Genomic Medicine HT 512. Data representation, transformation & modeling in genomics
|
|
|
- Ursula Stevenson
- 5 years ago
- Views:
Transcription
1 Harvard-MIT Division of Health Sciences and Technology HST.512: Genomic Medicine Prof. Alvin T.Kho Genomic Medicine HT 512 Data representation, transformation & modeling in genomics Lecture 11, Mar 18, 2004 Alvin T. Kho Children's Hospital Boston Dana-Farber Cancer Institute
2 Lecture Outline 2 prototypical study design Background concepts 2-way comparison Noise Time series Replicates, reproducibility Data representation (DR) Normalization What is DR? Fold Measurement device to Miscellany spreadsheet Dimensionality - scales Transformations / Changes-of-coordinates
3 Prototype study 1: 2-way comparison Molecular differences in adipocytes of type-2 Diabetes vs Normal humans. 27 type-2 diabetes patients. 11 normal patients. arrays. Pre-study reality check: Stratification clinical phenotype. Partial math formulation: Let D = chip data of j-th Diabetic. N j = j-th j Normal. D, N j are vectors/matrices whose dimensions depend j upon # genes/parameters/variables measured per sample. Next steps...
4 Prototype study 2: Time series RNA expression profile of a developing whole mouse pancreas at time points P1 to P60. Pre-study reality check: Stratification histology, regulation. Partial math formulation: Let T = chip data of j-th developmental j stage are vectors/matrices whose dimensions depend upon # genes/parameters measured per sample. Next steps... patterns
5 What is DR? A mathematical reformulation of a scientific, real-world problem. Mapping observations and measurements to a set of symbols (typically, real numbers) that can be acted upon by an algebra. The form of representing the data values in the integrated dataset, including any numerical type conversion or reclassification that was performed. Usually this will involve only type conversions, although occasionally actual numerical changes may have been necessary. The units and precision are also indicated. [ Something to do with database annotation and standards. Multimedia: Charts, graphs, plots.
6 Device to Spreadsheet: Know measuring device Understand general principles of measurement Relevance of scanner / device settings Phosphor imager mechanism 1-/2-channel (competitive hybridization)? Image to Device to Numbers / Spreadsheet Internal image analytic software? Pre-processing? Spot evaluation: Statistics, microscopic diffusion/thermodynamics Units / dimensions of the device output
7 DR: Dimensionality/scales; Transformations Typical dimensionality/scale 2-channel readout is fold/ratio = dimensionless 1-channel = absolute or relative intensity units Importance of dimensionality/scale in large datasets Different math techniques apply Guides formulation of null hypothetic distribution / state Gamma distributions for radiation measurements Power/scaling laws for gross error detection, e.g., Zipf's Why perform DR transformations? 1. Simplify mathematical manipulation. 2. Reveal existing intrinsic geometries in the data.
8 DR: Why transform data 1 1. Simplify mathematical manipulation Writing data/problem on paper, apply basic math rules All spreadsheets are essentially matrix - subject to formal math operations/theorems (linear algebra). Exp 1 Exp 2 Exp 3 Exp M Gene 1 G 1,1 G 1,2 G 1,3... G 1,M Gene 2 G 2,1 G 2,2 G 2,3... G 2,M Gene 3 G 3,1 G 3,2 G 3,3... G 3,M : : : : : : Gene N G N,1 G N,2 G N,3... G N,M Warning: Heterogeneity of matrix entries. Blind application of math, e.g., adding apples and oranges.
9 DR: Why transform data 2 2. Reveal intrinsic geometries in the data Q: What's meant by intrinsic? geometries? A: Data = set of numbers that may contain internal orderings or structures, explicit or implicit Explicit: A priori gene, sample labels / relations. Implicit: Higher-order gene-sample relations. Clues to existence of relational structures: Numeric changes, patterning, that in a graphical/geometric setting may become more obvious/intuitive. High dimensional space. Warning: Know prior assumptions. Explicit & Implicit. No study / analysis is ever hypothesis-free.
10 DR: Model idea Modeling summary diagram: D R Physical Quantity Measurab le Quanti ty / Numbers Physical System / Phenomenon Model Coherent Relati on Perturbations Numeric patterns / fluctuations
11 DR: Transformations example 0 Example 0: 2 diff patient populations X O. Each patient has 2 gene measurements: G1, G2. Principal component analysis (PCA) -> G1-G2 is discriminant
12 DR: Transformations example 1 Example 1: Simulated 2 diff gene populations R, B in 3 sample conditions genes in each population.
13 DR: Transformations example 1 Example 1: R, B in new coordinates after PCA of condition axes.
14 DR: Transformations example 2 Example 2: Prototype study 2 pancreas development time series. Principal component (PC) analysis = an affine change-ofcoordinate. Similarity/distance in Euclidean/L 2 sense. Sample-wise (as is): CLT-scaled genes Sample-wise (PC): CLT-scaled genes Gene-wise (as is): CLT-scaled samples Gene-wise (PC): CLT-scaled samples CLT ~ Central Limit theorem normalization -> mean 0, var 1. Later.
15 DR: Transformations example 2 Sample-wise (as is): CLT-scaled genes
16 DR: Transformations example 2 Sample-wise (PC): CLT-scaled genes Profile Profile Profile
17 DR: Transformations example 2 Gene-wise (as is): CLT-scaled samples
18 DR: Transformations example 2 Gene-wise (PC): CLT-scaled samples
19 DR: Transformations example 3 App li cat ion in sequence ana lys is: {A,T,C,G} -> {0,1,2,3} -> Four ier Example 3: Fourier decomposition. Individuals and sum of 3 sinosoids in freq space.
20 DR: Transformations summary Old Common transformations: x = a PCA (Euclidean) - finite bases. Rotation/translation. New j j j Fourier (Euclidean) - infinite bases (localize freq domain). Signal decomposed into sinusoids. Periodic boundary conditions. Wavelet (Euclidean) - infinite bases (localize time domain). Signal decomposed by discretized amplitudinal range. Different approaches emphasize different geometric/relational structures within the data. There is almost always a geometric interpretation. Secondary uses: Feature reduction, de-noiseing, etc. Basis
21 Make In Noise/Reproducibility: What is noise? Axiom 1 Nature makes no leaps (Tissot-Coke-Leibniz). Continuity of physical phenomena at a macroscopic level. Example separate measurements of room temperature within a 1- min interval at different locations in the room. High likelihood that measurements are not all identical. Working definition of Noise a narrow sense, noise is a/ny measurable divergence from Axiom 1, or more generally any applicable axiom, in a studied system. In ideal situations, math theorems apply: Central Limit, Large Numbers
22 Noise/Reproducibility: What is a replicate? What is a replicate?... a repeated measurement? Need a reference system. Grades of being a replicate. 3 cases: Separate RNA samples from pancreas of 2 ( normal ) mice: Age, gender, weight. RNA sample from 1 mouse pancreas, split into 2, aliquots. RNA pooled from 3 ( normal ) mice pancreas, and split into 2, aliquots. POOL
23 Noise/Reproducibility: Replicates & normalization Definition of replicate will have implication on noise definition. Biological versus measurement variation. Warning: Over-restrictive definition of replicate may hinder generalizability of result to larger population. Despite taking all precautions, it is unlikely that replicate assays will be numerically identical. One never steps into the same river twice. Normalization: Informally, averaging out differences between replicate assays.
24 Noise/Reproducibility: Reproducibility example E12 versus...
25 Noise/Reproducibility: Normalization To normalize or not? A priori assumptions about how system behaves. Common normalization techniques, a vector x against reference r CLT (central limit theorem) scaling: x -> (x mean(x))/std(x) What happens in normalized data/vector? Linear regression: x -> (x - a )/a 0 1 where a is the y-intercept, and a the slope of the regression of 0 1 x against reference r. What happens in normalized data/vector?
26 Fold Fold: While conventional in PCR/blots, the fold may not make sense in a 1-channel microarray setting. How to calculate? Limits? E.g., fold A = {-21.2, 14.9, -3.7} vs. B = {541.3, 596.6, 551.1}. Fold = ArithmeticAverage(A)/ArithmeticAverage(B); or GeometricAverage (A)/GeometricAverage(B)?
27 Outline: Recall 2 prototypical study design Background 2-way comparison Noise Time series Replicates, reproducibility Data representation (DR) Normalization What is DR? Fold Measurement device to Miscellany spreadsheet Dimensionality - scales Transformations / Changes-of-coordinates
28 A miscellany Our discussion so far makes nominal reference to biology. Approaches are general & apply in almost any setting. Math only provides the tools to biological discovery. Key point: The underlying biology is the point (at least our understanding of it). Lose this and the whole enterprise becomes purely technology/method driven. Biology dictates Experiment design Appropriate measure/similarity space to formulate the dual mathematical problem - representation, modeling. Reading and making sense of the model outcome. Corroboration with observed/measured phenomenon? No study is hypothesis free. Know your prior assumptions.
29 References/Epilogue Pattern Classification. 2nd ed. Duda, Hart & Stork. Wiley-Interscience, Applied Multivariate Statistical Analysis. Johnson & Wichern. Prentic-Hall, The discoveries that one can make with the microscope amount to very little, for one sees with the mind's eye and without the microscope, the real existence of all these little beings. George-Louis Leclerc Comte de Buffon,
Module 4 (Functional Genomics/FG) Lecture 1 recall
 Harvard-MIT Division of Health Sciences and Technology HST.508: Quantitative Genomics, Fall 2005 Instructors: Leonid Mirny, Robert Berwick, Alvin Kho, Isaac Kohane Module 4 (Functional Genomics/FG) Lecture
Harvard-MIT Division of Health Sciences and Technology HST.508: Quantitative Genomics, Fall 2005 Instructors: Leonid Mirny, Robert Berwick, Alvin Kho, Isaac Kohane Module 4 (Functional Genomics/FG) Lecture
Expression arrays, normalization, and error models
 1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
1 Epression arrays, normalization, and error models There are a number of different array technologies available for measuring mrna transcript levels in cell populations, from spotted cdna arrays to in
Relations in epidemiology-- the need for models
 Plant Disease Epidemiology REVIEW: Terminology & history Monitoring epidemics: Disease measurement Disease intensity: severity, incidence,... Types of variables, etc. Measurement (assessment) of severity
Plant Disease Epidemiology REVIEW: Terminology & history Monitoring epidemics: Disease measurement Disease intensity: severity, incidence,... Types of variables, etc. Measurement (assessment) of severity
4 Bias-Variance for Ridge Regression (24 points)
 Implement Ridge Regression with λ = 0.00001. Plot the Squared Euclidean test error for the following values of k (the dimensions you reduce to): k = {0, 50, 100, 150, 200, 250, 300, 350, 400, 450, 500,
Implement Ridge Regression with λ = 0.00001. Plot the Squared Euclidean test error for the following values of k (the dimensions you reduce to): k = {0, 50, 100, 150, 200, 250, 300, 350, 400, 450, 500,
Inferring Transcriptional Regulatory Networks from High-throughput Data
 Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Inferring Transcriptional Regulatory Networks from High-throughput Data Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday 12:00-1:20
Design of Microarray Experiments. Xiangqin Cui
 Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Design of Microarray Experiments Xiangqin Cui Experimental design Experimental design: is a term used about efficient methods for planning the collection of data, in order to obtain the maximum amount
Course content (will be adapted to the background knowledge of the class):
 Biomedical Signal Processing and Signal Modeling Lucas C Parra, parra@ccny.cuny.edu Departamento the Fisica, UBA Synopsis This course introduces two fundamental concepts of signal processing: linear systems
Biomedical Signal Processing and Signal Modeling Lucas C Parra, parra@ccny.cuny.edu Departamento the Fisica, UBA Synopsis This course introduces two fundamental concepts of signal processing: linear systems
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
microrna pseudo-targets
 microrna pseudo-targets Natalia Pinzón Restrepo Institut de Génétique Humaine (CNRS), Montpellier, France August, 2012 microrna target identification mir: target: 2 7 5 N NNNNNNNNNNNNNN NNNNNN 3 the seed
microrna pseudo-targets Natalia Pinzón Restrepo Institut de Génétique Humaine (CNRS), Montpellier, France August, 2012 microrna target identification mir: target: 2 7 5 N NNNNNNNNNNNNNN NNNNNN 3 the seed
Computational Biology Course Descriptions 12-14
 Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Combined Science Biology Academic Overview
 Combined Science Biology Academic Overview 2018-19 Science Term 1.1 Term 1.2 Term 2.1 Term 2.2 Term 3.1 Term 3.1 Year 9 Key Concepts in Biology Key Concepts in Biology cont. Cells & Control Cells & Control
Combined Science Biology Academic Overview 2018-19 Science Term 1.1 Term 1.2 Term 2.1 Term 2.2 Term 3.1 Term 3.1 Year 9 Key Concepts in Biology Key Concepts in Biology cont. Cells & Control Cells & Control
ECE521 week 3: 23/26 January 2017
 ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
High-Throughput Sequencing Course
 High-Throughput Sequencing Course DESeq Model for RNA-Seq Biostatistics and Bioinformatics Summer 2017 Outline Review: Standard linear regression model (e.g., to model gene expression as function of an
High-Throughput Sequencing Course DESeq Model for RNA-Seq Biostatistics and Bioinformatics Summer 2017 Outline Review: Standard linear regression model (e.g., to model gene expression as function of an
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
ALGEBRA GRADE 7 MINNESOTA ACADEMIC STANDARDS CORRELATED TO MOVING WITH MATH. Part B Student Book Skill Builders (SB)
 MINNESOTA ACADEMIC STANDARDS CORRELATED TO MOVING WITH MATH ALGEBRA GRADE 7 NUMBER AND OPERATION Read, write, represent and compare positive and negative rational numbers, expressed as integers, fractions
MINNESOTA ACADEMIC STANDARDS CORRELATED TO MOVING WITH MATH ALGEBRA GRADE 7 NUMBER AND OPERATION Read, write, represent and compare positive and negative rational numbers, expressed as integers, fractions
Lecture 13. Principal Component Analysis. Brett Bernstein. April 25, CDS at NYU. Brett Bernstein (CDS at NYU) Lecture 13 April 25, / 26
 Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Principal Component Analysis Brett Bernstein CDS at NYU April 25, 2017 Brett Bernstein (CDS at NYU) Lecture 13 April 25, 2017 1 / 26 Initial Question Intro Question Question Let S R n n be symmetric. 1
Mathematical Biology - Lecture 1 - general formulation
 Mathematical Biology - Lecture 1 - general formulation course description Learning Outcomes This course is aimed to be accessible both to masters students of biology who have a good understanding of the
Mathematical Biology - Lecture 1 - general formulation course description Learning Outcomes This course is aimed to be accessible both to masters students of biology who have a good understanding of the
Linear Regression (1/1/17)
 STA613/CBB540: Statistical methods in computational biology Linear Regression (1/1/17) Lecturer: Barbara Engelhardt Scribe: Ethan Hada 1. Linear regression 1.1. Linear regression basics. Linear regression
STA613/CBB540: Statistical methods in computational biology Linear Regression (1/1/17) Lecturer: Barbara Engelhardt Scribe: Ethan Hada 1. Linear regression 1.1. Linear regression basics. Linear regression
robustness: revisting the significance of mirna-mediated regulation
 : revisting the significance of mirna-mediated regulation Hervé Seitz IGH (CNRS), Montpellier, France October 13, 2012 microrna target identification .. microrna target identification mir: target: 2 7
: revisting the significance of mirna-mediated regulation Hervé Seitz IGH (CNRS), Montpellier, France October 13, 2012 microrna target identification .. microrna target identification mir: target: 2 7
Class 4: Classification. Quaid Morris February 11 th, 2011 ML4Bio
 Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Class 4: Classification Quaid Morris February 11 th, 211 ML4Bio Overview Basic concepts in classification: overfitting, cross-validation, evaluation. Linear Discriminant Analysis and Quadratic Discriminant
Mini Lecture 2.1 Introduction to Functions
 Mini Lecture.1 Introduction to Functions 1. Find the domain and range of a relation.. Determine whether a relation is a function. 3. Evaluate a function. 1. Find the domain and range of the relation. a.
Mini Lecture.1 Introduction to Functions 1. Find the domain and range of a relation.. Determine whether a relation is a function. 3. Evaluate a function. 1. Find the domain and range of the relation. a.
CURRICULUM MAP. Course/Subject: Honors Math I Grade: 10 Teacher: Davis. Month: September (19 instructional days)
 Month: September (19 instructional days) Numbers, Number Systems and Number Relationships Standard 2.1.11.A: Use operations (e.g., opposite, reciprocal, absolute value, raising to a power, finding roots,
Month: September (19 instructional days) Numbers, Number Systems and Number Relationships Standard 2.1.11.A: Use operations (e.g., opposite, reciprocal, absolute value, raising to a power, finding roots,
Bioinformatics. Genotype -> Phenotype DNA. Jason H. Moore, Ph.D. GECCO 2007 Tutorial / Bioinformatics.
 Bioinformatics Jason H. Moore, Ph.D. Frank Lane Research Scholar in Computational Genetics Associate Professor of Genetics Adjunct Associate Professor of Biological Sciences Adjunct Associate Professor
Bioinformatics Jason H. Moore, Ph.D. Frank Lane Research Scholar in Computational Genetics Associate Professor of Genetics Adjunct Associate Professor of Biological Sciences Adjunct Associate Professor
Prentice Hall Mathematics, Geometry 2009 Correlated to: Connecticut Mathematics Curriculum Framework Companion, 2005 (Grades 9-12 Core and Extended)
 Grades 9-12 CORE Algebraic Reasoning: Patterns And Functions GEOMETRY 2009 Patterns and functional relationships can be represented and analyzed using a variety of strategies, tools and technologies. 1.1
Grades 9-12 CORE Algebraic Reasoning: Patterns And Functions GEOMETRY 2009 Patterns and functional relationships can be represented and analyzed using a variety of strategies, tools and technologies. 1.1
Algebra I Study Topics
 This Algebra I Study Guide contains clear, straight-forward problems that represent the topics covered in a complete Algebra I course. After completing the study guide without a calculator, correct it
This Algebra I Study Guide contains clear, straight-forward problems that represent the topics covered in a complete Algebra I course. After completing the study guide without a calculator, correct it
SOLVING INEQUALITIES and 9.1.2
 SOLVING INEQUALITIES 9.1.1 and 9.1.2 To solve an inequality in one variable, first change it to an equation and solve. Place the solution, called a boundary point, on a number line. This point separates
SOLVING INEQUALITIES 9.1.1 and 9.1.2 To solve an inequality in one variable, first change it to an equation and solve. Place the solution, called a boundary point, on a number line. This point separates
Chapter 3: Statistical methods for estimation and testing. Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001).
 Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
Chapter 3: Statistical methods for estimation and testing Key reference: Statistical methods in bioinformatics by Ewens & Grant (2001). Chapter 3: Statistical methods for estimation and testing Key reference:
STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK. COURSE OUTLINE BIOL 310 The Human Genome. Prepared By: Ron Tavernier
 STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE BIOL 310 The Human Genome Prepared By: Ron Tavernier SCHOOL OF SCIENCE, HEALTH & CRIMINAL JUSTICE SCIENCE DEPARTMENT May
STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE BIOL 310 The Human Genome Prepared By: Ron Tavernier SCHOOL OF SCIENCE, HEALTH & CRIMINAL JUSTICE SCIENCE DEPARTMENT May
Causal Discovery by Computer
 Causal Discovery by Computer Clark Glymour Carnegie Mellon University 1 Outline 1. A century of mistakes about causation and discovery: 1. Fisher 2. Yule 3. Spearman/Thurstone 2. Search for causes is statistical
Causal Discovery by Computer Clark Glymour Carnegie Mellon University 1 Outline 1. A century of mistakes about causation and discovery: 1. Fisher 2. Yule 3. Spearman/Thurstone 2. Search for causes is statistical
Lecture 1 January 18
 STAT 263/363: Experimental Design Winter 2016/17 Lecture 1 January 18 Lecturer: Art B. Owen Scribe: Julie Zhu Overview Experiments are powerful because you can conclude causality from the results. In most
STAT 263/363: Experimental Design Winter 2016/17 Lecture 1 January 18 Lecturer: Art B. Owen Scribe: Julie Zhu Overview Experiments are powerful because you can conclude causality from the results. In most
Statistical methods research done as science rather than math: Estimates on the boundary in random regressions
 Statistical methods research done as science rather than math: Estimates on the boundary in random regressions This lecture is about how we study statistical methods. It uses as an example a problem that
Statistical methods research done as science rather than math: Estimates on the boundary in random regressions This lecture is about how we study statistical methods. It uses as an example a problem that
The Comparison Test & Limit Comparison Test
 The Comparison Test & Limit Comparison Test Math4 Department of Mathematics, University of Kentucky February 5, 207 Math4 Lecture 3 / 3 Summary of (some of) what we have learned about series... Math4 Lecture
The Comparison Test & Limit Comparison Test Math4 Department of Mathematics, University of Kentucky February 5, 207 Math4 Lecture 3 / 3 Summary of (some of) what we have learned about series... Math4 Lecture
Lecture 1: Systems of linear equations and their solutions
 Lecture 1: Systems of linear equations and their solutions Course overview Topics to be covered this semester: Systems of linear equations and Gaussian elimination: Solving linear equations and applications
Lecture 1: Systems of linear equations and their solutions Course overview Topics to be covered this semester: Systems of linear equations and Gaussian elimination: Solving linear equations and applications
Data representation and classification
 Representation and classification of data January 25, 2016 Outline Lecture 1: Data Representation 1 Lecture 1: Data Representation Data representation The phrase data representation usually refers to the
Representation and classification of data January 25, 2016 Outline Lecture 1: Data Representation 1 Lecture 1: Data Representation Data representation The phrase data representation usually refers to the
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling
 Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Expression Data Exploration: Association, Patterns, Factors & Regression Modelling Exploring gene expression data Scale factors, median chip correlation on gene subsets for crude data quality investigation
Statistical Methods for Analysis of Genetic Data
 Statistical Methods for Analysis of Genetic Data Christopher R. Cabanski A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
Statistical Methods for Analysis of Genetic Data Christopher R. Cabanski A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance
 Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Use of Agilent Feature Extraction Software (v8.1) QC Report to Evaluate Microarray Performance Anthea Dokidis Glenda Delenstarr Abstract The performance of the Agilent microarray system can now be evaluated
Microarray Data Analysis: Discovery
 Microarray Data Analysis: Discovery Lecture 5 Classification Classification vs. Clustering Classification: Goal: Placing objects (e.g. genes) into meaningful classes Supervised Clustering: Goal: Discover
Microarray Data Analysis: Discovery Lecture 5 Classification Classification vs. Clustering Classification: Goal: Placing objects (e.g. genes) into meaningful classes Supervised Clustering: Goal: Discover
Sample Size Estimation for Studies of High-Dimensional Data
 Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Sample Size Estimation for Studies of High-Dimensional Data James J. Chen, Ph.D. National Center for Toxicological Research Food and Drug Administration June 3, 2009 China Medical University Taichung,
Varieties of Count Data
 CHAPTER 1 Varieties of Count Data SOME POINTS OF DISCUSSION What are counts? What are count data? What is a linear statistical model? What is the relationship between a probability distribution function
CHAPTER 1 Varieties of Count Data SOME POINTS OF DISCUSSION What are counts? What are count data? What is a linear statistical model? What is the relationship between a probability distribution function
Statistics in medicine
 Statistics in medicine Lecture 4: and multivariable regression Fatma Shebl, MD, MS, MPH, PhD Assistant Professor Chronic Disease Epidemiology Department Yale School of Public Health Fatma.shebl@yale.edu
Statistics in medicine Lecture 4: and multivariable regression Fatma Shebl, MD, MS, MPH, PhD Assistant Professor Chronic Disease Epidemiology Department Yale School of Public Health Fatma.shebl@yale.edu
Statistical Analysis of Engineering Data The Bare Bones Edition. Precision, Bias, Accuracy, Measures of Precision, Propagation of Error
 Statistical Analysis of Engineering Data The Bare Bones Edition (I) Precision, Bias, Accuracy, Measures of Precision, Propagation of Error PRIOR TO DATA ACQUISITION ONE SHOULD CONSIDER: 1. The accuracy
Statistical Analysis of Engineering Data The Bare Bones Edition (I) Precision, Bias, Accuracy, Measures of Precision, Propagation of Error PRIOR TO DATA ACQUISITION ONE SHOULD CONSIDER: 1. The accuracy
MATH 0409: Foundations of Mathematics COURSE OUTLINE
 MATH 0409: Foundations of Mathematics COURSE OUTLINE Spring 2016 CRN91085 MW 5:30-7:30pm AM209 Professor Sherri Escobar sherri.escobar@hccs.edu 281-620-1115 Catalog Description: Foundations of Mathematics.
MATH 0409: Foundations of Mathematics COURSE OUTLINE Spring 2016 CRN91085 MW 5:30-7:30pm AM209 Professor Sherri Escobar sherri.escobar@hccs.edu 281-620-1115 Catalog Description: Foundations of Mathematics.
Prentice Hall Geometry (c) 2007 correlated to American Diploma Project, High School Math Benchmarks
 I1.1. Add, subtract, multiply and divide integers, fractions and decimals. I1.2. Calculate and apply ratios, proportions, rates and percentages to solve problems. I1.3. Use the correct order of operations
I1.1. Add, subtract, multiply and divide integers, fractions and decimals. I1.2. Calculate and apply ratios, proportions, rates and percentages to solve problems. I1.3. Use the correct order of operations
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
ACCESS TO SCIENCE, ENGINEERING AND AGRICULTURE: MATHEMATICS 1 MATH00030 SEMESTER / Lines and Their Equations
 ACCESS TO SCIENCE, ENGINEERING AND AGRICULTURE: MATHEMATICS 1 MATH00030 SEMESTER 1 017/018 DR. ANTHONY BROWN. Lines and Their Equations.1. Slope of a Line and its y-intercept. In Euclidean geometry (where
ACCESS TO SCIENCE, ENGINEERING AND AGRICULTURE: MATHEMATICS 1 MATH00030 SEMESTER 1 017/018 DR. ANTHONY BROWN. Lines and Their Equations.1. Slope of a Line and its y-intercept. In Euclidean geometry (where
Multivariate Analysis of Ecological Data using CANOCO
 Multivariate Analysis of Ecological Data using CANOCO JAN LEPS University of South Bohemia, and Czech Academy of Sciences, Czech Republic Universitats- uric! Lanttesbibiiothek Darmstadt Bibliothek Biologie
Multivariate Analysis of Ecological Data using CANOCO JAN LEPS University of South Bohemia, and Czech Academy of Sciences, Czech Republic Universitats- uric! Lanttesbibiiothek Darmstadt Bibliothek Biologie
UNIT PLAN th Grade Pre-Calculus
 UNIT PLAN Subject/Grade Level: 10-12 th Grade Pre-Calculus Unit #: 1 Unit Name: Trigonometric Functions Big Idea/Theme: Using trigonometric functions to model real-life situations will prepare students
UNIT PLAN Subject/Grade Level: 10-12 th Grade Pre-Calculus Unit #: 1 Unit Name: Trigonometric Functions Big Idea/Theme: Using trigonometric functions to model real-life situations will prepare students
Lecture 16 Solving GLMs via IRWLS
 Lecture 16 Solving GLMs via IRWLS 09 November 2015 Taylor B. Arnold Yale Statistics STAT 312/612 Notes problem set 5 posted; due next class problem set 6, November 18th Goals for today fixed PCA example
Lecture 16 Solving GLMs via IRWLS 09 November 2015 Taylor B. Arnold Yale Statistics STAT 312/612 Notes problem set 5 posted; due next class problem set 6, November 18th Goals for today fixed PCA example
This gives us an upper and lower bound that capture our population mean.
 Confidence Intervals Critical Values Practice Problems 1 Estimation 1.1 Confidence Intervals Definition 1.1 Margin of error. The margin of error of a distribution is the amount of error we predict when
Confidence Intervals Critical Values Practice Problems 1 Estimation 1.1 Confidence Intervals Definition 1.1 Margin of error. The margin of error of a distribution is the amount of error we predict when
Contents. Chapter 5 Elements and Compounds 129. Chapter 1 Living Cells 1. Chapter 6 Physical and Chemical Changes 161. Chapter 2 Organ Systems 25
 Contents Words to Watch iv Chapter 5 Elements and Compounds 129 1 1.1 Plant, animal and fungal cells 3 1.2 Structures within cells 7 1.3 Examining cells 9 1.4 Single-celled organisms 15 1.5 Cell division
Contents Words to Watch iv Chapter 5 Elements and Compounds 129 1 1.1 Plant, animal and fungal cells 3 1.2 Structures within cells 7 1.3 Examining cells 9 1.4 Single-celled organisms 15 1.5 Cell division
Discrete Multivariate Statistics
 Discrete Multivariate Statistics Univariate Discrete Random variables Let X be a discrete random variable which, in this module, will be assumed to take a finite number of t different values which are
Discrete Multivariate Statistics Univariate Discrete Random variables Let X be a discrete random variable which, in this module, will be assumed to take a finite number of t different values which are
Chapter 1. A Physics Toolkit
 Chapter 1 A Physics Toolkit Chapter 1 A Physics Toolkit In this chapter you will: Use mathematical tools to measure and predict. Apply accuracy and precision when measuring. Display and evaluate data graphically.
Chapter 1 A Physics Toolkit Chapter 1 A Physics Toolkit In this chapter you will: Use mathematical tools to measure and predict. Apply accuracy and precision when measuring. Display and evaluate data graphically.
Generalized Linear Models (1/29/13)
 STA613/CBB540: Statistical methods in computational biology Generalized Linear Models (1/29/13) Lecturer: Barbara Engelhardt Scribe: Yangxiaolu Cao When processing discrete data, two commonly used probability
STA613/CBB540: Statistical methods in computational biology Generalized Linear Models (1/29/13) Lecturer: Barbara Engelhardt Scribe: Yangxiaolu Cao When processing discrete data, two commonly used probability
ISyE 6416: Computational Statistics Spring Lecture 5: Discriminant analysis and classification
 ISyE 6416: Computational Statistics Spring 2017 Lecture 5: Discriminant analysis and classification Prof. Yao Xie H. Milton Stewart School of Industrial and Systems Engineering Georgia Institute of Technology
ISyE 6416: Computational Statistics Spring 2017 Lecture 5: Discriminant analysis and classification Prof. Yao Xie H. Milton Stewart School of Industrial and Systems Engineering Georgia Institute of Technology
Algebra 1 S1 Lesson Summaries. Lesson Goal: Mastery 70% or higher
 Algebra 1 S1 Lesson Summaries For every lesson, you need to: Read through the LESSON REVIEW which is located below or on the last page of the lesson and 3-hole punch into your MATH BINDER. Read and work
Algebra 1 S1 Lesson Summaries For every lesson, you need to: Read through the LESSON REVIEW which is located below or on the last page of the lesson and 3-hole punch into your MATH BINDER. Read and work
Lecture 1 Introduction to Multi-level Models
 Lecture 1 Introduction to Multi-level Models Course Website: http://www.biostat.jhsph.edu/~ejohnson/multilevel.htm All lecture materials extracted and further developed from the Multilevel Model course
Lecture 1 Introduction to Multi-level Models Course Website: http://www.biostat.jhsph.edu/~ejohnson/multilevel.htm All lecture materials extracted and further developed from the Multilevel Model course
Regression. Estimation of the linear function (straight line) describing the linear component of the joint relationship between two variables X and Y.
 Regression Bivariate i linear regression: Estimation of the linear function (straight line) describing the linear component of the joint relationship between two variables and. Generally describe as a
Regression Bivariate i linear regression: Estimation of the linear function (straight line) describing the linear component of the joint relationship between two variables and. Generally describe as a
Prerequisite: STATS 7 or STATS 8 or AP90 or (STATS 120A and STATS 120B and STATS 120C). AP90 with a minimum score of 3
 University of California, Irvine 2017-2018 1 Statistics (STATS) Courses STATS 5. Seminar in Data Science. 1 Unit. An introduction to the field of Data Science; intended for entering freshman and transfers.
University of California, Irvine 2017-2018 1 Statistics (STATS) Courses STATS 5. Seminar in Data Science. 1 Unit. An introduction to the field of Data Science; intended for entering freshman and transfers.
CMSC858P Supervised Learning Methods
 CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
CMSC858P Supervised Learning Methods Hector Corrada Bravo March, 2010 Introduction Today we discuss the classification setting in detail. Our setting is that we observe for each subject i a set of p predictors
Algebra I. CORE TOPICS (Key Concepts & Real World Context) UNIT TITLE
 CHAPTER 1 Using variables Exponents and Order of Operations Exploring real numbers Adding real numbers Subtracting real numbers Multiplying and dividing real numbers The Distributive Property Properties
CHAPTER 1 Using variables Exponents and Order of Operations Exploring real numbers Adding real numbers Subtracting real numbers Multiplying and dividing real numbers The Distributive Property Properties
Non-specific filtering and control of false positives
 Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
Non-specific filtering and control of false positives Richard Bourgon 16 June 2009 bourgon@ebi.ac.uk EBI is an outstation of the European Molecular Biology Laboratory Outline Multiple testing I: overview
TEKS Clarification Document. Mathematics Algebra
 TEKS Clarification Document Mathematics Algebra 2 2012 2013 111.31. Implementation of Texas Essential Knowledge and Skills for Mathematics, Grades 9-12. Source: The provisions of this 111.31 adopted to
TEKS Clarification Document Mathematics Algebra 2 2012 2013 111.31. Implementation of Texas Essential Knowledge and Skills for Mathematics, Grades 9-12. Source: The provisions of this 111.31 adopted to
LIST OF ABBREVIATIONS:
 KEY LINK BETWEEN REGRESSION COEFFICIENTS AND STRUCTURAL CHARACTERIZATION OF HYDROCARBONS IN GASOLINE (MOTOR SPIRIT) FOR CONTINUOUS BLENDING PROCESS APPLICATIONS Santanu Talukdar Manager-Analytical and
KEY LINK BETWEEN REGRESSION COEFFICIENTS AND STRUCTURAL CHARACTERIZATION OF HYDROCARBONS IN GASOLINE (MOTOR SPIRIT) FOR CONTINUOUS BLENDING PROCESS APPLICATIONS Santanu Talukdar Manager-Analytical and
Math 2 Unit 4 Quadratic Functions: Modeling
 Approximate Time Frame: 2 3 Weeks Connections to Previous Learning: In Math 1, students learn how to model functions and how to interpret models of functions. Students looked at multiple representations
Approximate Time Frame: 2 3 Weeks Connections to Previous Learning: In Math 1, students learn how to model functions and how to interpret models of functions. Students looked at multiple representations
Introduction to Machine Learning Midterm Exam
 10-701 Introduction to Machine Learning Midterm Exam Instructors: Eric Xing, Ziv Bar-Joseph 17 November, 2015 There are 11 questions, for a total of 100 points. This exam is open book, open notes, but
10-701 Introduction to Machine Learning Midterm Exam Instructors: Eric Xing, Ziv Bar-Joseph 17 November, 2015 There are 11 questions, for a total of 100 points. This exam is open book, open notes, but
Physical Science: Concepts in Action with Earth and Space Science 2009
 Prentice Hall Physical Science: Concepts in Action with Grades 9-12 C O R R E L A T E D T O PHYSICAL SCIENCE Course Description Physical Science is a laboratory course that explores the relationship between
Prentice Hall Physical Science: Concepts in Action with Grades 9-12 C O R R E L A T E D T O PHYSICAL SCIENCE Course Description Physical Science is a laboratory course that explores the relationship between
Information in Biology
 Lecture 3: Information in Biology Tsvi Tlusty, tsvi@unist.ac.kr Living information is carried by molecular channels Living systems I. Self-replicating information processors Environment II. III. Evolve
Lecture 3: Information in Biology Tsvi Tlusty, tsvi@unist.ac.kr Living information is carried by molecular channels Living systems I. Self-replicating information processors Environment II. III. Evolve
Name Class Date. t = = 10m. n + 19 = = 2f + 9
 1-4 Reteaching Solving Equations To solve an equation that contains a variable, find all of the values of the variable that make the equation true. Use the equality properties of real numbers and inverse
1-4 Reteaching Solving Equations To solve an equation that contains a variable, find all of the values of the variable that make the equation true. Use the equality properties of real numbers and inverse
Principal Component Analysis, A Powerful Scoring Technique
 Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Utah Core State Standards for Mathematics Secondary Mathematics I
 A Correlation of Integrated Math I Common Core 2014 to the Utah Core State for Mathematics Secondary Resource Title: : Common Core Publisher: Pearson Education publishing as Prentice Hall ISBN (10 or 13
A Correlation of Integrated Math I Common Core 2014 to the Utah Core State for Mathematics Secondary Resource Title: : Common Core Publisher: Pearson Education publishing as Prentice Hall ISBN (10 or 13
Application of mathematical, statistical, graphical or symbolic methods to maximize chemical information.
 Application of mathematical, statistical, graphical or symbolic methods to maximize chemical information. -However, this definition can be expanded to include: biology (biometrics), environmental science
Application of mathematical, statistical, graphical or symbolic methods to maximize chemical information. -However, this definition can be expanded to include: biology (biometrics), environmental science
Seymour Public Schools Curriculum
 Algebra II continues the study of functions. Students learn operations with functions and graphing of different types of functions. Students solve equations and perform operations with exponents, matrices
Algebra II continues the study of functions. Students learn operations with functions and graphing of different types of functions. Students solve equations and perform operations with exponents, matrices
Algebra and Trigonometry 2006 (Foerster) Correlated to: Washington Mathematics Standards, Algebra 2 (2008)
 A2.1. Core Content: Solving problems The first core content area highlights the type of problems students will be able to solve by the end of, as they extend their ability to solve problems with additional
A2.1. Core Content: Solving problems The first core content area highlights the type of problems students will be able to solve by the end of, as they extend their ability to solve problems with additional
An analytical model of the effect of plasmid copy number on transcriptional noise strength
 An analytical model of the effect of plasmid copy number on transcriptional noise strength William and Mary igem 2015 1 Introduction In the early phases of our experimental design, we considered incorporating
An analytical model of the effect of plasmid copy number on transcriptional noise strength William and Mary igem 2015 1 Introduction In the early phases of our experimental design, we considered incorporating
Chapter 6. Logistic Regression. 6.1 A linear model for the log odds
 Chapter 6 Logistic Regression In logistic regression, there is a categorical response variables, often coded 1=Yes and 0=No. Many important phenomena fit this framework. The patient survives the operation,
Chapter 6 Logistic Regression In logistic regression, there is a categorical response variables, often coded 1=Yes and 0=No. Many important phenomena fit this framework. The patient survives the operation,
Binary Logistic Regression
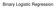 The coefficients of the multiple regression model are estimated using sample data with k independent variables Estimated (or predicted) value of Y Estimated intercept Estimated slope coefficients Ŷ = b
The coefficients of the multiple regression model are estimated using sample data with k independent variables Estimated (or predicted) value of Y Estimated intercept Estimated slope coefficients Ŷ = b
Comparison of Shannon, Renyi and Tsallis Entropy used in Decision Trees
 Comparison of Shannon, Renyi and Tsallis Entropy used in Decision Trees Tomasz Maszczyk and W lodzis law Duch Department of Informatics, Nicolaus Copernicus University Grudzi adzka 5, 87-100 Toruń, Poland
Comparison of Shannon, Renyi and Tsallis Entropy used in Decision Trees Tomasz Maszczyk and W lodzis law Duch Department of Informatics, Nicolaus Copernicus University Grudzi adzka 5, 87-100 Toruń, Poland
12 Statistical Justifications; the Bias-Variance Decomposition
 Statistical Justifications; the Bias-Variance Decomposition 65 12 Statistical Justifications; the Bias-Variance Decomposition STATISTICAL JUSTIFICATIONS FOR REGRESSION [So far, I ve talked about regression
Statistical Justifications; the Bias-Variance Decomposition 65 12 Statistical Justifications; the Bias-Variance Decomposition STATISTICAL JUSTIFICATIONS FOR REGRESSION [So far, I ve talked about regression
General Linear Model (Chapter 4)
 General Linear Model (Chapter 4) Outcome variable is considered continuous Simple linear regression Scatterplots OLS is BLUE under basic assumptions MSE estimates residual variance testing regression coefficients
General Linear Model (Chapter 4) Outcome variable is considered continuous Simple linear regression Scatterplots OLS is BLUE under basic assumptions MSE estimates residual variance testing regression coefficients
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
MTH Linear Algebra. Study Guide. Dr. Tony Yee Department of Mathematics and Information Technology The Hong Kong Institute of Education
 MTH 3 Linear Algebra Study Guide Dr. Tony Yee Department of Mathematics and Information Technology The Hong Kong Institute of Education June 3, ii Contents Table of Contents iii Matrix Algebra. Real Life
MTH 3 Linear Algebra Study Guide Dr. Tony Yee Department of Mathematics and Information Technology The Hong Kong Institute of Education June 3, ii Contents Table of Contents iii Matrix Algebra. Real Life
Multilevel Models in Matrix Form. Lecture 7 July 27, 2011 Advanced Multivariate Statistical Methods ICPSR Summer Session #2
 Multilevel Models in Matrix Form Lecture 7 July 27, 2011 Advanced Multivariate Statistical Methods ICPSR Summer Session #2 Today s Lecture Linear models from a matrix perspective An example of how to do
Multilevel Models in Matrix Form Lecture 7 July 27, 2011 Advanced Multivariate Statistical Methods ICPSR Summer Session #2 Today s Lecture Linear models from a matrix perspective An example of how to do
Uncertainty, Error, and Precision in Quantitative Measurements an Introduction 4.4 cm Experimental error
 Uncertainty, Error, and Precision in Quantitative Measurements an Introduction Much of the work in any chemistry laboratory involves the measurement of numerical quantities. A quantitative measurement
Uncertainty, Error, and Precision in Quantitative Measurements an Introduction Much of the work in any chemistry laboratory involves the measurement of numerical quantities. A quantitative measurement
COS513: FOUNDATIONS OF PROBABILISTIC MODELS LECTURE 10
 COS53: FOUNDATIONS OF PROBABILISTIC MODELS LECTURE 0 MELISSA CARROLL, LINJIE LUO. BIAS-VARIANCE TRADE-OFF (CONTINUED FROM LAST LECTURE) If V = (X n, Y n )} are observed data, the linear regression problem
COS53: FOUNDATIONS OF PROBABILISTIC MODELS LECTURE 0 MELISSA CARROLL, LINJIE LUO. BIAS-VARIANCE TRADE-OFF (CONTINUED FROM LAST LECTURE) If V = (X n, Y n )} are observed data, the linear regression problem
Linear, threshold units. Linear Discriminant Functions and Support Vector Machines. Biometrics CSE 190 Lecture 11. X i : inputs W i : weights
 Linear Discriminant Functions and Support Vector Machines Linear, threshold units CSE19, Winter 11 Biometrics CSE 19 Lecture 11 1 X i : inputs W i : weights θ : threshold 3 4 5 1 6 7 Courtesy of University
Linear Discriminant Functions and Support Vector Machines Linear, threshold units CSE19, Winter 11 Biometrics CSE 19 Lecture 11 1 X i : inputs W i : weights θ : threshold 3 4 5 1 6 7 Courtesy of University
DIFFERENTIAL EQUATIONS
 DIFFERENTIAL EQUATIONS Basic Concepts Paul Dawkins Table of Contents Preface... Basic Concepts... 1 Introduction... 1 Definitions... Direction Fields... 8 Final Thoughts...19 007 Paul Dawkins i http://tutorial.math.lamar.edu/terms.aspx
DIFFERENTIAL EQUATIONS Basic Concepts Paul Dawkins Table of Contents Preface... Basic Concepts... 1 Introduction... 1 Definitions... Direction Fields... 8 Final Thoughts...19 007 Paul Dawkins i http://tutorial.math.lamar.edu/terms.aspx
Data Preprocessing. Data Preprocessing
 Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Data Preprocessing 1 Data Preprocessing Normalization: the process of removing sampleto-sample variations in the measurements not due to differential gene expression. Bringing measurements from the different
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Singular Value Decomposition and Principal Component Analysis (PCA) I
 Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
Singular Value Decomposition and Principal Component Analysis (PCA) I Prof Ned Wingreen MOL 40/50 Microarray review Data per array: 0000 genes, I (green) i,i (red) i 000 000+ data points! The expression
OCR Biology Checklist
 Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Part 8: GLMs and Hierarchical LMs and GLMs
 Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
Part 8: GLMs and Hierarchical LMs and GLMs 1 Example: Song sparrow reproductive success Arcese et al., (1992) provide data on a sample from a population of 52 female song sparrows studied over the course
OCR Biology Checklist
 Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Bradley Broom Department of Bioinformatics and Computational Biology UT M. D. Anderson Cancer Center kabagg@mdanderson.org bmbroom@mdanderson.org
STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE MATH BEGINNING ALGEBRA
 STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE MATH 100 - BEGINNING ALGEBRA Prepared By: Mathematics Department CANINO SCHOOL OF ENGINEERING TECHNOLOGY MATHEMATICS DEPARTMENT
STATE UNIVERSITY OF NEW YORK COLLEGE OF TECHNOLOGY CANTON, NEW YORK COURSE OUTLINE MATH 100 - BEGINNING ALGEBRA Prepared By: Mathematics Department CANINO SCHOOL OF ENGINEERING TECHNOLOGY MATHEMATICS DEPARTMENT
Sample Size and Power I: Binary Outcomes. James Ware, PhD Harvard School of Public Health Boston, MA
 Sample Size and Power I: Binary Outcomes James Ware, PhD Harvard School of Public Health Boston, MA Sample Size and Power Principles: Sample size calculations are an essential part of study design Consider
Sample Size and Power I: Binary Outcomes James Ware, PhD Harvard School of Public Health Boston, MA Sample Size and Power Principles: Sample size calculations are an essential part of study design Consider
GS Analysis of Microarray Data
 GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
GS01 0163 Analysis of Microarray Data Keith Baggerly and Kevin Coombes Section of Bioinformatics Department of Biostatistics and Applied Mathematics UT M. D. Anderson Cancer Center kabagg@mdanderson.org
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
I L L I N O I S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN
 Introduction Edps/Psych/Stat/ 584 Applied Multivariate Statistics Carolyn J Anderson Department of Educational Psychology I L L I N O I S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN c Board of Trustees,
Introduction Edps/Psych/Stat/ 584 Applied Multivariate Statistics Carolyn J Anderson Department of Educational Psychology I L L I N O I S UNIVERSITY OF ILLINOIS AT URBANA-CHAMPAIGN c Board of Trustees,
