Canonical Wnt signaling through Lef1 is required for hypothalamic neurogenesis
|
|
|
- Garey Dawson
- 5 years ago
- Views:
Transcription
1 RESEARCH ARTICLE 4451 Development 133, (2006) doi: /dev Canonical Wnt signaling through Lef1 is required for hypothalamic neurogenesis Ji Eun Lee 1, Shan-Fu Wu 1, Lisa M. Goering 2 and Richard I. Dorsky 1, * Although the functional importance of the hypothalamus has been demonstrated throughout vertebrates, the mechanisms controlling neurogenesis in this forebrain structure are poorly understood. We report that canonical Wnt signaling acts through Lef1 to regulate neurogenesis in the zebrafish hypothalamus. We show that Lef1 is required for proneural and neuronal gene expression, and for neuronal differentiation in the posterior hypothalamus. Furthermore, we find that this process is dependent on Wnt8b, a ligand of the canonical pathway expressed in the posterior hypothalamus, and that both Wnt8b and Lef1 act to mediate -catenin-dependent transcription in this region. Finally, we show that Lef1 associates in vivo with the promoter of sox3, which depends on Lef1 for its expression and can rescue neurogenesis in the absence of Lef1. The conserved presence of this pathway in other vertebrates suggests a common mechanism for regulating hypothalamic neurogenesis. KEY WORDS: Zebrafish, Wnt, Lef1, Hypothalamus INTRODUCTION The hypothalamus is an evolutionarily conserved vertebrate brain structure responsible for regulation of the autonomic nervous system and endocrine hormone production. Although many specific neuronal populations in the adult hypothalamus have been well characterized, relatively little is known about the process through which these neurons are induced and specified during development. In zebrafish, where initial hypothalamus induction and patterning has been extensively studied, these events primarily occur in the first 18 hours of development (Varga et al., 1999; Woo and Fraser, 1995). The hypothalamus develops from the most ventral region of the anterior diencephalon, and is induced through identified molecular pathways such as Sonic Hedgehog and Nodal signaling. Specifically, Hedgehog signaling is required for inducing the anterior hypothalamus and Nodal signaling is required for the posterior hypothalamus (Chiang et al., 1996; Mathieu et al., 2002). After initial induction and patterning, the hypothalamus is regionalized into subdomains distinguished by specific gene expression patterns (Hauptmann and Gerster, 2000), but the upstream signals responsible for these subdivisions are unknown. In zebrafish, proneural markers begin to be expressed in specific regions of the hypothalamus by 18 hours post-fertilization (Mueller and Wullimann, 2002), but it is not clear which mature neuronal populations are labeled by these markers (Guo et al., 1999; Ross et al., 1992; Wilson et al., 1990). By contrast, the anatomical identities of specific neuronal populations in the hypothalamus of larval and adult zebrafish have been well characterized (Rink and Wullimann, 2001). Therefore, there is a gap in our understanding of the developmental signaling pathways between hypothalamic patterning and the eventual functional anatomy in the hypothalamus. We are interested in the function of Wnt/ -catenin signaling in hypothalamic neurogenesis. Canonical Wnt signaling plays important roles in embryonic patterning, cell-fate determination, cell 1 Department of Neurobiology and Anatomy, University of Utah School of Medicine, Salt Lake City, UT 84105, USA. 2 Department of Genetics, North Carolina State University, Raleigh, NC 27695, USA. *Author for correspondence ( richard.dorsky@neuro.utah.edu) Accepted 5 September 2006 proliferation and cell differentiation during vertebrate development. Several previous studies have demonstrated roles for Wnt signals in specific aspects of central nervous system (CNS) formation (Logan and Nusse, 2004). In neural induction, Wnt signals from the paraxial mesoderm are required for the specification of posterior neural character (Nordstrom et al., 2002) during initial anteroposterior (AP) patterning. Later, this patterning is further refined into smaller subdivisions that also require Wnt signals from the posterior (Houart et al., 2002). Importantly, Wnt signaling induces posteriorisation during development of the zebrafish hypothalamus (Kapsimali et al., 2004). However, the required functions of canonical Wnt signals in later developmental steps are poorly understood, partly because of functional redundancy (Lekven et al., 2003). Although the roles of some specific Wnt proteins in CNS development have been characterized (Brault et al., 2001; Buckles et al., 2004; Erter et al., 2001; Houart et al., 2002; Lee et al., 2000), they have primarily been defined in the context of general brain regions, such as the cerebellum or hippocampus. Wnt genes continue to be expressed in the brain at later embryonic stages, when they have been proposed to function in neuronal maturation, synapse formation, synaptic plasticity and axon guidance (Ciani and Salinas, 2005). However, the specific downstream targets of Wnt signaling during later embryogenesis remain unclear. In particular, there is little information on what functions Wnt signaling may have in the development of particular neuronal populations. The nuclear response to canonical Wnt signals is mediated by the Lef/Tcf family of transcription factors, including lymphoid enhancer factor 1 (Lef1), which activate downstream genes by association with -catenin (Eastman and Grosschedl, 1999). All Lef/Tcf proteins have highly similar DNA and -catenin interaction domains, and there are no known differences in their affinities for these targets. In the absence of -catenin, some members of the Lef/Tcf family can repress the transcription of target genes in cooperation with co-repressors such as Groucho and CtBP (Roose and Clevers, 1999). However, identified isoforms of Lef1 in zebrafish embryos lack a putative co-repressor interacting domain (Dorsky et al., 1999), and cannot substitute for the repressor function of Tcf3 in AP patterning (Dorsky et al., 2003), suggesting that Lef1 may function only as a transcriptional activator in the presence of -catenin. Of the identified Lef/Tcf
2 4452 RESEARCH ARTICLE Development 133 (22) family members, only Lef1 has thus far been shown to play a required role in CNS neurogenesis (Galceran et al., 2000; van Genderen et al., 1994). In zebrafish, Lef1 is expressed in multiple tissues during embryonic development, including the CNS (Dorsky et al., 1999). Removal of maternal and zygotic lef1 function using a translation blocking morpholino oligonucleotide (MO) results in tail truncations and paraxial mesoderm defects (Dorsky et al., 2002). However, the expression and function of Lef1 at later stages in zebrafish remain uncharacterized. In the present study, we have investigated the role of Lef1 in the developing zebrafish brain using splice-blocking MOs and mutants. We show that lef1 is expressed in the posterior hypothalamus after initial patterning but before the first neurons differentiate. In addition, we find that Wnt8b is expressed appropriately to function as a specific upstream modulator of Lef1 through the canonical pathway during hypothalamic development. We demonstrate through loss-offunction experiments that Wnt8b and Lef1 are required for the development of a specific neuronal population in the posterior hypothalamus, and signal through the canonical Wnt pathway in this region. These studies address for the first time the requirement for Wnt/ -catenin signaling in hypothalamic neurogenesis. In addition, analysis of downstream targets suggests a specific role for Lef1 in this later step of CNS development, in which it regulates a neurogenesis program by activating the expression of sox3, a gene required for neural competence. MATERIALS AND METHODS Fish strains and staging Embryos were obtained from natural spawning of wild-type (AB*) or TOPdGFP (Dorsky et al., 2002) zebrafish lines. X8 deletion mutant embryos (line provided by Dr B. Riley) were identified by consistent phenotypes in 25% of embryos from crosses of heterozygous parents. All developmental stages in this study are reported in hours post-fertilization (hpf) at 28.5 C (Kimmel et al., 1995). Morpholino injections The lef1 splice-blocking morpholino antisense oligonucleotide (MO) was obtained from Gene Tools (5 -ACTGCCTGGATGAAACACTT- ACATG-3 ). The wnt8b translation-blocking MO (Riley et al., 2004) was kindly provided by Dr B. Riley. Both MOs were injected into one-cell stage wild-type or transgenic embryos at doses of 2 ng and 0.5 ng, respectively. 1994); dlx2 (Akimenko et al., 1994); isl1 (Okamoto et al., 2000); wnt8b (Kelly et al., 1995); gfp (Dorsky et al., 2002); ngn1 (Blader et al., 1997); olig2 (Park et al., 2002). Antibodies were obtained from the following sources: anti-ph3 (Upstate Biotechnology, 1:500), anti-gfp (Molecular Probes, 1:5000), anti-huc/d (Molecular Probes, 1:500), anti-acetylated Tubulin (Sigma, 1:1000) and affinity-purified rabbit anti-lef1 (Open Biosystems, 1:500). For immunostaining, embryos were fixed with 4% paraformaldehyde (PFA) for 3 hours at room temperature, and incubated with primary and secondary antibodies at 4 C overnight. For whole-mount photography after all staining methods, yolks and eyes of embryos were dissected. Hu, ph3 and AT-stained embryos were imaged on a confocal microscope, all other embryos and cryosections were imaged on a compound microscope. TUNEL staining For TUNEL analysis, 19 and 24 hpf embryos were fixed with 4% PFA for 4 hours at room temperature. Embryos were permeabilized with acetone at 20 C and washed twice with PBC (0.001% Triton X-100, 0.1% sodium citrate in PBS) for 10 minutes. Labeling for apoptotic cells was performed using In situ Cell Death Detection Kit (Roche) at 37 C for 1 hour, washed and mounted for fluorescent microscopic imaging. ChIP ChIP analysis was performed as described previously (Weinmann et al., 2001) with the following modifications. One-hundred embryos at hpf were fixed in 1.85% formaldehyde for 15 minutes at room temperature, and then lysed in cell lysis buffer [10 mm Tris (ph 8.1), 10 mm NaCl, 0.5% NP- 40, and protease inhibitors] by pipetting. For each immunoprecipitation, 5 g of Lef1 antibody was conjugated to protein A beads. The following primers were used for PCR after immunoprecipitation: sox3, 5 -AATTAGCCTTGCAGCCAATG-3 and 5 -ATCGGAAGGG- GTTTCTCAAT-3 ; ngn1, 5 -GGGCTCATTGGAGCAAGTTTGATT-3 and 5 -CGCGGTAGCCTACATTACTGCACA-3 ; nacre, 5 -GCAAT- TACCAAAGGCCCATCAGAC-3 and 5 -ACTGGCTTACGGCTAA- CTAACGTT-3. Western blotting Dechorionated embryos were homogenized in 4 sample buffer, subjected to 8% SDS-PAGE electrophoresis, and blotted onto PVDF membrane. Affinity-purified rabbit anti-lef1 serum was applied at 1:2000 dilution, and anti-rabbit IgG-HRP (Molecular Probes) was applied at 1:10,000. The secondary antibody was visualized with an ECL reaction, using standard protocols. The same blot was stripped and re-probed with rabbit anti- -catenin at 1:5000 dilution (Sigma), and the same secondary antibody. RT-PCR Fifty wild-type embryos and lef1 morphants were used for preparing RNA. Total RNA was isolated using Trizol reagent and standard protocols. Total RNA (1-5 g/ l) was reverse transcribed by either random hexamers or a gene-specific primer using the Superscript first strand synthesis kit (Invitrogen) following the manufacturer s protocol. PCR was performed for cycles using an annealing temperature of 55 C, and reactions were visualized on 1% agarose gels in TAE. RNA injections The lef1 and sox3 mrnas were synthesized from lef1-pcs2+mt and sox3- pcs2+mt plasmids, respectively, using the SP6 mmessage mmachine transcription kit (Ambion). For mrna rescue experiments, 100 pg of lef1 mrna and 20 pg of sox3 mrna were injected into one-cell stage wild-type embryos together with or without 2 ng of lef1 MO. In situ hybridization and immunohistochemistry Probe synthesis and in situ hybridization were performed as described elsewhere (Oxtoby and Jowett, 1993). Single and double in situ hybridizations were carried out using digoxigenin- or fluorescein-labeled antisense RNA probes (Jowett, 2001) and visualized using BM Purple and Fast Red (Roche). The following RNA probes were used: lef1 (Dorsky et al., 1999); nk2.1a (Rohr et al., 2001); rx3 (Chuang et al., 1999); emx2 (Morita et al., 1995); sox3 (Kudoh et al., 2004); zash1a (Allende and Weinberg, RESULTS The expression of lef1 suggests a role in hypothalamic development Although lef1 is expressed both maternally and zygotically in early zebrafish embryos (Dorsky et al., 1999), the expression pattern at later embryonic stages has not been characterized. To assess later roles of lef1 during brain development, we examined mrna expression during late somitogenesis stages by in situ hybridization (Fig. 1). At 14 hpf, the only specific brain expression is in the midbrain and in the midbrain-hindbrain boundary (Fig. 1A). At 16 hpf, lef1 expression begins in the ventral forebrain (Fig. 1B); by 19 hpf, it becomes restricted to the posterior hypothalamus (Fig. 1C). Expression in all these brain regions continues until 30 hpf (Fig. 1D-F). By examining cross-sections through the posterior hypothalamus, we observed that lef1 is expressed in presumptive mitotic and post-mitotic cells located in the medial and lateral regions, respectively (Fig. 1G). Comparison of lef1 expression with other known hypothalamic markers, such as dlx2 (Fig. 2) and hlx1 (not shown) led us to conclude that its expression was limited to transverse domain 4 and the ventral part of domain 5 (Fig. 1H), as defined by Hauptmann and Gerster (Hauptmann and Gerster, 2000).
3 Lef1 regulates hypothalamic neurogenesis RESEARCH ARTICLE 4453 The specific expression pattern of lef1 therefore suggested that it may play an important role in the posterior hypothalamus following the initial steps of tissue induction and patterning. If Lef1 acts in the canonical Wnt signaling pathway, we would expect a Wnt ligand to be expressed in close proximity to the lef1- expressing hypothalamic cells. The wnt8b gene, which encodes a ligand of the canonical pathway, is expressed in the forebrain during late somitogenesis stages (Kelly et al., 1995). We observed wnt8b expression in the posterior hypothalamus at 19 hpf in a region overlapping with and adjacent to lef1 expression (Fig. 1I). Previous studies have shown that lef1 is itself a target of Wnt signaling (Kengaku et al., 1998), and we also observed that that lef1 was severely downregulated following injection with a previously published wnt8b MO (Riley et al., 2004). We therefore concluded that Wnt8b functions upstream of Lef1 expression during brain development. We next asked whether the expression of specific proneural and neuronal genes overlapped with lef1 in the posterior hypothalamus. We examined the expression of sox3, which encodes an HMG-box transcription factor in the SoxB1 family, members of which function at an early step in the process of neurogenesis (Kan et al., 2004). At 19 hpf, sox3 expression did not overlap with lef1 in the posterior hypothalamus (Fig. 2A,B). By 22 hpf, we observed co-expression of the two genes, which was maintained through 30 hpf (Fig. 2E,F). The zash1a gene (ascl1a Zebrafish Information Network), which encodes a proneural bhlh transcription factor, was previously shown to be expressed in the posterior hypothalamus (Allende and Weinberg, 1994). We found that zash1a was not co-expressed with lef1 in the posterior hypothalamus at 24 hpf (Fig. 2C,D), but coexpression was observed beginning at 26 hpf and continuing through 30 hpf (Fig. 2G,H). For both sox3 and zash1a, we observed coexpression with lef1 in medial progenitors and more lateral differentiated neurons. By contrast, two other genes are co-expressed with lef1 only in differentiated hypothalamic neurons at 30 hpf (Fig. 2I-L). The dlx2 gene, which is involved in forebrain regional specification, is also expressed in transverse domain 4 of the posterior hypothalamus (Hauptmann and Gerster, 2000). The expression of dlx2 primarily in postmitotic neurons suggests that it might act to regulate neuronal differentiation in this region, rather than playing an earlier role in progenitor specification. Finally, isl1 labels specific populations of differentiated neurons throughout the embryo, and was detected in the posterior hypothalamus at 30 hpf. Fig. 1. lef1 is expressed in the posterior hypothalamus during embryonic development. Lateral views are shown with anterior towards the left. White circles outline posterior hypothalamus. (A) At 14 hpf, lef1 is strongly expressed in the dorsal midbrain, but does not show specific expression in the developing hypothalamus. (B-F) At 16 hpf, lef1 expression first appears in the presumptive posterior hypothalamus, and this expression is maintained through 30 hpf. After 19 hpf, expression is present in dorsal and ventral regions of the posterior hypothalamus. Black line in F indicates plane of section in G. (G) Transverse section through the posterior hypothalamus (black oval) at 30 hpf, showing lef1 expression in both the medial mitotic cells and the lateral postmitotic cells of the posterior region. (H) Schematic depiction of lef1 expression domain in 30 hpf zebrafish hypothalamus. Numbered regions are based on those of Hauptmann and Gerster (Hauptmann and Gerster, 2000), and lef1 expression is shown in blue. (I) At 19 hpf, wnt8b (blue) and lef1 (red) show overlapping and adjacent expression in the posterior hypothalamus. (J) In wnt8b morphants, lef1 expression is reduced throughout the brain. Lef1 is not required for induction or AP patterning of the hypothalamus To determine the required function for lef1 during hypothalamic development, we used two methods to inactivate zygotic gene function. First, a splice-blocking MO was designed against an intron-exon boundary in the region encoding the DNA-binding HMG box. This region was targeted because exon-skipping, a potential outcome of splice-blocking MOs, would create a protein unable to bind DNA. In fact, RT-PCR analysis of injected embryos showed a smaller product, indicating the presence of a cryptic splice donor in the preceding exon (see Fig. S1 in the supplementary material). Sequencing of this product confirmed a small deletion, which resulted in a shifted open reading frame. Furthermore, RT- PCR (see Fig. S1 in the supplementary material) and in situ hybridization (not shown) indicated that lef1 mrna levels rapidly decreased following MO injection, an outcome that could be due to either nonsense-mediated decay or lack of auto-activation of lef1 transcription (Kengaku et al., 1998). Second, we examined embryos homozygous for the X8 deletion mutation, generated by Dr B. Riley. PCR and linkage analysis shows that X8 is a deletion in chromosome 1 with one end just distal to msxb and the other end proximal to lef1, a distance of 2-8 cm (Phillips et al., 2006). Although X8 probably removes many genes in addition to lef1, the only other identified gene in this region is msxb, which is not expressed in the developing hypothalamus. Importantly, in all following experiments both MOs and the X8 mutation produced identical hypothalamic phenotypes. Ventral midline CNS cells in the forebrain differentiate into hypothalamus anteriorly and floor plate posteriorly as a result of Nodal and Wnt signaling (Kapsimali et al., 2004). After the initial AP subdivision of ventral midline CNS fates, these signals also affect subsequent AP patterning within the hypothalamus (Kapsimali et al., 2004; Mathieu et al., 2002). To examine whether Lef1 is also required for AP patterning of the hypothalamus, we
4 4454 RESEARCH ARTICLE Development 133 (22) Fig. 2. lef1 is co-expressed with proneural and neuronal markers in the posterior hypothalamus. White circles outline posterior hypothalamus and inset panels show lef1 mrna expression from Fast Red fluorescence. Lines in whole-mount images show plane of corresponding cross-sections on the right. In cross-sections, black ovals outline posterior hypothalamus. (A,B) At 19 hpf, sox3 (blue) is not co-expressed with lef1 (red) in the posterior hypothalamus. (C,D) At 24 hpf, zash1a (blue) is not co-expressed with lef1 (red) in the posterior hypothalamus. (E-H) By 30 hpf, sox3 and zash1a are co-expressed with lef1 in both medial progenitors and lateral postmitotic neurons of the posterior hypothalamus. (I-L) By contrast, dlx2 and isl1 are co-expressed only with lef1 in lateral postmitotic neurons. performed in situ hybridization for specific patterning markers. We investigated hypothalamic AP patterning in wild-type embryos, lef1 morphants and X8 mutants by comparing the expression of nk2.1a (titf1a Zebrafish Information Network), rx3 and emx2. The nk2.1a gene is a marker for the entire hypothalamus (Rohr et al., 2001), whereas rx3 and emx2, respectively, mark the anterior and posterior hypothalamus (Chuang et al., 1999; Mathieu et al., 2002). As opposed to the severe defects observed in zebrafish axin1 mutants (Kapsimali et al., 2004), all three markers were still expressed appropriately at 30 hpf in lef1 morphants (Fig. 3) and X8 mutants (see Fig. S2 in the supplementary material), suggesting that the regional identity of posterior hypothalamus was unchanged. morphants (Fig. 4Q-T). Thus, expression of proneural and neuronal genes in the lef1-positive domain of posterior hypothalamus specifically requires lef1 function. These data suggest that Lef1 may act to regulate the development of a specific population of hypothalamic neurons. Lef1 is required for proneural and neuronal gene expression in the posterior hypothalamus The above observations, coupled with the expression pattern of lef1 in the hypothalamus, suggested that this gene may play a role in a later step of hypothalamic development. Such a role would be consistent with previous studies demonstrating that Wnt signals can regulate neurogenesis in the vertebrate midbrain and hindbrain (Amoyel et al., 2005; Castelo-Branco et al., 2003). To determine whether Lef1 function is required for expression of the marker genes listed previously, we analyzed loss-of-function embryos. We found that expression of sox3 and zash1a was absent in the posterior hypothalamus at 24 and 28 hpf, respectively, in both lef1 morphants and X8 mutants (Fig. 4A-H; see Fig. S2 in the supplementary material). In addition, dlx2 and isl1 were also not expressed in the posterior hypothalamus of lef1 morphants and X8 mutants at 30 hpf (Fig. 4I-P; see Fig. S2 in the supplementary material). All of these markers were expressed relatively normally in other forebrain regions. In addition, ngn1 (neurog1 Zebrafish Information Network) and olig2, which are expressed posterior and dorsal to lef1 in the posterior tuberculum, were not altered in lef1 Fig. 3. Lef1 is not required for molecular markers of hypothalamus identity or AP patterning. White circles outline posterior hypothalamus. (A,C,E) Uninjected embryos. (B,D,F) Embryos injected with 2 ng of lef1 MO. (A,B) nk2.1a expression in the entire hypothalamus is unaffected in lef1 morphants at 30 hpf. (C,D) rx3 is still expressed in the anterior hypothalamus in lef1 morphants at 30 hpf. (E,F) emx2 is still expressed in the posterior hypothalamus of lef1 morphants at 30 hpf.
5 Lef1 regulates hypothalamic neurogenesis RESEARCH ARTICLE 4455 Fig. 4. Lef1 is required for proneural and neuronal gene expression in the posterior hypothalamus. White circles outline posterior hypothalamus in whole-mount views, and black ovals outline posterior hypothalamus in cross-sections. Lines in whole-mount images show plane of corresponding cross-sections. (A-D) Expression of sox3 is absent in the posterior hypothalamus of lef1 morphants at 24 hpf. (E-H) Expression of zash1a is absent in the posterior hypothalamus of lef1 morphants at 28 hpf. (I-P) Expression of dlx2 and isl1 are absent in the posterior hypothalamus of lef1 morphants at 30 hpf. (Q-T) Expression of ngn1 and olig2, which are expressed in the posterior tuberculum, is unaffected in the posterior tuberculum of lef1 morphants. Rescue of marker gene expression by mrna injection To confirm that the phenotypes in lef1 morphants were specific to the lef1 gene and not due to other side-effects, we attempted to rescue hypothalamic gene expression by co-injection of lef1 mrna lacking the MO target sequence. We first titrated the dose of lef1 mrna to find a concentration that does not produce phenotypes when overexpressed, and observed normal development in embryos injected with 100 pg of mrna. Co-injection of 100 pg of lef1 mrna rescued the expression of zash1a, dlx2 and isl1 at 30 hpf in the posterior hypothalamus of lef1 morphants (Table 1; see Fig. S3 in the supplementary material). This result, combined with the similar phenotypes produced by MO injection and the X8 deletion, led us to conclude that the absence of gene expression in lef1 morphants is specific to lef1 loss of function. Of all the markers we analyzed, sox3 is expressed at the earliest time in the posterior hypothalamus and functions at the earliest step of neurogenesis (Kan et al., 2004). Furthermore, another soxb1 family gene, sox2, has been shown to be downstream of canonical Wnt signaling in the Xenopus retina (Van Raay et al., 2005). We therefore investigated whether sox3 mrna could rescue the expression of the other proneural and neuronal markers in lef1 morphants. Co-injection of 20 pg of sox3 mrna with the lef1 MO rescued zash1a, dlx2 and isl1 expression at 30 hpf in the posterior hypothalamus (Table 1; see Fig. S3 in the supplementary material). These results led us to conclude that Lef1 may act through Sox3 to establish a program of neurogenesis in the posterior hypothalamus, resulting in eventual differentiation of a discrete neuronal population. Lef1 is required for neurogenesis in the posterior hypothalamus We also examined later effects on neuronal differentiation in lef1 morphants. Hu proteins, which mark all postmitotic neurons, are expressed in the posterior hypothalamus at 36 hpf. We observed a specific and complete loss of Hu expression in the posterior Table 1. Rescue of lef1 MO phenotypes with sox3 or lef1 mrna Treatment zash1a (%) dlx2 (%) isl1 (%) None 100 (n=40) 100 (n=40) 100 (n=40) lef1 MO 9 (n=109) 9 (n=135) 9 (n=135) lef1 MO + lef1 mrna 61 (n=100) 60 (n=100) 61 (n=100) lef1 MO + sox3 mrna 70 (n=104) 69 (n=104) 68 (n=104) lef1 MO (2 ng), 20 pg of sox3 mrna and 100 pg of lef1 mrna were injected into one-cell stage embryos. Results of phenotypic rescue by the mrna were obtained performing in situ hybridization for zash1a, dlx2 and isl1. P-values were measured by Student s t-test comparing the results of single lef1 MO injection to co-injection with either sox3 mrna or lef1 mrna for each gene in situ hybridization (P<0.001 in all comparisons).
6 4456 RESEARCH ARTICLE Development 133 (22) hypothalamus of morphants (Fig. 5A-D). Interestingly, a subset of Hu-positive neurons in the posterior hypothalamus continue to express Lef1 protein even at 36 hpf (Fig. 5E), suggesting that Lef1 may play an additional later role in their differentiation or function. By 48 hpf, specific axonal populations are visible in the posterior hypothalamus by acetylated tubulin staining. We observed a loss of these axons in lef1 morphants (Fig. 5F,G), and although the staining method used did not allow us to identify these axons as afferents or efferents, this result was consistent with decreased neuronal differentiation in this region. Lef1 is not necessary for cell proliferation or survival in the posterior hypothalamus The changes we observed in gene expression and neuronal differentiation led us to investigate whether hypothalamic progenitors exhibited changes in proliferation or apoptosis when Lef1 function was lost. First, we found expression of pcna mrna (not shown) and phospho-histone H3 (ph3) in the posterior hypothalamus of morphants (Fig. 6A-L), indicating that proliferating cells were still present. The percentage of ph3-positive cells at 19 hpf (Fig. 6C,D) in the posterior hypothalamus was 7.5±2.0% (s.d., n=15) in controls compared with 5.4±1.1% (n=15) in morphants. At 24 hpf (Fig. 6G,H), there were 13±1.3% (n=15) positive cells in controls, compared with 11±2.1% (n=15) cells in morphants. At 30 hpf (Fig. 6K,L), we found 7.1±1.0% (n=15) ph3- positive cells in controls and 6.7±0.8% (n=15) in morphants. In addition, total cell number in the posterior hypothalamus was not significantly affected at any stage. Analysis of apoptosis by TUNEL staining showed no additional labeling in the hypothalamus of morphants at 19 or 24 hpf (Fig. 6M-P). These results, combined with the lack of Hu staining observed in morphants, suggest that the small decrease in ph3 index reflects a slower cell cycle time rather than premature differentiation or cell death. Therefore, although Lef1 may be required for the proper rate of cell division, it is not required for proliferation in general and the reduced proliferation rate alone cannot explain the complete lack of proneural and neuronal markers we observed in morphants. Our results are consistent with a model where in the absence of Lef1 function, dividing hypothalamic progenitors remain in an immature undifferentiated state and fail to acquire neural competence. Wnt8b regulates gene expression in the posterior hypothalamus similarly to Lef1 In addition to our finding that wnt8b was expressed near lef1 during early hypothalamic development (Fig. 1I), we observed continued wnt8b expression in the posterior hypothalamus at 30 hpf in a region overlapping with and adjacent to lef1 expression (Fig. 7A,B). To test whether wnt8b and lef1 function are similarly required for hypothalamic neurogenesis, we examined wnt8b morphants for proneural and neuronal markers. In support of our hypothesis that Wnt8b functions upstream of Lef1 in hypothalamic development, we found that injection of wnt8b MO led to absence of sox3, zash1a, dlx2 and isl1 expression in the posterior hypothalamus at 30 hpf (Fig. 7C-F). Fig. 5. Lef1 is required for neuronal differentiation in the posterior hypothalamus. (A,B) Confocal projections through the hypothalamus of whole-mount 36 hpf brains stained for HuC/D, a panneuronal marker. Owing to differences in mounting, neurons outside the hypothalamus may not be visible. (C,D) Cross-sections of 36 hpf brains through the plane marked by the lines in A,B. A specific population of Hu-positive neurons (yellow circles and ovals) is absent in the posterior hypothalamus of lef1 morphants (B,D). (E) Hu (green) is co-expressed with Lef1 protein (red) in a subset of posterior neurons at 36 hpf. (F,G) Acetylated tubulin (AT) staining labels specific axons in the posterior hypothalamus (yellow circles) that are reduced in lef1 morphants at 48 hpf. Confocal projections through the ventral side of the hypothalamus are shown, with anterior towards the top. Wnt8b and Lef1 regulate -catenin-mediated transcription in the posterior hypothalamus To test whether both Wnt8b and Lef1 act in the canonical Wnt pathway, we used a transgenic reporter for -catenin-mediated transcription (TOPdGFP), in which GFP is activated in a Lef1- dependent manner (Dorsky et al., 2002). First, we examined whether gfp mrna expression overlapped with wnt8b and lef1 expression in the posterior hypothalamus of TOPdGFP embryos at 24 hpf, shortly after sox3 expression first appears in this region. Double in situ hybridization on 24 hpf TOPdGFP embryos showed adjacent expression of gfp and wnt8b in the posterior hypothalamus. (Fig. 7G,H). In addition, gfp was expressed in a overlapping pattern with lef1 in the posterior hypothalamus (Fig. 7I). The relative expression patterns of these three genes were maintained until 30 hpf (not shown). We next asked whether TOPdGFP expression is disrupted in the posterior hypothalamus when either wnt8b or lef1 function is removed. To address this issue, we performed immunohistochemistry for GFP protein after injecting wnt8b or lef1 MOs into TOPdGFP embryos. Removal of both wnt8b and lef1 resulted in the loss of GFP in the posterior hypothalamus at 24 and 30 hpf (Fig. 7J-O). These results were consistent with our observations that lef1 expression coincides with gfp mrna in this region, and suggested that although Wnt8b may function as a secreted ligand, Lef1 probably functions cell-autonomously. Because lef1 expression was downregulated in wnt8b morphants, we cannot determine whether Wnt8b normally acts only to induce lef1,
7 Lef1 regulates hypothalamic neurogenesis RESEARCH ARTICLE 4457 Fig. 6. Analysis of proliferation and apoptosis in lef1 morphants. (A-L) In lef1 morphants, ph3-positive cells are observed in the posterior hypothalamus at 19, 24 and 30 hpf, indicating that proliferating cells are still present in this region. (A,B,E,F,I,J) Cross-sections through posterior hypothalamus (outlined by dotted circles). (C,D,G,H,K,L) Confocal projections of ph3-stained embryos used for quantitative analysis. The central region outlined by broken lines was defined as posterior hypothalamus, based on morphology in bright-field views. The region farthest to the right was excluded from counts because it does not express lef1 and is unaffected in lef1 morphants (Fig. 4Q-T). (M-P) Apoptotic cells identified by TUNEL staining are absent in the posterior hypothalamus at 19 hpf and 24 hpf in wild-type and lef1 MO-injected embryos (arrowheads). Asterisks indicate apoptotic cells in the dorsal brain of lef1 morphants. Chromatin immunoprecipitation identifies sox3 as a binding target of Lef1 in vivo Our analysis of proneural and neuronal markers indicating that progenitor genes may be downstream targets of lef1 gene function, and our observation that sox3 mrna could rescue the lef1 MO phenotype (Table 1; see Fig. S3 in the supplementary material), led to the hypothesis that sox3 transcription could be directly regulated by -catenin and Lef1. We therefore employed chromatin immunoprecipitation (ChIP) assays to determine whether the sox3 upstream regulatory region binds to Lef1 protein in vivo. For immunoprecipitation, we used a polyclonal antibody from our laboratory, raised against zebrafish Lef1 (also used in Fig. 5E). This antibody recognizes a single band of ~50-55 kda in lysates from wild-type 24 hpf embryos, and this band is absent in X8 deletion mutant embryo lysates, indicating that it specifically detects Lef1 protein (Fig. 8A). To identify putative Lef1-binding sites in upstream regulatory sequences of sox3, we analyzed genomic sequence data from the Sanger Centre zebrafish genome assembly and found several consensus sites within 10 kb upstream of the sox3 transcription start site. PCR primers designed to flank these putative sites were able to amplify products from chromatin extracts of 24 hpf embryos. Following sonication, immunoprecipitation and de-crosslinking, we were able to amplify one of these fragments from Lef1 antibody precipitated extracts, but not from controls with no antibody, or more importantly from X8 mutant extracts that lack Lef1 protein (Fig. 8B). The fragment that specifically interacted with Lef1 protein was located ~6.5 kb upstream of the sox3 transcription start, and contains a consensus binding site in a region that is conserved between zebrafish and Fugu. As a positive control, we were also able to immunoprecipitate binding sites in the promoters of nacre (mitfa Zebrafish Information Network) and ngn1 (Fig. 8B), both previously identified as -catenin target genes (Dorsky et al., 2000; Hirabayashi et al., 2004). We therefore conclude that Lef1 specifically interacts with an upstream regulatory region of sox3 in 24 hpf zebrafish embryos, and thus may be a direct transcriptional activator of this gene in the posterior hypothalamus. DISCUSSION In this study, we have revealed a specific role for canonical Wnt signaling during hypothalamic neurogenesis by analyzing the function of zebrafish Lef1, an essential mediator of the pathway. In Lef1 loss-of-function embryos, proneural and neuronal markers and differentiated neurons are absent from the posterior hypothalamus, indicating a requirement for Lef1 in the development of these cells. We have also shown that Wnt8b and Lef1 both act through the canonical Wnt pathway in this region. Our data indicate that the activation of sox3 downstream of canonical Wnt signaling drives a neurogenesis program during zebrafish posterior hypothalamus development, eventually leading to the differentiation of a specific neuronal population (Fig. 9). Specific expression of canonical Wnt pathway components in the posterior hypothalamus The expression of zebrafish lef1 begins at 16 hpf in the ventral forebrain, and continues until 30 hpf in the posterior hypothalamus. Of all known zebrafish lef/tcf family genes, lef1 is the only member expressed in this particular brain region, suggesting that it may have a unique function in hypothalamic development. We have found that lef1 is expressed in both the progenitors and neurons of the posterior hypothalamus, suggesting that it may function to regulate a program of neurogenesis. Examination of a canonical Wnt pathway reporter, or also to signal through Lef1 in target gene activation. However, our data suggest that both genes are required in the same pathway mediating hypothalamic neurogenesis.
8 4458 RESEARCH ARTICLE Development 133 (22) Fig. 7. Wnt8b and Lef1 act through the canonical Wnt pathway in the posterior hypothalamus. White circles outline posterior hypothalamus in whole-mount views, and black ovals outline posterior hypothalamus in cross-sections. (A,B) At 30 hpf, wnt8b (blue) and lef1 (red) show overlapping and adjacent expression in the posterior hypothalamus. (C-F) In wnt8b morphants, sox3, zash1a, dlx2 and isl1 expression are specifically reduced in the posterior hypothalamus at 30 hpf. (G-I) A transgenic reporter for -catenin-mediated transcription (TOPdGFP) shows mrna expression in the posterior hypothalamus (red) at 24 hpf. In the posterior hypothalamus, wnt8b expression (blue-h) partially overlaps with gfp, whereas lef1 (blue-i) almost completely overlaps with gfp. (J-O) In TOPdGFP embryos, GFP protein (brown) is expressed in the posterior hypothalamus. At 24 hpf (J-L) and 30 hpf (M-O), this expression is absent in wnt8b and lef1 morphants. TOPdGFP, indicates that this pathway is active where lef1 is expressed. Furthermore, we have identified a gene encoding an upstream ligand of the canonical Wnt pathway, wnt8b, expressed adjacent to lef1 in the posterior hypothalamus and required for lef1 expression. Together, these findings indicate that -catenin and Lef1 may interact in hypothalamic progenitors to activate target genes involved in neurogenesis. Lef1 is required for neurogenesis in the posterior hypothalamus Canonical Wnt signaling is required for the initial induction and subsequent AP pattering of the hypothalamus (Wilson and Houart, 2004). Our data indicate that Lef1 does not regulate either of these events, based on the unaffected expression of markers such as nk2.1a, rx3, emx2, ngn1 and olig2, and the presence of anterior hypothalamic neurons in morphants and mutants. By contrast, we find a specific loss of genes expressed in progenitors (sox3 and zash1a) and in postmitotic neurons (dlx2 and isl1) in the posterior hypothalamus of these embryos. In addition, we observed continued cell proliferation and no increase in apoptosis, suggesting that progenitor cells remain in an undifferentiated state and fail to be specified as neural precursors. Because we observed a complete loss of Hu-positive cells in the posterior hypothalamus in lef1 morphants, we conclude that neurons are not mis-specified to alternate fates by 36 hpf, and that Lef1 function is required for the entire process of neurogenesis in this region. We cannot determine whether neurogenesis has been inhibited or merely delayed in the absence of Lef1; however, analysis of neuronal markers at 36 hpf (not shown) indicates it is delayed by at least 6 hours. In either case, our data suggest that Lef1 regulates the proper timing of neurogenesis in the hypothalamus. Although it has been shown previously that Lef1 is required for the generation of neuronal populations in the mouse cortex and midbrain (Galceran et al., 2000; van Genderen et al., 1994), our data are the first to demonstrate that hypothalamic neurogenesis is regulated by Lef1. Lef1 functions downstream of Wnt8b to regulate neurogenesis through Sox3 Multiple Wnt proteins are involved in canonical signaling, but it has been difficult to study the specific function of individual Wnts because of their redundant functions in CNS development. In this case, we have found that Wnt8b is uniquely positioned to act upstream of Lef1 in the posterior hypothalamus. Our studies provide two sets of data indicating that canonical Wnt signaling through Wnt8b is required for neurogenesis. First, when we eliminated wnt8b function proneural and neuronal marker gene expression was lost, producing a similar phenotype to lef1 morphants. Second, expression of TOPdGFP was specifically eliminated in the posterior hypothalamus in both lef1 and wnt8b morphants. We were unable to rescue neurogenesis in wnt8b morphants by sox3 overexpression, perhaps owing to additional roles for wnt8b in embryonic development. Indeed, Wnt8b may signal through other Tcf proteins,
9 Lef1 regulates hypothalamic neurogenesis RESEARCH ARTICLE 4459 activate sox3 transcription in the posterior hypothalamus. A more detailed promoter analysis, including mutagenesis of potential binding sites, will be necessary to prove conclusively that sox3 directly requires Lef1 for its activation in posterior hypothalamic progenitors. However, we believe that this model most accurately explains the phenotypes we observe in our embryos. Evidence from other species indicates that SoxB1 factors are required for the acquisition of neural potential (Graham et al., 2003; Kan et al., 2004). Therefore, activation of sox3 by Lef1 could initiate a program of neurogenesis, and subsequently allow for cell cycle exit and neuronal differentiation. Our model would also be consistent with results obtained in the Xenopus retina, where elimination of Fz5 function results in loss of both sox2 and proneural gene expression (Van Raay et al., 2005). Fig. 8. Lef1 associates with the sox3 promoter in vivo. (A) Western blot of whole-embryo lysates probed with anti- -catenin and Lef1 polyclonal antibodies. The Lef1 band is specifically absent in X8 mutants, whereas the positive control -catenin band is present. (B) ChIP analysis of whole-embryo lysates shows that the Lef1 antibody can immunoprecipitate DNA fragments containing Lef-binding sites in the upstream regulatory region of sox3, nacre and ngn1. These fragments do not precipitate in controls lacking antibody or chromatin (mock). In lysates from X8 mutants that lack Lef1 protein, the antibody is unable to precipitate these fragments, suggesting that the interaction is specific to Lef1. including Tcf7 which is also expressed in the forebrain (Veien et al., 2005). Our experiments therefore cannot determine whether after Wnt8b initiates lef1 expression, a different Wnt signal then activates the pathway through Lef1. However the continued expression of Wnt8b in the posterior hypothalamus through 30 hpf suggests that it might be required for both functions. Our data indicate that Lef1 protein binds directly to upstream regulatory elements of the sox3 gene in 24 hpf zebrafish embryos. Combined with the requirement for lef1 in the hypothalamic expression of sox3, and the ability of sox3 to rescue the loss of Lef1 function, we propose that -catenin and Lef1 cooperate to directly Evolutionary conservation of Sox3 and Wnt function in posterior hypothalamic development In contrast to the larva and adult, hypothalamic neuronal identity and anatomy has been poorly characterized during zebrafish embryogenesis. In the present study, we have shown that a specific population of neurons in the posterior hypothalamus differentiates between 22 and 36 hpf, and that its development depends on Lef1 function. However, we do not know the ultimate fate of the neurons that we have identified in this study. Several neurotransmitters have been identified in the hypothalamus in zebrafish larvae and adults (Doldan et al., 1999; Poon et al., 2003; Ross et al., 1992; Teraoka et al., 2004). In addition, dopaminergic neurons project from the tuberal hypothalamus to the subpallium in adults, but it remains unclear when the projection arises (Rink and Wullimann, 2001). Previous studies have reported that neurotransmitter expression and axonogenesis in the posterior hypothalamus appears between 36 and 48 hpf, although proneural genes are expressed about one day earlier (Clemente et al., 2004; Hauptmann and Gerster, 2000; Wilson et al., 1990; Wullimann and Mueller, 2004). Intriguingly, a recent report shows specific expression of the secreted hormones AGRP and PMOC as early as 24 hpf in the posterior hypothalamus (Song et al., 2003). This result, combined with the similar expression of dlx2 in the mouse hypothalamus (Bulfone et al., 1993) and in zebrafish Lef1-dependent neurons, suggests that these cells may contribute to the infundibulum or neurohypophysis. Significantly, at least three of the genes analyzed in this study in addition to dlx2 show similar expression domains in the mammalian posterior hypothalamus. The expression of wnt8b has previously been reported in the mammillary and retromammillary hypothalamus of mice and humans (Lako et al., 1998). Both the endogenous lef1 gene and a reporter knocked into the mouse lef1 locus shows expression in the posterior hypothalamus, although the precise location has not been characterized (Galceran et al., 2000). Fig. 9. Summary and model. During A/P patterning of the hypothalamus (16-22 hpf), lef1 and wnt8b are expressed in posterior regions. Later, when neurogenesis begins in the posterior hypothalamus (22-30 hpf), lef1 is required for the expression of sox3, which in turn regulates proneural and neuronal gene expression. After 30 hpf, a specific population of Lef1-dependent neurons differentiates in this region. Morphology of the hypothalamus is adapted from Hauptmann and Gerster (Hauptmann and Gerster, 2000).
10 4460 RESEARCH ARTICLE Development 133 (22) Finally, not only is sox3 expressed in the mammalian infundibulum (Solomon et al., 2004), specific mutations in this gene lead to defects in infundibular hypoplasia and associated hypopituitarism in both mice and humans (Rizzoti et al., 2004; Woods et al., 2005). Our data suggest that a pathway containing Wnt8b, Lef1 and Sox3 may be an important conserved mechanism for driving a program of neurogenesis in the posterior hypothalamus and promoting normal endocrine hormone function. We thank Bruce Riley for sharing the X8 mutant fish, Arne Lekven for providing the wnt8b morpholino and probe, and Stephen Wilson for providing numerous in situ hybridization probes. R.I.D. is supported by the American Cancer Society and the Pew Scholars Program. Supplementary material Supplementary material for this article is available at References Akimenko, M. A., Ekker, M., Wegner, J., Lin, W. and Westerfield, M. (1994). Combinatorial expression of three zebrafish genes related to distal-less: part of a homeobox gene code for the head. J. Neurosci. 14, Allende, M. L. and Weinberg, E. S. (1994). The expression pattern of two zebrafish achaete-scute homolog (ash) genes is altered in the embryonic brain of the cyclops mutant. Dev. Biol. 166, Amoyel, M., Cheng, Y. C., Jiang, Y. J. and Wilkinson, D. G. (2005). Wnt1 regulates neurogenesis and mediates lateral inhibition of boundary cell specification in the zebrafish hindbrain. Development 132, Blader, P., Fischer, N., Gradwohl, G., Guillemot, F. and Strahle, U. (1997). The activity of neurogenin1 is controlled by local cues in the zebrafish embryo. Development 124, Brault, V., Moore, R., Kutsch, S., Ishibashi, M., Rowitch, D. H., McMahon, A. P., Sommer, L., Boussadia, O. and Kemler, R. (2001). Inactivation of the betacatenin gene by Wnt1-Cre-mediated deletion results in dramatic brain malformation and failure of craniofacial development. Development 128, Buckles, G. R., Thorpe, C. J., Ramel, M. C. and Lekven, A. C. (2004). Combinatorial Wnt control of zebrafish midbrain-hindbrain boundary formation. Mech. Dev. 121, Bulfone, A., Puelles, L., Porteus, M. H., Frohman, M. A., Martin, G. R. and Rubenstein, J. L. (1993). Spatially restricted expression of Dlx-1, Dlx-2 (Tes-1), Gbx-2, and Wnt-3 in the embryonic day 12.5 mouse forebrain defines potential transverse and longitudinal segmental boundaries. J. Neurosci. 13, Castelo-Branco, G., Wagner, J., Rodriguez, F. J., Kele, J., Sousa, K., Rawal, N., Pasolli, H. A., Fuchs, E., Kitajewski, J. and Arenas, E. (2003). Differential regulation of midbrain dopaminergic neuron development by Wnt-1, Wnt-3a, and Wnt-5a. Proc. Natl. Acad. Sci. USA 100, Chiang, C., Litingtung, Y., Lee, E., Young, K. E., Corden, J. L., Westphal, H. and Beachy, P. A. (1996). Cyclopia and defective axial patterning in mice lacking Sonic hedgehog gene function. Nature 383, Chuang, J. C., Mathers, P. H. and Raymond, P. A. (1999). Expression of three Rx homeobox genes in embryonic and adult zebrafish. Mech. Dev. 84, Ciani, L. and Salinas, P. C. (2005). WNTs in the vertebrate nervous system: from patterning to neuronal connectivity. Nat. Rev. Neurosci. 6, Clemente, D., Porteros, A., Weruaga, E., Alonso, J. R., Arenzana, F. J., Aijon, J. and Arevalo, R. (2004). Cholinergic elements in the zebrafish central nervous system: histochemical and immunohistochemical analysis. J. Comp. Neurol. 474, Doldan, M. J., Prego, B., Holmqvist, B. I. and de Miguel, E. (1999). Distribution of GABA-immunolabeling in the early zebrafish (Danio rerio) brain. Eur. J. Morphol. 37, Dorsky, R. I., Snyder, A., Cretekos, C. J., Grunwald, D. J., Geisler, R., Haffter, P., Moon, R. T. and Raible, D. W. (1999). Maternal and embryonic expression of zebrafish lef1. Mech. Dev. 86, Dorsky, R. I., Raible, D. W. and Moon, R. T. (2000). Direct regulation of nacre, a zebrafish MITF homolog required for pigment cell formation, by the Wnt pathway. Genes Dev. 14, Dorsky, R. I., Sheldahl, L. C. and Moon, R. T. (2002). A transgenic Lef1/betacatenin-dependent reporter is expressed in spatially restricted domains throughout zebrafish development. Dev. Biol. 241, Dorsky, R. I., Itoh, M., Moon, R. T. and Chitnis, A. (2003). Two tcf3 genes cooperate to pattern the zebrafish brain. Development 130, Eastman, Q. and Grosschedl, R. (1999). Regulation of LEF-1/TCF transcription factors by Wnt and other signals. Curr. Opin. Cell Biol. 11, Erter, C. E., Wilm, T. P., Basler, N., Wright, C. V. and Solnica-Krezel, L. (2001). Wnt8 is required in lateral mesendodermal precursors for neural posteriorization in vivo. Development 128, Galceran, J., Miyashita-Lin, E. M., Devaney, E., Rubenstein, J. L. and Grosschedl, R. (2000). Hippocampus development and generation of dentate gyrus granule cells is regulated by LEF1. Development 127, Graham, V., Khudyakov, J., Ellis, P. and Pevny, L. (2003). SOX2 functions to maintain neural progenitor identity. Neuron 39, Guo, S., Brush, J., Teraoka, H., Goddard, A., Wilson, S. W., Mullins, M. C. and Rosenthal, A. (1999). Development of noradrenergic neurons in the zebrafish hindbrain requires BMP, FGF8, and the homeodomain protein soulless/phox2a. Neuron 24, Hauptmann, G. and Gerster, T. (2000). Regulatory gene expression patterns reveal transverse and longitudinal subdivisions of the embryonic zebrafish forebrain. Mech. Dev. 91, Hirabayashi, Y., Itoh, Y., Tabata, H., Nakajima, K., Akiyama, T., Masuyama, N. and Gotoh, Y. (2004). The Wnt/beta-catenin pathway directs neuronal differentiation of cortical neural precursor cells. Development 131, Houart, C., Caneparo, L., Heisenberg, C., Barth, K., Take-Uchi, M. and Wilson, S. (2002). Establishment of the telencephalon during gastrulation by local antagonism of Wnt signaling. Neuron 35, Jowett, T. (2001). Double in situ hybridization techniques in zebrafish. Methods 23, Kan, L., Israsena, N., Zhang, Z., Hu, M., Zhao, L. R., Jalali, A., Sahni, V. and Kessler, J. A. (2004). Sox1 acts through multiple independent pathways to promote neurogenesis. Dev. Biol. 269, Kapsimali, M., Caneparo, L., Houart, C. and Wilson, S. W. (2004). Inhibition of Wnt/Axin/beta-catenin pathway activity promotes ventral CNS midline tissue to adopt hypothalamic rather than floorplate identity. Development 131, Kelly, G. M., Greenstein, P., Erezyilmaz, D. F. and Moon, R. T. (1995). Zebrafish wnt8 and wnt8b share a common activity but are involved in distinct developmental pathways. Development 121, Kengaku, M., Capdevila, J., Rodriguez-Esteban, C., De La Pena, J., Johnson, R. L., Belmonte, J. C. I. and Tabin, C. J. (1998). Distinct WNT pathways regulating AER formation and dorsoventral polarity in the chick limb bud. Science 280, Kimmel, C. B., Ballard, W. W., Kimmel, S. R., Ullmann, B. and Schilling, T. F. (1995). Stages of embryonic development of the zebrafish. Dev. Dyn. 203, Kudoh, T., Concha, M. L., Houart, C., Dawid, I. B. and Wilson, S. W. (2004). Combinatorial Fgf and Bmp signalling patterns the gastrula ectoderm into prospective neural and epidermal domains. Development 131, Lako, M., Lindsay, S., Bullen, P., Wilson, D. I., Robson, S. C. and Strachan, T. (1998). A novel mammalian wnt gene, WNT8B, shows brain-restricted expression in early development, with sharply delimited expression boundaries in the developing forebrain. Hum. Mol. Genet. 7, Lee, S. M., Tole, S., Grove, E. and McMahon, A. P. (2000). A local Wnt-3a signal is required for development of the mammalian hippocampus. Development 127, Lekven, A. C., Buckles, G. R., Kostakis, N. and Moon, R. T. (2003). Wnt1 and wnt10b function redundantly at the zebrafish midbrain-hindbrain boundary. Dev. Biol. 254, Logan, C. Y. and Nusse, R. (2004). The Wnt signaling pathway in development and disease. Annu. Rev. Cell Dev. Biol. 20, Mathieu, J., Barth, A., Rosa, F. M., Wilson, S. W. and Peyrieras, N. (2002). Distinct and cooperative roles for Nodal and Hedgehog signals during hypothalamic development. Development 129, Morita, T., Nitta, H., Kiyama, Y., Mori, H. and Mishina, M. (1995). Differential expression of two zebrafish emx homeoprotein mrnas in the developing brain. Neurosci. Lett. 198, Mueller, T. and Wullimann, M. F. (2002). Expression domains of neurod (nrd) in the early postembryonic zebrafish brain. Brain Res. Bull. 57, Nordstrom, U., Jessell, T. M. and Edlund, T. (2002). Progressive induction of caudal neural character by graded Wnt signaling. Nat. Neurosci. 5, Okamoto, H., Kikuchi, H. and Segawa, H. (2000). [Islet-1 family and neural differentiation: study using zebrafish embryos]. Tanpakushitsu Kakusan Koso 45, Oxtoby, E. and Jowett, T. (1993). Cloning of the zebrafish krox-20 gene (krx-20) and its expression during hindbrain development. Nucleic Acids Res. 21, Park, H. C., Mehta, A., Richardson, J. S. and Appel, B. (2002). olig2 is required for zebrafish primary motor neuron and oligodendrocyte development. Dev. Biol. 248, Phillips, B. T., Kwon, H. J., Melton, C., Houghtaling, P., Fritz, A. and Riley, B. B. (2006). Zebrafish msxb, msxc and msxe function together to refine the neural-nonneural border and regulate cranial placodes and neural crest development. Dev. Biol. 294, Poon, K. L., Richardson, M., Lam, C. S., Khoo, H. E. and Korzh, V. (2003). Expression pattern of neuronal nitric oxide synthase in embryonic zebrafish. Gene Expr. Patterns 3,
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Biology 218, practise Exam 2, 2011
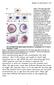 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Supplementary Figures
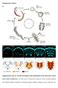 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
UNIVERSITY OF YORK BIOLOGY. Developmental Biology
 Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Maternal Control of GermLayer Formation in Xenopus
 Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Role of Organizer Chages in Late Frog Embryos
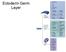 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Zebrafish Genes rx1 and rx2 Help Define the Region of Forebrain That Gives Rise to Retina
 Developmental Biology 231, 13 30 (2001) doi:10.1006/dbio.2000.0125, available online at http://www.idealibrary.com on Zebrafish Genes rx1 and rx2 Help Define the Region of Forebrain That Gives Rise to
Developmental Biology 231, 13 30 (2001) doi:10.1006/dbio.2000.0125, available online at http://www.idealibrary.com on Zebrafish Genes rx1 and rx2 Help Define the Region of Forebrain That Gives Rise to
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Interaction of Wnt and caudal-related genes in zebrafish posterior body formation
 Developmental Biology 279 (2005) 125 141 www.elsevier.com/locate/ydbio Interaction of Wnt and caudal-related genes in zebrafish posterior body formation Takashi Shimizu, Young-Ki Bae, Osamu Muraoka, Masahiko
Developmental Biology 279 (2005) 125 141 www.elsevier.com/locate/ydbio Interaction of Wnt and caudal-related genes in zebrafish posterior body formation Takashi Shimizu, Young-Ki Bae, Osamu Muraoka, Masahiko
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Zebrafish wnt8 and wnt8b share a common activity but are involved in distinct developmental pathways
 Development 121, 1787-1799 (1995) Printed in Great Britain The Company of Biologists Limited 1995 1787 Zebrafish wnt8 and wnt8b share a common activity but are involved in distinct developmental pathways
Development 121, 1787-1799 (1995) Printed in Great Britain The Company of Biologists Limited 1995 1787 Zebrafish wnt8 and wnt8b share a common activity but are involved in distinct developmental pathways
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
NIH Public Access Author Manuscript Int J Dev Biol. Author manuscript; available in PMC 2012 January 19.
 NIH Public Access Author Manuscript Published in final edited form as: Int J Dev Biol. 2011 ; 55(10-11-12): 917 921. doi:10.1387/ijdb.113288sh. XIer2 is required for convergent extension movements during
NIH Public Access Author Manuscript Published in final edited form as: Int J Dev Biol. 2011 ; 55(10-11-12): 917 921. doi:10.1387/ijdb.113288sh. XIer2 is required for convergent extension movements during
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Lhx5 promotes forebrain development and activates transcription of secreted Wnt antagonists
 RESEARCH ARTICLE 3191 Development 133, 3191-3200 (2006) doi:10.1242/dev.02485 Lhx5 promotes forebrain development and activates transcription of secreted Wnt antagonists Gang Peng and Monte Westerfield*
RESEARCH ARTICLE 3191 Development 133, 3191-3200 (2006) doi:10.1242/dev.02485 Lhx5 promotes forebrain development and activates transcription of secreted Wnt antagonists Gang Peng and Monte Westerfield*
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
A role for neural determination genes in specifying the dorsoventral identity of telencephalic neurons
 A role for neural determination genes in specifying the dorsoventral identity of telencephalic neurons Carol Fode, 1 Qiufu Ma, 2,3 Simona Casarosa, 1 Siew-Lan Ang, 1 David J. Anderson, 2 and François Guillemot
A role for neural determination genes in specifying the dorsoventral identity of telencephalic neurons Carol Fode, 1 Qiufu Ma, 2,3 Simona Casarosa, 1 Siew-Lan Ang, 1 David J. Anderson, 2 and François Guillemot
Zebrafish aussicht mutant embryos exhibit widespread overexpression of ace (fgf8) and coincident defects in CNS development
 Development 126, 2129-2140 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV1367 2129 Zebrafish aussicht mutant embryos exhibit widespread overexpression of ace (fgf8) and coincident
Development 126, 2129-2140 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV1367 2129 Zebrafish aussicht mutant embryos exhibit widespread overexpression of ace (fgf8) and coincident
Europe PMC Funders Group Author Manuscript Dev Cell. Author manuscript; available in PMC 2009 December 05.
 Europe PMC Funders Group Author Manuscript Published in final edited form as: Dev Cell. 2004 February ; 6(2): 167 181. Early Steps in the Development of the Forebrain Stephen W. Wilson 1,* and Corinne
Europe PMC Funders Group Author Manuscript Published in final edited form as: Dev Cell. 2004 February ; 6(2): 167 181. Early Steps in the Development of the Forebrain Stephen W. Wilson 1,* and Corinne
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Maternal VegT and b-catenin: Patterning the Xenopus Blastula
 CHAPTER 1 Maternal VegT and b-catenin: Patterning the Xenopus Blastula Matthew Kofron, Jennifer Xanthos, and Janet Heasman 1 1.1 Introduction Loss of the maternal T-box transcription factor VegT has a
CHAPTER 1 Maternal VegT and b-catenin: Patterning the Xenopus Blastula Matthew Kofron, Jennifer Xanthos, and Janet Heasman 1 1.1 Introduction Loss of the maternal T-box transcription factor VegT has a
Dorsal and intermediate neuronal cell types of the spinal cord are established by a BMP signaling pathway
 Development 127, 1209-1220 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV8681 1209 Dorsal and intermediate neuronal cell types of the spinal cord are established by a BMP signaling
Development 127, 1209-1220 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV8681 1209 Dorsal and intermediate neuronal cell types of the spinal cord are established by a BMP signaling
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice Supplementary Figure 2.
 Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
SUPPLEMENTARY FIGURES AND TABLES AND THEIR LEGENDS. Transient infection of the zebrafish notochord triggers chronic inflammation
 SUPPLEMENTARY FIGURES AND TABLES AND THEIR LEGENDS Transient infection of the zebrafish notochord triggers chronic inflammation Mai Nguyen-Chi 1,2,5, Quang Tien Phan 1,2,5, Catherine Gonzalez 1,2, Jean-François
SUPPLEMENTARY FIGURES AND TABLES AND THEIR LEGENDS Transient infection of the zebrafish notochord triggers chronic inflammation Mai Nguyen-Chi 1,2,5, Quang Tien Phan 1,2,5, Catherine Gonzalez 1,2, Jean-François
THE ROLE OF WNT8 IN POSTERIOR MESODERM FORMATION. A Thesis CATHRYN RENEE KELTON
 THE ROLE OF WNT8 IN POSTERIOR MESODERM FORMATION A Thesis by CATHRYN RENEE KELTON Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements for the
THE ROLE OF WNT8 IN POSTERIOR MESODERM FORMATION A Thesis by CATHRYN RENEE KELTON Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of the requirements for the
 DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
DOI: 10.1038/ncb2819 Gαi3 / Actin / Acetylated Tubulin Gαi3 / Actin / Acetylated Tubulin a a Gαi3 a Actin Gαi3 WT Gαi3 WT Gαi3 WT b b Gαi3 b Actin Gαi3 KO Gαi3 KO Gαi3 KO # # Figure S1 Loss of protein
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam
 Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Hedgehog signalling maintains the optic stalk-retinal interface through the regulation of Vax gene activity
 Development 130, 955-968 2003 The Company of Biologists Ltd doi:10.1242/dev.00305 955 Hedgehog signalling maintains the optic stalk-retinal interface through the regulation of Vax gene activity Masaya
Development 130, 955-968 2003 The Company of Biologists Ltd doi:10.1242/dev.00305 955 Hedgehog signalling maintains the optic stalk-retinal interface through the regulation of Vax gene activity Masaya
Tsukushi Modulates Xnr2, FGF and BMP Signaling: Regulation of Xenopus Germ Layer Formation
 Tsukushi Modulates Xnr2, FGF and BMP Signaling: Regulation of Xenopus Germ Layer Formation Samantha A. Morris 1 *, Alexandra D. Almeida 1, Hideaki Tanaka 2, Kunimasa Ohta 2, Shin-ichi Ohnuma 1 * 1 Department
Tsukushi Modulates Xnr2, FGF and BMP Signaling: Regulation of Xenopus Germ Layer Formation Samantha A. Morris 1 *, Alexandra D. Almeida 1, Hideaki Tanaka 2, Kunimasa Ohta 2, Shin-ichi Ohnuma 1 * 1 Department
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
A dynamic fate map of the forebrain shows how vertebrate eyes form and explains two causes of cyclopia
 4613 Development 133, 4613-4617 (2006) doi:10.1242/dev.02678 A dynamic fate map of the forebrain shows how vertebrate eyes form and explains two causes of cyclopia Samantha J. England 1, Guy B. Blanchard
4613 Development 133, 4613-4617 (2006) doi:10.1242/dev.02678 A dynamic fate map of the forebrain shows how vertebrate eyes form and explains two causes of cyclopia Samantha J. England 1, Guy B. Blanchard
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Exam 2 ID#: November 9, 2006
 Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila
 Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila Melissa M. Rolls and Chris Q. Doe, Inst. Neurosci and Inst. Mol. Biol., HHMI, Univ. Oregon, Eugene, Oregon 97403 Correspondence
Baz, Par-6 and apkc are not required for axon or dendrite specification in Drosophila Melissa M. Rolls and Chris Q. Doe, Inst. Neurosci and Inst. Mol. Biol., HHMI, Univ. Oregon, Eugene, Oregon 97403 Correspondence
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
10/15/09. Tetrapod Limb Development & Pattern Formation. Developing limb region is an example of a morphogenetic field
 Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
Tetrapod Limb Development & Pattern Formation Figure 16.5(1) Limb Bud Formation derived from lateral plate (somatic) & paraxial (myotome) Fig. 16.2 Prospective Forelimb Field of Salamander Ambystoma maculatum
TOXICOLOGICAL ENDPOINTS EASILY MEASURED IN DEVELOPING ZEBRAFISH. Robert L. Tanguay Environmental and Molecular Toxicology Oregon State University
 TOXICOLOGICAL ENDPOINTS EASILY MEASURED IN DEVELOPING ZEBRAFISH Robert L. Tanguay Environmental and Molecular Toxicology Oregon State University Why zebrafish for toxicity testing? Vertebrates.. Most importantly:
TOXICOLOGICAL ENDPOINTS EASILY MEASURED IN DEVELOPING ZEBRAFISH Robert L. Tanguay Environmental and Molecular Toxicology Oregon State University Why zebrafish for toxicity testing? Vertebrates.. Most importantly:
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Building the brain (1): Evolutionary insights
 Building the brain (1): Evolutionary insights Historical considerations! Initial insight into the general role of the brain in human behaviour was already attained in antiquity and formulated by Hippocrates
Building the brain (1): Evolutionary insights Historical considerations! Initial insight into the general role of the brain in human behaviour was already attained in antiquity and formulated by Hippocrates
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Nodal Signaling is Required for Closure of the Anterior Neural Tube in Zebrafish
 The University of Akron IdeaExchange@UAkron Biology Faculty Research Biology Department 11-8-2007 Nodal Signaling is Required for Closure of the Anterior Neural Tube in Zebrafish Allisan Aquilina-Beck
The University of Akron IdeaExchange@UAkron Biology Faculty Research Biology Department 11-8-2007 Nodal Signaling is Required for Closure of the Anterior Neural Tube in Zebrafish Allisan Aquilina-Beck
Reading. Lecture VI. Making Connections 9/17/12. Bio 3411 Lecture VI. Making Connections. Bio 3411 Monday September 17, 2012
 Lecture VI. Making Connections Bio 3411 Monday September 17, 2012!! 1! Reading NEUROSCIENCE: 5 th ed, pp!507?536! 4 th ed, pp 577-609 Bentley, D., & Caudy, M. (1983). Nature, 304(5921), 62-65. Dickson,
Lecture VI. Making Connections Bio 3411 Monday September 17, 2012!! 1! Reading NEUROSCIENCE: 5 th ed, pp!507?536! 4 th ed, pp 577-609 Bentley, D., & Caudy, M. (1983). Nature, 304(5921), 62-65. Dickson,
SUPPLEMENTARY INFORMATION
 DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
DOI: 10.1038/ncb3267 Supplementary Figure 1 A group of genes required for formation or orientation of annular F-actin bundles and aecm ridges: RNAi phenotypes and their validation by standard mutations.
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome
 Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Dynamic and Differential Regulation of Stem Cell Factor FoxD3 in the Neural Crest Is Encrypted in the Genome Marcos S. Simões-Costa 1., Sonja J. McKeown 1., Joanne Tan-Cabugao 1, Tatjana Sauka-Spengler
Cooperation of sonic hedgehog enhancers in midline expression
 Developmental Biology 301 (2007) 578 589 www.elsevier.com/locate/ydbio Genomes & Developmental Control Cooperation of sonic hedgehog enhancers in midline expression Raymond Ertzer a,b, Ferenc Müller a,b,
Developmental Biology 301 (2007) 578 589 www.elsevier.com/locate/ydbio Genomes & Developmental Control Cooperation of sonic hedgehog enhancers in midline expression Raymond Ertzer a,b, Ferenc Müller a,b,
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Name. Biology Developmental Biology Winter Quarter 2013 KEY. Midterm 3
 Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM
 Annu. Rev. Neurosci. 1999. 22:261 94 Copyright c 1999 by Annual Reviews. All rights reserved THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM Kevin J. Lee and Thomas M. Jessell
Annu. Rev. Neurosci. 1999. 22:261 94 Copyright c 1999 by Annual Reviews. All rights reserved THE SPECIFICATION OF DORSAL CELL FATES IN THE VERTEBRATE CENTRAL NERVOUS SYSTEM Kevin J. Lee and Thomas M. Jessell
IN the Drosophila ventral nerve cord, there are 20
 Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.075085 Regulation of Axon Guidance by Slit and Netrin Signaling in the Drosophila Ventral Nerve Cord Krishna Moorthi Bhat,
Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.107.075085 Regulation of Axon Guidance by Slit and Netrin Signaling in the Drosophila Ventral Nerve Cord Krishna Moorthi Bhat,
The Making of the Fittest: Evolving Switches, Evolving Bodies
 INTRODUCTION MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH The types and amounts of proteins produced by a given cell in the body are very important and carefully regulated. Transcribing
INTRODUCTION MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH The types and amounts of proteins produced by a given cell in the body are very important and carefully regulated. Transcribing
Wnt2b and Fgf10. The limb identity gene Tbx5 promotes limb initiation by interacting with
 Development 129, 5161-5170 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV2958 5161 The limb identity gene Tbx5 promotes limb initiation by interacting with Wnt2b and Fgf10 Jennifer
Development 129, 5161-5170 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV2958 5161 The limb identity gene Tbx5 promotes limb initiation by interacting with Wnt2b and Fgf10 Jennifer
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Morphogens in biological development: Drosophila example
 LSM5194 Morphogens in biological development: Drosophila example Lecture 29 The concept of morphogen gradients The concept of morphogens was proposed by L. Wolpert as a part of the positional information
LSM5194 Morphogens in biological development: Drosophila example Lecture 29 The concept of morphogen gradients The concept of morphogens was proposed by L. Wolpert as a part of the positional information
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Chapter 10 Development and Differentiation
 Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Novel retinoic acid generating activities in the neural tube and heart identified by conditional rescue of Raldh2 null mutant mice
 Development 129, 2271-2282 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV9819 2271 Novel retinoic acid generating activities in the neural tube and heart identified by conditional
Development 129, 2271-2282 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV9819 2271 Novel retinoic acid generating activities in the neural tube and heart identified by conditional
178 Part 3.2 SUMMARY INTRODUCTION
 178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
178 Part 3.2 Chapter # DYNAMIC FILTRATION OF VARIABILITY WITHIN EXPRESSION PATTERNS OF ZYGOTIC SEGMENTATION GENES IN DROSOPHILA Surkova S.Yu. *, Samsonova M.G. St. Petersburg State Polytechnical University,
Cell Death & Trophic Factors II. Steven McLoon Department of Neuroscience University of Minnesota
 Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
