Design and Analysis of Linear Cascade DNA Hybridization Chain Reactions. Using DNA Hairpins. Reif 1. Duke University. Durham, NC 27708, USA
|
|
|
- Annis Morris
- 5 years ago
- Views:
Transcription
1 Design and Analysis of inear Cascade DNA Hybridization Chain eactions Using DNA Hairpins Hieu Bui, Sudhanshu Garg, Vincent Miao, Tianqi Song, eem Mokhtar, John eif Department of mputer Science, Department of Biomedical Engineering, Duke University Durham, NC 7708, USA Supporting Information
2 S: Sequence Design via NUPACK In general, the design process begins with a set of target DNA nanostructures consisting of 9 hairpins and initiators. Each hairpin structure has a - bp stem, 8- nt loop, and 6- nt external toehold. Hairpin 9 has a slightly longer loop length of - nt. In order to displace the stem domain and form a thermodynamically stable structure, each target molecule forms a - bp duplex with a designated hairpin strand. The melting temperature of a - bp duplex is estimated to be 55 to 65 o C at.5 mm Mg + salt concentration and ph 8.0. The reporter complex consists of a 0- bp duplex with 7- nt toehold. All binding domains are assigned with nucleotides using three letter codes (A, T, C) in order to minimize downstream undesirable leaks. NUPACK has built- in functions to ) accommodate sequence symmetry minimization, ) find the average number of incorrect nucleotides relative to target structures at equilibrium, ) generate the probability map of adopting target structures at equilibrium, and ) apply DNA hybridization kinetics. A set of stringent stopping conditions ensures a higher probability of obtaining desirable target structures; however, it takes longer to generate such structures. In our system, a list of stopping conditions includes ) target initiator structures cannot form undesirable complexes with more than 5% of all generated structures, ) target structures cannot have more than three consecutive guanine bases or more than four consecutive bases of any other nucleotide. The number of trials is set to 0, which means that there are 0 different matching sets of target molecules from the given specification list. The actual
3 sequence composition can be assigned or unassigned. Since we adopted the same reporter complex reported by Zhang et al., the sequences for the reporter complex are known. As a result, a portion of binding domains in the linear cascade reaction share the same sequences as the reporter complex. The script to generate target DNA sequences for our system is shown below:
4 design an HC system of 9 distinct hairpins design material, temperature (C), and trials material = dna temperature = 5 trials = 0 magnesium = 0.05 sodium = 0.5 target structures structure hairpin = D U8 U6 structure hairpin = U6 D U8 structure hairpin = D U8 U6 structure hairpin = U6 D U8 structure hairpin5 = D U8 U6 structure hairpin6 = U6 D U8 structure hairpin7 = D U8 U6 structure hairpin8 = U6 D U8 structure hairpin9 = D U U6 structure initiator = U structure initiator = U structure initiator = U structure initiator = U structure initiator5 = U structure initiator6 = U structure initiator7 = U structure initiator8 = U structure initiator9 = U structure C = D0 (+ U7) structure initiatorc = U sequence domains domain s0 = N6 domain s = TACCTA domain s = CTCTTT domain s = CCACCC domain s = CCTCCT domain s5 = TCTTTC domain s6 = TCTTCC domain s7 = TTTCCT domain s8 = CTTCTC domain s9 = T CATTC domain r = ACATACATCA domain l = T domain l = AA domain l = GC domain ci = N domain c0 = CCAC domain c = TTCT domain c = CATT domain c = CAAC domain c = ACTC domain c5 = ATCG domain c6 = TTTA domain c7 = CCAC domain c8 = CT CC domain c9 = TATTCCC domain t = CATTC AA domain bm = CC ACATACATCA TATTCCC T
5 thread sequence domains onto target structures hairpin.seq = c0 r c s l ci* r* c0* s0* hairpin.seq = s* c* r* c0* l s c r c hairpin.seq = c r c s l c* r* c* s* hairpin.seq = s* c* r* c* l s c r c hairpin5.seq = c r c5 s5 l c* r* c* s* hairpin6.seq = s5* c5* r* c* l s6 c6 r c5 hairpin7.seq = c6 r c7 s7 l c5* r* c6* s6* hairpin8.seq = s7* c7* r* c6* l s8 c8 r c7 hairpin9.seq = c8 r c9 s9 l l c7* r* c8* s8* initiator.seq = s0 c0 r ci initiator.seq = c0 r c s initiator.seq = s c r c initiator.seq = c r c s initiator5.seq = s c r c initiator6.seq = c r c5 s5 initiator7.seq = s6 c6 r c5 initiator8.seq = c6 r c7 s7 initiator9.seq = s8 c8 r c7 initiatorc.seq = c8 r c9 s9 l l C.seq = bm t* bm* specify stop conditions for normalized ensemble defect default:.0 (percent) for each target structure initiator.stop = 5.0 larger defect for unpaired structures prevent = SSSS, AAAAA, AAAA, GGG Script : NUPACK code to generate designed C system S: Modeling the reaction kinetics of linear cascade DNA hybridization reactions via chemical reaction equations Each DNA hairpin in the proposed system is treated abstractly as an individual molecule and is assumed to only bind the designated DNA strand strictly via DNA hybridization and strand displacement thermodynamics. The rates of all the reactions in 9 hairpins system can be described based on the second order kinetic process. 5
6 I + H!! P () P + H!! P () P + H!! P () P + H!! P () P + H5!! P5 (5) P5 + H6!! P6 (6) P6 + H7!! P7 (7) P7 + H8!! P8 (8) P8 + H9!! P9 (9) P9 + C!! FQ + TET (0) We further assume that the reverse reaction is negligible, although it is possible in autocatalytic DNA systems reported by Zhang et al.[9]. Since the number of base pairs from each cascade reaction is equal, we assume the forward rate constants are relatively equivalent for reactions () to (9). For the rate constant from the reaction (0), we used the same rate constant reported by Zhang et al.[9]. 6
7 ncentration (nm) hairpin (x I) hairpins (x I) hairpins (x I) 6 hairpins (x I) 9 hairpins (x I) ncentration (nm) hairpin (x I) hairpins (x I) hairpins (x I) 6 hairpins (x I) 9 hairpins (x I) Time (minutes) Time (minutes) ncentration (nm) hairpin (x I) hairpins (x I) hairpins (x I) 6 hairpins (x I) 9 hairpins (x I) ncentration (nm) hairpin (x I) hairpins (x I) hairpins (x I) 6 hairpins (x I) 9 hairpins (x I) Time (minutes) Time (minutes) Figure S: Modeling linear cascade DNA hybridization reactions with different rate constants (k) (i.e 0.5 x 0 5, 0 5, 5 x 0 5, 0 6 /M/s). The length of linear duplex is a function of the number of hairpins. Simulation analysis shows results from implementing different rate constants S: Matlab scripts The following scripts consist of two parts. Part one contains a function to solve the linear cascade DNA hybridization reaction involving nine distinct hairpins. Here, the function consists of a list of 0 bimolecular reactions as shown in Section S. Part two contains a function to ) simulate DNA hairpin interactions with the given initial concentrations and rate constants or ) optimize the least squares fit between the experimental data and 7
8 simulation data to determine the rate constant, in similar fashion to the previously reported method by Zhang et al. [7]. %%% This function is used to solve cascade chain reactions %%% Date: July, 06 function dy = lcr9(t,y) % % I + H -> IH ( + -> ) % % IH + H -> IH ( + 5 -> 6) % % IH + H -> IH ( > 8) % % IH + H -> IH ( > 0) % % IH + H5 -> IH5 (0 + -> ) % % IH5 + H6 -> IH6 ( + -> ) % % IH6 + H7 -> IH7 ( + 5 -> 6) % % IH7 + H8 -> IH8 ( > 8) % % IH8 + H9 -> IH9 ( > 0) % % IH9 + C -> OF (0 + -> ) dy = zeros(,); krep =.E6*60; % ate constant (/M/min) from earlier studies by Zhang et al. dy() = - y() * y() * y(); dy() = - y() * y() * y(); dy() = y() * y() * y() - y() * y() * y(5); dy(5) = - y() * y() * y(5); dy(6) = y() * y() * y(5) - y() * y(6) * y(7); dy(7) = - y() * y(6) * y(7); dy(8) = y() * y(6) * y(7) - y() * y(8) * y(9); dy(9) = - y() * y(8) * y(9); dy(0) = y() * y(8) * y(9) - y() * y(0) * y(); dy() = - y() * y(0) * y(); dy() = y() * y(0) * y() - y() * y() * y(); dy() = - y() * y() * y(); dy() = y() * y() * y() - y() * y() * y(5); dy(5) = - y() * y() * y(5); dy(6) = y() * y() * y(5) - y() * y(6) * y(7); dy(7) = - y() * y(6) * y(7); dy(8) = y() * y(6) * y(7) - y() * y(8) * y(9); dy(9) = - y() * y(8) * y(9); dy(0) = y() * y(8) * y(9) - krep * y(0) * y(); dy() = - krep * y(0) * y(); dy() = krep * y(0) * y(); return Script : C function to solve bimolecular cascade reactions 8
9 function err_fun = rate(input) data = load('path-to-data-file ); k = exp(input()); scalingconst = exp(input()); err_fun = 0; options = odeset('eltol', e-, 'AbsTol', e-0); datasize = size(data,); t = data(:datasize,); y0=5e-9*[,, 0,, 0,, 0,, 0,, 0,, 0,, 0,, 0,, 0,., 0]; % initial concentrations [t y]=odes(@lcr9, t, [k, y0], options); % 9H %%% find minimum error between the ODE solution and data ye = y(:,size(y,)) * scalingconst + data(,); for i = :size(data,) err_fun = err_fun + (ye(i) - data(i,))^/ye(i); end return % Fitting the experimental data using the least squares method by calling the err_fun function. The err_fun function then uses the lcr9 function to solve the ODE with the given initial conditions. clear all close all % Use this one to figure out the rate k0 = log(e6*60); % initial guess scale0 = log(e); % initial guess [k, fval] = fminunc(@rate,[k0, scale0]); Script : Error function to determine the rate constant. 9
10 So C8 S9 C8 S9 Hairpin 9 H 9 Initiator I C complex Schematic B So So So C8 C0 C S C C0 S S9 Hairpin 9 H 9 Initiator I C C S Hairpin I H S9 So So S C8 C7 C8 S8 Hairpin 8 H 8 C complex C C C C S Hairpin H S8 C8 C6 S C C (A) C7 C7 S7 Hairpin 7 H 7 S7 C7 C5 C6 C6 S6 S6 C6 C C5 C5 S5 S5 C5 C C C S C C S Hairpin H Hairpin 6 H 6 Hairpin 5 H 5 Hairpin H Schematic C Figure S: Estimation of yield via maximum fluorescence intensity readout. All experimental data were collected at 5 nm concentration. Assume that the maximum fluorescence intensity at thermal equilibrium is correlated with the yield of hairpins participating in the linear cascade reaction. Black curve is the fluorescence emission of initiating the last hairpin and resulting in displacing the reporter complex (schematic B). Cyan curve is the fluorescence emission of initiating the first hairpin and resulting in displacing the reporter complex after the linear cascade reaction (schematic C). The maximum fluorescence at thermal equilibrium of the linear cascade consisting of a single hairpin is 7 a.u. The maximum fluorescence at thermal equilibrium of the linear cascade consisting of 9 distinct hairpins is 57 a.u. To the first order approximation, the yield of 9 hairpins is approximated 79%. 0
11 5.5 ncentration (nm) Data - hairpin (x I) Data - hairpins (x I) Data - hairpins (x I) Data - 6 hairpins (x I) Data - 9 hairpins (x I) Model - hairpin (x I) Model - hairpins (x I) Model - hairpins (x I) Model - 6 hairpins (x I) Model - 9 hairpins (x I) Time (minutes) Figure S: Fitting the experimental results using the model proposed in the Supporting Information Section S: Scatter plots are experimental data; ine curves are fitted data using the least squares method[7]. The experimental results from Figure 6A were fitted using the least squares method[7]. Matlab scripts were detailed in Section SI S. Each experimental result was fitted to the corresponding cascade simulation. For example, the experimental result of nine hairpins was fitted using the cascade simulation of nine hairpins as shown in Script. As mentioned from prior discussion, the hairpins were designed to have the same structure with different nucleotide composition. The model assumed that the rate constant of each cascade reaction was equal. To determine this rate constant, Script was run until a final best- fit curve was obtained. This best- fit curve was resulted from minimizing the error function, which was the least squares error between the experimental data and the simulation data. Figure S shows the results from fitting the experimental data to the corresponding cascade simulations. The rate constants for systems of,,, 6, and 9 hairpins were fitted to be 0.97 x 0 6 /M/s, 0.85 x 0 6 /M/s, 0.58 x 0 6 /M/s,.0 x 0 6 /M/s, and. x 0 6 /M/s, respectively. These rate constants are independent of the cascade length and are approximately similar to the rate constant of the reporter complex (. x 0 6 /M/s). Note that the reporter complex was adopted from Zhang et al. [9]. Note that the fitted data reached the saturation level earlier than the experimental data. This discrepancy may be due to one or both of the following: i) improper synthesis or defects of DNA, or ii) the model does account for the fact that the actual system is not 00% efficient (i.e. all molecules from the model are assumed to participate in the reaction and this notion seems to be far- off from the actual observed experimental data).
S1. Quasi-steady state (QSS) derivation of the BM rate constant
 Control of DNA Strand Displacement Kinetics using Toehold Exchange Supporting Materials David Yu Zhang 1 and Erik Winfree 1 1 California Institute of Technology, MC 136-93, 1200 E. California Blvd., Pasadena,
Control of DNA Strand Displacement Kinetics using Toehold Exchange Supporting Materials David Yu Zhang 1 and Erik Winfree 1 1 California Institute of Technology, MC 136-93, 1200 E. California Blvd., Pasadena,
Introduction to Polymer Physics
 Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Computational Approaches for determination of Most Probable RNA Secondary Structure Using Different Thermodynamics Parameters
 Computational Approaches for determination of Most Probable RNA Secondary Structure Using Different Thermodynamics Parameters 1 Binod Kumar, Assistant Professor, Computer Sc. Dept, ISTAR, Vallabh Vidyanagar,
Computational Approaches for determination of Most Probable RNA Secondary Structure Using Different Thermodynamics Parameters 1 Binod Kumar, Assistant Professor, Computer Sc. Dept, ISTAR, Vallabh Vidyanagar,
Supporting Information for. Accelerating DNA based computing on a supramolecular polymer
 Supporting Information for Accelerating DNA based computing on a supramolecular polymer Wouter Engelen, #, Sjors P.W. Wijnands # and Maarten Merkx #,,* * Corresponding Author # Institute for Complex Molecular
Supporting Information for Accelerating DNA based computing on a supramolecular polymer Wouter Engelen, #, Sjors P.W. Wijnands # and Maarten Merkx #,,* * Corresponding Author # Institute for Complex Molecular
Programmability and versatility are essential properties of
 pubs.acs.org/synthbio Analog Computation by DNA Strand Displacement Circuits Tianqi Song, Sudhanshu Garg, Reem Mokhtar, Hieu Bui, and John Reif* Department of Computer Science, Duke University, Durham,
pubs.acs.org/synthbio Analog Computation by DNA Strand Displacement Circuits Tianqi Song, Sudhanshu Garg, Reem Mokhtar, Hieu Bui, and John Reif* Department of Computer Science, Duke University, Durham,
Dot Bracket Notation for RNA and DNA nanostructures. Slides by Reem Mokhtar
 Dot Bracket Notation for RNA and DNA nanostructures Slides by Reem Mokhtar Graphical/Data Purpose: - Ease of interaction and design - Aid in validating designs Representations might include - GUI input
Dot Bracket Notation for RNA and DNA nanostructures Slides by Reem Mokhtar Graphical/Data Purpose: - Ease of interaction and design - Aid in validating designs Representations might include - GUI input
Videos. Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.
 Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Reconfigurable DNA Origami Nanocapsule for ph- Controlled Encapsulation and Display of Cargo
 Supporting Information Reconfigurable DNA Origami Nanocapsule for ph- Controlled Encapsulation and Display of Cargo Heini Ijäs, Iiris Hakaste, Boxuan Shen, Mauri A. Kostiainen, and Veikko Linko* Biohybrid
Supporting Information Reconfigurable DNA Origami Nanocapsule for ph- Controlled Encapsulation and Display of Cargo Heini Ijäs, Iiris Hakaste, Boxuan Shen, Mauri A. Kostiainen, and Veikko Linko* Biohybrid
Supplementary Information. for. Controlled Scalable Synthesis of Uniform, High-Quality Monolayer and Fewlayer
 Supplementary Information for Controlled Scalable Synthesis of Uniform, High-Quality Monolayer and Fewlayer MoS 2 Films Yifei Yu 1, Chun Li 1, Yi Liu 3, Liqin Su 4, Yong Zhang 4, Linyou Cao 1,2 * 1 Department
Supplementary Information for Controlled Scalable Synthesis of Uniform, High-Quality Monolayer and Fewlayer MoS 2 Films Yifei Yu 1, Chun Li 1, Yi Liu 3, Liqin Su 4, Yong Zhang 4, Linyou Cao 1,2 * 1 Department
Protein Synthesis. Unit 6 Goal: Students will be able to describe the processes of transcription and translation.
 Protein Synthesis Unit 6 Goal: Students will be able to describe the processes of transcription and translation. Protein Synthesis: Protein synthesis uses the information in genes to make proteins. 2 Steps
Protein Synthesis Unit 6 Goal: Students will be able to describe the processes of transcription and translation. Protein Synthesis: Protein synthesis uses the information in genes to make proteins. 2 Steps
SUPPLEMENTARY INFORMATION
 Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
Fig. 1 Influences of crystal lattice contacts on Pol η structures. a. The dominant lattice contact between two hpol η molecules (silver and gold) in the type 1 crystals. b. A close-up view of the hydrophobic
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM
 The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
The Riboswitch is functionally separated into the ligand binding APTAMER and the decision-making EXPRESSION PLATFORM Purine riboswitch TPP riboswitch SAM riboswitch glms ribozyme In-line probing is used
RNA secondary structure prediction. Farhat Habib
 RNA secondary structure prediction Farhat Habib RNA RNA is similar to DNA chemically. It is usually only a single strand. T(hyamine) is replaced by U(racil) Some forms of RNA can form secondary structures
RNA secondary structure prediction Farhat Habib RNA RNA is similar to DNA chemically. It is usually only a single strand. T(hyamine) is replaced by U(racil) Some forms of RNA can form secondary structures
Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India. 1 st November, 2013
 Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Hydration of protein-rna recognition sites Ranjit P. Bahadur Assistant Professor Department of Biotechnology Indian Institute of Technology Kharagpur, India 1 st November, 2013 Central Dogma of life DNA
Chapter 2. The Chemical Context of Life
 Chapter 2 The Chemical Context of Life 1 Matter Takes up space and has mass Exists as elements (pure form) and in chemical combinations called compounds 2 Elements Can t be broken down into simpler substances
Chapter 2 The Chemical Context of Life 1 Matter Takes up space and has mass Exists as elements (pure form) and in chemical combinations called compounds 2 Elements Can t be broken down into simpler substances
DNA as a Universal Substrate for Chemical Kinetics (Extended Abstract)
 DNA as a Universal Substrate for Chemical Kinetics (Extended Abstract) David Soloveichik, Georg Seelig, and Erik Winfree California Institute of Technology, Pasadena, CA, USA dsolov@caltech.edu, seelig@dna.caltech.edu,
DNA as a Universal Substrate for Chemical Kinetics (Extended Abstract) David Soloveichik, Georg Seelig, and Erik Winfree California Institute of Technology, Pasadena, CA, USA dsolov@caltech.edu, seelig@dna.caltech.edu,
The Design and Fabrication of a Fully Addressable 8-tile DNA Lattice
 The Design and Fabrication of a Fully Addressable 8-tile DNA Lattice Chris Dwyer 1,3, Sung Ha Park 2, Thomas LaBean 3, Alvin Lebeck 3 1 Dept. of Electrical and Computer Engineering, Duke University, Durham,
The Design and Fabrication of a Fully Addressable 8-tile DNA Lattice Chris Dwyer 1,3, Sung Ha Park 2, Thomas LaBean 3, Alvin Lebeck 3 1 Dept. of Electrical and Computer Engineering, Duke University, Durham,
Exquisite Sequence Selectivity with Small Conditional RNAs
 Supplementary Information Exquisite Sequence Selectivity with Small Conditional RNAs Jonathan B. Sternberg and Niles A. Pierce,, Division of Biology & Biological Engineering, Division of Engineering &
Supplementary Information Exquisite Sequence Selectivity with Small Conditional RNAs Jonathan B. Sternberg and Niles A. Pierce,, Division of Biology & Biological Engineering, Division of Engineering &
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Chapter 2: Chemistry. What does chemistry have to do with biology? Vocabulary BIO 105
 Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Chapter 2: Chemistry What does chemistry have to do with biology? BIO 105 Vocabulary 1. Matter anything that takes up space and has mass Atoms are the smallest units of matter that can participate in chemical
Homework Assignment 4 Root Finding Algorithms The Maximum Velocity of a Car Due: Friday, February 19, 2010 at 12noon
 ME 2016 A Spring Semester 2010 Computing Techniques 3-0-3 Homework Assignment 4 Root Finding Algorithms The Maximum Velocity of a Car Due: Friday, February 19, 2010 at 12noon Description and Outcomes In
ME 2016 A Spring Semester 2010 Computing Techniques 3-0-3 Homework Assignment 4 Root Finding Algorithms The Maximum Velocity of a Car Due: Friday, February 19, 2010 at 12noon Description and Outcomes In
Protein Synthesis. Unit 6 Goal: Students will be able to describe the processes of transcription and translation.
 Protein Synthesis Unit 6 Goal: Students will be able to describe the processes of transcription and translation. Types of RNA Messenger RNA (mrna) makes a copy of DNA, carries instructions for making proteins,
Protein Synthesis Unit 6 Goal: Students will be able to describe the processes of transcription and translation. Types of RNA Messenger RNA (mrna) makes a copy of DNA, carries instructions for making proteins,
Supporting Information
 Supporting Information α,β-d-cna Preorganization of unpaired loop moiety stabilize DNA hairpin Christelle Dupouy, Pierre Millard, Arnaud Boissonnet and Jean-Marc Escudier* Laboratoire de Synthèse et Physico-Chimie
Supporting Information α,β-d-cna Preorganization of unpaired loop moiety stabilize DNA hairpin Christelle Dupouy, Pierre Millard, Arnaud Boissonnet and Jean-Marc Escudier* Laboratoire de Synthèse et Physico-Chimie
Sample Question Solutions for the Chemistry of Life Topic Test
 Sample Question Solutions for the Chemistry of Life Topic Test 1. Enzymes play a crucial role in biology by serving as biological catalysts, increasing the rates of biochemical reactions by decreasing
Sample Question Solutions for the Chemistry of Life Topic Test 1. Enzymes play a crucial role in biology by serving as biological catalysts, increasing the rates of biochemical reactions by decreasing
Electronic Supplementary Information
 Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Label-free and enzyme-free platform with visible output
Electronic Supplementary Material (ESI) for Chemical Science. This journal is The Royal Society of Chemistry 2017 Electronic Supplementary Information Label-free and enzyme-free platform with visible output
Supplementary Information
 Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Solution of a System of ODEs with POLYMATH and MATLAB, Boundary Value Iterations with MATLAB
 dy Solution of a System of ODEs with POLYMATH and MATLAB, Boundary Value Iterations with MATLAB For a system of n simultaneous first-order ODEs: dy1 = f1( y1, y2, K yn, x) dx dy2 = f 2( y1, y2, K yn, x)
dy Solution of a System of ODEs with POLYMATH and MATLAB, Boundary Value Iterations with MATLAB For a system of n simultaneous first-order ODEs: dy1 = f1( y1, y2, K yn, x) dx dy2 = f 2( y1, y2, K yn, x)
The body has three primary lines of defense against changes in hydrogen ion concentration in the body fluids.
 ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
ph and Nucleic acids Hydrogen Ion (H+) concentration is precisely regulated. The H+ concentration in the extracellular fluid is maintained at a very low level, averaging 0.00000004Eq/L. normal variations
Supplemental Materials and Methods
 Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
Supplemental Materials and Methods Time-resolved FRET (trfret) to probe for changes in the Box A/A stem upon complex assembly U3 MINI was folded and the decay of Fl fluorescence was measured at 20 ºC (see
+ regulation. ribosomes
 central dogma + regulation rpl DNA tsx rrna trna mrna ribosomes tsl ribosomal proteins structural proteins transporters enzymes srna regulators RNAp DNAp tsx initiation control by transcription factors
central dogma + regulation rpl DNA tsx rrna trna mrna ribosomes tsl ribosomal proteins structural proteins transporters enzymes srna regulators RNAp DNAp tsx initiation control by transcription factors
Velocity distribution of ideal gas molecules
 August 25, 2006 Contents Review of thermal movement of ideal gas molecules. Contents Review of thermal movement of ideal gas molecules. Distribution of the velocity of a molecule in ideal gas. Contents
August 25, 2006 Contents Review of thermal movement of ideal gas molecules. Contents Review of thermal movement of ideal gas molecules. Distribution of the velocity of a molecule in ideal gas. Contents
Pseudo translation and Twinning
 Pseudo translation and Twinning Crystal peculiarities Pseudo translation Twin Order-disorder Pseudo Translation Pseudo translation b Real space a Reciprocal space Distance between spots: 1/a, 1/b Crystallographic
Pseudo translation and Twinning Crystal peculiarities Pseudo translation Twin Order-disorder Pseudo Translation Pseudo translation b Real space a Reciprocal space Distance between spots: 1/a, 1/b Crystallographic
Lesson Overview. Ribosomes and Protein Synthesis 13.2
 13.2 The Genetic Code The first step in decoding genetic messages is to transcribe a nucleotide base sequence from DNA to mrna. This transcribed information contains a code for making proteins. The Genetic
13.2 The Genetic Code The first step in decoding genetic messages is to transcribe a nucleotide base sequence from DNA to mrna. This transcribed information contains a code for making proteins. The Genetic
Using DNA strand displacement to control interactions in DNA-grafted colloids
 Using DNA strand displacement to control interactions in DNA-grafted colloids The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters
Using DNA strand displacement to control interactions in DNA-grafted colloids The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Types of RNA. 1. Messenger RNA(mRNA): 1. Represents only 5% of the total RNA in the cell.
 RNAs L.Os. Know the different types of RNA & their relative concentration Know the structure of each RNA Understand their functions Know their locations in the cell Understand the differences between prokaryotic
RNAs L.Os. Know the different types of RNA & their relative concentration Know the structure of each RNA Understand their functions Know their locations in the cell Understand the differences between prokaryotic
Lecture 8: RNA folding
 1/16, Lecture 8: RNA folding Hamidreza Chitsaz Colorado State University chitsaz@cs.colostate.edu Fall 2018 September 13, 2018 2/16, Nearest Neighbor Model 3/16, McCaskill s Algorithm for MFE Structure
1/16, Lecture 8: RNA folding Hamidreza Chitsaz Colorado State University chitsaz@cs.colostate.edu Fall 2018 September 13, 2018 2/16, Nearest Neighbor Model 3/16, McCaskill s Algorithm for MFE Structure
Bi 8 Lecture 11. Quantitative aspects of transcription factor binding and gene regulatory circuit design. Ellen Rothenberg 9 February 2016
 Bi 8 Lecture 11 Quantitative aspects of transcription factor binding and gene regulatory circuit design Ellen Rothenberg 9 February 2016 Major take-home messages from λ phage system that apply to many
Bi 8 Lecture 11 Quantitative aspects of transcription factor binding and gene regulatory circuit design Ellen Rothenberg 9 February 2016 Major take-home messages from λ phage system that apply to many
Palindromes in Viral Genomes. Ming-Ying Leung Department of Mathematical Sciences The University of Texas at El Paso (UTEP)
 Palindromes in Viral Genomes Ming-Ying Leung Department of Mathematical Sciences The University of Texas at El Paso (UTEP) Outline: Palindromes Viral Genomes Roles of palindromes in DNA and RNA viruses
Palindromes in Viral Genomes Ming-Ying Leung Department of Mathematical Sciences The University of Texas at El Paso (UTEP) Outline: Palindromes Viral Genomes Roles of palindromes in DNA and RNA viruses
Organic Chemistry Option II: Chemical Biology
 Organic Chemistry Option II: Chemical Biology Recommended books: Dr Stuart Conway Department of Chemistry, Chemistry Research Laboratory, University of Oxford email: stuart.conway@chem.ox.ac.uk Teaching
Organic Chemistry Option II: Chemical Biology Recommended books: Dr Stuart Conway Department of Chemistry, Chemistry Research Laboratory, University of Oxford email: stuart.conway@chem.ox.ac.uk Teaching
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
Controlling the Catalytic Functions of DNAzymes within Constitutional. Dynamic Networks of DNA Nanostructures
 Supporting Information Controlling the Catalytic Functions of DNAzymes within Constitutional Dynamic Networks of DNA Nanostructures Shan Wang,, Liang Yue,, Zohar Shpilt, Alessandro Cecconello, Jason S.
Supporting Information Controlling the Catalytic Functions of DNAzymes within Constitutional Dynamic Networks of DNA Nanostructures Shan Wang,, Liang Yue,, Zohar Shpilt, Alessandro Cecconello, Jason S.
Lecture Notes J: Chemical Equilibrium I
 Lecture Notes J: Chemical Equilibrium I 1) Motivation: Binding/Unbinding of biological molecules How do we think about reactions where two molecules stick together/ unstick together: Protein + Drug Protein:Drug
Lecture Notes J: Chemical Equilibrium I 1) Motivation: Binding/Unbinding of biological molecules How do we think about reactions where two molecules stick together/ unstick together: Protein + Drug Protein:Drug
Chemistry 112 Name Practice Exam 2C Section
 Chemistry 112 Name Practice Exam 2C Section email IMPORTANT: On the scantron (answer sheet), you MUST clearly fill your name, your student number, section number, and test form (white cover = test form
Chemistry 112 Name Practice Exam 2C Section email IMPORTANT: On the scantron (answer sheet), you MUST clearly fill your name, your student number, section number, and test form (white cover = test form
Specific Interactions between Silver(I) Ions and Cytosine Cytosine Pairs in DNA Duplexes
 Supporting Information Specific Interactions between Silver(I) Ions and Cytosine Cytosine Pairs in DNA Duplexes Akira Ono*, Shiqi Cao, Humika Togashi, Mitsuru Tashiro, Takashi Fujimoto, Tomoya Machinami,
Supporting Information Specific Interactions between Silver(I) Ions and Cytosine Cytosine Pairs in DNA Duplexes Akira Ono*, Shiqi Cao, Humika Togashi, Mitsuru Tashiro, Takashi Fujimoto, Tomoya Machinami,
Stochastic processes and
 Stochastic processes and Markov chains (part II) Wessel van Wieringen w.n.van.wieringen@vu.nl wieringen@vu nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University
Stochastic processes and Markov chains (part II) Wessel van Wieringen w.n.van.wieringen@vu.nl wieringen@vu nl Department of Epidemiology and Biostatistics, VUmc & Department of Mathematics, VU University
RNA-Strukturvorhersage Strukturelle Bioinformatik WS16/17
 RNA-Strukturvorhersage Strukturelle Bioinformatik WS16/17 Dr. Stefan Simm, 01.11.2016 simm@bio.uni-frankfurt.de RNA secondary structures a. hairpin loop b. stem c. bulge loop d. interior loop e. multi
RNA-Strukturvorhersage Strukturelle Bioinformatik WS16/17 Dr. Stefan Simm, 01.11.2016 simm@bio.uni-frankfurt.de RNA secondary structures a. hairpin loop b. stem c. bulge loop d. interior loop e. multi
CHAPTER 2: THE CHEMICAL CONTEXT OF LIFE AP Biology CASE STUDY: DEVIL S GARDEN MATTER. Figs. 2.1 & 2.2. Fig. 2.3
 CHAPTER 2: THE CHEMICAL CONTEXT OF LIFE AP Biology 1 CASE STUDY: DEVIL S GARDEN Ants use formic acid to maintain the garden of a single flowering tree called Duroia hirsuta Ants live in the hollow tree
CHAPTER 2: THE CHEMICAL CONTEXT OF LIFE AP Biology 1 CASE STUDY: DEVIL S GARDEN Ants use formic acid to maintain the garden of a single flowering tree called Duroia hirsuta Ants live in the hollow tree
More Protein Synthesis and a Model for Protein Transcription Error Rates
 More Protein Synthesis and a Model for Protein James K. Peterson Department of Biological Sciences and Department of Mathematical Sciences Clemson University October 3, 2013 Outline 1 Signal Patterns Example
More Protein Synthesis and a Model for Protein James K. Peterson Department of Biological Sciences and Department of Mathematical Sciences Clemson University October 3, 2013 Outline 1 Signal Patterns Example
Simulative and experimental characterization of a ph-dependent
 Simulative and experimental characterization of a ph-dependent clamp-like DNA triple-helix nanoswitch Federico Iacovelli, # Andrea Idili, # Alessandro Benincasa, Davide Mariottini, Alessio Ottaviani, Mattia
Simulative and experimental characterization of a ph-dependent clamp-like DNA triple-helix nanoswitch Federico Iacovelli, # Andrea Idili, # Alessandro Benincasa, Davide Mariottini, Alessio Ottaviani, Mattia
Structural basis for catalytically restrictive dynamics of a high-energy enzyme state
 Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supplementary Material Structural basis for catalytically restrictive dynamics of a high-energy enzyme state Michael Kovermann, Jörgen Ådén, Christin Grundström, A. Elisabeth Sauer-Eriksson, Uwe H. Sauer
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/9/e1500511/dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of
advances.sciencemag.org/cgi/content/full/1/9/e1500511/dc1 Supplementary Materials for Contractility parameters of human -cardiac myosin with the hypertrophic cardiomyopathy mutation R403Q show loss of
HMMs and biological sequence analysis
 HMMs and biological sequence analysis Hidden Markov Model A Markov chain is a sequence of random variables X 1, X 2, X 3,... That has the property that the value of the current state depends only on the
HMMs and biological sequence analysis Hidden Markov Model A Markov chain is a sequence of random variables X 1, X 2, X 3,... That has the property that the value of the current state depends only on the
Agampodi S. De Silva Indrasekara 1,2, Sean F Johnson 1, Ren A Odion 1,2, Tuan Vo-Dinh 1,2,3*
 Supporting Information: Manipulation of the geometry and modulation of the optical response of surfactant-free gold nanostars: a systematic bottom-up synthesis Agampodi S. De Silva Indrasekara 1,2, Sean
Supporting Information: Manipulation of the geometry and modulation of the optical response of surfactant-free gold nanostars: a systematic bottom-up synthesis Agampodi S. De Silva Indrasekara 1,2, Sean
From Gene to Protein
 From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
A rule of seven in Watson-Crick base-pairing of mismatched sequences
 A rule of seven in Watson-Crick base-pairing of mismatched sequences Ibrahim I. Cisse 1,3, Hajin Kim 1,2, Taekjip Ha 1,2 1 Department of Physics and Center for the Physics of Living Cells, University of
A rule of seven in Watson-Crick base-pairing of mismatched sequences Ibrahim I. Cisse 1,3, Hajin Kim 1,2, Taekjip Ha 1,2 1 Department of Physics and Center for the Physics of Living Cells, University of
Architecture based on the integration of intermolecular. G-quadruplex structure with sticky-end pairing and
 This journal is The Royal Society of Chemistry 0 Electronic Supplementary Information for: Architecture based on the integration of intermolecular G-quadruplex structure with sticky-end pairing and colorimetric
This journal is The Royal Society of Chemistry 0 Electronic Supplementary Information for: Architecture based on the integration of intermolecular G-quadruplex structure with sticky-end pairing and colorimetric
SUPPLEMENTARY INFORMATION
 Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Supplementary Results DNA binding property of the SRA domain was examined by an electrophoresis mobility shift assay (EMSA) using synthesized 12-bp oligonucleotide duplexes containing unmodified, hemi-methylated,
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
Problem Set 5 Question 1
 2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable
2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable
Chapter 9b: Numerical Methods for Calculus and Differential Equations. Initial-Value Problems Euler Method Time-Step Independence MATLAB ODE Solvers
 Chapter 9b: Numerical Methods for Calculus and Differential Equations Initial-Value Problems Euler Method Time-Step Independence MATLAB ODE Solvers Acceleration Initial-Value Problems Consider a skydiver
Chapter 9b: Numerical Methods for Calculus and Differential Equations Initial-Value Problems Euler Method Time-Step Independence MATLAB ODE Solvers Acceleration Initial-Value Problems Consider a skydiver
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Section 10/5/06. Junaid Malek, M.D.
 Section 10/5/06 Junaid Malek, M.D. Equilibrium Occurs when two interconverting states are stable This is represented using stacked arrows: State A State B, where State A and State B can represent any number
Section 10/5/06 Junaid Malek, M.D. Equilibrium Occurs when two interconverting states are stable This is represented using stacked arrows: State A State B, where State A and State B can represent any number
DINAMelt web server for nucleic acid melting prediction
 DINAMelt web server for nucleic acid melting prediction Nicholas R. Markham 1 and Michael Zuker 2, * Nucleic Acids Research, 2005, Vol. 33, Web Server issue W577 W581 doi:10.1093/nar/gki591 1 Department
DINAMelt web server for nucleic acid melting prediction Nicholas R. Markham 1 and Michael Zuker 2, * Nucleic Acids Research, 2005, Vol. 33, Web Server issue W577 W581 doi:10.1093/nar/gki591 1 Department
AP Biology. Chapter 2
 AP Biology Chapter 2 Matter is anything that has weight and takes up space 1. Mass is a measure of how much matter is present in a body 2. Weight is a measure of the gravitational force exerted on an object
AP Biology Chapter 2 Matter is anything that has weight and takes up space 1. Mass is a measure of how much matter is present in a body 2. Weight is a measure of the gravitational force exerted on an object
Lecture 7: Simple genetic circuits I
 Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
1. Contains the sugar ribose instead of deoxyribose. 2. Single-stranded instead of double stranded. 3. Contains uracil in place of thymine.
 Protein Synthesis & Mutations RNA 1. Contains the sugar ribose instead of deoxyribose. 2. Single-stranded instead of double stranded. 3. Contains uracil in place of thymine. RNA Contains: 1. Adenine 2.
Protein Synthesis & Mutations RNA 1. Contains the sugar ribose instead of deoxyribose. 2. Single-stranded instead of double stranded. 3. Contains uracil in place of thymine. RNA Contains: 1. Adenine 2.
Homework for Chapter 7 Chem 2310
 omework for Chapter 7 Chem 2310 Name I. Introduction to Reactions 1. Explain why the following fits the definition of a chemical reaction. C 3 Na C 3 Na 2. Using the chemical reaction above, give all compounds
omework for Chapter 7 Chem 2310 Name I. Introduction to Reactions 1. Explain why the following fits the definition of a chemical reaction. C 3 Na C 3 Na 2. Using the chemical reaction above, give all compounds
of the Guanine Nucleotide Exchange Factor FARP2
 Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Structure, Volume 21 Supplemental Information Structural Basis for Autoinhibition of the Guanine Nucleotide Exchange Factor FARP2 Xiaojing He, Yi-Chun Kuo, Tyler J. Rosche, and Xuewu Zhang Inventory of
Name Student number. UNIVERSITY OF GUELPH CHEM 4540 ENZYMOLOGY Winter 2002 Quiz #1: February 14, 2002, 11:30 13:00 Instructor: Prof R.
 UNIVERSITY OF GUELPH CHEM 4540 ENZYMOLOGY Winter 2002 Quiz #1: February 14, 2002, 11:30 13:00 Instructor: Prof R. Merrill Instructions: Time allowed = 90 minutes. Total marks = 30. This quiz represents
UNIVERSITY OF GUELPH CHEM 4540 ENZYMOLOGY Winter 2002 Quiz #1: February 14, 2002, 11:30 13:00 Instructor: Prof R. Merrill Instructions: Time allowed = 90 minutes. Total marks = 30. This quiz represents
Soft Matter PAPER. Using DNA strand displacement to control interactions in DNA-grafted colloids. 1 Introduction
 PAPER View Article Online View Journal View Issue Cite this:, 2018, 14, 969 Received 26th August 2017, Accepted 3rd January 2018 DOI: 10.1039/c7sm01722g rsc.li/soft-matter-journal 1 Introduction In this
PAPER View Article Online View Journal View Issue Cite this:, 2018, 14, 969 Received 26th August 2017, Accepted 3rd January 2018 DOI: 10.1039/c7sm01722g rsc.li/soft-matter-journal 1 Introduction In this
CORE MOLIT ACTIVITIES at a glance
 CORE MOLIT ACTIVITIES at a glance 1. Amplification of Biochemical Signals: The ELISA Test http://molit.concord.org/database/activities/248.html The shape of molecules affects the way they function. A test
CORE MOLIT ACTIVITIES at a glance 1. Amplification of Biochemical Signals: The ELISA Test http://molit.concord.org/database/activities/248.html The shape of molecules affects the way they function. A test
SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question.
 ch 2 chemical basis of life Name SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question. Fill in the blank or provide a short answer: 1) When a change in matter
ch 2 chemical basis of life Name SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question. Fill in the blank or provide a short answer: 1) When a change in matter
Computing Radioimmunoassay Analysis Dose-Response Curves
 Computing Radioimmunoassay Analysis Dose-Response Curves Gary D. Knott, Ph.D. Civilized Software, Inc. 12109 Heritage Park Circle Silver Spring MD 20906 Tel.: (301)-962-3711 email: csi@civilized.com URL:
Computing Radioimmunoassay Analysis Dose-Response Curves Gary D. Knott, Ph.D. Civilized Software, Inc. 12109 Heritage Park Circle Silver Spring MD 20906 Tel.: (301)-962-3711 email: csi@civilized.com URL:
Chemistry 12 June 2003 Provincial Examination
 Chemistry 12 June 2003 Provincial Examination ANSWER KEY / SCORING GUIDE CURRICULUM: Organizers 1. Reaction Kinetics 2. Dynamic Equilibrium 3. Solubility Equilibria 4. Acids, Bases, and Salts 5. Oxidation
Chemistry 12 June 2003 Provincial Examination ANSWER KEY / SCORING GUIDE CURRICULUM: Organizers 1. Reaction Kinetics 2. Dynamic Equilibrium 3. Solubility Equilibria 4. Acids, Bases, and Salts 5. Oxidation
Visible and IR Absorption Spectroscopy. Andrew Rouff and Kyle Chau
 Visible and IR Absorption Spectroscopy Andrew Rouff and Kyle Chau The Basics wavelength= (λ) original intensity= Ι o sample slab thickness= dl Final intensity= I f ε = molar extinction coefficient -di=
Visible and IR Absorption Spectroscopy Andrew Rouff and Kyle Chau The Basics wavelength= (λ) original intensity= Ι o sample slab thickness= dl Final intensity= I f ε = molar extinction coefficient -di=
Chapter 17. From Gene to Protein. Biology Kevin Dees
 Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Chapter 17 From Gene to Protein DNA The information molecule Sequences of bases is a code DNA organized in to chromosomes Chromosomes are organized into genes What do the genes actually say??? Reflecting
Name: Regents Chemistry Review Packet B1
 Name: Regents Chemistry Review Packet B1 1. Compared to an electron, which particle has a charge that is equal in magnitude but opposite in sign? an alpha particle a beta particle a neutron a proton 2.
Name: Regents Chemistry Review Packet B1 1. Compared to an electron, which particle has a charge that is equal in magnitude but opposite in sign? an alpha particle a beta particle a neutron a proton 2.
University of Alberta ENGM 541: Modeling and Simulation of Engineering Systems Laboratory #5
 University of Alberta ENGM 54: Modeling and Simulation of Engineering Systems Laboratory #5 M.G. Lipsett, Updated 00 Integration Methods with Higher-Order Truncation Errors with MATLAB MATLAB is capable
University of Alberta ENGM 54: Modeling and Simulation of Engineering Systems Laboratory #5 M.G. Lipsett, Updated 00 Integration Methods with Higher-Order Truncation Errors with MATLAB MATLAB is capable
Basic Synthetic Biology circuits
 Basic Synthetic Biology circuits Note: these practices were obtained from the Computer Modelling Practicals lecture by Vincent Rouilly and Geoff Baldwin at Imperial College s course of Introduction to
Basic Synthetic Biology circuits Note: these practices were obtained from the Computer Modelling Practicals lecture by Vincent Rouilly and Geoff Baldwin at Imperial College s course of Introduction to
CHEMISTRY 110 EXAM 2 Oct.
 EMISTRY 110 EXAM 2 ct. 15, 2012 FRM A 1. Which of the following is an incorrect formula for a neutral compound made from the given ions?! "#$%&'! #'%&'! (&)*+,#! -.!! aluminum oxide Al 2 3 /.!! magnesium
EMISTRY 110 EXAM 2 ct. 15, 2012 FRM A 1. Which of the following is an incorrect formula for a neutral compound made from the given ions?! "#$%&'! #'%&'! (&)*+,#! -.!! aluminum oxide Al 2 3 /.!! magnesium
Kinetics Mechanisms (2008-rev) Review and Examples
 II 25 Kinetics Mechanisms (2008-rev) Review and Examples Mechanism: series of elementary steps (uni-, bimolecular) that combine to give observed rate law elementary step - reaction order lie stoichiometry
II 25 Kinetics Mechanisms (2008-rev) Review and Examples Mechanism: series of elementary steps (uni-, bimolecular) that combine to give observed rate law elementary step - reaction order lie stoichiometry
CS-E5880 Modeling biological networks Gene regulatory networks
 CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
Time-dependence of key H-bond/electrostatic interaction distances in the sirna5-hago2 complexes... Page S14
 Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
Supporting Information Probing the Binding Interactions between Chemically Modified sirnas and Human Argonaute 2 Using Microsecond Molecular Dynamics Simulations S. Harikrishna* and P. I. Pradeepkumar*
I N N O V A T I O N L E C T U R E S (I N N O l E C)
 I N N O V A T I O N L E C T U R E S (I N N O l E C) Binding and Kinetics for Experimental Biologists Lecture 1 Numerical Models for Biomolecular Interactions Petr Kuzmič, Ph.D. BioKin, Ltd. WATERTOWN,
I N N O V A T I O N L E C T U R E S (I N N O l E C) Binding and Kinetics for Experimental Biologists Lecture 1 Numerical Models for Biomolecular Interactions Petr Kuzmič, Ph.D. BioKin, Ltd. WATERTOWN,
1985B4. A kilogram sample of a material is initially a solid at a temperature of 20 C. Heat is added to the sample at a constant rate of 100
 1985B4. A 0.020-kilogram sample of a material is initially a solid at a temperature of 20 C. Heat is added to the sample at a constant rate of 100 joules per second until the temperature increases to 60
1985B4. A 0.020-kilogram sample of a material is initially a solid at a temperature of 20 C. Heat is added to the sample at a constant rate of 100 joules per second until the temperature increases to 60
Bimolecular processes
 Bimolecular processes Electron transfer *A + B A + + B - *A + B A - + B + EA IP *EA *IP LUMO An excited state is a better oxidant and a better reductant than the ground state HOMO X X* Kinetic of electron
Bimolecular processes Electron transfer *A + B A + + B - *A + B A - + B + EA IP *EA *IP LUMO An excited state is a better oxidant and a better reductant than the ground state HOMO X X* Kinetic of electron
Sample Questions Chem 22 Student Chapters Page 1 of 5 Spring 2016
 Sample Questions Chem 22 Student Chapters 13-18 Page 1 of 5 1. The vapor pressure of a liquid is the pressure, at equilibrium, of the a) solid above its liquid. b) liquid above its solid. c) gas above
Sample Questions Chem 22 Student Chapters 13-18 Page 1 of 5 1. The vapor pressure of a liquid is the pressure, at equilibrium, of the a) solid above its liquid. b) liquid above its solid. c) gas above
The ideal fiber pattern exhibits 4-quadrant symmetry. In the ideal pattern the fiber axis is called the meridian, the perpendicular direction is
 Fiber diffraction is a method used to determine the structural information of a molecule by using scattering data from X-rays. Rosalind Franklin used this technique in discovering structural information
Fiber diffraction is a method used to determine the structural information of a molecule by using scattering data from X-rays. Rosalind Franklin used this technique in discovering structural information
Chapter 2: The Chemical Basis of Life
 Chapter 2: The Chemical Basis of Life I. Basic Chemistry A. Matter, Mass, and Weight 1. All living and nonliving things are composed of 2. represents the amount of matter. 3. is caused by the gravitational
Chapter 2: The Chemical Basis of Life I. Basic Chemistry A. Matter, Mass, and Weight 1. All living and nonliving things are composed of 2. represents the amount of matter. 3. is caused by the gravitational
(Lys), resulting in translation of a polypeptide without the Lys amino acid. resulting in translation of a polypeptide without the Lys amino acid.
 1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
1. A change that makes a polypeptide defective has been discovered in its amino acid sequence. The normal and defective amino acid sequences are shown below. Researchers are attempting to reproduce the
Supplementary Information for: Integrating DNA Strand Displacement Circuitry with DNA Tile Self-assembly
 Supplementary Information for: Integrating DN Strand Displacement Circuitry with DN Tile Self-assembly David Yu Zhang, Rizal F. Hariadi, Harry M. T. Choi, and Erik Winfree (Dated: June, 03) Table of Contents:
Supplementary Information for: Integrating DN Strand Displacement Circuitry with DN Tile Self-assembly David Yu Zhang, Rizal F. Hariadi, Harry M. T. Choi, and Erik Winfree (Dated: June, 03) Table of Contents:
Chapter Practice Test
 Name: Class: Date: Chapter 17-18 Practice Test Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Examining a chemical system before and after a reaction
Name: Class: Date: Chapter 17-18 Practice Test Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Examining a chemical system before and after a reaction
Chapter 1: Introduction
 Chapter 1: Introduction Definition: A differential equation is an equation involving the derivative of a function. If the function depends on a single variable, then only ordinary derivatives appear and
Chapter 1: Introduction Definition: A differential equation is an equation involving the derivative of a function. If the function depends on a single variable, then only ordinary derivatives appear and
LUMO + 1 LUMO. Tómas Arnar Guðmundsson Report 2 Reikniefnafræði G
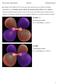 Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
Q1: Display all the MOs for N2 in your report and classify each one of them as bonding, antibonding or non-bonding, and say whether the symmetry of the orbital is σ or π. Sketch a molecular orbital diagram
SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question.
 Exam Name SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question. Figure 2.1 Using Figure 2.1, match the following: 1) Lipid. 2) Functional protein. 3) Nucleotide.
Exam Name SHORT ANSWER. Write the word or phrase that best completes each statement or answers the question. Figure 2.1 Using Figure 2.1, match the following: 1) Lipid. 2) Functional protein. 3) Nucleotide.
Chemistry. Essential Standards Chemistry
 Essential Standards Chemistry Chemistry Matter: Properties & Change 1.1 Students will analyze the structure of atoms and ions. 1.2 Student will understand the bonding that occurs in simple compounds in
Essential Standards Chemistry Chemistry Matter: Properties & Change 1.1 Students will analyze the structure of atoms and ions. 1.2 Student will understand the bonding that occurs in simple compounds in
AR-7781 (Physical Chemistry)
 Model Answer: B.Sc-VI th Semester-CBT-602 AR-7781 (Physical Chemistry) One Mark Questions: 1. Write a nuclear reaction for following Bethe s notation? 35 Cl(n, p) 35 S Answer: 35 17Cl + 1 1H + 35 16S 2.
Model Answer: B.Sc-VI th Semester-CBT-602 AR-7781 (Physical Chemistry) One Mark Questions: 1. Write a nuclear reaction for following Bethe s notation? 35 Cl(n, p) 35 S Answer: 35 17Cl + 1 1H + 35 16S 2.
downstream (0.8 kb) homologous sequences to the genomic locus of DIC. A DIC mutant strain (ro- 6
 A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
A B C D ts Figure S1 Generation of DIC- mcherry expressing N.crassa strain. A. N. crassa colony morphology. When a cot1 (top, left panel) strain is grown at permissive temperature (25 C), it exhibits straight
