The calpain system and skeletal muscle growth
|
|
|
- Pearl Kory McBride
- 6 years ago
- Views:
Transcription
1 The calpain system and skeletal muscle growth Darrel E. Goll, Valery F. Thompson, Richard G. Taylor, and Ahmed Ouali Muscle Biology Group, University of Arizona, Tucson, Arizona USA, and Institut National de la Recherche Agronomique, Station de Recherches sur la Viande, Theix, St. Genes Champanelle, France. Received 27 July 1998, 1 October Can. J. Anim. Sci. Downloaded from by on 03/09/18 Goll, D. E., Thompson, V. F., Taylor, R. G. and Ouali, A The calpain system and skeletal muscle growth. Can. J. Anim. Sci. 78: The first protein of a group of proteins now identified as belonging to the calpain system was purified in The calpain system presently is known to be constituted of three well-characterized proteins; several lesser studied proteins that have been isolated from invertebrates; and 10 mrnas, two each in Drosophila and C. elegans and six in vertebrates, that encode proteins, which, based on sequence homology, belong to the calpain family. The three well-characterized proteins in the calpain family include two Ca 2+ -dependent proteolytic enzymes, µ-calpain and m-calpain, and a protein, calpastatin, that has no known activity other than to inhibit the two calpains. A substantial amount of experimental evidence accumulated during the past 25 yr has shown that the calpain system has an important role both in rate of skeletal muscle growth and in rate and extent of postmortem tenderization. Calpastatin seems to be the variable component of the calpain system, and skeletal muscle calpastatin activity is highly related to rate of muscle protein turnover and rate of postmortem tenderization. The current paradigm is that high calpastatin activity: 1) decreases rate of muscle protein turnover and hence is associated with an increased rate of skeletal muscle growth; and 2) decreases calpain activity in postmortem muscle and hence is associated with a lower rate of postmortem tenderization. This article summarizes some of the known properties of the calpain system and discusses the potential importance of the calpain system to animal science. Key words: Calpain, calpastatin, postmortem tenderization, skeletal muscle growth Goll, D. E., Thompson, V. F., Taylor, R. G. et Ouali, A Le système des calpaïnes et la croissance des muscles squeletiques. Can. J. Anim. Sci. 78: La première protéine d un groupe désormais identifié comme appartenant au système des calpaïnes a été purifiée en On sait aujourd hui que ce système comprend trois protéines bien caractérisées. Plusieurs protéines moins étudiées, isolées chez les invertébrés et 10 ARNm codant pour les protéines,dont deux chacun trouvés chez Drosophila et chez C. elegans et six, chez les vertébrés, appartiendraient également d après l homologie séquentielle à la famille des calpaïnes. Les trois protéines bien caractérisées, mentionnées plus haut, comprennent deux enzymes protéolytiques Ca 2+ - dépendantes, la µ-calpaïne et la m-calpaïne, et une protéine, la calpastatine qui semble n avoir d autre fonction que d inhiber l activité des deux calpaïnes. Un corpus substantiel d évidence expérimentale accumulé au cours des 25 dernières années révèle que le système des calpaïnes joue un rôle important à la fois à l égard du taux de croissance des muscules squelettiques et du degré d attendrissage de la viande postmortem. La calpastatine semble être la composante variable du système et son activité dans les muscles squelettiques est étroitement reliée au taux de renouvellement des protéines musculaires et au taux d attendrissage postmortem. Le paradigme actuel est que la forte activité de la calpastatine: 1) ralentit le taux de renouvellement des protéines musculaires et, partant, s accompagne d un taux accru de la croissance des muscles squeletiques et 2) diminue l activité des calpaïnes dans le muscle après l abattage et, par là, ralentit le taux d attendrissage postmortem. Dans cette mise au point bibliographique, nous récapitulons certaines des propriétés connues du système des calpaïnes et nous en examinons l importance éventuelle pour la recherche zootechnique. Mots clés: Calpaïne, calpastatine, attendrissage postmortem, croissance des muscles squelettiques The rate and extent of skeletal muscle growth ultimately depends on only three factors: 1) rate of muscle protein synthesis; 2) rate of muscle protein degradation; and 3) the number and size of skeletal muscle cells. The numerous agents that have been reported to influence or be related to rate of muscle growth, including various hormones or growth factors such as insulin, the insulinlike growth factors I and II, growth hormone, etc. all exert their effects by influencing one or more of these three basic factors. Genetic selection for increased skeletal muscle mass involves these three factors. Double-muscled cattle have a larger number of muscle cells (fibers) (Kambadur et al. 1997) and a greater capacity for protein synthesis (Bjercke et al. 1984a,b) than normal cattle. A member of the TGF-β superfamily (GDF-8; also called myostatin) has recently been identified 503 as the gene involved in the double muscle phenotype (Kambadur et al. 1997; McPherron and Lee 1997; McPherron et al. 1997); this phenotype is inherited as a monogenic, autosomal trait that maps to the centromeric end of bovine chromosome 2 (Grobet et al. 1997). Disruption of the myostatin gene, caused either by deletion of an 11-bp region in the coding sequence for the carboxy-terminal domain of the protein (Grobet et al. 1997; Kambadur et al. 1997) or a G A mutation that converts C314 to Y314 (Kambadur et al. 1997; McPherron and Lee 1997), results in the pronounced muscle hypertrophy characteristic of double muscling. The 11-bp deletion is found in Belgian blue double-muscled cattle, and the cysteine to tyrosine conversion is Abbreviations: ERM, easily releasable filaments
2 504 CANADIAN JOURNAL OF ANIMAL SCIENCE found in Piedmontese double-muscled cattle. Hence, this TGF-β isoform is a negative regulator of growth. Myostatin is an extracellular molecule that binds to a receptor on the skeletal muscle cell to exert its effects. This binding evidently influences muscle cell number and rate of muscle protein synthesis; it will be interesting to learn whether rate of muscle protein degradation (calpain activity) is also affected. Many studies involving treatments designed to increase rate of muscle growth have simply used measurements of live animal or carcass weight to evaluate the efficacy of the treatments used, and the effects of these treatments on the three basic factors determining muscle growth have not been determined. Studies attempting to improve the nutritional status of animals are likely to increase the rate of muscle protein synthesis, but may also affect rate of muscle protein degradation (Goll et al. 1989; Reeds et al. 1986). Indeed, differences in rates of muscle growth in domestic animals often are due to differences in rate of muscle protein degradation with little or no change in rate of protein synthesis (Bohorov et al. 1987; Maruyama et al. 1978; Reeds et al. 1986). Because skeletal muscle growth is a complex process involving a large number of pathways, it is useful and often simplifying to consider the outcome of experiments in the context of which of these three factors have been affected. This review will focus on the role of the calpain system in skeletal muscle growth. It was originally proposed that the calpain system was responsible for initiating metabolic turnover of the myofibrillar proteins and that it therefore affected only rate of muscle protein degradation (Dayton et al. 1975; Goll et al. 1983, 1992b). Recent studies, however, have shown that calpain activity is required for myoblast fusion (Balcerzak et al. 1995; Barnoy et al. 1997; Kumar et al. 1992; Kwak et al. 1993a,b; Temm-Grove et al. 1999) and for cell proliferation (Mellgren 1997; Watanabe et al. 1989; Zhang et al. 1997) in addition to cell growth (Mellgren et al. 1994). Hence, the calpain system may also affect the number of skeletal muscle cells (fibers) in domestic animals by altering rate of myoblast proliferation and modulating myoblast fusion. First, some properties of the calpain system and the evidence that this system is involved both in normal skeletal muscle growth and in pathological loss of skeletal muscle mass will be summarized briefly. The unique effects of the calpains on skeletal muscle proteins and how these unique effects are related to the probable mechanism by which the calpains degrade myofibrillar proteins will then be described. Finally, some of the current questions related to activity of the calpains in living cells will be discussed, and possible studies that may clarify the role of the calpain system in skeletal muscle growth will be proposed. Table 1. General properties of the calpain system Occurence Found in all vertebrate cells that have been examined; calpain-like proteins have been isolated from Drosophila and from a blood fluke, Schistosoma; mrnas encoding for molecules having sequence homology to the calpains have been identified in C. elegans, Drosophila, and Schistosoma; complete genomic sequences of yeast (Saccharomyces) and Escherichia coli do not contain calpain-like sequences; relation of Ca 2+ -dependent proteases in plants to the calpains is still unclear. Ubiquitous Calpains [Ca 2+ ] required for 1 2 Name Polypeptides maximal activity µ-calpain 80, 28-kDa 3 50 µm m-calpain 80, 28-kDa µm µ/m-calpain 80, 28-kDa 420 µm Tissue-specific calpains Name Polypeptides Tissue skm-calpain, p94, calpain 3 94, 82-kDa skeletal muscle, rat lens n-calpain-2, ncl-2 80kDa stomach muscle n-calpain-3, ncl-3 43kDa stomach muscle All calpains that have been isolated in protein form are cysteine proteases with ph optima of Calpastatin Multiheaded protein inhibitor that inhibits only the calpains; expressed in several different isoforms that have one, three or four inhibitory domains and different N-terminal sequences. Cellular distribution The calpains and calpastatins studied thus far are located exclusively intracellularly; various proportions of the calpains are associated with subcellular organelles, which are primarily myofibrils in skeletal muscle, but may include the plasma membrane, mitochondria, and nuclei. PROPERTIES OF THE CALPAIN SYSTEM Some properties of the calpain system and the calpains are listed in Table 1 and Fig. 1. Incubation in a Ca 2+ -containing solution resulted in complete loss of Z-disks in skeletal muscle myofibrils (Busch et al. 1972) and led to purification of a Ca 2+ -dependent proteolytic enzyme, initially named CAF, in 1976 (Dayton et al. 1976a). Purification and characterization of CAF (Dayton et al. 1976b) was followed almost immediately by identification in skeletal muscle extracts of an inhibitor of the Ca 2+ -dependent proteolytic activity (Okitani et al. 1976), and then several years later by identification (Mellgren 1980) and purification (Dayton et al. 1981; Szpacenko et al. 1981) of a second Ca 2+ -dependent protease. The two Ca 2+ -dependent proteolytic activities have been named m-calpain (originally CAF) and µ-calpain (Table 1) and the specific inhibitor has been named calpastatin (Murachi 1989). In 1989, a mrna encoding a molecule with approximately 50% sequence homology to the two calpains was identified specifically in skeletal muscle (Sorimachi et al. 1989), and this discovery has been followed by identification of calpain-like mrnas in a number of other tissues and species (Sorimachi et al. 1994; Dear et al. 1997). With only a few exceptions, however, the molecules encoded by these mrnas have not been isolated in protein form, and nothing therefore is known about their catalytic properties or even in fact whether they are active proteolytic enzymes. Until more information is available on the properties of these molecules, studies on the role of the calpain system in skeletal muscle growth likely will continue to emphasize only µ- and m-calpain. Calpastatin has not been studied as extensively as the calpains, although it presently seems that levels of calpastatin vary more widely in response to different treatments than
3 GOLL ET AL. THE CALPAIN SYSTEM 505 Can. J. Anim. Sci. Downloaded from by on 03/09/18 levels of either µ- or m-calpain do. It has become evident during the past several years that a number of different calpastatin isoforms are expressed from a single calpastatin gene as the result of different start sites of translation/transcription or by alternative splicing mechanisms (Lee et al. 1992; Cong et al. 1998). The significance of these different calpastatin isoforms is unclear, but it is probable that their expression is regulated differently and that they differ in their ability to inhibit calpain activity. Calpastatin also exists in a phosphorylated form in vivo, with this phosphorylation occurring principally on serine residues (Adachi et al. 1991). Phosphorylation has been shown to alter the ability of calpastatin to inhibit proteolytic activity of µ- or m-calpain (Salamino et al. 1994). Hence, studies on the role of calpastatin in muscle growth need to determine the nature of the calpastatin present as well as the levels of this polypeptide. ROLE OF THE CALPAIN SYSTEM IN SKELETAL MUSCLE GROWTH There is ample evidence that the calpain system has an important role both in normal, postnatal skeletal muscle growth and in the muscle wasting observed in muscular dystrophies and other conditions accompanied by loss of muscle mass. The intracellular free Ca 2+ concentration, [Ca 2+ ], is elevated in the muscular dystrophies and other muscle pathologies, and this elevated intracellular [Ca 2+ ] evidently stimulates calpain activity (Mongini et al. 1988; Turner et al. 1988, 1991; Hopf et al. 1996). The structural changes observed in rapidly atrophying muscles (Cullen and Pluskal 1977; Cullen and Fulthorpe 1982) are mimicked closely by treatment of myofibrils from normal muscle with µ-calpain (Dayton et al. 1981; Goll et al. 1991, 1992b) or m-calpain (Dayton et al. 1975, 1976b; Goll et al. 1991, 1992b). SDS- PAGE has shown that the calpains cause the same degradative changes in myofibrils as those seen in Duchenne muscular dystrophy (Sugita et al. 1980) or other rapidly atrophying muscle (Dayton et al. 1979; Ishiura et al. 1980). A number of studies have shown that the calpain system also has a role in normal skeletal muscle growth. Fig. 1. Schematic diagram showing the structure of the µ- and m-calpain molecules as indicated by their amino acid sequences. The numbers shown are for the human calpains; the calpains are highly conserved among vertebrate species, and amino acid sequences of calpains from rabbit, pig, and porcine are approximately 90 95% homologous. The bars in domains III, IV, and VI represent EF-hand-like sequences and are potential Ca 2+ -binding sites. Administering various β-adrenergic agonists to animals results in a 10 30% increase in rate of accumulation of muscle mass (Yang and McElligott 1989). Although various studies using different species, different β-adrenergic agonists, and different conditions have produced slightly different results, most studies agree that administration of β-adrenergic agonists increases both the rate and the efficiency at which skeletal muscle protein is accumulated. Administration of β-adrenergic agonists also affects activity of the calpain system (Forsberg et al. 1989). Muscle calpastatin activity is significantly increased (Higgins et al. 1988; Wang and Beermann 1988; Forsberg et al. 1989; Kretchmar et al. 1989, 1990; Bardsley et al. 1992; Parr et al. 1992; Pringle et al. 1993), with this increase ranging from 52 (Forsberg et al. 1989) to 430% (Kretchmar et al. 1990). Muscle µ-calpain activity is either decreased or remains unchanged, whereas m-calpain activity seems to be increased (Higgins et al. 1988; Wang and Beermann 1988; Kretchmar et al. 1989, 1990; Koohmaraie and Shackelford 1991), although one study has found that activity of both µ- and m- calpain is decreased (Forsberg et al. 1989). The callipyge phenotype in sheep is inherited as an autosomal, dominant gene that maps to the ovine chromosome 18 (Cockett et al. 1994). Skeletal muscle mass in callipyge lambs is 30 40% greater than in half-siblings not expressing the callipyge trait, with most of this increase resulting from increases in muscles in the pelvic limbs and back and smaller or no increases in muscles from the fore limbs (Jackson et al. 1997; Koohmaraie et al. 1995). Calpastatin activities in the affected muscles from callipyge lambs are % higher than in the same muscles from normal lambs, whereas calpastatin activities in the unaffected callipyge muscles (e.g., supraspinatus) were the same as those in the supraspinatus muscle from normal lambs (Koohmaraie et al. 1995). These results suggest that increased rates of skeletal muscle growth can result from a decrease in rate of muscle protein degradation, and that this decreased rate of muscle protein degradation is associated with a decrease in activity of the calpain system, due principally to a large increase in calpastatin activity.
4 506 CANADIAN JOURNAL OF ANIMAL SCIENCE The recent studies indicating that calpain activity is required for cells (or myoblasts for this review) to progress through the G1 to S phase of the mitotic cycle (Mellgren 1997; Zhang et al. 1997) and for myoblast fusion (Kwak et al. 1993a, b; Cottin et al. 1994; Balcerzak et al. 1995; Barnoy et al. 1997; Temm-Grove et al. 1999) suggest that increased calpain activity during muscle development may be associated with an increased number of myoblasts that would result in a larger number of muscle nuclei and hence potentially larger muscle fibers in mature muscle. Developing C 2 C 12 myoblasts contain only m-calpain (Cottin et al. 1994; Temm-Grove et al. 1999), and because β-adrenergic agonist administration to growing animals seems to have less effect on m-calpain than on calpastatin, it is tempting to speculate that increasing m-calpain activity, especially in developing muscle, would result in an increased rate of skeletal muscle growth. EFFECTS OF THE CALPAINS ON MUSCLE PROTEINS A number of reports have shown that the calpains have very limited and specific effects on skeletal muscle proteins (Dayton et al. 1975; Goll et al. 1992b). Myosin, the major muscle protein, is degraded very slowly, and even this slow degradation is limited to a few cleavages of the light chains and a small amount of nibbling at the end of the large 200- kda heavy chain. Undenatured actin, the second most abundant protein in skeletal muscle myofibrils, is not cleaved by either µ- or m-calpain. Both µ- and m-calpain rapidly cleave troponin T, desmin, vinculin, talin, spectrin, nebulin, and titin; more slowly cleave troponin I, filamin, C-protein, dystrophin, and tropomyosin, and cleave α-actinin and M protein very slowly. Ultrastructurally, incubation with the calpains results first in loss of the N 2 line, and then in complete loss of Z-disks leaving a gap in the middle of the sarcomere; loss of periodicity in the I-band area, due likely to troponin and tropomyosin degradation (Goll et al. 1992b) occurs at the same time as loss of Z-disks. α-actinin is a major Z-disk protein, but it is degraded slowly by the calpains, and the loss of Z-disk structure is caused by release of α-actinin from Z-disks in a nearly intact form (Goll et al. 1991). Although several studies have reported that myosin, actin, and α-actinin are degraded by the calpains, it is unclear whether these studies have used undenatured proteins. Both µ- and m-calpain rapidly degrade denatured actin, myosin, and α-actinin. The calpains specifically cleave several cytoplasmic (sarcoplasmic) proteins including most protein kinases and phosphatases, but they do not cause bulk degradation of sarcoplasmic proteins to small fragments (Tan et al. 1988). Indeed, if a crude sarcoplasmic protein extract is incubated with the calpains, no proteolytic degradation of the crude fraction can be detected by measuring the release of TCAsoluble material (Tan et al. 1988). In addition to degrading a limited number of protein substrates, the calpains cleave relatively few peptide bonds in each protein and leave large polypeptide fragments rather than reducing the protein to small peptides and amino acids. This unique property of the calpains has important consequences when considering whether the calpains have a role in muscle protein turnover and what this role may be. Most studies attempting to measure muscle protein turnover use release of free amino acids (frequently tyrosine) or 3-methyl histidine to estimate rate of muscle protein degradation (Gopinath and Kitts 1984; Smith and Sugden 1986). Because the calpains do not degrade proteins to free amino acids, measurements of the release of free amino acids or 3- methyl histidine are not directly related to calpain activity, but rather reflect activity of other protease(s) that produce free amino acids. Consequently, studies attempting to determine the role of the calpain system in muscle protein turnover need to use measurements other than or in addition to the release of free amino acids or 3-methyl histidine if they are to assess the role of the calpain system. MUSCLE PROTEIN TURNOVER AND THE ROLE OF THE CALPAIN SYSTEM It is now evident that metabolic turnover of skeletal muscle proteins is a more complex process than turnover of proteins in other cells. This increased complexity is caused by the presence of myofibrillar proteins, which constitute 55 65% of the total protein in skeletal muscle cells, and which are present in highly ordered structures called myofibrils. Because the myofibrillar structure must remain intact for skeletal muscle cells to be functional, turnover of myofibrillar proteins, which represent the major fraction of total protein in muscle cells, must proceed via a different mechanism than turn over of the sarcoplasmic (cytoplasmic) proteins does. Although the mechanism by which myofibrillar proteins turn over is still unclear and remains an area of active research, it presently seems that this turnover proceeds in at least two steps: 1) disassembly or removal of the proteins from the myofibrillar structure; this disassembly must occur without disruption or severing of the myofibril which extends continuously from one end of the muscle cell (fiber) to the other (Fig. 2); and 2) degradation of the individual myofibrillar proteins to small peptides and free amino acids. Kinetic studies (Clark 1993) have shown that proteins in adult cardiac myocytes are degraded from two different pools; one that comprises approximately 10% of total muscle protein and that turns over rapidly with a mean half-life of 11.9 h; and a second pool that comprises the remaining 90% of the total muscle protein and that turns over more slowly with a half-life of 15.6 d. Similar studies have not been done on skeletal muscle cells, but the structure of cardiac myofibrils, which constitute a slightly smaller percentage of total muscle protein (45 55%) than in skeletal muscle, is very similar to the structure of skeletal muscle myofibrils, and the myofibrillar proteins probably turn over via the same mechanism in the two types of cells. It seems likely that the rapidly turning over pool contains, in addition to some of the sarcoplasmic proteins, those myofibrillar proteins that have been disassembled from the myofibril, and hence are available for degradation to amino acids/small peptides. The presence of two pools of proteins turning over at different rates in muscle cells does not prove that the myofibrillar proteins must be disassembled from the myofibril
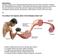
5 GOLL ET AL. THE CALPAIN SYSTEM 507 Can. J. Anim. Sci. Downloaded from by on 03/09/18 Fig. 2. A schematic diagram showing the structure of striated muscle beginning at the muscle fiber (cell) and extending in increasingly higher magnification to the actin and myosin molecules. Most of the interior of muscle cells (A) is occupied by protein threads called myofibrils (B) that extend continuously from one end of the cell to the other. If examined at high resolution in the electron microscope, it is possible to see that myofibrils contain an interdigitating array of smaller filaments called myofilaments (C). The two main filaments shown in this diagram (there is a third set of filaments composed principally of titin and nebulin not shown in this diagram) contain actin (thin filaments, D and E) and myosin (thick filaments; F and G). The thick myosin filaments connect to the thin actin filaments via myosin cross bridges that extend outwardly from the surface of the thick filaments (C). Muscle contraction is caused by a sliding of the thick and thin filaments past one another, resulting in a shortening of the myofibril. For muscle cells to remain functional, the continuity of the myofibril must remain intact. Consequently, myosin molecules, for example, cannot be removed from the interior of a thick filament disrupting the filament and breaking the connection of adjacent Z discs (C). Ostensibly, the only way to turn over myofibrillar proteins without disrupting the continuity of the myofibrillar structure would be to remove filaments from the outer surface of the myofibril, thereby gradually reducing its diameter. Diagram is reproduced from A Textbook of Histology (Bloom and Fawcett 1975) with permission of the authors and Edward Arnold Publishers. before they can be degraded to amino acids. A number of studies have shown, however, that approximately 5 15% of total myofibrillar protein in skeletal and cardiac muscle cells can be dissociated from intact myofibrils in the form of myofilaments by using gentle agitation in an ATP-containing solution (van der Westhuyzen et al. 1981). These easily releasable filaments (ERM) lack α-actinin, desmin, titin, and other cytoskeletal proteins such as filamin having molecular masses above 200 kda (Reville et al. 1994; van der Westhuyzen et al. 1981) but contain the major myofibrilar proteins, actin and myosin. Proteins in the ERM fraction turn over rapidly, indicating that they are in a pool of rapidly degraded proteins (van der Westhuyzen et al. 1981). ERM levels in muscle increase significantly in response to treatments that increase calpain activity (van der Westhuyzen et al. 1981; Dahlmann et al. 1986; Belcastro et al. 1991; Reville et al. 1994) and decrease in response to treatments that inhibit calpain activity (van der Westhuyzen et al. 1981; Dahlmann et al. 1986; Reville et al. 1994). Brief treatment with calpain increases the amount of ERM in myofibrillar preparations (van der Westhuyzen et al. 1981). As described in the preceding section, the calpains make specific cleavages in those cytoskeletal proteins that are involved in maintaining the myofibrillar structure: 1) degradation of desmin, vinculin, talin, dystrophin, and spectrin in addition to other minor proteins that are responsible for linking adjacent myofibrils together and to the sarcolemma; 2) loss of Z-disks and release of α-actinin evidently due to cleavage of the N-terminal (Z-disk) end of the large titin polypeptide and to the rapid cleavage of nebulin, because α- actinin has been reported to bind to both titin and nebulin in the Z-disk (Nave et al. 1990; Ohtsuka et al. 1997; Sorimachi et al. 1997); 3) degradation of C-protein, which encompasses thick filaments like staves around a barrel, and whose
6 508 CANADIAN JOURNAL OF ANIMAL SCIENCE degradation would favor the release of individual thick filaments from the surface of myofibrils; 4) degradation of troponin T and tropomyosin, which would contribute to weakening of the thin filament and increase the tendency for this structure to dissociate into actin monomers; and 5) degradation of M proteins, which would, along with the degradation of C protein, favor the release of myosin filaments. On the other hand, the calpains are unique among proteolytic enzymes in that they do not rapidly degrade the major muscle proteins, actin and myosin, which also are the major constituents of the ERM. The large size and highly ordered structure of intact myofibrils (1 2 µm in diameter and extending the entire length of the cell, up to several mm) would prevent them being taken up into lysosomes (Lowell et al. 1986) or from entering the central channel containing the active sites of the proteasome (multicatalytic protease). Indeed, neither myofibrils nor structurally recognizable fragments of myofibrils have been observed in lysosomal structures, even in rapidly atrophying muscle (Goll et al. 1989), and the proteasome has no effect on myofibrillar proteins when they are in the myofibrillar structure (Koohmaraie 1992; Solomon and Goldberg 1996). Both lysosomal cathepsins and the proteasome, however, rapidly degrade individual myofibrillar proteins under the appropriate conditions. Consequently, the available evidence suggests that the myofibrillar proteins in skeletal (and probably also in cardiac) muscle cells are first removed or released from the myofibril either in the form of filaments or as individual protein molecules and that these protein molecules or the proteins in the filaments are then degraded to amino acids by proteolytic systems in the cell cytoplasm. The mechanism by which myofilaments or individual proteins are released from myofibrils is still unclear. Some reports have suggested that individual proteins are removed from within myofibrils and that newly synthesized proteins are then incorporated into the area vacated by the departing molecule. The mechanism by which this exchange is accomplished without disrupting the myofibrillar structure, however, is not clear. The presence of ERM, on the other hand, suggests that myofibrillar proteins/filaments are released from the surface of myofibrils, resulting in a myofibril with an increasingly smaller diameter and a growing pool of individual thick and thin myofilaments and free myofibrillar proteins (Goll et al. 1992b). This mechanism would leave functionally intact, although smaller, myofibrils as turnover progressed. The released myofilaments/ myofibrillar proteins could either reassemble back onto the surface of the myofibril or be degraded to amino acids/small peptides by cytoplasmic proteinases (likely the proteasome and lysosomal cathepsins). According to this latter mechanism, the calpains would initiate disassembly of the myofibril by specific cleavages of Z-disk proteins at the surface of the myofibril (probably titin and nebulin), releasing the thin filaments from their attachments to the myofibril. The myosin thick filaments attached to the released thin filaments would dissociate in the presence of the ATP in the cell, and calpain-induced cleavage of C-protein and M-protein would lead to further dissociation of the thick filaments to individual myosin molecules that are degraded by the proteasome or taken up into lysosomes and degraded by lysosomal cathepsins. Similarly, calpain-induced cleavage of tropomyosin and troponin T and I together with degradation of nebulin would favor dissociation of thin filaments to actin monomers that can be degraded by the proteasome or that can be engulfed by lysosomes and degraded by lysosomal cathepsins. Although this disassembly and then degradation process is teleologically attractive and proposes definite roles for the calpains and the proteasome that are consistent with the known properties of these systems, there is little direct experimental evidence that either supports or refutes this mechanism. A study done over 25 years ago suggested that newly synthesized myofibrillar proteins were added exclusively to the surface of growing myofibrils (Morkin 1970), an observation consistent with assembly and disassembly occurring at the surface and not in the interior of myofibrils. On the other hand, this mechanism also indicates that the interior of myofibrils would be immortal unless the entire myofibril was turned over, and the implications of such immortality to physiological functioning of the myofibril are unclear. Hence, although there is considerable circumstantial evidence indicating that the calpains have an important role in initiating turnover of the myofibrillar proteins, very little information is available on the exact mechanism by which such initiation occurs. It is clear that the calpains cannot degrade the myofibrillar or any other class of proteins to amino acids, and it is equally clear that turnover of intact myofibrils cannot be initiated by either of the other two major proteolytic systems in muscle cells, the proteasome and the lysosomal cathepsins. Consequently, at least two and perhaps three proteolytic systems are involved in turnover of the myofibrillar proteins. It is highly likely that the calpain system and the proteasome are among these. OTHER PROPERTIES OF THE CALPAINS THAT ARE IMPORTANT TO ANIMAL SCIENCE The suggested roles of the calpain system in myoblast proliferation and fusion during muscle development have already been discussed. Other less direct evidence has indicated that the calpains have a role in signal transduction processes. The nature of this role is less well-defined, however, and is related to ability of the calpains to rapidly cleave many of the kinases and phosphatases involved in signal transduction. Calpain cleavage frequently ablates the regulation that normally governs activity of these kinases/phosphatases, and leaves constitutively active enzymes. The effects of constitutively active enzymes on signal transduction are unclear and are likely to be complex. It is ironic that the most clearly defined property of the calpains does not involve a function in living cells, but rather its role in postmortem tenderization (Boehm et al. 1998; Goll et al. 1992a; Taylor et al. 1995). An enormous amount of evidence acquired during the past 25 years has indicated that the calpains are responsible for up to 95% of all proteolytically induced postmortem tenderization that occurs during the first 7 14 d of postmortem storage at 2-4 C. Storage for longer periods postmortem or at higher
7 temperatures above 20 C may involve some catheptic proteolysis, but these conditions are not used frequently in normal processing. It is still unknown whether postmortem proteolysis involves primarily µ- or m-calpain or both (Boehm et al. 1998) and exactly what calpain cleavages are most important to tenderization. Studies on animals that have received β-agonists or animals having a Bos indicus or callipyge genetic background have indicated that less tender meat or meat that undergoes little postmortem tenderization has higher calpastatin activities than meat whose tenderness increases substantially during postmortem storage. Muscle calpain activity, however, seems to vary little among these different groups of animals. The currently accepted hypothesis, therefore, is that high calpastatin activity decreases ability of the calpains to degrade myofibrillar proteins during postmortem storage, and that because muscle calpastatin levels vary more widely in response to different treatments than muscle calpain activities do, muscle calpastatin and not muscle calpain activity is related to degree of postmortem tenderization. CURRENT QUESTIONS INVOLVING THE ROLE OF THE CALPAINS IN ANIMAL SCIENCE The presently available evidence indicates that the calpain system is implicated in two of the three factors that determine rate of skeletal muscle growth. 1) The calpains initiate metabolic turnover of the myofibrillar proteins and therefore have an important influence on rate of muscle protein degradation. The known properties of the calpains indicate that the calpain system does not have an important role in metabolic turnover of sarcoplasmic proteins, which constitute approximately 30 35% of total muscle protein, and that the calpains only initiate turnover of the myofibrillar proteins and do not degrade these proteins to amino acids. Consequently, other proteolytic systems are also involved in muscle protein turnover. It seems likely that the proteasome is responsible for degradation of the proteins released from myofibrils by the calpains and may also be involved in metabolic turnover of the sarcoplasmic proteins. Nevertheless, the myofibrillar proteins constitute the major fraction of total protein in muscle cells, and because their turnover cannot occur until they are released from the myofibrillar structure, calpain activity may be the rate limiting step in metabolic turnover of the myofibrillar proteins. 2) Because calpain activity is required for cell proliferation and for myoblast fusion, the calpain system also has an important influence on the number and size of muscle cells in mature skeletal muscle. Although the role of the calpain system in turnover of the myofibrillar proteins has been at least partly characterized, the role of calpains in muscle cell number and size is still poorly defined. Based on the evidence available thus far, it may be suggested that enhanced calpain activity in embryonic muscle results in an increased rate of myoblast proliferation and hence an increased number of muscle cell nuclei. It has been shown that mass/size of postnatal muscle fibers/cells is proportional to the number of nuclei in these cells. Hence, increased calpain activity in developing muscle may be expected to be associated with increased muscle fiber size. Because calpain activity is GOLL ET AL. THE CALPAIN SYSTEM 509 required for myoblast fusion, increased calpain activity in developing muscle may also be associated with an increased number of muscle fibers in mature skeletal muscle. It is interesting to note that increased calpain activity in developing muscle is associated with increased muscle mass, whereas increased calpain activity in mature muscle is associated with decreased muscle mass. There are several fundamental questions concerning calpain activity in skeletal muscle cells. Two of the currently most important questions are: 1) how is calpain activity regulated in living muscle cells? and 2) what are the protein substrates of the calpains in skeletal muscle cells? 1) Skeletal muscle cells contain sufficient calpain to destroy all Z-disks in these cells in 5 10 min. Hence, most of the calpain in muscle cells must be inactive most of the time. It seems likely that calpain activity is regulated by calpastatin levels and the nature of the calpastatin isoforms present and by the Ca 2+ requirement of the calpains (Goll et al. 1992c). The nature of this regulation, however, is almost completely unknown. The Ca 2+ concentrations required for calpain activity in in vitro assays (Table 1) are much higher than the µm free Ca 2+ concentrations that exist in living cells. Cells, therefore, must have a mechanism that lowers or abrogates this Ca 2+ requirement. The nature of this mechanism is a complete mystery at present. A number of different calpastatin isoforms exists (Cong et al. 1998), but the physiological significance of these different forms of calpastatin is unclear. β-agonist administration, for example, has been shown to induce expression of specific calpastatin mrnas (Parr et al. 1992; Killefer and Koohmaraie 1994). Do these different calpastatins differ in their ability to inhibit the calpains? The calpains autolyze rapidly in the presence of Ca 2+ in vitro. Although it was originally suggested that autolysis was required for activation of calpain proteolytic activity, recent studies involving mutation of the autolytic cleavage site (Elce et al. 1997) have proven that the unautolyzed calpains are fully active proteases. It seems unlikely, therefore, that autolysis per se regulates calpain activity in skeletal muscle cells. 2) Although in vitro assays have identified a number of muscle proteins that are cleaved by the calpains in these assays, it is unclear whether all these proteins are also calpain substrates in living cells. It seems likely that the proteins that are cleaved by the calpains in vivo will differ depending on the state of the cell, the proximity of the protein substrate to the calpain, and other conditions such as activation of the process that reduces the Ca 2+ requirement needed for activity. The mechanism(s) involved in this selective substrate cleavage also remain unknown. Several simple studies that could be initiated without difficulty would likely clarify some of the currently unanswered questions as to how the calpain system functions in living skeletal muscle cells. Determination of the calpastatin isoforms present in different muscles and whether these isoforms change in response to treatments that alter muscle growth (such as β-agonist administration) would show whether the relationship between calpastatin activity and rate of skeletal muscle growth in mature animals is the result simply of a change to a calpastatin isoform with different
8 510 CANADIAN JOURNAL OF ANIMAL SCIENCE calpain inhibitory properties. Characterization of the calpain system in developing muscle is needed to show which calpastatin isoforms are present and whether both µ- and m- calpain exist in differentiating myoblasts. The studies done thus far on developing muscle have indicated that only m- calpain is present before fusion and have produced differing results as to whether calpastatin exists. Information from studies such as these is needed to provide a basis for more detailed studies that would clarify the role of the calpain system in skeletal muscle growth. ACKNOWLEDGMENTS This work was supported by grants from the USDA National Research Initiative Competitive Grants Program, and ; the Muscular Dystrophy Association; the Arizona Agricultural Experiment Station, Project 28, a contributing project to USDA Regional Research Project NC-131; and INRA. We thank Janet Christner for her assistance in assembling the manuscript while the authors were in a different country. Adachi, Y., Ishida-Takahashi, A., Takahashi, C., Takano, E., Murachi, T. and Hatanaka, M Phosphorylation and subcellular distribution of calpastatin in human hematopoietic cells. J. Biol. Chem. 266: Balcerzak, D., Poussard, S., Brutis, J. J., Flamrani, N., Soriano, M., Cottin, P. and Ducastaing, A An antisense oligonucleotide to m-calpain mrna inhibits myoblast fusion. J. Cell Sci. 108: Bardsley, R. G., Allock, S. M. J., Dawson, J. M., Dumelow, J. M., Higgins, J. A., Lasslett, Y. V., Lockley, A. K., Parr, T. and Buttery, P. J Effect of β-agonists on expression of calpain and calpastatin activity in skeletal muscle. Biochemie 74: Barnoy, S., Glaser, T., Kosower, N. S Calpain and calpastatin in myoblast differentiation and fusion effects of inhibitors. Biochim. Biophys. Acta 1358: Belcastro, A. N., Machan, C. and Gilchrist, J. S Diabetes enhances calpain degradation of cardiac myofibrils and easily releasable myofilaments. Pages in M. Nagano and N.S. Dhalla, eds. The diabetis heart. Raven Press Ltd., New York, NY. Bjercke, R. J., Goll, D. E. and Robson, R. M. 1984a. Development of methods to measure activity of polysomes and cytoplasmic enyzmes from bovine skeletal muscle in in vitro protein synthesis assays. J. Anim. Sci. 59: Bjercke, R. J., Goll, D. E., Robson, R. M. and Dutson, T. R. 1984b. Relative roles of polysomes and cytoplasmic enzymes in regulating bovine skeletal muscle protein synthesis. J. Anim. Sci. 59: Bloom, W. and Fawcett, D. W Muscular tissue. Pages in A textbook of histology. 10th ed. W.B. Saunders Company, Philadelphia, PA. Boehm, M. L., Kendall, T. L., Thompson, V. F. and Goll, D. E Changes in the calpains and calpastatin during postmortem storage of bovine muscle. J. Anim. Sci. 76: Bohorov, O., Buttery, P. J., Correia, J. H. R. D. and Soar, J. B The effect of the β2-adrenergic agonist, clenbuterol, implantation with oestradiol plus trenbolone on protein metabolism in wether lambs. Brit. J. Nutr. 57: Busch, W. A., Stromer, M. H., Goll, D. E. and Suzuki, A Ca 2+ -specific removal of Z-lines from rabbit skeletal muscle. J. Cell Biol. 52: Clark, W. A Evidence for posttranslational kinetic compartmentation of protein turnover pools in isolated adult cardiac myocytes. J. Biol. Chem. 268: Cockett, N. F., Jackson, S. P., Shay, T.L., Nielsen, D., Moore, S. S., Stelle, M. R., Barendse, W., Green, R. D. and Georges, M Chromosomal localization of the callipyge gene in sheep (Ovis aries) using bovine DNA markers. Proc. Natl. Acad. Sci. 91: Cong, M., Thompson, V. F., Goll, D. E. and Antin, P The bovine calpastatin gene promoter and a new N-terminal region of the protein are targets for camp-dependent protein kinase activity. J. Biol. Chem. 273: Cottin, P., Brutis, J. J., Poussard, S., Elamrani, N., Broncard, S. and Ducastaing, A Ca 2+ -dependent proteinases (calpains) and muscle cell differentiation. Biochim. Biophys. Acta 1223: Cullen, M. J. and Fulthorpe, J. J Phagocytosis of the A band following Z line and I band loss. Its significance in skeletal muscle breakdown. J. Pathol.138: Cullen, M. J. and Pluskal, M. G Early changes in the ultrastructure of denervated rat skeletal muscle. Exp. Neurol. 56: Dahlmannn, B., Rutschmann, M. and Reinauer, H Effect of starvation or treatment with corticosterone on the amount of easily releasable myofilaments in rat skeletal muscle. Biochem. J. 234: Dayton, W. R., Goll, D. E., Stromer, M. H., Reville, W. J., Zeece, M. G. and Robson, R. M Some properties of a Ca 2+ -activated protease that may be involved in myofibrillar protein turnover. Pages in E. Reich, D. B. Rifkin, and E. Shaw, eds. Cold Spring Harbor Conferences on Cell Proliferation. Vol. 2. Proteases and Biological Control. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY. Dayton, W. R., Goll, D. E., Zeece, M. G., Robson, R. M. and Reville, W. J. 1976a. A Ca 2+ -activated protease possibly involved in myofibrilar protein turnover. Purification from porcine muscle. Biochemistry 15: Dayton, W. R., Reville, W. J., Goll, D. E. and Stromer, M. H. 1976b. A Ca 2+ -activated protease possibly involved in myofibrillar protein turnover. Partial characterization of the purified enzyme. Biochemistry 15: Dayton, W. R., Schollmeyer, J. V., Chan, A. C. and Allen, C.E Elevated levels of a calcium-activated muscle protease in rapidly atrophying muscles from vitamin E-deficient rabbits. Biochim. Biophys. Acta 584: Dayton, W. R., Schollmeyer, J. V., Lepley, R. A. and Cortes, L. R A calcium-activated protease possibly involved in myofibrillar protein turnover. Isolation of a low-calcium-requiring form of the protease. Biochim. Biophys. Acta 659: Dear, N., Matena, K., Vingron, M. and Boehm, T A new subfamily of vertebrate calpains lacking a calmodulin-like domain: implications for calpain regulation and evolution. Genomics 45: Elce, J. S., Hegadorn, C. and Arthur, J. S. C Autolysis, Ca 2+ requirement, and heterodimer stability in m-calpain. J. Biol. Chem. 272: Forsberg, N. E., Ilian, M. A., AlBar, A., Checke, P. R. and Wehr, N. B Effects of cimaterol on rabbit growth and myofibrillar protein degradation and on calcium-dependent proteinase and calpastatin activities in skeletal muscle. J. Anim. Sci. 67: Goll, D. E., Dayton, W. R., Singh, I. and Robson, R. M Studies of the α-actinin/actin interaction in the Z-disk by using calpain. J. Biol. Chem. 266:
9 GOLL ET AL. THE CALPAIN SYSTEM 511 Goll, D. E., Kleese, W. C. and Szpacenko, A Skeletal muscle proteases and protein turnover. Pages in D. R. Campion, G. J. Hausman, and R. J. Martin, eds. Animal growth regulation. Plenum Publishing Corp., New York, NY. Goll, D. E., Otsuka, Y., Nagainis, P. A., Shannon, J. D., Sathe, S. K. and Muguruma, M Role of muscle proteinases in maintenance of muscle integrity and mass. J. Food. Biochem. 7: Goll, D. E., Taylor, R. G., Christiansen, J. A. and Thompson, V. F. 1992a. Role of proteinases and protein turnover in muscle growth and meat quality. Proc. 44th Annual Reciprocal Meat Conf., National Live Stock and Meat Board, Chicago, IL. pp Goll, D. E., Thompson, V. F., Taylor, R. G. and Christiansen, J. A. 1992b. Role of the calpain system in muscle growth. Biochimie 74: Goll, D. E., Thompson, V. F., Taylor, R. G. and Zalewksa, T. 1992c. Is calpain activity regulated by membranes and autolysis or by calcium and calpastatin? BioEssays 14: Gopinath, R. and Kitts, W. D Growth, N-methylhistidine excretion and muscle protein degradation in growing beef steers. J. Anim. Sci. 59: Grobet, L., Martin, L. J. R., Poncelet, D., Pirottin, D., Brouwers, B., Riquet, S., Schoeberlin, A., Dunner, S., Menissier, F., Massabanda, J., Fries, R., Hanset, R. and Georges, M A deletion in the bovine myostatin gene causes the double-muscled phenotype in cattle. Nature Genetics 17: Higgins, J. A., Lasslet, Y., Bardsley, R. G. and Buttery, P. J The relation between dietary restriction or clenbuterol (a selective β2 agonist) treatment on skeletal muscle growth and calpain proteinase (EC ) and calpastatin activities in lambs. Br. J. Nutr. 60: Hopf, F. W., Turner, P. R., Denetclaw, W. F. Jr., Reddy, P. and Steinhardt, R. A A critical evaluation of resting intracellular free calcium regulation in dystrophic mdx muscle. Am. J. Physiol. 271: C1325 C1339. Ishiura, S., Nonaka, I. and Suita, H Ca 2+ -activated neutral protease: its degradative role in muscle cells. Pages in S. Ebashi, ed. Proc. Intern. Symp. Muscular Dystrophy, University of Tokyo Press, Tokyo, Japan. Jackson, S. P., Miller, M. F. and Green, R. D Phenotypic characterization of Rambouillet sheep expressing the callipyge gene: III. Muscle weights and muscle weight distribution. J. Anim. Sci. 75: Kambadur, R., Sharma, M., Smith, T. P. L. and Bass, J. J Mutations in myostastin (GDF8) in double-muscled Belgium Blue and Piedmontese cattle. Genomic Res. 7: Killefer, J. and Koohmaraie, M Bovine skeletal muscle calpastatin: cloning, sequence analysis and steady-state mrna expression. J. Anim. Sci. 72: Koohmaraie, M Ovine skeletal muscle multicatalytic proteinase complex (proteasome) purification, characterization, and comparison of its effects on myofibrils with µ-calpain. J. Anim. Sci. 70: Koohmaraie, M. and Shackelford, S. D Effect of calcium chloride infusion on the tenderness of lambs fed a β-adrenergic agonist. J. Anim. Sci. 69: Koohmaraie, M., Shackelford, S. D., Wheeler, T. L. Lonergan, S. M. and Doumit, M. E A muscle hypertrophy condition in lamb (callipyge): characterization of effects on muscle growth and meat quality traits. J. Anim Sci. 73: Kretchmar, D. H., Hathaway, M. R., Epley, R. J. and Dayton, W. R In vivo effect of a β-adrenergic agonist on activity of calcium-dependent proteinases, their specific inhibitor, and cathepsins B and H in skeletal muscle. Arch. Biochem. Biophys. 275: Kretchmar, D. H., Hathaway, M. R., Epley, R. J. and Dayton, W. R Alterations in postmortem degradation of myofibrillar proteins in muscle of lambs fed a β-adrenergic agonist. J. Anim. Sci. 69: Kumar, A., Shafiq, S., Wadgaonkar, R. and Stracher, A The effect of protease inhibitors, leupeptin and E64, on differentiation of C 2 C 12 myoblasts in tissue culture. Cell. Mol. Biol. 38: Kwak, K. B., Chung, S. S., Kim, O-M., Kang, M-S., Ha, D. B. and Chung, C. H. 1993a. Increase in the level of m-calpain correlates with the elevated cleavage of filamin during myogenic differentiation of embryonic muscle cells. Biochim. Biophys. Acta 1175: Kwak, K. B., Kambayashi, J-i., Kang, M. S., Ha, D. B. and Chung, C. H. 1993b. Cell-penetrating inhibitors of calpain block both membrane fusion and filamin cleavage in chick embryonic myoblasts. FEBS Lett. 323: Lee, W. J., Mu, H., Takano, E., Yang, H. Q., Hatanaka, M. and Maki, M Molecular diversity in amino-terminal domains of human calpastatin by exon skipping. J. Biol. Chem. 267: Lowell, B. B., Ruderman, N. B. and Goodman, M. N Evidence that lysosomes are not involved in the degradation of myofibrillar proteins in rat skeletal muscle. Biochem. J. 234: Maruyama, K., Sunde, M. L. and Swick, R. W Growth and muscle protein turnover in the chick. Biochem. J. 716: McPherron, A. C. and Lee, S-J Double muscling in cattle due to mutations in the myostatin gene. Proc. Natl. Acad. Sci. 94: McPherron, A. C., Lawler, A. M. and Lee, S-J Regulation of skeletal muscle mass in mice by a new TGF-β superfamily member. Nature 387: Mellgren, R. L Canine cardiac calcium-dependent proteases: resolution of two forms with different requirements for calcium. FEBS Lett. 109: Mellgren, R. L Evidence for participation of a calpain-like cysteine protease in cell cycle progression through the late G1 phase. Biochem. Biophys. Res. Comm 236: Mellgren, R. L., Shaw, E. and Mericle, M. T Inhibition of growth of human TEL and C-33A cells by the cell permeant inhibitor, benzyloxycarbonyl-leu-leu-tyr diazomethyl ketone. Exp. Cell Res. 215: Mongini, T., Ghigo, D., Doriguzzi, C., Bussolino, F., Pescarmona, G., Pollo, B., Schiffer, D. and Bosia, A Free cytoplasmic Ca 2+ at rest and after cholinergic stimulus is increased in cultured muscle cells from Duchenne muscular dystrophy patients. Neurology 38: Morkin, E Postnatal muscle fiber assembly: localization of newly synthesized myofibrillar proteins. Science 167: Murachi, T Intracellular regulatory system involving calpain and calpastatin. Biochem. Int. 18: Nave, R., Furst, D. O., and Weber, K Interaction of α- actinin and nebulin in vitro. FEBS Lett. 269: Ohtsuka, H., Yajima, H., Maruyama, K. and Kimura, S The N-terminal Z-repeat 5 of connectin/titin binds to the C-terminal end of α-actinin. Biochem. Biophys. Res. Commun. 235: 1 3. Okitani, A., Goll, D. E., Stromer, M. H. and Robson, R. M Intracellular inhibitor of a Ca 2+ -activated protease involved in myofibrillar protein turnover. Federation Proc. 35: Parr, T., Bardsley, R. G., Gilmour, R. S. and Buttery, P. J.
The Caspase System: a potential role in muscle proteolysis and meat quality? Tim Parr
 The Caspase System: a potential role in muscle proteolysis and meat quality? Tim Parr Caroline Kemp, Ron Bardsley,, Peter Buttery Division of Nutritional Sciences, School of Biosciences, University of
The Caspase System: a potential role in muscle proteolysis and meat quality? Tim Parr Caroline Kemp, Ron Bardsley,, Peter Buttery Division of Nutritional Sciences, School of Biosciences, University of
MUSCLE BIOLOGY OVERVIEW OF THE SKELETAL MUSCLE CYTOSKELETON
 MUSCLE BIOLOGY OVERVIEW OF THE SKELETAL MUSCLE CYTOSKELETON Ted Huiatt, Ph.D. Associate Professor, Department of Animal Science and Roy J. Carver Department of Biochemistry, Biophysics and Molecular Biology,
MUSCLE BIOLOGY OVERVIEW OF THE SKELETAL MUSCLE CYTOSKELETON Ted Huiatt, Ph.D. Associate Professor, Department of Animal Science and Roy J. Carver Department of Biochemistry, Biophysics and Molecular Biology,
Lecture 13, 05 October 2004 Chapter 10, Muscle. Vertebrate Physiology ECOL 437 University of Arizona Fall instr: Kevin Bonine t.a.
 Lecture 13, 05 October 2004 Chapter 10, Muscle Vertebrate Physiology ECOL 437 University of Arizona Fall 2004 instr: Kevin Bonine t.a.: Nate Swenson Vertebrate Physiology 437 18 1. Muscle A. Sarcomere
Lecture 13, 05 October 2004 Chapter 10, Muscle Vertebrate Physiology ECOL 437 University of Arizona Fall 2004 instr: Kevin Bonine t.a.: Nate Swenson Vertebrate Physiology 437 18 1. Muscle A. Sarcomere
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Growth and Muscle Production
 Growth and Muscle Production The goal of these lectures is to discuss basic physiology associated with the control of growth and muscle production. 32 The sections for this lecture are: Animal growth and
Growth and Muscle Production The goal of these lectures is to discuss basic physiology associated with the control of growth and muscle production. 32 The sections for this lecture are: Animal growth and
PHYSIOLOGY CHAPTER 9 MUSCLE TISSUE Fall 2016
 PHYSIOLOGY CHAPTER 9 MUSCLE TISSUE Fall 2016 2 Chapter 9 Muscles and Muscle Tissue Overview of Muscle Tissue types of muscle: are all prefixes for muscle Contractility all muscles cells can Smooth & skeletal
PHYSIOLOGY CHAPTER 9 MUSCLE TISSUE Fall 2016 2 Chapter 9 Muscles and Muscle Tissue Overview of Muscle Tissue types of muscle: are all prefixes for muscle Contractility all muscles cells can Smooth & skeletal
Chapter 16. Cellular Movement: Motility and Contractility. Lectures by Kathleen Fitzpatrick Simon Fraser University Pearson Education, Inc.
 Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
Chapter 16 Cellular Movement: Motility and Contractility Lectures by Kathleen Fitzpatrick Simon Fraser University Two eukaryotic motility systems 1. Interactions between motor proteins and microtubules
According to the diagram, which of the following is NOT true?
 Instructions: Review Chapter 44 on muscular-skeletal systems and locomotion, and then complete the following Blackboard activity. This activity will introduce topics that will be covered in the next few
Instructions: Review Chapter 44 on muscular-skeletal systems and locomotion, and then complete the following Blackboard activity. This activity will introduce topics that will be covered in the next few
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement
 1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
1 Muscle regulation and Actin Topics: Tropomyosin and Troponin, Actin Assembly, Actin-dependent Movement In the last lecture, we saw that a repeating alternation between chemical (ATP hydrolysis) and vectorial
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation of gene expression. Premedical - Biology
 Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
An Introduction to Metabolism
 An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
3.a.2- Cell Cycle and Meiosis
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. 3.a.2- Cell Cycle and Meiosis EU 3.A: Heritable information provides for continuity of life.
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. 3.a.2- Cell Cycle and Meiosis EU 3.A: Heritable information provides for continuity of life.
CC2.04_t.txt The Caspase System and Muscle Degradation Tim Parr
 The Caspase System and Muscle Degradation Tim Parr [1]Our last speaker of the day is Dr. Tim Parr. Dr. Parr graduated in 1987 from the University of Southampton with a degree in physiology, biochemistry,
The Caspase System and Muscle Degradation Tim Parr [1]Our last speaker of the day is Dr. Tim Parr. Dr. Parr graduated in 1987 from the University of Southampton with a degree in physiology, biochemistry,
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
The diagram below represents levels of organization within a cell of a multicellular organism.
 STATION 1 1. Unlike prokaryotic cells, eukaryotic cells have the capacity to a. assemble into multicellular organisms b. establish symbiotic relationships with other organisms c. obtain energy from the
STATION 1 1. Unlike prokaryotic cells, eukaryotic cells have the capacity to a. assemble into multicellular organisms b. establish symbiotic relationships with other organisms c. obtain energy from the
Name # Class Date Regents Review: Cells & Cell Transport
 Name # Class Date Regents Review: Cells & Cell Transport 1. All of the following are true regarding cells except? A) All cells have genetic material B) All cells have cell walls C) All cells have plasma
Name # Class Date Regents Review: Cells & Cell Transport 1. All of the following are true regarding cells except? A) All cells have genetic material B) All cells have cell walls C) All cells have plasma
Which row in the chart correctly identifies the functions of structures A, B, and C? A) 1 B) 2 C) 3 D) 4
 1. What is a similarity between all bacteria and plants? A) They both have a nucleus B) They are both composed of cells C) They both have chloroplasts D) They both lack a cell wall 2. Which statement is
1. What is a similarity between all bacteria and plants? A) They both have a nucleus B) They are both composed of cells C) They both have chloroplasts D) They both lack a cell wall 2. Which statement is
Richik N. Ghosh, Linnette Grove, and Oleg Lapets ASSAY and Drug Development Technologies 2004, 2:
 1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Our patient for the day...
 Muscles Ch.12 Our patient for the day... Name: Eddy Age: Newborn Whole-body muscle contractions No relaxation Severe difficulty breathing due to inadequate relaxation of breathing muscles Diagnosed with
Muscles Ch.12 Our patient for the day... Name: Eddy Age: Newborn Whole-body muscle contractions No relaxation Severe difficulty breathing due to inadequate relaxation of breathing muscles Diagnosed with
Fundamentals of Neurosciences. Smooth Muscle. Dr. Kumar Sambamurti 613-SEI; ;
 Fundamentals of Neurosciences Smooth Muscle Dr. Kumar Sambamurti 613-SEI; 792-4315; sambak@musc.edu 1 Smooth Muscle Structure Cells much smaller than skeletal muscle (2-5µM diam, 100-400µM long) Single
Fundamentals of Neurosciences Smooth Muscle Dr. Kumar Sambamurti 613-SEI; 792-4315; sambak@musc.edu 1 Smooth Muscle Structure Cells much smaller than skeletal muscle (2-5µM diam, 100-400µM long) Single
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Modeling. EC-Coupling and Contraction
 Bioeng 6460 Electrophysiology and Bioelectricity Modeling of EC-Coupling and Contraction Frank B. Sachse fs@cvrti.utah.edu Overview Quiz Excitation-Contraction Coupling Anatomy Cross Bridge Binding Coupling
Bioeng 6460 Electrophysiology and Bioelectricity Modeling of EC-Coupling and Contraction Frank B. Sachse fs@cvrti.utah.edu Overview Quiz Excitation-Contraction Coupling Anatomy Cross Bridge Binding Coupling
UNIT 6 THE MUSCULAR SYSTEM
 UNIT 6 THE MUSCULAR SYSTEM I. Functions of Muscular System A. Produces Movement Internal vs. External «locomotion & manipulation «circulate blood & maintain blood pressure «move fluids, food, baby B. Maintaining
UNIT 6 THE MUSCULAR SYSTEM I. Functions of Muscular System A. Produces Movement Internal vs. External «locomotion & manipulation «circulate blood & maintain blood pressure «move fluids, food, baby B. Maintaining
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells
 Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
Life Sciences 1a: Section 3B. The cell division cycle Objectives Understand the challenges to producing genetically identical daughter cells Understand how a simple biochemical oscillator can drive the
9/25/2011. Outline. Overview: The Energy of Life. I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V.
 Chapter 8 Introduction to Metabolism Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes Overview: The Energy of Life Figure 8.1 The living cell is a miniature
Chapter 8 Introduction to Metabolism Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes Overview: The Energy of Life Figure 8.1 The living cell is a miniature
Study Guide 11 & 12 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 Study Guide 11 & 12 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) The receptors for a group of signaling molecules known as growth factors are
Study Guide 11 & 12 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) The receptors for a group of signaling molecules known as growth factors are
Metabolism: Energy and Enzymes. February 24 th, 2012
 Metabolism: Energy and Enzymes February 24 th, 2012 1 Outline Forms of Energy Laws of Thermodynamics Metabolic Reactions ATP Metabolic Pathways Energy of Activation Enzymes Photosynthesis Cellular Respiration
Metabolism: Energy and Enzymes February 24 th, 2012 1 Outline Forms of Energy Laws of Thermodynamics Metabolic Reactions ATP Metabolic Pathways Energy of Activation Enzymes Photosynthesis Cellular Respiration
Enzyme Enzymes are proteins that act as biological catalysts. Enzymes accelerate, or catalyze, chemical reactions. The molecules at the beginning of
 Enzyme Enzyme Enzymes are proteins that act as biological catalysts. Enzymes accelerate, or catalyze, chemical reactions. The molecules at the beginning of the process are called substrates and the enzyme
Enzyme Enzyme Enzymes are proteins that act as biological catalysts. Enzymes accelerate, or catalyze, chemical reactions. The molecules at the beginning of the process are called substrates and the enzyme
Muscle tissue. Types. Functions. Cardiac, Smooth, and Skeletal
 Types Cardiac, Smooth, and Skeletal Functions movements posture and body position Support soft tissues Guard openings body temperature nutrient reserves Muscle tissue Special Characteristics of Muscle
Types Cardiac, Smooth, and Skeletal Functions movements posture and body position Support soft tissues Guard openings body temperature nutrient reserves Muscle tissue Special Characteristics of Muscle
Chapter 2 Concepts of Chemistry
 Anatomy Physiology and Disease for the Health Professions 3rd Edition Booth Test Bank Full Download: http://testbanklive.com/download/anatomy-physiology-and-disease-for-the-health-professions-3rd-edition-booth-te
Anatomy Physiology and Disease for the Health Professions 3rd Edition Booth Test Bank Full Download: http://testbanklive.com/download/anatomy-physiology-and-disease-for-the-health-professions-3rd-edition-booth-te
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
BIOMECHANICS 3 Origins and consequences of forces in biological systems
 BIOMECHANICS 3 Origins and consequences of forces in biological systems MOLECULAR MECHANISMS OF BIOLOGICAL MOVEMENT AT THE LEVELOF ORGANISMS MOLECULAR BASIS OF MUSCLE CONTRACTION DR. BEÁTA BUGYI - BIOPHYSICS
BIOMECHANICS 3 Origins and consequences of forces in biological systems MOLECULAR MECHANISMS OF BIOLOGICAL MOVEMENT AT THE LEVELOF ORGANISMS MOLECULAR BASIS OF MUSCLE CONTRACTION DR. BEÁTA BUGYI - BIOPHYSICS
BIOLOGY 10/11/2014. An Introduction to Metabolism. Outline. Overview: The Energy of Life
 8 An Introduction to Metabolism CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes
8 An Introduction to Metabolism CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson Outline I. Forms of Energy II. Laws of Thermodynamics III. Energy and metabolism IV. ATP V. Enzymes
An Introduction to Metabolism
 CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 6 An Introduction to Metabolism Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: The Energy of Life The
CAMPBELL BIOLOGY IN FOCUS Urry Cain Wasserman Minorsky Jackson Reece 6 An Introduction to Metabolism Lecture Presentations by Kathleen Fitzpatrick and Nicole Tunbridge Overview: The Energy of Life The
CELL CYCLE AND DIFFERENTIATION
 CELL CYCLE AND DIFFERENTIATION Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id WHAT IS CELL CYCLE? 09/12/14 d.purnomosari@ugm.ac.id
CELL CYCLE AND DIFFERENTIATION Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id WHAT IS CELL CYCLE? 09/12/14 d.purnomosari@ugm.ac.id
Metabolism and enzymes
 Metabolism and enzymes 4-11-16 What is a chemical reaction? A chemical reaction is a process that forms or breaks the chemical bonds that hold atoms together Chemical reactions convert one set of chemical
Metabolism and enzymes 4-11-16 What is a chemical reaction? A chemical reaction is a process that forms or breaks the chemical bonds that hold atoms together Chemical reactions convert one set of chemical
Modelling Muscle Contraction a multiscale approach
 Porto Ercole, M&MKT 2016 Multiscale Systems from Particles to Continuum: Modelling and Computation Modelling Muscle Contraction a multiscale approach Giovanni Naldi Dipartimento di Matematica ``F. Enriques
Porto Ercole, M&MKT 2016 Multiscale Systems from Particles to Continuum: Modelling and Computation Modelling Muscle Contraction a multiscale approach Giovanni Naldi Dipartimento di Matematica ``F. Enriques
Biology: Life on Earth
 Biology: Life on Earth Eighth Edition Lecture for Chapter 11 The Continuity of Life: Cellular Reproduction Cellular Reproduction Intracellular activity between one cell division to the next is the cell
Biology: Life on Earth Eighth Edition Lecture for Chapter 11 The Continuity of Life: Cellular Reproduction Cellular Reproduction Intracellular activity between one cell division to the next is the cell
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Lipniacki 2004 Ground Truth
 Abstract Lipniacki 2004 Ground Truth The two-feedback-loop regulatory module of nuclear factor kb (NF-kB) signaling pathway is modeled by means of ordinary differential equations. signaling pathway: https://en.wikipedia.org/wiki/signaling_pathway
Abstract Lipniacki 2004 Ground Truth The two-feedback-loop regulatory module of nuclear factor kb (NF-kB) signaling pathway is modeled by means of ordinary differential equations. signaling pathway: https://en.wikipedia.org/wiki/signaling_pathway
The Caspase System: a Potential Role in Muscle Proteolysis and Meat Quality?
 CURRENT TOPICS IN MUSCLE GROWTH & DEVELOPMENT The Caspase System: a Potential Role in Muscle Proteolysis and Meat Quality? Tim Parr, Ron Bardsley, Peter Buttery & Caroline Kemp Introduction Meat quality
CURRENT TOPICS IN MUSCLE GROWTH & DEVELOPMENT The Caspase System: a Potential Role in Muscle Proteolysis and Meat Quality? Tim Parr, Ron Bardsley, Peter Buttery & Caroline Kemp Introduction Meat quality
Cell Biology Review. The key components of cells that concern us are as follows: 1. Nucleus
 Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Base your answers to questions 1 and 2 on the diagram below which represents a typical green plant cell and on your knowledge of biology.
 Base your answers to questions 1 and 2 on the diagram below which represents a typical green plant cell and on your knowledge of biology. 5. Which letter corresponds to that of the endoplasmic reticulum?
Base your answers to questions 1 and 2 on the diagram below which represents a typical green plant cell and on your knowledge of biology. 5. Which letter corresponds to that of the endoplasmic reticulum?
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11
 UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
UNIT 6 PART 3 *REGULATION USING OPERONS* Hillis Textbook, CH 11 REVIEW: Signals that Start and Stop Transcription and Translation BUT, HOW DO CELLS CONTROL WHICH GENES ARE EXPRESSED AND WHEN? First of
Gene Control Mechanisms at Transcription and Translation Levels
 Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Metabolism and Enzymes
 Energy Basics Metabolism and Enzymes Chapter 5 Pgs. 77 86 Chapter 8 Pgs. 142 162 Energy is the capacity to cause change, and is required to do work. Very difficult to define quantity. Two types of energy:
Energy Basics Metabolism and Enzymes Chapter 5 Pgs. 77 86 Chapter 8 Pgs. 142 162 Energy is the capacity to cause change, and is required to do work. Very difficult to define quantity. Two types of energy:
BIOH111. o Cell Biology Module o Tissue Module o Integumentary system o Skeletal system o Muscle system o Nervous system o Endocrine system
 BIOH111 o Cell Biology Module o Tissue Module o Integumentary system o Skeletal system o Muscle system o Nervous system o Endocrine system Endeavour College of Natural Health endeavour.edu.au 1 Textbook
BIOH111 o Cell Biology Module o Tissue Module o Integumentary system o Skeletal system o Muscle system o Nervous system o Endocrine system Endeavour College of Natural Health endeavour.edu.au 1 Textbook
S1 Gene ontology (GO) analysis of the network alignment results
 1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
1 Supplementary Material for Effective comparative analysis of protein-protein interaction networks by measuring the steady-state network flow using a Markov model Hyundoo Jeong 1, Xiaoning Qian 1 and
Name: Date: Answer: Answer:
 Name: Date: 5 6 7 8 9 0 Scoring Guide: Scoring Guide: 5 6 7 8 9 0 5 6 7 8 9 0 Scoring Guide: Scoring Guide: 5 Scoring Guide: 6 7 8 9 0 5 6 7 8 9 50 Scoring Guide: 5 Scoring Guide: Standard(s):..0.F,...F,..0.D,...D,..0.C,...C,..0.E,...E,.5.0.F,.5..F
Name: Date: 5 6 7 8 9 0 Scoring Guide: Scoring Guide: 5 6 7 8 9 0 5 6 7 8 9 0 Scoring Guide: Scoring Guide: 5 Scoring Guide: 6 7 8 9 0 5 6 7 8 9 50 Scoring Guide: 5 Scoring Guide: Standard(s):..0.F,...F,..0.D,...D,..0.C,...C,..0.E,...E,.5.0.F,.5..F
Biology Kevin Dees. Chapter 8 Introduction to Metabolism
 Chapter 8 Introduction to Metabolism Defined as the sum total of the chemical reactions that occur in a living thing. Think of metabolism as a road map of thousands of different chemical reactions regulate
Chapter 8 Introduction to Metabolism Defined as the sum total of the chemical reactions that occur in a living thing. Think of metabolism as a road map of thousands of different chemical reactions regulate
CELL PRACTICE TEST
 Name: Date: 1. As a human red blood cell matures, it loses its nucleus. As a result of this loss, a mature red blood cell lacks the ability to (1) take in material from the blood (2) release hormones to
Name: Date: 1. As a human red blood cell matures, it loses its nucleus. As a result of this loss, a mature red blood cell lacks the ability to (1) take in material from the blood (2) release hormones to
Biochemical and Structural Properties of Titin, Nebulin and Intermediate Filaments in Muscle
 MUSCLE BIOCHEMISTRY Biochemical and Structural Properties of Titin, Nebulin and Intermediate Filaments in Muscle Richard M. Robson* Ted W. Huiatt Frederick C. Parrish, Jr. Introduction In a generic vertebrate
MUSCLE BIOCHEMISTRY Biochemical and Structural Properties of Titin, Nebulin and Intermediate Filaments in Muscle Richard M. Robson* Ted W. Huiatt Frederick C. Parrish, Jr. Introduction In a generic vertebrate
Analyze the roles of enzymes in biochemical reactions
 ENZYMES and METABOLISM Elements: Cell Biology (Enzymes) Estimated Time: 6 7 hours By the end of this course, students will have an understanding of the role of enzymes in biochemical reactions. Vocabulary
ENZYMES and METABOLISM Elements: Cell Biology (Enzymes) Estimated Time: 6 7 hours By the end of this course, students will have an understanding of the role of enzymes in biochemical reactions. Vocabulary
Chapter Cells and the Flow of Energy A. Forms of Energy 1. Energy is capacity to do work; cells continually use energy to develop, grow,
 Chapter 6 6.1 Cells and the Flow of Energy A. Forms of Energy 1. Energy is capacity to do work; cells continually use energy to develop, grow, repair, reproduce, etc. 2. Kinetic energy is energy of motion;
Chapter 6 6.1 Cells and the Flow of Energy A. Forms of Energy 1. Energy is capacity to do work; cells continually use energy to develop, grow, repair, reproduce, etc. 2. Kinetic energy is energy of motion;
Energy Transformation and Metabolism (Outline)
 Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Cells to Tissues. Peter Takizawa Department of Cell Biology
 Cells to Tissues Peter Takizawa Department of Cell Biology From one cell to ensembles of cells. Multicellular organisms require individual cells to work together in functional groups. This means cells
Cells to Tissues Peter Takizawa Department of Cell Biology From one cell to ensembles of cells. Multicellular organisms require individual cells to work together in functional groups. This means cells
Reading Assignments. A. Genes and the Synthesis of Polypeptides. Lecture Series 7 From DNA to Protein: Genotype to Phenotype
 Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Lecture Series 7 From DNA to Protein: Genotype to Phenotype Reading Assignments Read Chapter 7 From DNA to Protein A. Genes and the Synthesis of Polypeptides Genes are made up of DNA and are expressed
Problem Set # 3
 20.320 Problem Set # 3 October 1 st, 2010 Due on October 8 th, 2010 at 11:59am. No extensions, no electronic submissions. General Instructions: 1. You are expected to state all your assumptions and provide
20.320 Problem Set # 3 October 1 st, 2010 Due on October 8 th, 2010 at 11:59am. No extensions, no electronic submissions. General Instructions: 1. You are expected to state all your assumptions and provide
Reception The target cell s detection of a signal coming from outside the cell May Occur by: Direct connect Through signal molecules
 Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
Why Do Cells Communicate? Regulation Cells need to control cellular processes In multicellular organism, cells signaling pathways coordinate the activities within individual cells that support the function
C a h p a t p e t r e r 6 E z n y z m y e m s
 Chapter 6 Enzymes 4. Examples of enzymatic reactions acid-base catalysis: give and take protons covalent catalysis: a transient covalent bond is formed between the enzyme and the substrate metal ion catalysis:
Chapter 6 Enzymes 4. Examples of enzymatic reactions acid-base catalysis: give and take protons covalent catalysis: a transient covalent bond is formed between the enzyme and the substrate metal ion catalysis:
(Be sure to clearly state the principles addressed in your discussion.)
 CELL QUESTION 1992: AP BIOLOGY A laboratory assistant prepared solutions of 0.8 M, 0.6 M, 0.4 M, and 0.2 M sucrose, but forgot to label them. After realizing the error, the assistant randomly labeled the
CELL QUESTION 1992: AP BIOLOGY A laboratory assistant prepared solutions of 0.8 M, 0.6 M, 0.4 M, and 0.2 M sucrose, but forgot to label them. After realizing the error, the assistant randomly labeled the
2 The Proteome. The Proteome 15
 The Proteome 15 2 The Proteome 2.1. The Proteome and the Genome Each of our cells contains all the information necessary to make a complete human being. However, not all the genes are expressed in all
The Proteome 15 2 The Proteome 2.1. The Proteome and the Genome Each of our cells contains all the information necessary to make a complete human being. However, not all the genes are expressed in all
Chem Lecture 10 Signal Transduction
 Chem 452 - Lecture 10 Signal Transduction 111202 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Chem 452 - Lecture 10 Signal Transduction 111202 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
4. Which of the following organelles digests waste using hydrolytic enzymes:
 Multichoice questions section. You must answer ALL questions. 1. A cell contains many organelles, each of which has a specific function. What is function of mitochondria? a) production of plasma membrane
Multichoice questions section. You must answer ALL questions. 1. A cell contains many organelles, each of which has a specific function. What is function of mitochondria? a) production of plasma membrane
Chapter 8 Introduction to Metabolism. Metabolism. The sum total of the chemical reactions that occur in a living thing.
 Chapter 8 Introduction to Metabolism Metabolism The sum total of the chemical reactions that occur in a living thing. Think of metabolism as a road map of thousands of different chemical reactions Enzymes
Chapter 8 Introduction to Metabolism Metabolism The sum total of the chemical reactions that occur in a living thing. Think of metabolism as a road map of thousands of different chemical reactions Enzymes
Chapter Chemical Uniqueness 1/23/2009. The Uses of Principles. Zoology: the Study of Animal Life. Fig. 1.1
 Fig. 1.1 Chapter 1 Life: Biological Principles and the Science of Zoology BIO 2402 General Zoology Copyright The McGraw Hill Companies, Inc. Permission required for reproduction or display. The Uses of
Fig. 1.1 Chapter 1 Life: Biological Principles and the Science of Zoology BIO 2402 General Zoology Copyright The McGraw Hill Companies, Inc. Permission required for reproduction or display. The Uses of
Ch 4: Cellular Metabolism, Part 1
 Developed by John Gallagher, MS, DVM Ch 4: Cellular Metabolism, Part 1 Energy as it relates to Biology Energy for synthesis and movement Energy transformation Enzymes and how they speed reactions Metabolism
Developed by John Gallagher, MS, DVM Ch 4: Cellular Metabolism, Part 1 Energy as it relates to Biology Energy for synthesis and movement Energy transformation Enzymes and how they speed reactions Metabolism
CHAPTER 3. Cell Structure and Genetic Control. Chapter 3 Outline
 CHAPTER 3 Cell Structure and Genetic Control Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell Nucleus and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division
CHAPTER 3 Cell Structure and Genetic Control Chapter 3 Outline Plasma Membrane Cytoplasm and Its Organelles Cell Nucleus and Gene Expression Protein Synthesis and Secretion DNA Synthesis and Cell Division
Solution Authoring Guidelines Version 9.4 September 2016
 Solution Authoring Guidelines Version 9.4 September 2016 Subject-specific Guidelines- Biology Table of Contents B1. Technology... 3 B2. Special points/others... 3 List of changes made over Version 9.1
Solution Authoring Guidelines Version 9.4 September 2016 Subject-specific Guidelines- Biology Table of Contents B1. Technology... 3 B2. Special points/others... 3 List of changes made over Version 9.1
BIOLOGY. An Introduction to Metabolism CAMPBELL. Reece Urry Cain Wasserman Minorsky Jackson
 CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 8 An Introduction to Metabolism Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick The Energy of Life The living
CAMPBELL BIOLOGY TENTH EDITION Reece Urry Cain Wasserman Minorsky Jackson 8 An Introduction to Metabolism Lecture Presentation by Nicole Tunbridge and Kathleen Fitzpatrick The Energy of Life The living
An Introduction to Metabolism
 Chapter 8 An Introduction to Metabolism PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
Chapter 8 An Introduction to Metabolism PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley with contributions from
An Introduction to Metabolism
 An Introduction to Metabolism The living cell is a microscopic factory where life s giant processes can be performed: -sugars to amino acids to proteins and vise versa -reactions to dismantle polymers
An Introduction to Metabolism The living cell is a microscopic factory where life s giant processes can be performed: -sugars to amino acids to proteins and vise versa -reactions to dismantle polymers
Molecular Cell Biology 5068 In Class Exam 2 November 8, 2016
 Molecular Cell Biology 5068 In Class Exam 2 November 8, 2016 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
Molecular Cell Biology 5068 In Class Exam 2 November 8, 2016 Exam Number: Please print your name: Instructions: Please write only on these pages, in the spaces allotted and not on the back. Write your
An Introduction to Metabolism
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 8 An Introduction to Metabolism
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 8 An Introduction to Metabolism
Ch. 3 Metabolism and Enzymes
 Ch. 3 Metabolism and Enzymes Originally prepared by Kim B. Foglia. Revised and adapted by Nhan A. Pham Flow of energy through life Life is built on chemical reactions that enable energy to flow through
Ch. 3 Metabolism and Enzymes Originally prepared by Kim B. Foglia. Revised and adapted by Nhan A. Pham Flow of energy through life Life is built on chemical reactions that enable energy to flow through
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
ADAM FAMILY. ephrin A INTERAZIONE. Eph ADESIONE? PROTEOLISI ENDOCITOSI B A RISULTATO REPULSIONE. reverse. forward
 ADAM FAMILY - a family of membrane-anchored metalloproteases that are known as A Disintegrin And Metalloprotease proteins and are key components in protein ectodomain shedding Eph A INTERAZIONE B ephrin
ADAM FAMILY - a family of membrane-anchored metalloproteases that are known as A Disintegrin And Metalloprotease proteins and are key components in protein ectodomain shedding Eph A INTERAZIONE B ephrin
Chapter 5. Energy Flow in the Life of a Cell
 Chapter 5 Energy Flow in the Life of a Cell Including some materials from lectures by Gregory Ahearn University of North Florida Ammended by John Crocker Copyright 2009 Pearson Education, Inc.. Review
Chapter 5 Energy Flow in the Life of a Cell Including some materials from lectures by Gregory Ahearn University of North Florida Ammended by John Crocker Copyright 2009 Pearson Education, Inc.. Review
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
WILLIAM R. DAYTON, D. E. GOLL AND W. Iowa State University INTRODUCTION
 2 14 CONTROL OF MUSCL;E PROTEN DEGRADATON AND TS ROLE N ACCUMULATON OF MUSCLE MASS* WLLAM R DAYTON, D E GOLL AND W owa State University J REVLLF: NTRODUCTON t has been known for over 30 years that intracellular
2 14 CONTROL OF MUSCL;E PROTEN DEGRADATON AND TS ROLE N ACCUMULATON OF MUSCLE MASS* WLLAM R DAYTON, D E GOLL AND W owa State University J REVLLF: NTRODUCTON t has been known for over 30 years that intracellular
An Introduction to Metabolism
 Chapter 8 An Introduction to Metabolism Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 8 An Introduction to Metabolism Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Cell Theory. Cell Structure. Chapter 4. Cell is basic unit of life. Cells discovered in 1665 by Robert Hooke
 Cell Structure Chapter 4 Cell is basic unit of life Cell Theory Cells discovered in 1665 by Robert Hooke Early cell studies conducted by - Mathias Schleiden (1838) - Theodor Schwann (1839) Schleiden &
Cell Structure Chapter 4 Cell is basic unit of life Cell Theory Cells discovered in 1665 by Robert Hooke Early cell studies conducted by - Mathias Schleiden (1838) - Theodor Schwann (1839) Schleiden &
15.2 Prokaryotic Transcription *
 OpenStax-CNX module: m52697 1 15.2 Prokaryotic Transcription * Shannon McDermott Based on Prokaryotic Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
OpenStax-CNX module: m52697 1 15.2 Prokaryotic Transcription * Shannon McDermott Based on Prokaryotic Transcription by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative Commons
The EGF Signaling Pathway! Introduction! Introduction! Chem Lecture 10 Signal Transduction & Sensory Systems Part 3. EGF promotes cell growth
 Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 3 Question of the Day: Who is the son of Sevenless? Introduction! Signal transduction involves the changing of a cell s metabolism or gene
Chem 452 - Lecture 10 Signal Transduction & Sensory Systems Part 3 Question of the Day: Who is the son of Sevenless? Introduction! Signal transduction involves the changing of a cell s metabolism or gene
Chapter 2 Cells and Cell Division
 Chapter 2 Cells and Cell Division MULTIPLE CHOICE 1. The process of meiosis results in. A. the production of four identical cells B. no change in chromosome number from parental cells C. a doubling of
Chapter 2 Cells and Cell Division MULTIPLE CHOICE 1. The process of meiosis results in. A. the production of four identical cells B. no change in chromosome number from parental cells C. a doubling of
RANK. Alternative names. Discovery. Structure. William J. Boyle* SUMMARY BACKGROUND
 RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
RANK William J. Boyle* Department of Cell Biology, Amgen, Inc., One Amgen Center Drive, Thousand Oaks, CA 91320-1799, USA * corresponding author tel: 805-447-4304, fax: 805-447-1982, e-mail: bboyle@amgen.com
Chapter 15 Active Reading Guide Regulation of Gene Expression
 Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Name: AP Biology Mr. Croft Chapter 15 Active Reading Guide Regulation of Gene Expression The overview for Chapter 15 introduces the idea that while all cells of an organism have all genes in the genome,
Warm Up. What are some examples of living things? Describe the characteristics of living things
 Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
Warm Up What are some examples of living things? Describe the characteristics of living things Objectives Identify the levels of biological organization and explain their relationships Describe cell structure
An Introduction to Metabolism
 An Introduction to Metabolism Chapter 8 Objectives Distinguish between the following pairs of terms: catabolic and anabolic pathways; kinetic and potential energy; open and closed systems; exergonic and
An Introduction to Metabolism Chapter 8 Objectives Distinguish between the following pairs of terms: catabolic and anabolic pathways; kinetic and potential energy; open and closed systems; exergonic and
Ribosome readthrough
 Ribosome readthrough Starting from the base PROTEIN SYNTHESIS Eukaryotic translation can be divided into four stages: Initiation, Elongation, Termination and Recycling During translation, the ribosome
Ribosome readthrough Starting from the base PROTEIN SYNTHESIS Eukaryotic translation can be divided into four stages: Initiation, Elongation, Termination and Recycling During translation, the ribosome
An Introduction to Metabolism
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 8 An Introduction to Metabolism
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 8 An Introduction to Metabolism
Energy and Cellular Metabolism
 1 Chapter 4 About This Chapter Energy and Cellular Metabolism 2 Energy in biological systems Chemical reactions Enzymes Metabolism Figure 4.1 Energy transfer in the environment Table 4.1 Properties of
1 Chapter 4 About This Chapter Energy and Cellular Metabolism 2 Energy in biological systems Chemical reactions Enzymes Metabolism Figure 4.1 Energy transfer in the environment Table 4.1 Properties of
TISSUE. A) Types. (i)
 MUSCLES & MUSCLE TISSUE I. OVERVIEW - Muscle ( little mouse ) - tissue designed to cause movementt thru contraction ( shortening ). A) Types - There are some SIMILARITIES between muscle types: (i) All
MUSCLES & MUSCLE TISSUE I. OVERVIEW - Muscle ( little mouse ) - tissue designed to cause movementt thru contraction ( shortening ). A) Types - There are some SIMILARITIES between muscle types: (i) All
Name Period The Control of Gene Expression in Prokaryotes Notes
 Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
Bacterial DNA contains genes that encode for many different proteins (enzymes) so that many processes have the ability to occur -not all processes are carried out at any one time -what allows expression
