Free energy calculations
|
|
|
- Neal Heath
- 5 years ago
- Views:
Transcription
1 Free energy calculations Jochen Hub & David van der Spoel
2 Overview Free energies and Probabilities Thermodynamic cycles (Free energy perturbation (FEP)) Thermodynamic integration (TI) (Jarzynski equality Crooks fluctuation theorem) Potential of mean force (PMF)
3 Free Energies and Probabilities Free energies are related to probabilities of states Partition function Z(N,V,T) = d 3N x d 3N p e βe(x,p) Probability to be in state s P s = 1 Z e βe s Helmholtz free energy Normalization with Z probability to be in any state = one A(N,V,T) :=U TS = k B T ln Z(N,V,T)
4 Thermodynamic Cycles Which ligand binds better? What you often want to know: ΔΔG = ΔG1 - ΔG2 Instead of measuring ΔG1 and ΔG2, it is often easier to computationally measure ΔG3 and ΔG4 Alchemical transformation in bulk and in binding site Other important example: How does a mutation destabilize a folded compared to the unfolded protein?
5 Thermodynamic Integration Instead of FEP, better do a TI! Idea: Parameter λ smoothly switches from state A to B H(λ) = (1 λ)h A + λh B G(A B) = 1 0 dλ H(λ) λ λ Switch slowly from λ=0 to λ=1 and integrate the dgdl.xvg output of mdrun. Problem: non-equilibrium/hysteresis effects Choose 20 to 50 discrete λ-steps (init-lambda), sample each λ-step at constant lambda (delta-lambda=0), integrate the averages (discrete TI)
6 Example TI: Solvation of Ethanol Van der Waals Coulomb dh/dλ λ
7 Example TI: Solvation of Ethanol Van der Waals Coulomb Integral of VdW Integral of Coul dh/dλ λ Total DGsolv ~ kj/mol
8 Constrained free energies Constrained free energies. Some observable f constrained to ξ Partition function Z(ξ) = d 3 x d 3N pδ(f(x, p) ξ)e βe(x,p) Z(x 1 )= dy 1 dz 1 dx 2 dx N d N p e βe(x,p) Helmholtz Free Energy A(ξ) = k B T ln Z(ξ) Probability to be in state with f=ξ P (ξ) = Z(ξ) Z = 1 Z e βg(ξ) Relative probabilities of states ξ1 and ξ2 P (ξ 2 )=P (ξ 1 ) e β[a(ξ 2) A(ξ 1 )]
9 Typical reaction coordinates ξ Continuous reaction coordinates Distance between two ions/molecules Ion position along an ion channel Tilting angle of helix in membrane Discrete reaction coordinates ligand bound or unbound alchemical transformation (mutation)
10 Potential of mean force (PMF) The negative derivative of a constrained free energy is the mean force along the reaction coordinate G(a) a G(b) =G(a) = F (a) b a dξ F (ξ) Constrain system at various positions ξ1,...,ξn along reaction coordinate, measure the average force, and integrate the force A potential of mean force is a free energy profile!
11 Potential of mean force (PMF) Alternative: Umbrella sampling From Justin Lemkul s umbrella sampling tutorial (gmx website) Restrain system with harmonic potential wi(ξ) along ξ Check pull_xxx mdp-options in GROMACS generate trajectories (probabilities) referring to V(x)+wi(ξ) Use Weighted Histogram Analysis Method to compute G(ξ) (g_wham)
12 Barriers are related to rates Experimentalists measure rates But: many biological processes hardly ever occur in simulation under equilibrium k = ν e β G
13 Computing PMFs from simulation How do I get a PMF (or free energy profile) from a simulation? Method 1: Simulate very long, get probability along reaction coordinate, compute G(ξ) via G(ξ) = k B T ln P (ξ) Method 2: Constrain system at various positions ξ1,...,ξn along reaction coordinate, measure the average (mean) force, and integrate the mean force Method 3: Restrain (bias) system at various positions ξ1,...,ξn along reaction coordinate using additional harmonic potentials ( umbrealla sampling ) V i (ξ) = 1 2 (ξ ξ i) 2 Compute unbiased probabilities using WHAM. E.g. the RDF of water
14 Umbrella sampling - simple example Unknown PMF Histograms from simulations with restrained system PMF constructed from histograms
15 NH3 permeation across lipid membrane 40 mol% cholesterol
16 Permeation of different molecules through different membranes Pull multiple molecules through in one simulation
17 Permeation of different molecules through different membranes
18 Permeation of different molecules through different membranes Results correlate somewhat with Khex
19 Effect of Cholesterol in Membrane
20 Neopentane in membrane Cholesterol makes membrane rigid Neopentane disrupts favorable Van der Waals interactions in DMPC/Chol
21 Summary Membrane Permeability PMF calculations give Gibbs energy as a function of a coordinate Often these coordinates correspond to a physical variable - e.g. the membrane orthogonal Very useful to extract thermodynamic parameters from MD simulations Very complex relation between phospholid, cholesterol concentration, solute molecule and energetics.
22 Solvation of Ions and Organic Molecules in Water Droplets David van der Spoel
23 Atmospheric chemistry: Surface solubilities influence reactions and processes in air or vacuum.
24 Why do some ions prefer to be on the surface? Alkali ions are always completely solvated F- is solvated Cl-, Br-, I- prefer the surface
25 Simulations Polarizable models from the Roux group for water and ions GROMACS Potential of mean force (PMF) = free energy profile ΔG(r) ΔG = ΔH - TΔS
26 ΔH is related to Etot The total energy in the system, or in case of constant temperature simulations the potential energy is directly related to the enthalpy For a profile we are interested in the relative enthalpy and entropy
27
28 Potential of mean force
29 80 60 A B H(r) [kj/mol] I II III I II III Thermodynamics 0 Minimum at surface T S(r) [kj/mol] C F Cl Br I D Li + Na + K + Rb + Cs + V water-water (r) [kj/mol] E F V ion-water (r) [kj/mol] G H 0,0 0,5 1,0 1,5 r [nm] 0,0 0,5 1,0 1,5 2,0 r [nm]
30 There is no simple structural explanation. Energetics does explain the surface preference. Caleman, Hub,Van Maaren,Van der Spoel, Proc. Natl. Acad. Sci. 108 (2011) p. 6838
31 Organic molecules in droplets:
32 Surface orientation
33 20 PMF [kj mol -1 ] PMF B [kj/mol] ΔH(r) [kj/mol] A TΔS(r) methanol ethanol propanoic acid diethyl ether n-butylamine neopentane All molecules have minimum at the surface C Enthalpic barrier to droplet entry r [nm] Hub, Caleman, Van der Spoel Phys. Chem. Chem. Phys. 14 (2012) pp. 9537
34 Summary Droplets Small to medium droplet and water systems studied by polarizable models It is possible to get detailed energetics (< 4 kj/mol resolution) New results in atmospheric chemistry Sampling is bottleneck
35 Embrace Complexity Simple methods often give incorrect answers - even trends Simulation can give well-founded results within an inherent error margin
36 Acknowledgments Jochen Hub Carl Caleman Paul van Maaren Maarten Wolf
37 Further reading Umbrella sampling introduction B. Roux. The calculation of the potential of mean force using computer simulations. Comp. Phys. Comm., 91: , Free energies from non-equilibrium pulling, excellent summary I. Kosztin, B. Barz, and L. Janosi. Calculating potentials of mean force and diffusion coefficients from nonequilibrium processes without jarzynski s equality. J Chem Phys, 124 pp (2006). Error analysis when using Crooks Götte and Grubmüller, J Comp Chem 30: (2009) WHAM with Bayesian error estimates: Hub et al. J. Chem. Theor. Comput. 6 pp (2010) Wennberg et al. Large Effect of Cholesterol on Membrane Permeability, J. Amer. Chem. Soc. 134 (2012) 5351 Hub et al. Organic molecules on the surface of water droplets - An energetic perspective Phys. Chem. Chem. Phys. 14 pp (2012)
Free energy calculations
 Free energy calculations Berk Hess May 5, 2017 Why do free energy calculations? The free energy G gives the population of states: ( ) P 1 G = exp, G = G 2 G 1 P 2 k B T Since we mostly simulate in the
Free energy calculations Berk Hess May 5, 2017 Why do free energy calculations? The free energy G gives the population of states: ( ) P 1 G = exp, G = G 2 G 1 P 2 k B T Since we mostly simulate in the
Free energy simulations
 Free energy simulations Marcus Elstner and Tomáš Kubař January 14, 2013 Motivation a physical quantity that is of most interest in chemistry? free energies Helmholtz F or Gibbs G holy grail of computational
Free energy simulations Marcus Elstner and Tomáš Kubař January 14, 2013 Motivation a physical quantity that is of most interest in chemistry? free energies Helmholtz F or Gibbs G holy grail of computational
Biomolecular modeling. Theoretical Chemistry, TU Braunschweig (Dated: December 10, 2010)
 Biomolecular modeling Marcus Elstner and Tomáš Kubař Theoretical Chemistry, TU Braunschweig (Dated: December 10, 2010) IX. FREE ENERGY SIMULATIONS When searching for a physical quantity that is of most
Biomolecular modeling Marcus Elstner and Tomáš Kubař Theoretical Chemistry, TU Braunschweig (Dated: December 10, 2010) IX. FREE ENERGY SIMULATIONS When searching for a physical quantity that is of most
Lecture 19: Free Energies in Modern Computational Statistical Thermodynamics: FEP and Related Methods
 Statistical Thermodynamics Lecture 19: Free Energies in Modern Computational Statistical Thermodynamics: FEP and Related Methods Dr. Ronald M. Levy ronlevy@temple.edu Free energy calculations Free energy
Statistical Thermodynamics Lecture 19: Free Energies in Modern Computational Statistical Thermodynamics: FEP and Related Methods Dr. Ronald M. Levy ronlevy@temple.edu Free energy calculations Free energy
Free energy calculations using molecular dynamics simulations. Anna Johansson
 Free energy calculations using molecular dynamics simulations Anna Johansson 2007-03-13 Outline Introduction to concepts Why is free energy important? Calculating free energy using MD Thermodynamical Integration
Free energy calculations using molecular dynamics simulations Anna Johansson 2007-03-13 Outline Introduction to concepts Why is free energy important? Calculating free energy using MD Thermodynamical Integration
Free energy calculations with alchemlyb
 Free energy calculations with alchemlyb Oliver Beckstein Arizona State University SPIDAL Teleconference 2019-02-01 Binding free energy measure of how strong a protein P and a ligand X stick together key
Free energy calculations with alchemlyb Oliver Beckstein Arizona State University SPIDAL Teleconference 2019-02-01 Binding free energy measure of how strong a protein P and a ligand X stick together key
From Dynamics to Thermodynamics using Molecular Simulation
 From Dynamics to Thermodynamics using Molecular Simulation David van der Spoel Computational Chemistry Physical models to describe molecules Software to evaluate models and do predictions - GROMACS Model
From Dynamics to Thermodynamics using Molecular Simulation David van der Spoel Computational Chemistry Physical models to describe molecules Software to evaluate models and do predictions - GROMACS Model
Chem 112 Dr. Kevin Moore
 Chem 112 Dr. Kevin Moore Gas Liquid Solid Polar Covalent Bond Partial Separation of Charge Electronegativity: H 2.1 Cl 3.0 H Cl δ + δ - Dipole Moment measure of the net polarity in a molecule Q Q magnitude
Chem 112 Dr. Kevin Moore Gas Liquid Solid Polar Covalent Bond Partial Separation of Charge Electronegativity: H 2.1 Cl 3.0 H Cl δ + δ - Dipole Moment measure of the net polarity in a molecule Q Q magnitude
3. Solutions W = N!/(N A!N B!) (3.1) Using Stirling s approximation ln(n!) = NlnN N: ΔS mix = k (N A lnn + N B lnn N A lnn A N B lnn B ) (3.
 3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
AP CHEMISTRY 2007 SCORING GUIDELINES (Form B)
 AP CHEMISTRY 2007 SCORING GUIDELINES (Form B) Question 1 A sample of solid U O 8 is placed in a rigid 1.500 L flask. Chlorine gas, Cl 2 (g), is added, and the flask is heated to 862 C. The equation for
AP CHEMISTRY 2007 SCORING GUIDELINES (Form B) Question 1 A sample of solid U O 8 is placed in a rigid 1.500 L flask. Chlorine gas, Cl 2 (g), is added, and the flask is heated to 862 C. The equation for
Modeling the Free Energy Landscape for Janus Particle Self-Assembly in the Gas Phase. Andy Long Kridsanaphong Limtragool
 Modeling the Free Energy Landscape for Janus Particle Self-Assembly in the Gas Phase Andy Long Kridsanaphong Limtragool Motivation We want to study the spontaneous formation of micelles and vesicles Applications
Modeling the Free Energy Landscape for Janus Particle Self-Assembly in the Gas Phase Andy Long Kridsanaphong Limtragool Motivation We want to study the spontaneous formation of micelles and vesicles Applications
= (-22) = +2kJ /mol
 Lecture 8: Thermodynamics & Protein Stability Assigned reading in Campbell: Chapter 4.4-4.6 Key Terms: DG = -RT lnk eq = DH - TDS Transition Curve, Melting Curve, Tm DH calculation DS calculation van der
Lecture 8: Thermodynamics & Protein Stability Assigned reading in Campbell: Chapter 4.4-4.6 Key Terms: DG = -RT lnk eq = DH - TDS Transition Curve, Melting Curve, Tm DH calculation DS calculation van der
MCB100A/Chem130 MidTerm Exam 2 April 4, 2013
 MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Statistical Mechanics for Proteins
 The Partition Function From Q all relevant thermodynamic properties can be obtained by differentiation of the free energy F: = kt q p E q pd d h T V Q ), ( exp 1! 1 ),, ( 3 3 3 ),, ( ln ),, ( T V Q kt
The Partition Function From Q all relevant thermodynamic properties can be obtained by differentiation of the free energy F: = kt q p E q pd d h T V Q ), ( exp 1! 1 ),, ( 3 3 3 ),, ( ln ),, ( T V Q kt
Chapter 1. Topic: Overview of basic principles
 Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Entropy and Free Energy in Biology
 Entropy and Free Energy in Biology Energy vs. length from Phillips, Quake. Physics Today. 59:38-43, 2006. kt = 0.6 kcal/mol = 2.5 kj/mol = 25 mev typical protein typical cell Thermal effects = deterministic
Entropy and Free Energy in Biology Energy vs. length from Phillips, Quake. Physics Today. 59:38-43, 2006. kt = 0.6 kcal/mol = 2.5 kj/mol = 25 mev typical protein typical cell Thermal effects = deterministic
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Free Energy Simulation Methods
 Free Energy Simulation Methods Free energy simulation methods Many methods have been developed to compute (relative) free energies on the basis of statistical mechanics Free energy perturbation Thermodynamic
Free Energy Simulation Methods Free energy simulation methods Many methods have been developed to compute (relative) free energies on the basis of statistical mechanics Free energy perturbation Thermodynamic
Entropy. Spontaneity. Entropy. Entropy mol of N 2 at 1 atm or 1 mol of N 2 at atm. process a process that occurs without intervention
 Entropy Spontaneity process a process that occurs without intervention can be fast or slow Entropy (s) the measure of molecular randomness or disorder Think of entropy as the amount of chaos Entropy Predict
Entropy Spontaneity process a process that occurs without intervention can be fast or slow Entropy (s) the measure of molecular randomness or disorder Think of entropy as the amount of chaos Entropy Predict
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
MCB100A/Chem130 MidTerm Exam 2 April 4, 2013
 MCB1A/Chem13 MidTerm Exam 2 April 4, 213 Name Student ID True/False (2 points each). 1. The Boltzmann constant, k b T sets the energy scale for observing energy microstates 2. Atoms with favorable electronic
MCB1A/Chem13 MidTerm Exam 2 April 4, 213 Name Student ID True/False (2 points each). 1. The Boltzmann constant, k b T sets the energy scale for observing energy microstates 2. Atoms with favorable electronic
Lecture 20. Chemical Potential
 Lecture 20 Chemical Potential Reading: Lecture 20, today: Chapter 10, sections A and B Lecture 21, Wednesday: Chapter 10: 10 17 end 3/21/16 1 Pop Question 7 Boltzmann Distribution Two systems with lowest
Lecture 20 Chemical Potential Reading: Lecture 20, today: Chapter 10, sections A and B Lecture 21, Wednesday: Chapter 10: 10 17 end 3/21/16 1 Pop Question 7 Boltzmann Distribution Two systems with lowest
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Measuring The Binding Energy Of Glucose To The Glucose/Galactose Binding Protein Computationally
 Proceedings of The National Conference On Undergraduate Research (NCUR) 2017 University of Memphis, TN April 7-9, 2017 Measuring The Binding Energy Of Glucose To The Glucose/Galactose Binding Protein Computationally
Proceedings of The National Conference On Undergraduate Research (NCUR) 2017 University of Memphis, TN April 7-9, 2017 Measuring The Binding Energy Of Glucose To The Glucose/Galactose Binding Protein Computationally
BIOC : Homework 1 Due 10/10
 Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Second Law of Thermodynamics
 Second Law of Thermodynamics First Law: the total energy of the universe is a constant Second Law: The entropy of the universe increases in a spontaneous process, and remains unchanged in a process at
Second Law of Thermodynamics First Law: the total energy of the universe is a constant Second Law: The entropy of the universe increases in a spontaneous process, and remains unchanged in a process at
7/19/2011. Models of Solution. State of Equilibrium. State of Equilibrium Chemical Reaction
 Models of Solution Chemistry- I State of Equilibrium A covered cup of coffee will not be colder than or warmer than the room temperature Heat is defined as a form of energy that flows from a high temperature
Models of Solution Chemistry- I State of Equilibrium A covered cup of coffee will not be colder than or warmer than the room temperature Heat is defined as a form of energy that flows from a high temperature
Entropy and Free Energy in Biology
 Entropy and Free Energy in Biology Energy vs. length from Phillips, Quake. Physics Today. 59:38-43, 2006. kt = 0.6 kcal/mol = 2.5 kj/mol = 25 mev typical protein typical cell Thermal effects = deterministic
Entropy and Free Energy in Biology Energy vs. length from Phillips, Quake. Physics Today. 59:38-43, 2006. kt = 0.6 kcal/mol = 2.5 kj/mol = 25 mev typical protein typical cell Thermal effects = deterministic
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
a) Write the reaction that occurs (pay attention to and label ends correctly) 5 AGCTG CAGCT > 5 AGCTG 3 3 TCGAC 5
 Chem 315 Applications Practice Problem Set 1.As you learned in Chem 315, DNA higher order structure formation is a two step process. The first step, nucleation, is entropically the least favorable. The
Chem 315 Applications Practice Problem Set 1.As you learned in Chem 315, DNA higher order structure formation is a two step process. The first step, nucleation, is entropically the least favorable. The
The PLUMED plugin and free energy methods in electronic-structure-based molecular dynamics
 The PLUMED plugin and free energy methods in electronic-structure-based molecular dynamics Davide Branduardi, Theoretical Molecular Biophysics Group Max Planck for Biophysics, Frankfurt am Main, Germany
The PLUMED plugin and free energy methods in electronic-structure-based molecular dynamics Davide Branduardi, Theoretical Molecular Biophysics Group Max Planck for Biophysics, Frankfurt am Main, Germany
Some Accelerated/Enhanced Sampling Techniques
 Some Accelerated/Enhanced Sampling Techniques The Sampling Problem Accelerated sampling can be achieved by addressing one or more of the MD ingredients Back to free energy differences... Examples of accelerated
Some Accelerated/Enhanced Sampling Techniques The Sampling Problem Accelerated sampling can be achieved by addressing one or more of the MD ingredients Back to free energy differences... Examples of accelerated
Alchemical free energy calculations in OpenMM
 Alchemical free energy calculations in OpenMM Lee-Ping Wang Stanford Department of Chemistry OpenMM Workshop, Stanford University September 7, 2012 Special thanks to: John Chodera, Morgan Lawrenz Outline
Alchemical free energy calculations in OpenMM Lee-Ping Wang Stanford Department of Chemistry OpenMM Workshop, Stanford University September 7, 2012 Special thanks to: John Chodera, Morgan Lawrenz Outline
Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
 Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Unit 12. Thermochemistry
 Unit 12 Thermochemistry A reaction is spontaneous if it will occur without a continuous input of energy However, it may require an initial input of energy to get it started (activation energy) For Thermochemistry
Unit 12 Thermochemistry A reaction is spontaneous if it will occur without a continuous input of energy However, it may require an initial input of energy to get it started (activation energy) For Thermochemistry
Chemistry and the material world Unit 4, Lecture 4 Matthias Lein
 Chemistry and the material world 123.102 Unit 4, Lecture 4 Matthias Lein Gibbs ree energy Gibbs ree energy to predict the direction o a chemical process. Exergonic and endergonic reactions. Temperature
Chemistry and the material world 123.102 Unit 4, Lecture 4 Matthias Lein Gibbs ree energy Gibbs ree energy to predict the direction o a chemical process. Exergonic and endergonic reactions. Temperature
Phase Equilibria and Molecular Solutions Jan G. Korvink and Evgenii Rudnyi IMTEK Albert Ludwig University Freiburg, Germany
 Phase Equilibria and Molecular Solutions Jan G. Korvink and Evgenii Rudnyi IMTEK Albert Ludwig University Freiburg, Germany Preliminaries Learning Goals Phase Equilibria Phase diagrams and classical thermodynamics
Phase Equilibria and Molecular Solutions Jan G. Korvink and Evgenii Rudnyi IMTEK Albert Ludwig University Freiburg, Germany Preliminaries Learning Goals Phase Equilibria Phase diagrams and classical thermodynamics
Computing free energies with PLUMED 2.0
 Computing free energies with PLUMED 2.0 Davide Branduardi Formerly at MPI Biophysics, Frankfurt a.m. TyOutline of the talk Relevance of free energy computation Free-energy from histograms: issues Tackling
Computing free energies with PLUMED 2.0 Davide Branduardi Formerly at MPI Biophysics, Frankfurt a.m. TyOutline of the talk Relevance of free energy computation Free-energy from histograms: issues Tackling
A general overview of. Free energy methods. Comer et al. (2014) JPCB 119:1129. James C. (JC) Gumbart Georgia Institute of Technology, Atlanta
 A general overview of Free energy methods Comer et al. (2014) JPCB 119:1129. James C. (JC) Gumbart Georgia Institute of Technology, Atlanta Computational Biophysics Workshop Georgia Tech Nov. 16 2016 Outline
A general overview of Free energy methods Comer et al. (2014) JPCB 119:1129. James C. (JC) Gumbart Georgia Institute of Technology, Atlanta Computational Biophysics Workshop Georgia Tech Nov. 16 2016 Outline
MARTINI simulation details
 S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
Lecture Notes 1: Physical Equilibria Vapor Pressure
 Lecture Notes 1: Physical Equilibria Vapor Pressure Our first exploration of equilibria will examine physical equilibria (no chemical changes) in which the only changes occurring are matter changes phases.
Lecture Notes 1: Physical Equilibria Vapor Pressure Our first exploration of equilibria will examine physical equilibria (no chemical changes) in which the only changes occurring are matter changes phases.
Thermodynamics: Free Energy and Entropy. Suggested Reading: Chapter 19
 Thermodynamics: Free Energy and Entropy Suggested Reading: Chapter 19 System and Surroundings System: An object or collection of objects being studied. Surroundings: Everything outside of the system. the
Thermodynamics: Free Energy and Entropy Suggested Reading: Chapter 19 System and Surroundings System: An object or collection of objects being studied. Surroundings: Everything outside of the system. the
W tot (z) = W (z) k B T ln e i q iφ mp(z i )/k B T, [1]
![W tot (z) = W (z) k B T ln e i q iφ mp(z i )/k B T, [1] W tot (z) = W (z) k B T ln e i q iφ mp(z i )/k B T, [1]](/thumbs/87/95587399.jpg) Supporting Text Simulation Details. Simulations were carried out with the program charmm (1) using the PARAM27 force field with standard lipid (2), protein (3), and TIP3P water model (4) parameters. The
Supporting Text Simulation Details. Simulations were carried out with the program charmm (1) using the PARAM27 force field with standard lipid (2), protein (3), and TIP3P water model (4) parameters. The
This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail.
 This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Author(s): Hub, Jochen S.; Groot, Bert L. De; Grubmüller, Helmut;
This is an electronic reprint of the original article. This reprint may differ from the original in pagination and typographic detail. Author(s): Hub, Jochen S.; Groot, Bert L. De; Grubmüller, Helmut;
Chemistry 201. Working with K. NC State University. Lecture 11
 Chemistry 201 Lecture 11 Working with K NC State University Working With K What is the relationship between pressure and concentration in K? How does one calculate K or components of K? How does one calculate
Chemistry 201 Lecture 11 Working with K NC State University Working With K What is the relationship between pressure and concentration in K? How does one calculate K or components of K? How does one calculate
SCORING. The exam consists of 5 questions totaling 100 points as broken down in this table:
 UNIVERSITY OF CALIFORNIA, BERKELEY CHEM C130/MCB C100A MIDTERM EXAMINATION #2 OCTOBER 20, 2016 INSTRUCTORS: John Kuriyan and David Savage THE TIME LIMIT FOR THIS EXAMINATION: 1 HOUR 50 MINUTES SIGNATURE:
UNIVERSITY OF CALIFORNIA, BERKELEY CHEM C130/MCB C100A MIDTERM EXAMINATION #2 OCTOBER 20, 2016 INSTRUCTORS: John Kuriyan and David Savage THE TIME LIMIT FOR THIS EXAMINATION: 1 HOUR 50 MINUTES SIGNATURE:
Thermochemistry Chapter 8
 Thermochemistry Chapter 8 Thermochemistry First law of thermochemistry: Internal energy of an isolated system is constant; energy cannot be created or destroyed; however, energy can be converted to different
Thermochemistry Chapter 8 Thermochemistry First law of thermochemistry: Internal energy of an isolated system is constant; energy cannot be created or destroyed; however, energy can be converted to different
Chemical Thermodynamics. Chapter 18
 Chemical Thermodynamics Chapter 18 Thermodynamics Spontaneous Processes Entropy and Second Law of Thermodynamics Entropy Changes Gibbs Free Energy Free Energy and Temperature Free Energy and Equilibrium
Chemical Thermodynamics Chapter 18 Thermodynamics Spontaneous Processes Entropy and Second Law of Thermodynamics Entropy Changes Gibbs Free Energy Free Energy and Temperature Free Energy and Equilibrium
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Gibbs Free Energy. Evaluating spontaneity
 Gibbs Free Energy Evaluating spontaneity Predicting Spontaneity An increase in entropy; Changing from a more structured to less structured physical state: Solid to liquid Liquid to gas Increase in temperature
Gibbs Free Energy Evaluating spontaneity Predicting Spontaneity An increase in entropy; Changing from a more structured to less structured physical state: Solid to liquid Liquid to gas Increase in temperature
Biological Thermodynamics
 Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Free energy, electrostatics, and the hydrophobic effect
 Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Computational Studies of the Photoreceptor Rhodopsin. Scott E. Feller Wabash College
 Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
Computational Studies of the Photoreceptor Rhodopsin Scott E. Feller Wabash College Rhodopsin Photocycle Dark-adapted Rhodopsin hn Isomerize retinal Photorhodopsin ~200 fs Bathorhodopsin Meta-II ms timescale
= (+)206 (kj mol 1) 206 scores 1 only Units not essential if ans in kj mol 1 but penalise incorrect units
 M.(a) (i) ΔH = Σ(enthalpies formation products) Σ(enthalpies formation reactants) Or correct cycle with enthalpy changes labelled = ( 75 242) = (+)206 (kj mol ) 206 scores only Units not essential if ans
M.(a) (i) ΔH = Σ(enthalpies formation products) Σ(enthalpies formation reactants) Or correct cycle with enthalpy changes labelled = ( 75 242) = (+)206 (kj mol ) 206 scores only Units not essential if ans
A proposed mechanism for the decomposition of hydrogen peroxide by iodide ion is: slow fast (D) H 2 O
 Chemistry 112, Spring 2007 Prof. Metz Exam 2 Practice Use the following information to answer questions 1 through 3 A proposed mechanism for the decomposition of hydrogen peroxide by iodide ion is: H 2
Chemistry 112, Spring 2007 Prof. Metz Exam 2 Practice Use the following information to answer questions 1 through 3 A proposed mechanism for the decomposition of hydrogen peroxide by iodide ion is: H 2
Orthogonal Space Sampling of Slow Environment Responses
 IMA University of Minnesota 2015 Orthogonal Space Sampling of Slow Environment Responses Lianqing Zheng, Chao Lv, Dongsheng Wu, William Harris, Xubin Li, Erick Aitchison, and Wei Yang Institute of Molecular
IMA University of Minnesota 2015 Orthogonal Space Sampling of Slow Environment Responses Lianqing Zheng, Chao Lv, Dongsheng Wu, William Harris, Xubin Li, Erick Aitchison, and Wei Yang Institute of Molecular
arxiv: v1 [physics.chem-ph] 30 Jul 2012
![arxiv: v1 [physics.chem-ph] 30 Jul 2012 arxiv: v1 [physics.chem-ph] 30 Jul 2012](/thumbs/92/110504311.jpg) Is ion channel selectivity mediated by confined water? Diego Prada-Gracia and Francesco Rao School of Soft Matter, Freiburg Institute for Advanced Studies, Freiburg, Germany arxiv:27.6953v [physics.chem-ph]
Is ion channel selectivity mediated by confined water? Diego Prada-Gracia and Francesco Rao School of Soft Matter, Freiburg Institute for Advanced Studies, Freiburg, Germany arxiv:27.6953v [physics.chem-ph]
q 2 Da This looks very similar to Coulomb doesn t it? The only difference to Coulomb is the factor of 1/2.
 Born Lets now think about a somewhat different, but related problem that is associated with the name Born. Using a name (usually the name of the guy who first discovered or solved the problem) to describe
Born Lets now think about a somewhat different, but related problem that is associated with the name Born. Using a name (usually the name of the guy who first discovered or solved the problem) to describe
Chapter 2 Experimental sources of intermolecular potentials
 Chapter 2 Experimental sources of intermolecular potentials 2.1 Overview thermodynamical properties: heat of vaporization (Trouton s rule) crystal structures ionic crystals rare gas solids physico-chemical
Chapter 2 Experimental sources of intermolecular potentials 2.1 Overview thermodynamical properties: heat of vaporization (Trouton s rule) crystal structures ionic crystals rare gas solids physico-chemical
Topic 3 Periodicity 3.2 Physical Properties. IB Chemistry T03D02
 Topic 3 Periodicity 3.2 Physical Properties IB Chemistry T03D02 3.1 Physical Properties hrs 3.2.1 Define the terms first ionization energy and electronegativity. (1) 3.2.2 Describe and explain the trends
Topic 3 Periodicity 3.2 Physical Properties IB Chemistry T03D02 3.1 Physical Properties hrs 3.2.1 Define the terms first ionization energy and electronegativity. (1) 3.2.2 Describe and explain the trends
5th CCPN Matt Crump. Thermodynamic quantities derived from protein dynamics
 5th CCPN 2005 -Matt Crump Thermodynamic quantities derived from protein dynamics Relaxation in Liquids (briefly!) The fluctuations of each bond vector can be described in terms of an angular correlation
5th CCPN 2005 -Matt Crump Thermodynamic quantities derived from protein dynamics Relaxation in Liquids (briefly!) The fluctuations of each bond vector can be described in terms of an angular correlation
Free-energy change ( G) and entropy change ( S)
 Free-energy change ( G) and entropy change ( S) A SPONTANEOUS PROCESS (e.g. diffusion) will proceed on its own without any external influence. A problem with H A reaction that is exothermic will result
Free-energy change ( G) and entropy change ( S) A SPONTANEOUS PROCESS (e.g. diffusion) will proceed on its own without any external influence. A problem with H A reaction that is exothermic will result
Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality
 JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 6 8 AUGUST 2003 Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality Sanghyun Park and Fatemeh Khalili-Araghi Beckman
JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 6 8 AUGUST 2003 Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality Sanghyun Park and Fatemeh Khalili-Araghi Beckman
Protein-Ligand Interactions* and Energy Evaluation Methods
 Protein-Ligand Interactions* and Energy Evaluation Methods *with a revealing look at roles of water Glen E. Kellogg Department of Medicinal Chemistry Institute for Structural Biology, Drug Discovery &
Protein-Ligand Interactions* and Energy Evaluation Methods *with a revealing look at roles of water Glen E. Kellogg Department of Medicinal Chemistry Institute for Structural Biology, Drug Discovery &
Free energy calculations and the potential of mean force
 Free energy calculations and the potential of mean force IMA Workshop on Classical and Quantum Approaches in Molecular Modeling Mark Tuckerman Dept. of Chemistry and Courant Institute of Mathematical Science
Free energy calculations and the potential of mean force IMA Workshop on Classical and Quantum Approaches in Molecular Modeling Mark Tuckerman Dept. of Chemistry and Courant Institute of Mathematical Science
Second law of thermodynamics
 Second law of thermodynamics It is known from everyday life that nature does the most probable thing when nothing prevents that For example it rains at cool weather because the liquid phase has less energy
Second law of thermodynamics It is known from everyday life that nature does the most probable thing when nothing prevents that For example it rains at cool weather because the liquid phase has less energy
Some properties of water
 Some properties of water Hydrogen bond network Solvation under the microscope 1 Water solutions Oil and water does not mix at equilibrium essentially due to entropy Substances that does not mix with water
Some properties of water Hydrogen bond network Solvation under the microscope 1 Water solutions Oil and water does not mix at equilibrium essentially due to entropy Substances that does not mix with water
UC Berkeley. Chem 130A. Spring nd Exam. March 10, 2004 Instructor: John Kuriyan
 UC Berkeley. Chem 130A. Spring 2004 2nd Exam. March 10, 2004 Instructor: John Kuriyan (kuriyan@uclink.berkeley.edu) Enter your name & student ID number above the line, in ink. Sign your name above the
UC Berkeley. Chem 130A. Spring 2004 2nd Exam. March 10, 2004 Instructor: John Kuriyan (kuriyan@uclink.berkeley.edu) Enter your name & student ID number above the line, in ink. Sign your name above the
Liquid-vapour oscillations of water in hydrophobic nanopores
 Alicante 2003 4th European Biophysics Congress Liquid-vapour oscillations of water in hydrophobic nanopores 7 th July 2003 Oliver Beckstein and Mark S. P. Sansom Department of Biochemistry, Laboratory
Alicante 2003 4th European Biophysics Congress Liquid-vapour oscillations of water in hydrophobic nanopores 7 th July 2003 Oliver Beckstein and Mark S. P. Sansom Department of Biochemistry, Laboratory
The Second Law of Thermodynamics (Chapter 4)
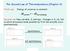 The Second Law of Thermodynamics (Chapter 4) First Law: Energy of universe is constant: ΔE system = - ΔE surroundings Second Law: New variable, S, entropy. Changes in S, ΔS, tell us which processes made
The Second Law of Thermodynamics (Chapter 4) First Law: Energy of universe is constant: ΔE system = - ΔE surroundings Second Law: New variable, S, entropy. Changes in S, ΔS, tell us which processes made
When does TMAO fold a polymer chain and urea unfold it?
 When does TMAO fold a polymer chain and urea unfold it? Jagannath Mondal, Guillaume Stirnemann, and B. J. Berne Department of Chemistry, Columbia University, New York, NY 7 (Dated: March 1, 1) Other studies
When does TMAO fold a polymer chain and urea unfold it? Jagannath Mondal, Guillaume Stirnemann, and B. J. Berne Department of Chemistry, Columbia University, New York, NY 7 (Dated: March 1, 1) Other studies
Chapters 11 and 12: Intermolecular Forces of Liquids and Solids
 1 Chapters 11 and 12: Intermolecular Forces of Liquids and Solids 11.1 A Molecular Comparison of Liquids and Solids The state of matter (Gas, liquid or solid) at a particular temperature and pressure depends
1 Chapters 11 and 12: Intermolecular Forces of Liquids and Solids 11.1 A Molecular Comparison of Liquids and Solids The state of matter (Gas, liquid or solid) at a particular temperature and pressure depends
OCR Chemistry A H432
 All the energy changes we have considered so far have been in terms of enthalpy, and we have been able to predict whether a reaction is likely to occur on the basis of the enthalpy change associated with
All the energy changes we have considered so far have been in terms of enthalpy, and we have been able to predict whether a reaction is likely to occur on the basis of the enthalpy change associated with
Aspects of nonautonomous molecular dynamics
 Aspects of nonautonomous molecular dynamics IMA, University of Minnesota, Minneapolis January 28, 2007 Michel Cuendet Swiss Institute of Bioinformatics, Lausanne, Switzerland Introduction to the Jarzynski
Aspects of nonautonomous molecular dynamics IMA, University of Minnesota, Minneapolis January 28, 2007 Michel Cuendet Swiss Institute of Bioinformatics, Lausanne, Switzerland Introduction to the Jarzynski
Name AP CHEM / / Collected AP Exam Essay Answers for Chapter 16
 Name AP CHEM / / Collected AP Exam Essay Answers for Chapter 16 1980 - #7 (a) State the physical significance of entropy. Entropy (S) is a measure of randomness or disorder in a system. (b) From each of
Name AP CHEM / / Collected AP Exam Essay Answers for Chapter 16 1980 - #7 (a) State the physical significance of entropy. Entropy (S) is a measure of randomness or disorder in a system. (b) From each of
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Fundamentals of Distribution Separations (III)
 Fundamentals of Distribution Separations (III) (01/16/15) K = exp -Δμ 0 ext i - Δμ i RT distribution coefficient C i = exp -Δμ 0 RT - - Δμ i = ΔH i TΔS i 0 0 0 solubility q A---B A + B 0 0 0 ΔH i = ΔH
Fundamentals of Distribution Separations (III) (01/16/15) K = exp -Δμ 0 ext i - Δμ i RT distribution coefficient C i = exp -Δμ 0 RT - - Δμ i = ΔH i TΔS i 0 0 0 solubility q A---B A + B 0 0 0 ΔH i = ΔH
1 What is energy?
 http://www.intothecool.com/ 1 What is energy? the capacity to do work? (Greek: en-, in; + ergon, work) the capacity to cause change to produce an effect? a certain quantity that does not change in the
http://www.intothecool.com/ 1 What is energy? the capacity to do work? (Greek: en-, in; + ergon, work) the capacity to cause change to produce an effect? a certain quantity that does not change in the
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
ONETEP PB/SA: Application to G-Quadruplex DNA Stability. Danny Cole
 ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
ONETEP PB/SA: Application to G-Quadruplex DNA Stability Danny Cole Introduction Historical overview of structure and free energy calculation of complex molecules using molecular mechanics and continuum
FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION
 PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 08, 5(), p. 75 79 FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION C h emistr y H. A. SHILAJYAN, K. R. GRIGORYAN
PROCEEDINGS OF THE YEREVAN STATE UNIVERSITY C h e m i s t r y a n d B i o l o g y 08, 5(), p. 75 79 FLUORESCENCE STUDY ON TRYPTOPHAN POTASSIUM IODIDE INTERACTION C h emistr y H. A. SHILAJYAN, K. R. GRIGORYAN
6 Hydrophobic interactions
 The Physics and Chemistry of Water 6 Hydrophobic interactions A non-polar molecule in water disrupts the H- bond structure by forcing some water molecules to give up their hydrogen bonds. As a result,
The Physics and Chemistry of Water 6 Hydrophobic interactions A non-polar molecule in water disrupts the H- bond structure by forcing some water molecules to give up their hydrogen bonds. As a result,
Lattice energy of ionic solids
 1 Lattice energy of ionic solids Interatomic Forces Solids are aggregates of atoms, ions or molecules. The bonding between these particles may be understood in terms of forces that play between them. Attractive
1 Lattice energy of ionic solids Interatomic Forces Solids are aggregates of atoms, ions or molecules. The bonding between these particles may be understood in terms of forces that play between them. Attractive
THERMODYNAMICS I. TERMS AND DEFINITIONS A. Review of Definitions 1. Thermodynamics = Study of the exchange of heat, energy and work between a system
 THERMODYNAMICS I. TERMS AND DEFINITIONS A. Review of Definitions 1. Thermodynamics = Study of the exchange of heat, energy and work between a system and its surroundings. a. System = That part of universe
THERMODYNAMICS I. TERMS AND DEFINITIONS A. Review of Definitions 1. Thermodynamics = Study of the exchange of heat, energy and work between a system and its surroundings. a. System = That part of universe
2/18/2013. Spontaneity, Entropy & Free Energy Chapter 16
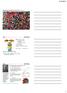 Spontaneity, Entropy & Chapter 16 G = H T S G is Gibbs free energy H is enthalpy T is temperture in Kelvin S is entropy Refers to the system Proved on p. 760 from S = H/T S univ = - G T at constant T &
Spontaneity, Entropy & Chapter 16 G = H T S G is Gibbs free energy H is enthalpy T is temperture in Kelvin S is entropy Refers to the system Proved on p. 760 from S = H/T S univ = - G T at constant T &
DSC Characterization of the Structure/Function Relationship for Proteins
 DSC Characterization of the Structure/Function Relationship for Proteins Differential Scanning Calorimetry (DSC) DSC is recognized as Gold Std technique for measuring molecular thermal stability and structure
DSC Characterization of the Structure/Function Relationship for Proteins Differential Scanning Calorimetry (DSC) DSC is recognized as Gold Std technique for measuring molecular thermal stability and structure
Do now: Write equations for the following expressions:
 Do now: Write equations for the following expressions: ½ N 2(g) + 1 ½ H 2(g) NH 3(g) Δ f H (NH 3(g) ) = -46 kj.mol -1 ½ H 2(g) + ½ Cl 2(g) HCl (g) Δ f H (HCl (g) ) = -92 kj.mol -1 ½ N 2(g) + 2 H 2(g) +
Do now: Write equations for the following expressions: ½ N 2(g) + 1 ½ H 2(g) NH 3(g) Δ f H (NH 3(g) ) = -46 kj.mol -1 ½ H 2(g) + ½ Cl 2(g) HCl (g) Δ f H (HCl (g) ) = -92 kj.mol -1 ½ N 2(g) + 2 H 2(g) +
Advanced in silico drug design
 Advanced in silico drug design RNDr. Martin Lepšík, Ph.D. Lecture: Advanced scoring Palacky University, Olomouc 2016 1 Outline 1. Scoring Definition, Types 2. Physics-based Scoring: Master Equation Terms
Advanced in silico drug design RNDr. Martin Lepšík, Ph.D. Lecture: Advanced scoring Palacky University, Olomouc 2016 1 Outline 1. Scoring Definition, Types 2. Physics-based Scoring: Master Equation Terms
Exp.3 Determination of the Thermodynamic functions for the Borax Solution
 Exp.3 Determination of the Thermodynamic functions for the Borax Solution Theory: The relationship between Gibb s energy (ΔG), Enthalpy (ΔH), Entropy (ΔS) and the equilibrium constant (K) for a chemical
Exp.3 Determination of the Thermodynamic functions for the Borax Solution Theory: The relationship between Gibb s energy (ΔG), Enthalpy (ΔH), Entropy (ΔS) and the equilibrium constant (K) for a chemical
The change in free energy on transferring an ion from a medium of low dielectric constantε1 to one of high dielectric constant ε2:
 The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
CHEM Exam 3 - March 31, 2017
 CHEM 3530 - Exam 3 - March 31, 2017 Constants and Conversion Factors NA = 6.02x10 23 mol -1 R = 8.31 J/mol-K = 8.31 kpa-l/mol-k 1 bar = 100 kpa = 750 torr 1 kpa = 7.50 torr 1 J = 1 kpa-l 1 kcal = 4.18
CHEM 3530 - Exam 3 - March 31, 2017 Constants and Conversion Factors NA = 6.02x10 23 mol -1 R = 8.31 J/mol-K = 8.31 kpa-l/mol-k 1 bar = 100 kpa = 750 torr 1 kpa = 7.50 torr 1 J = 1 kpa-l 1 kcal = 4.18
Microcalorimetry for the Life Sciences
 Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Electronic Supplementary Information Effective lead optimization targeted for displacing bridging water molecule
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Effective lead optimization targeted for displacing
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Effective lead optimization targeted for displacing
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro. Karel Berka, Ph.D.
 4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
4 th Advanced in silico Drug Design KFC/ADD Molecular Modelling Intro Karel Berka, Ph.D. UP Olomouc, 21.1.-25.1. 2019 Motto A theory is something nobody believes, except the person who made it An experiment
Hydrophobic and Ionic Interactions in Nanosized Water Droplets
 Published on Web 09/23/2006 Hydrophobic and Ionic Interactions in Nanosized Water Droplets S. Vaitheeswaran and D. Thirumalai*,, Contribution from the Biophysics Program, Institute for Physical Science
Published on Web 09/23/2006 Hydrophobic and Ionic Interactions in Nanosized Water Droplets S. Vaitheeswaran and D. Thirumalai*,, Contribution from the Biophysics Program, Institute for Physical Science
Chapter 2 - Water 9/8/2014. Water exists as a H-bonded network with an average of 4 H-bonds per molecule in ice and 3.4 in liquid. 104.
 Chapter 2 - Water Water exists as a -bonded network with an average of 4 -bonds per molecule in ice and 3.4 in liquid. 104.5 o -bond: An electrostatic attraction between polarized molecules containing
Chapter 2 - Water Water exists as a -bonded network with an average of 4 -bonds per molecule in ice and 3.4 in liquid. 104.5 o -bond: An electrostatic attraction between polarized molecules containing
Unit 5: Spontaneity of Reaction. You need to bring your textbooks everyday of this unit.
 Unit 5: Spontaneity of Reaction You need to bring your textbooks everyday of this unit. THE LAWS OF THERMODYNAMICS 1 st Law of Thermodynamics Energy is conserved ΔE = q + w 2 nd Law of Thermodynamics A
Unit 5: Spontaneity of Reaction You need to bring your textbooks everyday of this unit. THE LAWS OF THERMODYNAMICS 1 st Law of Thermodynamics Energy is conserved ΔE = q + w 2 nd Law of Thermodynamics A
