Lecture 29: Cosmology Cosmology Reading: Weinberg, Ch A metric tensor appropriate to infalling matter In general (see, eg, Weinberg, Ch ) we may write a spherically symmetric, time-dependent metric in
Energy Transformation, Cellular Energy & Enzymes (Outline)
 Energy Transformation, Cellular Energy & Enzymes (Outline) Energy conversions and recycling of matter in the ecosystem. Forms of energy: potential and kinetic energy The two laws of thermodynamic and definitions
Energy Transformation, Cellular Energy & Enzymes (Outline) Energy conversions and recycling of matter in the ecosystem. Forms of energy: potential and kinetic energy The two laws of thermodynamic and definitions
DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS. Scientific Background on the Nobel Prize in Medicine 2013
 DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS Scientific Background on the Nobel Prize in Medicine 2013 Daniela Scalet 6/12/2013 The Nobel Prize in Medicine
DISCOVERIES OF MACHINERY REGULATING VESICLE TRAFFIC, A MAJOR TRANSPORT SYSTEM IN OUR CELLS Scientific Background on the Nobel Prize in Medicine 2013 Daniela Scalet 6/12/2013 The Nobel Prize in Medicine
Basic Principles of Kinetics and Thermodynamics Arthur L. Haas, Ph.D. Department of Biochemistry and Molecular Biology
 Basic Principles of Kinetics and Thermodynamics Arthur L. Haas, Ph.D. Department of Biochemistry and Molecular Biology Review: Garrett and Grisham, Enzyme Kinetics (Chapt. 13), in Biochemistry, Third Edition,
Basic Principles of Kinetics and Thermodynamics Arthur L. Haas, Ph.D. Department of Biochemistry and Molecular Biology Review: Garrett and Grisham, Enzyme Kinetics (Chapt. 13), in Biochemistry, Third Edition,
TOPIC 6: Chemical kinetics
 TOPIC 6: Chemical kinetics Reaction rates Reaction rate laws Integrated reaction rate laws Reaction mechanism Kinetic theories Arrhenius law Catalysis Enzimatic catalysis Fuente: Cedre http://loincognito.-iles.wordpress.com/202/04/titanic-
TOPIC 6: Chemical kinetics Reaction rates Reaction rate laws Integrated reaction rate laws Reaction mechanism Kinetic theories Arrhenius law Catalysis Enzimatic catalysis Fuente: Cedre http://loincognito.-iles.wordpress.com/202/04/titanic-
BAE 820 Physical Principles of Environmental Systems
 BAE 820 Physical Principles of Environmental Systems Catalysis of environmental reactions Dr. Zifei Liu Catalysis and catalysts Catalysis is the increase in the rate of a chemical reaction due to the participation
BAE 820 Physical Principles of Environmental Systems Catalysis of environmental reactions Dr. Zifei Liu Catalysis and catalysts Catalysis is the increase in the rate of a chemical reaction due to the participation
Quantifying the Affinities and Kinetics of Protein Interactions Using Silicon Nanowire Biosensors
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.82 Quantifying the Affinities and Kinetics of Protein Interactions Using Silicon Nanowire Biosensors Xuexin Duan, Yue Li, Nitin K. Rajan, David A. Routenberg,
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.82 Quantifying the Affinities and Kinetics of Protein Interactions Using Silicon Nanowire Biosensors Xuexin Duan, Yue Li, Nitin K. Rajan, David A. Routenberg,
First Invader Dynamics in Diffusion-Controlled Absorption
 First Invader Dynamics in Diffusion-Controlled Absorption S. Redner Department of Physics, Boston University, Boston, MA 2215, USA and Santa Fe Institute, 1399 Hyde Park Road, Santa Fe, New Mexico 8751,
First Invader Dynamics in Diffusion-Controlled Absorption S. Redner Department of Physics, Boston University, Boston, MA 2215, USA and Santa Fe Institute, 1399 Hyde Park Road, Santa Fe, New Mexico 8751,
Mathematical Modelling of Partially Absorbing Boundaries in Biological Systems
 Mathematical Modelling of Partially Absorbing Boundaries in Biological Systems by Sarafa Adewale Iyaniwura B.Sc., University of Ilorin, 2012 M.Sc., AIMS-Stellenbosch University, 2014 A THESIS SUBMITTED
Mathematical Modelling of Partially Absorbing Boundaries in Biological Systems by Sarafa Adewale Iyaniwura B.Sc., University of Ilorin, 2012 M.Sc., AIMS-Stellenbosch University, 2014 A THESIS SUBMITTED
Study of Tricyclic Cascade Networks using Dynamic Optimization
 Study of Tricyclic Cascade Networks using Dynamic Optimization Systems Biology interdisciplinary field that focuses on the systematic study of complex interactions in biological systems Signal transduction
Study of Tricyclic Cascade Networks using Dynamic Optimization Systems Biology interdisciplinary field that focuses on the systematic study of complex interactions in biological systems Signal transduction
Compartmental Modelling: Eigenvalues, Traps and Nonlinear Models
 Compartmental Modelling: Eigenvalues, Traps and Nonlinear Models Neil D. Evans neil.evans@warwick.ac.uk School of Engineering, University of Warwick Systems Pharmacology - Pharmacokinetics Vacation School,
Compartmental Modelling: Eigenvalues, Traps and Nonlinear Models Neil D. Evans neil.evans@warwick.ac.uk School of Engineering, University of Warwick Systems Pharmacology - Pharmacokinetics Vacation School,
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins
 13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
Network Biology: Understanding the cell s functional organization. Albert-László Barabási Zoltán N. Oltvai
 Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
3.1 Cell Theory. KEY CONCEPT Cells are the Basic unit of life.
 3.1 Cell Theory KEY CONCEPT Cells are the Basic unit of life. 3.1 Cell Theory The cell theory grew out of the work of many scientists and improvements in the microscope. Many scientists contributed to
3.1 Cell Theory KEY CONCEPT Cells are the Basic unit of life. 3.1 Cell Theory The cell theory grew out of the work of many scientists and improvements in the microscope. Many scientists contributed to
BOUNDARY HOMOGENIZATION AND CAPTURE TIME DISTRIBUTIONS OF SEMI-PERMEABLE MEMBRANES WITH PERIODIC PATTERNS OF REACTIVE SITES.
 BOUNDARY HOMOGENIZATION AND CAPTURE TIME DISTRIBUTIONS OF SEMI-PERMEABLE MEMBRANES WITH PERIODIC PATTERNS OF REACTIVE SITES. ANDREW J. BERNOFF, ALAN E. LINDSAY, AND DANIEL D. SCHMIDT Abstract. In this
BOUNDARY HOMOGENIZATION AND CAPTURE TIME DISTRIBUTIONS OF SEMI-PERMEABLE MEMBRANES WITH PERIODIC PATTERNS OF REACTIVE SITES. ANDREW J. BERNOFF, ALAN E. LINDSAY, AND DANIEL D. SCHMIDT Abstract. In this
An augmented Keller-Segal model for E. coli chemotaxis in fast-varying environments
 An augmented Keller-Segal model for E. coli chemotaxis in fast-varying environments Tong Li Min Tang Xu Yang August 6, 5 Abstract This is a continuous study on E. coli chemotaxis under the framework of
An augmented Keller-Segal model for E. coli chemotaxis in fast-varying environments Tong Li Min Tang Xu Yang August 6, 5 Abstract This is a continuous study on E. coli chemotaxis under the framework of
Part One: Reaction Rates. 1. Rates of chemical reactions. (how fast products are formed and/or reactants are used up)
 A. Chemical Kinetics deals with: CHAPTER 13: RATES OF REACTION Part One: Reaction Rates 1. Rates of chemical reactions. (how fast products are formed and/or reactants are used up) 2. Mechanisms of chemical
A. Chemical Kinetics deals with: CHAPTER 13: RATES OF REACTION Part One: Reaction Rates 1. Rates of chemical reactions. (how fast products are formed and/or reactants are used up) 2. Mechanisms of chemical
BE/APh161: Physical Biology of the Cell Homework 2 Due Date: Wednesday, January 18, 2017
 BE/APh161: Physical Biology of the Cell Homework 2 Due Date: Wednesday, January 18, 2017 Doubt is the father of creation. - Galileo Galilei 1. Number of mrna in a cell. In this problem, we are going to
BE/APh161: Physical Biology of the Cell Homework 2 Due Date: Wednesday, January 18, 2017 Doubt is the father of creation. - Galileo Galilei 1. Number of mrna in a cell. In this problem, we are going to
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
AP3162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions
 AP162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions Hyun Youk Delft University of Technology (Dated: March 15, 2018) In this lecture, we will derive the Berg-Purcell
AP162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions Hyun Youk Delft University of Technology (Dated: March 15, 2018) In this lecture, we will derive the Berg-Purcell
Part One: Reaction Rates. 1. Even though a reaction is thermodynamically favorable it may not occur at all if it is kinetically very slow.
 CHAPTER 13: RATES OF REACTION Part One: Reaction Rates A. Chemical Kinetics deals with: 1. 2. B. Importance: 1. Even though a reaction is thermodynamically favorable it may not occur at all if it is kinetically
CHAPTER 13: RATES OF REACTION Part One: Reaction Rates A. Chemical Kinetics deals with: 1. 2. B. Importance: 1. Even though a reaction is thermodynamically favorable it may not occur at all if it is kinetically
For one such collision, a new particle, "k", is formed of volume v k = v i + v j. The rate of formation of "k" particles is, N ij
 Coagulation Theory and The Smoluchowski Equation: Coagulation is defined as growth of particles by collisions among particles. It is usually associated with dense, 3-d growth, in contrast to aggregation
Coagulation Theory and The Smoluchowski Equation: Coagulation is defined as growth of particles by collisions among particles. It is usually associated with dense, 3-d growth, in contrast to aggregation
Lecture 27. Transition States and Enzyme Catalysis
 Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Bacterial Physiology Lec -3
 Bacterial Physiology Lec -3 Uptake of nutrients by the cell The first step in nutrient use is uptake of the required nutrients by the microbial cell,uptake mechanism must be specific-that is the necessary
Bacterial Physiology Lec -3 Uptake of nutrients by the cell The first step in nutrient use is uptake of the required nutrients by the microbial cell,uptake mechanism must be specific-that is the necessary
Introduction. Dagmar Iber Jörg Stelling. CSB Deterministic, SS 2015, 1.
 Introduction Dagmar Iber Jörg Stelling joerg.stelling@bsse.ethz.ch CSB Deterministic, SS 2015, 1 Origins of Systems Biology On this assumption of the passage of blood, made as a basis for argument, and
Introduction Dagmar Iber Jörg Stelling joerg.stelling@bsse.ethz.ch CSB Deterministic, SS 2015, 1 Origins of Systems Biology On this assumption of the passage of blood, made as a basis for argument, and
Universality of sensory-response systems
 excite.org(anism): Electrical Signaling Universality of sensory-response systems Three step process: sensation-integration-response Bacterial chemotaxis Madigan et al. Fig. 8.24 Rick Stewart (CBMG) Human
excite.org(anism): Electrical Signaling Universality of sensory-response systems Three step process: sensation-integration-response Bacterial chemotaxis Madigan et al. Fig. 8.24 Rick Stewart (CBMG) Human
