Problem Set 5 Question 1
|
|
|
- Christiana West
- 6 years ago
- Views:
Transcription
1 2.32 Problem Set 5 Question As discussed in class, drug discovery often involves screening large libraries of small molecules to identify those that have favorable interactions with a certain druggable target. One of the first tests to determine whether a small molecule could potentially have a therapeutic effect is to measure its binding affinity to the target of interest. These are typically performed in highthroughput assays to select for those compounds that bind the target with the highest affinity. A) Briefly discuss the role of binding affinity in determining the overall therapeutic effect of a small molecule. At this stage in the drug discovery process, given a list of compounds and their affinities to a particular target, what can you conclude about the relative effectiveness of each compound as a drug? Binding affinity between a small molecule and its target is an important factor for determining therapeutic effect. In order for a small molecule to affect a target, it must associate with the target long enough to produce the desired change. However, binding affinity is not the sole parameter that must be optimized. In other words, binding tighter to the target does not guarantee a better therapeutic outcome. This stage of the drug discovery process seeks to identify compounds that associate with the target molecule with at least a certain affinity. Choosing between compounds that meet this criteria happens based on results of preclinical and clinical trials. Downstream screening steps reveal a small-molecule lead (code-named SML, MW = 34 Da) to be a promising candidate. Results from the initial high-throughput screen show that SML binds to its target TAR2 (MW = 98 kda) with K d = 29 nm. You are interested in improving the affinity of this interaction. In order to do this, you decide to perform an isothermal titration calorimetry (ITC) experiment to deduce more detail about the thermodynamics of the binding reaction. B) Design the ITC experiment you would use to determine the entropic vs. enthalpic contributions to binding affinity between SML and TAR2. Be sure to specify/calculate the following quantities, based on guidelines discussed in class. - The concentrations of TAR2 and SML - The number of injections and the injection volume - The volume of the sample cell - The total amount [i.e. the mass] of purified target protein your experiment will require - The total amount [i.e. the mass] of SML your experiment will use. ITC experiments use two cells filled with solution a reference cell (where no reaction is taking place), and a sample cell, with target protein in solution. In this case, TAR2 (at a concentration x the K d of the binding reaction) would be placed in a ml sample cell, and µl of SML would be injected 2 times at a concentration x that of TAR2. Because the system aims to keep both cells at the same temperature throughout the experiment (and because SML binding TAR2 is an exothermic reaction), the amount of heat applied to the sample cell would decrease briefly when SML is injected into the sample cell. Integration of these peaks yields points on a binding isotherm, from which binding data can be extracted.
2 2.32 Problem Set 5 Question As discussed in lecture, ITC experiments typically require a protein concentration roughly - fold above the K d of the drug-protein interaction. Assuming a cell volume of ml: [TAR2] = 29 nm = 2.9 µm mol L 3 L = mol TAR mol 98, g mol = 284 µg TAR2 For this design, twice as much SML is required compared to TAR2. Therefore twice as many moles of SML are required mol 34 g mol =82 µg SML Students answers do not need to match these solutions exactly, but their calculations should be consistent with the values they specified in their design. C) For the experimental set-up you described in Part B), calculate the amount of heat absorbed or released after the third injection of ligand, given H = -.7 kcal/mol. Assume that each injection is µl, and that the injected solution consists of SML at a concentration of 29 µm (i.e. x the concentration of TAR2). Using the equation below, [C] = y[p] = (K d + [L] + [P] ) (K d + [L] + [P] 2 ) 2 4[L] [P] In the above setup, we chose a value of [P] (i.e. [TAR2] ) that was x the K d of TAR2 binding SML. After the second injection, 2 µl of a 29 µm solution of SML has been added to a starting volume of ml. Therefore: L) 2 6 L) (29 6 mol ( = M,2 3 L L [L] = [P] = M Therefore: [ C] = y[ P] = 2 + d [ ] + [ ] + d [ ] + (K L P ) (K L P ) 2 2 [ ] 4[L] [P] Similarly, after the third injection: [ L] = [C] = (29 6 mol 6 L)(3 L) = 7 M 3 L L,3 8.4 y[p] = 3 (K d + [L] K + [ P] ) ( + d [L] + [ 2 = 5. 7 M P] 2 ) 4[L] [P] = M 2
3 2.32 Problem Set 5 Question Therefore: Q 3 = ΔH([C 3 ] [C 2 ])V cell = (.7 kcal mol)( mol L)( 3 L) Q 3 = kcal, or cal D) Describe the pseudo-first order approximation (both mathematically and physically) and discuss whether this is a valid assumption for ITC. The pseudo-first order approximation can be used when ligand is in excess, and the concentration of free ligand in the solution can be approximated as constant as the system approaches equilibrium. In other words, binding protein to excess ligand causes a negligible decrease in the amount of free ligand in solution. Mathematically, this corresponds to setting [L] = [L]. eq The equation for fractional saturation of protein at equilibrium is: y eq = [L] eq [L], and applying the pseudo-first order approximation, yeq [L] K + eq d [L] + K d This is not appropriate for use in ITC calculations, because ITC involves injecting small amounts of ligand into a solution of excess free protein and measuring the heat evolved as the system reaches equilibrium. Therefore ligand is not in excess when heat evolution is measured, and the depletion of ligand must be considered when performing ITC calculations. E) Why are ITC experiments only accurate within a certain range of binding affinities? For very strong interactions, saturation can be reached very rapidly and abruptly without providing data points in the middle to allow us to determine the K d accurately. For weak binders, the opposite problem occurs: each additional injection may not generate enough new complex formation to register on the detector and give reliable readout. Therefore, the protein-ligand affinity must be such that injections of ligand will result in a full range of fractional saturations. 3
4 2.32 Problem Set 5 Question 2 Dasatinib (Sprycell ) is a small molecule tyrosine kinase inhibitor developed by Bristol- Myers Squibb with fairly exquisite specificity for Src family kinases and Abl family kinases, with low affinity for most other kinases. This drug is currently used as secondline therapy for chronic myeloid leukemia patients that have developed resistance to Gleevec, the first-line therapeutic. Some initial data has emerged that Dasatinib might have an off-target effect in which it is binding to and inhibiting the EphB4 receptor tyrosine kinase. Initial estimates of the K d for this interaction are 5 x -8 M. To better quantify the affinity and kinetics of this interaction, a Surface Plasmon Resonance (SPR) experiment was performed in which 45 fmoles of Dasatinib was immobilized onto a sensor chip with dimensions.25 cm.25 cm. EphB4 at a concentration of 5 nm was then washed over the immobilized Dasatinib, and the reaction was allowed to reach equilibrium. A) Why was Dasatinib (rather than EphB4) immobilized onto the SPR sensor chip? This configuration was chosen to maximize signal. Immobilizing the small molecule on the surface of the sensor chip results in a large change in mass when a large macromolecule like an enzyme binds. Since the change in resonance angle (and therefore, the RU) is proportional to the change in mass of each immobilized species upon binding, this configuration maximizes the dynamic range, and therefore the sensitivity of the experiment. B) Given k on = 8.4 x 4 L mol - sec - and k off = 3. x -3 sec -, calculate the time required for this system to reach equilibrium. ln2 ln2 τ = = = 59 sec 2 k on [L ] + koff L 9 mol 3 mol s [5 L ] + 3. s C) How much of the Dasatinib on the surface is bound by EphB4 at equilibrium? 9 [L] [L] + 5 M y eq.23 K = 5 9 M M = d.23(45 fmol) = 4 fmol 4
5 2.32 Problem Set 5 Question 2 D) Determine the maximum Resonance Units (RUs) measured for this binding interaction. Assume a molecular weight of 488 Da for Dasatinib, and 8 kda for EphB4. The maximum signal (in RU) will result when every immobilized Dasatinib molecule is bound to one molecule of EphB4. Assuming that this is a : binding reaction, the total amount of material bound to the chip can be calculated. (45 5 mol complex)( molecules mol) = 2.7 molecules complex Da.66 g pg 2.7 molecules) = pg complex molecule complex Da g ( pg pg RUmax = = mm 2 mm
6 2.32 Problem Set 5 Question 3 You are interested in investigating the interaction between a protein kinase KIN and its substrate SUB. While it is well established that KIN is primarily responsible for phosphorylating SUB, the mechanism of SUB-mediated signaling is poorly understood. To date, twenty other proteins have been identified as potential interacting partners for SUB, which has been shown to possess six accessible phosphorylation sites. You decide that the next step in probing this signaling pathway is to determine which phosphorylation sites on SUB are bound by the SH2 domains of these 2 proteins. A) Design a protein microarray experiment to address this question. In your experimental design, be sure to [briefly] discuss: - Experimental setup: What will be immobilized on your microarray, and how will you detect binding? - Expected results: Draw what you anticipate your data will look like for a representative cell in your microarray. What parameter will your data help you calculate? - Interpretation: How do these calculations help you determine the relative specificity of the SH2 domains for different tyrosine residues on SUB? For this experiment, SH2 domains of the 2 proteins of interest will be expressed recombinantly and immobilized on the surface of the microarray. Fluorescently labeled peptides from SUB, each containing the phosphorylated residue of interest, will be washed over the microarray, and fluorescent spots will be read by an imager. Representative data would resemble a saturation curve for each cell, which allows a K d to be calculated. Fobs [L] Comparing the K d of SH2 domains between different pairs of proteins and phosphorylation sites will determine which protein-phosphosite interactions are most favorable. Proteins whose SH2 domain has the lowest K d value for a specific phosphorylation site are likely to be involved with signaling pathways that are affected by the phosphorylation of a specific residue. 6
7 2.32 Problem Set 5 Question 3 A small subset of the data from this experiment is posted on the Course website. The file PS5Q3data.mat contains raw microarray readouts from the SH2 domain of SUB7 (a candidate SUB interaction partner) binding to each phosphorylation site on SUB. The names of the phosphorylation sites are stored in the cell array phosphosites. Each row represents a series of binding experiments using a single phosphorylation site with the columns representing nine initial ligand concentrations. These concentrations (in nm) are stored in the vector L. B) Begin your analysis by loading these results into MATLAB and transforming each data point into a fractional saturation [Hint: assume that the maximum measured value in the entire experiment corresponds to full saturation]. In a single figure, plot the fractional saturation of each phosphorylation site with respect to initial ligand concentration. Each site should be represented by a subplot in your figure, and should have consistent axes to facilitate direct comparison. To receive full credit for this problem, you must automate the data transformation and plotting processes (i.e. you should only need one line of code to transform all your data, and you should only call the plot function once for this part of the problem). You may need to employ the max command and use a for loop. Note: phosphosites is a cell array and does not require quotation marks or apostrophes when called from the ylabel, xlabel, or title commands in MATLAB. load PS5Q3data.mat % Transform data into fractional saturations by dividing by the absolute maximum y = Data./(max(max(Data))); % Plot fractional saturations in individual subplots figure() for i = [:6] subplot(2,3,i) plot(l, y(i,:),'o') ylabel('') xlabel('[l]_ ') title(phosphosites(i)) axis([ 25 ]) end 7
8 2.32 Problem Set 5 Question 3.5 py 2 [L] py4.5 2 [L].5 py26 2 [L] py [L].5 py37 2 [L] py [L] C) Using nlinfit, calculate the K d for the SH2 domain of SUB7 interacting with each phosphorylation site on SUB. You will need to call nlinfit individually for each data set. Be sure to follow the coding guidelines established in Part B). For each phosphorylation site (py, py26, py37, py4, py49, and py58): Kd = See Part D) for code. D) In a new figure, overlay the binding curves based on the K d values you calculated with the values you obtained in Part B). Use the linspace command to give you an appropriate basis of x-values from which to compute the points for your binding curves. % Initialize array of Kd values which will be filled later Kd = zeros(,6); % Plot fractional saturations with binding isotherms overlaid figure(2) x = linspace(, 25); % Basis vector for drawing binding curves for i = [:6] subplot(2,3,i) Kd(,i) = nlinfit(l, ); % Fit data to model y_fit = isotherm(kd(,i), x); % Calculate points for binding curves plot(l, y(i,:), 'o', x, y_fit); % Plot data and binding curves ylabel('') xlabel('[l]_ ') title(phosphosites(i)) axis([ 25 ]) end 8
9 2.32 Problem Set 5 Question 3 Kd function out = isotherm(kd, L) out = L./(L + Kd); py 2 [L] py [L] py26 2 [L] py [L] py37 2 [L] py [L] E) Based on your analysis, which phosphorylation site on SUB is likely responsible for the most downstream signaling through SUB7? py4 on SUB has the highest affinity for the SH2 domain of SUB7, and therefore is most likely involved with SUB7-mediated signaling. F) A key step in evaluating the fit of a model is examining the residuals between the model and the data. In a new figure, plot these residuals. Was your model a good fit for your binding data? Again, be sure to follow the coding guidelines established in Part B). % Calculate and plot residuals figure(3) for i = [:6] y_resids = isotherm(kd(,i), L) - y(i,:); % Calculate residuals subplot(2,3,i) plot(l, y_resids) ylabel('') xlabel('[l]_ ') title(phosphosites(i)) end 9
10 2.32 Problem Set 5 Question 3 $% 89"" $% 89#: $% 89;< $% " # 234 / %" $% $% " # 234 /5-67 $% " # 234 / %= $% $% " # 234 /5-67 &'()*+,-(./(*'(*+,- &'()*+,-(./(*'(*+,- &'()*+,-(./(*'(*+,- &'()*+,-(./(*'(*+,- &'()*+,-(./(*'(*+,- &'()*+,-(./(*'(*+,- $% " # 234 / >? $% $% " # 234 /5-67 The model appears to be a good fit for the binding data, as the residuals are small ( 3% variation from the model) and do not show systematic error.
11 MIT OpenCourseWare Analysis of Biomolecular and Cellular Systems Fall 22 For information about citing these materials or our Terms of Use, visit:
Problem Set # 1
 20.320 Problem Set # 1 September 17 th, 2010 Due on September 24 th, 2010 at 11:59am. No extensions will be granted. General Instructions: 1. You are expected to state all your assumptions and provide
20.320 Problem Set # 1 September 17 th, 2010 Due on September 24 th, 2010 at 11:59am. No extensions will be granted. General Instructions: 1. You are expected to state all your assumptions and provide
MicroCal itc 200. System MicroCal Auto-iTC 200. System. GE Healthcare Life Sciences. System design and description. provide:
 GE Healthcare Life Sciences Data file 28-97822 AC MicroCal label-free interaction analysis MicroCal itc 2 System MicroCal Auto-iTC 2 System MicroCal itc 2 and MicroCal Auto-iTC 2 isothermal titration calorimetry
GE Healthcare Life Sciences Data file 28-97822 AC MicroCal label-free interaction analysis MicroCal itc 2 System MicroCal Auto-iTC 2 System MicroCal itc 2 and MicroCal Auto-iTC 2 isothermal titration calorimetry
Isothermal Titration Calorimetry in Drug Discovery. Geoff Holdgate Structure & Biophysics, Discovery Sciences, AstraZeneca October 2017
 Isothermal Titration Calorimetry in Drug Discovery Geoff Holdgate Structure & Biophysics, Discovery Sciences, AstraZeneca October 217 Introduction Introduction to ITC Strengths / weaknesses & what is required
Isothermal Titration Calorimetry in Drug Discovery Geoff Holdgate Structure & Biophysics, Discovery Sciences, AstraZeneca October 217 Introduction Introduction to ITC Strengths / weaknesses & what is required
ISoTherMal TITraTIon Calorimetry
 ISoTherMal TITraTIon Calorimetry With the Nano ITC, heat effects as small as 1 nanojoules are detectable using one nanomole or less of biopolymer. The Nano ITC uses a solid-state thermoelectric heating
ISoTherMal TITraTIon Calorimetry With the Nano ITC, heat effects as small as 1 nanojoules are detectable using one nanomole or less of biopolymer. The Nano ITC uses a solid-state thermoelectric heating
Problem solving steps
 Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
Problem solving steps Determine the reaction Write the (balanced) equation ΔG K v Write the equilibrium constant v Find the equilibrium constant using v If necessary, solve for components K K = [ p ] ν
schematic diagram; EGF binding, dimerization, phosphorylation, Grb2 binding, etc.
 Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
Lecture 1: Noncovalent Biomolecular Interactions Bioengineering and Modeling of biological processes -e.g. tissue engineering, cancer, autoimmune disease Example: RTK signaling, e.g. EGFR Growth responses
17. Biomolecular Interaction
 17. Biomolecular Interaction Methods for characterizing biomolecular interactions Sequence-specific DNA binding ligands Molecular mechanisms of drug action and drug resistance In silico compound design
17. Biomolecular Interaction Methods for characterizing biomolecular interactions Sequence-specific DNA binding ligands Molecular mechanisms of drug action and drug resistance In silico compound design
From Friday s material
 5.111 Lecture 35 35.1 Kinetics Topic: Catalysis Chapter 13 (Section 13.14-13.15) From Friday s material Le Chatelier's Principle - when a stress is applied to a system in equilibrium, the equilibrium tends
5.111 Lecture 35 35.1 Kinetics Topic: Catalysis Chapter 13 (Section 13.14-13.15) From Friday s material Le Chatelier's Principle - when a stress is applied to a system in equilibrium, the equilibrium tends
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer
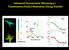 Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Advanced Fluorescence Microscopy I: Fluorescence (Foster) Resonance Energy Transfer 3.0 Pax 2.5 2.0 1200 800 400 GFP- Pax GFP-Pax + FATmCherry FAT FAT 1.5 Lifetime (ns) 0 1500 2000 2500 3000 Average lifetime
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Problem Set # 3
 20.320 Problem Set # 3 October 1 st, 2010 Due on October 8 th, 2010 at 11:59am. No extensions, no electronic submissions. General Instructions: 1. You are expected to state all your assumptions and provide
20.320 Problem Set # 3 October 1 st, 2010 Due on October 8 th, 2010 at 11:59am. No extensions, no electronic submissions. General Instructions: 1. You are expected to state all your assumptions and provide
Calorimetry: differential scanning calorimetry (DSC), isothermal titration calorimetry (ITC)
 Calorimetry: differential scanning calorimetry (DSC), isothermal titration calorimetry (ITC) Dr. Yin Li Department of Biophysics, Medical School University of Pecs Thermal Analysis IUPAC definition - a
Calorimetry: differential scanning calorimetry (DSC), isothermal titration calorimetry (ITC) Dr. Yin Li Department of Biophysics, Medical School University of Pecs Thermal Analysis IUPAC definition - a
Kinetic & Affinity Analysis
 Kinetic & Affinity Analysis An introduction What are kinetics and affinity? Kinetics How fast do things happen? Time-dependent Association how fast molecules bind Dissociation how fast complexes fall apart
Kinetic & Affinity Analysis An introduction What are kinetics and affinity? Kinetics How fast do things happen? Time-dependent Association how fast molecules bind Dissociation how fast complexes fall apart
Microcalorimetry for the Life Sciences
 Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Microcalorimetry for the Life Sciences Why Microcalorimetry? Microcalorimetry is universal detector Heat is generated or absorbed in every chemical process In-solution No molecular weight limitations Label-free
Presentation Microcalorimetry for Life Science Research
 Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
Presentation Microcalorimetry for Life Science Research MicroCalorimetry The Universal Detector Heat is either generated or absorbed in every chemical process Capable of thermal measurements over a wide
Fragment-Based Drug Discovery (FBDD) Using the dispr Technique on Pioneer Systems with OneStep and NeXtStep Injection Methodologies
 APPLICATION NOTE 21 Fragment-Based Drug Discovery (FBDD) Using the dispr Technique on Pioneer Systems with OneStep and NeXtStep Injection Methodologies Eric L. Reese, Ph.D, SensiQ Technologies, Aaron Martin
APPLICATION NOTE 21 Fragment-Based Drug Discovery (FBDD) Using the dispr Technique on Pioneer Systems with OneStep and NeXtStep Injection Methodologies Eric L. Reese, Ph.D, SensiQ Technologies, Aaron Martin
Richik N. Ghosh, Linnette Grove, and Oleg Lapets ASSAY and Drug Development Technologies 2004, 2:
 1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
1 3/1/2005 A Quantitative Cell-Based High-Content Screening Assay for the Epidermal Growth Factor Receptor-Specific Activation of Mitogen-Activated Protein Kinase Richik N. Ghosh, Linnette Grove, and Oleg
Problem Set 3 September 25, 2009
 September 25, 2009 General Instructions: 1. You are expected to state all your assumptions and provide step-by-step solutions to the numerical problems. Unless indicated otherwise, the computational problems
September 25, 2009 General Instructions: 1. You are expected to state all your assumptions and provide step-by-step solutions to the numerical problems. Unless indicated otherwise, the computational problems
Isothermal titration calorimetry (ITC)
 Isothermal titration calorimetry (ITC) Peter.gimeson@malvern.com Why microcalorimetry? Label-free Broad dynamic range Information rich Ease-of-use Direct measurement of heat change (ITC) Direct measurement
Isothermal titration calorimetry (ITC) Peter.gimeson@malvern.com Why microcalorimetry? Label-free Broad dynamic range Information rich Ease-of-use Direct measurement of heat change (ITC) Direct measurement
Irreversible Inhibition Kinetics
 1 Irreversible Inhibition Kinetics Automation and Simulation Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Automate the determination of biochemical parameters 2. PK/PD simulations with multiple injections Irreversible
1 Irreversible Inhibition Kinetics Automation and Simulation Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Automate the determination of biochemical parameters 2. PK/PD simulations with multiple injections Irreversible
Free Energy. because H is negative doesn't mean that G will be negative and just because S is positive doesn't mean that G will be negative.
 Biochemistry 462a Bioenergetics Reading - Lehninger Principles, Chapter 14, pp. 485-512 Practice problems - Chapter 14: 2-8, 10, 12, 13; Physical Chemistry extra problems, free energy problems Free Energy
Biochemistry 462a Bioenergetics Reading - Lehninger Principles, Chapter 14, pp. 485-512 Practice problems - Chapter 14: 2-8, 10, 12, 13; Physical Chemistry extra problems, free energy problems Free Energy
Lecture 27. Transition States and Enzyme Catalysis
 Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Lecture 27 Transition States and Enzyme Catalysis Reading for Today: Chapter 15 sections B and C Chapter 16 next two lectures 4/8/16 1 Pop Question 9 Binding data for your thesis protein (YTP), binding
Characterization of Reversible Kinase Inhibitors using Microfluidic Mobility-Shift Assays
 Application Note 211 Characterization of Reversible Kinase Inhibitors using Microfluidic Mobility-Shift Assays Introduction Current drug discovery efforts typically focus on developing small molecule inhibitors
Application Note 211 Characterization of Reversible Kinase Inhibitors using Microfluidic Mobility-Shift Assays Introduction Current drug discovery efforts typically focus on developing small molecule inhibitors
ENZYME KINETICS. Medical Biochemistry, Lecture 24
 ENZYME KINETICS Medical Biochemistry, Lecture 24 Lecture 24, Outline Michaelis-Menten kinetics Interpretations and uses of the Michaelis- Menten equation Enzyme inhibitors: types and kinetics Enzyme Kinetics
ENZYME KINETICS Medical Biochemistry, Lecture 24 Lecture 24, Outline Michaelis-Menten kinetics Interpretations and uses of the Michaelis- Menten equation Enzyme inhibitors: types and kinetics Enzyme Kinetics
Introduction to FBDD Fragment screening methods and library design
 Introduction to FBDD Fragment screening methods and library design Samantha Hughes, PhD Fragments 2013 RSC BMCS Workshop 3 rd March 2013 Copyright 2013 Galapagos NV Why fragment screening methods? Guess
Introduction to FBDD Fragment screening methods and library design Samantha Hughes, PhD Fragments 2013 RSC BMCS Workshop 3 rd March 2013 Copyright 2013 Galapagos NV Why fragment screening methods? Guess
DSC Characterization of the Structure/Function Relationship for Proteins
 DSC Characterization of the Structure/Function Relationship for Proteins Differential Scanning Calorimetry (DSC) DSC is recognized as Gold Std technique for measuring molecular thermal stability and structure
DSC Characterization of the Structure/Function Relationship for Proteins Differential Scanning Calorimetry (DSC) DSC is recognized as Gold Std technique for measuring molecular thermal stability and structure
BSc and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BSc and MSc Degree Examinations 2018-9 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes Marking
Examination Candidate Number: Desk Number: BSc and MSc Degree Examinations 2018-9 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes Marking
LanthaScreen Eu Kinase Binding Assay Validation Packet. Optimization of a LanthaScreen Eu Kinase Binding Assay for AURKB
 Page 1 of 18 LanthaScreen Eu Kinase Binding Assay for AURKB Overview This protocol describes how to perform a LanthaScreen Eu Kinase Binding Assay designed to detect and characterize kinase inhibitors.
Page 1 of 18 LanthaScreen Eu Kinase Binding Assay for AURKB Overview This protocol describes how to perform a LanthaScreen Eu Kinase Binding Assay designed to detect and characterize kinase inhibitors.
Small-Molecule Kinetics
 Application Note No. 1 / September 1, 2014 Small-Molecule Kinetics Creoptix WAVE Small-Molecule Kinetics: Binding of Sulfonamides to Carbonic Anhydrase II Summary Label-free interaction analysis of biomolecules
Application Note No. 1 / September 1, 2014 Small-Molecule Kinetics Creoptix WAVE Small-Molecule Kinetics: Binding of Sulfonamides to Carbonic Anhydrase II Summary Label-free interaction analysis of biomolecules
BA, BSc, and MSc Degree Examinations
 Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Examination Candidate Number: Desk Number: BA, BSc, and MSc Degree Examinations 2017-8 Department : BIOLOGY Title of Exam: Molecular Biology and Biochemistry Part I Time Allowed: 1 hour and 30 minutes
Small-Molecule Kinetics
 Application Note No. 1 / February 4, 2015 Small-Molecule Kinetics Creoptix WAVE Small-Molecule Kinetics: Binding of Sulfonamides to Carbonic Anhydrase II Summary Label-free interaction analysis of biomolecules
Application Note No. 1 / February 4, 2015 Small-Molecule Kinetics Creoptix WAVE Small-Molecule Kinetics: Binding of Sulfonamides to Carbonic Anhydrase II Summary Label-free interaction analysis of biomolecules
Microcalorimetric techniques
 Microcalorimetric techniques Isothermal titration calorimetry (ITC) Differential scanning calorimetry (DSC) Filip Šupljika Filip.Supljika@irb.hr Laboratory for the study of interactions of biomacromolecules
Microcalorimetric techniques Isothermal titration calorimetry (ITC) Differential scanning calorimetry (DSC) Filip Šupljika Filip.Supljika@irb.hr Laboratory for the study of interactions of biomacromolecules
Application Note. Authors. Introduction. Lauren E. Frick and William A. LaMarr Agilent Technologies, Inc. Wakefield, MA, USA
 Fragment-Based Drug Discovery: Comparing Labeled and Label-Free Screening of β-amyloid Secretase (BACE-1) Using Fluorescence Spectroscopy and Ultrafast SPE/MS/MS Application Note Authors Lauren E. Frick
Fragment-Based Drug Discovery: Comparing Labeled and Label-Free Screening of β-amyloid Secretase (BACE-1) Using Fluorescence Spectroscopy and Ultrafast SPE/MS/MS Application Note Authors Lauren E. Frick
Kinetic and Thermodynamic Analysis of Ligand Receptor Interactions: SPR Applications in Drug Development
 CHAPTER 5 Kinetic and Thermodynamic Analysis of Ligand Receptor Interactions: SPR Applications in Drug Development NICO J. DE MOL AND MARCEL J.E. FISCHER Department of Medicinal Chemistry and Chemical
CHAPTER 5 Kinetic and Thermodynamic Analysis of Ligand Receptor Interactions: SPR Applications in Drug Development NICO J. DE MOL AND MARCEL J.E. FISCHER Department of Medicinal Chemistry and Chemical
Principles of Drug Design
 Advanced Medicinal Chemistry II Principles of Drug Design Tentative Course Outline Instructors: Longqin Hu and John Kerrigan Direct questions and enquiries to the Course Coordinator: Longqin Hu I. Introduction
Advanced Medicinal Chemistry II Principles of Drug Design Tentative Course Outline Instructors: Longqin Hu and John Kerrigan Direct questions and enquiries to the Course Coordinator: Longqin Hu I. Introduction
Biological Thermodynamics
 Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
Biological Thermodynamics Classical thermodynamics is the only physical theory of universal content concerning which I am convinced that, within the framework of applicability of its basic contents, will
NMR Solutions for drug discovery
 NMR Solutions for drug discovery Dr. Matteo Pennestri London, UK Bruker Users Meeting Innovation with Integrity The principle of Fragment Based Screening from efficient fragments to Drug candidates Fragment
NMR Solutions for drug discovery Dr. Matteo Pennestri London, UK Bruker Users Meeting Innovation with Integrity The principle of Fragment Based Screening from efficient fragments to Drug candidates Fragment
Monolith NT.Automated Product Information
 Monolith NT.Automated Product Information Monolith Instruments for MicroScale Thermophoresis www.nanotemper-technologies.com Monolith NT.Automated Product Information 2 www.nanotemper-technologies.com
Monolith NT.Automated Product Information Monolith Instruments for MicroScale Thermophoresis www.nanotemper-technologies.com Monolith NT.Automated Product Information 2 www.nanotemper-technologies.com
S2004 Methods for characterization of biomolecular interactions - classical versus modern
 S2004 Methods for characterization of biomolecular interactions - classical versus modern Isothermal Titration Calorimetry (ITC) Eva Dubská email: eva.dubska@ceitec.cz Outline Calorimetry - history + a
S2004 Methods for characterization of biomolecular interactions - classical versus modern Isothermal Titration Calorimetry (ITC) Eva Dubská email: eva.dubska@ceitec.cz Outline Calorimetry - history + a
FRAGMENT SCREENING IN LEAD DISCOVERY BY WEAK AFFINITY CHROMATOGRAPHY (WAC )
 FRAGMENT SCREENING IN LEAD DISCOVERY BY WEAK AFFINITY CHROMATOGRAPHY (WAC ) SARomics Biostructures AB & Red Glead Discovery AB Medicon Village, Lund, Sweden Fragment-based lead discovery The basic idea:
FRAGMENT SCREENING IN LEAD DISCOVERY BY WEAK AFFINITY CHROMATOGRAPHY (WAC ) SARomics Biostructures AB & Red Glead Discovery AB Medicon Village, Lund, Sweden Fragment-based lead discovery The basic idea:
In silico pharmacology for drug discovery
 In silico pharmacology for drug discovery In silico drug design In silico methods can contribute to drug targets identification through application of bionformatics tools. Currently, the application of
In silico pharmacology for drug discovery In silico drug design In silico methods can contribute to drug targets identification through application of bionformatics tools. Currently, the application of
protein interaction analysis bulletin 6300
 protein interaction analysis bulletin 6300 Guide to SPR Data Analysis on the ProteOn XPR36 System Ruben Luo, Bio-Rad Laboratories, Inc., 2000 Alfred Nobel Drive, Hercules, CA 94547 Kinetic Analysis To
protein interaction analysis bulletin 6300 Guide to SPR Data Analysis on the ProteOn XPR36 System Ruben Luo, Bio-Rad Laboratories, Inc., 2000 Alfred Nobel Drive, Hercules, CA 94547 Kinetic Analysis To
Isothermal titration calorimetry to determine association constants for high-affinity ligands
 Isothermal titration calorimetry to determine association constants for high-affinity ligands Adrian Velazquez-Campoy 1 & Ernesto Freire 2 1 Institute of Biocomputation and Complex Systems Physics, Corona
Isothermal titration calorimetry to determine association constants for high-affinity ligands Adrian Velazquez-Campoy 1 & Ernesto Freire 2 1 Institute of Biocomputation and Complex Systems Physics, Corona
Binding Theory Equations for Affinity and Kinetics Analysis
 Technology Note #101 Binding Theory Equations for Affinity and Kinetics Analysis This technology note summarizes important equations underlying the theory of binding of solute analytes to surface-tethered
Technology Note #101 Binding Theory Equations for Affinity and Kinetics Analysis This technology note summarizes important equations underlying the theory of binding of solute analytes to surface-tethered
LanthaScreen Eu Kinase Binding Assay Validation Packet. Optimization of a LanthaScreen Eu Kinase Binding Assay for RPS6KA1
 Page 1 of 18 Optimization of a Assay for RPS6KA1 Assay for RPS6KA1 Overview This protocol describes how to perform a Assay designed to detect and characterize kinase inhibitors. Procedure 1 describes an
Page 1 of 18 Optimization of a Assay for RPS6KA1 Assay for RPS6KA1 Overview This protocol describes how to perform a Assay designed to detect and characterize kinase inhibitors. Procedure 1 describes an
It is generally believed that the catalytic reactions occur in at least two steps.
 Lecture 16 MECHANISM OF ENZYME ACTION A chemical reaction such as A ----> P takes place because a certain fraction of the substrate possesses enough energy to attain an activated condition called the transition
Lecture 16 MECHANISM OF ENZYME ACTION A chemical reaction such as A ----> P takes place because a certain fraction of the substrate possesses enough energy to attain an activated condition called the transition
Binding Studies on Trafficking Proteins Using Microcalorimetry McMahon lab
 Binding Studies on Trafficking Proteins Using Microcalorimetry McMahon lab Neurobiology Division Laboratory of Molecular Biology Cambridge Clathrin Mediated Endocytosis Binding Recruitment Coating Budding
Binding Studies on Trafficking Proteins Using Microcalorimetry McMahon lab Neurobiology Division Laboratory of Molecular Biology Cambridge Clathrin Mediated Endocytosis Binding Recruitment Coating Budding
2054, Chap. 8, page 1
 2054, Chap. 8, page 1 I. Metabolism: Energetics, Enzymes, and Regulation (Chapter 8) A. Energetics and work 1. overview a. energy = ability to do work (1) chemical, transport, mechanical (2) ultimate source
2054, Chap. 8, page 1 I. Metabolism: Energetics, Enzymes, and Regulation (Chapter 8) A. Energetics and work 1. overview a. energy = ability to do work (1) chemical, transport, mechanical (2) ultimate source
7.012 Problem Set 1. i) What are two main differences between prokaryotic cells and eukaryotic cells?
 ame 7.01 Problem Set 1 Section Question 1 a) What are the four major types of biological molecules discussed in lecture? Give one important function of each type of biological molecule in the cell? b)
ame 7.01 Problem Set 1 Section Question 1 a) What are the four major types of biological molecules discussed in lecture? Give one important function of each type of biological molecule in the cell? b)
Receptor Based Drug Design (1)
 Induced Fit Model For more than 100 years, the behaviour of enzymes had been explained by the "lock-and-key" mechanism developed by pioneering German chemist Emil Fischer. Fischer thought that the chemicals
Induced Fit Model For more than 100 years, the behaviour of enzymes had been explained by the "lock-and-key" mechanism developed by pioneering German chemist Emil Fischer. Fischer thought that the chemicals
MSc Drug Design. Module Structure: (15 credits each) Lectures and Tutorials Assessment: 50% coursework, 50% unseen examination.
 Module Structure: (15 credits each) Lectures and Assessment: 50% coursework, 50% unseen examination. Module Title Module 1: Bioinformatics and structural biology as applied to drug design MEDC0075 In the
Module Structure: (15 credits each) Lectures and Assessment: 50% coursework, 50% unseen examination. Module Title Module 1: Bioinformatics and structural biology as applied to drug design MEDC0075 In the
Concept review: Binding equilibria
 Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Concept review: Binding equilibria 1 Binding equilibria and association/dissociation constants 2 The binding of a protein to a ligand at equilibrium can be written as: P + L PL And so the equilibrium constant
Discussion Exercise 5: Analyzing Graphical Data
 Discussion Exercise 5: Analyzing Graphical Data Skill 1: Use axis labels to describe a phenomenon as a function of a variable Some value y may be described as a function of some variable x and used to
Discussion Exercise 5: Analyzing Graphical Data Skill 1: Use axis labels to describe a phenomenon as a function of a variable Some value y may be described as a function of some variable x and used to
Table 1. Kinetic data obtained from SPR analysis of domain 11 mutants interacting with IGF-II. Kinetic parameters K D 1.
 Kinetics and Thermodynamics of the Insulin-like Growth Factor II (IGF-II) Interaction with IGF-II/Mannose 6-phosphate Receptor and the function of CD and AB Loop Solvent-exposed Residues. Research Team:
Kinetics and Thermodynamics of the Insulin-like Growth Factor II (IGF-II) Interaction with IGF-II/Mannose 6-phosphate Receptor and the function of CD and AB Loop Solvent-exposed Residues. Research Team:
Chem/Biochem 471 Exam 2 11/14/07 Page 1 of 7 Name:
 Page 1 of 7 Please leave the exam pages stapled together. The formulas are on a separate sheet. This exam has 5 questions. You must answer at least 4 of the questions. You may answer all 5 questions if
Page 1 of 7 Please leave the exam pages stapled together. The formulas are on a separate sheet. This exam has 5 questions. You must answer at least 4 of the questions. You may answer all 5 questions if
An Introduction to Metabolism
 An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
Characterizing Binding Interactions by ITC
 Characterizing Binding Interactions by ITC Christin T. Choma TA Instruments, 19 Lukens Drive, New Castle, DE 1972, USA All biochemical reactions involve recognition, binding and the formation of noncovalent
Characterizing Binding Interactions by ITC Christin T. Choma TA Instruments, 19 Lukens Drive, New Castle, DE 1972, USA All biochemical reactions involve recognition, binding and the formation of noncovalent
CHAPTER 8 Analysis of FP Binding Data
 CHAPTER 8 Analysis of FP Binding Data Determination of Binding Constants............................................................8-2 Definitions.........................................................................8-2
CHAPTER 8 Analysis of FP Binding Data Determination of Binding Constants............................................................8-2 Definitions.........................................................................8-2
Chapter 6. The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR
 The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
The interaction of Src SH2 with the focal adhesion kinase catalytic domain studied by NMR 103 Abstract The interaction of the Src SH2 domain with the catalytic domain of FAK, including the Y397 SH2 domain
Analysis of Macromolecular Interactions by Isothermal Titration Calorimetry
 Analysis of Macromolecular Interactions by Isothermal Titration Calorimetry A primer on the experimental thermodynamic charaterisation of binding of biomolecules Gonzalo Obal Protein Biophysics Unit Structural
Analysis of Macromolecular Interactions by Isothermal Titration Calorimetry A primer on the experimental thermodynamic charaterisation of binding of biomolecules Gonzalo Obal Protein Biophysics Unit Structural
ITC Expert User s Manual
 ITC Expert User s Manual 1 Section 1: ITC Expert Background... 3 Minimal Heats and Injections... 3 C Parameter... 3 C Limitations... 4 High C... 4 Low C... 6 Concentrations Ratio... 6 Section 2: ITC Expert
ITC Expert User s Manual 1 Section 1: ITC Expert Background... 3 Minimal Heats and Injections... 3 C Parameter... 3 C Limitations... 4 High C... 4 Low C... 6 Concentrations Ratio... 6 Section 2: ITC Expert
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Chapter 8: An Introduction to Metabolism
 Chapter 8: An Introduction to Metabolism Key Concepts 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 8.2 The free-energy change of a reaction tells us
Chapter 8: An Introduction to Metabolism Key Concepts 8.1 An organism s metabolism transforms matter and energy, subject to the laws of thermodynamics 8.2 The free-energy change of a reaction tells us
Transient kinetic methods. Biophysics seminars Kinga Futó
 Transient kinetic methods Biophysics seminars Kinga Futó Fast kinetics Reaction kinetics: temporal description of reactions reactionmechanisms reaction dynamics (processes on molecular level) rate of reaction
Transient kinetic methods Biophysics seminars Kinga Futó Fast kinetics Reaction kinetics: temporal description of reactions reactionmechanisms reaction dynamics (processes on molecular level) rate of reaction
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/3/4/e1600663/dc1 Supplementary Materials for A dynamic hydrophobic core orchestrates allostery in protein kinases Jonggul Kim, Lalima G. Ahuja, Fa-An Chao, Youlin
advances.sciencemag.org/cgi/content/full/3/4/e1600663/dc1 Supplementary Materials for A dynamic hydrophobic core orchestrates allostery in protein kinases Jonggul Kim, Lalima G. Ahuja, Fa-An Chao, Youlin
Computational Biology 1
 Computational Biology 1 Protein Function & nzyme inetics Guna Rajagopal, Bioinformatics Institute, guna@bii.a-star.edu.sg References : Molecular Biology of the Cell, 4 th d. Alberts et. al. Pg. 129 190
Computational Biology 1 Protein Function & nzyme inetics Guna Rajagopal, Bioinformatics Institute, guna@bii.a-star.edu.sg References : Molecular Biology of the Cell, 4 th d. Alberts et. al. Pg. 129 190
BCH 3023 Fall 2008 Exam 2, Form C Name: ANSWER KEY
 Name: ANSWER KEY In class, we discussed one method to linearize the Michaelis-Menton equation. There are other methods to do this, one being an Eadie-Hofstee plot. Given the Eadie-Hofstee plot below, answer
Name: ANSWER KEY In class, we discussed one method to linearize the Michaelis-Menton equation. There are other methods to do this, one being an Eadie-Hofstee plot. Given the Eadie-Hofstee plot below, answer
Supplementary Figures
 1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
1 Supplementary Figures Supplementary Figure 1 Type I FGFR1 inhibitors (a) Chemical structures of a pyrazolylaminopyrimidine inhibitor (henceforth referred to as PAPI; PDB-code of the FGFR1-PAPI complex:
Thermodynamic measurements
 Thermoynamic measurements When stuying protein structure an function, it is essential to measure various physical quantities, incluing: thermal an chemical stability bining affinity to a ligan solubility
Thermoynamic measurements When stuying protein structure an function, it is essential to measure various physical quantities, incluing: thermal an chemical stability bining affinity to a ligan solubility
Biophysics Service at the MPIB Biochemistry Core Facility Stephan Uebel, Biochemistry Core Facility
 Biophysics Service at the MPIB Biochemistry Core Facility 30.11.2015 Stephan Uebel, Biochemistry Core Facility uebel@biochem.mpg.de Overview Peptide Chemistry - Peptide synthesis -Amino acid analysis -
Biophysics Service at the MPIB Biochemistry Core Facility 30.11.2015 Stephan Uebel, Biochemistry Core Facility uebel@biochem.mpg.de Overview Peptide Chemistry - Peptide synthesis -Amino acid analysis -
COMBINATORIAL CHEMISTRY IN A HISTORICAL PERSPECTIVE
 NUE FEATURE T R A N S F O R M I N G C H A L L E N G E S I N T O M E D I C I N E Nuevolution Feature no. 1 October 2015 Technical Information COMBINATORIAL CHEMISTRY IN A HISTORICAL PERSPECTIVE A PROMISING
NUE FEATURE T R A N S F O R M I N G C H A L L E N G E S I N T O M E D I C I N E Nuevolution Feature no. 1 October 2015 Technical Information COMBINATORIAL CHEMISTRY IN A HISTORICAL PERSPECTIVE A PROMISING
EMPIRICAL VS. RATIONAL METHODS OF DISCOVERING NEW DRUGS
 EMPIRICAL VS. RATIONAL METHODS OF DISCOVERING NEW DRUGS PETER GUND Pharmacopeia Inc., CN 5350 Princeton, NJ 08543, USA pgund@pharmacop.com Empirical and theoretical approaches to drug discovery have often
EMPIRICAL VS. RATIONAL METHODS OF DISCOVERING NEW DRUGS PETER GUND Pharmacopeia Inc., CN 5350 Princeton, NJ 08543, USA pgund@pharmacop.com Empirical and theoretical approaches to drug discovery have often
BI/CH421 Biochemistry I Exam 1 09/29/2014
 Part I: Multiple choice for each question, circle the choice that best answers the question. Do not write the letter in the margin to indicate your answer, circle it. 3 points each. 1. For a reaction with
Part I: Multiple choice for each question, circle the choice that best answers the question. Do not write the letter in the margin to indicate your answer, circle it. 3 points each. 1. For a reaction with
13 TH NEW ENZYMOLOGY KINETICS WORKSHOP
 13 TH NEW ENZYMOLOGY KINETICS WORKSHOP OFFERED BY KENNETH A. JOHNSON, PH.D. 26 May 30 May 2019 Hotel International BRNO, Czech Republic This figure shows the simultaneous fitting six experiments to define
13 TH NEW ENZYMOLOGY KINETICS WORKSHOP OFFERED BY KENNETH A. JOHNSON, PH.D. 26 May 30 May 2019 Hotel International BRNO, Czech Republic This figure shows the simultaneous fitting six experiments to define
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
How antibody surface coverage on nanoparticles determines the. activity and kinetics of antigen capturing for biosensing
 How antibody surface coverage on nanoparticles determines the activity and kinetics of antigen capturing for biosensing Bedabrata Saha, Toon H. Evers, and Menno W. J. Prins Philips Research, High Tech
How antibody surface coverage on nanoparticles determines the activity and kinetics of antigen capturing for biosensing Bedabrata Saha, Toon H. Evers, and Menno W. J. Prins Philips Research, High Tech
Page 1 of 16. Final Exam 5.111
 Final Exam 5.111 Page 1 of 16 Write your name and your TA's name below. Do not open the exam until the start of the exam is announced. 1) Read each part of a problem thoroughly. Many of a problem s latter
Final Exam 5.111 Page 1 of 16 Write your name and your TA's name below. Do not open the exam until the start of the exam is announced. 1) Read each part of a problem thoroughly. Many of a problem s latter
The Institute of Cancer Research PHD STUDENTSHIP PROJECT PROPOSAL
 The Institute of Cancer Research PHD STUDENTSHIP PROJECT PROPOSAL PROJECT DETAILS Project Title: Design and synthesis of libraries of bifunctional degraders for the discovery of new cancer targets SUPERVISORY
The Institute of Cancer Research PHD STUDENTSHIP PROJECT PROPOSAL PROJECT DETAILS Project Title: Design and synthesis of libraries of bifunctional degraders for the discovery of new cancer targets SUPERVISORY
Life Sciences 1a Lecture Slides Set 10 Fall Prof. David R. Liu. Lecture Readings. Required: Lecture Notes McMurray p , O NH
 Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Life ciences 1a Lecture lides et 10 Fall 2006-2007 Prof. David R. Liu Lectures 17-18: The molecular basis of drug-protein binding: IV protease inhibitors 1. Drug development and its impact on IV-infected
Supplementary information for cloud computing approaches for prediction of ligand binding poses and pathways
 Supplementary information for cloud computing approaches for prediction of ligand binding poses and pathways Morgan Lawrenz 1, Diwakar Shukla 1,2 & Vijay S. Pande 1,2 1 Department of Chemistry, Stanford
Supplementary information for cloud computing approaches for prediction of ligand binding poses and pathways Morgan Lawrenz 1, Diwakar Shukla 1,2 & Vijay S. Pande 1,2 1 Department of Chemistry, Stanford
LanthaScreen Eu Kinase Binding Assay Validation Packet
 Page of 7 LanthaScreen Eu kinase binding assay for RF Overview This protocol describes how to perform a LanthaScreen Eu Kinase indng ssay designed to detect and characterize kinase inhibitors. Procedure
Page of 7 LanthaScreen Eu kinase binding assay for RF Overview This protocol describes how to perform a LanthaScreen Eu Kinase indng ssay designed to detect and characterize kinase inhibitors. Procedure
Energy Changes, Reaction Rates and Equilibrium. Thermodynamics: study of energy, work and heat. Kinetic energy: energy of motion
 Energy Changes, Reaction Rates and Equilibrium Thermodynamics: study of energy, work and heat Kinetic energy: energy of motion Potential energy: energy of position, stored energy Chemical reactions involve
Energy Changes, Reaction Rates and Equilibrium Thermodynamics: study of energy, work and heat Kinetic energy: energy of motion Potential energy: energy of position, stored energy Chemical reactions involve
Structural Analysis of an Unknown Compound and Determination of its pk a by NMR Spectroscopy
 Structural Analysis of an Unknown Compound and Determination of its pk a by NMR Spectroscopy Yoshitaka Ishii, Dan McElheny, and Isamu Matsuda, August 31, 2013 (revised January 14, 2014) 1. Theoretical
Structural Analysis of an Unknown Compound and Determination of its pk a by NMR Spectroscopy Yoshitaka Ishii, Dan McElheny, and Isamu Matsuda, August 31, 2013 (revised January 14, 2014) 1. Theoretical
Problem Set No. 4 Due: Monday, 11/18/10 at the start of class
 Department of Chemical Engineering ChE 170 University of California, Santa Barbara Fall 2010 Problem Set No. 4 Due: Monday, 11/18/10 at the start of class Objective: To understand the thermodynamic and
Department of Chemical Engineering ChE 170 University of California, Santa Barbara Fall 2010 Problem Set No. 4 Due: Monday, 11/18/10 at the start of class Objective: To understand the thermodynamic and
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing
 Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Supplementary Figure 1. SDS-PAGE analysis of GFP oligomer variants with different linkers. Oligomer mixtures were applied to a PAGE gel containing 0.1% SDS without boiling. The gel was analyzed by a fluorescent
Protein assay. Absorbance Fluorescence Emission Colorimetric detection BIO/MDT 325. Absorbance
 Protein assay Absorbance Fluorescence Emission Colorimetric detection BIO/MDT 325 Absorbance Using A280 to Determine Protein Concentration Determination of protein concentration by measuring absorbance
Protein assay Absorbance Fluorescence Emission Colorimetric detection BIO/MDT 325 Absorbance Using A280 to Determine Protein Concentration Determination of protein concentration by measuring absorbance
Objectives INTRODUCTION TO METABOLISM. Metabolism. Catabolic Pathways. Anabolic Pathways 3/6/2011. How to Read a Chemical Equation
 Objectives INTRODUCTION TO METABOLISM. Chapter 8 Metabolism, Energy, and Life Explain the role of catabolic and anabolic pathways in cell metabolism Distinguish between kinetic and potential energy Distinguish
Objectives INTRODUCTION TO METABOLISM. Chapter 8 Metabolism, Energy, and Life Explain the role of catabolic and anabolic pathways in cell metabolism Distinguish between kinetic and potential energy Distinguish
Material Relationships - distributor details
 Malvern Instruments Limited Grovewood Road, Malvern, Worcestershire, UK, WR14 1XZ Material Relationships - distributor details Tel +44 1684 892456 Fax +44 1684 892789 www.malvern.com Malvern Instruments
Malvern Instruments Limited Grovewood Road, Malvern, Worcestershire, UK, WR14 1XZ Material Relationships - distributor details Tel +44 1684 892456 Fax +44 1684 892789 www.malvern.com Malvern Instruments
Chem 204. Mid-Term Exam I. July 21, There are 3 sections to this exam: Answer ALL questions
 Chem 204 Mid-Term Exam I July 21, 2009 Name: Answer Key Student ID: There are 3 sections to this exam: Answer ALL questions Section I: Multiple-Choice 20 questions, 2 pts each Section II: Fill-in-the-Blank
Chem 204 Mid-Term Exam I July 21, 2009 Name: Answer Key Student ID: There are 3 sections to this exam: Answer ALL questions Section I: Multiple-Choice 20 questions, 2 pts each Section II: Fill-in-the-Blank
MICROCAL ITC SYSTEMS UNDERSTANDING BIOMOLECULAR INTERACTIONS LABEL-FREE BINDING ANALYSIS MICROCALORIMETRY
 LABEL-FREE BINDING ANALYSIS MICROCALORIMETRY MICROCAL ITC SYSTEMS UNDERSTANDING BIOMOLECULAR INTERACTIONS MEASURE MULTIPLE BINDING PARAMETERS IN A SINGLE EXPERIMENT Isothermal titration microcalorimetry
LABEL-FREE BINDING ANALYSIS MICROCALORIMETRY MICROCAL ITC SYSTEMS UNDERSTANDING BIOMOLECULAR INTERACTIONS MEASURE MULTIPLE BINDING PARAMETERS IN A SINGLE EXPERIMENT Isothermal titration microcalorimetry
CALORIMETRIA. strumenti ed applicazioni
 09/11/2010 CALORIMETRIA strumenti ed applicazioni Università di Parma, 9 novembre 2010 R. Pepi TA Instruments Thermal Analysis Thermal analysis refers to a group of techniques in which a physical property
09/11/2010 CALORIMETRIA strumenti ed applicazioni Università di Parma, 9 novembre 2010 R. Pepi TA Instruments Thermal Analysis Thermal analysis refers to a group of techniques in which a physical property
Biochemical / Biophysical Kinetics Made Easy
 Biochemical / Biophysical Kinetics Made Easy Software DYNAFIT in drug discovery research Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Theory: differential equation models - DYNAFIT software 2. Example: lanthascreen
Biochemical / Biophysical Kinetics Made Easy Software DYNAFIT in drug discovery research Petr Kuzmič, Ph.D. BioKin, Ltd. 1. Theory: differential equation models - DYNAFIT software 2. Example: lanthascreen
Energy Transformation and Metabolism (Outline)
 Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Energy Transformation and Metabolism (Outline) - Definitions & Laws of Thermodynamics - Overview of energy flow ecosystem - Biochemical processes: Anabolic/endergonic & Catabolic/exergonic - Chemical reactions
Protein-Ligand Interactions Are Responsible for Signal Transduction
 Proteinigand Interactions Are Responsible for Signal ransduction ypes of Interactions: 1. ProteinProtein 2. ProteinDNA (RNA) 3. Proteinsmall molecule Dynamic Proteinigand Interaction Dynamic interaction
Proteinigand Interactions Are Responsible for Signal ransduction ypes of Interactions: 1. ProteinProtein 2. ProteinDNA (RNA) 3. Proteinsmall molecule Dynamic Proteinigand Interaction Dynamic interaction
Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase
 SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
SUPPLEMENTARY INFORMATION Analysis of nucleotide binding to p97 reveals the properties of a tandem AAA hexameric ATPase Louise C Briggs, Geoff S Baldwin, Non Miyata, Hisao Kondo, Xiaodong Zhang, Paul S
Lecture 7. Surface Reaction Kinetics on a
 Lecture 7 Data processing in SPR Surface Reaction Kinetics on a Biochip Surface plasmon sensor Principle of affinity SP biosensor Data processing Processing steps: zero response to the base line before
Lecture 7 Data processing in SPR Surface Reaction Kinetics on a Biochip Surface plasmon sensor Principle of affinity SP biosensor Data processing Processing steps: zero response to the base line before
Unlocking the potential of your drug discovery programme
 Unlocking the potential of your drug discovery programme Innovative screening The leading fragment screening platform with MicroScale Thermophoresis at its core Domainex expertise High quality results
Unlocking the potential of your drug discovery programme Innovative screening The leading fragment screening platform with MicroScale Thermophoresis at its core Domainex expertise High quality results
CHEM 4170 Problem Set #1
 CHEM 4170 Problem Set #1 0. Work problems 1-7 at the end of Chapter ne and problems 1, 3, 4, 5, 8, 10, 12, 17, 18, 19, 22, 24, and 25 at the end of Chapter Two and problem 1 at the end of Chapter Three
CHEM 4170 Problem Set #1 0. Work problems 1-7 at the end of Chapter ne and problems 1, 3, 4, 5, 8, 10, 12, 17, 18, 19, 22, 24, and 25 at the end of Chapter Two and problem 1 at the end of Chapter Three
Table 1. Data for Heat Capacity Trial 1 Trial 2
 Thermochemistry: Measuring Enthalpy Change in Chemical Reactions Experiment created by the UMaine InterChemNet Team. Adapted with permission. Print this form and bring it with you to lab. You will complete
Thermochemistry: Measuring Enthalpy Change in Chemical Reactions Experiment created by the UMaine InterChemNet Team. Adapted with permission. Print this form and bring it with you to lab. You will complete
Identifying Interaction Hot Spots with SuperStar
 Identifying Interaction Hot Spots with SuperStar Version 1.0 November 2017 Table of Contents Identifying Interaction Hot Spots with SuperStar... 2 Case Study... 3 Introduction... 3 Generate SuperStar Maps
Identifying Interaction Hot Spots with SuperStar Version 1.0 November 2017 Table of Contents Identifying Interaction Hot Spots with SuperStar... 2 Case Study... 3 Introduction... 3 Generate SuperStar Maps
