BIOINFORMATICS Structures. Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.
|
|
|
- Bertha Crawford
- 5 years ago
- Views:
Transcription
1 BIOINFORMATICS Structures Mark Gerstein, Yale University bioinfo.mbb.yale.edu/mbb452a 1 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
2 Contents: Structures What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 2 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
3 Molecular Biology Information: Macromolecular Structure DNA/RNA/Protein Almost all protein (RNA Adapted From D Soll Web Page, Right Hand Top Protein from M Levitt web page) 3 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
4 Molecular Biology Information: Protein Structure Details Statistics on Number of XYZ triplets 200 residues/domain -> 200 CA atoms, separated by 3.8 A Avg. Residue is Leu: 4 backbone atoms + 4 sidechain atoms, 150 cubic A o => ~1500 xyz triplets (=8x200) per protein domain 10 K known domain, ~300 folds ATOM 1 C ACE GKY 67 ATOM 2 O ACE GKY 68 ATOM 3 CH3 ACE GKY 69 ATOM 4 N SER GKY 70 ATOM 5 CA SER GKY 71 ATOM 6 C SER GKY 72 ATOM 7 O SER GKY 73 ATOM 8 CB SER GKY 74 ATOM 9 OG SER GKY 75 ATOM 10 N ARG GKY 76 ATOM 11 CA ARG GKY 77 ATOM 12 C ARG GKY ATOM 1444 CB LYS GKY1510 ATOM 1445 CG LYS GKY1511 ATOM 1446 CD LYS GKY1512 ATOM 1447 CE LYS GKY1513 ATOM 1448 NZ LYS GKY1514 ATOM 1449 OXT LYS GKY1515 TER 1450 LYS 186 1GKY (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
5 Other Aspects of Structure, Besides just Comparing Atom Positions Atom Position, XYZ triplets Lines, Axes, Angles Surfaces, Volumes 5 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
6 What is Protein Geometry? Coordinates (X, Y, Z s) Derivative Concepts Distance, Surface Area, Volume, Cavity, Groove, Axes, Angle, &c Relationto Function, Energies (E(x)), Dynamics (dx/dt) 6 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
7 Depicting Protein Structure: Sperm Whale Myoglobin 7 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
8 8 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Incredulase
9 Structure alignment - Method What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surface I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 9 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
10 10 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu Sperm Whale Myoglobin
11 Structural Alignment of Two Globins 11 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
12 Hb Automatic Alignment to Build Fold Library Alignment of Individual Structures Fusing into a Single Fold Template Mb Hb VLSPADKTNVKAAWGKVGAHAGEYGAEALERMFLSFPTTKTYFPHF-DLS-----HGSAQVKGHGKKVADALTNAV Mb VLSEGEWQLVLHVWAKVEADVAGHGQDILIRLFKSHPETLEKFDRFKHLKTEAEMKASEDLKKHGVTVLTALGAIL Hb AHVD-DMPNALSALSDLHAHKLRVDPVNFKLLSHCLLVTLAAHLPAEFTPAVHASLDKFLASVSTVLTSKYR Mb KK-KGHHEAELKPLAQSHATKHKIPIKYLEFISEAIIHVLHSRHPGDFGADAQGAMNKALELFRKDIAAKYKELGYQG Elements: Domain definitions; Aligned structures, collecting together Non-homologous Sequences; Core annotation Previous work: Remington, Matthews 80; Taylor, Orengo 89, 94; Artymiuk, Rice, Willett 89; Sali, Blundell, 90; Vriend, Sander 91; Russell, Barton 92; Holm, Sander 93; Godzik, Skolnick 94; Gibrat, Madej, Bryant 96; Falicov, F Cohen, 96; Feng, Sippl 96; G Cohen 97; Singh & Brutlag, 98 Structure Sequence Core Core 12 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu 2hhb HA HU - D P N A L S A L S D L H A H K L - F - - R V D P V N K L L S H C L L V T L A A H < HADG - D LP GA L S A L S D L H A YKL - F - - R V D P V NKL L S H C L L V T L ACH HATS - D LP T A L S A L S D L H A H K L - F - - R V D P A NKL L S H C I L V T L ACH HABOKA - D LP GA L S D L S D L H A H K L - F - - R V D P V NKL L S H S L L V T L A S H HTOR - D LP H A L S A L S H L H ACQ L - F - - R V D P A S Q L L G H C L L V T L A R H HBA_CAIMO - D I A G A L S K L S D L H A QKL - F - - R V D P V NKF L G H C F LVVVA I H HBAT_HO - E LP R A L S A L R H R H V R E L - L - - R V D P A S Q L L G H C L L V T P A R H 1ecd GGICE3 P N I E A D V N T F V A S H K P R - L - N - - T H D N N F R A F V S Y K A H < CTTEE P N I G K H V DAL V A T H K P R G - F - N - - T H A QNNFRAA F I A Y L K G H GGICE1 P T I L A K A K D F G KS H KS RA - L - T - - S P A Q D N F R K S LVVY L K G A 1mbd MY WHP - K - H H E A E L K P L A S H A T K H - L - H K I P I K Y E F I S E A I I H V L H S R < MYG_ CAS FI - K - G HHE A E I K P L A QS H A TKH - L - H K I P I KY E F I S EA I I H V L QS K MYHU - K - G HHE A E I K P L A QS H A TKH - L - H K I P V KY E F I S EC I I Q V L QS K MYBAO - K - G HHE A E I K P L A QS H A TKH - L - H K I P V KY E L I S E S I I Q V L QS K Consensus Profile - c - - d L P A E h p A h p h? H A? K h - h - d c h p h c Y p h h S? C h L V v L h p p <
13 Automatically Comparing Protein Structures Given 2 Structures (A & B), 2Basic Comparison Operations 1 Given an alignment optimally SUPERIMPOSE AontoB Find Best R & T to move A onto B 2 Find an Alignment between A andbbasedontheir3d coordinates 13 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
14 RMS Superposition (1) B A 14 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
15 RMS Superposition (2): Distance Between an Atom in 2 Structures 15 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
16 RMS Superposition (3): RMS Distance Between Aligned Atoms in 2 Structures 16 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
17 RMS Superposition (4): Rigid-Body Rotation and Translation of One Structure (B) 17 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
18 RMS Superposition (5): Optimal Movement of One Structure to Minimize the RMS Methods of Solution: springs (F ~ kx) SVD Kabsch 18 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
19 Alignment (1) Make a Similarity Matrix (Like Dot Plot) A B C N Y R Q C L C R P M A 1 Y 1 C Y 1 N 1 R 1 1 C K C R 1 1 B 1 P 1 19 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
20 PAM(A,V) = 0.5 Structural Alignment (1b) Make a Similarity Matrix (Generalized Similarity Matrix) Applies at every position i, J) Specific Matrix for each pair of residues i in protein 1 and Jinprotein2 Example is Y near N-term. matches any C-term. residue (Y at J=2) S(i,J) Doesn t need to depend on a.a. identities at all! Just need to make up a score for matching residue i in protein 1withresidueJinprotein2 J A B C N Y R Q C L C R P M 1 A 1 2 Y C Y 1 5 N 1 6 R C K 9 C R B 1 12 P 1 i 20 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
21 Structural Alignment (1c*) Similarity Matrix for Structural Alignment Structural Alignment d = Similarity Matrix S(i,J) depends on the 3D coordinates of residues i and J Distance between CA of i and J 2 2 ( xi xj ) + ( yi yj ) + ( zi zj M(i,j) = 100 / (5 + d 2 ) Threading S(i,J) depends on the how well the amino acid at position i in protein 1 fits into the 3D structural environment at position J of protein 2 ) 2 21 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
22 Alignment (2): Dynamic Programming, Start Computing the Sum Matrix new_value_cell(r,c) <= cell(r,c) { Old value, either 1 or 0 } + Max[ cell (R+1, C+1), { Diagonally Down, no gaps } cells(r+1, C+2 to C_max),{ Down a row, making col. gap } cells(r+2 to R_max, C+2) { Down a col., making row gap } ] A B C N Y R Q C L C R P M A 1 Y 1 C Y 1 N 1 R 1 1 C K C R 1 1 B 1 P 1 A B C N Y R Q C L C R P M A 1 Y 1 C Y 1 N 1 R 1 1 C K C R B P (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
23 Alignment (3):Dynamic Programming, Keep Going A B C N Y R Q C L C R P M A 1 Y 1 C Y 1 N 1 R 1 1 C K C R B P A B C N Y R Q C L C R P M A 1 Y 1 C Y 1 N 1 R C K C R B P (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
24 Alignment (4): Dynamic Programming, Sum Matrix All Done A B C N Y R Q C L C R P M A 1 Y 1 C Y 1 N 1 R C K C R B P A B C N Y R Q C L C R P M A Y C Y N R C K C R B P (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
25 Alignment (5): Traceback Find Best Score (8) and Trace Back A B C N Y - R Q C L C R - P M A Y C - Y N R - C K C R B P A B C N Y R Q C L C R P M A Y C Y N R C K C R B P (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
26 In Structural Alignment, Not Yet Done (Step 6*) Use Alignment to LSQ Fit Structure B onto Structure A However, movement of B will now change the Similarity Matrix This Violates Fundamental Premise of Dynamic Programming Way Residue at i is aligned can now affect previously optimal alignment of residues (from 1 to i-1) ACSQRP--LRV-SH -R SENCV A-SNKPQLVKLMTH VK DFCV- 26 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
27 Structural Alignment (7*), Iterate Until Convergence 1 Compute Sim. Matrix 2 Align via Dyn. Prog. 3 RMS Fit Based on Alignment 4 Move Structure B 5Re-computeSim.Matrix 6 If changed from #1, GOTO #2 27 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
28 Structure alignment - Scoring What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surface I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 28 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
29 Score S at End Just Like SW Score, but also have final RMS S=TotalScore S(i,j) = similarity matrix score for aligning i and j Sumiscarriedoutoverallalignediandj n = number of gaps (assuming no gap ext. penalty) G = gap penalty S = S( i, j) ng i, j 29 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
30 Some Similarities are Readily Apparent others are more Subtle Easy: Globins 125 res., ~1.5 Å Tricky: Ig C & V 85 res., ~3 Å Very Subtle: G3P-dehydrogenase, C-term. Domain >5 Å 30 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
31 Some Similarities are Readily Apparent others are more Subtle Easy: Globins 125 res., ~1.5 Å Tricky: Ig C & V 85 res., ~3 Å Very Subtle: G3P-dehydrogenase, C-term. Domain >5 Å 31 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
32 P-values 1 2 [ e.g. P(score s>392) = 1% chance] 3 Significance Statistics For sequences, originally used in Blast (Karlin-Altschul). Then in FASTA, &c. Extrapolated Percentile Rank: How does a Score Rank Relative to all Other Scores? Our Strategy: Fit to Observed Distribution 1)All-vs-All comparison 2)Graph Distribution of Scores in 2D (N dependence); 1K x 1K families -> ~1M scores; ~2K included TPs 3)Fit a function ρ(s) to TN distribution (TNs from scop); Integrating ρ gives P(s>S), the CDF, chance of getting a score better than threshold S randomly 4) Use same formalism for sequence & structure 32 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
33 Statistics on Range of Similarities For 2107 pairs, only 2% Outliers (with subtle similarity) RMS RMS For 2107 scop superfamily pairs N (aligned residues) Num. Aligned Frequency Histogram of RMS or RMS' values RMS RMS' RMS or RMS' 33 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
34 Scores from Structural Alignment Distributed Just Like Ones from Sequence Alignment (E.V.D.) 34 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
35 Same Results for Sequence & Structure Structure 3 Free Parm. fit to EVD involving: a, b,σ. These are the only difference betw. sequence and structure. S Z = M ( i, i, j = j) S G ρ( z) = exp ( a ln N + b) σ ( z z e ) N, G, M also defined differently for sequence and structure. N = number of residues matched. G = total gap penalty. M(i,j) = similarity matrix (Blossum for seq. or M str (i,j), struc.) Sequence 35 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
36 Score Significance (P-value) derived from Extreme Value Distribution (just like BLAST, FASTA) F(s) = E.V.D of scores F(s) = exp(-z(s) - exp(-z(s))) Z(s) = As + ln(n) + B s = Score from random alignment N length of sequence matched A & B are fit parameters P(s>S) = CDF = integral[ F(s) ] P(s>S) = 1 - exp(-exp(-z(s))) Given Score S (1%), P (s > S) is the chance that a given random score s is greater than the threshold i.e. P-value gives chance score would occur randomly Exactly like Sequence Matching Statistics (BLAST and FASTA) 36 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
37 RMS is a similarity Score S str d 2 vs + 2 di i Also, RMS doesn t work instead of structural alignment (no EVD fit) RMS RMS penalizes worst fitting atoms, easily skewed 37 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
38 Structure alignment - Other methods What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surface I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 38 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
39 Refine Method 39 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
40 Significance Ignoring Crucial Features in Structural Similarity 40 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
41 Other Methods of Structural Alignment RMS fitting used universally, but other alignment methods Comparison of Distance Matrices Holm & Sander, DALI Taylor & Orengo Structure Hashing Bryant, VAST Rice, Artymiuk Others Cohen (Soap) Sippl Godzik (Lattice) 41 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
42 DADE WALKER CHATTOOGA FLOYD POLK CATOOSA HARALSON CARROLL HEARD TROUP CLAY GORDON PAULDING HARRIS EARLY COWETA CHATTA- HOOCHEE STEWART MILLER MURRAY BARTOW DOUGLAS MUSCOGEE CALHOUN DECATUR FANNIN GILMER PICKENS CHEROKEE COBB FULTON TALBOT MARION TERRELL RANDOLPH BAKER SPALDING PIKE UPSON TAYLOR SCHLEY DOUGHERTY MITCHELL GRADY UNION LUMPKIN DAWSON DEKALB SUMTER LEE GWINNETT HENRY LAMAR BUTTS MACON THOMAS TOWNS RABUN WHITE HALL NEWTON MONROE CRAWFORD PEACH DOOLY WORTH COLQUITT JACKSON HOUSTON CRISP BARROW WALTON JASPER BIBB JONES TURNER MORGAN PULASKI TIFT BROOKS WILCOX COOK STEPHENS CLARKE PUTNAM TWIGGS MADISON BALDWIN IRWIN BERRIEN LOWNDES OGLETHORPE GREENE WILKINSON DODGE BEN HILL ELBERT WILKES WARREN HANCOCK WASHINGTON LAURENS TELFAIR COFFEE ATKINSON ECHOLS JOHNSON WHEELER JEFF DAVIS CLINCH LINCOLN JEFFERSON TREUTLEN COLUMBIA EMANUEL RICHMOND BURKE JENKINS SCREVEN BULLOCH CANDLER EFFINGHAM EVANS TOOMBS BRYAN TATTNALL LIBERTY APPLING LONG BACON WAYNE McINTOSH PIERCE GLYNN BRANTLEY WARE CHARLTON CAMDEN CHATHAM Fold Library vs. Other Fundamental Data structures Parts List Database; Statistical, rather than mathematical relationships and conclusions Folds in Molecular Biology const. mant. exp. unit e 1.60 e 8C F 9.65 e 4 C/mol ε e -12 F/m µ e -6 H/m h 6.63 e -34 J s k 1.38 e -23 J/K me 9.11 e -31 kg mp 1.67 e -27 kg mn 1.68 e -27 kg a e -11 m λc 2.43 e -12 m c 3.00 e -19 m/s G 6.67 e -11 m 3 /kg s 2 NA 6.02 e 23 mol Physics H He Li Be B C N O F Ne Na Mg Al Si P S Cl Ar K Ca Ga Ge As Se Br Kr Sc Ti V Cr Mn Fe Co Ni Cu Zn Rb Sr Y Zr Nb Mo Tc Ru Rh Pd Ag Cd In Sn Sb Te I Xe Cs Ba Lu Hf Ta W Re Os Ir Pt Au Hg Tl Pb Bi Po At Rn * Fr Ra ** Lr Rf Db Sg Bh Hs Mt Uun Uuu Uub La Ce Pr Nd Pm Sm Eu Gd Tb Dy Ho Er Tm Yb Ac Th Pa U Np Pu Am Cm Bk Cf Es Fm Md No DJ Industrials 4/6/98 4/13/98 4/20/98 4/27/98 5/4/98 5/11/98 5/18/98 5/25/98 6/1/98 6/8/98 6/15/98 6/22/98 6/29/98 7/6/ > Chemistry Finance Politics (Large than physics and chemistry, Similar to Finance (Exact Finite Number of Objects (3,056 on NYSE by 1/98), descrip. by Standardized Statistics (even abbrevs, INTC) and groups (sectors)) Smaller than Social Surveys, Indefinite Number of People, Not Well Defined Vocabulary and statistics. QUITMAN WHITFIELD SEMINOLE MERIWETHER WEBSTER FORSYTH ATLANTA FAYETTE CLAYTON ROCKDALE ALBANY HABERSHAM BANKS FRANKLIN HART OCONEE BLECKLEY ATHENS TALIAFERRO MACON LANIER McDUFFIE GLASCOCK MONTGOMERY 42 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
43 Fold Scop Classifications Chothia, Murzin (Cambridge) Manual classification, auto-alignments available Evolutionary clusters Cath Thornton (London) semi-automatic classification with alignments class, arch, topo., homol. FSSP Sander, Holm (Cambridge) totally automatic with DALI objective but not always interpretable clusters VAST 43 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
44 Sequence-structure Relationships What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surface I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 44 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
45 Adding Structure to Functional Genomics, Function to Structural Genomics? %ID v. RMS? Function Folds v. Genomes Purely Seq. Based Analysis -- e.g. EcoCyc, ENZYME, GenProtEC, COGs, MIPS Why Structure? Do we really need it? 1 Most Highly Conserved 2 Precisely Defined Modules 3 Seq. Struc. Clearer than Seq. Func. 4 Link to Chemistry, Drugs RMSD %ID Drug 45 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
46 Chothia & Lesk, points EMBO J 4: 823 (1986) The relation between the divergence of sequence and structure in proteins 32 pairs of homologous proteins RMS, percent identity =0.40e 1.87H Now redo with >16,000 pairs in scop + auto-alignments (pdb95d) (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
47 Chothia and Lesk, revisited 16K points C&L 86: =.4 exp(1.9 H) Here: =.2exp(1.3H) =.2exp(1.9H) 47 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
48 100 Problems with RMS Dominated by worstfitting atoms Trimming is arbitrary (50%) Bunching up between 20% and 0% identity mean RMS Chothia & Lesk Chothia & Lesk weighted fit 1 =0.40e 1.87H weighted fit 1: weighted fit 2 weighted fit 3 =0.24e 1.41H weighted fit 2: CL Fit =(6.6E-5)e 12.6H median weighted fit 3: =0.22e 1.68H CL Fit: =0.20e 1.87H 80 Weighted fit 1 and CL Fit are fit to percent sequence identity from %, fit 2 to % seq id from 17-25%; fit 3 to 17%-100%. CL Fit was calculated by varying the 0.40 parameter in the Chothia & Lesk function % sequence identity RMS (50% trim) 100 mean RMS Chothia & Lesk weighted fit 1 weighted fit 2 weighted fit 3 CL Fit median leskfity leskdatay 80 Chothia & Lesk =0.40e 1.87H weighted fit 1: =0.78e 2.15H weighted fit 2: =.036e 6.40H weighted fit 3: =0.68e 2.47H CL Fit: =0.96e 1.87H W eighted fit 1 and CL Fit are fit to percent sequence identity from %, fit 2 to % seq id from 17-25%; fit 3 to 17%-100%. CL Fit was calculated by varying the 0.40 parameter in the Chothia & Lesk function (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
49 Structural Comp. Score vs.smith- Waterman Score overcomes zero bunching, trimming problem Sstr = 100(21-11 exp ( SWS) 49 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
50 Problems with Structural Alignment Score Different Lengths give different scores. Scores follow equation of the form: y=an+mx+b 50 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
51 Modern statistical language ~in TZ: P str =10-10 P.05 seq in TZ P str =10-6 P.274 seq overcomes length dependency 51 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
52 Focus on Twilight Zone Sequence Sig. without structure signif. Protein motions small proteins low-res, NMR Struc. Sig. without Seq. signif. More in bottom-right than top-left 52 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
53 Hierarchy of Protein Functions All of SCOP entries ENZYME NON-ENZYME Alcohol dehydro genase NAD and NADP acceptor 1.1 Acting on CH-OH Homo serine dehydro genase 1 Oxidoreductases 1.5 Acting on CH-NH Carboxyl esterase Precise functional similarity 3.1 Acting on ester bonds Carboxylic ester hydrolases 3 Hydrolases 3.4 Acting on peptide bonds Choline esterase 1.2 Nucleotide metab Starch metab. General similarity 1 Metabolism 1.1 Carb. metab Polysach. metab Glycogen metab. 3.1 Nucleus 3 Cell structure Fibronectin 3.8 Extracel. matrix Extracel. matrix glycoprotein Tenascin Functional class similarity 53 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
54 Relationship of Similarity in Sequence & Structuretothatin Function 54 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
55 Relationship of Similarity in Sequence & Structure, & Function - Summary Traditional Scores Aligment Similarity Scores Modern Probabilistic Scores Sequence Similarity Percent sequence identity Structural Similarity RMS C α separation Features Well understood, in use S seq S str Analogous similarity scores, S str depends most highly on best matches P seq P str Statistical significance, unified framework for different comparisons Limitations RMS depends most highly on worst matches, requiring arbitrary trimming Dependence on alignment length Not as familiar as RMS and percent identity, some residual length-dependency 55 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
56 Surfaces I What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 56 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
57 Why calculate? Protein is solid object. Surface is where action takes place. Surface useful for docking and drugdesign Hydrophobic energy proportional to surface area Various Types of Protein Surfaces Accessible Surface Molecular Surface Hydration Surface Accessible Surface Roll sphere (water) on surface and look at locus of sphere centers. Usually represented as a dot surface Not smooth and continuously differentiable (relevant for energy calculations). It has sharp cusps. Protein Surfaces -- Accessible Surface 57 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
58 Molecular Surface Cusps in the Accessible Surface Solution: the smooth molecular surface. M.S. = contact surface + reentrant surface C.S. = points of tangency between probe sphere and protein when probe sphere is only touching one atom R.S. = solid angle of probe sphere when tangent to two protein atoms First proposed by Richards, but hard to calculate. First numeric calc. by Connelly. Later analytic calculation by Connelly. Analytic version is continuously differentiable. 58 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
59 Richards Molecular and Accessible Surfaces Probe Radius Part of Probe Sphere Type of Surface 0 Center (or Tangent) Van der Waals Surface (vdws) 1.4 Å Center Solvent Accessible Surface (SAS) "" Tangent (1 atom) Contact Surface (CS, from parts of atoms) "" Tangent (2 or 3 atoms) Reentrant Surface (RS, from parts of Probe) "" Tangent (1,2, or 3 atoms) Molecular Surface (MS = CS + RS) 10 Å Center A Ligand or Reagent Accessible Surface Tangent Minimum limit of MS (related to convex hull ) "" Center Undefined 59 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
60 How to Calculate Accessible Surface Area Lee & Richards algorithm (first method, 1970) Pick an arbitrary direction from which to view the protein. Slice it into many sections perpendicular to this direction. In each section, cycle over all the atoms. Each atom is represented as a sphere with a radius that is the sumofitsvdwradiusplusthatofa probe solvent -- i.e. 1.4 for water. For each atom determine the circle corresponding to the intersection of this sphere with the sectioning plane. Remove all parts (i.e. arcs) of this circle occluded by the circles of other atoms. Multiply the total amount of non-occluded arc length by the sectioning width to get the surface area for atom. Sum over all atoms and all sections to get total area. 60 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
61 Shrake & Rupley algorithm (easier) Surround each atom with sphere of uniformly spaced dots (e.g. 92). Remove dots contained in other atoms spheres. Total number of remaining dots is accessible surface. 61 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
62 Calculation of Volumes What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 62 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
63 Voronoi Volumes Each atom surrounded by a single convex polyhedron and allocated space within it Allocation of all space (large V implies cavities) 2 methods of determination Find planes separating atoms, intersection of these is polyhedron Locate vertices, which are equidistant from 4 atoms 63 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
64 Classic Papers Lee, B. & Richards, F. M. (1971). The Interpretation of Protein Structures: Estimation of Static Accessibility, J. Mol. Biol. 55, Richards, F. M. (1974). The Interpretation of Protein Structures: Total Volume, Group Volume Distributions and Packing Density, J. Mol. Biol. 82, Richards, F. M. (1977). Areas, Volumes, Packing, and Protein Structure, Ann. Rev. Biophys. Bioeng. 6, (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
65 Calculating Volumes with Voronoi polyhedra In 1908 Voronoi found a way of partitioning all space amongst a collection of points using specially constructed polyhedra. Here we refer to a collection of "atom centers" rather than "points. In 3D, each atom is surrounded by a unique limiting polyhedron such that all points within an atom's polyhedron are closer to this atom than all other atoms. Likewise, points equidistant from 2 atoms form planes (lines in 2D). Those equidistant from 3 atoms form lines, and those equidistant form 4 centers form vertices. 65 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
66 Determining Voronoi Volumes Integrating on a Grid The simplest method for calculating volumes with Voronoi polyhedra is to put all atoms in the system on a fine grid. Then go to each grid-point (i.e., voxel) and add its infinitesimal volume to the atom center closest to it. This is prohibitively slow for a real protein structure, but it can be made somewhat faster by randomly sampling grid-points. It is, furthermore, a useful approach for high-dimensional integration. Solving for the Vertices In the basic Voronoi construction, each atom is surrounded by a unique limiting polyhedron such that all points within an atom s polyhedron are closer to this atom than all other atoms. Points equidistant from 2 atoms lie on a dividing plane; those equidistant from 3 atoms are on a line, and those equidistant from 4 centers form a vertex. It is straightforward to solve for possible vertex coordinates using the equation of a sphere. (That is, one uses four sets of coordinates (x,y,z) and the equation (x-a) 2 +(y-b) 2 + (z-c) 2 =r 2 to solve for the center (a,b,c) and radius (r) of the sphere.) One then checks whether this putative vertex is closer to these four atoms than any other atom; if so, it is a real vertex. 66 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
67 Collecting Vertices and Calculating Volumes To systematically collect the vertices associated with an atom, label each one by the indices of the four atoms with which it is associated. To traverse the vertices on one face of a polyhedron, find all vertices that share two indices and thus have two atoms in common e.g., a central atom (atom 0) and another atom (atom 1). Arbitrarily pick a vertex to start and walk around the perimeter of the face. One can tell which vertices are connected by edges because they will have a third atom in common (in addition to atom 0 and atom 1). This sequential walking procedure also provides a way to draw polyhedra on a graphics device. More importantly, with reference to the starting vertex, the face can be divided into triangles, for which it is trivial to calculate areas and volumes. 67 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
68 Atoms have different sizes Difficulty with Voronoi Meth. Not all atoms created equal Solutions Bisection -- plane midway between atoms Method B (Richards) Positions the dividing plane accordingtoratio Radical Plane VDW Radii Set 68 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
69 Complexity from different atom sizes requires new ways to calculate polyhedra Vertex Error Chopping Down Method of Calculating Polyhedra 69 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
70 Representing Lines, Planes, and their Intersection 70 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
71 Calculating Areas and Volumes from Vectors 71 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
72 Delauney Triangulation, the Natural Way to Define Packing Neighbors Related to Voronoi polyhedra (dual) What coordination number does an atom have? Doesn t depend on distance alpha shape threading 72 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
73 Properties of Voronoi Polyhedra If Voronoi polyhedra are constructed around atoms in a periodic system, such as in a crystal, all the volume in the unit cell will be apportioned to the atoms. There will be no gaps or cavities as there would be if one, for instance, simply drew spheres around the atoms. Voronoi volume of an atom is a weighted average of distances to all its neighbors, where the weighting factor is the contact area with the neighbor. 73 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
74 Voronoi diagrams are generally useful, beyond proteins Border of D.T. is Convex Hull D.T. produces "fatest" possible triangles which makes it convenient for things such as finite element analysis. Nearest neighbor problems. The nearest neighbor of a query point in center of the Voronoi diagram in which it resides Largest empty circle in a collection of points has center at a Voronoi vertex Voronoi volume of "something" often is a useful weighting factor. This fact can be used, for instance, to weight sequences in alignment to correct for over or under-representation 74 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
75 Voronoi Volumes & Packing What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 75 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
76 Voronoi Volumes, the Natural Way to Measure Packing Packing Efficiency = Volume-of-Object Space-it-occupies = V(VDW) / V(Voronoi) Absolute v relative eff. V1 / V2 Other methods Measure Cavity Volume (grids, constructions, &c) 76 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
77 Close-Packing of Spheres Efficiency Volume Spheres / Volume of space Close packed spheres 74% volume filled Coordination of 12 Two Ways of laying out Fcc cubic close packing ABC layers hcp Hexagonally close packed ABABAB Illustration Credits: Atkins, Pchem, (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
78 Other Well Known Sphere Arrangements Simple cubic packing 8 nbrs 52% efficiency bcc cubic packing one sphere sits in middle of 8 others (body-centered) 8 nbrs 68% efficiency fcc -> bcc -> simple apx 3/4, 2/3, 1/2 78 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
79 Optimal Packing Finally Proved Illustration Credits: Singh, New York Times 79 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
80 Water v. Argon More Complex Systems -- what to do? 80 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
81 Small Packing Changes Significant Exponential dependence Bounded within a range of 0.5 (.8 and.3) Many observations in standard volumes gives small error about the mean (SD/sqrt(N)) 81 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
82 Packing ~ VDW force Longer-range isotropic attractive tail provides general cohesion Shorter-ranged repulsion determines detailed geometry of interaction Billiard Ball model, WCA Theory 82 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
83 Close-packing is Default No tight packing when highly directional interactions (such as H-bonds) need to be satisfied Packing spheres (.74), hexagonal Water (~.35), Open tetrahedral, H-bonds 83 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
84 Standard Radii & Volumes What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 84 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
85 Different Sets of Radii Atom Type & Symbol Bondi Lee & Richards Shrake & Rupley Richards Chothia Richmond & Richards Gelin Dunfield ENCAD & Karplus et al. derived CHARMM derived Tsai et al CH 3 Aliphatic, methyl CH 2 - Aliphatic, methyl >CH- Aliphatic, CH =CH Aromatic, CH * >C= Trigonal, aromatic * NH 3 + Amino, protonated NH 2 Amino or amide >NH Peptide, NH or N =O Carbonyl Oxygen OH Alcoholic hydroxyl OM Carboxyl Oxygen SH Sulfhydryl S- Thioether or S-S (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
86 ProtOr Parameter Set Consistent Radii, Typing, and Volumes for Packing Calculations 86 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
87 Factors Affecting Volume Calculations 87 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
88 Set of VDW Radii Great differences in a sensitive parameter (Radii for carbon 1.87 vs 2.00) Complex calculation: minimizing SD, iterative procedure, from protein structures Look for common distances in CCD Preliminary Solution Atom Bondi New C C3H C3H O1HO O2H N S (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
89 Standard Residue Volumes Database of many hi-res structures (~100, 2 Å) Volumes statistics for buried residues (various selections, resample, &c) Standard atomic volumes harder parameter set development... G 64 c 105 T 120 V 139 H 159 M 168 R 194 A 90 C 113 P 124 E 140 L 165 K 170 Y 198 S 94 D 117 N 128 N 150 I 165 F 193 W (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
90 Standard Core Volumes (Prelim.) Atom Types Num. Volume Error (Å 3 ) (%) Mainchain Atoms carbonyl carbon (except G) C alpha carbon (except G) CA nitrogen (except P) N carbonyl oxygen O Gly C Gly CA Pro N Sidechain atoms trigonal or aromatic carbon >C= aromatic CH (H,F,W,Y) -CH= aliphatic CH >CH methylene group -CH methyl group (A,V,L,I) CH hydroxyl oxygen (S,T) OH carbonyl oxygen (N,Q) =O carboxyl oxygen (D,E) O amine (R,H,W) -NH amine or amide (R,N,Q) NH tetrahedral nitrogen (K) NH thioether or disulfide (C,M) -S sulfhydryl (C) SH (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
91 Clustering into a set of Atom Types I Which atoms are equivalent? How many types valid? 18 types, [CNOS][34]H[123][bsu] Chemical Single-link Multi-link 91 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
92 Clustering into a set of Atom Types II Which atoms are equivalent? How many types valid? 18 types, [CNOS][34]H[123][bsu] E statistic to tell apart 92 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
93 Compare Different Structure Sets 93 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
94 Overlap of Volumes of Aromatic C3H1 and Aliphatic C4H2 94 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
95 Surfaces II What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 95 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
96 Packing at Interfaces Voronoi volumes (and D. triangulation) to measure packing Tight core packing v. Loose surface packing Grooves & ridges: closepacking v. H-bonding How packing defines a surface (hydration surface) Implications for Motions 96 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
97 Packing defines the Correct Definition of the Protein Surface Voronoi polyhedra are the Natural way to study packing! How reasonable is a geometric definition of the surface in light of what we know about packing The relationship between accessible surface molecular surface Delauney Triangulation (Convex Hull) polyhedra faces hydration surface 97 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
98 Surface and Volume Definitions Linked 98 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
99 Problem of Protein Surface for Voronoi Construction 99 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
100 Sensitivity of Voronoi Construction to Surface Structure 100 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
101 Hydration Surface Bring together two helices Unusually low water density in grooves and crevices especially, as compared to uncharged water Fit line through second shell 101 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
102 Defining Surfaces from Packing: Convex Hull and Layers of Waters 102 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
103 Defining a Surface from the Faces of Voronoi Polyhedra 103 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
104 Accessible Surface as a Time-averaged Water Layer 104 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
105 The Hydration Surface: Trying to Model Real Water 105 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
106 Other Applications of Volumes -- Motions, Docking What Structures Look Like? Structural Alignment by Iterated Dynamic Programming RMS Superposition Scoring Structural Similarity Other Aspects of Structural Alignment Distance Matrix based methods Fold Library Relation of Sequence Similarity to Structural and Functional Similarity Protein Geometry Surfaces I (Calculation) Calculation of Volume Voronoi Volumes & Packing Standard Volumes & Radii Surfaces II (Relationship to Volumes) Other Applications of Volumes -- Motions, Docking 106 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
107 Intercalcating Interface, Knobs into Holes Packing is a strong constraint on motions Domain or loop motions havetobefast(~10ps 100 ns) Can t cross big energy barriers involved in repacking an interface Not applicable to allosteric motions, which are much slower (~1ms)anddoinvolve repacking interfaces Interface Packing and Motions 107 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
108 Packing Based Classification: Hinge v Shear Hinge Mechanism involves absence of steric constraints (continuously maintained interface), esp. at hinge 108 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
109 Absence of Tight Packing at Hinge Chain Topology is not important 109 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
110 Docking Enzyme Active Sites Lock and Key Induced Fit Theactivesiteofanenzymeis constituted from a relatively small part of thethetotalvolumeofanenzyme The active site is three-dimensional and formed from distant parts of the linear amino-acid or nucleic acid sequence Substrates are bound to enzymes by multiple weak interactions Active sites are usually clefts or crevices in the enzyme that maximize interaction with the substrate and exclude water The active site creates an unusual microenvironment that specifically stabilizes the chemical transition state The specificity of substrate binding depends upon the precise arrangements of atoms within the active site Theactivesitecanbeprearranged(rigid lock and key mechanism) or have a dynamic interaction with the substrate (induced fit mechanism) 110 (c) Mark Gerstein, 1999, Yale, bioinfo.mbb.yale.edu
Local Sales Taxes Collected on Motor Fuel Sales from July 1, 2013 to June 30, 2014
 Appling Gas 10,072,603.00 431,500.00 9,641,103.00 0.03 882,420.46 Diesel 3,917,584.00 158,884.00 3,758,700.00 0.03 390,826.12 Aviation Gas 23,902.00 0.00 23,902.00 0.03 3,921.75 LP.G. 605,405.00 451,313.00
Appling Gas 10,072,603.00 431,500.00 9,641,103.00 0.03 882,420.46 Diesel 3,917,584.00 158,884.00 3,758,700.00 0.03 390,826.12 Aviation Gas 23,902.00 0.00 23,902.00 0.03 3,921.75 LP.G. 605,405.00 451,313.00
The University of Georgia
 The University of Georgia Center for Agribusiness and Economic Development College of Agricultural and Environmental Sciences Economic Vitality Index and Human Vitality Index Data Summaries and Calculations
The University of Georgia Center for Agribusiness and Economic Development College of Agricultural and Environmental Sciences Economic Vitality Index and Human Vitality Index Data Summaries and Calculations
Constructing a Human Development Index for Georgia s Counties. Jeffrey L. Jordan FS-04-08
 Constructing a Human Development Index for Georgia s Counties Jeffrey L. Jordan FS-04-08 Agricultural & Applied Economics University of Georgia, Griffin Campus 1109 Experiment Street Griffin, GA 30223-1797
Constructing a Human Development Index for Georgia s Counties Jeffrey L. Jordan FS-04-08 Agricultural & Applied Economics University of Georgia, Griffin Campus 1109 Experiment Street Griffin, GA 30223-1797
Bioinformatics. Molecular Biophysics & Biochemistry 447b3 / 747b3. Class 3, 1/19/98. Mark Gerstein. Yale University
 Molecular Biophysics & Biochemistry 447b3 / 747b3 Bioinformatics Mark Gerstein Class 3, 1/19/98 Yale University 1 Aligning Text Strings Raw Data??? T C A T G C A T T G 2 matches, 0 gaps T C A T G C A T
Molecular Biophysics & Biochemistry 447b3 / 747b3 Bioinformatics Mark Gerstein Class 3, 1/19/98 Yale University 1 Aligning Text Strings Raw Data??? T C A T G C A T T G 2 matches, 0 gaps T C A T G C A T
RABUN HABERSHAM STEPHENS TALIAFERRO CHANGING RICHMOND SPALDING BUTTS HANCOCK WASHINGTON UPSON LANDSCAPE: JENKINS HARRIS TALBOT WILKINSON JOHNSON
 DADE CATOOSA FANNIN TOWNS MURRAY UNION WALKER GILMER WHITE CHATTOOGA LUMPKIN GORDON PICKENS WHITFIELD DAWSON HALL BANKS FRANKLIN HART FLOYD CHEROKEE BARTOW FORSYTH RABUN HABERSHAM OCONEE STEPHENS TALIAFERRO
DADE CATOOSA FANNIN TOWNS MURRAY UNION WALKER GILMER WHITE CHATTOOGA LUMPKIN GORDON PICKENS WHITFIELD DAWSON HALL BANKS FRANKLIN HART FLOYD CHEROKEE BARTOW FORSYTH RABUN HABERSHAM OCONEE STEPHENS TALIAFERRO
Last 4 Digits of USC ID:
 Chemistry 05 B Practice Exam Dr. Jessica Parr First Letter of last Name PLEASE PRINT YOUR NAME IN BLOCK LETTERS Name: Last 4 Digits of USC ID: Lab TA s Name: Question Points Score Grader 8 2 4 3 9 4 0
Chemistry 05 B Practice Exam Dr. Jessica Parr First Letter of last Name PLEASE PRINT YOUR NAME IN BLOCK LETTERS Name: Last 4 Digits of USC ID: Lab TA s Name: Question Points Score Grader 8 2 4 3 9 4 0
Made the FIRST periodic table
 Made the FIRST periodic table 1869 Mendeleev organized the periodic table based on the similar properties and relativities of certain elements Later, Henri Moseley organized the elements by increasing
Made the FIRST periodic table 1869 Mendeleev organized the periodic table based on the similar properties and relativities of certain elements Later, Henri Moseley organized the elements by increasing
Nucleus. Electron Cloud
 Atomic Structure I. Picture of an Atom Nucleus Electron Cloud II. Subatomic particles Particle Symbol Charge Relative Mass (amu) protons p + +1 1.0073 neutrons n 0 1.0087 electrons e - -1 0.00054858 Compare
Atomic Structure I. Picture of an Atom Nucleus Electron Cloud II. Subatomic particles Particle Symbol Charge Relative Mass (amu) protons p + +1 1.0073 neutrons n 0 1.0087 electrons e - -1 0.00054858 Compare
The Periodic Table. Periodic Properties. Can you explain this graph? Valence Electrons. Valence Electrons. Paramagnetism
 Periodic Properties Atomic & Ionic Radius Energy Electron Affinity We want to understand the variations in these properties in terms of electron configurations. The Periodic Table Elements in a column
Periodic Properties Atomic & Ionic Radius Energy Electron Affinity We want to understand the variations in these properties in terms of electron configurations. The Periodic Table Elements in a column
CHEM 130 Exp. 8: Molecular Models
 CHEM 130 Exp. 8: Molecular Models In this lab, we will learn and practice predicting molecular structures from molecular formulas. The Periodic Table of the Elements IA 1 H IIA IIIA IVA VA VIA VIIA 3 5
CHEM 130 Exp. 8: Molecular Models In this lab, we will learn and practice predicting molecular structures from molecular formulas. The Periodic Table of the Elements IA 1 H IIA IIIA IVA VA VIA VIIA 3 5
CHEM 10113, Quiz 5 October 26, 2011
 CHEM 10113, Quiz 5 October 26, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
CHEM 10113, Quiz 5 October 26, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
Radiometric Dating (tap anywhere)
 Radiometric Dating (tap anywhere) Protons Neutrons Electrons Elements on the periodic table are STABLE Elements can have radioactive versions of itself called ISOTOPES!! Page 1 in your ESRT has your list!
Radiometric Dating (tap anywhere) Protons Neutrons Electrons Elements on the periodic table are STABLE Elements can have radioactive versions of itself called ISOTOPES!! Page 1 in your ESRT has your list!
Ch. 9 NOTES ~ Chemical Bonding NOTE: Vocabulary terms are in boldface and underlined. Supporting details are in italics.
 Ch. 9 NOTES ~ Chemical Bonding NOTE: Vocabulary terms are in boldface and underlined. Supporting details are in italics. I. Review: Comparison of ionic and molecular compounds Molecular compounds Ionic
Ch. 9 NOTES ~ Chemical Bonding NOTE: Vocabulary terms are in boldface and underlined. Supporting details are in italics. I. Review: Comparison of ionic and molecular compounds Molecular compounds Ionic
Speed of light c = m/s. x n e a x d x = 1. 2 n+1 a n π a. He Li Ne Na Ar K Ni 58.
 Physical Chemistry II Test Name: KEY CHEM 464 Spring 18 Chapters 7-11 Average = 1. / 16 6 questions worth a total of 16 points Planck's constant h = 6.63 1-34 J s Speed of light c = 3. 1 8 m/s ħ = h π
Physical Chemistry II Test Name: KEY CHEM 464 Spring 18 Chapters 7-11 Average = 1. / 16 6 questions worth a total of 16 points Planck's constant h = 6.63 1-34 J s Speed of light c = 3. 1 8 m/s ħ = h π
(C) Pavel Sedach and Prep101 1
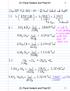 (C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach
(C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 1 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 2 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach and Prep101 3 (C) Pavel Sedach
Using the Periodic Table
 MATH SKILLS TRANSPARENCY WORKSHEET Using the Periodic Table 6 Use with Chapter 6, Section 6.2 1. Identify the number of valence electrons in each of the following elements. a. Ne e. O b. K f. Cl c. B g.
MATH SKILLS TRANSPARENCY WORKSHEET Using the Periodic Table 6 Use with Chapter 6, Section 6.2 1. Identify the number of valence electrons in each of the following elements. a. Ne e. O b. K f. Cl c. B g.
Lecture 6 - Bonding in Crystals
 Lecture 6 onding in Crystals inding in Crystals (Kittel Ch. 3) inding of atoms to form crystals A crystal is a repeated array of atoms Why do they form? What are characteristic bonding mechanisms? How
Lecture 6 onding in Crystals inding in Crystals (Kittel Ch. 3) inding of atoms to form crystals A crystal is a repeated array of atoms Why do they form? What are characteristic bonding mechanisms? How
The Periodic Table of Elements
 The Periodic Table of Elements 8 Uuo Uus Uuh (9) Uup (88) Uuq (89) Uut (8) Uub (8) Rg () 0 Ds (9) 09 Mt (8) 08 Hs (9) 0 h () 0 Sg () 0 Db () 0 Rf () 0 Lr () 88 Ra () 8 Fr () 8 Rn () 8 At (0) 8 Po (09)
The Periodic Table of Elements 8 Uuo Uus Uuh (9) Uup (88) Uuq (89) Uut (8) Uub (8) Rg () 0 Ds (9) 09 Mt (8) 08 Hs (9) 0 h () 0 Sg () 0 Db () 0 Rf () 0 Lr () 88 Ra () 8 Fr () 8 Rn () 8 At (0) 8 Po (09)
9/20/2017. Elements are Pure Substances that cannot be broken down into simpler substances by chemical change (contain Only One Type of Atom)
 CAPTER 6: TE PERIODIC TABLE Elements are Pure Substances that cannot be broken down into simpler substances by chemical change (contain Only One Type of Atom) The Periodic Table (Mendeleev) In 1872, Dmitri
CAPTER 6: TE PERIODIC TABLE Elements are Pure Substances that cannot be broken down into simpler substances by chemical change (contain Only One Type of Atom) The Periodic Table (Mendeleev) In 1872, Dmitri
 lectures accompanying the book: Solid State Physics: An Introduction, by Philip ofmann (2nd edition 2015, ISBN-10: 3527412824, ISBN-13: 978-3527412822, Wiley-VC Berlin. www.philiphofmann.net 1 Bonds between
lectures accompanying the book: Solid State Physics: An Introduction, by Philip ofmann (2nd edition 2015, ISBN-10: 3527412824, ISBN-13: 978-3527412822, Wiley-VC Berlin. www.philiphofmann.net 1 Bonds between
Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Secondary Support Pack. be introduced to some of the different elements within the periodic table;
 Secondary Support Pack INTRODUCTION The periodic table of the elements is central to chemistry as we know it today and the study of it is a key part of every student s chemical education. By playing the
Secondary Support Pack INTRODUCTION The periodic table of the elements is central to chemistry as we know it today and the study of it is a key part of every student s chemical education. By playing the
02/05/09 Last 4 Digits of USC ID: Dr. Jessica Parr
 Chemistry 05 B First Letter of PLEASE PRINT YOUR NAME IN BLOCK LETTERS Exam last Name Name: 02/05/09 Last 4 Digits of USC ID: Dr. Jessica Parr Lab TA s Name: Question Points Score Grader 2 2 9 3 9 4 2
Chemistry 05 B First Letter of PLEASE PRINT YOUR NAME IN BLOCK LETTERS Exam last Name Name: 02/05/09 Last 4 Digits of USC ID: Dr. Jessica Parr Lab TA s Name: Question Points Score Grader 2 2 9 3 9 4 2
Lab Day and Time: Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Chemistry 431 Practice Final Exam Fall Hours
 Chemistry 431 Practice Final Exam Fall 2018 3 Hours R =8.3144 J mol 1 K 1 R=.0821 L atm mol 1 K 1 R=.08314 L bar mol 1 K 1 k=1.381 10 23 J molecule 1 K 1 h=6.626 10 34 Js N A = 6.022 10 23 molecules mol
Chemistry 431 Practice Final Exam Fall 2018 3 Hours R =8.3144 J mol 1 K 1 R=.0821 L atm mol 1 K 1 R=.08314 L bar mol 1 K 1 k=1.381 10 23 J molecule 1 K 1 h=6.626 10 34 Js N A = 6.022 10 23 molecules mol
5 questions, 3 points each, 15 points total possible. 26 Fe Cu Ni Co Pd Ag Ru 101.
 Physical Chemistry II Lab CHEM 4644 spring 2017 final exam KEY 5 questions, 3 points each, 15 points total possible h = 6.626 10-34 J s c = 3.00 10 8 m/s 1 GHz = 10 9 s -1. B= h 8π 2 I ν= 1 2 π k μ 6 P
Physical Chemistry II Lab CHEM 4644 spring 2017 final exam KEY 5 questions, 3 points each, 15 points total possible h = 6.626 10-34 J s c = 3.00 10 8 m/s 1 GHz = 10 9 s -1. B= h 8π 2 I ν= 1 2 π k μ 6 P
If anything confuses you or is not clear, raise your hand and ask!
 CHM 1045 Dr. Light s Section December 10, 2002 FINAL EXAM Name (please print) Recitation Section Meeting Time This exam consists of six pages. Make sure you have one of each. Print your name at the top
CHM 1045 Dr. Light s Section December 10, 2002 FINAL EXAM Name (please print) Recitation Section Meeting Time This exam consists of six pages. Make sure you have one of each. Print your name at the top
Atoms and the Periodic Table
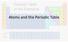 Atoms and the Periodic Table Parts of the Atom Proton Found in the nucleus Number of protons defines the element Charge +1, mass 1 Parts of the Atom Neutron Found in the nucleus Stabilizes the nucleus
Atoms and the Periodic Table Parts of the Atom Proton Found in the nucleus Number of protons defines the element Charge +1, mass 1 Parts of the Atom Neutron Found in the nucleus Stabilizes the nucleus
Lab Day and Time: Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
INSTRUCTIONS: Exam III. November 10, 1999 Lab Section
 CHEM 1215 Exam III John III. Gelder November 10, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
CHEM 1215 Exam III John III. Gelder November 10, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
Atomic Structure & Interatomic Bonding
 Atomic Structure & Interatomic Bonding Chapter Outline Review of Atomic Structure Atomic Bonding Atomic Structure Atoms are the smallest structural units of all solids, liquids & gases. Atom: The smallest
Atomic Structure & Interatomic Bonding Chapter Outline Review of Atomic Structure Atomic Bonding Atomic Structure Atoms are the smallest structural units of all solids, liquids & gases. Atom: The smallest
30 Zn(s) 45 Rh. Pd(s) Ag(s) Cd(s) In(s) Sn(s) white. 77 Ir. Pt(s) Au. Hg(l) Tl. 109 Mt. 111 Uuu. 112 Uub. 110 Uun. 65 Tb. 62 Sm. 64 Gd. 63 Eu.
 Enthalpy changes: experimentally it is much easier to measure heat flow at const pressure - this is enthalpy q p = )H : also nearly all chemical reactions are done at constant pressure. Enthalpy (heat)
Enthalpy changes: experimentally it is much easier to measure heat flow at const pressure - this is enthalpy q p = )H : also nearly all chemical reactions are done at constant pressure. Enthalpy (heat)
Guide to the Extended Step-Pyramid Periodic Table
 Guide to the Extended Step-Pyramid Periodic Table William B. Jensen Department of Chemistry University of Cincinnati Cincinnati, OH 452201-0172 The extended step-pyramid table recognizes that elements
Guide to the Extended Step-Pyramid Periodic Table William B. Jensen Department of Chemistry University of Cincinnati Cincinnati, OH 452201-0172 The extended step-pyramid table recognizes that elements
Solutions and Ions. Pure Substances
 Class #4 Solutions and Ions CHEM 107 L.S. Brown Texas A&M University Pure Substances Pure substance: described completely by a single chemical formula Fixed composition 1 Mixtures Combination of 2 or more
Class #4 Solutions and Ions CHEM 107 L.S. Brown Texas A&M University Pure Substances Pure substance: described completely by a single chemical formula Fixed composition 1 Mixtures Combination of 2 or more
HANDOUT SET GENERAL CHEMISTRY II
 HANDOUT SET GENERAL CHEMISTRY II Periodic Table of the Elements 1 2 3 4 5 6 7 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 IA VIIIA 1 2 H He 1.00794 IIA IIIA IVA VA VIA VIIA 4.00262 3 Li 6.941 11 Na 22.9898
HANDOUT SET GENERAL CHEMISTRY II Periodic Table of the Elements 1 2 3 4 5 6 7 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 IA VIIIA 1 2 H He 1.00794 IIA IIIA IVA VA VIA VIIA 4.00262 3 Li 6.941 11 Na 22.9898
Why all the repeating Why all the repeating Why all the repeating Why all the repeating
 Why all the repeating Why all the repeating Why all the repeating Why all the repeating Patterns What Patterns have you observed in your life? Where to Get Help If you don t understand concepts in chapter
Why all the repeating Why all the repeating Why all the repeating Why all the repeating Patterns What Patterns have you observed in your life? Where to Get Help If you don t understand concepts in chapter
Modified from: Larry Scheffler Lincoln High School IB Chemistry 1-2.1
 Modified from: Larry Scheffler Lincoln High School IB Chemistry 1-2.1 The development of the periodic table brought a system of order to what was otherwise an collection of thousands of pieces of information.
Modified from: Larry Scheffler Lincoln High School IB Chemistry 1-2.1 The development of the periodic table brought a system of order to what was otherwise an collection of thousands of pieces of information.
DO NOW: Retrieve your projects. We will be reviewing them again today. Textbook pg 23, answer questions 1-3. Use the section 1.2 to help you.
 DO NOW: Retrieve your projects. We will be reviewing them again today. Textbook pg, answer questions. Use the section. to help you. Chapter test is FRIDAY. The Periodic Table of Elements 8 Uuo Uus Uuh
DO NOW: Retrieve your projects. We will be reviewing them again today. Textbook pg, answer questions. Use the section. to help you. Chapter test is FRIDAY. The Periodic Table of Elements 8 Uuo Uus Uuh
8. Relax and do well.
 CHEM 15 Exam II John II. Gelder March 4, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last two pages includes a periodic table, a solubility
CHEM 15 Exam II John II. Gelder March 4, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last two pages includes a periodic table, a solubility
What is the periodic table?
 The periodic table of the elements represents one of the greatest discoveries in the history of science that certain elements, the basic chemical substances from which all matter is made, resemble each
The periodic table of the elements represents one of the greatest discoveries in the history of science that certain elements, the basic chemical substances from which all matter is made, resemble each
PERIODIC TABLE OF THE ELEMENTS
 Useful Constants and equations: K = o C + 273 Avogadro's number = 6.022 x 10 23 d = density = mass/volume R H = 2.178 x 10-18 J c = E = h = hc/ h = 6.626 x 10-34 J s c = 2.998 x 10 8 m/s E n = -R H Z 2
Useful Constants and equations: K = o C + 273 Avogadro's number = 6.022 x 10 23 d = density = mass/volume R H = 2.178 x 10-18 J c = E = h = hc/ h = 6.626 x 10-34 J s c = 2.998 x 10 8 m/s E n = -R H Z 2
Lab Day and Time: Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Marks for each question are as indicated in [] brackets.
![Marks for each question are as indicated in [] brackets. Marks for each question are as indicated in [] brackets.](/thumbs/77/75878580.jpg) Name Student Number CHEMISTRY 140 FINAL EXAM December 10, 2002 Numerical answers must be given with appropriate units and significant figures. Please place all answers in the space provided for the question.
Name Student Number CHEMISTRY 140 FINAL EXAM December 10, 2002 Numerical answers must be given with appropriate units and significant figures. Please place all answers in the space provided for the question.
single-layer transition metal dichalcogenides MC2
 single-layer transition metal dichalcogenides MC2 Period 1 1 H 18 He 2 Group 1 2 Li Be Group 13 14 15 16 17 18 B C N O F Ne 3 4 Na K Mg Ca Group 3 4 5 6 7 8 9 10 11 12 Sc Ti V Cr Mn Fe Co Ni Cu Zn Al Ga
single-layer transition metal dichalcogenides MC2 Period 1 1 H 18 He 2 Group 1 2 Li Be Group 13 14 15 16 17 18 B C N O F Ne 3 4 Na K Mg Ca Group 3 4 5 6 7 8 9 10 11 12 Sc Ti V Cr Mn Fe Co Ni Cu Zn Al Ga
Essential Chemistry for Biology
 1 Chapter 2 Essential Chemistry for Biology Biology and Society: More Precious than Gold A drought is a period of abnormally dry weather that changes the environment and one of the most devastating disasters.
1 Chapter 2 Essential Chemistry for Biology Biology and Society: More Precious than Gold A drought is a period of abnormally dry weather that changes the environment and one of the most devastating disasters.
MANY ELECTRON ATOMS Chapter 15
 MANY ELECTRON ATOMS Chapter 15 Electron-Electron Repulsions (15.5-15.9) The hydrogen atom Schrödinger equation is exactly solvable yielding the wavefunctions and orbitals of chemistry. Howev er, the Schrödinger
MANY ELECTRON ATOMS Chapter 15 Electron-Electron Repulsions (15.5-15.9) The hydrogen atom Schrödinger equation is exactly solvable yielding the wavefunctions and orbitals of chemistry. Howev er, the Schrödinger
NAME: FIRST EXAMINATION
 1 Chemistry 64 Winter 1994 NAME: FIRST EXAMINATION THIS EXAMINATION IS WORTH 100 POINTS AND CONTAINS 4 (FOUR) QUESTIONS THEY ARE NOT EQUALLY WEIGHTED! YOU SHOULD ATTEMPT ALL QUESTIONS AND ALLOCATE YOUR
1 Chemistry 64 Winter 1994 NAME: FIRST EXAMINATION THIS EXAMINATION IS WORTH 100 POINTS AND CONTAINS 4 (FOUR) QUESTIONS THEY ARE NOT EQUALLY WEIGHTED! YOU SHOULD ATTEMPT ALL QUESTIONS AND ALLOCATE YOUR
8. Relax and do well.
 CHEM 1225 Exam I John I. Gelder February 4, 1999 Name KEY TA's Name Lab Section Please sign your name below to give permission to post your course scores on homework, laboratories and exams. If you do
CHEM 1225 Exam I John I. Gelder February 4, 1999 Name KEY TA's Name Lab Section Please sign your name below to give permission to post your course scores on homework, laboratories and exams. If you do
8. Relax and do well.
 CHEM 1215 Exam III John III. Gelder November 11, 1998 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
CHEM 1215 Exam III John III. Gelder November 11, 1998 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
CHM 101 PRACTICE TEST 1 Page 1 of 4
 CHM 101 PRACTICE TEST 1 Page 1 of 4 Please show calculations (stuffed equations) on all mathematical problems!! On the actual test, "naked answers, with no work shown, will receive no credit even if correct.
CHM 101 PRACTICE TEST 1 Page 1 of 4 Please show calculations (stuffed equations) on all mathematical problems!! On the actual test, "naked answers, with no work shown, will receive no credit even if correct.
UNIVERSITY OF CALGARY FACULTY OF SCIENCE MIDTERM EXAMINATION CHEMISTRY 353 READ ALL THE INSTRUCTIONS CAREFULLY
 WEDNESDAY MARCH 9th, 2016 UNIVERSITY OF CALGARY FACULTY OF SCIENCE MIDTERM EXAMINATION CHEMISTRY 353 Version 1 Time: 2 Hours READ ALL THE INSTRUCTIONS CAREFULLY PLEASE WRITE YOUR NAME, STUDENT I.D. NUMBER
WEDNESDAY MARCH 9th, 2016 UNIVERSITY OF CALGARY FACULTY OF SCIENCE MIDTERM EXAMINATION CHEMISTRY 353 Version 1 Time: 2 Hours READ ALL THE INSTRUCTIONS CAREFULLY PLEASE WRITE YOUR NAME, STUDENT I.D. NUMBER
The Periodic Table of the Elements
 The Periodic Table of the Elements All matter is composed of elements. All of the elements are composed of atoms. An atom is the smallest part of an element which still retains the properties of that element.
The Periodic Table of the Elements All matter is composed of elements. All of the elements are composed of atoms. An atom is the smallest part of an element which still retains the properties of that element.
Chemistry Standard level Paper 1
 Chemistry Standard level Paper 1 Thursday 12 May 2016 (morning) 45 minutes Instructions to candidates Do not open this examination paper until instructed to do so. Answer all the questions. For each question,
Chemistry Standard level Paper 1 Thursday 12 May 2016 (morning) 45 minutes Instructions to candidates Do not open this examination paper until instructed to do so. Answer all the questions. For each question,
(please print) (1) (18) H IIA IIIA IVA VA VIA VIIA He (2) (13) (14) (15) (16) (17)
 CHEM 10113, Quiz 3 September 28, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
CHEM 10113, Quiz 3 September 28, 2011 Name (please print) All equations must be balanced and show phases for full credit. Significant figures count, show charges as appropriate, and please box your answers!
PART 1 Introduction to Theory of Solids
 Elsevier UK Job code: MIOC Ch01-I044647 9-3-2007 3:03p.m. Page:1 Trim:165 240MM TS: Integra, India PART 1 Introduction to Theory of Solids Elsevier UK Job code: MIOC Ch01-I044647 9-3-2007 3:03p.m. Page:2
Elsevier UK Job code: MIOC Ch01-I044647 9-3-2007 3:03p.m. Page:1 Trim:165 240MM TS: Integra, India PART 1 Introduction to Theory of Solids Elsevier UK Job code: MIOC Ch01-I044647 9-3-2007 3:03p.m. Page:2
CHEM 172 EXAMINATION 1. January 15, 2009
 CHEM 17 EXAMINATION 1 January 15, 009 Dr. Kimberly M. Broekemeier NAME: Circle lecture time: 9:00 11:00 Constants: c = 3.00 X 10 8 m/s h = 6.63 X 10-34 J x s J = kg x m /s Rydberg Constant = 1.096776 x
CHEM 17 EXAMINATION 1 January 15, 009 Dr. Kimberly M. Broekemeier NAME: Circle lecture time: 9:00 11:00 Constants: c = 3.00 X 10 8 m/s h = 6.63 X 10-34 J x s J = kg x m /s Rydberg Constant = 1.096776 x
Advanced Chemistry. Mrs. Klingaman. Chapter 5: Name:
 Advanced Chemistry Mrs. Klingaman Chapter 5: The Periodic Law Name: _ Mods: Chapter 5: The Periodic Law Reading Guide 5.1 History of the Periodic Table (pgs. 125-129) 1) What did Dimitri Mendeleev notice
Advanced Chemistry Mrs. Klingaman Chapter 5: The Periodic Law Name: _ Mods: Chapter 5: The Periodic Law Reading Guide 5.1 History of the Periodic Table (pgs. 125-129) 1) What did Dimitri Mendeleev notice
1 Genesis 1:1. Chapter 10 Matter. Lesson. Genesis 1:1 In the beginning God created the heavens and the earth. (NKJV)
 1 Genesis 1:1 Genesis 1:1 In the beginning God created the heavens and the earth. (NKJV) 1 Vocabulary Saturated having all the solute that can be dissolved at that temperature Neutron a particle with no
1 Genesis 1:1 Genesis 1:1 In the beginning God created the heavens and the earth. (NKJV) 1 Vocabulary Saturated having all the solute that can be dissolved at that temperature Neutron a particle with no
Faculty of Natural and Agricultural Sciences Chemistry Department. Semester Test 1. Analytical Chemistry CMY 283. Time: 120 min Marks: 100 Pages: 6
 Faculty of Natural and Agricultural Sciences Chemistry Department Semester Test 1 Analytical Chemistry CMY 283 Date: 5 September 2016 Lecturers : Prof P Forbes, Dr Laurens, Mr SA Nsibande Time: 120 min
Faculty of Natural and Agricultural Sciences Chemistry Department Semester Test 1 Analytical Chemistry CMY 283 Date: 5 September 2016 Lecturers : Prof P Forbes, Dr Laurens, Mr SA Nsibande Time: 120 min
Chemistry 2 Exam Roane State Academic Festival. Name (print neatly) School
 Name (print neatly) School There are fifteen question on this exam. Each question is weighted equally. n the answer sheet, write your name in the space provided and your answers in the blanks provided.
Name (print neatly) School There are fifteen question on this exam. Each question is weighted equally. n the answer sheet, write your name in the space provided and your answers in the blanks provided.
Chem Exam 1. September 26, Dr. Susan E. Bates. Name 9:00 OR 10:00
 Chem 1711 Exam 1 September 26, 2013 Dr. Susan E. Bates Name 9:00 OR 10:00 N A = 6.022 x 10 23 mol 1 I A II A III B IV B V B VI B VII B VIII I B II B III A IV A V A VI A VII A inert gases 1 H 1.008 3 Li
Chem 1711 Exam 1 September 26, 2013 Dr. Susan E. Bates Name 9:00 OR 10:00 N A = 6.022 x 10 23 mol 1 I A II A III B IV B V B VI B VII B VIII I B II B III A IV A V A VI A VII A inert gases 1 H 1.008 3 Li
8. Relax and do well.
 CHEM 1515 Exam II John II. Gelder October 14, 1993 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 8 different pages. The last two pages include a periodic table, a
CHEM 1515 Exam II John II. Gelder October 14, 1993 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 8 different pages. The last two pages include a periodic table, a
Unit 1 Part 2 Atomic Structure and The Periodic Table Introduction to the Periodic Table UNIT 1 ATOMIC STRUCTURE AND THE PERIODIC TABLE
 UNIT 1 ATOMIC STRUCTURE AND THE PERIODIC TABLE PART 2 INTRODUCTION TO THE PERIODIC TABLE Contents 1. The Structure of the Periodic Table 2. Trends in the Periodic Table Key words: group, period, block,
UNIT 1 ATOMIC STRUCTURE AND THE PERIODIC TABLE PART 2 INTRODUCTION TO THE PERIODIC TABLE Contents 1. The Structure of the Periodic Table 2. Trends in the Periodic Table Key words: group, period, block,
K. 27 Co. 28 Ni. 29 Cu Rb. 46 Pd. 45 Rh. 47 Ag Cs Ir. 78 Pt.
 1 IA 1 ydrogen 1.01 Atomic number Element symbol Element name Atomic mass VIIIA 1 1.01 IIA IIIA IVA VA VIA VIIA 2 e 4.00 Metalloids 3 Li 6.94 4 Be 9.01 5 B 10.81 6 C 12.01 7 N 14.01 8 O 16.00 9 F 19.00
1 IA 1 ydrogen 1.01 Atomic number Element symbol Element name Atomic mass VIIIA 1 1.01 IIA IIIA IVA VA VIA VIIA 2 e 4.00 Metalloids 3 Li 6.94 4 Be 9.01 5 B 10.81 6 C 12.01 7 N 14.01 8 O 16.00 9 F 19.00
Lab Day and Time: Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
CLASS TEST GRADE 11. PHYSICAL SCIENCES: CHEMISTRY Test 4: Matter and materials 1
 CLASS TEST GRADE PHYSICAL SCIENCES: CHEMISTRY Test 4: Matter and materials MARKS: 45 TIME: hour INSTRUCTIONS AND INFORMATION. Answer ALL the questions. 2. You may use non-programmable calculators. 3. You
CLASS TEST GRADE PHYSICAL SCIENCES: CHEMISTRY Test 4: Matter and materials MARKS: 45 TIME: hour INSTRUCTIONS AND INFORMATION. Answer ALL the questions. 2. You may use non-programmable calculators. 3. You
CHEM 108 (Spring-2008) Exam. 3 (105 pts)
 CHEM 08 (Spring-008) Exam. (05 pts) Name: --------------------------------------------------------------------------, CLID # -------------------------------- LAST NAME, First (Circle the alphabet segment
CHEM 08 (Spring-008) Exam. (05 pts) Name: --------------------------------------------------------------------------, CLID # -------------------------------- LAST NAME, First (Circle the alphabet segment
Chapter 12 The Atom & Periodic Table- part 2
 Chapter 12 The Atom & Periodic Table- part 2 Electrons found outside the nucleus; negatively charged Protons found in the nucleus; positive charge equal in magnitude to the electron s negative charge Neutrons
Chapter 12 The Atom & Periodic Table- part 2 Electrons found outside the nucleus; negatively charged Protons found in the nucleus; positive charge equal in magnitude to the electron s negative charge Neutrons
8. Relax and do well.
 CHEM 1314 3;30 pm Theory Exam III John III. Gelder November 13, 2002 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 8 different pages. The last page include a periodic
CHEM 1314 3;30 pm Theory Exam III John III. Gelder November 13, 2002 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 8 different pages. The last page include a periodic
8. Relax and do well.
 CHEM 1014 Exam I John I. Gelder September 16, 1999 Name TA's Name Lab Section Please sign your name below to give permission to post your course scores on homework, laboratories and exams. If you do not
CHEM 1014 Exam I John I. Gelder September 16, 1999 Name TA's Name Lab Section Please sign your name below to give permission to post your course scores on homework, laboratories and exams. If you do not
CMSC 313 Lecture 17 Postulates & Theorems of Boolean Algebra Semiconductors CMOS Logic Gates
 CMSC 313 Lecture 17 Postulates & Theorems of Boolean Algebra Semiconductors CMOS Logic Gates UMBC, CMSC313, Richard Chang Last Time Overview of second half of this course Logic gates &
CMSC 313 Lecture 17 Postulates & Theorems of Boolean Algebra Semiconductors CMOS Logic Gates UMBC, CMSC313, Richard Chang Last Time Overview of second half of this course Logic gates &
UNIVERSITY OF CALGARY FACULTY OF SCIENCE MIDTERM EXAMINATION CHEMISTRY 353 READ ALL THE INSTRUCTIONS CAREFULLY
 TUESDAY MARCH 3rd, 2015 UNIVERSITY OF CALGARY FACULTY OF SCIENCE MIDTERM EXAMINATION CHEMISTRY 353 Version 1 Time: 2 Hours READ ALL THE INSTRUCTIONS CAREFULLY PLEASE WRITE YOUR NAME, STUDENT I.D. NUMBER
TUESDAY MARCH 3rd, 2015 UNIVERSITY OF CALGARY FACULTY OF SCIENCE MIDTERM EXAMINATION CHEMISTRY 353 Version 1 Time: 2 Hours READ ALL THE INSTRUCTIONS CAREFULLY PLEASE WRITE YOUR NAME, STUDENT I.D. NUMBER
Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
INSTRUCTIONS: CHEM Exam I. September 13, 1994 Lab Section
 CHEM 1314.05 Exam I John I. Gelder September 13, 1994 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
CHEM 1314.05 Exam I John I. Gelder September 13, 1994 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
CHEM Come to the PASS workshop with your mock exam complete. During the workshop you can work with other students to review your work.
 It is most beneficial to you to write this mock midterm UNDER EXAM CONDITIONS. This means: Complete the midterm in 1.5 hours. Work on your own. Keep your notes and textbook closed. Attempt every question.
It is most beneficial to you to write this mock midterm UNDER EXAM CONDITIONS. This means: Complete the midterm in 1.5 hours. Work on your own. Keep your notes and textbook closed. Attempt every question.
7. Relax and do well.
 CHEM 1014 Exam III John III. Gelder November 18, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
CHEM 1014 Exam III John III. Gelder November 18, 1999 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 7 different pages. The last page includes a periodic table and
Physical Chemistry I CHEM 4641 Final Exam 13 questions, 30 points
 Physical Chemistry I CHEM 4641 Final Exam 13 questions, 30 points Name: KEY Gas constant: R = 8.314 J mol -1 K -1 = 0.008314 kj mol -1 K -1. Boltzmann constant k = 1.381 10-23 J/K = 0.6950 cm -1 /K h =
Physical Chemistry I CHEM 4641 Final Exam 13 questions, 30 points Name: KEY Gas constant: R = 8.314 J mol -1 K -1 = 0.008314 kj mol -1 K -1. Boltzmann constant k = 1.381 10-23 J/K = 0.6950 cm -1 /K h =
CHEM 107 (Spring-2004) Exam 2 (100 pts)
 CHEM 107 (Spring-2004) Exam 2 (100 pts) Name: ------------------------------------------------------------------------, SSN -------------------------------- LAST NAME, First (Circle the alphabet segment
CHEM 107 (Spring-2004) Exam 2 (100 pts) Name: ------------------------------------------------------------------------, SSN -------------------------------- LAST NAME, First (Circle the alphabet segment
K. 27 Co. 28 Ni. 29 Cu Rb. 46 Pd. 45 Rh. 47 Ag Cs Ir. 78 Pt.
 1 IA 1 H Hydrogen 1.01 Atomic number Element symbol Element name Atomic mass VIIIA 1 H 1.01 IIA IIIA IVA VA VIA VIIA 2 He 4.00 Metalloids 3 Li 6.94 4 Be 9.01 5 B 10.81 6 C 12.01 7 N 14.01 8 O 16.00 9 F
1 IA 1 H Hydrogen 1.01 Atomic number Element symbol Element name Atomic mass VIIIA 1 H 1.01 IIA IIIA IVA VA VIA VIIA 2 He 4.00 Metalloids 3 Li 6.94 4 Be 9.01 5 B 10.81 6 C 12.01 7 N 14.01 8 O 16.00 9 F
CHEM 10123/10125, Exam 2
 CHEM 10123/10125, Exam 2 March 7, 2012 (50 minutes) Name (please print) Please box your answers, and remember that significant figures, phases (for chemical equations), and units do count! 1. (13 points)
CHEM 10123/10125, Exam 2 March 7, 2012 (50 minutes) Name (please print) Please box your answers, and remember that significant figures, phases (for chemical equations), and units do count! 1. (13 points)
Element Cube Project (x2)
 Element Cube Project (x2) Background: As a class, we will construct a three dimensional periodic table by each student selecting two elements in which you will need to create an element cube. Helpful Links
Element Cube Project (x2) Background: As a class, we will construct a three dimensional periodic table by each student selecting two elements in which you will need to create an element cube. Helpful Links
Chemistry 126 Final Examination, Prof. Hanson, May, Section B or D (circle one) Seat Coordinate Name
 Chemistry 126 Final Examination, Prof. Hanson, May, 2004 Section B or D (circle one) Seat Coordinate Name DO NOT OPEN THIS EXAM UNTIL INSTRUCTED TO DO SO Each asterisk () is 5 points. There are 40 s, for
Chemistry 126 Final Examination, Prof. Hanson, May, 2004 Section B or D (circle one) Seat Coordinate Name DO NOT OPEN THIS EXAM UNTIL INSTRUCTED TO DO SO Each asterisk () is 5 points. There are 40 s, for
Faculty of Natural and Agricultural Sciences Chemistry Department. Semester Test 1 MEMO. Analytical Chemistry CMY 283
 Faculty of Natural and Agricultural Sciences Chemistry Department Semester Test 1 MEMO Analytical Chemistry CMY 283 Date: 5 September 2016 Lecturers : Prof P Forbes, Dr Laurens, Mr SA Nsibande Time: 90
Faculty of Natural and Agricultural Sciences Chemistry Department Semester Test 1 MEMO Analytical Chemistry CMY 283 Date: 5 September 2016 Lecturers : Prof P Forbes, Dr Laurens, Mr SA Nsibande Time: 90
8. Relax and do well.
 CHEM 1314.03 Exam I John I. Gelder September 25, 1997 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
CHEM 1314.03 Exam I John I. Gelder September 25, 1997 Name TA's Name Lab Section Please sign your name below to give permission to post, by the last 4 digits of your student I.D. number, your course scores
FY 2002 Annual Report
 FY 2002 Annual Report Judicial Council of Georgia Administrative Office of the Courts July 1, 2001 June 30, 2002 Published by the Judicial Council of Georgia and the Administrative Office of the Courts
FY 2002 Annual Report Judicial Council of Georgia Administrative Office of the Courts July 1, 2001 June 30, 2002 Published by the Judicial Council of Georgia and the Administrative Office of the Courts
Lab Day and Time: Instructions. 1. Do not open the exam until you are told to start.
 Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
Name: Lab Day and Time: Instructions 1. Do not open the exam until you are told to start. 2. This exam is closed note and closed book. You are not allowed to use any outside material while taking this
CHEM 167 FINAL EXAM MONDAY, MAY 2 9:45 11:45 A.M GILMAN HALL
 PROF. JOHN VERKADE SPRING 2005 THIS EXAM CONSISTS OF 12 QUESTIONS ON 9 PAGES CHEM 167 HOUR EXAM IV APRIL 20, 2005 SEAT NO. NAME RECIT. INSTR. RECIT. SECT. GRADING PAGE Page 2 Page 3 Page 4 Page 5 Page
PROF. JOHN VERKADE SPRING 2005 THIS EXAM CONSISTS OF 12 QUESTIONS ON 9 PAGES CHEM 167 HOUR EXAM IV APRIL 20, 2005 SEAT NO. NAME RECIT. INSTR. RECIT. SECT. GRADING PAGE Page 2 Page 3 Page 4 Page 5 Page
POLYTECHNIC OF NAMIBIA
 POLYTECHNIC OF NAMIBIA DEPARTMENT OF HEALTH SCIENCES BACHELOR OF ENVIRONMENTAL HEALTH SCIENCES HEALTH SCIENCE CHEMISTRY (HSC 511S) NQF level 5 SECOND OPPORTUNITY EXAMINATION November 2014 TIME: MARKS:
POLYTECHNIC OF NAMIBIA DEPARTMENT OF HEALTH SCIENCES BACHELOR OF ENVIRONMENTAL HEALTH SCIENCES HEALTH SCIENCE CHEMISTRY (HSC 511S) NQF level 5 SECOND OPPORTUNITY EXAMINATION November 2014 TIME: MARKS:
7. Relax and do well.
 CHEM 1215 Exam II John II. Gelder October 7, 1998 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 5 different pages. The last page includes a periodic table and a solubility
CHEM 1215 Exam II John II. Gelder October 7, 1998 Name TA's Name Lab Section INSTRUCTIONS: 1. This examination consists of a total of 5 different pages. The last page includes a periodic table and a solubility
M10/4/CHEMI/SPM/ENG/TZ2/XX+ CHEMISTRY. Wednesday 12 May 2010 (afternoon) 45 minutes INSTRUCTIONS TO CANDIDATES
 M10/4/CHEMI/SPM/ENG/TZ/XX+ 106116 CHEMISTRY standard level Paper 1 Wednesday 1 May 010 (afternoon) 45 minutes INSTRUCTIONS TO CANDIDATES Do not open this examination paper until instructed to do so. Answer
M10/4/CHEMI/SPM/ENG/TZ/XX+ 106116 CHEMISTRY standard level Paper 1 Wednesday 1 May 010 (afternoon) 45 minutes INSTRUCTIONS TO CANDIDATES Do not open this examination paper until instructed to do so. Answer
נושא מס' 8: המבנה האלקטרוני של אטומים. Electronic Structure of Atoms. 1 Prof. Zvi C. Koren
 נושא מס' 8: המבנה האלקטרוני של אטומים Electronic Structure of Atoms 1 Prof. Zvi C. Koren 19.07.10 The Electron Spin From further experiments, it was evident that the e had additional magnetic properties
נושא מס' 8: המבנה האלקטרוני של אטומים Electronic Structure of Atoms 1 Prof. Zvi C. Koren 19.07.10 The Electron Spin From further experiments, it was evident that the e had additional magnetic properties
Advanced Placement. Chemistry. Integrated Rates
 Advanced Placement Chemistry Integrated Rates 204 47.90 9.22 78.49 (26) 50.94 92.9 80.95 (262) 52.00 93.94 83.85 (263) 54.938 (98) 86.2 (262) 55.85 0. 90.2 (265) 58.93 02.9 92.2 (266) H Li Na K Rb Cs Fr
Advanced Placement Chemistry Integrated Rates 204 47.90 9.22 78.49 (26) 50.94 92.9 80.95 (262) 52.00 93.94 83.85 (263) 54.938 (98) 86.2 (262) 55.85 0. 90.2 (265) 58.93 02.9 92.2 (266) H Li Na K Rb Cs Fr
610B Final Exam Cover Page
 1 st Letter of Last Name NAME: 610B Final Exam Cover Page No notes or calculators of any sort allowed. You have 3 hours to complete the exam. CHEM 610B, 50995 Final Exam Fall 2003 Instructor: Dr. Brian
1 st Letter of Last Name NAME: 610B Final Exam Cover Page No notes or calculators of any sort allowed. You have 3 hours to complete the exam. CHEM 610B, 50995 Final Exam Fall 2003 Instructor: Dr. Brian
A little history. When and How? Sir William Ramsey. ü 12/5/13. ü 1. Who put together the first useable Periodic Table?
 ü // A little history Johahann Dobereiner (80-89) o Triads John Newlands (8-898) o Law of Octaves Who put together the first useable ic Table? Mendeleev you remember him right? When and How? You know it
ü // A little history Johahann Dobereiner (80-89) o Triads John Newlands (8-898) o Law of Octaves Who put together the first useable ic Table? Mendeleev you remember him right? When and How? You know it
CHEM 107 (Spring-2005) Exam 3 (100 pts)
 CHEM 107 (Spring-2005) Exam 3 (100 pts) Name: ------------------------------------------------------------------------, Clid # ------------------------------ LAST NAME, First (Circle the alphabet segment
CHEM 107 (Spring-2005) Exam 3 (100 pts) Name: ------------------------------------------------------------------------, Clid # ------------------------------ LAST NAME, First (Circle the alphabet segment
M11/4/CHEMI/SPM/ENG/TZ2/XX CHEMISTRY STANDARD LEVEL PAPER 1. Monday 9 May 2011 (afternoon) 45 minutes INSTRUCTIONS TO CANDIDATES
 M11/4/CHEMI/SPM/ENG/TZ/XX 116116 CHEMISTRY STANDARD LEVEL PAPER 1 Monday 9 May 011 (afternoon) 45 minutes INSTRUCTIONS TO CANDIDATES Do not open this examination paper until instructed to do so. Answer
M11/4/CHEMI/SPM/ENG/TZ/XX 116116 CHEMISTRY STANDARD LEVEL PAPER 1 Monday 9 May 011 (afternoon) 45 minutes INSTRUCTIONS TO CANDIDATES Do not open this examination paper until instructed to do so. Answer
Chem GENERAL CHEMISTRY I MIDTERM EXAMINATION
 Concordia University CHEM 205 Fall 2009, B LAST NAME: FIRST NAME: STUDENT ID: Chem 205 - GENERAL CHEMISTRY I MIDTERM EXAMINATION PLEASE READ THIS BOX WHILE WAITING TO START INSTRUCTIONS: Calculators are
Concordia University CHEM 205 Fall 2009, B LAST NAME: FIRST NAME: STUDENT ID: Chem 205 - GENERAL CHEMISTRY I MIDTERM EXAMINATION PLEASE READ THIS BOX WHILE WAITING TO START INSTRUCTIONS: Calculators are
Chem 51, Spring 2015 Exam 8 (Chp 8) Use your Scantron to answer Questions There is only one answer for each Question. Questions are 2 pt each.
 Chem 51, Spring 2015 Exam 8 (Chp 8) Name 120 pt Use your Scantron to answer Questions 1-32. There is only one answer for each Question. Questions are 2 pt each. CHP 8.1 Solutions are Mixtures 1. A saturated
Chem 51, Spring 2015 Exam 8 (Chp 8) Name 120 pt Use your Scantron to answer Questions 1-32. There is only one answer for each Question. Questions are 2 pt each. CHP 8.1 Solutions are Mixtures 1. A saturated
Chapter 3: Stoichiometry
 Chapter 3: Stoichiometry Chem 6A Michael J. Sailor, UC San Diego 1 Announcements: Thursday (Sep 29) quiz: Bring student ID or we cannot accept your quiz! No notes, no calculators Covers chapters 1 and
Chapter 3: Stoichiometry Chem 6A Michael J. Sailor, UC San Diego 1 Announcements: Thursday (Sep 29) quiz: Bring student ID or we cannot accept your quiz! No notes, no calculators Covers chapters 1 and
Topic 3: Periodicity OBJECTIVES FOR TODAY: Fall in love with the Periodic Table, Interpret trends in atomic radii, ionic radii, ionization energies &
 Topic 3: Periodicity OBJECTIVES FOR TODAY: Fall in love with the Periodic Table, Interpret trends in atomic radii, ionic radii, ionization energies & electronegativity The Periodic Table What is the periodic
Topic 3: Periodicity OBJECTIVES FOR TODAY: Fall in love with the Periodic Table, Interpret trends in atomic radii, ionic radii, ionization energies & electronegativity The Periodic Table What is the periodic
