Simulating Gene Expression Patterns of Neurogenesis, Somitogenesis, and Morphogenesis During Zebrafish Embryo Development
|
|
|
- Rolf Nelson
- 5 years ago
- Views:
Transcription
1 Simulating Gene Expression Patterns of Neurogenesis, Somitogenesis, and Morphogenesis During Zebrafish Embryo Development Ei-Ei Gaw Computational Biosciences Program Arizona State University Tempe, Arizona Ajay B Chitnis Internship Supervisor Laboratory of Molecular Genetics NICHD, NIH Bethesda, Maryland April 2004 Internship: June August 2003
2 Simulating gene expression patterns of neurogenesis, somitogenesis, and morphogenesis during zebrafish embryo development Ei-Ei Gaw and Ajay B Chitnis Abstract During embryo development, it is essential that relatively homogenous groups of cells undergo differentiation to form spatially different patterns and eventually take on many different functions Intercellular communication and morphogen gradients are two aspects that have been shown to play roles in determining cell fate To understand better how these activities result in pattern formation, we utilized the NetLogo programming environment to simulate these processes Using this program, we were able to observe visually the possible pattern formation of gene expression from activation and inhibition of genes, intercellular interactions, and the exposure to morphogen gradients The three developmental phenomena of zebrafish embryos we studied were neurogenesis, somitogenesis, and morphogenesis during anterior/posterior patterning The mo for neurogenesis examines how autocatalysis and lateral inhibition (notch signaling) are required to form stable patterns In addition, we investigated to what extent the geometry of the domains and the initial noise in the her gene expression help determine a cell s fate To study somitogenesis, we explored how transcription and translation ays coupled with notch pathway and independent moving wave-front activity of fgf may contribute to the oscillation and synchronization of gene expression during formation of somites Lastly, we used our mos to examine how time and concentration of a morphogen gradient may signal gene activation and eventually form patterns of cells with stable and differing gene expression Although, there is much room to study in more detail the systems of equations and the numerical analysis we used, overall this method is a good addition to the traditional methods of studying how gene-expression patterns may develop 1 Gaw, 2004
3 Introduction Zebrafish as a mo organism has become popular for studying developmental biology for many reasons: zebrafish are vertebrates, they are inexpensive and easy to maintain and breed, they are experimentally accessible, and are transparent during early development The ease with which the embryo s DNA can be manipulated and the short period of embryogenesis make them an ideal system for studying different phenomena during development Furthermore, there is a large community of resources for zebrafish information ( In the process of going from a uniform mass of cells to cells with differing gene expressions and function, communication between cells and exposure to different cellular environments are vital in determination of a cell s fate We studied the role of these actions in neurogenesis, somitogenesis, and morphogenesis for anterior/posterior patterning during zebrafish embryo development Neurogenesis In early zebrafish development, the neural plate forms domains containing cells that eventually become neurons It has been shown that neurogenin1 (ngn1) is a proneural gene that is expressed in these domains [1-9] However, the number of cells within these proneural domains that become neurons is only a fraction of the total number of cells Lateral inhibition coupled with autocatalysis is believed to play a key role in the cell s fate within the clusters of cells, see Fig 1 Autocatalysis alone is not sufficient to differentiate cells within a neighborhood; if this was the case, all cells that express ngn1, no matter how small initially, would eventually become neurons Lateral inhibition, a competitive process where one cell s autocatalysis process inhibits the same process in the neighboring cells, is necessary to prevent the over production of neurons The Delta/Notch signaling provides the lateral inhibition that prevents neurogenesis in the adjacent cells by promoting hairy (her4) gene, which in turn inhibits the production of ngn1 Somitogenesis In many vertebrates, somites are formed synchronously on both sides of the body axis from a group of cells that buds off sequentially at the anterior end of the unsegmented presomitic mesoderm (PSM) [10-13] Somites give rise to skeletal muscles, vertebrae, and some dermis An essential part of this periodic process is the segmentation clock, the oscillator that produces the cyclic expression of specific genes There have been many theoretical mos developed to explain the mechanisms of this oscillator: cell cycle mo [14], clock and induction mo [15], clock and trail mo [16], and the oscillator and traveling waves [16-24, 29] Of all the references cited, only the cell cycle mo [14] and Lewis s oscillator mo [29] provided mathematical analysis of their theoretical mo There have been findings that support that the notch signaling plays a critical role [20], see Fig 2, and that fgf is important in determining somite boundary position [23] Furthermore, the target genes,, and tac expressions were found to oscillate during somitogenesis However, tac alone cannot oscillate It requires the oscillation of and, thus and are excellent candidates for the central mechanism of the oscillator [24] Anterior/Posterior Morphogenesis 2 Gaw, 2004
4 The morphogen Wnt, acting through -catenin and its antagonists secreted by the vertebrate organizer, regulate the compartmentalization of gene expression along the rostral-caudal axis during the development of the vertebrate nervous system Wnt signaling, regulated by -catenin, is responsible for posteriorizing activities that facilitate the expression of caudalizing genes The ectopic expression of wnt has led to a loss of rostral neural domains and expansion of caudal domains, while ectopic suppression of wnt signaling has resulted in the opposite The study of zebrafish headless (hdl) mutants reveals that the loss of head phenotype is due to mutation in the tcf3 gene, a member of the Tcf/Lef family In hdl mutants, the hindbrain and spinal cord are relatively unaffected, but there is an increase in the size of the midbrain-hindbrain boundary (MHB) and a loss of forebrain This is consistent with tcf3 as the repressor of caudal target genes Furthermore, tcf3b, a homolog of tcf3, has been shown to limit caudalization activity of hdl mutants [25-26] Methods NetLogo NetLogo is the next generation in multi-agent moing languages that started with StarLogo Daniel Bobrow and Wallace Feurzeig at Bolt, Beranek and Newman, Inc, and Seymour Papert, at MIT in the 1960 s, originally developed StarLogo In 2002, a team at Northwestern University led by Uri Wilensky and Seth Tisue developed NetLogo As with StarLogo, it has a programmable environment ideal for studying the emergence properties of complex systems developing over time This environment utilizes Logo language, which supports agents and parallelism Three different types of agents make up the NetLogo world: turtles, patches, and observers The world is divided into grids called patches, the turtles are agents that can move about the patches, and the observer is looking at the world All three agents may have unlimited variables and can carry out activities simultaneously As one can see, this environment is ideal to study the world of zebrafish embryogenesis Information and programs may be downloaded from Mos Neurogenesis Our lateral inhibition mo is based on Meinhardt s mo for self-enhancement (autocatalysis) and long-range inhibition [27-28], the nonlinear system of differential equations A ngn t A t A t her P P P ngn her her A 1+ S C ngn 2 ( A + 1)( 1+ S A ) notch A ngn A A tot _ 2 R R ngn her A ngn A her R f 3 2 ngn A f, and ngn f, 1 (1) (2) (3) The A variables are the amount of expression The parameters P and R are the rates of production and the rates of degradation, respectively The subscripts ngn,, and her represent genes ngn, ta, and her The parameter S ngn is the rate of saturation of ngn, and S is the rate of self-inhibition of ta The variable A _ is the total ta in the neighboring cells The tot 3 Gaw, 2004
5 constant, C notch, is the relative importance of A tot _ and represents the effect of notch signaling We made the assumption that diffusion does not play a role since only the immediately neighboring cells ta promotes the production of her through notch signaling This mo simulates the expression of ngn, ta, and her through the notch-signaling pathway The expression of ngn is driven by non-linear autocatalysis and is inhibited by self-saturation and her The total amount of ngn is equal to the production minus the removal of ngn which is proportional to the number of ngn present Since the term for basal production of ngn is removed, the initial amount of ngn must be greater than zero The expression of ta is promoted by ngn but is regulated by self-inhibition The total amount of ta is calculated by subtracting the amount of ta removed, R, from the ta produced The total her A product is linearly proportional to the total ta in the neighboring cells, A tot _, multiplied by the notch factor, C notch The following is the algorithm used to simulate the mo in NetLogo Euler s method of integration is implemented with step size of one, t 1 Neurogenesis NetLogo Algorithm: 1 Set up the proneuro domains: motorneurons, interneurons, and sensoryneurons See Fig 4a 2 Set production (P), removal (R), and saturation and self-inhibition (S) rates Set notch factor C notch 3 Set t 0 and t 1 4 Set the A ) of the patches in the domains to user specified value ngn (0 5 Optional: Add noise distributed normally with mean and variance set by user, A ( 0) A (0) + α α ngn ngn 1 ( 0) P Angn 6 If t 0, A (0) and if t > 0, A ( t) A ( t 1) + t f 2 8 _ ( t) i i 1 her ( 0) Pher Cnotch Atot _ 7 Calculate A tot ( t) A, where i represents the eight neighboring cells 8 If t 0, A (0) and if t > 0, Aher ( t) Aher ( t 1) + t f 3 9 If A ngn (t) is greater than the threshold specified by user, mark the patch as neuron 10 Set t t Calculate Angn ( t) Angn ( t 1) + t f1 12 Optional: Add noise, normally distributed random number with mean 0 and variance specified by user, Angn ( t) Angn ( t) + α 2 13 Repeat steps 6 12 Somitogenesis A Lewis mo with auto-inhibition and transcription and translation ays was used for the study of somitogenesis [29] The amount of proteins are calculated as follows, 4 Gaw, 2004
6 dp dt dp dt dp dt a a a [ m ] b [ p ] t Tmher 1 [ m ] b [ p ] t Tmher 7 t [ m ] b [ p ] f t Tm t t p f p f, p, and (4) (5) (6) The subscripts,,, and represent the gene,, and ta, respectively The variable p is the amount of protein (indicated by the subscript) at time t Constants a and b are the rate of protein synthesis and rate of protein degradation, respectively And, T m* is the transcription ay of the * protein The change in the amount of protein is proportional to the amount of mrna at time t T p and minus the amount of protein removed which is proportional to the amount of protein, b[p] t The amounts of,, and ta mrna are calculated using the following equations: dm dt dm dt dm dt [ f ( p, p, p )] c [ m ] [ f ( p, p, p )] c [ m ] her 7 [ f ( p, p, p )] c [ m ] f her 7 n _ n _ n _ t T p t T pher 7 t T p her 7 t t t f m m f, m, and (7) (8) (9) The subscripts are the same as above with an addition of n_ta, which is the total ta in the neighboring cells The constant c is the degradation rate of mrna The amount of mrna production is a function of the amount of,, and neighboring ta at time t T p The variable T p* is the transcription ay The mrna production functions are given below f f f k k k φ r0 + r d 1+ φ φ r 0 + rd 1+ φ φ s 0 + sd 1+ φ n _ n _ n _ n _ n _ n _ + r h + r + s h h 1 1+ φ φ 1 1+ φ φ her φ φ her 7 + r + r + s hd hd hd φ 1 ( 1+ φ ) ( 1+ φ φ ) φ her 7 1 ( 1+ φ ) ( 1+ φ φ ) φ her 7 1 ( 1+ φ ) ( 1+ φ φ ),, and (10) (11) (12) The transcription of,, and ta of a cell is positively regulated by the neighboring cells ta and negatively regulated by its own and The constant k is the rate of mrna synthesis The relative fractional importance on transcription of and that is regulated by ta alone, by and only, and by all three (ta,, and ) are given by r d, r h, and r dh, respectively For ta transcription, they are given by s d, s h, and s dh The parameters r 0 and s 0 are for basal or unregulated transcription The sum of r 0, r d, r h, and r dh and the sum of s 0, s d, s h, and s dh should both equal one The variable φ is the ratio of the current level of protein to the critical protein level when it starts the inhibition of its own production of protein, 5 Gaw, 2004
7 φ φ φ n _ p p p p p p c _ her 7 c _ n _ c _,, and (13) (14) (15) The rate of mrna degradation is directly proportional to the amount of mrna In our analysis, the amount of mrna at (t-1) is used To achieve the wave-like oscillation of and, we set up a linear gradient that is a function of the position In addition, to halt the oscillation and produce spatial patterning, we use an independent moving sigmodial gradient of fgf In our mo, once the fgf level falls below the critical fgf level and is above a threshold, the expression of,, and ta are stabilized at those current levels The fgf level controls when somites begin to form (stabilized gene expression), and the threshold allows us to skip the small peak in the beginning of the oscillation to avoid peaks that occur prior to the development of full oscillation To integrate this system of differential equations, we used Euler s method with t 1 The implementation of this mo in NetLogo is described below Somitogenesis NetLogo Algorithm: 1 Set the synthesis (a and k), degradation (b and c), and ay (T m and T p ) constants for mrna and protein of,, and ta Set the critical protein value (p crit ), of the three genes 2 Set r 0, r d, r h, s 0, s d, and s h Calculate r dh 1- r 0 - r d - r h and s dh 1 - s 0 - s d - s h 3 Set t 0 and t 1 4 Set p pher 7 p 0 5 Set up the domain For the patches in the domain do the following steps 6 Set up the initial linear gradient of and, m her C1x + C2, where m is the number of mrna, C 1 is the slope of the gradient, x is x-coordinate of the patch, and C 2 is the maximum value of /7 C3 7 Set up the initial sinusoidal fgf gradient, Fgf ( x+ ( C4 tm ) / C , where C 3 is the maximum fgf, C 4 determines the location, C 5 determines the shape of the gradient, and M determines the rate of the gradient movement 8 Set the rate of domain extension, E 9 Optional: add noise The current number of m [0] and m [0] are replaced by the absolute value of a normal random number generated with mean of current number and variance specified by user 10 If t < ( T m + T p ), set m( t) m( t 1) cm(t -1) If t < ( T m + T p ), set m( t) m( t 1) cm(t -1) If t < ( T m + T p ), set m ( t) m ( t 1) cm (t -1) 6 Gaw, 2004
8 p( t Tp) 11 If t > ( T m + T p ), set φ and m ( t) m( t 1) + t f m p c _ p ( t T p ) If t > ( T m + T p ), set φher 7 and mher 7 ( t) m ( t 1) + t f mher 7 p If c _ p ( t Tp ) t > ( T m + T p ), set φ and m ( t) m ( t 1) + t f p 12 If T mher 1 c _ ( t) p( t 1) + t f pher her 7 ( t) p ( t 1) + t f pher t >, set p 1 If t > T mher 7, set p 7 If t > Tm, set p t) p ( t 1) + t f ( 13 If fgf level is less than the critical fgf level and p < threshold, both set by user, stop m and p calculations 14 Extend the domain by one each time the rounded value of t / E increased by one 15 Move fgf gradient with the rate specified by the user 16 Set t t Repeat steps Anterior/Posterior Patterning The anterior/posterior patterning under the influence of a morphogen gradient simulation is based on Meinhardt s mo [27-28] Meinhardt proposed that for only one gene to maintain primary expression in a cell, it must exhibit autocatalysis and inhibit all alternative genes In addition, genes should subsequently initiate the expression of another gene in the presence of a morphogen: gene A initiates gene B, which initiates gene C, which initiates gene D dg dt dg dt a dg dt b dg dt c d 2 Sa ( Ga + M tm a ) R aga Ga + Gb + Gc + Gd + 1 f a, 2 Sb ( Gb + M t M bga ) R bgb Ga + Gb + Gc + Gd + 1 fb, 2 Sc ( Gc + M t M cgb ) R cgc Ga + Gb + Gc + Gd + 1 f c, and 2 Sd ( Gd + M tm dgc ) R dgd f d G + G + G + G + 1 a b c d (16) (17) (18) (19) The parameter G is the amount of gene expressed and the subscripts a, b, c, and d represent gene A, gene B, gene C, and gene D The variables S and R are the rate of synthesis and the rate of removal or degradation, respectively M t is the amount of morphogen and M is the fraction of importance the morphogen plays in promoting a particular gene Exposure to morphogen is a key factor in determining the pattern of gene expression It is known that stable, discrete compartments of gene expression can be achieved by activities of a morphogen such as wnt However, the question remains whether the concentration gradient determines gene expression patterns or is this gradient too inefficient and instead time plays the 7 Gaw, 2004
9 key role [30] To study the spatial pattern developed based on a stable and non-moving morphogen, a gradient was set up on a circular domain with the maximum concentration on the edge and minimum in the center The morphogen level was calculated with the following equation: M M ( 20 ) M max ( x) t 2 C1 ( x + 1) + max M max is the maximum morphogen level, C determines the shape and the steepness of the gradient, and x is the distance from the center of the circle For our simulation we used M max 1 and C 001 To examine if a similar pattern is obtainable by the effect of time exposure to morphogen, another circular domain was set up Initially, the whole region was set to the maximum level of morphogen; as time increases, the morphogen near the center will decay moving outward until all the morphogen levels are zero The decay is calculated by M t [ t] M t[ t] D M t[ t 1], ( 21) where D is the rate of decay The time when the decay began is determined by distance from the center of the domain When Dist C 2 t, where C 2 is the rate of movement and t is the time, the morphogen will begin to decay until the level reaches zero Again, we used Euler s method of integration with t 1 for morphogenesis system of equations Region 1 (Stable and non-moving morphogen) NetLogo Algorithm: 1 Set t 0, t 1, and G a G b G c G d 0 2 Set synthesis, degradation, and morphogen constants (S and M) to user specified values 3 Setup the morphogen gradient 4 Set G ( t) G ( t 1) + t f, a b a G ( t) G ( t 1) + t f, c c b G ( t) G ( t 1) + t f, and Gd ( t) Gd ( t 1) + t f d 5 Set t t Repeat step 3 and 4 a c b Region 2 (Time exposure) NetLogo Algorithm: 1 Set t 0, t 1, and G a G b G c G d 0 2 Set synthesis, degradation, and morphogen constants (S and M) to user specified values 3 Set M t to the maximum level 4 Calculate gene levels, same as step 4 for Region1 5 Set t t Recalculate M t, start decay from center moving toward the edge with each iteration 7 Repeat step 3 and 5 8 Gaw, 2004
10 Results and Discussion Neurogenesis The values used for rates of synthesis, degradation, saturation, self-inhibition, and ngn threshold used for the simulation are listed in Table 1 These values were empirically determined In studying the effect of notch signaling, the number of cells fated to become neurons was determined as a function of C notch (See Table 2) Using our specified values in Table 1 and initial P ngn 06, the range of C notch was found to be small, 0 03, where C notch 0 fated all cells to become neurons and C notch 03 fated only 2 cells to become neurons Since there was no noise in the initial P ngn or in subsequent iterations, the neuron-fated cells are determined solely by the cell s location and the geometry of the domains as illustrated in Fig 4 This is obvious since cells on the edge of the domains are inhibited less by the neighboring cells We chose C notch 075 to study the effect the initial amount of ngn (P ngn [0]), Type 1 noise, and Type 2 noise have on the pattern The effects of the P ngn initial condition is given in Table 3 and Fig 5 Upon visual inspection of the pattern with different P ngn [0] and comparing the change in the number of neuron-fated cells, two critical points were observed, P ngn [0] 053 and P ngn [0] 063 The introduction of Type 1 and Type 2 noise (described below) slightly decreases the overall number of neuron-fated cells, but increases the number of neuron-fated cells in the inner regions of the domains The effects of Type 1 and Type 2 noise were tested with P ngn [0] 058, since it was the midpoint of the two critical points; results are given in Table 4, Table 5, and Fig 6 Type 1 noise is variance introduced only to the P ngn [0] This is achieved by replacing the initial P ngn with a random number generated by a normal distribution with mean P ngn and different variance Type 2 noise is Type 1 noise plus normally distributed noise (mean 0 and variance specified by user) between iterations The variance should be small to represent small differences in gene expression and therefore the variance of 002, 003, and 004 were chosen From the visual inspection, it can be concluded that Type 1 noise affects the pattern the most, while Type 2 noise has a secondary effect Of course, increasing the mean of the noise added during iteration will increase the importance of the Type 2 noise Somitogenesis The synthesis and degradation rates, ay, critical protein levels, threshold, and relative importance of regulation used in our simulation were taken from Lewis s mo and are listed in Table 7a & 7b We use the following equations for / gradients and fgf gradient, m( her 1, ) 1* x + 18 and 50 fgf ( x+ (80 03t)) / The variable x is the x-coordinate of the cell The initial domain and the initial gradients are from x -80 to x 0 The fgf gradient moves independently through the domain to the right at the rate of 03 per iteration Simultaneously, the domain also extends at the rate of 01 per iteration and takes on the condition of the cell immediately to the left The critical point for fgf is 9 Gaw, 2004
11 set at 10 Therefore, the oscillation of genes is halted for patches with fgf less than 10 and greater than 200 (to skip the first peak) We began by examining the role of the initial / gradients and the initial fgf gradients on the formation of somites The / gradients are found not to play a role in the size or the formation time of somites (Table 8) It does provide the mechanism to produce the sweeping wave-like motion of / expressions across the domain However, this motion was only sustained for a couple of cycles To have the motion last longer, it is highly probable that the parameters listed in Table 7 could be tweaked, but this was not examined Although the shape of the fgf gradient is not significant in somite formation time, it does play a role in somite size determination It can be assumed from Lewis s mo that the formation time is highly dependent on the critical protein levels and the transcription and translation ays The importance of notch signaling in synchronizing a gene s oscillation of the neighboring cells was examined Noise was introduced to the initial and, determined from the gradient equation, by changing it with a random number generated from a normal distribution with a mean of current her and variance ranging from 1 5, see Table 9 It shows that when the transcription is highly regulated by ta,, and (r dh 099), synchronization occurs even with a variance of 5 Although it seems to synchronize at lower r dh (See Fig 7), it is hard to make any conclusions since oscillation ceases at r dh below 098 Anterior/Posterior Patterning Similar patterns of gene expressions under the influence of a morphogen can be obtain from either of the following cases: 1 S a > S b > S c > S d and M a < M b < M c < M d or 2 S a < S b < S c < S d and M a > M b > M c > M d The values used for the remaining parameters are given in Table 10a and 10b In both cases, gene A is first expressed, then gene B is promoted except for a small region at the center of the circle As time increases, in the gene B expressing region, gene C is promoted except for a region around the stable gene A expressing region Finally, gene D is promoted starting at the edge of the circle (gene C expressing region) and continues to increase until the expressions are all stabilized, see Fig 8 and Fig 9 Furthermore, patterns due to time exposure to morphogen can produce patterns similar to those obtained by a stable exposure to the morphogen gradient by tweaking the decay rate of morphogen and speed of the increase in the decay region of the time-exposure study Future Work First, our study of the systems of equations presented here is very basic Numerical analyses were performed for all these systems using Euler s method with step size of one We did not study the error associated with this method It would be valuable to examine just how much of the result is due to bias in the numerical method and how much is actually due to the biological process mo Perhaps, use the Runge-Kutta method which is known to be a more accurate numerical procedure for obtaining approximate solutions to differential equations Furthermore, 10 Gaw, 2004
12 it would be ideal to perform dynamical systems analysis to better mo and understand these systems Secondly, our simulation can be modified to be more robust, perhaps by extending the mos to incorporate more factors It is important that enough aspects of the system are taken into account and are accurately known in the mos For neurogenesis, the mo can be modified to include a factor to study the unregulated expression of ngn The noise added during iteration can be construed as the basal expression, but separating the two will allow users to better study and manipulate the mo system The independent moving gradient in somitogenesis simulation is to halt the oscillation seems to be a brute force way to achieve our objective It may be more reasonable to incorporate fgf in the equations to stop the oscillation, then to reach and maintain, her2, and ta expressions In the morphogenesis mo, two things can be added to improve the mo The first one is a minor one; instead of setting up the regions from the start, allow the region to grow with time This may give an insight into how cell growth plays a role in the pattern formation The second one, the effect of the antagonist on the morphogen should be further studied This has been studied and in the anterior/posterior patterning, tcf has been shown to play a critical role in determining the sizes of midbrain and hindbrain This has been studied for the morphogen gradient case but not thoroughly studied for the time-exposure case There have been little or no mathematical-mo studies for neurogenesis, somitogenesis, and morphogenesis Although our study here is basic, overall, moing is an excellent way to study biological processes The NetLogo environment provides a world where cell-cell communication alone with gene interaction among itself and the environment can be scrutinized and studied effectively However, moing alone is not enough to make conclusions regarding biological systems It is necessary to use simulation in parallel with bench work to design better experiments and to present compelling evidence for conclusions drawn 11 Gaw, 2004
13 Table 1 These are the values used in the analysis for the rate of synthesis (P) and rate of degradation (R) for ngn, ta, and her The last column is the rate of saturation for ngn and rate of self-inhibition of ta The threshold of 4 was used for ngn Parameter Gene Ngn ta her Rate of synthesis (P) Rate of Degradation (R) Rate of Saturation or Self- Inhibition (S) Threshold None Table 2 Number of cells that become neurons by varying notch factor with P ngn [0] 06, Threshold 4, and no noise in the initial P ngn The number of iterations required to reach stabilized gene expression is given in the last column Notch Factor Neuron (C notch ) Number Percent % % % % % 03 2 < 1 % 12 Gaw, 2004
14 Table 3 In determining what initial condition (P ngn [0]) to use, two critical points were found, * and ** To examine the role of noise, P ngn [0] 058 (midpoint between the two critical points) and C notch 0075 were used The number of neurons with Type 1 (P ngn [0] is distributed with mean 058 and variance 003) and Type 2 (Type 1 noise and noise distributed normally with mean 0 and variance 003 is added to P ngn [t] at every iteration) See Table 4 and Table 5 for details Number of neurons for Type 1 and Type 2 noise is the average of 10 samples P ngn Number of neurons No noise Type 1 noise (avg) Type 2 noise (avg) * ** Table 4 Number of neurons with Type 1 noise Average and standard deviation are rounded up to next integer Conditions 1 P ngn [0] C notch Noise in only P ngn [0] with mean P ngn [0] 4 Iterate until stabilized Number of neurons Variance 002 Variance 003 Variance Average Standard Deviation Gaw, 2004
15 Table 5 Number of neuron with Type 2 Noise Average and standard deviation are rounded up to next integer Conditions 1 P ngn [0] C notch Noise in only P ngn [0] with mean P ngn [0] and variance Add noise with normal distribution (mean 0) to P ngn during every iterations 5 t 100 Number of neurons Variance 002 Variance 003 Variance Average Standard Deviation Table 6 Average number of neurons with varying C notch and Type 1 (mean Pngn[0] and variance 003) and Type 2 (mean 0 and variance 003) noise conditions See Table 4 and Table 5 for details Sample size is 10 Notch Average Number of Neurons Factor (C notch ) No Noise Type 1 Noise Type 2 Noise Gaw, 2004
16 Table 7 Parameters and their values used in somitogenesis simulation a Synthesis and degradation rates, transcription and translation ays, critical protein level, and threshold of Parameter Gene ta Rate of protein synthesis (a) Rate of protein degradation (b) Transcription ay (T m ) Rate of mrna synthesis (k) Rate of mrna gradation (c) Translation ay (T p ) Critical protein level (P crit ) Threshold 200 b The values of relative importance of regulated and unregulated transcription of,, and ta The subscribes 0, d, h, and dh represent unregulated, negatively regulated by ta, positively regulated by /7, and regulated both by ta and /7 Parameter and ta R r d - 0 r h - 0 r dh S s d 0 - s h 1 - s dh 0-15 Gaw, 2004
17 Table 8 The average time required for formation of 1 somite after 7 somites at varying initial / gradients (C 1 ) and varying rates of fgf movement, M Slope of the initial / gradients, C 1 (C 2 18) Rate of fgf movement, M (C 3 50, C 4 80, and C 5 382) Time (formation of 1 somite) Size (1 somite, number of rows) Table 9 The expression levels of p for 3 neighboring cells lined up in a column at different important levels of regulated transcription (r dh ), variance (var) and time (t) r dh 099 Var Expression of (p ) for Three Neighbor Cells t 10 t 30 t Gaw, 2004
18 Table 10 Values used in the morphogenesis simulation a Parameters where S a > S b > S c > S d and M a < M b < M c < M d Parameter Gene A B C D Rate synthesis (S) Rate gradation (R) Importance of morphogen (M) Morphogen Region 1 Region 2 Max Morphgen (M max ) C Decay Rate (D) C b Parameters where S a < S b < S c < S d and M a > M b > M c > M d Parameter Gene A B C D Rate synthesis (S) Rate gradation (R) Importance of morphogen (M) Morphogen Region 1 Region 2 Max Morphogen (M max ) C Decay Rate (D) C Gaw, 2004
19 Figure 1 Lateral inhibition through notch signaling ta ngn 1 her4 Notch Notch her4 ngn 1 ta Figure 2 Diagram of the notch signaling pathway for somitogenesis /7 Transcription ay Translation ay Transcription ay Translation ay /7 Transcription ay ta ta Transcription ay Translation ay Notch Notch Translation ay 18 Gaw, 2004
20 Figure 3 Schematic used in the simulation of spatially differentiated genes under the influence of morphogen morphogen Gene A Gene B Gene C Gene D Figure 4 Distributions of cells that are fated to become neurons at different notch factors, C notch, with parameters listed in Table 1, Pngn[0] 06, and no noise At C notch 0025, all the cells are fated to become neurons a C notch 0025 b C notch 005 c C notch 0075 patches 1171 (100%) patches 785 (67%) patches 568 (49%) 19 Gaw, 2004
21 Figure 5 Distributions of cells that are fated to become neurons with different initial ngn levels, P ngn [0] and no noise The parameters listed in Table 1 were used, along with C notch 075 Under these conditions, P ngn [0] 053 and P ngn [0] 063 are two critical points CRITICAL POINT 1 P ngn [0] 052 P ngn [0] 053 CRITICAL POINT 2 P ngn [0] 062 P ngn [0] Gaw, 2004
22 Figure 6 Effects of Type 1 noise where P ngn [0] is a normal distribution with mean 058 and different variance (var) Parameters in Table 1 and C notch 0075 were used Effects of Type 2 noise where P ngn [0] is a normal distribution with mean 058 and var 003 and noise that is normally distributed with mean 0 and different variance was added to P ngn [t] at each interval Parameters in Table 1 and C notch 0075 were used TYPE 1 NOISE var 002 var 003 var 004 TYPE 2 NOISE var 002 var 003 var Gaw, 2004
23 Figure 7 Somites formation simulation using NetLogo The upper pictures represent the presomitic mesoderm region budding off somites represented by expression of, white is low expression and red is high expression The bottom graphs shows the synchronization of in 3 neighboring patches located in a column for different r hd values a r hd 099 b r hd 098 c r hd 097 d r hd Gaw, 2004
24 Figure 8 Graphical representation of the gene expression for different regions using S a < S b < S c < S d and M a > M b > M c > M d, and R a R b R c R d (see Table 10b for values) a Region where gene A is expressed, b Region where B is expressed, c Region where C is expressed, and d Region where D is express Overtime the expression of these genes are stable and are able to inhibit the expression of alternate genes within a specific region a b c d 23 Gaw, 2004
25 Figure 9 Pattern formation in gene expression under the influence of a morphogen a and b are patterns obtained using parameter values from Table 10a, and c and d are patterns using parameter values from Table 10b a b Stable-Morphogen Concentration Study Morphogen Time-Exposure Study c d Stable-Morphogen Concentration Study Morphogen Time-Exposure Study 24 Gaw, 2004
26 Reference 1 Chitnis, Ajay B (1995) The Role of Notch in Lateral Inhibition and Cell Fate Specification Mol Cell Neurosci 6: Blader, Patrick, Fischer, Nadine, Gradwohl, Gerard, Guillemot, François and Strägke (1997) The activity of Neurogenin 1 is controlled by local cues in the zebrafish Development 124: Haddon, Catherine, Jiang, Yun-Jin, Smithers, Lucy and Lewis, Julian (1998) Delta- Notch signaling and the patterning of sensory cell differentiation in the zebrafish ear: evidence from the mind bomb mutant Development 125: Haddon, Catherine, Smithers, Lucy, Schneider-Maunoury, Sylvie, Coche, Thierry, Henrique, Domingos and Lewis, Julian (1998) Multiple ta genes and lateral inhibition in zebrafish primary neurogenesis Development 125: Itoh, Motoyuki and Chitnis, Ajay B (2001) Expression of proneural and neurogenic genes in the zebrafish lateral line primordium correlates with selection of hair cell fate in neuromasts Mech Dev 102: Cornell, Robert A and Eisen, Judith S (2002) Delta/Notch signaling promotes formation of zebrafish neural crest by repressing Neurogenin 1 function Development 129: Hans, Stefan and Campos-Ortega, José A (2002) On the organization of the regulatory region of the zebrafish tad gene Development 129: Hicks, Carol, Laki, Ena, Lindsell, Claire, Hshieh, James J-D, Hatward, Diane, Collazo, Andres and Weinmaster, Gerry (2002) A Secreted Delta1-Fc Fusion Protein Functions Both As An Activator and Inhibitor of Notch1 Signaling J Neurosci Research 68: Itoh, Motoyuki, Kim Cheol_Hee, Palardy, Gregory, Oda, Takaya, Jiang, Yun-Jin, Maust, Donovan, Yeo, Dang-Yeob, Lorick, Kevin, Wright, Gavin J, Ariza-McNaughton, Linda, Weissman, Allan M, Lewis, Julian, Chandrasekharappa, Settara C and Chitnis, Ajay B (2003) Mind Bomb Is a ubiquitin Ligase that Is Essential for Efficient Activation of Notch Signaling by Delta Dev Cell 4: Saga, Yumiko and Takeda, Hiroyuki (2001) The making of the somite: molecular events in vertebrate segmentation Genetics 2: Kulesa, Paul M and Fraser, Scott E (2002) Cell Dynamics During Somite Boundary Formation revealed by Time-Lapse Analysis Science 298: Gaw, 2004
27 12 Pourquié, Olivier (2003) The Segmentation Clock: Converting Embryonic Time into Spatial Pattern Science 301: Sawada, Atsushi, Fritz, Andreas, Jiang, Yun-Jin, Yomamoto, Akihito, Yamasu, Kyo, Kuroiwa, Atsushi, Saga, Yumiko and Takeda, Hiroyuki (2000) Zebrafish Mesp family genes, mesp-a and mesp-b are segmentally expressed in the presomitic mesoderm, and Mesp-b confers the anterior identity to the developing somites Development 127: Collier, J R, McInerney, D, Schnell, S, Maini, P K, Gavaghan, D J, Houston, P and Stern C D (2000) A Cell Cycle Mo for Somitogenesis: Mathematical Formulation and Numerical Simulation J theor Biol 207: Schnell, Santiago and Maini Philip K (2000) Clock and Induction Mo for Somitogenesis Dev Dynam 217: Kerszberg, Michel and Wolpert, Lewis (2000) A Clock and Trail Mo for Somite Formation, Specialization and Polarization J theor Biol 205: Kærn, Mads, Menzinger, Michael and Hunding Axel (2000) Segmentation and Somitogenesis Derived from Phase Dynamics in Growing Oscillatory Media J theor Biol 207: Holley, Scott A, Geisler, Robert and Nüsslein-Volhard, Christiane (2000) Control of expression during zebrafish somitogenesis by a Delta-dependent oscillator and an independent wave-front activity Genes Dev 14: Holley, Scott A and Takeda, Hyroyuki (2002) Catching a wave: the oscillator and wavefront that create the zebrafish somite Cell Dev Bio 13: Jiang, Yun-Jin, Aerne, Birgit, Smithers, Lucy, Haddon, Catherine, Ish-Horowicz, David and Lewis, Julian (2000) Notch signaling and the synchronization of the somite segmentation clock Nature 408: Serth, Katrin, Schuster-Gossler, Karin, Cordes, Ralf and Gossler, Achim (2003) Transcriptional oscillation of Lunatic fringe is essential for somitogenesis Genes Dev 17: Jouve, Caroline, Iimura, Tadahiro and Pourquié, Olivier (2002) Onset of the segmentation clock in the chick embryo: evidence for oscillations in the somite precursors in the primitive streak Development 129: Dubrulle, Julien, McGrew Michael J and Pourquié, Olivier (2001) FGF Signaling Controls Somite Boundary Position and Regulates Segmentation Clock Control of Spatiotemporal Hox Gene Activation Cell 106: Gaw, 2004
28 24 Holley, Scott A, Jülich, Dörthe, Rauch, Gerd-Jörg, Geisler, Robert and Nüsslein-Volhard, Christiane (2002) and the notch pathway function within the oscillator mechanism that regulates zebrafish somiteogenesis Development 129: Kim, Cheol-Hee, Oda, Takaya, Itoh, Motoyuki, Jiang, Di, Artinger, Kristin Bruk, Chandrasekharappa, Settara C, Driever, Wolfgang and Chitnis, Ajay B (2000) Repressor activity of Headless/Tcf3 is essential for vertebrate head formation Nature 407: Dorsky, Richard I, Itoh, Motoyuki, Moon, Randall T and Chitnis, Ajay (1937) Two tcf3 genes cooperate to pattern the zebrafish brain Development 130: Meinhardt, Hans (1982) Mos of Biological Pattern Formation 28 Meinhardt, Hans, Fowler, Deborah R and Prusinkiewicz, Przemyslaw (1998) The Algorithmic Beauty of Sea Shells 29 Lewis, Julian (2003) Autoinhibition with transcriptional ay: a simple mechanism for the zebrafish somitogenesis oscillator Unpublished as of Aug 2003 julianlewis@cancerorguk 30 Pagè, Françoise and Kerridge Stephen (2000) Morphogen gradients a question of time or concentration? TIG 16: Gaw, 2004
Simulating Gene Expression Patterns During Zebrafish Embryo Development
 Simulting ene Expression Ptterns During Zebrfish Embryo Development Ei-Ei w Ajy B. Chitnis Ntionl Institute of Helth (NIH) Ntionl Institute of Child Helth nd Humn Development (NICHD) Lbortory of Moleculr
Simulting ene Expression Ptterns During Zebrfish Embryo Development Ei-Ei w Ajy B. Chitnis Ntionl Institute of Helth (NIH) Ntionl Institute of Child Helth nd Humn Development (NICHD) Lbortory of Moleculr
arxiv: v1 [q-bio.qm] 23 Apr 2007
![arxiv: v1 [q-bio.qm] 23 Apr 2007 arxiv: v1 [q-bio.qm] 23 Apr 2007](/thumbs/95/124440410.jpg) Converting genetic networ oscillations into somite spatial pattern K. I. Mazzitello, 1,2 C. M. Arizmendi, 2 and H. G. E. Hentschel 3 1 CONICET 2 Facultad de Ingeniería, Universidad Nacional de Mar del
Converting genetic networ oscillations into somite spatial pattern K. I. Mazzitello, 1,2 C. M. Arizmendi, 2 and H. G. E. Hentschel 3 1 CONICET 2 Facultad de Ingeniería, Universidad Nacional de Mar del
A systems approach to biology
 A systems approach to biology SB200 Lecture 7 7 October 2008 Jeremy Gunawardena jeremy@hms.harvard.edu Recap of Lecture 6 In phage lambda, cooperativity leads to bistability and hysteresis In HIV-1, sequestration
A systems approach to biology SB200 Lecture 7 7 October 2008 Jeremy Gunawardena jeremy@hms.harvard.edu Recap of Lecture 6 In phage lambda, cooperativity leads to bistability and hysteresis In HIV-1, sequestration
Modeling neural patterning; exploring the gene regulatory networks in. developing zebrafish embryos
 Modeling neural patterning; exploring the gene regulatory networks in developing zebrafish embryos by Charu Gaur An Internship Report presented in partial fulfillment of the requirements for the degree
Modeling neural patterning; exploring the gene regulatory networks in developing zebrafish embryos by Charu Gaur An Internship Report presented in partial fulfillment of the requirements for the degree
2/23/09. Regional differentiation of mesoderm. Morphological changes at early postgastrulation. Segments organize the body plan during embryogenesis
 Regional differentiation of mesoderm Axial Paraxial Intermediate Somatic Splanchnic Chick embryo Morphological changes at early postgastrulation stages Segments organize the body plan during embryogenesis
Regional differentiation of mesoderm Axial Paraxial Intermediate Somatic Splanchnic Chick embryo Morphological changes at early postgastrulation stages Segments organize the body plan during embryogenesis
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
MODELLING PERIODIC OSCILLATIONS DURING SOMITOGENESIS. Peng Feng. Menaka Navaratna. (Communicated by Yang Kuang)
 MATHEMATICAL BIOSCIENCES http://www.mbejournal.org/ AND ENGINEERING Volume 4, Number 4, October 2007 pp. 661 673 MODELLING PERIODIC OSCILLATIONS DURING SOMITOGENESIS Peng Feng Department of Physical Sciences
MATHEMATICAL BIOSCIENCES http://www.mbejournal.org/ AND ENGINEERING Volume 4, Number 4, October 2007 pp. 661 673 MODELLING PERIODIC OSCILLATIONS DURING SOMITOGENESIS Peng Feng Department of Physical Sciences
Role of Organizer Chages in Late Frog Embryos
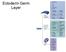 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
ASpatio-TemporalModelofNotchSignallinginthe Zebrafish Segmentation Clock: Conditions for Synchronised Oscillatory Dynamics
 ASpatio-TemporalModelofNotchSignallinginthe Zebrafish Segmentation Clock: Conditions for Synchronised Oscillatory Dynamics Alan J. Terry 1 *, Marc Sturrock 1, J. Kim Dale 2, Miguel Maroto 2, Mark A. J.
ASpatio-TemporalModelofNotchSignallinginthe Zebrafish Segmentation Clock: Conditions for Synchronised Oscillatory Dynamics Alan J. Terry 1 *, Marc Sturrock 1, J. Kim Dale 2, Miguel Maroto 2, Mark A. J.
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
Practical Lessons from Theoretical Models about the Somitogenesis
 REVIEW Practical Lessons from Theoretical Models about the Somitogenesis Aitor González and Ryoichiro Kageyama Institute for Virus Research, Kyoto University, and Japan Science and Technology Agency, CREST
REVIEW Practical Lessons from Theoretical Models about the Somitogenesis Aitor González and Ryoichiro Kageyama Institute for Virus Research, Kyoto University, and Japan Science and Technology Agency, CREST
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
THE MAKING OF THE SOMITE: MOLECULAR EVENTS IN VERTEBRATE SEGMENTATION. Yumiko Saga* and Hiroyuki Takeda
 THE MAKING OF THE SOMITE: MOLECULAR EVENTS IN VERTEBRATE SEGMENTATION Yumiko Saga* and Hiroyuki Takeda The reiterated structures of the vertebrate axial skeleton, spinal nervous system and body muscle
THE MAKING OF THE SOMITE: MOLECULAR EVENTS IN VERTEBRATE SEGMENTATION Yumiko Saga* and Hiroyuki Takeda The reiterated structures of the vertebrate axial skeleton, spinal nervous system and body muscle
Somitogenesis Clock-Wave Initiation Requires Differential Decay and Multiple Binding Sites for Clock Protein
 Requires Differential Decay and Multiple Binding Sites for Clock Protein Mark Campanelli 1, Tomáš Gedeon 2 * 1 Department of Mathematics and Computer Science, Southwest Minnesota State University, Marshall,
Requires Differential Decay and Multiple Binding Sites for Clock Protein Mark Campanelli 1, Tomáš Gedeon 2 * 1 Department of Mathematics and Computer Science, Southwest Minnesota State University, Marshall,
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11804 a Tailbud after cutting PSM after cutting b 3500 3000 2500 mean intensity 2000 1500 1000 ROI1 (TB) ROI2 (PSM) 500 0 0 1 2 3 4 5 6 7 8 9 time (h) Supplementary Fig.1 Lfng gene activity
doi:10.1038/nature11804 a Tailbud after cutting PSM after cutting b 3500 3000 2500 mean intensity 2000 1500 1000 ROI1 (TB) ROI2 (PSM) 500 0 0 1 2 3 4 5 6 7 8 9 time (h) Supplementary Fig.1 Lfng gene activity
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Autoinhibition with Transcriptional Delay: A Simple Mechanism for the Zebrafish Somitogenesis Oscillator
 Current Biology, Vol. 13, 1398 1408, August 19, 2003, 2003 Elsevier Science Ltd. All rights reserved. DOI 10.1016/S0960-9822(03)00534-7 Autoinhibition with Transcriptional Delay: A Simple Mechanism for
Current Biology, Vol. 13, 1398 1408, August 19, 2003, 2003 Elsevier Science Ltd. All rights reserved. DOI 10.1016/S0960-9822(03)00534-7 Autoinhibition with Transcriptional Delay: A Simple Mechanism for
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
A mathematical investigation of a Clock and Wavefront model for somitogenesis
 J. Math. Biol. 52, 458 482 (6) Mathematical Biology Digital Object Identifier (DOI): 1.17/s285-5-362-2 R.E. Baker S. Schnell P.K. Maini A mathematical investigation of a Clock and Wavefront model for somitogenesis
J. Math. Biol. 52, 458 482 (6) Mathematical Biology Digital Object Identifier (DOI): 1.17/s285-5-362-2 R.E. Baker S. Schnell P.K. Maini A mathematical investigation of a Clock and Wavefront model for somitogenesis
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
6 Mathematical Models for Somite Formation
 6 Mathematical Models for Somite Formation Ruth E. Baker, Santiago Schnell, and Philip K. Maini, Centre for Mathematical Biology, Mathematical Institute, University of Oxford, 24-29 St. Giles, Oxford OX1
6 Mathematical Models for Somite Formation Ruth E. Baker, Santiago Schnell, and Philip K. Maini, Centre for Mathematical Biology, Mathematical Institute, University of Oxford, 24-29 St. Giles, Oxford OX1
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Avian hairy Gene Expression Iden4fies a Molecular Clock Linked to Vertebrate Segmenta4on and Somitogenesis
 Avian hairy Gene Expression Iden4fies a Molecular Clock Linked to Vertebrate Segmenta4on and Somitogenesis Pourquie, et. al., 1997 Swetha Mummini, Victoria Fong What is a somite? Defini4on: Segmental block
Avian hairy Gene Expression Iden4fies a Molecular Clock Linked to Vertebrate Segmenta4on and Somitogenesis Pourquie, et. al., 1997 Swetha Mummini, Victoria Fong What is a somite? Defini4on: Segmental block
A Proposed Mechanism for the Interaction of the Segmentation Clock and the Determination Front in Somitogenesis
 A Proposed Mechanism for the Interaction of the Segmentation Clock and the Determination Front in Somitogenesis Moisés Santillán 1,2 *, Michael C. Mackey 1 1 Campus Monterrey, Centro de Investigación y
A Proposed Mechanism for the Interaction of the Segmentation Clock and the Determination Front in Somitogenesis Moisés Santillán 1,2 *, Michael C. Mackey 1 1 Campus Monterrey, Centro de Investigación y
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Simulation of Gene Regulatory Networks
 Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
In ovo time-lapse analysis after dorsal neural tube ablation shows rerouting of chick hindbrain neural crest
 In ovo time-lapse analysis after dorsal neural tube ablation shows rerouting of chick hindbrain neural crest Paul Kulesa, Marianne Bronner-Fraser and Scott Fraser (2000) Presented by Diandra Lucia Background
In ovo time-lapse analysis after dorsal neural tube ablation shows rerouting of chick hindbrain neural crest Paul Kulesa, Marianne Bronner-Fraser and Scott Fraser (2000) Presented by Diandra Lucia Background
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
presumptiv e germ layers during Gastrulatio n and neurulation Somites
 Vertebrate embryos are similar at the phylotypic stage Patterning the Vertebrate Body Plan II: Mesoderm & Early Nervous System Wolpert L, Beddington R, Jessell T, Lawrence P, Meyerowitz E, Smith J. (2001)
Vertebrate embryos are similar at the phylotypic stage Patterning the Vertebrate Body Plan II: Mesoderm & Early Nervous System Wolpert L, Beddington R, Jessell T, Lawrence P, Meyerowitz E, Smith J. (2001)
Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension
 Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension Nigel C. Harrison 1, Ruth Diez del Corral 2 *, Bakhtier Vasiev 1 * 1 Department of Mathematical
Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension Nigel C. Harrison 1, Ruth Diez del Corral 2 *, Bakhtier Vasiev 1 * 1 Department of Mathematical
Revisiting the involvement of signaling gradients in somitogenesis
 VIEWPOINT Revisiting the involvement of signaling gradients in somitogenesis Moises Mallo Instituto Gulbenkian de Ciencia, Oeiras, Portugal Keywords development; mesoderm; segmentation; signaling; somite
VIEWPOINT Revisiting the involvement of signaling gradients in somitogenesis Moises Mallo Instituto Gulbenkian de Ciencia, Oeiras, Portugal Keywords development; mesoderm; segmentation; signaling; somite
Rui Dilão NonLinear Dynamics Group, IST
 1st Conference on Computational Interdisciplinary Sciences (CCIS 2010) 23-27 August 2010, INPE, São José dos Campos, Brasil Modeling, Simulating and Calibrating Genetic Regulatory Networks: An Application
1st Conference on Computational Interdisciplinary Sciences (CCIS 2010) 23-27 August 2010, INPE, São José dos Campos, Brasil Modeling, Simulating and Calibrating Genetic Regulatory Networks: An Application
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Periodic Pattern Formation In Developmental Biology: A Study Of The Mechanisms Underlying Somitogenesis
 Periodic Pattern Formation In Developmental Biology: A Study Of The Mechanisms Underlying Somitogenesis Ruth E. Baker WADHAM COLLEGE University of Oxford A thesis submitted for the degree of Doctor of
Periodic Pattern Formation In Developmental Biology: A Study Of The Mechanisms Underlying Somitogenesis Ruth E. Baker WADHAM COLLEGE University of Oxford A thesis submitted for the degree of Doctor of
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Name KEY. Biology Developmental Biology Winter Quarter Midterm 3 KEY
 Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2009 Midterm 3 KEY All of the 25 multi-choice questions are single-answer. Choose the best answer. (4 pts each) Place
Name KEY 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2009 Midterm 3 KEY All of the 25 multi-choice questions are single-answer. Choose the best answer. (4 pts each) Place
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Onset of the segmentation clock in the chick embryo: evidence for oscillations in the somite precursors in the primitive streak
 Development 129, 1107-1117 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV2742 1107 Onset of the segmentation clock in the chick embryo: evidence for oscillations in the somite
Development 129, 1107-1117 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV2742 1107 Onset of the segmentation clock in the chick embryo: evidence for oscillations in the somite
Somites without a clock
 Dias, AS; de Almeida, I; Belmonte, JM; Glazier, JA; Stern, CD; (2014) Somites Without a Clock. Science 10.1126/science.1247575. ARTICLE Somites without a clock Ana S. Dias 1,, Irene de Almeida 1,, Julio
Dias, AS; de Almeida, I; Belmonte, JM; Glazier, JA; Stern, CD; (2014) Somites Without a Clock. Science 10.1126/science.1247575. ARTICLE Somites without a clock Ana S. Dias 1,, Irene de Almeida 1,, Julio
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Drosophila Life Cycle
 Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
UNIVERSITY OF YORK BIOLOGY. Developmental Biology
 Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
CS-E5880 Modeling biological networks Gene regulatory networks
 CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
CS-E5880 Modeling biological networks Gene regulatory networks Jukka Intosalmi (based on slides by Harri Lähdesmäki) Department of Computer Science Aalto University January 12, 2018 Outline Modeling gene
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Biology 218, practise Exam 2, 2011
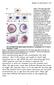 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
A synthetic oscillatory network of transcriptional regulators
 A synthetic oscillatory network of transcriptional regulators Michael B. Elowitz & Stanislas Leibler, Nature, 403, 2000 igem Team Heidelberg 2008 Journal Club Andreas Kühne Introduction Networks of interacting
A synthetic oscillatory network of transcriptional regulators Michael B. Elowitz & Stanislas Leibler, Nature, 403, 2000 igem Team Heidelberg 2008 Journal Club Andreas Kühne Introduction Networks of interacting
The oscillation of Notch activation, but not its boundary, is required for somite border formation and rostral-caudal patterning within a somite
 Access the Development most First recent posted version epress online at http://dev.biologists.org/lookup/doi/10.1242/dev.044545 on online 24 March publication 2010 as 10.1242/dev.044545 date March 2010
Access the Development most First recent posted version epress online at http://dev.biologists.org/lookup/doi/10.1242/dev.044545 on online 24 March publication 2010 as 10.1242/dev.044545 date March 2010
Examining the role of Lunatic fringe dosage in somitogenesis. A Senior Honors Thesis. Jason D. Lather
 Examining the role of Lunatic fringe dosage in somitogenesis A Senior Honors Thesis Presented in partial fulfillment of the requirements for graduation with distinction in Molecular Genetics in the undergraduate
Examining the role of Lunatic fringe dosage in somitogenesis A Senior Honors Thesis Presented in partial fulfillment of the requirements for graduation with distinction in Molecular Genetics in the undergraduate
StarLogo Simulation of Streaming Aggregation. Demonstration of StarLogo Simulation of Streaming. Differentiation & Pattern Formation.
 StarLogo Simulation of Streaming Aggregation 1. chemical diffuses 2. if cell is refractory (yellow) 3. then chemical degrades 4. else (it s excitable, colored white) 1. if chemical > movement threshold
StarLogo Simulation of Streaming Aggregation 1. chemical diffuses 2. if cell is refractory (yellow) 3. then chemical degrades 4. else (it s excitable, colored white) 1. if chemical > movement threshold
ARTICLE IN PRESS YDBIO-01976; No of pages: 11; 4C: 4, 5, 6, 8
 YDBIO-01976; No of pages: 11; 4C: 4, 5, 6, 8 DTD 5 Developmental Biology xx (2005) xxx xxx www.elsevier.com/locate/ydbio Generation of segment polarity in the paraxial mesoderm of the zebrafish through
YDBIO-01976; No of pages: 11; 4C: 4, 5, 6, 8 DTD 5 Developmental Biology xx (2005) xxx xxx www.elsevier.com/locate/ydbio Generation of segment polarity in the paraxial mesoderm of the zebrafish through
Coupling segmentation to axis formation
 5783 Coupling segmentation to axis formation Julien Dubrulle* and Olivier Pourquié Stowers Institute for Medical Research, 1000E 50th street, Kansas City, MO 64110, USA *Present address: Skirball Institute
5783 Coupling segmentation to axis formation Julien Dubrulle* and Olivier Pourquié Stowers Institute for Medical Research, 1000E 50th street, Kansas City, MO 64110, USA *Present address: Skirball Institute
BE 159: Signal Transduction and Mechanics in Morphogenesis
 BE 159: Signal Transduction and Mechanics in Morphogenesis Justin Bois Caltech Winter, 2018 2018 Justin Bois. This work is licensed under a Creative Commons Attribution License CC-BY 4.0. 5 Delta-Notch
BE 159: Signal Transduction and Mechanics in Morphogenesis Justin Bois Caltech Winter, 2018 2018 Justin Bois. This work is licensed under a Creative Commons Attribution License CC-BY 4.0. 5 Delta-Notch
Three different fusions led to three basic ideas: 1) If one fuses a cell in mitosis with a cell in any other stage of the cell cycle, the chromosomes
 Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
The Chick Embryo Model System
 The Chick Embryo Model System Guest Editor Claudio Stern Department of Cell & Developmental Biology, University College London, London, U.K. http://orcid.org/0000-0002-9907-889x Volume 62 Nos. 1/2/3 Special
The Chick Embryo Model System Guest Editor Claudio Stern Department of Cell & Developmental Biology, University College London, London, U.K. http://orcid.org/0000-0002-9907-889x Volume 62 Nos. 1/2/3 Special
10/2/2015. Chapter 4. Determination and Differentiation. Neuroanatomical Diversity
 Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Chapter 4 Determination and Differentiation Neuroanatomical Diversity 1 Neurochemical diversity: another important aspect of neuronal fate Neurotransmitters and their receptors Excitatory Glutamate Acetylcholine
Dr. Rosemary Renaut, Director, Computational Biosciences
 Dr. Rosemary Renaut, renaut@asu.edu Director, Computational Biosciences www.asu.edu/compbiosci A Professional Masters Program The School of Life Sciences Mathematics and Statistics Computer Science Engineering
Dr. Rosemary Renaut, renaut@asu.edu Director, Computational Biosciences www.asu.edu/compbiosci A Professional Masters Program The School of Life Sciences Mathematics and Statistics Computer Science Engineering
A clock-work somite. Kim J. Dale and Olivier Pourquié* Review articles. 72 BioEssays 22.1 BioEssays 22:72 83, 2000 John Wiley & Sons, Inc.
 A clock-work somite Kim J. Dale and Olivier Pourquié* Summary Somites are transient structures which represent the most overt segmental feature of the vertebrate embryo. The strict temporal regulation
A clock-work somite Kim J. Dale and Olivier Pourquié* Summary Somites are transient structures which represent the most overt segmental feature of the vertebrate embryo. The strict temporal regulation
Cellular Systems Biology or Biological Network Analysis
 Cellular Systems Biology or Biological Network Analysis Joel S. Bader Department of Biomedical Engineering Johns Hopkins University (c) 2012 December 4, 2012 1 Preface Cells are systems. Standard engineering
Cellular Systems Biology or Biological Network Analysis Joel S. Bader Department of Biomedical Engineering Johns Hopkins University (c) 2012 December 4, 2012 1 Preface Cells are systems. Standard engineering
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Determination of Spatiotemporal Interactions Regulating Head/Trunk Axial Pattern in the Chick Embryo
 Eukaryon, Vol. 2, January 2006, Lake Forest ollege Senior Thesis Determination of Spatiotemporal Interactions Regulating Head/Trunk Axial Pattern in the hick Embryo David Mann * Department of Biology,
Eukaryon, Vol. 2, January 2006, Lake Forest ollege Senior Thesis Determination of Spatiotemporal Interactions Regulating Head/Trunk Axial Pattern in the hick Embryo David Mann * Department of Biology,
From DNA to Diversity
 From DNA to Diversity Molecular Genetics and the Evolution of Animal Design Sean B. Carroll Jennifer K. Grenier Scott D. Weatherbee Howard Hughes Medical Institute and University of Wisconsin Madison,
From DNA to Diversity Molecular Genetics and the Evolution of Animal Design Sean B. Carroll Jennifer K. Grenier Scott D. Weatherbee Howard Hughes Medical Institute and University of Wisconsin Madison,
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
Morphogens in biological development: Drosophila example
 LSM5194 Morphogens in biological development: Drosophila example Lecture 29 The concept of morphogen gradients The concept of morphogens was proposed by L. Wolpert as a part of the positional information
LSM5194 Morphogens in biological development: Drosophila example Lecture 29 The concept of morphogen gradients The concept of morphogens was proposed by L. Wolpert as a part of the positional information
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Chapter 10 Development and Differentiation
 Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Name. Biology Developmental Biology Winter Quarter 2013 KEY. Midterm 3
 Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Name 100 Total Points Open Book Biology 411 - Developmental Biology Winter Quarter 2013 KEY Midterm 3 Read the Following Instructions: * Answer 20 questions (5 points each) out of the available 25 questions
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Developmental System System(Mathematical Topics in Biolo. Citation 数理解析研究所講究録 (1993), 827:
 Title Development of Developmental System System(Mathematical Topics in Biolo Author(s) Takeda, Yasuhiko Citation 数理解析研究所講究録 (1993), 827: 57-74 Issue Date 1993-03 URL http://hdl.handle.net/2433/83294 Right
Title Development of Developmental System System(Mathematical Topics in Biolo Author(s) Takeda, Yasuhiko Citation 数理解析研究所講究録 (1993), 827: 57-74 Issue Date 1993-03 URL http://hdl.handle.net/2433/83294 Right
Activator-Inhibitor Systems
 Activator-Inhibitor Systems Activator-Inhibitor Systems Sabine Pilari P. Kazzer, S. Pilari, M. Schulz (FU Berlin) Pattern Formation July 16, 2007 1 / 21 Activator-Inhibitor Systems Activator-Inhibitor
Activator-Inhibitor Systems Activator-Inhibitor Systems Sabine Pilari P. Kazzer, S. Pilari, M. Schulz (FU Berlin) Pattern Formation July 16, 2007 1 / 21 Activator-Inhibitor Systems Activator-Inhibitor
From Segment to Somite: Segmentation to Epithelialization Analyzed Within Quantitative Frameworks
 DEVELOPMENTAL DYNAMICS 236:1392 1402, 2007 SPECIAL ISSUE REVIEWS A PEER REVIEWED FORUM From Segment to Somite: Segmentation to Epithelialization Analyzed Within Quantitative Frameworks Paul M. Kulesa,
DEVELOPMENTAL DYNAMICS 236:1392 1402, 2007 SPECIAL ISSUE REVIEWS A PEER REVIEWED FORUM From Segment to Somite: Segmentation to Epithelialization Analyzed Within Quantitative Frameworks Paul M. Kulesa,
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Lecture 7: Simple genetic circuits I
 Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
Diffusion, Reaction, and Biological pattern formation
 Diffusion, Reaction, and Biological pattern formation Morphogenesis and positional information How do cells know what to do? Fundamental questions How do proteins in a cell segregate to front or back?
Diffusion, Reaction, and Biological pattern formation Morphogenesis and positional information How do cells know what to do? Fundamental questions How do proteins in a cell segregate to front or back?
Exploring Nonlinear Oscillator Models for the Auditory Periphery
 Exploring Nonlinear Oscillator Models for the Auditory Periphery Andrew Binder Dr. Christopher Bergevin, Supervisor October 31, 2008 1 Introduction Sound-induced vibrations are converted into electrical
Exploring Nonlinear Oscillator Models for the Auditory Periphery Andrew Binder Dr. Christopher Bergevin, Supervisor October 31, 2008 1 Introduction Sound-induced vibrations are converted into electrical
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
The segmentation clock mechanism moves up a notch
 Review The segmentation clock mechanism moves up a notch Sarah Gibb, Miguel Maroto and J. Kim Dale College of Life Sciences, University of Dundee, Dundee, DD1 5EH, Scotland, UK The vertebrate segmentation
Review The segmentation clock mechanism moves up a notch Sarah Gibb, Miguel Maroto and J. Kim Dale College of Life Sciences, University of Dundee, Dundee, DD1 5EH, Scotland, UK The vertebrate segmentation
4. Why not make all enzymes all the time (even if not needed)? Enzyme synthesis uses a lot of energy.
 1 C2005/F2401 '10-- Lecture 15 -- Last Edited: 11/02/10 01:58 PM Copyright 2010 Deborah Mowshowitz and Lawrence Chasin Department of Biological Sciences Columbia University New York, NY. Handouts: 15A
1 C2005/F2401 '10-- Lecture 15 -- Last Edited: 11/02/10 01:58 PM Copyright 2010 Deborah Mowshowitz and Lawrence Chasin Department of Biological Sciences Columbia University New York, NY. Handouts: 15A
Morphogens, modeling and patterning the neural tube: an interview with James Briscoe
 Briscoe BMC Biology (2015) 13:5 DOI 10.1186/s12915-014-0105-1 INTERVIEW Open Access Morphogens, modeling and patterning the neural tube: an interview with James Briscoe James Briscoe Abstract James Briscoe
Briscoe BMC Biology (2015) 13:5 DOI 10.1186/s12915-014-0105-1 INTERVIEW Open Access Morphogens, modeling and patterning the neural tube: an interview with James Briscoe James Briscoe Abstract James Briscoe
Vertebrate Segmentation: From Cyclic Gene Networks to Scoliosis
 Leading Edge Review Vertebrate Segmentation: From Cyclic Gene Networks to Scoliosis Olivier Pourquié 1, * 1 Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), CNRS (UMR 7104), Inserm
Leading Edge Review Vertebrate Segmentation: From Cyclic Gene Networks to Scoliosis Olivier Pourquié 1, * 1 Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), CNRS (UMR 7104), Inserm
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
