BIOINF 4371 Drug Design 1 Oliver Kohlbacher & Jens Krüger
|
|
|
- Phyllis Patterson
- 6 years ago
- Views:
Transcription
1 BIIF 4371 Drug Design 1 liver Kohlbacher & Jens Krüger Winter 2013/ Receptor- Ligand InteracAons
2 verview Receptor structure Structure elucida0on of proteins Structural change upon binding Receptor- ligand interac0ons Types of interac0ons Rela0ve strength of interac0ons Structure- based drug design Virtual screening, docking, de novo design 1
3 Structure- Based Drug Design Given 3D structure of a target Find Ligands binding to the target Binding strength of these ligands HS H? HS H H HS H H HS CH 3 H CH 3 H H CH 3 SH H 2
4 Structure- Based Drug Design Prerequisites Geometric and chemical complementarity of ligand and receptor High- quality target structure (high resolu0on) HS H? HS H H HS H H HS CH 3 H CH 3 H H CH 3 SH H 3
5 Structure- Based Drug Design Protein Candidate Structures 3D Structure Candidates Active Ligands Structure Elucidation Molecular Modeling Syn- thesis In vitro Testing Preclinical Tests Structure Elucidation (complex) HS H HS H H H H CH 3 SH H CH 3 H HS CH 3 H HS H SH H 4
6 verview of Methods Key Problems Which ligands will bind? (op0miza0on of libraries for HTS) ) Virtual Screening (in silico HTS) RelaAve binding strength of some ligands? (lead op0miza0on) ) Docking Which new ligands could be made? (no known ligands, avoid IP issues) ) de novo Design 5
7 Virtual Screening Test many(!) ligand structures quickly for one target ApplicaAons Improve enrichment in HTS, find new scaffolds Speed is more important than accuracy Large number of ligands have to be processed (millions) Precise predic0on of binding affinity not essen0al CH CH 3 HS CH 3 H H 3 C HS H H H CH 3 H H H CH 3 H HS H H CH 3 H? H H RH SH H H H CH 3 H S H CH H 2 CH 3 CH 3 Cl H H H H 2 S H F 3 C H CH 3 CH 3 CH 3 CH 3 HS H H H H HS CH 3 H RH S CH 3 CH 3 CH H 6
8 Docking Predict binding conforma0on (pose) and affinity (ΔG) of a small number of ligands for one target ApplicaAons Predic0on/understanding of binding mode, lead op0miza0on Accuracy is more important than speed Small number of compounds (hundreds?) Precise ordering of compounds is essen0al for lead op0miza0on H 2 -C CH 3 H 2 H C-? ΔG =... 7
9 De ovo Design Construct a novel compound fixng into the (known) binding pocket of a target ApplicaAons Try to come up with new leads/scaffolds if HTS did not come up with any viable hits, escape patented structure space Structures are constructed from scratch in a way that ensures op0mal interac0on with the target Problem: synthesizability of the structures H H H H HS H 2 HS H 2 8
10 Prerequisites Docking/VS/de novo design require Accurate receptor geometries Conforma0on genera0on that is complementary to the receptor Predic0on of binding free energy for the ligand Requirements Protein structure elucidaaon Conforma0onal analysis Modeling of receptor- ligand interacaons 9
11 Protein Structure Since about 1960: determina0on of protein structures using X- ray crystallography (Kendrew, Perutz) Later also nuclear magneac resonance spectroscopy (MR) Receptor structure is the basis for structure- based drug design 10
12 Protein Structure Structure reveals shape (e.g., binding pockets) Structure also reveals atomic details (interac0ons) 11
13 X- Ray Crystallography X-ray source Protein Crystal Detector Analysis 12
14 X- Rays 13
15 Protein Crystals Proteins are difficult to crystallize Irregular shape large holes in the crystal Rather large crystals required ( mm) Large amounts of protein necessary Protein needs to be very pure Crystal growth is very slow (weeks to months) Some proteins do not crystallize at all (membrane proteins!) Branden, Tooze, p
16 CrystallizaAon Hanging Drop ölting, p. 70, Branden/Tooze, p
17 DiffracAon Bragg s Law 2d sin θ = λ ConstrucAve interference occurs under angles corresponding to reflec0on at the laxce planes of the crystal d: distance between laxce planes θ: angle λ: wavelength θ d 16
18 DiffracAon Pa_ern of a Protein 17
19 verview X- Ray Crystallography 18
20 Electron Density Map 19
21 Electron Density Map 20
22 ResoluAon Resolu0on of a structure determines informa0on content Determined by quality of the crystal: Purity Inclusions Water content Stability under irradia0on Resolu0on can be es0mated from diffrac0on paiern 21
23 ResoluAon Resolu0on determines which atomic details are recognizable Poor resolu0on (large value) blurs the details of the structure Resolu0on is measured in Å Resolu0on of 2 Å does not mean, that the error for the atom coordinates is about 2 Å! Error in the atom coordinates would be about 0,3 Å in that case 22
24 uclear MagneAc Resonance 1 H nuclei possess nuclear magneac moment In an external magne0c field B 0, every nucleus assumes one of two possible states (spins) : α or β The two states differ in energy, spin state α (parallel to B 0 ) is energe0cally more favorable β B 0 α ΔE 23
25 uclear MagneAc Resonance 1 H nuclei possess nuclear magneac moment In an external magne0c field B 0, every nucleus assumes one of two possible states (spins) : α or β The two states differ in energy, spin state α (parallel to B 0 ) is energe0cally more favorable Addi0on of energy can invert the spin state β h ν α ΔE 24
26 uclear MagneAc Resonance 1 H nuclei possess nuclear magneac moment In an external magne0c field B 0, every nucleus assumes one of two possible states (spins) : α or β The two states differ in energy, spin state α (parallel to B 0 ) is energe0cally more favorable Addi0on of energy can invert the spin state β α ΔE 25
27 MR Hardware 26
28 MR Hardware 27
29 1 H- MR Spectrum of a Protein 28
30 2D- MR Spectrum Peaks on the diagonal correspond to the shins in 1D spectrum Cross peaks (off- diagonal) are caused by transfer of magne0za0on between two nuclei, i.e., interac0on between these nuclei δ 1 δ A δ B It usual implies closeness of these nuclei δ B δ A δ 2 29
31 (H,H)- CSY 30
32 Result: Structure Family MR data is compa0ble with a whole range of structures (proteins in solu0on are moving!) The result is thus a structure family Data from 1SVQ 31
33 Flexibility Different regions of the proteins show different flexibility Flexibility can be measured, for example, as RMSD between individual structure Figure on the right displays backbone flexibility as backbone thickness Termini usually highly flexible 32
34 Comparison XRD MR XRD MR Also for large proteins Requires crystals Hydrogen atoms invisible Unlabeled protein Higher spa0al resolu0on < 30 kda From solu0on Hydrogen atoms are essen0al Isotope- labeled protein required Informa0on on flexibility 33
35 Databases PDB PDB (Protein Data Bank) h_p:// Database for biomolecular structures Maintained by the RCSB (Research Collaboratory for Structural Bioinforma=cs) Deposi0on of structures in the PDB is prerequisite for the publica0on of the structure in a journal Each structure is given a unique iden0fier (PDB ID) 4 characters 1st character version 2nd 4th character structure ID Example: 2PTI, 3PTI, 4PTI are different structures of protein BPTI 2PTI: 1973, 3PTI: 1976, 4PTI:
36 PDB Contents & Growth yearly total Data from: 0
37 Receptor Structures Very good structures are required to analyze and model receptor- ligand interac0ons (resolu0on beier than 2.5 Å) Receptor structure onen reveals binding pockets and interacaons with the ligand Difficult: integral membrane proteins (ion channels, GPCRs,...) Cannot be crystallized in most cases Too large for MR, insoluble However, we onen need only the structure of an individual domain to understand receptor- ligand interac0on, not the whole receptor XRD/MR of individual (globular) domains onen sufficient and feasible 36
38 Bound Structures Details of the binding mode can be obtained from the complex structure (receptor with bound ligand) Interac0ons can be iden0fied through contacts in the structure (e.g., spa0al proximity of acceptor/donor) Good resolu0on of the structure is essen0al to iden0fy interac0ons reliably nen the receptor structure differs (slightly) between the bound and unbound state The same is true for the ligand: the bound conforma0on is not necessarily the lowest- energy conforma0on 37
39 Structural Flexibility Comparison of the unbound receptor (blue) and bound complex (red) reveals structural changes upon binding (here: MTX/DHFR, PDB: 1DF7/1DG8) 38
40 Receptor- Ligand InteracAons Two possible modes of interac0on for ligands: Irreversible covalent binding of the ligand to the receptor ) difficult to model Reversible receptor and ligand are in equilibrium with the receptor- ligand complex 39
41 Reversible/Irreversible InteracAons Irreversible (e.g., CX- 1 and aspirin) Reversible (e.g., DHFR and methotrexate) 40
42 Receptor- Ligand InteracAons Binding determined by binding free energy ΔG Entropic and enthalpic contribuaons ΔG = ΔH TΔS ΔH: binding enthalpy ΔS: binding entropy T: absolute temperature The smaller (more nega0ve) ΔG, the stronger the interac0on, the 0ghter the binding ΔH and ΔS are sums of several enthalpic/entropic contribuaons arising from different physical interacaons 41
43 Hydrogen Bonds Hydrogen bound to strongly electronega0ve atoms are polar (e.g., bound to,, F) Polar H atom at a donor atom D interacts with electron pair (lone pair) of an hydrogen bond acceptor A D H A Energe0cally, H- bonds are somewhere between covalent bonds and most other intermolecular interac0ons H 42
44 Hydrogen Bonds Can be described accurately using quantum mechanics 4- electron 3- center bond: 2 e - from lone pair of A 2 e - from D H bond Direc0onality Depends on orienta0on of acceptor lone pair Depends on direc0on of D H bond Typical distances well below the sum of van der Waals radii of paracipaang atoms \Å(X A H) H \Å(D H A) d(d A) \Å(D H A) \Å(X A H) d(d A) Å 43
45 Hydrogen Bonds EnergeAcs Hydrogen bonds possess opamal angle and length The greater the devia0on from ideal geometry, the smaller the hydrogen bonding energy Free energy contributed by a single H- bond typically 2-20 kj/mol Important acceptor groups C= H C - with lone pair with lone pair Important donor groups H H 44
46 Salt Bridges Ionic interac0on Electrosta0c interac0on between posi0vely and nega0vely charged groups - + H 3 Very strong interacaon: large charges (+/- 1 e 0 ) at a small distance ( Å) Can also involve metal ions (e.g., Zn 2+ ) Counterparts in proteins: charged side chains (Arg, Lys, Asp, Glu, His) H - + H H H 45
47 ComplexaAon of Metal Ions Many proteins contain metal ions as cofactors (Zn, Fe, Mg,...) Apart from the electrosta0c interac0on, metal ions can also form coordinaaon bonds to ligands and receptor (complexaaon) Thiols (- SH) Hydroxyamate (- CHH) - heterocycles with a lone pair Zn complexa0on by a ligand (SC 12155) in LF (lethal factor) from Bacillus anthracis Panchal et al., ature Struct. Biol., 2004, 11, 67 46
48 Hydrophobic InteracAon Favorable interac0on between apolar side chains of the receptor and lipophilic groups in the ligand ot really an interac0on o real pairwise interac0on between atoms (only vdw interac0ons and those are rather weak) Hydrophobic groups cannot form polar interac0ons Thus no polar interac0on with water possible hydrophobic groups disturb the hydrogen bond network of surrounding water Energe0c contribu0on of hydrophobic interac0on is based on reduced disturbance of the surrounding water (entropic effect) 47
49 Hydrophobic Effect Entropy comes from the solvent Apolar groups/side chains cannot form H bonds with water Water is forced to assume cage- like structures Loss of disorder in the solvent ) ΔS < 0 Free energy contribu0on approximately proporaonal to hydrophobic surface area 48
50 ther Solvent Effects Receptor and ligand are both surrounded by water and interact with water Water modulates interac0on Forms H bonds with acceptors/donors in receptor/ligand Shields electrosta0c interac0ons (dielectric constant of water: 78!) Ligand binding requires Breaking the solvent shell surrounding receptor and ligand Releasing bound waters in the receptor- ligand interface 49
51 Explizites Wasser 50
52 Solvent Effects Water forms H bonds with acceptors/donors of receptor and ligand For each H bond formed between receptor and ligand, an H bond to water has to be broken ) relaave strength of H bond is important, not their number! H H H H H + + H H H H H 51
53 Water- Mediated Hydrogen Bonds Fixed water molecules are part of many receptor- ligand complexes Example: methotrexate forms an H bond with a water molecule that is kept in place by two addi0onal H bonds with the receptor (DHFR) 52
54 Entropic ContribuAons Loss/gain of degrees of freedom (DFs) corresponds to entropic contribu0on InteracAons reduce degrees of freedom of receptor and ligand Ligand frozen in place Loses transla0onal and rota0onal degrees of freedom Loses internal degrees of freedom (flexibility) Loss of entropy energeacally unfavorable ΔG = ΔH TΔS ΔS < 0 ) TΔS > 0 53
55 Entropic ContribuAons Ligand freezing Loss of rota0onal/transla0onal DFs Loss of internal DFs Displacement of water from binding site Solvent hull of ligand and receptor stripped Water molecules displaced from binding site are released into the bulk solvent and gain rota0onal/ transla0on DFs Klebe, p
56 Strength of InteracAons Typical binding free energies of receptor- ligand complexes are between - 40 and - 10 kj/mol Each complex has its unique mix of interac0ons contribu0ng Depending on receptor and ligand, different interac0ons can dominate the binding Some complexes are dominated by enthalpic contribu0ons, some by entropic contribu0ons However: only sum of ALL contribuaons is decisive ) We need to model ALL interacaons accurately! 55
57 Example: Thermolysine Inhibitors Binding constants were determined for 80 different ligands of thermolysine Binding strength is shown as a func0on of the number of H bonds formed nly weakly correlated Reasons Desolva0on effects Strength of H bonds varies Klebe, p
58 Example: Avidin and BioAn Complex of avidin and biotin is one of the strongest interactions known: ΔG = -78 kj/mol Two dominant interactions: Seven hydrogen bonds Hydrophobic interaction BKK, p.?? 57
59 Example: Avidin and BioAn Complex of avidin and biotin is one of the strongest interactions known: ΔG = -78 kj/mol Two dominant interactions: Seven hydrogen bonds Hydrophobic interaction PDB: 2AVI 58
60 Example: CyAdine Deaminase Cy0dine deaminase (CD) catalyzes conversion of cy0dine to uridine Reac0on involves a tetrahedral transiaon state TransiAon state analogs are good inhibitors for CD Cy0dine H 2 H 2 H H Uridine H H H Ribose K i = 1.2 pmol/l H H H Ribose Ribose Tetrahedral transiaon state H Ribose Ribose K i = 30 µmol/l BKK, p
61 Example: CyAdine Deaminase Extreme difference between the two inhibitors can be explained by their ability to displace a water molecule Hydroxyl group can take the place of a water in the binding site and form favorable interac0ons with the receptor CH 2 group cannot displace the water BKK, p
62 Summary Structure- based drug design tries to iden0fy drug candidates based on the 3D structure of the receptor Structure of these receptors is elucidated using XRD or MR High- resolu0on structures reveal detailed interac0ons between receptor and ligands Key interac0ons: hydrogen bonds, hydrophobic interac0ons, electrosta0c interac0ons Ligand displaces water molecules upon binding Water plays a key role and its influence can dominate the binding free energy Ligand binds in a fixed conforma0on and thus loses internal degrees of freedom upon binding 61
63 References Books [Klebe] Klebe, Wirkstoffentwurf, 2 nd ed., Spektrum, 2009 [BKK] Böhm, Klebe, Kubinyi: Wirkstoffdesign, Spektrum 2002 [HSRF] H.- D. Höltje, W. Sippl, D. Rognan, G. Folkers: Molecular Modeling Basic Principles and Applica0ons, 2nd ed., Wiley, 2003 More on XRD: Gale Rhodes: Crystallography made crystal clear, Academic Press,
BIOINF 4371 Drug Design 1 Oliver Kohlbacher & Jens Krüger
 BIOINF 4371 Drug Design 1 Oliver Kohlbacher & Jens Krüger Winter 2013/2014 11. Docking Part IV: Receptor Flexibility Overview Receptor flexibility Types of flexibility Implica5ons for docking Examples
BIOINF 4371 Drug Design 1 Oliver Kohlbacher & Jens Krüger Winter 2013/2014 11. Docking Part IV: Receptor Flexibility Overview Receptor flexibility Types of flexibility Implica5ons for docking Examples
Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
 Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Electronic Supplementary Information Effective lead optimization targeted for displacing bridging water molecule
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Effective lead optimization targeted for displacing
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 2018 Electronic Supplementary Information Effective lead optimization targeted for displacing
Carbohydrate- Protein interac;ons are Cri;cal in Life and Death. Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
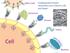 ther Cells Carbohydrate- Protein interac;ons are Cri;cal in Life and Death ormones Viruses Toxins Cell Bacteria ow to Model Protein- ligand interac;ons? Protein Protein Protein DNA/RNA Protein Carbohydrate
ther Cells Carbohydrate- Protein interac;ons are Cri;cal in Life and Death ormones Viruses Toxins Cell Bacteria ow to Model Protein- ligand interac;ons? Protein Protein Protein DNA/RNA Protein Carbohydrate
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Docking. GBCB 5874: Problem Solving in GBCB
 Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Docking Benzamidine Docking to Trypsin Relationship to Drug Design Ligand-based design QSAR Pharmacophore modeling Can be done without 3-D structure of protein Receptor/Structure-based design Molecular
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
BIOINF 4120 Bioinformatics 2 - Structures and Systems -
 BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer 2014 5. Structure Elucidation Overview Protein structure elucidation X-ray diffraction (XRD) Crystallization Physics of
BIOINF 4120 Bioinformatics 2 - Structures and Systems - Oliver Kohlbacher Summer 2014 5. Structure Elucidation Overview Protein structure elucidation X-ray diffraction (XRD) Crystallization Physics of
Life Sciences 1a Lecture Slides Set 10 Fall Prof. David R. Liu. Lecture Readings. Required: Lecture Notes McMurray p , O NH
 Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Aqueous solutions. Solubility of different compounds in water
 Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Basics of Thermodynamics: Easy learning by Dr. Anjana Sen
 Basics of Thermodynamics: Easy learning by Dr. Anjana Sen Part 1: Theory and concept Part 2: Definitions and equations Part 3: Laws of Thermodynamics Part 1: theory and concept Thermodynamics means conversion
Basics of Thermodynamics: Easy learning by Dr. Anjana Sen Part 1: Theory and concept Part 2: Definitions and equations Part 3: Laws of Thermodynamics Part 1: theory and concept Thermodynamics means conversion
Receptor Based Drug Design (1)
 Induced Fit Model For more than 100 years, the behaviour of enzymes had been explained by the "lock-and-key" mechanism developed by pioneering German chemist Emil Fischer. Fischer thought that the chemicals
Induced Fit Model For more than 100 years, the behaviour of enzymes had been explained by the "lock-and-key" mechanism developed by pioneering German chemist Emil Fischer. Fischer thought that the chemicals
CADD Pipeline Track. Docking, structure selec6on, and ensemble docking
 CADD Pipeline Track Docking, structure selec6on, and ensemble docking Jesper Sørensen, Ph.D. jesorensen@ucsd.edu NBCR Summer Ins6tute 2012 Tuesday 7/31/2012 Structure Based Drug Discovery hmp://www.proxychem.com/images/sbdd_diagram.png
CADD Pipeline Track Docking, structure selec6on, and ensemble docking Jesper Sørensen, Ph.D. jesorensen@ucsd.edu NBCR Summer Ins6tute 2012 Tuesday 7/31/2012 Structure Based Drug Discovery hmp://www.proxychem.com/images/sbdd_diagram.png
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Due in class on Thursday Sept. 8 th
 Problem Set #1 Chem 391 Due in class on Thursday Sept. 8 th ame Solutions A note: To help me read quickly, I structure the problem set like an exam. I have made small spaces for answers where I d like
Problem Set #1 Chem 391 Due in class on Thursday Sept. 8 th ame Solutions A note: To help me read quickly, I structure the problem set like an exam. I have made small spaces for answers where I d like
Some properties of water
 Some properties of water Hydrogen bond network Solvation under the microscope 1 Water solutions Oil and water does not mix at equilibrium essentially due to entropy Substances that does not mix with water
Some properties of water Hydrogen bond network Solvation under the microscope 1 Water solutions Oil and water does not mix at equilibrium essentially due to entropy Substances that does not mix with water
Back to molecular interac/ons Quantum theory and molecular structure
 Back to molecular interac/ons Quantum theory and molecular structure Atoms are arranged in 3D to cons1tute a molecule. Atoms in one molecule are connected by strong covalent bonds that are not easily broken
Back to molecular interac/ons Quantum theory and molecular structure Atoms are arranged in 3D to cons1tute a molecule. Atoms in one molecule are connected by strong covalent bonds that are not easily broken
Physiology Unit 1 CHEMISTRY REVIEW
 Physiology Unit 1 CHEMISTRY REVIEW Defini7ons Types of energy Kine7c vs. poten7al Forms of energy Chemical Ex: ATP Ma0er and Energy Electrical Ex: Ac7on poten7al of an neuron Mechanical Ex: Ac7on of muscles
Physiology Unit 1 CHEMISTRY REVIEW Defini7ons Types of energy Kine7c vs. poten7al Forms of energy Chemical Ex: ATP Ma0er and Energy Electrical Ex: Ac7on poten7al of an neuron Mechanical Ex: Ac7on of muscles
Virtual screening for drug discovery. Markus Lill Purdue University
 Virtual screening for drug discovery Markus Lill Purdue University mlill@purdue.edu Lecture material http://people.pharmacy.purdue.edu/~mlill/teaching/eidelberg/ I.1 Drug discovery Cl N Disease I.1 Drug
Virtual screening for drug discovery Markus Lill Purdue University mlill@purdue.edu Lecture material http://people.pharmacy.purdue.edu/~mlill/teaching/eidelberg/ I.1 Drug discovery Cl N Disease I.1 Drug
Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine. JAK2 Selective Mechanism Combined Molecular Dynamics Simulation
 Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
Electronic Supplementary Material (ESI) for Molecular BioSystems. This journal is The Royal Society of Chemistry 2015 Supporting Information Enhancing Specificity in the Janus Kinases: A Study on the Thienopyridine
PH 3 + H + à PH 4 PH 4. H 3 O + is pyramidal, bond angle 107 ; steric 4, 1LP Lone pair from O donated to H +
 Metallic bonding AS Bonding & Periodicity Test: Answers Ionic Bonding Ti is a giant metallic lacce. There is strong electrosta2c afrac2on between posi2ve metal ions and sea of delocalised electrons. Layers
Metallic bonding AS Bonding & Periodicity Test: Answers Ionic Bonding Ti is a giant metallic lacce. There is strong electrosta2c afrac2on between posi2ve metal ions and sea of delocalised electrons. Layers
schematic diagram; EGF binding, dimerization, phosphorylation, Grb2 binding, etc.
 Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
17. Biomolecular Interaction
 17. Biomolecular Interaction Methods for characterizing biomolecular interactions Sequence-specific DNA binding ligands Molecular mechanisms of drug action and drug resistance In silico compound design
17. Biomolecular Interaction Methods for characterizing biomolecular interactions Sequence-specific DNA binding ligands Molecular mechanisms of drug action and drug resistance In silico compound design
A. Reaction Mechanisms and Catalysis (1) proximity effect (2) acid-base catalysts (3) electrostatic (4) functional groups (5) structural flexibility
 (P&S Ch 5; Fer Ch 2, 9; Palm Ch 10,11; Zub Ch 9) A. Reaction Mechanisms and Catalysis (1) proximity effect (2) acid-base catalysts (3) electrostatic (4) functional groups (5) structural flexibility B.
(P&S Ch 5; Fer Ch 2, 9; Palm Ch 10,11; Zub Ch 9) A. Reaction Mechanisms and Catalysis (1) proximity effect (2) acid-base catalysts (3) electrostatic (4) functional groups (5) structural flexibility B.
Ch. 2 Chemical Context of Life BIOL 222
 Ch. 2 Chemical Context of Life BIOL 222 Ma1er Organisms are composed of ma1er Ma8er is anything that takes up space and has mass Ma8er is made up of elements Lowest end of the structural organiza@on of
Ch. 2 Chemical Context of Life BIOL 222 Ma1er Organisms are composed of ma1er Ma8er is anything that takes up space and has mass Ma8er is made up of elements Lowest end of the structural organiza@on of
Water. 2.1 Weak Interactions in Aqueous Sy stems Ionization of Water, Weak Acids, and Weak Bases 58
 Home http://www.macmillanhighered.com/launchpad/lehninger6e... 1 of 1 1/6/2016 3:07 PM 2 Printed Page 47 Water 2.1 Weak Interactions in Aqueous Sy stems 47 2.2 Ionization of Water, Weak Acids, and Weak
Home http://www.macmillanhighered.com/launchpad/lehninger6e... 1 of 1 1/6/2016 3:07 PM 2 Printed Page 47 Water 2.1 Weak Interactions in Aqueous Sy stems 47 2.2 Ionization of Water, Weak Acids, and Weak
Dr. Sander B. Nabuurs. Computational Drug Discovery group Center for Molecular and Biomolecular Informatics Radboud University Medical Centre
 Dr. Sander B. Nabuurs Computational Drug Discovery group Center for Molecular and Biomolecular Informatics Radboud University Medical Centre The road to new drugs. How to find new hits? High Throughput
Dr. Sander B. Nabuurs Computational Drug Discovery group Center for Molecular and Biomolecular Informatics Radboud University Medical Centre The road to new drugs. How to find new hits? High Throughput
Creating a Pharmacophore Query from a Reference Molecule & Scaffold Hopping in CSD-CrossMiner
 Table of Contents Creating a Pharmacophore Query from a Reference Molecule & Scaffold Hopping in CSD-CrossMiner Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Features
Table of Contents Creating a Pharmacophore Query from a Reference Molecule & Scaffold Hopping in CSD-CrossMiner Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Features
EXAM 1 Fall 2009 BCHS3304, SECTION # 21734, GENERAL BIOCHEMISTRY I Dr. Glen B Legge
 EXAM 1 Fall 2009 BCHS3304, SECTION # 21734, GENERAL BIOCHEMISTRY I 2009 Dr. Glen B Legge This is a Scantron exam. All answers should be transferred to the Scantron sheet using a #2 pencil. Write and bubble
EXAM 1 Fall 2009 BCHS3304, SECTION # 21734, GENERAL BIOCHEMISTRY I 2009 Dr. Glen B Legge This is a Scantron exam. All answers should be transferred to the Scantron sheet using a #2 pencil. Write and bubble
Ligand-receptor interactions
 University of Silesia, Katowice, Poland 11 22 March 2013 Ligand-receptor interactions Dr. Pavel Polishchuk A.V. Bogatsky Physico-Chemical Institute of National Academy of Sciences of Ukraine Odessa, Ukraine
University of Silesia, Katowice, Poland 11 22 March 2013 Ligand-receptor interactions Dr. Pavel Polishchuk A.V. Bogatsky Physico-Chemical Institute of National Academy of Sciences of Ukraine Odessa, Ukraine
Covalent Bonds. single bond, or single covalent bond. sharing of one pair of valence electrons. double bond, or double covalent bond
 Covalent Bonds Molecule two or more atoms held together by covalent bonds single bond, or single covalent bond sharing of one pair of valence electrons double bond, or double covalent bond sharing of two
Covalent Bonds Molecule two or more atoms held together by covalent bonds single bond, or single covalent bond sharing of one pair of valence electrons double bond, or double covalent bond sharing of two
Ch. 2 Chemical Context of Life BIOL 222
 Ch. 2 Chemical Context of Life BIOL 222 Ma1er Organisms are composed of ma1er Ma8er anything that takes up space and has mass Ma8er is made up of elements Lowest end of the structural organiza@on of life
Ch. 2 Chemical Context of Life BIOL 222 Ma1er Organisms are composed of ma1er Ma8er anything that takes up space and has mass Ma8er is made up of elements Lowest end of the structural organiza@on of life
DOCKING TUTORIAL. A. The docking Workflow
 2 nd Strasbourg Summer School on Chemoinformatics VVF Obernai, France, 20-24 June 2010 E. Kellenberger DOCKING TUTORIAL A. The docking Workflow 1. Ligand preparation It consists in the standardization
2 nd Strasbourg Summer School on Chemoinformatics VVF Obernai, France, 20-24 June 2010 E. Kellenberger DOCKING TUTORIAL A. The docking Workflow 1. Ligand preparation It consists in the standardization
Chem 112 Dr. Kevin Moore
 Chem 112 Dr. Kevin Moore Gas Liquid Solid Polar Covalent Bond Partial Separation of Charge Electronegativity: H 2.1 Cl 3.0 H Cl δ + δ - Dipole Moment measure of the net polarity in a molecule Q Q magnitude
Chem 112 Dr. Kevin Moore Gas Liquid Solid Polar Covalent Bond Partial Separation of Charge Electronegativity: H 2.1 Cl 3.0 H Cl δ + δ - Dipole Moment measure of the net polarity in a molecule Q Q magnitude
Pedro Alexandrino Fernandes, Dep. Chemistry & Biochemistry, University of Porto, Portugal
 Pedro Alexandrino Fernandes, Dep. Chemistry & Biochemistry, University of Porto, Portugal pedro.fernandes@fc.up.pt 1. Introduc3on Intermolecular Associa3ons 1. Introduc3on What type of forces govern these
Pedro Alexandrino Fernandes, Dep. Chemistry & Biochemistry, University of Porto, Portugal pedro.fernandes@fc.up.pt 1. Introduc3on Intermolecular Associa3ons 1. Introduc3on What type of forces govern these
Introduction to Chemoinformatics and Drug Discovery
 Introduction to Chemoinformatics and Drug Discovery Irene Kouskoumvekaki Associate Professor February 15 th, 2013 The Chemical Space There are atoms and space. Everything else is opinion. Democritus (ca.
Introduction to Chemoinformatics and Drug Discovery Irene Kouskoumvekaki Associate Professor February 15 th, 2013 The Chemical Space There are atoms and space. Everything else is opinion. Democritus (ca.
Why Proteins Fold. How Proteins Fold? e - ΔG/kT. Protein Folding, Nonbonding Forces, and Free Energy
 Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Details of Protein Structure
 Details of Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Anne Mølgaard, Kemisk Institut, Københavns Universitet Learning Objectives
Details of Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Anne Mølgaard, Kemisk Institut, Københavns Universitet Learning Objectives
Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India. 1 st November, 2013
 Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Determining Protein Structure BIBC 100
 Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Structural Bioinformatics (C3210) Molecular Docking
 Structural Bioinformatics (C3210) Molecular Docking Molecular Recognition, Molecular Docking Molecular recognition is the ability of biomolecules to recognize other biomolecules and selectively interact
Structural Bioinformatics (C3210) Molecular Docking Molecular Recognition, Molecular Docking Molecular recognition is the ability of biomolecules to recognize other biomolecules and selectively interact
HW 1 CHEM 362. Available: Jan. 16, 2008 Due: Jan. 25, 2008
 HW 1 CHEM 362 Available: Jan. 16, 2008 Due: Jan. 25, 2008 1. Write an equation that can be used to define the mean S-F bond energy in SF 6. How is this value likely to be related in magnitude to the energy
HW 1 CHEM 362 Available: Jan. 16, 2008 Due: Jan. 25, 2008 1. Write an equation that can be used to define the mean S-F bond energy in SF 6. How is this value likely to be related in magnitude to the energy
Enzyme function: the transition state. Enzymes & Kinetics V: Mechanisms. Catalytic Reactions. Margaret A. Daugherty A B. Lecture 16: Fall 2003
 Lecture 16: Enzymes & Kinetics V: Mechanisms Margaret A. Daugherty Fall 2003 Enzyme function: the transition state Catalytic Reactions A B Catalysts (e.g. enzymes) act by lowering the transition state
Lecture 16: Enzymes & Kinetics V: Mechanisms Margaret A. Daugherty Fall 2003 Enzyme function: the transition state Catalytic Reactions A B Catalysts (e.g. enzymes) act by lowering the transition state
Catalytic Reactions. Intermediate State in Catalysis. Lecture 16: Catalyzed reaction. Uncatalyzed reaction. Enzymes & Kinetics V: Mechanisms
 Enzyme function: the transition state Catalytic Reactions Lecture 16: Enzymes & Kinetics V: Mechanisms Margaret A. Daugherty Fall 2003 A B Catalysts (e.g. enzymes) act by lowering the transition state
Enzyme function: the transition state Catalytic Reactions Lecture 16: Enzymes & Kinetics V: Mechanisms Margaret A. Daugherty Fall 2003 A B Catalysts (e.g. enzymes) act by lowering the transition state
Chimica Farmaceutica
 Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Biological Thermodynamics
 Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
High Specificity and Reversibility
 Lecture #8 The Cell as a Machine High Specificity and Reversibility In considering the problem of transcription factor binding in the nucleus and the great specificity that is called for to transcribe
Lecture #8 The Cell as a Machine High Specificity and Reversibility In considering the problem of transcription factor binding in the nucleus and the great specificity that is called for to transcribe
Protein-Ligand Interactions* and Energy Evaluation Methods
 Protein-Ligand Interactions* and Energy Evaluation Methods *with a revealing look at roles of water Glen E. Kellogg Department of Medicinal Chemistry Institute for Structural Biology, Drug Discovery &
Protein-Ligand Interactions* and Energy Evaluation Methods *with a revealing look at roles of water Glen E. Kellogg Department of Medicinal Chemistry Institute for Structural Biology, Drug Discovery &
12A Entropy. Entropy change ( S) N Goalby chemrevise.org 1. System and Surroundings
 12A Entropy Entropy change ( S) A SPONTANEOUS PROCESS (e.g. diffusion) will proceed on its own without any external influence. A problem with H A reaction that is exothermic will result in products that
12A Entropy Entropy change ( S) A SPONTANEOUS PROCESS (e.g. diffusion) will proceed on its own without any external influence. A problem with H A reaction that is exothermic will result in products that
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Softwares for Molecular Docking. Lokesh P. Tripathi NCBS 17 December 2007
 Softwares for Molecular Docking Lokesh P. Tripathi NCBS 17 December 2007 Molecular Docking Attempt to predict structures of an intermolecular complex between two or more molecules Receptor-ligand (or drug)
Softwares for Molecular Docking Lokesh P. Tripathi NCBS 17 December 2007 Molecular Docking Attempt to predict structures of an intermolecular complex between two or more molecules Receptor-ligand (or drug)
It s the amino acids!
 Catalytic Mechanisms HOW do enzymes do their job? Reducing activation energy sure, but HOW does an enzyme catalysis reduce the energy barrier ΔG? Remember: The rate of a chemical reaction of substrate
Catalytic Mechanisms HOW do enzymes do their job? Reducing activation energy sure, but HOW does an enzyme catalysis reduce the energy barrier ΔG? Remember: The rate of a chemical reaction of substrate
Macromolecular X-ray Crystallography
 Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
7/19/2011. Models of Solution. State of Equilibrium. State of Equilibrium Chemical Reaction
 Models of Solution Chemistry- I State of Equilibrium A covered cup of coffee will not be colder than or warmer than the room temperature Heat is defined as a form of energy that flows from a high temperature
Models of Solution Chemistry- I State of Equilibrium A covered cup of coffee will not be colder than or warmer than the room temperature Heat is defined as a form of energy that flows from a high temperature
Kd = koff/kon = [R][L]/[RL]
![Kd = koff/kon = [R][L]/[RL] Kd = koff/kon = [R][L]/[RL]](/thumbs/96/127564193.jpg) Taller de docking y cribado virtual: Uso de herramientas computacionales en el diseño de fármacos Docking program GLIDE El programa de docking GLIDE Sonsoles Martín-Santamaría Shrödinger is a scientific
Taller de docking y cribado virtual: Uso de herramientas computacionales en el diseño de fármacos Docking program GLIDE El programa de docking GLIDE Sonsoles Martín-Santamaría Shrödinger is a scientific
Inorganic Pharmaceutical Chemistry
 Inorganic Pharmaceutical Chemistry Lecture No. 4 Date :25/10 /2012 Dr. Mohammed Hamed --------------------------------------------------------------------------------------------------------------------------------------
Inorganic Pharmaceutical Chemistry Lecture No. 4 Date :25/10 /2012 Dr. Mohammed Hamed --------------------------------------------------------------------------------------------------------------------------------------
Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes
 Introduction Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes The production of new drugs requires time for development and testing, and can result in large prohibitive costs
Introduction Chemical properties that affect binding of enzyme-inhibiting drugs to enzymes The production of new drugs requires time for development and testing, and can result in large prohibitive costs
Bonding in Chemistry. Chemical Bonds All chemical reactions involve breaking of some bonds and formation of new ones where new products are formed.
 CHEMICAL BONDS Atoms or ions are held together in molecules or compounds by chemical bonds. The type and number of electrons in the outer electronic shells of atoms or ions are instrumental in how atoms
CHEMICAL BONDS Atoms or ions are held together in molecules or compounds by chemical bonds. The type and number of electrons in the outer electronic shells of atoms or ions are instrumental in how atoms
MCB100A/Chem130 MidTerm Exam 2 April 4, 2013
 MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
MCBA/Chem Miderm Exam 2 April 4, 2 Name Student ID rue/false (2 points each).. he Boltzmann constant, k b sets the energy scale for observing energy microstates 2. Atoms with favorable electronic configurations
A Gentle Introduction to (or Review of ) Fundamentals of Chemistry and Organic Chemistry
 Wright State University CORE Scholar Computer Science and Engineering Faculty Publications Computer Science and Engineering 2003 A Gentle Introduction to (or Review of ) Fundamentals of Chemistry and Organic
Wright State University CORE Scholar Computer Science and Engineering Faculty Publications Computer Science and Engineering 2003 A Gentle Introduction to (or Review of ) Fundamentals of Chemistry and Organic
Water. Dr. Diala Abu-Hassan, DDS, PhD Lecture 2 MD summer Dr. Diala Abu-Hassan
 Water, DDS, PhD Dr.abuhassand@gmail.com Lecture 2 MD summer 2014 1 Lecture Content Importance of water in biological systems Noncovalent interactions Water structure Water properties Water as a solvent
Water, DDS, PhD Dr.abuhassand@gmail.com Lecture 2 MD summer 2014 1 Lecture Content Importance of water in biological systems Noncovalent interactions Water structure Water properties Water as a solvent
Protein Structure Determination using NMR Spectroscopy. Cesar Trinidad
 Protein Structure Determination using NMR Spectroscopy Cesar Trinidad Introduction Protein NMR Involves the analysis and calculation of data collected from multiple NMR techniques Utilizes Nuclear Magnetic
Protein Structure Determination using NMR Spectroscopy Cesar Trinidad Introduction Protein NMR Involves the analysis and calculation of data collected from multiple NMR techniques Utilizes Nuclear Magnetic
Class XI: Chemistry Chapter 4: Chemical Bonding and Molecular Structure Top Concepts
 1 Class XI: Chemistry Chapter 4: Chemical Bonding and Molecular Structure Top Concepts 1. The attractive force which holds together the constituent particles (atoms, ions or molecules) in chemical species
1 Class XI: Chemistry Chapter 4: Chemical Bonding and Molecular Structure Top Concepts 1. The attractive force which holds together the constituent particles (atoms, ions or molecules) in chemical species
2.2.2 Bonding and Structure
 2.2.2 Bonding and Structure Ionic Bonding Definition: Ionic bonding is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form
2.2.2 Bonding and Structure Ionic Bonding Definition: Ionic bonding is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form
1. What is an ångstrom unit, and why is it used to describe molecular structures?
 1. What is an ångstrom unit, and why is it used to describe molecular structures? The ångstrom unit is a unit of distance suitable for measuring atomic scale objects. 1 ångstrom (Å) = 1 10-10 m. The diameter
1. What is an ångstrom unit, and why is it used to describe molecular structures? The ångstrom unit is a unit of distance suitable for measuring atomic scale objects. 1 ångstrom (Å) = 1 10-10 m. The diameter
György M. Keserű H2020 FRAGNET Network Hungarian Academy of Sciences
 Fragment based lead discovery - introduction György M. Keserű H2020 FRAGET etwork Hungarian Academy of Sciences www.fragnet.eu Hit discovery from screening Druglike library Fragment library Large molecules
Fragment based lead discovery - introduction György M. Keserű H2020 FRAGET etwork Hungarian Academy of Sciences www.fragnet.eu Hit discovery from screening Druglike library Fragment library Large molecules
Downloaded from
 Points to Remember Class: XI Chapter Name: Chemical Bonding and Molecular Structure Top Concepts 1. The attractive force which holds together the constituent particles (atoms, ions or molecules) in chemical
Points to Remember Class: XI Chapter Name: Chemical Bonding and Molecular Structure Top Concepts 1. The attractive force which holds together the constituent particles (atoms, ions or molecules) in chemical
Figure 1. Molecules geometries of 5021 and Each neutral group in CHARMM topology was grouped in dash circle.
 Project I Chemistry 8021, Spring 2005/2/23 This document was turned in by a student as a homework paper. 1. Methods First, the cartesian coordinates of 5021 and 8021 molecules (Fig. 1) are generated, in
Project I Chemistry 8021, Spring 2005/2/23 This document was turned in by a student as a homework paper. 1. Methods First, the cartesian coordinates of 5021 and 8021 molecules (Fig. 1) are generated, in
Lecture outline: Chapter 7 Periodic properties
 Lecture outline: Chapter 7 Periodic properties 1. Electrostatic effects 2. Atomic size 3. Ionization energy 4. Electron affinity it 5. Summarize some periodic properties 1 Some important terms Electron
Lecture outline: Chapter 7 Periodic properties 1. Electrostatic effects 2. Atomic size 3. Ionization energy 4. Electron affinity it 5. Summarize some periodic properties 1 Some important terms Electron
Q1 current best prac2ce
 Group- A Q1 current best prac2ce Star2ng from some common molecular representa2on with bond orders, configura2on on chiral centers (e.g. ChemDraw, SMILES) NEW! PDB should become resource for refinement
Group- A Q1 current best prac2ce Star2ng from some common molecular representa2on with bond orders, configura2on on chiral centers (e.g. ChemDraw, SMILES) NEW! PDB should become resource for refinement
Performing a Pharmacophore Search using CSD-CrossMiner
 Table of Contents Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Searching with a Pharmacophore... 4 Performing a Pharmacophore Search using CSD-CrossMiner Version 2.0
Table of Contents Introduction... 2 CSD-CrossMiner Terminology... 2 Overview of CSD-CrossMiner... 3 Searching with a Pharmacophore... 4 Performing a Pharmacophore Search using CSD-CrossMiner Version 2.0
Molecular Modeling -- Lecture 15 Surfaces and electrostatics
 Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Lecture 14 (10/18/17) Lecture 14 (10/18/17)
 Lecture 14 (10/18/17) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 7, 24 Ch6 (study guide-facts); 4, 13 NEXT Reading: Ch6; 198-203 Ch6; Box 6-1 Problems: Ch6 (text); 8, 9, 10, 11, 12,
Lecture 14 (10/18/17) Reading: Ch6; 190-191, 194-195, 197-198 Problems: Ch6 (text); 7, 24 Ch6 (study guide-facts); 4, 13 NEXT Reading: Ch6; 198-203 Ch6; Box 6-1 Problems: Ch6 (text); 8, 9, 10, 11, 12,
MOLECULAR DRUG TARGETS
 MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
MOLECULAR DRUG TARGETS LEARNING OUTCOMES At the end of this session student shall be able to: List different types of druggable targets Describe forces involved in drug-receptor interactions Describe theories
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
concentrations (molarity) rate constant, (k), depends on size, speed, kind of molecule, temperature, etc.
 #80 Notes Ch. 12, 13, 16, 17 Rates, Equilibriums, Energies Ch. 12 I. Reaction Rates NO 2(g) + CO (g) NO (g) + CO 2(g) Rate is defined in terms of the rate of disappearance of one of the reactants, but
#80 Notes Ch. 12, 13, 16, 17 Rates, Equilibriums, Energies Ch. 12 I. Reaction Rates NO 2(g) + CO (g) NO (g) + CO 2(g) Rate is defined in terms of the rate of disappearance of one of the reactants, but
Concept review: Binding equilibria
 Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
User Guide for LeDock
 User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
User Guide for LeDock Hongtao Zhao, PhD Email: htzhao@lephar.com Website: www.lephar.com Copyright 2017 Hongtao Zhao. All rights reserved. Introduction LeDock is flexible small-molecule docking software,
Central Dogma. modifications genome transcriptome proteome
 entral Dogma DA ma protein post-translational modifications genome transcriptome proteome 83 ierarchy of Protein Structure 20 Amino Acids There are 20 n possible sequences for a protein of n residues!
entral Dogma DA ma protein post-translational modifications genome transcriptome proteome 83 ierarchy of Protein Structure 20 Amino Acids There are 20 n possible sequences for a protein of n residues!
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
For the following intermolecular forces:
 Lecturenotes 1 unit6_review_exercise_2017.odt Lecturenotes 2 unit6_review_exercise_2017.odt Lecturenotes 3 unit6_review_exercise_2017.odt Lecturenotes 4 unit6_review_exercise_2017.odt Answers: 1. Ionic
Lecturenotes 1 unit6_review_exercise_2017.odt Lecturenotes 2 unit6_review_exercise_2017.odt Lecturenotes 3 unit6_review_exercise_2017.odt Lecturenotes 4 unit6_review_exercise_2017.odt Answers: 1. Ionic
Lecture 11: Protein Folding & Stability
 Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall Protein Folding: What we know. Protein Folding
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
= (+)206 (kj mol 1) 206 scores 1 only Units not essential if ans in kj mol 1 but penalise incorrect units
 M.(a) (i) ΔH = Σ(enthalpies formation products) Σ(enthalpies formation reactants) Or correct cycle with enthalpy changes labelled = ( 75 242) = (+)206 (kj mol ) 206 scores only Units not essential if ans
M.(a) (i) ΔH = Σ(enthalpies formation products) Σ(enthalpies formation reactants) Or correct cycle with enthalpy changes labelled = ( 75 242) = (+)206 (kj mol ) 206 scores only Units not essential if ans
CHAPTER 2. Structure and Reactivity: Acids and Bases, Polar and Nonpolar Molecules
 CHAPTER 2 Structure and Reactivity: Acids and Bases, Polar and Nonpolar Molecules 2-1 Kinetics and Thermodynamics of Simple Chemical Processes Chemical thermodynamics: Is concerned with the extent that
CHAPTER 2 Structure and Reactivity: Acids and Bases, Polar and Nonpolar Molecules 2-1 Kinetics and Thermodynamics of Simple Chemical Processes Chemical thermodynamics: Is concerned with the extent that
Computational Analyses of Protein-Ligand Interactions
 Computational Analyses of Protein-Ligand Interactions Muhammad Kamran Haider Submitted for the Degree of Doctor of Philosophy University of York Department of Chemistry September 2010 Abstract Protein-ligand
Computational Analyses of Protein-Ligand Interactions Muhammad Kamran Haider Submitted for the Degree of Doctor of Philosophy University of York Department of Chemistry September 2010 Abstract Protein-ligand
Joana Pereira Lamzin Group EMBL Hamburg, Germany. Small molecules How to identify and build them (with ARP/wARP)
 Joana Pereira Lamzin Group EMBL Hamburg, Germany Small molecules How to identify and build them (with ARP/wARP) The task at hand To find ligand density and build it! Fitting a ligand We have: electron
Joana Pereira Lamzin Group EMBL Hamburg, Germany Small molecules How to identify and build them (with ARP/wARP) The task at hand To find ligand density and build it! Fitting a ligand We have: electron
Electronega+vity Review
 Electronega+vity Review Remember from the first course that electronega+vity is an es+mate of how atoms pull electrons towards themselves in a molecule. The higher the electron affinity, the more the element
Electronega+vity Review Remember from the first course that electronega+vity is an es+mate of how atoms pull electrons towards themselves in a molecule. The higher the electron affinity, the more the element
Nuclear Magnetic Resonance (NMR)
 Nuclear Magnetic Resonance (NMR) Nuclear Magnetic Resonance (NMR) The Nuclear Magnetic Resonance Spectroscopy (NMR) is one of the most important spectroscopic methods to explore the structure and dynamic
Nuclear Magnetic Resonance (NMR) Nuclear Magnetic Resonance (NMR) The Nuclear Magnetic Resonance Spectroscopy (NMR) is one of the most important spectroscopic methods to explore the structure and dynamic
Lecture 15: Enzymes & Kinetics. Mechanisms ROLE OF THE TRANSITION STATE. H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl. Margaret A. Daugherty.
 Lecture 15: Enzymes & Kinetics Mechanisms Margaret A. Daugherty Fall 2004 ROLE OF THE TRANSITION STATE Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products
Lecture 15: Enzymes & Kinetics Mechanisms Margaret A. Daugherty Fall 2004 ROLE OF THE TRANSITION STATE Consider the reaction: H-O-H + Cl - H-O δ- H Cl δ- HO - + H-Cl Reactants Transition state Products
Microcalorimetry for the Life Sciences
 Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Atomic Structure and Bonding. Chapter 1 Organic Chemistry, 8 th Edition John McMurry
 Atomic Structure and Bonding Chapter 1 Organic Chemistry, 8 th Edition John McMurry 1 Common Elements Groups First row Second row In most organic molecules carbon is combined with relatively few elements
Atomic Structure and Bonding Chapter 1 Organic Chemistry, 8 th Edition John McMurry 1 Common Elements Groups First row Second row In most organic molecules carbon is combined with relatively few elements
Version 1.0: 1006 abc. General Certificate of Education. CHM5 Thermodynamics and Further Inorganic Chemistry examination - June series
 Version.0: 006 abc General Certificate of Education Chemistry 642 CM5 Thermodynamics and Further Inorganic Chemistry Mark Scheme 2006 examination - June series Mark schemes are prepared by the Principal
Version.0: 006 abc General Certificate of Education Chemistry 642 CM5 Thermodynamics and Further Inorganic Chemistry Mark Scheme 2006 examination - June series Mark schemes are prepared by the Principal
CH1010 Exam #2 Study Guide For reference see Chemistry: An Atoms-focused Approach by Gilbert, Kirss, and Foster
 CH1010 Exam #2 Study Guide For reference see Chemistry: An Atoms-focused Approach by Gilbert, Kirss, and Foster Chapter 3: Atomic Structure, Explaining the Properties of Elements Trends to know (and be
CH1010 Exam #2 Study Guide For reference see Chemistry: An Atoms-focused Approach by Gilbert, Kirss, and Foster Chapter 3: Atomic Structure, Explaining the Properties of Elements Trends to know (and be
Chapter 8 Basic Concepts of Chemical Bonding
 Sec$on 8.1 Types of Chemical Bonds Chapter 8 Basic Concepts of Chemical Bonding Chapter 8 Ques$ons to Consider What is meant by the term chemical bond? Why do atoms bond with each other to form compounds?
Sec$on 8.1 Types of Chemical Bonds Chapter 8 Basic Concepts of Chemical Bonding Chapter 8 Ques$ons to Consider What is meant by the term chemical bond? Why do atoms bond with each other to form compounds?
Cheminformatics platform for drug discovery application
 EGI-InSPIRE Cheminformatics platform for drug discovery application Hsi-Kai, Wang Academic Sinica Grid Computing EGI User Forum, 13, April, 2011 1 Introduction to drug discovery Computing requirement of
EGI-InSPIRE Cheminformatics platform for drug discovery application Hsi-Kai, Wang Academic Sinica Grid Computing EGI User Forum, 13, April, 2011 1 Introduction to drug discovery Computing requirement of
2-1 The Nature of Matter
 2-1 The Nature of Matter Small Atoms Placed side by side, 100 million atoms would make a row only about 1 centimeter long. contain subatomic particles Atoms What three subatomic particles make up atoms?
2-1 The Nature of Matter Small Atoms Placed side by side, 100 million atoms would make a row only about 1 centimeter long. contain subatomic particles Atoms What three subatomic particles make up atoms?
Waves. λ = c / f ω = 2π f = 2π / T. E = hf = ω E mole = N A. E X ray. ~ ~ J / mol! Wilhelm Röntgen
 Waves An X-ray picture (radiograph), taken by Wilhelm Röntgen in 1896, of Albert von Kölliker's hand. Wilhelm Röntgen λ c / f ω 2π f 2π / T E hf ω E mole N A ω E X ray ~ 6 10 23 10 34 10 18 ~ 6 10 7 J
Waves An X-ray picture (radiograph), taken by Wilhelm Röntgen in 1896, of Albert von Kölliker's hand. Wilhelm Röntgen λ c / f ω 2π f 2π / T E hf ω E mole N A ω E X ray ~ 6 10 23 10 34 10 18 ~ 6 10 7 J
