Molecular Forces in Biological Systems - Electrostatic Interactions; - Shielding of charged objects in solution
|
|
|
- Alison Carroll
- 6 years ago
- Views:
Transcription
1 Molecular Forces in Biological Systems - Electrostatic Interactions; - Shielding of charged objects in solution
2 Electrostatic self-energy, effects of size and dielectric constant q q r r ε ε 1 ε 2? δq ΔU = ΔW Born = δu el = qδq 4πε 0 εr brought from infinity U el = 1 4πεε 0 q2 * 1 # 1 1 & % ( = cz2 4πε 0 2r $ ε 2 ε 1 ' 2r q q 2 qdq = 8πε 0 εr 0 # 1 1 & % ( $ ε 2 ε 1 ' where c=e 2 /4πε 0 = 14.4 ev*å = 2.3*10-28 J*m For a typical small ion (r = 2 4Å) the energy of transfer between water (ε=80) and interior of cellular membrane (ε=2) is about kcal/mol. The larger the ion, the easier is to transfer it to low dielectric medium
3 Biological macromolecules are stabilized by physical interactions: Strong Weak kj/mol ( kcal/mol) Covalent bonds kj/mol ( kcal/mol) Ionic interactions Ion-dipole interactions Hydrogen bonds Van der Waals interactions Hydrophobic effect
4 Why care about electrostatics? Ø Longest-range biologicaly relevant interactions. Ø A lot of biological molecules are charged. Amino acids: Asp-, Glu-, Lys+ Arg+, His+; DNA: phosphates- in backbone, lipids, salt ions, etc. Sugars Membranes trna (dense phosphate packing) dsdna
5 Electrostatic field V(r) = q A 4πεε 0 r q E(r) = A 4πεε 0 r 2 r r! E V ( r)! V ( r)! V ( r)! = V ( r) = i + j + k dx dy dz Energy! = q B V(r)! F ( r) = q B E Assumptions: homogeneous dielectric medium point charges no mobile ions infinite boundaries
6 Electrostatic force F(r) = q A q B 4πεε 0 r 2 r r Force on charge B due to charge A. Superposition U = q ref 4πε 0 ε F = q ref 4πε 0 ε i i q i x i x ref q i x i x ref 2 x i x ref x i x ref
7 A mean field theory replaces the interaction between elements in the system by the interaction of a single element with an effective field, which is the sum of the external field and the internal field.
8 Bjerrum length. The distance at which two unit charges interact with kt of energy. k l B B T = = 2 e 4πDε l 2 e 4πDε k 0 0 B B T Approximately 7 Å for water (ε = 80) at 298 o K. Electrostatic interactions are much stronger than most other nonbonded interactions; e.g., gravitational. U U U U grav elec elec grav Gm = r r e = 4πε r r 2 64 p c J m J m
9 Flux of an electric field Fluxes arise from - Sources: positive charges - Sinks: negative charges Electric field flux: integral of electric displacement over a surface. Φ = Boundary surface of volume Ω Ω εe( s) d s Electric displacement Jacobian; points in surface normal direction
10 Gauss law The integral of field flux through a closed, simple surface is equal to the total charge inside the surface This is true for both homogeneous and inhomogeneous dielectric media Point charge in a sphere Point charge has spherically-symmetric field Field is constant on sphere surface Flux is independent of sphere diameter Φ = εe(r)4πr 2 = ε q 4πε 0 εr 2 4πr2 = q ε 0 Φ = εe( s) d s Ω = 1 ρ ( x)d x ε 0 Ω
11 Field from a line charge Assumptions: Homogeneous medium Line of length L, where L is very big (radial symmetry) Linear charge density of λ Φ = 2π 0 εe( s) d s (cylin der) L 0 = εe (r )r dz d θ = 2πrLεE (r ) 2πrLεE (r ) = λl ε 0 λ E (r ) = 2πεε 0 r
12 Field around DNA B-DNA shape: 2 phosphates every 3.4 Å water dielectric constant of 80 What is the field 40 Å away from a very long B-DNA molecule? E ( r ) = = λ 2πε 0 εr ( C) ( m) 2π ( C 2 J -1 m -1 ) 80 = V m -1 ( ) ( ) m
13 Field from a charged plane Assumptions: Homogeneous medium Surface of area A, Surface charge density of σ Φ = 2π 0 εe( s) d s ( pillbox ) z 0 = 2 εe (r )r dz d θ = 4π R 2 εe (r ) = 2AεE (r ) 2AεE (r ) = σ A ε 0 E (r ) = σ 2εε 0 What is the field at distance r from the source?
14 Field around a membrane POPS membrane -1 e charge per lipid 1 lipid per 55 Å 2 What is the field 20 Å away from the membrane (in water)? Bigger than DNA!! 1 e $! σ = # & C$! Å 2 $ # &# & " 55 Å 2 %" e %" m % = 0.29 C m -2 E ( z ) = σ 2ε 0 ε = ( 0.29 C m -2 ) ( ) C 2 J -1 m -1 = V m -1 ( ) Mukhopadhyay P, et al. Biophys J 86 (3) , 2004.
15 p = 3.8 Debye Electric dipole moment! p =! qr Interaction energy of a dipole with a field U p! = q r! i i i! = p! E p 1 Debye = 0.2 electron-angstroms = 3.3x10 23 Cm
16 Charge-Dipole and Dipole-Dipole interactions q charge - dipole + q θ a r - q U U = = static qd cosθ 2 4πε εr with Brownian tumbling q ( 4πε 0ε ) 3kTr 0 d 2 static U = d1d2k 4πε εr 0 3 K orientation factor dependent on angles d 1 d 2 r with Brownian motion U 2 2 d1 d2 = 2 3kT (4πε ε ) 0 2 r 6
17 Partial Charges N-Acetyl-N -Methylserinyl amide
18 Electrostatics of maromolecules
19 A continuum dielectric medium Ø Reduces the strength of electrostatic interactions relative to a vacuum. Ø It has no atomic detail. An isotropic dielectric continuum exhibits the same response in all directions q the dielectric tensor can be reduced to a scalar q for a homogeneous isotropic Coulomb s law takes a very simple, scaled form [ε(r) ϕ(r)] = ρ(r) ε 0 V(r) = ρ(r) ε 0 ε u(r) = q 1q 2 4πεε 0 r
20 Dielectric Boundaries Boundary can be modeled as a step function change in dielectric constant. The relevant form of the Poisson equation is: [ε(r) ϕ(r)] = ρ(r) ε 0 The dielectric discontinuity can serve as a source of E- field lines even if there are no source charges there (ρ = 0).
21 Image Charges & Forces When a charge approaches a dielectric discontinuity: " q image = ε ε % 2 1 $ 'Q = kq # ε 2 +ε 1 & (charge is in medium with ε 1 ) Force = kq 2 /(4πε 0 ε 1 (2d) 2 ) k < 0 when ε 1 < ε 2 (charge in low dielectric) image charge of opposite sign, force is attractive. k > 0 when ε 1 > ε 2 (charge in high dielectric) image charge of same sign, force is repulsive.
22 Induced dipole! p =αε 0! E A α polarizability of the molecule Interaction energy between the induced dipole and the inducing field U = E!! p de E αε!! = EdE = αε E 0 2 Induced dipoles: dependent upon polarizability of molecule, how easily electrons can shuffle around to react to a charge
23 Induced dipoles and Van der Waals (dispersion) forces E - d = αe d dipole moment + a - polarizability d r U = αd ( 4π ) ε 0εr neutral molecule in the field constant dipole induced dipole r induced dipoles (all polarizable molecules are attracted by dispersion forces) U I1I2α1α 2 = 4 ( I + I )3n r 1 I 1,2 ionization energies α 1,2 polarizabilities 2 n refractive index of the medium 6 Large planar assemblies of dipoles are capable of generating long-range interactions
24 van der Waals interactions this is a general term for the favorable interactions that occur between uncharged atoms van der Waals forces include: permanent dipole-permanent dipole interactions permanent dipole-induced dipole interactions dispersion interactions all have energies that depend on 1/distance 6 we consider London dispersion interactions here: these are common to all atoms even atoms that have no permanent dipoles (e.g. Ar)
25 London dispersion interactions also known as temporary dipole-induced dipole interactions consider an argon atom on average, its electron distribution will be spherically symmetrical: but at any instant, it will not be perfectly symmetric, e.g.: when the distribution is like this, there will be an electronic dipole: δ+ δ
26 London dispersion interactions now consider a second atom adjacent to the first: it will be polarized by the first atom: this will lead to a favorable dipole-dipole interaction between the two atoms the interaction occurs because the electron distributions become correlated
27 London dispersion interactions London J Chem Soc 33, 8-26 (1937) the magnitude of this effect depends on the distance between the two atoms: r E dispersion 1 / r 6 (negative energy à favorable) it also depends on the atoms polarizabilities (a): E dispersion a atom1 x a atom2 a describes how much an electron distribution can fluctuate or respond to an applied electric field
28 London dispersion interactions a (of atoms and molecules) generally increases with more electrons: relative polarizability He Ne Ar Kr Xe boiling point stronger dispersion interactions so more energy required to do this: gas phase ΔH vaporization liquid phase
29 London dispersion interactions the dispersion interaction between many atoms is usually approximated as a sum of London terms: E dispersion = Σ E London (r ij ) for N atoms, the total energy will be a sum of N(N-1)/2 terms this is the pairwise additivity assumption
30 London dispersion + sterics we can now consider the sum of the two types of interactions that we ve seen so far consider approach of two uncharged atoms (e.g. Ar) E dispersion + repulsion (kcal/mol) term r 12 1 term r 6 total 3 3,5 4 4,5 5 5,5 6 distance between atoms(a) unfavorable electron overlap dominates at short distance favorable dispersion interaction dominates at long distance
31 Dispersion interactions & protein stability so favorable interactions with close-packed atoms in the folded state are partly balanced by: favorable interactions with water molecules in the unfolded state the density of atoms is higher in a protein s folded state than in water (proteins are very tightly packed) so dispersion interactions will stabilize proteins
32 Dispersion interactions in nature kg 000 (theoretically) nanohairs Photo: M. Moffet Geckos get a grip using Van-der-Waalsforces Autumn et al. PNAS 99, (2002)
33 The seta has nanohairs The Gecko toe has microhairs (setae) Nanostructure of the Gecko toe
34 Technical surface 1 Contact area Adhesion The Gecko effect through Van-der-Waals-forces Technical surface Technical surface 2 Nanohairs! Small Large contact area small large adhesion force Microhair If all of a gecko s setae were stuck to a surface at the same time it would be able to support 140 kg!
35 ( From micro to nano contacts in biological attachment devices, Arzt et al., PNAS 100(19) 2003)) lizards spiders Small insects: Compliant pads (to shape themselves to rough surfaces) + sweat Medium insects: Multiple pads per leg + sweat Spiders and lizards: Lots of pads (hair) per foot - but dry / no secretions flies bugs beetles
36 Gecko-Tape Stanford University s Sticky Bot
37 Electrostatic Steering System Potential dependence on distance Energy [kj/mol] ion-ion r ion-dipole r 2 15 dipol dipol r 3 2 London r 6 1 r -1 r -2 r -3 r ,9 0,8 0,7 0,6 0,5 0,4 0,3 0,2 0,
38 Interaction between ions and macromolecules Non-specific screening effects Depends only on ionic strength (not species) Results of damped electrostatic potential Described by Debye-Hückel and Poisson-Boltzmann theory for low concentration Site-specific binding Ionic specific (concentration of specific ion, not necessarily ionic strength) Site geometry, electrostatics, coordination, etc. enables favorable binding Influences Co-factors Allosteric activation Folding (RNA)
39 Polar solvation: Poisson equation Poisson equation Classic equation for continuum electrostatics Can be derived from hard sphere dipolar solvent Assumptions No dielectric saturation (linear response) No solvent-solvent correlation (local response) ε ( ) u( ) πρ( ) u( ) u ( ) x x = 4 x for x Ω x = x for x Ω 0
40 Polar solvation: Boltzmann equation Mean field model Include mobile charges Boltzmann distribution Ignore charge-charge correlations Finite ion size Weak ion-ion electrostatics Low ion correlations Missing ion chemistry No detailed ion-solvent interactions No ion coordination, etc ( x) = f ( x) + m( x) = ρ f ( x) + i i ρ ρ ρ i qce qu i ( x) V( x) i
41 Poisson-Boltzmann equation Biomolecular The space dielectric charge distribution properies 2 4πec ε(x) u(x) z kt i δ( x x Mobile charge distribution 4πρ( x) i i ) i ε(x) ϕ(x) = 4π q i δ(x x i ) + 4πρ(x) φ( ) = 0
42 Mobile ion charge distribution form: Boltzmann distribution no explicit ion-ion interaction No detailed structure for atom (de)solvation Result: Nonlinear partial differential equation
43 For not very high ion concentrations κa << 1 V ( r) = z+ e 4πεε r 0 e κr q Viewed from beyond the counterion cloud, the macroion appears neutral. q Macroions will not feel each other until they re nearby q Once they are nearby, the detailed surface pattern of charges can be felt by its neighbor, not just overall charge - streospecificity.
44 Hydrogen bonds The hydrogen bonds is in the range of 3-40 kcal mol -1. The covalent bond between H and O in water is about 492 kj mol -1. The van der Waals interaction is about 5.5 kj mol -1.
45 A hydrogen bond consists of a hydrogen atom lying between two small, strongly electronegative atoms with lone pairs of electrons (N, O, F). Strength of an H-bond is related to the D-H---A - Distance - The D-H-A angle. The hydrogen bond is stronger than typical electrostatic interactions between partial charges, but it is easily disassociated by heat or by interaction with other atoms.
46 Distance Van der Waals radius of H: 1.1Å, O 1.5 Å. The closest approach should be 2.6 Å. Separation is about 1 Å less! It is 1.76 Å. Intermediate between VdW distance and typical O-H covalent bond of 0.96Å. The shorter the distance between D & A the stronger the interaction.
47 Hydrogen bond is directional Hydrogen bond potential energy U = A r B r A' B' cosθ + cosθ 12 6 r r 12 6 ( 1 )
48 The hydrogen bonds define secondary structure of proteins. They are formed between the backbone oxygens and amide hydrogens. Hydrogen bonds define protein binding specificity
49
Atomic and molecular interaction forces in biology
 Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Atomic and molecular interaction forces in biology 1 Outline Types of interactions relevant to biology Van der Waals interactions H-bond interactions Some properties of water Hydrophobic effect 2 Types
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
2 Structure. 2.1 Coulomb interactions
 2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
2 Structure 2.1 Coulomb interactions While the information needed for reproduction of living systems is chiefly maintained in the sequence of macromolecules, any practical use of this information must
The change in free energy on transferring an ion from a medium of low dielectric constantε1 to one of high dielectric constant ε2:
 The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
Aqueous solutions. Solubility of different compounds in water
 Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Aqueous solutions Solubility of different compounds in water The dissolution of molecules into water (in any solvent actually) causes a volume change of the solution; the size of this volume change is
Molecular Modeling -- Lecture 15 Surfaces and electrostatics
 Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Molecular Modeling -- Lecture 15 Surfaces and electrostatics Molecular surfaces The Hydrophobic Effect Electrostatics Poisson-Boltzmann Equation Electrostatic maps Electrostatic surfaces in MOE 15.1 The
Fundamental Interactions: 6 Forces
 Fundamental Interactions: 6 Forces In nuclear and high-energy physics 6 fundamental forces are recognized, which describe the structure of matter. - the strong interaction - the weak interaction act inside
Fundamental Interactions: 6 Forces In nuclear and high-energy physics 6 fundamental forces are recognized, which describe the structure of matter. - the strong interaction - the weak interaction act inside
1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian. which is given in spherical coordinates by (2)
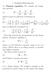 1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian operator (1) 2 = 2 x + 2 2 y + 2 2 z 2 which is given in spherical coordinates by (2) 2 = 1 ( r 2 ) + 1 r 2 r r r 2 sin θ θ and in
1. Poisson-Boltzmann 1.1. Poisson equation. We consider the Laplacian operator (1) 2 = 2 x + 2 2 y + 2 2 z 2 which is given in spherical coordinates by (2) 2 = 1 ( r 2 ) + 1 r 2 r r r 2 sin θ θ and in
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
INTERMOLECULAR AND SURFACE FORCES
 INTERMOLECULAR AND SURFACE FORCES SECOND EDITION JACOB N. ISRAELACHVILI Department of Chemical & Nuclear Engineering and Materials Department University of California, Santa Barbara California, USA ACADEMIC
INTERMOLECULAR AND SURFACE FORCES SECOND EDITION JACOB N. ISRAELACHVILI Department of Chemical & Nuclear Engineering and Materials Department University of California, Santa Barbara California, USA ACADEMIC
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Biochemistry 530: Introduction to Structural Biology. Autumn Quarter 2014 BIOC 530
 Biochemistry 530: Introduction to Structural Biology Autumn Quarter 2014 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic
Biochemistry 530: Introduction to Structural Biology Autumn Quarter 2014 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic
Molecular interactions. Levente Novák István Bányai Zoltán Nagy Department of Physical Chemistry
 Molecular interactions Levente Novák István Bányai Zoltán Nagy Department of Physical Chemistry Characterization of colloidal systems Degree of dispersion (=size) Morphology (shape and internal structure)
Molecular interactions Levente Novák István Bányai Zoltán Nagy Department of Physical Chemistry Characterization of colloidal systems Degree of dispersion (=size) Morphology (shape and internal structure)
Intermolecular Forces I
 I How does the arrangement of atoms differ in the 3 phases of matter (solid, liquid, gas)? Why doesn t ice just evaporate into a gas? Why does liquid water exist at all? There must be some force between
I How does the arrangement of atoms differ in the 3 phases of matter (solid, liquid, gas)? Why doesn t ice just evaporate into a gas? Why does liquid water exist at all? There must be some force between
Chimica Farmaceutica
 Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Chimica Farmaceutica Drug Targets Why should chemicals, some of which have remarkably simple structures, have such an important effect «in such a complicated and large structure as a human being? The answer
Water. Dr. Diala Abu-Hassan, DDS, PhD Lecture 2 MD summer Dr. Diala Abu-Hassan
 Water, DDS, PhD Dr.abuhassand@gmail.com Lecture 2 MD summer 2014 1 Lecture Content Importance of water in biological systems Noncovalent interactions Water structure Water properties Water as a solvent
Water, DDS, PhD Dr.abuhassand@gmail.com Lecture 2 MD summer 2014 1 Lecture Content Importance of water in biological systems Noncovalent interactions Water structure Water properties Water as a solvent
Intermolecular and Intramolecular Forces. Introduction
 Intermolecular and Intramolecular Forces Introduction Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule)
Intermolecular and Intramolecular Forces Introduction Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule)
General Physical Chemistry II
 General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
General Physical Chemistry II Lecture 13 Aleksey Kocherzhenko October 16, 2014" Last time " The Hückel method" Ø Used to study π systems of conjugated molecules" Ø π orbitals are treated separately from
Colloid Chemistry. La chimica moderna e la sua comunicazione Silvia Gross.
 Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Colloid Chemistry La chimica moderna e la sua comunicazione Silvia Gross Istituto Dipartimento di Scienze di e Scienze Tecnologie Chimiche Molecolari ISTM-CNR, Università Università degli Studi degli Studi
Solutions and Intermolecular Forces
 Solutions and Intermolecular Forces REVIEW Chemical Bonds Three basic types of bonds: Ionic Electrostatic attraction between ions Covalent Sharing of electrons Metallic Metal atoms bonded to several other
Solutions and Intermolecular Forces REVIEW Chemical Bonds Three basic types of bonds: Ionic Electrostatic attraction between ions Covalent Sharing of electrons Metallic Metal atoms bonded to several other
The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals.
 Physical Metallurgy The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals. Crystal Binding In our discussions
Physical Metallurgy The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals. Crystal Binding In our discussions
IMFA s. intermolecular forces of attraction Chez Chem, LLC All rights reserved.
 IMFA s intermolecular forces of attraction 2014 Chez Chem, LLC All rights reserved. **London Dispersion Forces Also know as Van der Waals forces A momentary non symmetrical electron distribution that can
IMFA s intermolecular forces of attraction 2014 Chez Chem, LLC All rights reserved. **London Dispersion Forces Also know as Van der Waals forces A momentary non symmetrical electron distribution that can
Atoms can form stable units called molecules by sharing electrons.
 Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule) e.g. ionic, covalent. Forces that cause the aggregation
Atoms can form stable units called molecules by sharing electrons. The formation of molecules is the result of intramolecular bonding (within the molecule) e.g. ionic, covalent. Forces that cause the aggregation
CE 530 Molecular Simulation
 1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
1 CE 530 Molecular Simulation Lecture 14 Molecular Models David A. Kofke Department of Chemical Engineering SUNY Buffalo kofke@eng.buffalo.edu 2 Review Monte Carlo ensemble averaging, no dynamics easy
Intermolecular Forces
 Intermolecular Forces Molecular Compounds The simplest molecule is H 2 : Increased electron density draws nuclei together The pair of shared electrons constitutes a covalent bond. Intermolecular Forces
Intermolecular Forces Molecular Compounds The simplest molecule is H 2 : Increased electron density draws nuclei together The pair of shared electrons constitutes a covalent bond. Intermolecular Forces
Mirror Charges, Hydrogen Bonds, And the Water-Dielectric Interface
 Mirror Charges, ydrogen Bonds, And the Water-Dielectric Interface Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ 1 Field Energy The energy E of an electric field E( x, t) is the space integral of
Mirror Charges, ydrogen Bonds, And the Water-Dielectric Interface Kevin Cahill cahill@unm.edu http://dna.phys.unm.edu/ 1 Field Energy The energy E of an electric field E( x, t) is the space integral of
Chapter 11. Intermolecular Forces, Liquids, and Solids
 Chapter 11. Intermolecular Forces, Liquids, and Solids A Molecular Comparison of Gases, Liquids, and Solids Physical properties of substances are understood in terms of kinetic-molecular theory: Gases
Chapter 11. Intermolecular Forces, Liquids, and Solids A Molecular Comparison of Gases, Liquids, and Solids Physical properties of substances are understood in terms of kinetic-molecular theory: Gases
Intermolecular forces
 Intermolecular forces World of Chemistry, 2000 Updated: August 29, 2013 The attractions of molecules to each other are known as intermolecular forces to distinguish them from intramolecular forces, such
Intermolecular forces World of Chemistry, 2000 Updated: August 29, 2013 The attractions of molecules to each other are known as intermolecular forces to distinguish them from intramolecular forces, such
BIOC 530 Fall, 2011 BIOC 530
 Fall, 2011 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic acids, control of enzymatic reactions. Please see the course
Fall, 2011 Course Information Course Description Graduate-level discussion of the structure, function, and chemistry of proteins and nucleic acids, control of enzymatic reactions. Please see the course
2.2.2 Bonding and Structure
 2.2.2 Bonding and Structure Ionic Bonding Definition: Ionic bonding is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form
2.2.2 Bonding and Structure Ionic Bonding Definition: Ionic bonding is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form
Material Properties & Characterization - Surfaces
 1) XPS Spectrum analysis: The figure below shows an XPS spectrum measured on the surface of a clean insoluble homo-polyether. Using the formulas and tables in this document, answer the following questions:
1) XPS Spectrum analysis: The figure below shows an XPS spectrum measured on the surface of a clean insoluble homo-polyether. Using the formulas and tables in this document, answer the following questions:
CS 2, HCN, BeF 2 Trigonal planar. Cl 120 BF 3, AlCl 3, SO 3, NO 3-, CO NCl 3,PF 3,ClO 3,H 3 O + ...
 Shape of molecules Name No bonding pairs No lone pairs Diagram Bond angle Examples linear 2 0 l Be l 180 2, S 2, N, Be 2 Trigonal planar 3 0 l l 120 B 3, All 3, S 3, N 3-, 2-3 B Tetrahedral 4 0 109.5 Sil
Shape of molecules Name No bonding pairs No lone pairs Diagram Bond angle Examples linear 2 0 l Be l 180 2, S 2, N, Be 2 Trigonal planar 3 0 l l 120 B 3, All 3, S 3, N 3-, 2-3 B Tetrahedral 4 0 109.5 Sil
Intermolecular Forces & Condensed Phases
 Intermolecular Forces & Condensed Phases CHEM 107 T. Hughbanks READING We will discuss some of Chapter 5 that we skipped earlier (Van der Waals equation, pp. 145-8), but this is just a segue into intermolecular
Intermolecular Forces & Condensed Phases CHEM 107 T. Hughbanks READING We will discuss some of Chapter 5 that we skipped earlier (Van der Waals equation, pp. 145-8), but this is just a segue into intermolecular
States of matter Part 1
 Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
BIOC : Homework 1 Due 10/10
 Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
Contact information: Name: Student # BIOC530 2012: Homework 1 Due 10/10 Department Email address The following problems are based on David Baker s lectures of forces and protein folding. When numerical
States of matter Part 1. Lecture 1. University of Kerbala. Hamid Alghurabi Assistant Lecturer in Pharmaceutics. Physical Pharmacy
 Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Physical pharmacy I 1. States of matter (2 Lectures) 2. Thermodynamics (2 Lectures) 3. Solution of non-electrolyte 4. Solution of electrolyte 5. Ionic equilibria 6. Buffered and isotonic solution Physical
Chapter 3. Crystal Binding
 Chapter 3. Crystal Binding Energy of a crystal and crystal binding Cohesive energy of Molecular crystals Ionic crystals Metallic crystals Elasticity What causes matter to exist in three different forms?
Chapter 3. Crystal Binding Energy of a crystal and crystal binding Cohesive energy of Molecular crystals Ionic crystals Metallic crystals Elasticity What causes matter to exist in three different forms?
POGIL 7 KEY Intermolecular Forces
 Honors Chem Block Name POGIL 7 KEY Intermolecular Forces In chemistry we talk a lot about properties of substances, since the object of chemistry is substances and their properties. After learning different
Honors Chem Block Name POGIL 7 KEY Intermolecular Forces In chemistry we talk a lot about properties of substances, since the object of chemistry is substances and their properties. After learning different
3. Solutions W = N!/(N A!N B!) (3.1) Using Stirling s approximation ln(n!) = NlnN N: ΔS mix = k (N A lnn + N B lnn N A lnn A N B lnn B ) (3.
 3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
3. Solutions Many biological processes occur between molecules in aqueous solution. In addition, many protein and nucleic acid molecules adopt three-dimensional structure ( fold ) in aqueous solution.
Intermolecular Forces
 Intermolecular Forces H covalent bond (stronger) Cl H Cl intermolecular attraction (weaker) The attractions between molecules are not nearly as strong as the covalent bonds that hold atoms together. They
Intermolecular Forces H covalent bond (stronger) Cl H Cl intermolecular attraction (weaker) The attractions between molecules are not nearly as strong as the covalent bonds that hold atoms together. They
Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia
 University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
Biochemistry 675, Lecture 8 Electrostatic Interactions
 Biochemistry 675, Lecture 8 Electrostatic Interactions Previous Classes Hydrophobic Effect Hydrogen Bonds Today: Electrostatic Interactions Reading: Handout:P.R. Bergethon & E.R. Simons, Biophysical Chemistry,
Biochemistry 675, Lecture 8 Electrostatic Interactions Previous Classes Hydrophobic Effect Hydrogen Bonds Today: Electrostatic Interactions Reading: Handout:P.R. Bergethon & E.R. Simons, Biophysical Chemistry,
The dative covalent bond acts like an ordinary covalent bond when thinking about shape so in NH 4. the shape is tetrahedral
 1.3 Bonding Definition Ionic bonding is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form ve ions. Non-metal atoms gain
1.3 Bonding Definition Ionic bonding is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form ve ions. Non-metal atoms gain
One Q partial negative, the other partial negative Ø H- bonding particularly strong. Abby Carroll 2
 Chemistry Notes v Polarity Experiment Ø Things involved Polarity Solubility Dispersion Ø Polarity Shaving cream has soap steric acid Water is polar Food coloring is polar/ionic because dissolved Like dissolves
Chemistry Notes v Polarity Experiment Ø Things involved Polarity Solubility Dispersion Ø Polarity Shaving cream has soap steric acid Water is polar Food coloring is polar/ionic because dissolved Like dissolves
Chapter 13: Phenomena
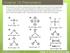 Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Chapter 1. Topic: Overview of basic principles
 Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Chapter 1 Topic: Overview of basic principles Four major themes of biochemistry I. What are living organism made from? II. How do organism acquire and use energy? III. How does an organism maintain its
Why Is Molecular Interaction Important in Our Life
 Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Why Is Molecular Interaction Important in ur Life QuLiS and Graduate School of Science iroshima University http://www.nabit.hiroshima-u.ac.jp/iwatasue/indexe.htm Suehiro Iwata Sept. 29, 2007 Department
Complicated, short range. þq 1 Q 2 /4p3 0 r (Coulomb energy) Q 2 u 2 /6(4p3 0 ) 2 ktr 4. u 2 1 u2 2 =3ð4p3 0Þ 2 ktr 6 ðkeesom energyþ
 Bonding ¼ Type of interaction Interaction energy w(r) Covalent, metallic Complicated, short range Charge charge þq 1 Q 2 /4p3 0 r (Coulomb energy) Charge dipole Qu cos q/4p3 0 r 2 Q 2 u 2 /6(4p3 0 ) 2
Bonding ¼ Type of interaction Interaction energy w(r) Covalent, metallic Complicated, short range Charge charge þq 1 Q 2 /4p3 0 r (Coulomb energy) Charge dipole Qu cos q/4p3 0 r 2 Q 2 u 2 /6(4p3 0 ) 2
Physical Chemistry - Problem Drill 01: Chemistry and Physics Review
 Physical Chemistry - Problem Drill 01: Chemistry and Physics Review No. 1 of 10 1. Chemical bonds are considered to be the interaction of their electronic structures of bonding atoms involved, with the
Physical Chemistry - Problem Drill 01: Chemistry and Physics Review No. 1 of 10 1. Chemical bonds are considered to be the interaction of their electronic structures of bonding atoms involved, with the
Why Proteins Fold. How Proteins Fold? e - ΔG/kT. Protein Folding, Nonbonding Forces, and Free Energy
 Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Why Proteins Fold Proteins are the action superheroes of the body. As enzymes, they make reactions go a million times faster. As versatile transport vehicles, they carry oxygen and antibodies to fight
Lecture: P1_Wk1_L1 IntraMolecular Interactions. Ron Reifenberger Birck Nanotechnology Center Purdue University 2012
 Lecture: IntraMolecular Interactions Distinguish between IntraMolecular (within a molecule) and InterMolecular (between molecules) Ron Reifenberger Birck Nanotechnology Center Purdue University 2012 1
Lecture: IntraMolecular Interactions Distinguish between IntraMolecular (within a molecule) and InterMolecular (between molecules) Ron Reifenberger Birck Nanotechnology Center Purdue University 2012 1
- intermolecular forces forces that exist between molecules
 Chapter 11: Intermolecular Forces, Liquids, and Solids - intermolecular forces forces that exist between molecules 11.1 A Molecular Comparison of Liquids and Solids - gases - average kinetic energy of
Chapter 11: Intermolecular Forces, Liquids, and Solids - intermolecular forces forces that exist between molecules 11.1 A Molecular Comparison of Liquids and Solids - gases - average kinetic energy of
Module17: Intermolecular Force between Surfaces and Particles. Lecture 23: Intermolecular Force between Surfaces and Particles
 Module17: Intermolecular Force between Surfaces and Particles Lecture 23: Intermolecular Force between Surfaces and Particles 1 We now try to understand the nature of spontaneous instability in a confined
Module17: Intermolecular Force between Surfaces and Particles Lecture 23: Intermolecular Force between Surfaces and Particles 1 We now try to understand the nature of spontaneous instability in a confined
Lectures 11-13: Electrostatics of Salty Solutions
 Lectures 11-13: Electrostatics of Salty Solutions Lecturer: Brigita Urbanc Office: 1-909 (E-mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_011-01/ 1 Water as
Lectures 11-13: Electrostatics of Salty Solutions Lecturer: Brigita Urbanc Office: 1-909 (E-mail: brigita@drexel.edu) Course website: www.physics.drexel.edu/~brigita/courses/biophys_011-01/ 1 Water as
Atomic Structure and Bonding. Chapter 1 Organic Chemistry, 8 th Edition John McMurry
 Atomic Structure and Bonding Chapter 1 Organic Chemistry, 8 th Edition John McMurry 1 Common Elements Groups First row Second row In most organic molecules carbon is combined with relatively few elements
Atomic Structure and Bonding Chapter 1 Organic Chemistry, 8 th Edition John McMurry 1 Common Elements Groups First row Second row In most organic molecules carbon is combined with relatively few elements
READING. Review of Intermolecular Forces & Liquids (Chapter 12) Ion-Ion Forces. Ion-Dipole Energies
 Review of Intermolecular Forces & Liquids (Chapter 12) CEM 102 T. ughbanks READIG We will very briefly review the underlying concepts from Chapters 12 on intermolecular forces since it is relevant to Chapter
Review of Intermolecular Forces & Liquids (Chapter 12) CEM 102 T. ughbanks READIG We will very briefly review the underlying concepts from Chapters 12 on intermolecular forces since it is relevant to Chapter
DEFINITION. The electrostatic force of attraction between oppositely charged ions
 DEFINITION The electrostatic force of attraction between oppositely charged ions Usually occurs when a metal bonds with a non-metal Ions are formed by complete electron transfer from the metal atoms to
DEFINITION The electrostatic force of attraction between oppositely charged ions Usually occurs when a metal bonds with a non-metal Ions are formed by complete electron transfer from the metal atoms to
BIBC 100. Structural Biochemistry
 BIBC 100 Structural Biochemistry http://classes.biology.ucsd.edu/bibc100.wi14 Papers- Dialogue with Scientists Questions: Why? How? What? So What? Dialogue Structure to explain function Knowledge Food
BIBC 100 Structural Biochemistry http://classes.biology.ucsd.edu/bibc100.wi14 Papers- Dialogue with Scientists Questions: Why? How? What? So What? Dialogue Structure to explain function Knowledge Food
All-atom Molecular Mechanics. Trent E. Balius AMS 535 / CHE /27/2010
 All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
All-atom Molecular Mechanics Trent E. Balius AMS 535 / CHE 535 09/27/2010 Outline Molecular models Molecular mechanics Force Fields Potential energy function functional form parameters and parameterization
Surface interactions part 1: Van der Waals Forces
 CHEM-E150 Interfacial Phenomena in Biobased Systems Surface interactions part 1: Van der Waals Forces Monika Österberg Spring 018 Content Colloidal stability van der Waals Forces Surface Forces and their
CHEM-E150 Interfacial Phenomena in Biobased Systems Surface interactions part 1: Van der Waals Forces Monika Österberg Spring 018 Content Colloidal stability van der Waals Forces Surface Forces and their
Polar molecules vs. Nonpolar molecules A molecule with separate centers of positive and negative charge is a polar molecule.
 CHM 123 Chapter 8 8.5 8.6 Polar covalent Bonds and Dipole moments Depending on the relative electronegativities of the two atoms sharing electrons, there may be partial transfer of electron density from
CHM 123 Chapter 8 8.5 8.6 Polar covalent Bonds and Dipole moments Depending on the relative electronegativities of the two atoms sharing electrons, there may be partial transfer of electron density from
Liquids and Solids The Condensed States of Matter
 Liquids and Solids The Condensed States of Matter AP Chemistry Ms. Grobsky Where We Have Been And Where We Are Going In the last few chapters, we saw that atoms can form stable units called molecules by
Liquids and Solids The Condensed States of Matter AP Chemistry Ms. Grobsky Where We Have Been And Where We Are Going In the last few chapters, we saw that atoms can form stable units called molecules by
Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation
 Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation Ion:ion interactions What is the free energy of ion:ion interactions ΔG i-i? Consider an ion in a solution
Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation Ion:ion interactions What is the free energy of ion:ion interactions ΔG i-i? Consider an ion in a solution
Saba Al Fayoumi. Tamer Barakat. Dr. Mamoun Ahram + Dr. Diala Abu-Hassan
 1 Saba Al Fayoumi Tamer Barakat Dr. Mamoun Ahram + Dr. Diala Abu-Hassan What is BIOCHEMISTRY??? Biochemistry = understanding life Chemical reactions are what makes an organism (An organism is simply atoms
1 Saba Al Fayoumi Tamer Barakat Dr. Mamoun Ahram + Dr. Diala Abu-Hassan What is BIOCHEMISTRY??? Biochemistry = understanding life Chemical reactions are what makes an organism (An organism is simply atoms
LECTURE 2 STRUCTURE AND PROPERTIES OF ORGANIC MOLECULES
 LECTURE 2 STRUCTURE AND PROPERTIES OF ORGANIC MOLECULES 1. Atomic wave functions and orbitals. LCAO. The important thing to know is that atomic orbitals are represented by wave functions, and they have
LECTURE 2 STRUCTURE AND PROPERTIES OF ORGANIC MOLECULES 1. Atomic wave functions and orbitals. LCAO. The important thing to know is that atomic orbitals are represented by wave functions, and they have
Biomolecules: The Molecules of Life. 1.2 Biomolecules: The Molecules of Life. Covalent bonding alternatives
 Biomolecules: The Molecules of Life What property unites H, O, C and N and renders these atoms so appropriate to the chemistry of life? Answer: Their ability to form covalent bonds by electron-pair sharing.
Biomolecules: The Molecules of Life What property unites H, O, C and N and renders these atoms so appropriate to the chemistry of life? Answer: Their ability to form covalent bonds by electron-pair sharing.
CHAPTER 2 INTERATOMIC FORCES. atoms together in a solid?
 CHAPTER 2 INTERATOMIC FORCES What kind of force holds the atoms together in a solid? Interatomic Binding All of the mechanisms which cause bonding between the atoms derive from electrostatic interaction
CHAPTER 2 INTERATOMIC FORCES What kind of force holds the atoms together in a solid? Interatomic Binding All of the mechanisms which cause bonding between the atoms derive from electrostatic interaction
Chapters 11 and 12: Intermolecular Forces of Liquids and Solids
 1 Chapters 11 and 12: Intermolecular Forces of Liquids and Solids 11.1 A Molecular Comparison of Liquids and Solids The state of matter (Gas, liquid or solid) at a particular temperature and pressure depends
1 Chapters 11 and 12: Intermolecular Forces of Liquids and Solids 11.1 A Molecular Comparison of Liquids and Solids The state of matter (Gas, liquid or solid) at a particular temperature and pressure depends
1. I can use Collision Theory to explain the effects of concentration, particle size, temperature, and collision geometry on reaction rates.
 Chemical Changes and Structure Learning Outcomes SECTION 1 Controlling the Rate. Subsection (a) Collision Theory 1. I can use Collision Theory to explain the effects of concentration, particle size, temperature,
Chemical Changes and Structure Learning Outcomes SECTION 1 Controlling the Rate. Subsection (a) Collision Theory 1. I can use Collision Theory to explain the effects of concentration, particle size, temperature,
When intermolecular forces are strong, the atoms, molecules, or ions are strongly attracted to each other, and draw closer together.
 INTERMOLECULAR FORCES: THE FORCE BEHIND VARIOUS PROPERTIES WHY? Intermolecular forces are largely responsible for the properties of affinity, solubility, volatility, melting/ boiling point, and viscosity.
INTERMOLECULAR FORCES: THE FORCE BEHIND VARIOUS PROPERTIES WHY? Intermolecular forces are largely responsible for the properties of affinity, solubility, volatility, melting/ boiling point, and viscosity.
Phys 102 Lecture 2 Coulomb s Law & Electric Dipoles
 Phys 102 Lecture 2 Coulomb s Law & Electric Dipoles 1 Today we will... Get practice using Coulomb s law & vector addition Learn about electric dipoles Apply these concepts! Molecular interactions Polar
Phys 102 Lecture 2 Coulomb s Law & Electric Dipoles 1 Today we will... Get practice using Coulomb s law & vector addition Learn about electric dipoles Apply these concepts! Molecular interactions Polar
SAM Teachers Guide Intermolecular Forces Overview Learning Objectives: Possible student pre/misconceptions
 SAM Teachers Guide Intermolecular Forces Overview This unit focuses on the attractive forces felt between molecules. Both London Dispersion and Dipole Dipole intermolecular attractions will be discussed
SAM Teachers Guide Intermolecular Forces Overview This unit focuses on the attractive forces felt between molecules. Both London Dispersion and Dipole Dipole intermolecular attractions will be discussed
Molecular Geometries. Molecular Geometries. Remember that covalent bonds are formed when electrons in atomic orbitals are shared between two nuclei.
 Molecular Geometries Lewis dot structures are very useful in determining the types of bonds in a molecule, but they may not provide the best insight into the spatial geometry of a molecule, i.e., how the
Molecular Geometries Lewis dot structures are very useful in determining the types of bonds in a molecule, but they may not provide the best insight into the spatial geometry of a molecule, i.e., how the
16 years ago TODAY (9/11) at 8:46, the first tower was hit at 9:03, the second tower was hit. Lecture 2 (9/11/17)
 16 years ago TODAY (9/11) at 8:46, the first tower was hit at 9:03, the second tower was hit By Anthony Quintano - https://www.flickr.com/photos/quintanomedia/15071865580, CC BY 2.0, https://commons.wikimedia.org/w/index.php?curid=38538291
16 years ago TODAY (9/11) at 8:46, the first tower was hit at 9:03, the second tower was hit By Anthony Quintano - https://www.flickr.com/photos/quintanomedia/15071865580, CC BY 2.0, https://commons.wikimedia.org/w/index.php?curid=38538291
MOLECULAR INTERACTIONS NOTES
 - 1 - MOLECULAR INTERACTIONS NOTES Summary of Fundamental Molecular Interactions Q1Q Ion-Ion U ( r) = 4 r πεε o µ Q cosθ U ( r) = 4πεε o r µ 1µ U ( r) 3 r µ 1 µ U ( r) 6 r µ 1 α U ( r) 6 r Ion-Dipole Dipole-Dipole
- 1 - MOLECULAR INTERACTIONS NOTES Summary of Fundamental Molecular Interactions Q1Q Ion-Ion U ( r) = 4 r πεε o µ Q cosθ U ( r) = 4πεε o r µ 1µ U ( r) 3 r µ 1 µ U ( r) 6 r µ 1 α U ( r) 6 r Ion-Dipole Dipole-Dipole
Chapter 11. Liquids and Intermolecular Forces
 Chapter 11. Liquids and Intermolecular Forces 11.1 A Molecular Comparison of Gases, Liquids, and Solids Gases are highly compressible and assume the shape and volume of their container. Gas molecules are
Chapter 11. Liquids and Intermolecular Forces 11.1 A Molecular Comparison of Gases, Liquids, and Solids Gases are highly compressible and assume the shape and volume of their container. Gas molecules are
Atomic structure & interatomic bonding. Chapter two
 Atomic structure & interatomic bonding Chapter two 1 Atomic Structure Mass Charge Proton 1.67 х 10-27 kg + 1.60 х 10-19 C Neutron 1.67 х 10-27 kg Neutral Electron 9.11 х 10-31 kg - 1.60 х 10-19 C Electron
Atomic structure & interatomic bonding Chapter two 1 Atomic Structure Mass Charge Proton 1.67 х 10-27 kg + 1.60 х 10-19 C Neutron 1.67 х 10-27 kg Neutral Electron 9.11 х 10-31 kg - 1.60 х 10-19 C Electron
Step 1: Solute particles must separate from each other. Since energy must be absorbed to overcome the forces of attraction between solute particles,
 Step 1: Solute particles must separate from each other. Since energy must be absorbed to overcome the forces of attraction between solute particles, this process is endothermic. Step 2: Solvent particles
Step 1: Solute particles must separate from each other. Since energy must be absorbed to overcome the forces of attraction between solute particles, this process is endothermic. Step 2: Solvent particles
Chapter 13: Phenomena
 Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Chapter 13: Phenomena Phenomena: Scientists measured the bond angles of some common molecules. In the pictures below each line represents a bond that contains 2 electrons. If multiple lines are drawn together
Intermolecular Forces and Monte-Carlo Integration 열역학특수연구
 Intermolecular Forces and Monte-Carlo Integration 열역학특수연구 2003.3.28 Source of the lecture note. J.M.Prausnitz and others, Molecular Thermodynamics of Fluid Phase Equiliria Atkins, Physical Chemistry Lecture
Intermolecular Forces and Monte-Carlo Integration 열역학특수연구 2003.3.28 Source of the lecture note. J.M.Prausnitz and others, Molecular Thermodynamics of Fluid Phase Equiliria Atkins, Physical Chemistry Lecture
Water, water everywhere,; not a drop to drink. Consumption resulting from how environment inhabited Deforestation disrupts water cycle
 Chapter 3 Water: The Matrix of Life Overview n n n Water, water everywhere,; not a drop to drink Only 3% of world s water is fresh How has this happened Consumption resulting from how environment inhabited
Chapter 3 Water: The Matrix of Life Overview n n n Water, water everywhere,; not a drop to drink Only 3% of world s water is fresh How has this happened Consumption resulting from how environment inhabited
Ionic Bonding. Example: Atomic Radius: Na (r = 0.192nm) Cl (r = 0.099nm) Ionic Radius : Na (r = 0.095nm) Cl (r = 0.181nm)
 Ionic Bonding Ion: an atom or molecule that gains or loses electrons (acquires an electrical charge). Atoms form cations (+charge), when they lose electrons, or anions (- charge), when they gain electrons.
Ionic Bonding Ion: an atom or molecule that gains or loses electrons (acquires an electrical charge). Atoms form cations (+charge), when they lose electrons, or anions (- charge), when they gain electrons.
Primary bonding: e- are transferred or shared Strong ( KJ/mol or 1-10 ev/atom) Secondary Bonding: no e -
 Types of Bondings Primary bonding: e- are transferred or shared Strong (100-1000 KJ/mol or 1-10 ev/atom) Ionic: Strong Coulomb interaction among negative atoms (have an extra electron each) and positive
Types of Bondings Primary bonding: e- are transferred or shared Strong (100-1000 KJ/mol or 1-10 ev/atom) Ionic: Strong Coulomb interaction among negative atoms (have an extra electron each) and positive
Topic 04 Bonding 4.3 Intermolecular Forces. IB Chemistry T04D05
 Topic 04 Bonding 4.3 Intermolecular orces IB Chemistry T04D05 Intermolecular orces 2 hrs 4.3.1 Describe the types of intermolecular forces (attractions between molecules that have temporary dipoles, permanent
Topic 04 Bonding 4.3 Intermolecular orces IB Chemistry T04D05 Intermolecular orces 2 hrs 4.3.1 Describe the types of intermolecular forces (attractions between molecules that have temporary dipoles, permanent
Definition: An Ionic bond is the electrostatic force of attraction between oppositely charged ions formed by electron transfer.
 3 Bonding Definition An Ionic bond is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form +ve ions. on-metal atoms gain
3 Bonding Definition An Ionic bond is the electrostatic force of attraction between oppositely charged ions formed by electron transfer. Metal atoms lose electrons to form +ve ions. on-metal atoms gain
Kinetic Molecular Theory of Gases used to account for Ideal Gas Behavior when gases approach high temperatures and low pressures
 LIQUIDS AND SOLIDS Kinetic Molecular Theory of Gases used to account for Ideal Gas Behavior when gases approach high temperatures and low pressures GASES are very different from solids and liquids. We
LIQUIDS AND SOLIDS Kinetic Molecular Theory of Gases used to account for Ideal Gas Behavior when gases approach high temperatures and low pressures GASES are very different from solids and liquids. We
Other Cells. Hormones. Viruses. Toxins. Cell. Bacteria
 Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Other Cells Hormones Viruses Toxins Cell Bacteria ΔH < 0 reaction is exothermic, tells us nothing about the spontaneity of the reaction Δ H > 0 reaction is endothermic, tells us nothing about the spontaneity
Intermolecular forces are classified into four major types.
 Intermolecular forces are classified into four major types. 1. Ion-dipole: IMF s that occur between neighboring an ion solution and a polar molecule (dipole) also in solution. Na+ 2. Dipole-dipole: IMF
Intermolecular forces are classified into four major types. 1. Ion-dipole: IMF s that occur between neighboring an ion solution and a polar molecule (dipole) also in solution. Na+ 2. Dipole-dipole: IMF
States of Matter. Intermolecular Forces. The States of Matter. Intermolecular Forces. Intermolecular Forces
 Intermolecular Forces Have studied INTRAmolecular forces the forces holding atoms together to form compounds. Now turn to forces between molecules INTERmolecular forces. Forces between molecules, between
Intermolecular Forces Have studied INTRAmolecular forces the forces holding atoms together to form compounds. Now turn to forces between molecules INTERmolecular forces. Forces between molecules, between
Rama Abbady. Zina Smadi. Diala Abu-Hassan
 1 Rama Abbady Zina Smadi Diala Abu-Hassan (00:00) (10:00) Types of Molecules in the Cell 1. Water Molecules: a large portion of the cell mass is water (70% of total cell mass). 2. Organic molecules (carbon
1 Rama Abbady Zina Smadi Diala Abu-Hassan (00:00) (10:00) Types of Molecules in the Cell 1. Water Molecules: a large portion of the cell mass is water (70% of total cell mass). 2. Organic molecules (carbon
Structural Bioinformatics (C3210) Molecular Mechanics
 Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Structural Bioinformatics (C3210) Molecular Mechanics How to Calculate Energies Calculation of molecular energies is of key importance in protein folding, molecular modelling etc. There are two main computational
Supplemental Activities. Module: States of Matter. Section: Intermolecular Forces - Key
 Supplemental Activities Module: States of Matter Section: Intermolecular Forces - Key Electrostatic Forces ACTIVITY 1 The purpose of this activity is to practice recognizing the nature of the forces important
Supplemental Activities Module: States of Matter Section: Intermolecular Forces - Key Electrostatic Forces ACTIVITY 1 The purpose of this activity is to practice recognizing the nature of the forces important
Lecture 11: Protein Folding & Stability
 Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall Protein Folding: What we know. Protein Folding
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture C2 Microscopic to Macroscopic, Part 2: Intermolecular Interactions. Let's get together.
 Lecture C2 Microscopic to Macroscopic, Part 2: Intermolecular Interactions Let's get together. Most gases are NOT ideal except at very low pressures: Z=1 for ideal gases Intermolecular interactions come
Lecture C2 Microscopic to Macroscopic, Part 2: Intermolecular Interactions Let's get together. Most gases are NOT ideal except at very low pressures: Z=1 for ideal gases Intermolecular interactions come
Bioengineering 215. An Introduction to Molecular Dynamics for Biomolecules
 Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
Bioengineering 215 An Introduction to Molecular Dynamics for Biomolecules David Parker May 18, 2007 ntroduction A principal tool to study biological molecules is molecular dynamics simulations (MD). MD
The Chemistry and Energy of Life
 2 The Chemistry and Energy of Life Chapter 2 The Chemistry and Energy of Life Key Concepts 2.1 Atomic Structure Is the Basis for Life s Chemistry 2.2 Atoms Interact and Form Molecules 2.3 Carbohydrates
2 The Chemistry and Energy of Life Chapter 2 The Chemistry and Energy of Life Key Concepts 2.1 Atomic Structure Is the Basis for Life s Chemistry 2.2 Atoms Interact and Form Molecules 2.3 Carbohydrates
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall How do we go from an unfolded polypeptide chain to a
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
CHAPTER 3. Crystallography
 CHAPTER 3 Crystallography Atomic Structure Atoms are made of Protons: mass 1.00728 amu, +1 positive charge Neutrons: mass of 1.00867 amu, neutral Electrons: mass of 0.00055 amu, -1 negative charge (1 amu
CHAPTER 3 Crystallography Atomic Structure Atoms are made of Protons: mass 1.00728 amu, +1 positive charge Neutrons: mass of 1.00867 amu, neutral Electrons: mass of 0.00055 amu, -1 negative charge (1 amu
File ISM04. Properties of Colloids I
 File ISM04 Properties of Colloids I 1 Colloids Small things, but much bigger than a molecule Background: Foundations of Colloid Science Vol I & II, R.J. Hunter Physical Chemistry, P.W. Atkins, Chapter
File ISM04 Properties of Colloids I 1 Colloids Small things, but much bigger than a molecule Background: Foundations of Colloid Science Vol I & II, R.J. Hunter Physical Chemistry, P.W. Atkins, Chapter
