Conformational Free-Energy Differences by Confinement Simulations
|
|
|
- Evangeline Palmer
- 6 years ago
- Views:
Transcription
1 Conformational Free-Energy Differences by Confinement Simulations Marco Cecchini Laboratoire d Ingeniérie des Fonctions Moléculaires (ISIS) UMR 76 - Université de Strasbourg mcecchini@unistra.fr HPC days - Strasbourg, Jan 214
2 Outline Conformational Free Energy Confinement approach: theory and methodology Test-case application: the β-hairpin of protein G Numerical validation of the predictions Free-energy components analysis Perspective: Energy storage in myosin Cecchini et al., J Phys Chem B (29)
3 Free-Energy Landscape Conformational Free Energy HH TR!G [kcal/mol] FS [FS] 2 [HH] [FS] 3 [TR] Rao Caflisch, J Mol Biol (24) Free-energy basins correspond to unique molecular states Conformational free energy (G) is a measure of structural stability Conformational free-energy differences (ΔG) determine the relative populations of microstates
4 the Confinement Approach A AA A * G AB G NMA A B B BB B * G AB = AA BB + G NMA A B
5 Confinement: Theory Thermodynamic Integration 1 G = U(X, λ)/ λ Energy Function U(X, λ) = U ff (X) λ k f λ dλ kf has to be large X X 2 A B AA BB A * B * G NMA A B Confinement Free-Energy = 1 kf X X 2 2 k dk = 1 2 kf X k dk Atomic Fluctuations X k = 2 U cons k k or better X k = N RMSD 2 k Tyka et al., J Phys Chem B (26)
6 Confinement: the idea 1 A1 B2 1 k 1 k 2 k 3 k f χ k 1 1 X k = N RMSD 2 k kf.1 1e-5 1e G A =G A AA Confinement G [kcal/mol] B A = 1 2 kf kf k [kcal/mol A 2 ] X k dk
7 Confinement: Normal Mode Analysis A B AA BB A * B * G NMA A B Classical Partition Function Z = exp ( E /k B T ) i Absolute Free Energy k B T hν i harmonic oscillator G A = k B T log Z A confined = k B T log Z B states G B Free-Energy Difference (ΔG NMA ) G NMA A B = E B E A k B T i ln ( ωi A ω i B ) NMA leg enthalpic contribution entropic contribution G NM = H NM T S NM E from energy minimization ωi from Hessian diagonalization
8 Test case: the β-hairpin of protein Krivov Karplus, PNAS (24) 2 3
9 Confinement Simulation Setup free-energy basins 1 and 2 of the β-hairpin of protein G 23 MD simulations per confinement with harmonic restraints ranging from to 82. kcal/mol/a 2 1 ns sampling per run (2.3 μs per confinement) the reference was a deeply minimized conformation MD Setup: Langevin dynamics (γ=1. ps K, best-fit harmonic restraint, EEF1implicit solvation model, time step of 2 fs with SHAKE a Cα-RMSD cutoff of 2. A from the reference was used to define the boundaries of the free-energy basins
10 1 Confinement Simulations A1 B2 Results: Confinement Legs 1 A B χ k 1 Confinement G [kcal/mol] 1 kf.1 1e-5 1e X k = N RMSD 2 k A1 B2 = 1 2 kf kf X k dk Max strength of 82 kcal/mol/a 2 A B AA BB A * G NMA A B B * k [kcal/mol A 2 ]
11 A B Results: NMA Leg Energy [kcal/mol] Normal Mode Analysis H T S G Max strength of 82 kcal/mol/a 2 A AA A * 2.16 G NMA A B -1 kf k [kcal/mol A 2 ] B BB B * ΔGAB = = 1.88 kcal/mol
12 A B Results: Conformational ΔG!G AB [kcal/mol] !G [kcal/mol] Confinement Result A B 1.88 ±.13 ΔGAB = 1.88 ±.13 kcal/mol 1e k f [kcal/mol/a 2 ]
13 Numerical validation by equilibrium MD 5 independent MD 36 K (each 4 μs) total sampling of 2 μs a Cα-RMSD cutoff of 2. A from the reference to define the boundaries of the free-energy basins round-trip decomposition of the individual MD runs (184 roundtrips sampled in total) boot-strapping analysis of the roundtrips to estimate the free-energy difference along with its statistical error forward 1 2 backward G 12 = k B T log(# 2 /# 1 ) σ N = 1 M M i=1 ( ) 2 G i G
14 confinement approach is ~5 times more efficient than CTMD Results: Numerical Validation.12 N=184.6 y = 1.73 # x -.2 number of roundtrips.1.5 Probability !!G [Kcal/mol] !G /"N ΔG12 = 1.86 ±.12 kcal/mol ΔG12 = 1.88 ±.13 kcal/mol from 2 µs CTMD from 4 µs confinement MD
15 Take Home Messages the confinement approach is a useful method to compute the free-energy difference between conformers of a biomolecule it allows for predictions of mutational effects through free-energy components analyses it may ultimately open up to a microscopic interpretation of the free energy
16 Transformation of chemical energy into mechanical work Molecular Motors: Myosin Muscle Contraction (MyoII) Vale & Milligan, Science (2)
17 Energy Storage in Myosin Málnási-Csimadia et al., Biochemistry (21)
18 Acknowledgments Prof. Martin Karplus Dr. Sergei Krivov Dr. Martin Spichty ISIS (Strasbourg) Laboratoire d Ingeniérie des Fonctions Moléculaires
Statistical Mechanics for Proteins
 The Partition Function From Q all relevant thermodynamic properties can be obtained by differentiation of the free energy F: = kt q p E q pd d h T V Q ), ( exp 1! 1 ),, ( 3 3 3 ),, ( ln ),, ( T V Q kt
The Partition Function From Q all relevant thermodynamic properties can be obtained by differentiation of the free energy F: = kt q p E q pd d h T V Q ), ( exp 1! 1 ),, ( 3 3 3 ),, ( ln ),, ( T V Q kt
Routine access to millisecond timescale events with accelerated molecular dynamics
 Routine access to millisecond timescale events with accelerated molecular dynamics Levi C.T. Pierce, Romelia Salomon-Ferrer, Cesar Augusto F. de Oliveira #, J. Andrew McCammon #, Ross C. Walker * SUPPORTING
Routine access to millisecond timescale events with accelerated molecular dynamics Levi C.T. Pierce, Romelia Salomon-Ferrer, Cesar Augusto F. de Oliveira #, J. Andrew McCammon #, Ross C. Walker * SUPPORTING
Organization of NAMD Tutorial Files
 Organization of NAMD Tutorial Files .1.1. RMSD for individual residues Objective: Find the average RMSD over time of each residue in the protein using VMD. Display the protein with the residues colored
Organization of NAMD Tutorial Files .1.1. RMSD for individual residues Objective: Find the average RMSD over time of each residue in the protein using VMD. Display the protein with the residues colored
Theoretical/computational study of complex chemistry in biological systems
 Theoretical/computational study of complex chemistry in biological systems Preliminary insights into the mechanochemical coupling in myosin with molecular simulations Qiang Cui Department of Chemistry
Theoretical/computational study of complex chemistry in biological systems Preliminary insights into the mechanochemical coupling in myosin with molecular simulations Qiang Cui Department of Chemistry
Slow Folding of Cross-Linked r-helical Peptides Due to Steric Hindrance
 J. Phys. Chem. B 2010, 114, 2023 2027 2023 Slow Folding of Cross-Linked r-helical Peptides Due to Steric Hindrance B. Paoli, R. Pellarin, and A. Caflisch* Department of Biochemistry, UniVersity of Zurich,
J. Phys. Chem. B 2010, 114, 2023 2027 2023 Slow Folding of Cross-Linked r-helical Peptides Due to Steric Hindrance B. Paoli, R. Pellarin, and A. Caflisch* Department of Biochemistry, UniVersity of Zurich,
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data
 Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Sampling the free energy surfaces of collective variables
 Sampling the free energy surfaces of collective variables Jérôme Hénin Enhanced Sampling and Free-Energy Calculations Urbana, 12 September 2018 Please interrupt! struct bioinform phys chem theoretical
Sampling the free energy surfaces of collective variables Jérôme Hénin Enhanced Sampling and Free-Energy Calculations Urbana, 12 September 2018 Please interrupt! struct bioinform phys chem theoretical
Resonant optical rectification. Theory and application to bacteriorhodopsin
 Resonant optical rectification. Theory and application to bacteriorhodopsin Institute of Biophysics Biological Research Centre of the Hungarian Academy of Sciences Szeged, Hungary Laboratory for Optical
Resonant optical rectification. Theory and application to bacteriorhodopsin Institute of Biophysics Biological Research Centre of the Hungarian Academy of Sciences Szeged, Hungary Laboratory for Optical
Amyloid fibril polymorphism is under kinetic control. Supplementary Information
 Amyloid fibril polymorphism is under kinetic control Supplementary Information Riccardo Pellarin Philipp Schütz Enrico Guarnera Amedeo Caflisch Department of Biochemistry, University of Zürich, Winterthurerstrasse
Amyloid fibril polymorphism is under kinetic control Supplementary Information Riccardo Pellarin Philipp Schütz Enrico Guarnera Amedeo Caflisch Department of Biochemistry, University of Zürich, Winterthurerstrasse
Medical Research, Medicinal Chemistry, University of Leuven, Leuven, Belgium.
 Supporting Information Towards peptide vaccines against Zika virus: Immunoinformatics combined with molecular dynamics simulations to predict antigenic epitopes of Zika viral proteins Muhammad Usman Mirza
Supporting Information Towards peptide vaccines against Zika virus: Immunoinformatics combined with molecular dynamics simulations to predict antigenic epitopes of Zika viral proteins Muhammad Usman Mirza
Infrared spectra of small biomolecules from first-principle molecular dynamics simulations and effective normal mode analysis
 Infrared spectra of small biomolecules from first-principle molecular dynamics simulations and effective normal mode analysis R. Vuilleumier, M.-P. Gaigeot and D. Borgis Département de chimie, Ecole Normale
Infrared spectra of small biomolecules from first-principle molecular dynamics simulations and effective normal mode analysis R. Vuilleumier, M.-P. Gaigeot and D. Borgis Département de chimie, Ecole Normale
Nonequilibrium thermodynamics at the microscale
 Nonequilibrium thermodynamics at the microscale Christopher Jarzynski Department of Chemistry and Biochemistry and Institute for Physical Science and Technology ~1 m ~20 nm Work and free energy: a macroscopic
Nonequilibrium thermodynamics at the microscale Christopher Jarzynski Department of Chemistry and Biochemistry and Institute for Physical Science and Technology ~1 m ~20 nm Work and free energy: a macroscopic
Free energy calculations with alchemlyb
 Free energy calculations with alchemlyb Oliver Beckstein Arizona State University SPIDAL Teleconference 2019-02-01 Binding free energy measure of how strong a protein P and a ligand X stick together key
Free energy calculations with alchemlyb Oliver Beckstein Arizona State University SPIDAL Teleconference 2019-02-01 Binding free energy measure of how strong a protein P and a ligand X stick together key
Protein Simulations in Confined Environments
 Critical Review Lecture Protein Simulations in Confined Environments Murat Cetinkaya 1, Jorge Sofo 2, Melik C. Demirel 1 1. 2. College of Engineering, Pennsylvania State University, University Park, 16802,
Critical Review Lecture Protein Simulations in Confined Environments Murat Cetinkaya 1, Jorge Sofo 2, Melik C. Demirel 1 1. 2. College of Engineering, Pennsylvania State University, University Park, 16802,
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Statistical thermodynamics for MD and MC simulations
 Statistical thermodynamics for MD and MC simulations knowing 2 atoms and wishing to know 10 23 of them Marcus Elstner and Tomáš Kubař 22 June 2016 Introduction Thermodynamic properties of molecular systems
Statistical thermodynamics for MD and MC simulations knowing 2 atoms and wishing to know 10 23 of them Marcus Elstner and Tomáš Kubař 22 June 2016 Introduction Thermodynamic properties of molecular systems
Free energy calculations and the potential of mean force
 Free energy calculations and the potential of mean force IMA Workshop on Classical and Quantum Approaches in Molecular Modeling Mark Tuckerman Dept. of Chemistry and Courant Institute of Mathematical Science
Free energy calculations and the potential of mean force IMA Workshop on Classical and Quantum Approaches in Molecular Modeling Mark Tuckerman Dept. of Chemistry and Courant Institute of Mathematical Science
What can be learnt from MD
 - 4.0 H-bond energy (kcal/mol) 0 What can be learnt from MD Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis ATPase, a molecular motor
- 4.0 H-bond energy (kcal/mol) 0 What can be learnt from MD Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis ATPase, a molecular motor
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Limitations of temperature replica exchange (T-REMD) for protein folding simulations
 Limitations of temperature replica exchange (T-REMD) for protein folding simulations Jed W. Pitera, William C. Swope IBM Research pitera@us.ibm.com Anomalies in protein folding kinetic thermodynamic 322K
Limitations of temperature replica exchange (T-REMD) for protein folding simulations Jed W. Pitera, William C. Swope IBM Research pitera@us.ibm.com Anomalies in protein folding kinetic thermodynamic 322K
Keto-Enol Thermodynamics of Breslow Intermediates
 Keto-Enol Thermodynamics of Breslow Intermediates Mathias Paul, Martin Breugst, Jörg-Martin Neudörfl, Raghavan B. Sunoj, and Albrecht Berkessel Computational Section of the Supporting Information 1 Computational
Keto-Enol Thermodynamics of Breslow Intermediates Mathias Paul, Martin Breugst, Jörg-Martin Neudörfl, Raghavan B. Sunoj, and Albrecht Berkessel Computational Section of the Supporting Information 1 Computational
Computing the Relative Stabilities and the Per-Residue Components in Protein Conformational Changes
 Resource Computing the Relative Stabilities and the Per-Residue Components in Protein Conformational Changes Arijit Roy, 1 Alberto Perez, 1 Ken A. Dill, 1,2,3, * and Justin L. MacCallum 1 1 Laufer Center
Resource Computing the Relative Stabilities and the Per-Residue Components in Protein Conformational Changes Arijit Roy, 1 Alberto Perez, 1 Ken A. Dill, 1,2,3, * and Justin L. MacCallum 1 1 Laufer Center
Free-Energy Landscapes of Proteins in the Presence and Absence of Force
 16902 J. Phys. Chem. B 2008, 112, 16902 16907 Free-Energy Landscapes of Proteins in the Presence and Absence of Force Zu Thur Yew, Sergei Krivov, and Emanuele Paci*, Institute of Molecular and Cellular
16902 J. Phys. Chem. B 2008, 112, 16902 16907 Free-Energy Landscapes of Proteins in the Presence and Absence of Force Zu Thur Yew, Sergei Krivov, and Emanuele Paci*, Institute of Molecular and Cellular
Supporting Materials
 Supporting Materials Figure S1 Experimental Setup Page Figure S (a) (b) (c) Feynman Diagrams Page 3-6 Figure S3 D IR Spectra Page 7 Figure S4 Kinetic Model Page 8 Figure S5 Van t Hoff Plots Page 9 1 k
Supporting Materials Figure S1 Experimental Setup Page Figure S (a) (b) (c) Feynman Diagrams Page 3-6 Figure S3 D IR Spectra Page 7 Figure S4 Kinetic Model Page 8 Figure S5 Van t Hoff Plots Page 9 1 k
Many proteins spontaneously refold into native form in vitro with high fidelity and high speed.
 Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Normal Mode Analysis. Jun Wan and Willy Wriggers School of Health Information Sciences & Institute of Molecular Medicine University of Texas Houston
 Normal Mode Analysis Jun Wan and Willy Wriggers School of Health Information Sciences & Institute of Molecular Medicine University of Teas Houston Contents Introduction --- protein dynamics (lowfrequency
Normal Mode Analysis Jun Wan and Willy Wriggers School of Health Information Sciences & Institute of Molecular Medicine University of Teas Houston Contents Introduction --- protein dynamics (lowfrequency
Targeted Molecular Dynamics Simulations of Protein Unfolding
 J. Phys. Chem. B 2000, 104, 4511-4518 4511 Targeted Molecular Dynamics Simulations of Protein Unfolding Philippe Ferrara, Joannis Apostolakis, and Amedeo Caflisch* Department of Biochemistry, UniVersity
J. Phys. Chem. B 2000, 104, 4511-4518 4511 Targeted Molecular Dynamics Simulations of Protein Unfolding Philippe Ferrara, Joannis Apostolakis, and Amedeo Caflisch* Department of Biochemistry, UniVersity
Density Functional Theory: from theory to Applications
 Density Functional Theory: from theory to Applications Uni Mainz May 27, 2012 Large barrier-activated processes time-dependent bias potential extended Lagrangian formalism Basic idea: during the MD dynamics
Density Functional Theory: from theory to Applications Uni Mainz May 27, 2012 Large barrier-activated processes time-dependent bias potential extended Lagrangian formalism Basic idea: during the MD dynamics
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Application of the Markov State Model to Molecular Dynamics of Biological Molecules. Speaker: Xun Sang-Ni Supervisor: Prof. Wu Dr.
 Application of the Markov State Model to Molecular Dynamics of Biological Molecules Speaker: Xun Sang-Ni Supervisor: Prof. Wu Dr. Jiang Introduction Conformational changes of proteins : essential part
Application of the Markov State Model to Molecular Dynamics of Biological Molecules Speaker: Xun Sang-Ni Supervisor: Prof. Wu Dr. Jiang Introduction Conformational changes of proteins : essential part
Hierarchical Metastable States and Kinetic Transition Networks: Trajectory Mapping and Clustering
 Hierarchical Metastable States and Kinetic Transition Networks: Trajectory Mapping and Clustering Xin Zhou xzhou@gucas.ac.cn Graduate University of Chinese Academy of Sciences, Beijing 2012.6.5 KITP, Santa
Hierarchical Metastable States and Kinetic Transition Networks: Trajectory Mapping and Clustering Xin Zhou xzhou@gucas.ac.cn Graduate University of Chinese Academy of Sciences, Beijing 2012.6.5 KITP, Santa
Overview & Applications. T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 04 June, 2015
 Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Linear Motors. Nanostrukturphysik II, Manuel Bastuck
 Molecular l Motors I: Linear Motors Nanostrukturphysik II, Manuel Bastuck Why can he run so fast? Usain Bolt 100 m / 9,58 s Usain Bolt: http://www.wallpaperdev.com/stock/fantesty-usain-bolt.jpg muscle:
Molecular l Motors I: Linear Motors Nanostrukturphysik II, Manuel Bastuck Why can he run so fast? Usain Bolt 100 m / 9,58 s Usain Bolt: http://www.wallpaperdev.com/stock/fantesty-usain-bolt.jpg muscle:
Lecture 20. Chemical Potential
 Lecture 20 Chemical Potential Reading: Lecture 20, today: Chapter 10, sections A and B Lecture 21, Wednesday: Chapter 10: 10 17 end 3/21/16 1 Pop Question 7 Boltzmann Distribution Two systems with lowest
Lecture 20 Chemical Potential Reading: Lecture 20, today: Chapter 10, sections A and B Lecture 21, Wednesday: Chapter 10: 10 17 end 3/21/16 1 Pop Question 7 Boltzmann Distribution Two systems with lowest
Femtosecond laser microfabrication in. Prof. Dr. Cleber R. Mendonca
 Femtosecond laser microfabrication in polymers Prof. Dr. Cleber R. Mendonca laser microfabrication focus laser beam on material s surface laser microfabrication laser microfabrication laser microfabrication
Femtosecond laser microfabrication in polymers Prof. Dr. Cleber R. Mendonca laser microfabrication focus laser beam on material s surface laser microfabrication laser microfabrication laser microfabrication
Protein Dynamics, Allostery and Function
 Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Characterizing Structural Transitions of Membrane Transport Proteins at Atomic Detail Mahmoud Moradi
 Characterizing Structural Transitions of Membrane Transport Proteins at Atomic Detail Mahmoud Moradi NCSA Blue Waters Symposium for Petascale Science and Beyond Sunriver, Oregon May 11, 2015 Outline Introduction
Characterizing Structural Transitions of Membrane Transport Proteins at Atomic Detail Mahmoud Moradi NCSA Blue Waters Symposium for Petascale Science and Beyond Sunriver, Oregon May 11, 2015 Outline Introduction
Supporting Information
 Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
Conformational sampling of macrocycles in solution and in the solid state
 Conformational sampling of macrocycles in solution and in the solid state Paul Hawkins, Ph.D. Head of Scientific Solutions Stanislaw Wlodek, Ph.D. Senior Scientific Developer 6/6/2018 https://berkonomics.com/?p=2437
Conformational sampling of macrocycles in solution and in the solid state Paul Hawkins, Ph.D. Head of Scientific Solutions Stanislaw Wlodek, Ph.D. Senior Scientific Developer 6/6/2018 https://berkonomics.com/?p=2437
Optically Triggered Stepwise Double Proton Transfer in an Intramolecular Proton Relay: A Case Study of 1,8-Dihydroxy-2-naphthaldehyde (DHNA)
 Supporting Information Optically Triggered Stepwise Double Proton Transfer in an Intramolecular Proton Relay: A Case Study of 1,8-Dihydroxy-2-naphthaldehyde (DHNA) Chia-Yu Peng,, Jiun-Yi Shen,, Yi-Ting
Supporting Information Optically Triggered Stepwise Double Proton Transfer in an Intramolecular Proton Relay: A Case Study of 1,8-Dihydroxy-2-naphthaldehyde (DHNA) Chia-Yu Peng,, Jiun-Yi Shen,, Yi-Ting
Equilibrium Distribution from Distributed Computing (Simulations of Protein Folding)
 pubs.acs.org/jpcb Equilibrium Distribution from Distributed Computing (Simulations of Protein Folding) Riccardo Scalco and Amedeo Caflisch* Department of Biochemistry, University of Zurich, Winterthurerstrasse
pubs.acs.org/jpcb Equilibrium Distribution from Distributed Computing (Simulations of Protein Folding) Riccardo Scalco and Amedeo Caflisch* Department of Biochemistry, University of Zurich, Winterthurerstrasse
Free energy simulations
 Free energy simulations Marcus Elstner and Tomáš Kubař January 14, 2013 Motivation a physical quantity that is of most interest in chemistry? free energies Helmholtz F or Gibbs G holy grail of computational
Free energy simulations Marcus Elstner and Tomáš Kubař January 14, 2013 Motivation a physical quantity that is of most interest in chemistry? free energies Helmholtz F or Gibbs G holy grail of computational
Liquid-vapour oscillations of water in hydrophobic nanopores
 Alicante 2003 4th European Biophysics Congress Liquid-vapour oscillations of water in hydrophobic nanopores 7 th July 2003 Oliver Beckstein and Mark S. P. Sansom Department of Biochemistry, Laboratory
Alicante 2003 4th European Biophysics Congress Liquid-vapour oscillations of water in hydrophobic nanopores 7 th July 2003 Oliver Beckstein and Mark S. P. Sansom Department of Biochemistry, Laboratory
Free energy calculations
 Free energy calculations Jochen Hub & David van der Spoel Overview Free energies and Probabilities Thermodynamic cycles (Free energy perturbation (FEP)) Thermodynamic integration (TI) (Jarzynski equality
Free energy calculations Jochen Hub & David van der Spoel Overview Free energies and Probabilities Thermodynamic cycles (Free energy perturbation (FEP)) Thermodynamic integration (TI) (Jarzynski equality
Principles and Applications of Molecular Dynamics Simulations with NAMD
 Principles and Applications of Molecular Dynamics Simulations with NAMD Nov. 14, 2016 Computational Microscope NCSA supercomputer JC Gumbart Assistant Professor of Physics Georgia Institute of Technology
Principles and Applications of Molecular Dynamics Simulations with NAMD Nov. 14, 2016 Computational Microscope NCSA supercomputer JC Gumbart Assistant Professor of Physics Georgia Institute of Technology
Intracellular transport
 Transport in cells Intracellular transport introduction: transport in cells, molecular players etc. cooperation of motors, forces good and bad transport regulation, traffic issues, Stefan Klumpp image
Transport in cells Intracellular transport introduction: transport in cells, molecular players etc. cooperation of motors, forces good and bad transport regulation, traffic issues, Stefan Klumpp image
Potential Energy (hyper)surface
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = " d dx U(x) Conformation
An introduction to Molecular Dynamics. EMBO, June 2016
 An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
An introduction to Molecular Dynamics EMBO, June 2016 What is MD? everything that living things do can be understood in terms of the jiggling and wiggling of atoms. The Feynman Lectures in Physics vol.
Non equilibrium thermodynamics: foundations, scope, and extension to the meso scale. Miguel Rubi
 Non equilibrium thermodynamics: foundations, scope, and extension to the meso scale Miguel Rubi References S.R. de Groot and P. Mazur, Non equilibrium Thermodynamics, Dover, New York, 1984 J.M. Vilar and
Non equilibrium thermodynamics: foundations, scope, and extension to the meso scale Miguel Rubi References S.R. de Groot and P. Mazur, Non equilibrium Thermodynamics, Dover, New York, 1984 J.M. Vilar and
Convergence of replica exchange molecular dynamics
 THE JOURNAL OF CHEMICAL PHYSICS 123, 154105 2005 Convergence of replica exchange molecular dynamics Wei Zhang and Chun Wu Department of Chemistry and Biochemistry, University of Delaware, Newark, Delaware
THE JOURNAL OF CHEMICAL PHYSICS 123, 154105 2005 Convergence of replica exchange molecular dynamics Wei Zhang and Chun Wu Department of Chemistry and Biochemistry, University of Delaware, Newark, Delaware
Free energy, electrostatics, and the hydrophobic effect
 Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Supplementary Figures:
 Supplementary Figures: Supplementary Figure 1: The two strings converge to two qualitatively different pathways. A) Models of active (red) and inactive (blue) states used as end points for the string calculations
Supplementary Figures: Supplementary Figure 1: The two strings converge to two qualitatively different pathways. A) Models of active (red) and inactive (blue) states used as end points for the string calculations
The Dominant Interaction Between Peptide and Urea is Electrostatic in Nature: A Molecular Dynamics Simulation Study
 Dror Tobi 1 Ron Elber 1,2 Devarajan Thirumalai 3 1 Department of Biological Chemistry, The Hebrew University, Jerusalem 91904, Israel 2 Department of Computer Science, Cornell University, Ithaca, NY 14853
Dror Tobi 1 Ron Elber 1,2 Devarajan Thirumalai 3 1 Department of Biological Chemistry, The Hebrew University, Jerusalem 91904, Israel 2 Department of Computer Science, Cornell University, Ithaca, NY 14853
Transition Theory Abbreviated Derivation [ A - B - C] # E o. Reaction Coordinate. [ ] # æ Æ
![Transition Theory Abbreviated Derivation [ A - B - C] # E o. Reaction Coordinate. [ ] # æ Æ Transition Theory Abbreviated Derivation [ A - B - C] # E o. Reaction Coordinate. [ ] # æ Æ](/thumbs/79/79511213.jpg) Transition Theory Abbreviated Derivation A + BC æ Æ AB + C [ A - B - C] # E A BC D E o AB, C Reaction Coordinate A + BC æ æ Æ æ A - B - C [ ] # æ Æ æ A - B + C The rate of reaction is the frequency of
Transition Theory Abbreviated Derivation A + BC æ Æ AB + C [ A - B - C] # E A BC D E o AB, C Reaction Coordinate A + BC æ æ Æ æ A - B - C [ ] # æ Æ æ A - B + C The rate of reaction is the frequency of
Supporting Information
 Supporting Information Constant ph molecular dynamics reveals ph-modulated binding of two small-molecule BACE1 inhibitors Christopher R. Ellis 1,, Cheng-Chieh Tsai 1,, Xinjun Hou 2, and Jana Shen 1, 1
Supporting Information Constant ph molecular dynamics reveals ph-modulated binding of two small-molecule BACE1 inhibitors Christopher R. Ellis 1,, Cheng-Chieh Tsai 1,, Xinjun Hou 2, and Jana Shen 1, 1
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Recovering Kinetics from a Simplified Protein Folding Model Using Replica Exchange Simulations: A Kinetic Network and Effective Stochastic Dynamics
 11702 J. Phys. Chem. B 2009, 113, 11702 11709 Recovering Kinetics from a Simplified Protein Folding Model Using Replica Exchange Simulations: A Kinetic Network and Effective Stochastic Dynamics Weihua
11702 J. Phys. Chem. B 2009, 113, 11702 11709 Recovering Kinetics from a Simplified Protein Folding Model Using Replica Exchange Simulations: A Kinetic Network and Effective Stochastic Dynamics Weihua
The PLUMED plugin and free energy methods in electronic-structure-based molecular dynamics
 The PLUMED plugin and free energy methods in electronic-structure-based molecular dynamics Davide Branduardi, Theoretical Molecular Biophysics Group Max Planck for Biophysics, Frankfurt am Main, Germany
The PLUMED plugin and free energy methods in electronic-structure-based molecular dynamics Davide Branduardi, Theoretical Molecular Biophysics Group Max Planck for Biophysics, Frankfurt am Main, Germany
Multi-Ensemble Markov Models and TRAM. Fabian Paul 21-Feb-2018
 Multi-Ensemble Markov Models and TRAM Fabian Paul 21-Feb-2018 Outline Free energies Simulation types Boltzmann reweighting Umbrella sampling multi-temperature simulation accelerated MD Analysis methods
Multi-Ensemble Markov Models and TRAM Fabian Paul 21-Feb-2018 Outline Free energies Simulation types Boltzmann reweighting Umbrella sampling multi-temperature simulation accelerated MD Analysis methods
Supporting Information
 Supporting Information Allosteric communication disrupted by small molecule binding to the Imidazole glycerol phosphate synthase protein-protein interface. Ivan Rivalta*,#, George P. Lisi #, Ning-Shiuan
Supporting Information Allosteric communication disrupted by small molecule binding to the Imidazole glycerol phosphate synthase protein-protein interface. Ivan Rivalta*,#, George P. Lisi #, Ning-Shiuan
Relative binding enthalpies from molecular dynamics simulations. using a direct method
 Relative binding enthalpies from molecular dynamics simulations using a direct method Amitava Roy, 1 Duy P. Hua, 1 Joshua M. Ward, 1 and Carol Beth Post* Department of Medicinal Chemistry, Markey Center
Relative binding enthalpies from molecular dynamics simulations using a direct method Amitava Roy, 1 Duy P. Hua, 1 Joshua M. Ward, 1 and Carol Beth Post* Department of Medicinal Chemistry, Markey Center
Free energy calculations
 Free energy calculations Berk Hess May 5, 2017 Why do free energy calculations? The free energy G gives the population of states: ( ) P 1 G = exp, G = G 2 G 1 P 2 k B T Since we mostly simulate in the
Free energy calculations Berk Hess May 5, 2017 Why do free energy calculations? The free energy G gives the population of states: ( ) P 1 G = exp, G = G 2 G 1 P 2 k B T Since we mostly simulate in the
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Lots of thanks for many reasons!
 Lots of thanks for many reasons! Structure and function of glutamate transporters from free energy simulations Turgut Baştuğ TOBB ETU Transporter structures Transporters have larger structures, which
Lots of thanks for many reasons! Structure and function of glutamate transporters from free energy simulations Turgut Baştuğ TOBB ETU Transporter structures Transporters have larger structures, which
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
doi:10.1038/nature17991 Supplementary Discussion Structural comparison with E. coli EmrE The DMT superfamily includes a wide variety of transporters with 4-10 TM segments 1. Since the subfamilies of the
Are the native states of proteins kinetic traps? Abstract
 Are the native states of proteins kinetic traps? Leonor Cruzeiro CCMAR and FCT, Universidade do Algarve, 85-139 Faro, Portugal Paulo A. Lopes CITI, Departamento de Informática, Faculdade de Ciências e
Are the native states of proteins kinetic traps? Leonor Cruzeiro CCMAR and FCT, Universidade do Algarve, 85-139 Faro, Portugal Paulo A. Lopes CITI, Departamento de Informática, Faculdade de Ciências e
Govardhan Reddy. John E. Straub. D. Thirumalai*
 1162 J. Phys. Chem. B 2009, 113, 1162 1172 Influence of Preformed Asp23-Lys28 Salt Bridge on the Conformational Fluctuations of Monomers and Dimers of Aβ Peptides with Implications for ates of Fibril Formation
1162 J. Phys. Chem. B 2009, 113, 1162 1172 Influence of Preformed Asp23-Lys28 Salt Bridge on the Conformational Fluctuations of Monomers and Dimers of Aβ Peptides with Implications for ates of Fibril Formation
Essential dynamics sampling of proteins. Tuorial 6 Neva Bešker
 Essential dynamics sampling of proteins Tuorial 6 Neva Bešker Relevant time scale Why we need enhanced sampling? Interconvertion between basins is infrequent at the roomtemperature: kinetics and thermodynamics
Essential dynamics sampling of proteins Tuorial 6 Neva Bešker Relevant time scale Why we need enhanced sampling? Interconvertion between basins is infrequent at the roomtemperature: kinetics and thermodynamics
Microcalorimetry for the Life Sciences
 Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
MCB100A/Chem130 MidTerm Exam 2 April 4, 2013
 MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
Computational Studies of the Photoreceptor Rhodopsin. Scott E. Feller Wabash College
 Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
On the calculation of solvation free energy from Kirkwood- Buff integrals: A large scale molecular dynamics study
 On the calculation of solvation free energy from Kirkwood- Buff integrals: A large scale molecular dynamics study Wynand Dednam and André E. Botha Department of Physics, University of South Africa, P.O.
On the calculation of solvation free energy from Kirkwood- Buff integrals: A large scale molecular dynamics study Wynand Dednam and André E. Botha Department of Physics, University of South Africa, P.O.
Wang-Landau Monte Carlo simulation. Aleš Vítek IT4I, VP3
 Wang-Landau Monte Carlo simulation Aleš Vítek IT4I, VP3 PART 1 Classical Monte Carlo Methods Outline 1. Statistical thermodynamics, ensembles 2. Numerical evaluation of integrals, crude Monte Carlo (MC)
Wang-Landau Monte Carlo simulation Aleš Vítek IT4I, VP3 PART 1 Classical Monte Carlo Methods Outline 1. Statistical thermodynamics, ensembles 2. Numerical evaluation of integrals, crude Monte Carlo (MC)
Supporting Information for: Structure and Stability Studies of. Pharmacologically Relevant S-nitrosothiols: A Theoretical.
 Supporting Information for: Structure and Stability Studies of Pharmacologically Relevant S-nitrosothiols: A Theoretical Approach Benjamin Meyer (1,2), Alessandro Genoni (1,2) *, Ariane Boudier (3), Pierre
Supporting Information for: Structure and Stability Studies of Pharmacologically Relevant S-nitrosothiols: A Theoretical Approach Benjamin Meyer (1,2), Alessandro Genoni (1,2) *, Ariane Boudier (3), Pierre
Guessing the upper bound free-energy difference between native-like structures. Jorge A. Vila
 1 Guessing the upper bound free-energy difference between native-like structures Jorge A. Vila IMASL-CONICET, Universidad Nacional de San Luis, Ejército de Los Andes 950, 5700- San Luis, Argentina Use
1 Guessing the upper bound free-energy difference between native-like structures Jorge A. Vila IMASL-CONICET, Universidad Nacional de San Luis, Ejército de Los Andes 950, 5700- San Luis, Argentina Use
Lecture 34 Protein Unfolding Thermodynamics
 Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Physics 172H Modern Mechanics
 Physics 172H Modern Mechanics Instructor: Dr. Mark Haugan Office: PHYS 282 haugan@purdue.edu TAs: Alex Kryzwda John Lorenz akryzwda@purdue.edu jdlorenz@purdue.edu Lecture 22: Matter & Interactions, Ch.
Physics 172H Modern Mechanics Instructor: Dr. Mark Haugan Office: PHYS 282 haugan@purdue.edu TAs: Alex Kryzwda John Lorenz akryzwda@purdue.edu jdlorenz@purdue.edu Lecture 22: Matter & Interactions, Ch.
Methods of Computer Simulation. Molecular Dynamics and Monte Carlo
 Molecular Dynamics Time is of the essence in biological processes therefore how do we understand time-dependent processes at the molecular level? How do we do this experimentally? How do we do this computationally?
Molecular Dynamics Time is of the essence in biological processes therefore how do we understand time-dependent processes at the molecular level? How do we do this experimentally? How do we do this computationally?
Supplementary Information
 Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
Supplementary Information Resveratrol Serves as a Protein-Substrate Interaction Stabilizer in Human SIRT1 Activation Xuben Hou,, David Rooklin, Hao Fang *,,, Yingkai Zhang Department of Medicinal Chemistry
The Sensitivity Limits of Nanowire Biosensors
 The Sensitivity Limits of Nanowire Biosensors Xuan Gao Dept of Chemistry and Chemical Biology, Harvard University Jan. 15 th, 2007 Texas A&M University Why Nano for Bio-detection? Protein/DNA Virus Cell
The Sensitivity Limits of Nanowire Biosensors Xuan Gao Dept of Chemistry and Chemical Biology, Harvard University Jan. 15 th, 2007 Texas A&M University Why Nano for Bio-detection? Protein/DNA Virus Cell
Alchemical free energy calculations in OpenMM
 Alchemical free energy calculations in OpenMM Lee-Ping Wang Stanford Department of Chemistry OpenMM Workshop, Stanford University September 7, 2012 Special thanks to: John Chodera, Morgan Lawrenz Outline
Alchemical free energy calculations in OpenMM Lee-Ping Wang Stanford Department of Chemistry OpenMM Workshop, Stanford University September 7, 2012 Special thanks to: John Chodera, Morgan Lawrenz Outline
Unfolding CspB by means of biased molecular dynamics
 Chapter 4 Unfolding CspB by means of biased molecular dynamics 4.1 Introduction Understanding the mechanism of protein folding has been a major challenge for the last twenty years, as pointed out in the
Chapter 4 Unfolding CspB by means of biased molecular dynamics 4.1 Introduction Understanding the mechanism of protein folding has been a major challenge for the last twenty years, as pointed out in the
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Equilibration of experimentally determined protein structures for molecular dynamics simulation
 Equilibration of experimentally determined protein structures for molecular dynamics simulation Emily B. Walton and Krystyn J. VanVliet* Department of Materials Science and Engineering, Massachusetts Institute
Equilibration of experimentally determined protein structures for molecular dynamics simulation Emily B. Walton and Krystyn J. VanVliet* Department of Materials Science and Engineering, Massachusetts Institute
Supplementary Information
 1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein
 Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Temperature-Induced Dissociation of Ab Monomers from Amyloid Fibril
 1758 Biophysical Journal Volume 95 August 2008 1758 1772 Temperature-Induced Dissociation of Ab Monomers from Amyloid Fibril Takako Takeda and Dmitri K. Klimov Department of Bioinformatics and Computational
1758 Biophysical Journal Volume 95 August 2008 1758 1772 Temperature-Induced Dissociation of Ab Monomers from Amyloid Fibril Takako Takeda and Dmitri K. Klimov Department of Bioinformatics and Computational
w REXAMD: A Hamiltonian Replica Exchange Approach to Improve Free Energy Calculations for Systems with Kinetically Trapped Conformations
 pubs.acs.org/jctc Downloaded via 148.251.232.83 on March 8, 2019 at 14:33:02 (UTC). See https://pubs.acs.org/sharingguidelines for options on how to legitimately share published articles. w REXAMD: A Hamiltonian
pubs.acs.org/jctc Downloaded via 148.251.232.83 on March 8, 2019 at 14:33:02 (UTC). See https://pubs.acs.org/sharingguidelines for options on how to legitimately share published articles. w REXAMD: A Hamiltonian
Exploring the anomalous behavior of metal nanocatalysts with finite temperature AIMD and x-ray spectra
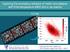 Exploring the anomalous behavior of metal nanocatalysts with finite temperature AIMD and x-ray spectra F.D. Vila DOE grant DE-FG02-03ER15476 With computer support from DOE - NERSC. Importance of Theoretical
Exploring the anomalous behavior of metal nanocatalysts with finite temperature AIMD and x-ray spectra F.D. Vila DOE grant DE-FG02-03ER15476 With computer support from DOE - NERSC. Importance of Theoretical
Steered Molecular Dynamics Introduction and Examples
 Steered Molecular Dynamics Introduction and Examples Klaus Schulten Justin Gullingsrud Hui Lu avidin Sergei Izrailev biotin Rosemary Braun Ferenc Molnar Dorina Kosztin Barry Isralewitz Why Steered Molecular
Steered Molecular Dynamics Introduction and Examples Klaus Schulten Justin Gullingsrud Hui Lu avidin Sergei Izrailev biotin Rosemary Braun Ferenc Molnar Dorina Kosztin Barry Isralewitz Why Steered Molecular
Molecular simulation and structure prediction using CHARMM and the MMTSB Tool Set Free Energy Methods
 Molecular simulation and structure prediction using CHARMM and the MMTSB Tool Set Free Energy Methods Charles L. Brooks III MMTSB/CTBP 2006 Summer Workshop CHARMM Simulations The flow of data and information
Molecular simulation and structure prediction using CHARMM and the MMTSB Tool Set Free Energy Methods Charles L. Brooks III MMTSB/CTBP 2006 Summer Workshop CHARMM Simulations The flow of data and information
Computational Predictions of 1-Octanol/Water Partition Coefficient for Imidazolium based Ionic Liquids.
 Computational Predictions of 1-Octanol/Water Partition Coefficient for Imidazolium based Ionic Liquids. Ganesh Kamath,* a Navendu Bhatnagar b, Gary A. Baker a, Sheila N. Baker c and Jeffrey J. Potoff b
Computational Predictions of 1-Octanol/Water Partition Coefficient for Imidazolium based Ionic Liquids. Ganesh Kamath,* a Navendu Bhatnagar b, Gary A. Baker a, Sheila N. Baker c and Jeffrey J. Potoff b
3. Solutions W = N!/(N A!N B!) (3.1) Using Stirling s approximation ln(n!) = NlnN N: ΔS mix = k (N A lnn + N B lnn N A lnn A N B lnn B ) (3.
 3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/330/6002/341/dc1 Supporting Online Material for Atomic-Level Characterization of the Structural Dynamics of Proteins David E. Shaw,* Paul Maragakis, Kresten Lindorff-Larsen,
www.sciencemag.org/cgi/content/full/330/6002/341/dc1 Supporting Online Material for Atomic-Level Characterization of the Structural Dynamics of Proteins David E. Shaw,* Paul Maragakis, Kresten Lindorff-Larsen,
Integration of Rational Functions by Partial Fractions
 Integration of Rational Functions by Partial Fractions Part 2: Integrating Rational Functions Rational Functions Recall that a rational function is the quotient of two polynomials. x + 3 x + 2 x + 2 x
Integration of Rational Functions by Partial Fractions Part 2: Integrating Rational Functions Rational Functions Recall that a rational function is the quotient of two polynomials. x + 3 x + 2 x + 2 x
Lecture 14: Advanced Conformational Sampling
 Lecture 14: Advanced Conformational Sampling Dr. Ronald M. Levy ronlevy@temple.edu Multidimensional Rough Energy Landscapes MD ~ ns, conformational motion in macromolecules ~µs to sec Interconversions
Lecture 14: Advanced Conformational Sampling Dr. Ronald M. Levy ronlevy@temple.edu Multidimensional Rough Energy Landscapes MD ~ ns, conformational motion in macromolecules ~µs to sec Interconversions
