The dynamics of fluorescently labeled endogenous gurken mrna in Drosophila
|
|
|
- Joella Fisher
- 5 years ago
- Views:
Transcription
1 Research Article 887 The dynamics of fluorescently labeled endogenous gurken mrna in Drosophila Angela M. Jaramillo 1,2, Timothy T. Weil 2, Joseph Goodhouse 2, Elizabeth R. Gavis 2 and Trudi Schupbach 1,2, * 1 Howard Hughes Medical Institute and 2 Department of Molecular Biology, Princeton University, NJ 08544, USA *Author for correspondence ( schupbac@princeton.edu) Accepted 11 December , Published by The Company of Biologists 2008 doi: /jcs Summary During Drosophila oogenesis, the targeted localization of gurken (grk) mrna leads to the establishment of the axis polarity of the egg. In early stages of oogenesis, grk mrna is found at the posterior of the oocyte, whereas in the later stages grk mrna is positioned at the dorsal anterior corner of the oocyte. In order to visualize the real-time localization and anchorage of endogenous grk mrna in living oocytes, we have utilized the MS2-MCP system. We show that MCP-GFP-tagged endogenous grk mrna localizes properly within wild-type oocytes and behaves aberrantly in mutant backgrounds. Fluorescence recovery after photobleaching (FRAP) experiments of localized grk mrna in egg chambers reveal a difference in the dynamics of grk mrna between young and older egg chambers. grk mrna particles, as a population, are highly dynamic molecules Introduction Asymmetric messenger RNA localization provides a means to spatially restrict protein synthesis. In many cell types, on-site translation serves to maintain and elaborate a polarized architecture. However, the fundamental questions of how a transcript moves to its destination, how it remains translationally silent while in transit and how it is anchored at its target site remain unanswered for many mrnas (reviewed in St Johnston, 2005). In Drosophila, several asymmetrically localized transcripts have been identified in the developing oocyte. bicoid mrna is localized at the anterior of the oocyte, whereas oskar can be found spatially restricted at the posterior of the oocyte (Berleth et al., 1988; Ephrussi et al., 1991; Lehmann and Nusslein-Volhard, 1986). These transcripts, along with gurken (grk) mrna, are necessary for the patterning of the Drosophila embryo. gurken encodes a transforming growth factorα (TGFα)-like protein that is secreted by the oocyte and activates the epidermal growth factor receptor (EGFR) in the overlying somatic follicle cells. In early oogenesis, grk mrna accumulates at the posterior of the oocyte. Gurken protein, synthesized from this RNA, induces posterior cell fates in the overlying follicle cells. Soon thereafter, during mid-to-late oogenesis, grk mrna and protein are localized to the dorsal anterior corner of the oocyte, establishing dorsal cell fates in lateral follicle cells (Neuman- Silberberg and Schupbach, 1996; St Johnston, 2005; Van Buskirk and Schupbach, 1999). Mutations in several genes have been identified that result in mislocalization of grk mrna. Some of these genes, such as squid and hrb27c, encode heterogeneous nuclear ribonucleoproteins (hnrnps) (Goodrich et al., 2004; Neuman-Silberberg and Schupbach, 1993), whereas others include genes necessary for cytoskeletal integrity, such as cappuccino, spire (an actin nucleator) and spindle-f (microtubule organizer) (Abdu et al., 2006; Manseau that steadily lose their dynamic nature as oogenesis progresses. This difference in dynamics is attenuated in K10 and sqd 1 mutants such that mislocalized grk mrna in older stages is much more dynamic compared with that in wild-type controls. By contrast, in flies with compromised dynein activity, properly localized grk mrna is much more static. Taken together, we have observed the nature of localized grk mrna in live oocytes and propose that its maintenance changes from a dynamic to a static process as oogenesis progresses. Supplementary material available online at Key words: grk, Oogenesis, FRAP, RNA localization, Drosophila and Schupbach, 1989; Neuman-Silberberg and Schupbach, 1993). In addition, it has been shown that the motor enzymes kinesin and dynein are required for correct grk mrna localization (Brendza et al., 2002; Clark et al., 2007; Delanoue et al., 2007; Duncan and Warrior, 2002; Januschke et al., 2002). These requirements suggest a model whereby grk mrna is assembled into an RNP complex that is transported on filaments by molecular motors. In fact, grk mrna synthesized in vitro and injected during mid-oogenesis assembles into nonmembranous transport particles that move first to the anterior of the oocyte and then dorsally towards the nucleus, and these movements are dependent on dynein. Furthermore, microtubule depolymerization disrupts the directionality of injected grk mrna, implying active transport along microtubules (Clark et al., 2007; Delanoue et al., 2007; MacDougall et al., 2003). However, little is known about how grk mrna is maintained or anchored at its destination. Recent ultrastructural analysis suggest that grk mrna is not only transported but also anchored by dynein to large cytoplasmic structures called sponge bodies at the dorsal-anterior corner (Delanoue et al., 2007). We have used a system for fluorescently labeling mrna in vivo to visualize the real-time localization and anchoring of endogenous grk mrna in living oocytes. We show that GFP-labeled endogenous grk mrna properly localizes within wild-type oocytes and behaves aberrantly in mutant backgrounds. Interestingly, pharmacological studies show that the anchoring of grk mrna might not be dependent on an intact cytoskeleton or, alternatively, it might utilize a subpopulation of microtubules that are very drug resistant. Fluorescence recovery after photobleaching (FRAP) experiments of grk mrna particles in live egg chambers reveal a difference in the dynamic state of localized grk mrna between early- and midstage egg chambers. grk mrna particles, as a population, are highly dynamic molecules, exhibiting high fluorescence recovery, during
2 (6) early stages. As oogenesis progresses, grk mrna steadily loses its dynamic nature. This difference in grk mrna mobility is attenuated in K10 and sqd 1 mutants such that mislocalized grk mrna in older stages remains much more dynamic. By contrast, in flies with compromised dynein activity, properly localized grk mrna is more static. Taken together, we have observed the changing nature of localized grk mrna in live oocytes and have demonstrated that its maintenance can be a dynamic process. We show that localized grk mrna, as a population, can be highly dynamic or static, depending on the stage of oogenesis. Investigating the changing nature of grk mrna allows us to gain insight into the mechanisms involved in the maintenance of localized transcripts. Results In vivo labeling of gurken mrna in egg chambers In order to visualize grk mrna in live egg chambers, we took advantage of a system for fluorescent labeling of mrna in vivo. We have utilized the MS2-MS2 coat protein (MS2-MCP) system pioneered by Singer and colleagues in yeast (Bertrand et al., 1998) and adapted for Drosophila by Gavis and colleagues (Forrest and Gavis, 2003; Weil et al., 2006). Twelve stem-loop binding sites for MCP were inserted into the 3 UTR of grk mrna. The grk-(ms2) 12 transgene rescues the grk 2B mutant phenotype, indicating that the stem loops do not significantly interfere with the function of the RNA. grk-(ms2) 12 was coexpressed with either the hsp83-mcp- GFP or the hsp83-mcp-rfp transgene, which encode MCP fused to either green fluorescent protein (GFP) or red fluorescent protein (RFP) (Weil et al., 2006). Eggs from flies expressing both transgenes, grk-(ms2) 12 together with hsp83-mcp-gfp or hsp83- MCP-RFP (referred to as grk*gfp and grk*rfp), show no obvious alterations in morphology. Moreover, the labeled RNAs, grk*gfp and grk*rfp, have localization patterns identical to that of wildtype grk mrna. As previously described, grk mrna is expressed in early egg chambers (stages 1-7) and localized to the posterior end of the oocyte. Starting at stage 8, the oocyte nucleus moves from the center to the anterior edge of the oocyte. As the egg chamber grows, this edge will become the future dorsal anterior corner. During this stage, grk mrna begins to accumulate around the new position of the nucleus as well as the anterior of the oocyte, creating a transient cortical ring. From stages 9 to 10B, grk mrna remains in tight association with the dorsal anterior corner, forming a cap over the nucleus (Neuman-Silberberg and Schupbach, 1993). We used laser scanning confocal microscopy for live imaging of grk*gfp and grk*rfp during oogenesis (Fig. 1). Egg chambers were dissected and kept within a culture dish with nutrient media and then immediately imaged. Throughout early stages 4-7, grk*gfp and grk*rfp can be seen lining the posterior of the oocyte (Fig. 1A). During stage 7, when the oocyte nucleus is centered or shifting to one side, live images of grk*gfp show it to accumulate around the anterior cortex as well as continuing to line the posterior. At stage 8, the oocyte nucleus moves entirely to one side. As soon as the nucleus is asymmetric, the amount of grk*gfp at the posterior and ventral lateral sides of the oocyte decreases and, instead, grk*gfp appears to line the lateral wall posterior to the nucleus (Fig. 1B). At this stage, the dorsal-anterior cap of grk*gfp above the nucleus becomes increasingly prominent. While stage 8 progresses, grk*gfp continues to be visible anteriorly in a prominent ring, and meanwhile little to no grk*gfp can be seen at the posterior. As the egg chamber develops through stage 9, less and less grk*rfp accumulates in the ring, whereas grk*rfp in the Fig. 1. Live imaging of endogenous grk mrna. (A-C) grk*gfp expressed in the younger stages of mid-oogenesis. (A) Stage 7 shows posterior accumulation. (B) In early stage 8, there is often a lateral-cortical lining of grk*gfp on the side of the oocyte nucleus (asterisk). (C) During late 8, there is no longer a posterior lining of grk*gfp but instead the beginnings of an anterior ring with significant accumulation at the ventral anterior. (D,E) grk*rfp expressed in older stages of mid-oogenesis. From stages 9 to 10, there is a prominent dorsal-anterior cap and a strong reduction of grk*rfp at the ventral-anterior corner. (F) Control line expressing MCP-RFP alone. MCP-GFP and MCP-RFP have a nuclear localization sequence, and, as a consequence, when no grk mrna with stem loops is present, the protein will enter the follicle cell nuclei (arrowhead) and nurse cell nuclei (arrow). Bars, 50 m. ventral-anterior corner persists a bit longer, as previously reported by MacDougall and colleagues using fluorescent in situ hybridization (MacDougall et al., 2003). During this time, grk*rfp becomes more prominently localized to the dorsal-anterior cap above the nucleus. From late stage 9 to 10B, grk*rfp continues to be found above the nucleus and has a fainter appearance. Actin-destabilizing drugs do not disrupt the maintenance of localized grk mrna in live egg chambers An intact actin cytoskeleton has been shown to be important for asymmetric localization of RNA and protein in Drosophila oocytes (reviewed by Hudson and Cooley, 2002). Treatment with actindestabilizing drugs, such as cytochalasin D, disrupts the anchoring of nanos and bicoid mrna (Forrest and Gavis, 2003; Weil et al., 2006). Also, elimination or reduction of actin-binding proteins, such as moesin and tropomyosin, disrupts the posterior maintenance of Osk protein and mrna (Jankovics et al., 2002; Polesello et al., 2002). In order to investigate whether grk mrna requires an intact actin framework, we treated live egg chambers expressing grk*rfp with the actin-destabilizing drugs latrunculin A and cytochalasin D, either individually or in combination. To test the effectiveness of these drugs, we stained egg chambers with phalloidin, which revealed very reduced, barely detectable F-actin, except for prominent ring canals. The drug concentrations that we used were capable of blocking nurse cell dumping, which results from actindependent contractions of the nurse cells (Gutzeit, 1982). Timelapse imaging of egg chambers from drug-treated GFP-actin flies show rapid depolymerization. We were therefore confident that actin was severely disrupted. Interestingly, we found that grk*rfp is well maintained during stages 6-10 (Fig. 2). grk*rfp remains tightly anchored both early at the posterior and later at the dorsal-anterior corner, and this anchoring is disrupted only when the drug
3 Dynamics of endogenous gurken in Drosophila 889 Fig. 2. Actin is not necessary for anchorage of grk mrna. (A-C) Dissected live grk*rfp egg chambers treated with DMSO, the solvent used to dissolve the actin-destabilizing drugs. grk*rfp is localized properly from young to late stages. (D-F) Egg chambers were treated with the actin-destabilizing drugs cytochalasin D and latrunculin A for minutes. There is no obvious disruption of grk*rfp. concentrations are high enough to cause the oocyte to collapse. In these cases, grk*rfp spreads along the cortex of the oocyte. Additionally, we fed adult female grk*rfp flies latrunculin A and cytochalasin D and observed that grk*rfp was properly localized (data not shown). Similarly, Saunders and Cohen (Saunders and Cohen, 1999) saw no changes in the localization of the grk transcript from flies fed cytochalasin D. Resistant microtubules might explain why grk mrna remains localized after using microtubule-destabilizing drugs Microtubules are central for the localization of several mrnas in Drosophila oocytes (St Johnston, 2005). With respect to maintenance, disruption of microtubules has been shown to release bicoid, fs(1)k10, orb and Bicaudal-D from the anterior of the oocyte (Pokrywka and Stephenson, 1995). In order to investigate whether an intact microtubule network is needed to maintain grk mrna localization, we treated live egg chambers from flies expressing grk*rfp with a combination of the microtubule-destabilizing drugs colchicine and colcemid. After drug treatments, grk*rfp localization appeared not to have been disrupted, whether positioned at the posterior or at the dorsal anterior corner during stages 6-10 (Fig. 3). Time-lapse imaging of drug-treated egg chambers over intervals of minutes confirmed these results. These results suggest that anchoring of grk*rfp is not dependent on microtubules or that the anchoring is meditated by a population of microtubules that are resistant to these inhibitors. To ensure that certain populations of microtubules were indeed disrupted, we monitored the microtubule-dependent ooplasmic streaming that occurs before nurse cell dumping (Gutzeit, 1982). This unidirectional flow ceased within 3 minutes of applying our treatment of colcemid and colchicine, indicating that the drug treatment was very efficient at disrupting microtubules that promote cytoplasmic streaming. We also imaged drug-treated tau-gfp-expressing fly egg chambers. Tau-GFP decorates microtubules and allowed us to follow depolymerization in real time. Interestingly, while depolymerization of microtubules in the oocyte and nurse cells was plainly evident, the microtubule basket surrounding the oocyte nucleus [previously reported at stage 9 by MacDougall and colleagues (MacDougall et al., 2003)] was clearly resistant to the combined effects of colchicine and colcemid (see supplementary material Movie 1). This suggests that there might be different types of microtubules present in the oocyte that have different sensitivities to depolymerizing drugs. Whether grk*rfp is anchored to these resistant microtubules is unclear, but our data clearly show that grk*rfp is not disrupted by the treatment. We also examined the combined effect of colcemid and colchicine on grk*rfp in egg chambers from females fed with these inhibitors. We observed a range of defects, including loss of grk*rfp, mislocalized grk*rfp in the oocyte and accumulation of grk*rfp in the nurse cells (supplementary material Fig. S1). By contrast, it has been reported that there is an accumulation of grk transcripts in the oocytes rather than the nurse cells from colchicine-fed flies (Saunders and Cohen, 1999). The combination of two inhibitors used in our experiments might account for the differences. Nevertheless, under these conditions, it is difficult to distinguish between the roles microtubules might play in mrna transport versus anchoring within the oocyte. grk mrna is a dynamic molecule In some cases, mrnas are stably anchored at their destinations. For example, nanos mrna diffuses to the posterior of the oocyte and becomes stably anchored in an actin-dependent manner (Forrest and Gavis, 2003). Dynein acts as a static anchor for the apical transcripts runt and fushi tarazu in Drosophila blastoderm embryos (Delanoue and Davis, 2005). However, recent work indicates that the maintenance of localized mrnas might also be a dynamic process. bicoid mrna is maintained at the anterior oocyte cortex by continual active transport on microtubules (Weil et al., 2006). To examine whether grk mrna is stably anchored during different stages of oogenesis, we performed FRAP experiments. grk*gfp was photobleached, either in the region posterior to the oocyte nucleus during stages 6 and 7 or on the dorsal-lateral side of the cap above the oocyte nucleus during stages 8 and 9 (see supplementary material Movies 2-4). Our FRAP conditions cause photobleaching of a 2.5 μm 2 area of grk*gfp in the focal plane, as well as grk*gfp as far out of focus as 2-3 μm above and below Fig. 3. grk*rfp appears to remain unaltered after treatment with microtubuledestabilizing drugs. (A-C) Dissected live grk*rfp egg chambers treated with ethanol show properly maintained grk*rfp. (D-F) Egg chambers were treated with the microtubule-destabilizing drugs colchicine and colcemid for minutes. grk*rfp does not appear to be disrupted in either young or older egg chambers.
4 (6) this plane (supplementary material Fig. S2D-F). Quantitative analysis shows 70% recovery of fluorescence, with a half-time of ~80 seconds (t 1/2 =τ*ln2, where τ is the time constant of recovery) during stage 6. By contrast, by late stage 9, recovery is reduced to 30%, with a t 1/2 of >120 seconds (Fig. 4). Thus, the steady-state population of grk*gfp is highly dynamic during early stages of oogenesis, but this dynamic behavior subsides at later stages. What is the source of fluorescence recovery shown by grk mrna? Fluorescence recovery could result from lateral movement of grk*gfp that is already localized. To address this possibility, we measured the fluorescence intensity values of the neighboring pools of localized non-bleached grk*gfp that surround the bleached area during a FRAP experiment for a corresponding decrease in fluorescence. The fluorescence of these neighboring regions was not decreased, however, implying that recovery does not primarily involve prelocalized sources of grk*gfp present at the cortex (Fig. 5). In addition, we performed inverse FRAP experiments on posterior localized grk*gfp. We bleached two areas of cortical grk*gfp and measured the fluorescence intensity of the localized, non-bleached grk*gfp in between the two bleached spots. In these cases, there was no appreciable loss of fluorescence to the nonbleached localized grk*gfp, while high fluorescence recovery occurred within the neighboring bleached spots (data not shown). Unless constant and rapid accumulation of new cytoplasmic grk*gfp obscures the fluorescence loss from the cortex, our results indicate that the recovery of fluorescence is due to movement of grk*gfp from the cytoplasm to the cortex. To determine whether our fast fluorescent recovery is due to newly synthesized grk*gfp from the nurse cells, we bleached the entire oocyte at stage 7. We Fig. 4. grk mrna is a dynamic molecule. FRAP experiments were performed on grk*gfp egg chambers at stages 6-9. (A-C) Images of grk*gfp egg chambers at stages 6-9, with a small box representing the area that is bleached during a FRAP experiment. (D,E) Line graphs representing the average relative fluorescence of grk*gfp during several FRAP experiments. (D) During stage 6, a defined area of fluorescent grk*gfp at the oocyte posterior was bleached and about 70% was recovered. (E) grk*gfp localized at the dorsal-anterior cap during late stage 9 was bleached and showed reduced recovery. (F) Bar graph representing the percentage of fluorescence recovery from stages 6 to late stage 9. There is a steady decrease in the amount of recovery as the oocyte progresses through development. (G) Bar graph representing the half-time of recovery (t 1/2 ) for stages 6 to late 9. Although recovery steadily decreased, the rate did not, suggesting there could be a similar mechanism occurring during stages 6 to early 9. A minimum of five FRAP experiments were performed for each data point. Bars represent the standard deviation; e, early; l, late. observed partial grk*gfp recovery within 20 minutes, more than ten times longer than the recovery observed after bleaching the cortex only (data not shown). Thus, we conclude that, at stage 7, mrna newly transported from the nurse cells cannot be the major source of cortical recovery and, instead, recovery most probably reflects exchange from the nearby cytoplasm. Mislocalized grk mrna in K10 and sqd 1 mutant oocytes is dynamic We considered the possibility that mislocalized grk*gfp in mutant backgrounds could manifest altered dynamics. In K10 and sqd 1 mutants (Kelley, 1993; Wieschaus, 1978), grk mrna appears to have normal localization patterns during younger stages of oogenesis, but, in mid-to-late stages, grk mrna is mislocalized along the entire anterior circumference of the oocyte instead of being restricted to the dorsal-anterior corner (Neuman-Silberberg and Schupbach, 1993). In addition, in these mutants, grk mrna is translated along the anterior cortex, resulting in ectopic EGFR activation. Excess receptor activation results in the induction of more dorsal cell fates, leading to expanded dorsal appendages on the egg (Kelley, 1993; Neuman-Silberberg and Schupbach, 1993; Neuman- Silberberg and Schupbach, 1994; Norvell et al., 1999). grk*gfp reproducibly shows a similar pattern of localization in either K10 or sqd 1 mutant backgrounds (Fig. 6B-D). To examine the behavior of localized and mislocalized grk mrna in the K10 and sqd 1 mutant oocytes, we performed FRAP experiments. We photobleached the dorsal and ventral anterior corners of K10 and sqd 1 oocytes (Fig. 6D-F; supplementary material Fig. S3). Interestingly, in mid-to-late stage 9 egg chambers, we found
5 Dynamics of endogenous gurken in Drosophila 891 Fig. 5. grk mrna recovery does not specifically involve depletion of nearby localized sources. (A,B) Images of stage 7 (A) and late stage 8 (B) egg chambers from flies expressing grk*gfp illustrating the region bleached during the FRAP experiment (black box), plus arrows pointing to the unbleached areas of grk*gfp monitored for fluorescence loss. (C,D) Line graphs representing average relative fluorescence of grk*gfp and two unbleached neighboring pools of grk*gfp during FRAP experiments. In panel C, the two posterior pools that surrounded the bleached region of grk*gfp during stage 7 show no dramatic decrease in fluorescence intensity. The pool on the right of the bleached region is represented by the dark gray line, whereas the pool on the left is the pale gray line. (D) Likewise, the surrounding sources of localized grk*gfp found at the dorsal-anterior cap during late stage 8 also displayed no measurable decrease while recovery occurred (right, dark gray; left, pale gray). This suggests that recovery is not due to nearby localized sources of grk*gfp. grk*gfp at both the dorsal and ventral sides to be dynamic. In fact, grk*gfp is more dynamic (~45% recovery) compared with grk*gfp in a wild-type background (~25%) at this stage (Fig. 6E and supplementary material Movie 5). Accordingly, the recovery rates (t 1/2 ) for grk*gfp during mid-to-late stage 9 egg chambers in the mutant backgrounds are much faster. The nature of grk*gfp dynamics resembles that of a younger stage with faster and higher recovery, significantly different from the control, which displayed a slow rate with little recovery (Fig. 6F). We also performed FRAP experiments on K10 and sqd 1 mutants at stages 6 and 7, when grk mrna is properly localized at the posterior (supplementary material Fig. S4). Surprisingly, we found a substantial amount of variability in K10 mutants and a general increase of recovery in sqd 1 mutants. Compromised dynein activity interferes with the recovery of grk mrna Recent evidence for direct binding between grk mrna and the dynein light chain adds to the mounting support for the potential role of the dynein motor enzyme in grk mrna localization (Rom et al., 2007). Given the dynamic nature of grk mrna, we examined the behavior of grk*gfp under conditions that compromise dynein motor activity. First, we overexpressed dynamitin (Dmn), which is an essential component of the dynactin complex and is necessary for dynein activity (Kamal and Goldstein, 2002). Overexpression of Dmn results in an increase of anteriorly localized grk mrna during mid-stages of oogenesis, as seen by in situ hybridization, and mispositioning of the oocyte nucleus (Duncan and Warrior, 2002; Januschke et al., 2002). Second, we took advantage of hypomorphic dynein heavy chain alleles (dhc) alleles, which permit oogenesis but have been shown to reduce transport (Clark et al., 2007; Weil et al., 2006). FRAP experiments were performed on localized grk*gfp in oocytes where dynein function was reduced by either of the above methods. In both situations, we observed mislocalized grk*gfp in the oocyte cytoplasm accompanied by large aggregates, from stages 5 to 10. These large aggregates are most clearly visible in the nurse cell cytoplasm and are very reminiscent of the aggregates we observed in colchicine-fed grk*gfp flies (supplementary material Fig. S1B). Upon overexpression of Dmn, a significant portion of grk*gfp is still capable of reaching its proper destination, both at the posterior and dorsal-anterior corner (Fig. 7C,D). In dhc 6-10/6-12 egg chambers (Fig. 7E,F), grk*gfp no longer accumulates in a thin crescent along the posterior cortex during stages 6 and 7 and instead is very diffuse throughout the oocyte. During mid-stages of oogenesis, there is an accumulation of grk*gfp at the anterior cortex beginning as early as stage 7. A defined area of fluorescent grk mrna at the posterior and dorsalanterior was bleached during FRAP experiments. With overexpression of Dmn, fluorescence recovery is reduced to 34% in stages 6-7 and 28% in stage late 8 to early 9 (l8-e9). Similarly, in dynein mutant egg chambers, grk*gfp recovery is reduced to 24% at stage l8-9. This is in direct contrast to control fluorescence recovery values of 67% for stages 6-7 and 46% for l8-e9 stages (Fig. 7G). Taken together, our results show that, when dynein is compromised by two different methods, we observe a significant reduction in the dynamics of grk mrna. Discussion In order to visualize grk mrna in vivo, we used the MS2-MCP system. Unlike in situ hybridization or FISH techniques that reveal static images of RNA, the MS2-MCP system has allowed us to observe live samples and film grk mrna during various stages of oogenesis. Using this new detection method, we have described the live localization patterns of fluorescent grk mrna. Our description is similar to those originally published (Neuman-Silberberg and Schupbach, 1993) and confirms observations recently published using FISH (MacDougall et al., 2003). We also describe grk mrna accumulating at the anterior before the posterior localization has
6 (6) Fig. 6. Mislocalized grk mrna at the anterior is dynamic. FRAP experiments were performed on K10; grk*gfp and grk*gfp; sqd 1 egg chambers. Images of grk*gfp (A), K10; grk*gfp (B,D) and grk*gfp; sqd 1 (C) egg chambers during midto-late stage 9. (B-D) Mislocalized grk*gfp forms an anterior ring at the edge of the oocyte. A defined area of fluorescent grk mrna at the dorsal (arrowhead) and ventral-anterior (arrow) sides was bleached during FRAP experiments. (E) Previously we noted that grk*gfp localized at the dorsal-anterior cap during mid-to-late stage 9 had very reduced recovery (~25% recovery) during FRAP experiments. By contrast, grk*gfp found both at the dorsal and ventral-anterior sides of K10 and sqd 1 mutants was more dynamic (~45%). (F) Bar graph representing the half-time of recovery (t 1/2 ) for grk*gfp during mid-to-late stage 9 from controls and mutant lines. Recovery rates are much faster in the mutants, similar to the younger stages in the control line (Fig. 4). A minimum of five FRAP experiments were taken for each data point. Error bars represent standard deviations. disappeared. Moreover, in early stage 8, we observe a lateral accumulation of grk mrna that persists on the side of the oocyte where the nucleus is found (the future dorsal side), whereas, at the same time, the opposite side (the future ventral side) no longer has grk mrna except for a small accumulation at the anterior cortex. At this transitional stage, large amounts of grk mrna line the entire anterior and dorsal cortex of the oocyte. This configuration eventually resolves to the well-defined dorsal-anterior cap, by first restricting grk mrna on the dorsal side, followed by restricting grk mrna at the anterior. The question arises as to whether grk mrna is transported along the lateral wall from the posterior to the dorsal-anterior corner or whether the lateral RNA decays and new grk mrna transcripts populate the corner from the nurse cells. As endogenous grk mrna does not form particles large enough to be resolved and tracked with our imaging system, we are presently unable to address this question. Using photoswitchable fluorescent molecules in combination with the MS2-MCP system could provide some answers to these questions in the future. The process of RNA localization is likely to involve several of the following steps: RNP assembly, nuclear export, cytoplasmic transport, anchorage, translation and decay (St Johnston, 2005). Stable anchorage versus continuous transport have been proposed as two different mechanisms for the maintenance of mrnas before translation occurs. Possible anchors can be microfilaments, microtubules, proteins and even noncoding RNAs, such as the Xlsirts in Xenopus oocytes (Kloc and Etkin, 2005). Continuous transport can give the appearance of an mrna having a static anchor because this dynamic mechanism can result in a net gain of accumulation. In fact, bicoid mrna has recently been shown to be maintained in late oocytes by continual active transport (Weil et al., 2006). Unexpectedly, it has been demonstrated that dynein, separate from its motor function, can act as a static anchor for apical transcripts in Drosophila embryos (Delanoue and Davis, 2005). In an attempt to study whether the cytoskeleton is a necessary factor for the maintenance of already localized grk transcripts, we used cytoskeleton-destabilizing drugs. Our results suggest that actin does not play a role in the anchorage. This also corresponds to data obtained by moesin-deficient flies, which have defects in localization of oskar mrna because of a loose and detached actin network, but in which grk mrna remains undisrupted (Jankovics et al., 2002; Polesello et al., 2002). Interestingly however, Babu and colleagues (Babu et al., 2004) showed that osk mrna has two redundant anchoring processes. They noted that latrunculin A treatments did not disrupt the maintenance of osk unless other proteins, such as Homer, were absent (Babu et al., 2004). It is therefore possible that the actin cytoskeleton is similarly involved in the maintenance of grk mrna in a manner redundant with that of other proteins, but so far no actin-binding proteins have been found to bind directly to the grk mrna. Using microtubule-destabilizing drugs, we see no obvious disruption of localized grk mrna throughout stages This agrees with injection studies using grk mrna synthesized in vitro that show that show localized grk mrna is not disrupted by the later injection of colcemid (MacDougall et al., 2003). Interestingly, we observed that the microtubule network around the oocyte nucleus of stage 9-10 egg chambers was resistant to our drug treatments. It is not uncommon to observe populations of microtubules to be resistant to depolymerizing drugs. Often it is a characteristic of stable versus dynamic populations of microtubules (Baas et al., 1994; Bannigan et al., 2006; Guillaud et al., 1998; Palazzo et al., 2003). These microtubules persist even when nuclear migration is disrupted in grk mutants, or when the nucleus is displaced in various other mutants (Guichet et al., 2001; Januschke et al., 2002). It is possible that grk mrna might attach itself to these MTs. Moreover, overexpressing Dmn, a component of the dynein-dynactin complex, disrupts this oocyte nucleus microtubule scaffold, resulting in grk mrna being dispersed in the oocyte within stage 10 egg chambers (Januschke et al., 2002), very similar to what we saw in our experiments. During stages 9 and 10, when we observed the resistant microtubule basket, our FRAP studies show little fluorescence recovery. During these later stages, grk mrna therefore appears to have a more stable anchorage and might no longer be in transit, whereas the high percentage and fast rate of recovery during younger stages suggests a high transport phase. The t 1/2 is remarkably similar for stages 6 to early 9, suggesting a similar mechanism might be occurring during these stages. However, recovery slows down at mid-to-late stage 9 and is likely to be indicative of a shift in an equilibrium to a more stably anchored grk mrna. Recovery does
7 Dynamics of endogenous gurken in Drosophila 893 Fig. 7. Reducing dynein activity leads to reduced grk mrna recovery. FRAP experiments were performed on grk*gfp; αtubgal4/+;uas-dmn/+ and grk*gfp; dhc 6-10/6-12 egg chambers. Images of grk*gfp (A,B), grk*gfp; αtubgal4/+;uas-dmn/+ egg (C,D) and grk*gfp; dhc 6-10/6-12 (E,F) egg chambers during stages 6-7 (A,C,E) and late 8 to early 9 (B,D,F). (A,B) Properly localized grk*gfp lines the posterior cortex of young egg chambers, whereas grk*gfp forms a crescent around the oocyte nucleus at the dorsal anterior corner starting stage 8. In a UAS-Dmn background (C,D), there is mislocalized grk*gfp in the oocyte cytoplasm accompanied by large aggregates that can also be seen in the nurse cells (arrow). Under these conditions, a significant proportion of grk*gfp is capable of reaching its destination both at the posterior and dorsal-anterior corner. In hypomorphic alleles (E,F), grk*gfp has a very diffuse localization during young stages and accumulates at the anterior cortex in later stages. Similar to (C,D), there are aggregates in both the nurse cell and oocyte cytoplasm. A defined area of fluorescent grk at the posterior and dorsal-anterior was bleached during FRAP experiments. (G) In control early stages (6-7), grk*gfp has on average 67% fluorescence recovery and during mid-stages (l8 to e9) a 46% recovery during FRAP experiments, whereas by contrast a significant reduction occurs in both young and mid stages when dynein activity is compromised. When Dmn is overexpressed, recovery is reduced to 34% in stages 6-7 and 28% in l8-e9. Similarly, in the dynein hypomorphs, recovery is reduced to 24% in stage l8-e9 egg chambers. A minimum of five FRAP experiments were performed for each data point. Error bars represent standard deviations. not appear to be from prelocalized sources of grk mrna. It appears to occur instead from nearby cytoplasm. At least at stage 7, the recovery is not due to nurse cells. At later stages, the low amount of recovery could represent grk mrna in transit from the nurse cells (already close to its destination) (Clark et al., 2007) or this could also be explained by continual active transport. Further studies are needed to distinguish between these possibilities. The difference in dynamics of grk mrna between young and old oocytes is altered in K10 and sqd 1 mutants. In fact, during the later stages, we found an increase in recovery in the mutants, suggesting the existence of a more dynamic grk mrna at the anterior of the oocytes. The rate of recovery also does not slow down and is more representative of grk mrna in a younger oocyte. It might be that, in addition to a defect in the machinery needed for proper localization (Norvell et al., 1999), the transition to a more stable anchorage is impaired in these mutants. In fact, in a recent publication, Delanoue and colleagues (Delanoue et al., 2007) suggest that Squid is a necessary component for the conversion of grk transport particles to a more stable structure. In addition, we found that compromising the activity of dynein by either overexpressing Dmn or using hypomorphic alleles of the dynein heavy chain led to less fluorescence recovery in both young and mid stages. Although we could not directly distinguish the roles of dynein in either long-range transport or anchorage, we have demonstrated that, when dynein activity is reduced, localized grk mrna is less dynamic. Using the MS2-MCP system, we have been able to visualize grk mrna in live egg chambers and have begun to dissect the processes involved in the maintenance of localized transcripts. We show that grk mrna has a remarkably different set of dynamics between young and older egg chambers and we also measured differences in various mutant backgrounds, which allows us to define the various stages involved in the process. Materials and Methods Drosophila stocks hsp83-mcp-rfp and hsp83-mcp-gfp transgenic flies have been described previously (Forrest and Gavis, 2003; Weil et al., 2006). We used fs(1)k10, sqd 1, dhc 6 10 and dhc 6 12 mutant strains (Kelley, 1993; Li et al., 1994; Wieschaus, 1978). Antoine Guichet provided the UAS-Dmn flies (Guichet et al., 2001). Overexpression of Dmn was achieved by using the αtubgal4 driver that begins to be active during stage 4 (matαgal4-67). For marker mutations, Gal4 lines and balancer chromosomes, see flybase@indiana.edu. Transgene construction A PmlI site was introduced at 1603 bp of the mrna, within the 3 UTR of grk mrna. This site is 343 bp after the stop codon. The mutagenesis was performed on a 5050 bp genomic fragment containing the complete grk locus (Queenan et al., 1999) with adjacent 5 and 3 sequences within the pbluescript II SK+ phagemid (Stratagene). The psl-ms2-12 construct contains 12 MS2 stem loops (a gift from K. Forrest). We used a mutated version of the stem loop that results in a very strong affinity (K d =10 11 ) of MS2 coat protein to the MS2 stem loop (Johansson et al., 1998; Valegard et al., 1997). The 693 bp BamHI-EcoRV fragment from psl-ms2-12 was end-filled with klenow (NE Biolabs) and placed into the PmlI site of grk. A XhoI-EagI grk-ms2-12 fragment was cloned into pcasper4 using XhoI-NotI. P-element-mediated transformation P-element transformation was performed according to standard procedures (Spradling and Rubin, 1982) using an Eppendorf Transjector 5246 with Eppendorf Femtotips (Eppendorf). Transgene constructs were injected at a concentration of 0.4 μg/μl along with the helper plasmid pturbo at a concentration of 0.1 μg/μl. Live imaging and inhibitor treatment Ovaries were dissected in Schneider s Drosophila Medium (Gibco). Egg chambers were placed in No. 1.5 glass-bottom culture dishes (MatTek) with 200 μl of medium. A No. 1.5 micro cover glass (VWR) was cut with a diamond tip to a coverslip of about 4 mm 2 and placed on top of the dissected egg chambers. Egg chambers were treated with either 200 μg/ml colchicine and 100 μg/ml colcemid (Sigma) or 4.2
8 (6) μg/ml latrunculin A and 100 μg/ml cytochalasin D (Sigma) for minutes while in the glass-bottom dishes. FRAP experiments FRAP experiments were performed using a Zeiss LSM510 confocal microscope with a NA water objective lens. Imaging was performed with a 488 nm argon laser. The following conditions gave minimal photobleaching: 70% laser output, 1.5% AOTF, zoom 4, scan speed 6, 10 seconds per frame. For each FRAP experiment, 1-3 pre-bleach images were taken. A defined region of interest (ROI), a 2.25 μm 2 region, was bleached at 80% power with 15 iterations (~4 seconds). After bleaching, a time series was taken every 10 seconds for a total of 10 minutes. Analysis was done using MATLAB and ImageJ (NIH). In order to test for the survival and transport properties of the egg chamber, we bleached the entire oocyte of stage 8 GFP-Spn-F egg chambers (Abdu et al., 2006) and monitored the recovery of Spn-F protein for 20 minutes. GFP-Spn-F is transported from nurse cells to the oocyte and, under our experimental conditions, significant recovery occurs (data not shown). Feeding inhibitor drugs Flies were starved for 24 hours. After starvation, the flies were placed in a vial for 24 hours containing fly food media topped with a coated layer of yeast and inhibitor drugs (200 μg/ml colchicine and 100 μg/ml colcemid or 4.2 μg/ml latrunculin A and 100 μg/ml cytochalasin D). Ovaries were then dissected and fixed with 4% paraformaldehyde. We thank Kevin Forrest for technical advice and reagents, Thomas Gregor for help with MATLAB analysis and Antoine Guichet for flies. We also thank the members of the Wieschaus and Schupbach labs for their insightful critiques of the work involved in the manuscript. This work was supported by the Howard Hughes Medical Institute (A.M.J. and T.S.) and by a grant from the National Institutes of Health (GM067758) to E.R.G. References Abdu, U., Bar, D. and Schupbach, T. (2006). spn-f encodes a novel protein that affects oocyte patterning and bristle morphology in Drosophila. Development 133, Baas, P. W., Pienkowski, T. P., Cimbalnik, K. A., Toyama, K., Bakalis, S., Ahmad, F. J. and Kosik, K. S. (1994). Tau confers drug stability but not cold stability to microtubules in living cells. J. Cell Sci. 107, Babu, K., Cai, Y., Bahri, S., Yang, X. and Chia, W. (2004). Roles of Bifocal, Homer, and F-actin in anchoring Oskar to the posterior cortex of Drosophila oocytes. Genes Dev. 18, Bannigan, A., Wiedemeier, A. M., Williamson, R. E., Overall, R. L. and Baskin, T. I. (2006). Cortical microtubule arrays lose uniform alignment between cells and are oryzalin resistant in the Arabidopsis mutant, radially swollen 6. Plant Cell Physiol. 47, Berleth, T., Burri, M., Thoma, G., Bopp, D., Richstein, S., Frigerio, G., Noll, M. and Nusslein-Volhard, C. (1988). The role of localization of bicoid RNA in organizing the anterior pattern of the Drosophila embryo. EMBO J. 7, Bertrand, E., Chartrand, P., Schaefer, M., Shenoy, S. M., Singer, R. H. and Long, R. M. (1998). Localization of ASH1 mrna particles in living yeast. Mol. Cell 2, Brendza, R. P., Serbus, L. R., Saxton, W. M. and Duffy, J. B. (2002). Posterior localization of dynein and dorsal-ventral axis formation depend on kinesin in Drosophila oocytes. Curr. Biol. 12, Clark, A., Meignin, C. and Davis, I. (2007). A Dynein-dependent shortcut rapidly delivers axis determination transcripts into the Drosophila oocyte. Development 134, Delanoue, R. and Davis, I. (2005). Dynein anchors its mrna cargo after apical transport in the Drosophila blastoderm embryo. Cell 122, Delanoue, R., Herpers, B., Soetaert, J., Davis, I. and Rabouille, C. (2007). Drosophila Squid/hnRNP helps Dynein switch from a gurken mrna transport motor to an ultrastructural static anchor in sponge bodies. Dev. Cell 13, Duncan, J. E. and Warrior, R. (2002). The cytoplasmic dynein and kinesin motors have interdependent roles in patterning the Drosophila oocyte. Curr. Biol. 12, Ephrussi, A., Dickinson, L. K. and Lehmann, R. (1991). Oskar organizes the germ plasm and directs localization of the posterior determinant nanos. Cell 66, Forrest, K. M. and Gavis, E. R. (2003). Live imaging of endogenous RNA reveals a diffusion and entrapment mechanism for nanos mrna localization in Drosophila. Curr. Biol. 13, Goodrich, J. S., Clouse, K. N. and Schupbach, T. (2004). Hrb27C, Sqd and Otu cooperatively regulate gurken RNA localization and mediate nurse cell chromosome dispersion in Drosophila oogenesis. Development 131, Guichet, A., Peri, F. and Roth, S. (2001). Stable anterior anchoring of the oocyte nucleus is required to establish dorsoventral polarity of the Drosophila egg. Dev. Biol. 237, Guillaud, L., Bosc, C., Fourest-Lieuvin, A., Denarier, E., Pirollet, F., Lafanechere, L. and Job, D. (1998). STOP proteins are responsible for the high degree of microtubule stabilization observed in neuronal cells. J. Cell Biol. 142, Gutzeit, H. O. (1982). Time-lapse film analysis of cytoplasmic streaming during late oogenesis of Drosophila. J. Embryol. Exp. Morphol. 67, Hudson, A. M. and Cooley, L. (2002). Understanding the function of actin-binding proteins through genetic analysis of Drosophila oogenesis. Annu. Rev. Genet. 36, Jankovics, F., Sinka, R., Lukacsovich, T. and Erdelyi, M. (2002). MOESIN crosslinks actin and cell membrane in Drosophila oocytes and is required for OSKAR anchoring. Curr. Biol. 12, Januschke, J., Gervais, L., Dass, S., Kaltschmidt, J. A., Lopez-Schier, H., St Johnston, D., Brand, A. H., Roth, S. and Guichet, A. (2002). Polar transport in the Drosophila oocyte requires Dynein and Kinesin I cooperation. Curr. Biol. 12, Johansson, H. E., Dertinger, D., LeCuyer, K. A., Behlen, L. S., Greef, C. H. and Uhlenbeck, O. C. (1998). A thermodynamic analysis of the sequence-specific binding of RNA by bacteriophage MS2 coat protein. Proc. Natl. Acad. Sci. USA 95, Kamal, A. and Goldstein, L. S. (2002). Principles of cargo attachment to cytoplasmic motor proteins. Curr. Opin. Cell Biol. 14, Kelley, R. L. (1993). Initial organization of the Drosophila dorsoventral axis depends on an RNA-binding protein encoded by the squid gene. Genes Dev. 7, Kloc, M. and Etkin, L. D. (2005). RNA localization mechanisms in oocytes. J. Cell Sci. 118, Lehmann, R. and Nusslein-Volhard, C. (1986). Abdominal segmentation, pole cell formation, and embryonic polarity require the localized activity of oskar, a maternal gene in Drosophila. Cell 47, Li, M., McGrail, M., Serr, M. and Hays, T. S. (1994). Drosophila cytoplasmic dynein, a microtubule motor that is asymmetrically localized in the oocyte. J. Cell Biol. 126, MacDougall, N., Clark, A., MacDougall, E. and Davis, I. (2003). Drosophila gurken (TGFalpha) mrna localizes as particles that move within the oocyte in two dyneindependent steps. Dev. Cell 4, Manseau, L. J. and Schupbach, T. (1989). cappuccino and spire: two unique maternaleffect loci required for both the anteroposterior and dorsoventral patterns of the Drosophila embryo. Genes Dev. 3, Neuman-Silberberg, F. S. and Schupbach, T. (1993). The Drosophila dorsoventral patterning gene gurken produces a dorsally localized RNA and encodes a TGF alphalike protein. Cell 75, Neuman-Silberberg, F. S. and Schupbach, T. (1994). Dorsoventral axis formation in Drosophila depends on the correct dosage of the gene gurken. Development 120, Neuman-Silberberg, F. S. and Schupbach, T. (1996). The Drosophila TGF-alpha-like protein Gurken: expression and cellular localization during Drosophila oogenesis. Mech. Dev. 59, Norvell, A., Kelley, R. L., Wehr, K. and Schupbach, T. (1999). Specific isoforms of squid, a Drosophila hnrnp, perform distinct roles in Gurken localization during oogenesis. Genes Dev. 13, Palazzo, A., Ackerman, B. and Gundersen, G. G. (2003). Cell biology: tubulin acetylation and cell motility. Nature 421, 230. Pokrywka, N. J. and Stephenson, E. C. (1995). Microtubules are a general component of mrna localization systems in Drosophila oocytes. Dev. Biol. 167, Polesello, C., Delon, I., Valenti, P., Ferrer, P. and Payre, F. (2002). Dmoesin controls actin-based cell shape and polarity during Drosophila melanogaster oogenesis. Nat. Cell Biol. 4, Queenan, A. M., Barcelo, G., Van Buskirk, C. and Schupbach, T. (1999). The transmembrane region of Gurken is not required for biological activity, but is necessary for transport to the oocyte membrane in Drosophila. Mech. Dev. 89, Rom, I., Faicevici, A., Almog, O. and Neuman-Silberberg, F. S. (2007). Drosophila Dynein light chain (DDLC1) binds to gurken mrna and is required for its localization. Biochim. Biophys. Acta 1773, Saunders, C. and Cohen, R. S. (1999). The role of oocyte transcription, the 5 UTR, and translation repression and derepression in Drosophila gurken mrna and protein localization. Mol. Cell 3, Spradling, A. C. and Rubin, G. M. (1982). Transposition of cloned P elements into Drosophila germ line chromosomes. Science 218, St Johnston, D. (2005). Moving messages: the intracellular localization of mrnas. Nat. Rev. Mol. Cell Biol. 6, Valegard, K., Murray, J. B., Stonehouse, N. J., van den Worm, S., Stockley, P. G. and Liljas, L. (1997). The three-dimensional structures of two complexes between recombinant MS2 capsids and RNA operator fragments reveal sequence-specific protein- RNA interactions. J. Mol. Biol. 270, Van Buskirk, C. and Schupbach, T. (1999). Versatility in signaling: multiple responses to EGF receptor activation during Drosophila oogenesis. Trends Cell Biol. 9, 1-4. Weil, T. T., Forrest, K. M. and Gavis, E. R. (2006). Localization of bicoid mrna in late oocytes is maintained by continual active transport. Dev. Cell 11, Wieschaus, E., Marsh, J. L. and Gehring, W. (1978). fs(1)k10, a germline-dependent female sterile mutation causing abnormal chorion morphology in Drosophila melanogaster. Roux s Arch. Dev. Biol. 184,
Lighting up mrna localization in Drosophila oogenesis
 REVIEW 2493 Development 136, 2493-2503 (2009) doi:10.1242/dev.032391 Lighting up mrna localization in Drosophila oogenesis Agata N. Becalska and Elizabeth R. Gavis* The asymmetric localization of four
REVIEW 2493 Development 136, 2493-2503 (2009) doi:10.1242/dev.032391 Lighting up mrna localization in Drosophila oogenesis Agata N. Becalska and Elizabeth R. Gavis* The asymmetric localization of four
Gratuitous mrna localization in the Drosophila oocyte
 Development 121, 3013-3021(1995) Printed in Great Britain The Company of Biologists Limited 1995 3013 Gratuitous mrna localization in the Drosophila oocyte Thomas L. Serano 1,2, and Robert S. Cohen 1,
Development 121, 3013-3021(1995) Printed in Great Britain The Company of Biologists Limited 1995 3013 Gratuitous mrna localization in the Drosophila oocyte Thomas L. Serano 1,2, and Robert S. Cohen 1,
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
In Vivo Imaging of oskar mrna Transport Reveals the Mechanism of Posterior Localization
 In Vivo Imaging of oskar mrna Transport Reveals the Mechanism of Posterior Localization Vitaly L. Zimyanin, 1,3,6 Katsiaryna Belaya, 1,6 Jacques Pecreaux, 3 Michael J. Gilchrist, 1 Alejandra Clark, 2,4
In Vivo Imaging of oskar mrna Transport Reveals the Mechanism of Posterior Localization Vitaly L. Zimyanin, 1,3,6 Katsiaryna Belaya, 1,6 Jacques Pecreaux, 3 Michael J. Gilchrist, 1 Alejandra Clark, 2,4
Drosophila Life Cycle
 Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Multiple steps in the localization of bicoid RNA to the anterior pole of the Drosophila oocyte
 Development 1989 Supplement, 13-19 Printed in Great Britain The Company of Biologists Limited 1989 13 Multiple steps in the localization of bicoid RNA to the anterior pole of the Drosophila oocyte DANIEL
Development 1989 Supplement, 13-19 Printed in Great Britain The Company of Biologists Limited 1989 13 Multiple steps in the localization of bicoid RNA to the anterior pole of the Drosophila oocyte DANIEL
Autonomous concentration-dependent activation and repression of Krüppel by hunchback in the Drosophila embryo
 Development 120, 3043-3049 (1994) Printed in Great Britain The Company of Biologists Limited 1994 3043 Autonomous concentration-dependent activation and repression of Krüppel by hunchback in the Drosophila
Development 120, 3043-3049 (1994) Printed in Great Britain The Company of Biologists Limited 1994 3043 Autonomous concentration-dependent activation and repression of Krüppel by hunchback in the Drosophila
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Analysis of Active Transport by Fluorescence Recovery after Photobleaching. MV Ciocanel, JA Kreiling, JA Gagnon, KL Mowry, B Sandstede
 Analysis of Active Transport by Fluorescence Recovery after Photobleaching MV Ciocanel, JA Kreiling, JA Gagnon, KL Mowry, B Sandstede Analysis of Active Transport by Fluorescence Recovery after Photobleaching
Analysis of Active Transport by Fluorescence Recovery after Photobleaching MV Ciocanel, JA Kreiling, JA Gagnon, KL Mowry, B Sandstede Analysis of Active Transport by Fluorescence Recovery after Photobleaching
RNA sorting in Drosophila oocytes and embryos
 RNA sorting in Drosophila oocytes and embryos PAUL LASKO 1 Departments of Biology and Anatomy and Cell Biology, McGill University, Montréal, Québec, Canada H3A 1B1 ABSTRACT Many RNAs involved in determination
RNA sorting in Drosophila oocytes and embryos PAUL LASKO 1 Departments of Biology and Anatomy and Cell Biology, McGill University, Montréal, Québec, Canada H3A 1B1 ABSTRACT Many RNAs involved in determination
Profilin is required for posterior patterning of the Drosophila oocyte
 Development 122, 2109-2116 (1996) Printed in Great Britain The Company of Biologists Limited 1996 DEV5067 2109 Profilin is required for posterior patterning of the Drosophila oocyte Lynn Manseau*, John
Development 122, 2109-2116 (1996) Printed in Great Britain The Company of Biologists Limited 1996 DEV5067 2109 Profilin is required for posterior patterning of the Drosophila oocyte Lynn Manseau*, John
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Role for mrna localization in translational activation but not spatial restriction of nanos RNA
 Development 126, 659-669 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV7673 659 Role for mrna localization in translational activation but not spatial restriction of nanos RNA
Development 126, 659-669 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV7673 659 Role for mrna localization in translational activation but not spatial restriction of nanos RNA
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Neurite formation & neuronal polarization. The cytoskeletal components of neurons have characteristic distributions and associations
 Mechanisms of neuronal migration & Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 The cytoskeletal
Mechanisms of neuronal migration & Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 The cytoskeletal
TRANSLATIONAL REGULATION AND RNA LOCALIZATION IN DROSOPHILA OOCYTES AND EMBRYOS
 Annu. Rev. Genet. 2001. 35:365 406 Copyright c 2001 by Annual Reviews. All rights reserved TRANSLATIONAL REGULATION AND RNA LOCALIZATION IN DROSOPHILA OOCYTES AND EMBRYOS Oona Johnstone and Paul Lasko
Annu. Rev. Genet. 2001. 35:365 406 Copyright c 2001 by Annual Reviews. All rights reserved TRANSLATIONAL REGULATION AND RNA LOCALIZATION IN DROSOPHILA OOCYTES AND EMBRYOS Oona Johnstone and Paul Lasko
MCB 141 Midterm I Feb. 14, 2012
 Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 14, 2012 Question #1 Question #2 Question #3 Question #4 BONUS / 28 pts / 27 pts / 25 pts / 20 pts / 1 pt TOTAL / 100 pts
Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 14, 2012 Question #1 Question #2 Question #3 Question #4 BONUS / 28 pts / 27 pts / 25 pts / 20 pts / 1 pt TOTAL / 100 pts
Nuclear import of DNA: genetic modification of plants
 Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
Nuclear import of DNA: genetic modification of plants gene delivery by Agrobacterium tumefaciens T. Tzfira & V. Citovsky. 2001. Trends in Cell Biol. 12: 121-129 VirE2 binds ssdna in vitro forms helical
DIFFERENTIATION MORPHOGENESIS GROWTH HOW CAN AN IDENTICAL SET OF GENETIC INSTRUCTIONS PRODUCE DIFFERENT TYPES OF CELLS?
 DIFFERENTIATION HOW CAN AN IDENTICAL SET OF GENETIC INSTRUCTIONS PRODUCE DIFFERENT TYPES OF CELLS? MORPHOGENESIS HOW CAN CELLS FORM ORDERED STRUCTURES? GROWTH HOW DO OUR CELLS KNOW WHEN TO STOP DIVIDING
DIFFERENTIATION HOW CAN AN IDENTICAL SET OF GENETIC INSTRUCTIONS PRODUCE DIFFERENT TYPES OF CELLS? MORPHOGENESIS HOW CAN CELLS FORM ORDERED STRUCTURES? GROWTH HOW DO OUR CELLS KNOW WHEN TO STOP DIVIDING
MOVING MESSAGES: THE INTRACELLULAR LOCALIZATION OF mrnas
 REVIEWS MOVING MESSAGES: THE INTRACELLULAR LOCALIZATION OF mrnas Daniel St Johnston Abstract mrna localization is a common mechanism for targeting proteins to regions of the cell where they are required.
REVIEWS MOVING MESSAGES: THE INTRACELLULAR LOCALIZATION OF mrnas Daniel St Johnston Abstract mrna localization is a common mechanism for targeting proteins to regions of the cell where they are required.
AP3162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions
 AP162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions Hyun Youk Delft University of Technology (Dated: March 15, 2018) In this lecture, we will derive the Berg-Purcell
AP162D: Lecture 4 - Basic modelling frameworks for developmental biology and cell-fate decisions Hyun Youk Delft University of Technology (Dated: March 15, 2018) In this lecture, we will derive the Berg-Purcell
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
The role of membranes and membrane trafficking in RNA localization
 Biol. Cell (2005) 97, 5 18 (Printed in Great Britain) Review The role of membranes and membrane trafficking in RNA localization Robert S. Cohen 1 Department of Molecular Biosciences, University of Kansas,
Biol. Cell (2005) 97, 5 18 (Printed in Great Britain) Review The role of membranes and membrane trafficking in RNA localization Robert S. Cohen 1 Department of Molecular Biosciences, University of Kansas,
REVIEW mrna Localisation during Development
 DEVELOPMENTAL BIOLOGY 172, 377 395 (1995) REVIEW mrna Localisation during Development David R. Micklem Wellcome/CRC Institute and Department of Genetics, University of Cambridge, Tennis Court Road, Cambridge
DEVELOPMENTAL BIOLOGY 172, 377 395 (1995) REVIEW mrna Localisation during Development David R. Micklem Wellcome/CRC Institute and Department of Genetics, University of Cambridge, Tennis Court Road, Cambridge
Cellular automata for exploring gene regulation in Drosophila segmentation
 Cellular automata for exploring gene regulation in Drosophila segmentation Matthew J. Berryman a, Andrew Allison a, and Derek Abbott a a Centre for Biomedical Engineering and School of Electrical and Electronic
Cellular automata for exploring gene regulation in Drosophila segmentation Matthew J. Berryman a, Andrew Allison a, and Derek Abbott a a Centre for Biomedical Engineering and School of Electrical and Electronic
Pathways for mrna localization in the cytoplasm
 Review TRENDS in Biochemical Sciences Vol.31 No.12 Pathways for mrna localization in the cytoplasm Kevin Czaplinski and Robert H. Singer Department of Anatomy and Structural Biology, Albert Einstein College
Review TRENDS in Biochemical Sciences Vol.31 No.12 Pathways for mrna localization in the cytoplasm Kevin Czaplinski and Robert H. Singer Department of Anatomy and Structural Biology, Albert Einstein College
Cell Biology Review. The key components of cells that concern us are as follows: 1. Nucleus
 Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Chapter 16. Cellular Movement: Motility and Contractility. Lectures by Kathleen Fitzpatrick Simon Fraser University Pearson Education, Inc.
 Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Drosophila germline stem cells
 Development 125, 679-690 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3776 679 Nanos and Pumilio have critical roles in the development and function of Drosophila germline
Development 125, 679-690 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3776 679 Nanos and Pumilio have critical roles in the development and function of Drosophila germline
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Neurite formation & neuronal polarization
 Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 An immature neuron in cell culture first sprouts
Neurite formation & neuronal polarization Paul Letourneau letou001@umn.edu Chapter 16; The Cytoskeleton; Molecular Biology of the Cell, Alberts et al. 1 An immature neuron in cell culture first sprouts
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
Extranuclear Inheritance
 Extranuclear Inheritance Extranuclear Inheritance The past couple of lectures, we ve been exploring exceptions to Mendel s principles of transmission inheritance. Scientists have observed inheritance patterns
Extranuclear Inheritance Extranuclear Inheritance The past couple of lectures, we ve been exploring exceptions to Mendel s principles of transmission inheritance. Scientists have observed inheritance patterns
Principles of Experimental Embryology
 Biology 4361 Developmental Biology Principles of Experimental Embryology June 16, 2008 Overview What forces affect embryonic development? The embryonic environment: external and internal How do forces
Biology 4361 Developmental Biology Principles of Experimental Embryology June 16, 2008 Overview What forces affect embryonic development? The embryonic environment: external and internal How do forces
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
The Microscopic Observation of Mitosis in Plant and Animal Cells
 The Microscopic Observation of Mitosis in Plant and Animal Cells Prelab Assignment Before coming to lab, read carefully the introduction and the procedures for each part of the experiment, and then answer
The Microscopic Observation of Mitosis in Plant and Animal Cells Prelab Assignment Before coming to lab, read carefully the introduction and the procedures for each part of the experiment, and then answer
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
DOI: 10.1038/ncb2647 Figure S1 Other Rab GTPases do not co-localize with the ER. a, Cos-7 cells cotransfected with an ER luminal marker (either KDEL-venus or mch-kdel) and mch-tagged human Rab5 (mch-rab5,
LIST of SUPPLEMENTARY MATERIALS
 LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
LIST of SUPPLEMENTARY MATERIALS Mir et al., Dense Bicoid Hubs Accentuate Binding along the Morphogen Gradient Supplemental Movie S1 (Related to Figure 1). Movies corresponding to the still frames shown
Neurite initiation. Neurite formation begins with a bud that sprouts from the cell body. One or several neurites can sprout at a time.
 Neurite initiation. Neuronal maturation initiation f-actin polarization and maturation tubulin stage 1: "spherical" neuron stage 2: neurons extend several neurites stage 3: one neurite accelerates its
Neurite initiation. Neuronal maturation initiation f-actin polarization and maturation tubulin stage 1: "spherical" neuron stage 2: neurons extend several neurites stage 3: one neurite accelerates its
Report. Live Imaging of Bicoid-Dependent Transcription in Drosophila Embryos
 Current Biology 23, 2135 2139, November 4, 2013 ª2013 Elsevier Ltd All rights reserved http://dx.doi.org/10.1016/j.cub.2013.08.053 Live Imaging of Bicoid-Dependent Transcription in Drosophila Embryos Report
Current Biology 23, 2135 2139, November 4, 2013 ª2013 Elsevier Ltd All rights reserved http://dx.doi.org/10.1016/j.cub.2013.08.053 Live Imaging of Bicoid-Dependent Transcription in Drosophila Embryos Report
GENES AND CHROMOSOMES III. Lecture 5. Biology Department Concordia University. Dr. S. Azam BIOL 266/
 GENES AND CHROMOSOMES III Lecture 5 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION OPERONS Introduction All cells in a multi-cellular
GENES AND CHROMOSOMES III Lecture 5 BIOL 266/4 2014-15 Dr. S. Azam Biology Department Concordia University CELL NUCLEUS AND THE CONTROL OF GENE EXPRESSION OPERONS Introduction All cells in a multi-cellular
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins
 13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
13-3. Synthesis-Secretory pathway: Sort lumenal proteins, Secrete proteins, Sort membrane proteins Molecular sorting: specific budding, vesicular transport, fusion 1. Why is this important? A. Form and
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Reciprocal localization of Nod and kinesin fusion proteins indicates microtubule polarity in the Drosophila oocyte, epithelium, neuron and muscle
 Development 124, 461-470 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV9498 461 Reciprocal localization of Nod and kinesin fusion proteins indicates microtubule polarity in
Development 124, 461-470 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV9498 461 Reciprocal localization of Nod and kinesin fusion proteins indicates microtubule polarity in
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
AP Biology. Biology is the only subject in which multiplication is the same thing as division. The Cell Cycle: Cell Growth, Cell Division
 QuickTime and and a TIFF TIFF (Uncompressed) decompressor are are needed needed to to see see this this picture. picture. Biology is the only subject in which multiplication is the same thing as division
QuickTime and and a TIFF TIFF (Uncompressed) decompressor are are needed needed to to see see this this picture. picture. Biology is the only subject in which multiplication is the same thing as division
Supplemental material
 Supplemental material THE JOURNAL OF CELL BIOLOGY Civelekoglu-Scholey et al., http://www.jcb.org/cgi/content/full/jcb.200908150/dc1 Figure S1. Spindle dynamics in Drosophila embryos and in vitro gliding
Supplemental material THE JOURNAL OF CELL BIOLOGY Civelekoglu-Scholey et al., http://www.jcb.org/cgi/content/full/jcb.200908150/dc1 Figure S1. Spindle dynamics in Drosophila embryos and in vitro gliding
7.32/7.81J/8.591J. Rm Rm (under construction) Alexander van Oudenaarden Jialing Li. Bernardo Pando. Rm.
 Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
Introducing... 7.32/7.81J/8.591J Systems Biology modeling biological networks Lectures: Recitations: ti TR 1:00-2:30 PM W 4:00-5:00 PM Rm. 6-120 Rm. 26-204 (under construction) Alexander van Oudenaarden
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha
 Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Gene expression in prokaryotic and eukaryotic cells, Plasmids: types, maintenance and functions. Mitesh Shrestha Plasmids 1. Extrachromosomal DNA, usually circular-parasite 2. Usually encode ancillary
Eukaryotic Gene Expression
 Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Stability and Nuclear Dynamics of the Bicoid Morphogen Gradient
 Stability and Nuclear Dynamics of the Bicoid Morphogen Gradient Thomas Gregor, 1,2,3,4, * Eric F. Wieschaus, 3,4 Alistair P. McGregor, 5 William Bialek, 1,2 and David W. Tank 1,2,3 1 Lewis-Sigler Institute
Stability and Nuclear Dynamics of the Bicoid Morphogen Gradient Thomas Gregor, 1,2,3,4, * Eric F. Wieschaus, 3,4 Alistair P. McGregor, 5 William Bialek, 1,2 and David W. Tank 1,2,3 1 Lewis-Sigler Institute
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and
 Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
Supplementary Figure 1. Real time in vivo imaging of SG secretion. (a) SGs from Drosophila third instar larvae that express Sgs3-GFP (green) and Lifeact-Ruby (red) were imaged in vivo to visualize secretion
AP Biology Unit 6 Practice Test 1. A group of cells is assayed for DNA content immediately following mitosis and is found to have an average of 8
 AP Biology Unit 6 Practice Test Name: 1. A group of cells is assayed for DNA content immediately following mitosis and is found to have an average of 8 picograms of DNA per nucleus. How many picograms
AP Biology Unit 6 Practice Test Name: 1. A group of cells is assayed for DNA content immediately following mitosis and is found to have an average of 8 picograms of DNA per nucleus. How many picograms
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes
 9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
Chromosome Chr Duplica Duplic t a ion Pixley
 Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila
 Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila Melissa M. Rolls and Chris Q. Doe, Inst. Neurosci and Inst. Mol. Biol., HHMI, Univ. Oregon, Eugene, Oregon 97403 Correspondence
Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila Melissa M. Rolls and Chris Q. Doe, Inst. Neurosci and Inst. Mol. Biol., HHMI, Univ. Oregon, Eugene, Oregon 97403 Correspondence
Name Chapter 10: Chromosomes, Mitosis, and Meiosis Mrs. Laux Take home test #7 DUE: MONDAY, NOVEMBER 16, 2009 MULTIPLE CHOICE QUESTIONS
 MULTIPLE CHOICE QUESTIONS 1. A bacterial chromosome consists of: A. a linear DNA molecule many times larger than the cell. B. a circular DNA molecule many times larger than the cell. C. a circular DNA
MULTIPLE CHOICE QUESTIONS 1. A bacterial chromosome consists of: A. a linear DNA molecule many times larger than the cell. B. a circular DNA molecule many times larger than the cell. C. a circular DNA
Name 8 Cell Cycle and Meiosis Test Date Study Guide You must know: The structure of the replicated chromosome. The stages of mitosis.
 Name 8 Cell Cycle and Meiosis Test Date Study Guide You must know: The structure of the replicated chromosome. The stages of mitosis. The role of kinases and cyclin in the regulation of the cell cycle.
Name 8 Cell Cycle and Meiosis Test Date Study Guide You must know: The structure of the replicated chromosome. The stages of mitosis. The role of kinases and cyclin in the regulation of the cell cycle.
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Caenorhabditis elegans
 Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
178 Part 3.2 SUMMARY INTRODUCTION
 178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
FREEMAN MEDIA INTEGRATION GUIDE Chapter 7: Inside the Cell
 FREEMAN MEDIA INTEGRATION GUIDE Chapter 7: Inside the Cell All media is on the Instructors Resource CD/DVD JPEG Resources Figures, Photos, and Tables PowerPoint Resources Chapter Outline with Figures Lecture
FREEMAN MEDIA INTEGRATION GUIDE Chapter 7: Inside the Cell All media is on the Instructors Resource CD/DVD JPEG Resources Figures, Photos, and Tables PowerPoint Resources Chapter Outline with Figures Lecture
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Three different fusions led to three basic ideas: 1) If one fuses a cell in mitosis with a cell in any other stage of the cell cycle, the chromosomes
 Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Figure 1. Automated 4D detection and tracking of endosomes and polarity markers.
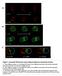 A t = 0 z t = 90 B t = 0 z t = 90 C t = 0-90 t = 0-180 t = 0-300 Figure 1. Automated 4D detection and tracking of endosomes and polarity markers. A: Three different z planes of a dividing SOP shown at
A t = 0 z t = 90 B t = 0 z t = 90 C t = 0-90 t = 0-180 t = 0-300 Figure 1. Automated 4D detection and tracking of endosomes and polarity markers. A: Three different z planes of a dividing SOP shown at
Shavenbaby Couples Patterning to Epidermal Cell Shape Control. Chanut-Delalande H, Fernandes I, Roch F, Payre F, Plaza S (2006) PLoS Biol 4(9): e290
 Shavenbaby Couples Patterning to Epidermal Cell Shape Control. Chanut-Delalande H, Fernandes I, Roch F, Payre F, Plaza S (2006) PLoS Biol 4(9): e290 Question (from Introduction): How does svb control the
Shavenbaby Couples Patterning to Epidermal Cell Shape Control. Chanut-Delalande H, Fernandes I, Roch F, Payre F, Plaza S (2006) PLoS Biol 4(9): e290 Question (from Introduction): How does svb control the
7-2 Eukaryotic Cell Structure
 1 of 49 Comparing the Cell to a Factory Eukaryotic Cell Structures Structures within a eukaryotic cell that perform important cellular functions are known as organelles. Cell biologists divide the eukaryotic
1 of 49 Comparing the Cell to a Factory Eukaryotic Cell Structures Structures within a eukaryotic cell that perform important cellular functions are known as organelles. Cell biologists divide the eukaryotic
Supplementary Figure 1. Phenotype of the HI strain.
 Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Supplementary Figure 1. Phenotype of the HI strain. (A) Phenotype of the HI and wild type plant after flowering (~1month). Wild type plant is tall with well elongated inflorescence. All four HI plants
Cell Structure and Function. The Development of Cell Theory
 Cell Structure and Function The Development of Cell Theory Hooke's original microscope, that he used at Oxford University, in 1665. Robert Hooke first examined a thin piece of cork (see below), and coined
Cell Structure and Function The Development of Cell Theory Hooke's original microscope, that he used at Oxford University, in 1665. Robert Hooke first examined a thin piece of cork (see below), and coined
MCB 141 Midterm I Feb. 19, 2009
 Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 19, 2009 Circle the name of your TA Jessica Lyons Alberto Stolfi Question #1 Question #2 Question #3 Question #4 TOTAL
Write your name and student ID# on EVERY PAGE of your exam MCB 141 Midterm I Feb. 19, 2009 Circle the name of your TA Jessica Lyons Alberto Stolfi Question #1 Question #2 Question #3 Question #4 TOTAL
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Supplementary Information for. Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria
 Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Supplementary Information for Single-cell dynamics of the chromosome replication and cell division cycles in mycobacteria Isabella Santi 1 *, Neeraj Dhar 1, Djenet Bousbaine 1, Yuichi Wakamoto, John D.
Pulling forces in Cell Division
 Pulling forces in Cell Division Frank Jülicher Max Planck Institute for the Physics of Complex Systems Dresden, Germany Max Planck Institute for the Physics of Complex Systems A. Zumdieck A. J.-Dalmaroni
Pulling forces in Cell Division Frank Jülicher Max Planck Institute for the Physics of Complex Systems Dresden, Germany Max Planck Institute for the Physics of Complex Systems A. Zumdieck A. J.-Dalmaroni
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Supplementary Figure 1.
 Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Supplementary Figure 1. Characterisation of IHG-1 overexpressing and knockdown cell lines. (A) Total cellular RNA was prepared from HeLa cells stably overexpressing IHG-1 or mts-ihg-1. IHG-1 mrna was quantified
Development Team. Developmental Biology Axis Specification in Drosophila. Head, Department of Zoology, University of Delhi
 Paper No. : 11 Module : 6 Development Team Principal Investigator: Prof. Neeta Sehgal Head, Department of Zoology, University of Delhi Paper Coordinator: Prof. Namita Agrawal Department of Zoology, University
Paper No. : 11 Module : 6 Development Team Principal Investigator: Prof. Neeta Sehgal Head, Department of Zoology, University of Delhi Paper Coordinator: Prof. Namita Agrawal Department of Zoology, University
Homeotic genes in flies. Sem 9.3.B.6 Animal Science
 Homeotic genes in flies Sem 9.3.B.6 Animal Science So far We have seen that identities of each segment is determined by various regulators of segment polarity genes In arthopods, and in flies, each segment
Homeotic genes in flies Sem 9.3.B.6 Animal Science So far We have seen that identities of each segment is determined by various regulators of segment polarity genes In arthopods, and in flies, each segment
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle. Chapter 12
 The Cell Cycle Chapter 12 Why are cells small? As cells get bigger they don t work as well WHY? Difficulties Larger Cells Have: More demands on its DNA Less efficient in moving nutrients/waste across its
The Cell Cycle Chapter 12 Why are cells small? As cells get bigger they don t work as well WHY? Difficulties Larger Cells Have: More demands on its DNA Less efficient in moving nutrients/waste across its
E. Incorrect! At telophase II, cells are nearly completed with meiosis, with no cross-over.
 OAT Biology - Problem Drill 06: Mitosis and Meiosis Question No. 1 of 10 1. During meiosis, cross-over between homologous chromosomes occurs at the end of. Question #01 (A) Anaphase II (B) Metaphase I
OAT Biology - Problem Drill 06: Mitosis and Meiosis Question No. 1 of 10 1. During meiosis, cross-over between homologous chromosomes occurs at the end of. Question #01 (A) Anaphase II (B) Metaphase I
