Examining the role of Lunatic fringe dosage in somitogenesis. A Senior Honors Thesis. Jason D. Lather
|
|
|
- Maurice Miles
- 5 years ago
- Views:
Transcription
1 Examining the role of Lunatic fringe dosage in somitogenesis A Senior Honors Thesis Presented in partial fulfillment of the requirements for graduation with distinction in Molecular Genetics in the undergraduate colleges of The Ohio State University By Jason D. Lather The Ohio State University June 2008 Project Advisor: Doctor Susan Cole, Department of Molecular Genetics Thesis Committee: Dr. Susan Cole Dr. Harold Fisk Dr. Heithem El-Hodiri
2 Abstract The repeating structures of the vertebrate axial skeleton are produced by the process of somitogenesis. Somites bud from the presomitic mesoderm (PSM) in an anterior to posterior direction, and are the embryonic precursors of the vertebrae and ribs. This process is highly regulated by one signaling pathway in particular: Notch signaling. This study focuses on the functions of Lunatic fringe (Lfng) which encodes a glycosyltransferase that inhibits Notch signaling. The PSM is divided into two regions: one in the posterior region (region I), a clock that times somitogenesis. In the anterior PSM (region II), rostral-caudal (R/C) patterning of somites occurs. Lfng and Notch signaling play roles in both regions of the PSM. Our lab created a novel Lfng allele that perturbs Lfng function in the clock, but preserves its function in R/C somite patterning, and found that while cyclic Lfng actually is important in anterior skeletal development, the Lfng patterning function is specifically required for tail development. This project focuses on the roles played by Lfng and somite patterning in tail development. We examined skeletal development in embryos expressing different amounts of Lfng during R/C patterning. In the absence of anterior Lfng tail development is severely affected and tail truncation is observed. As the expression of Lfng in the anterior PSM increases, less severe tail phenotypes are observed. Data obtained from this experiment suggests that anterior expression of Lfng in the rostral-caudal patterning region is critical for development of the tail and that tail development is sensitive to Lfng dosage. In order to test this hypothesis, a novel transgene expressing exogenous Lfng only in the anterior PSM has been created. We hypothesize that this anterior-specific Lfng will rescue tail development in Lfng null mice. We are currently investigating the effects of this transgene by first analyzing Lfng expression by whole mount in situ hybridization. Currently, mice are being bred in order to provide us with data to support our hypothesis. i
3 Table of Contents Abstract... Table of Contents... List of Figures... List of Tables... i ii iii iv Chapters Chapter 1 Introduction... 1 Overview of Somitogenesis... 1 Primary vs. Secondary Body Formation... 1 Two regions in the PSM: Clock vs. Patterning... 3 Notch Pathway and somitogenesis... 3 Lfng plays many roles in somitogenesis... 6 Chapter 2 Results Examining the requirements for anterior development Quantifying the posterior defects in Lfng mutant mice Production of an anterior specific Lfng transgene Chapter 3 Discussion Clock function in somitogenesis is required for proper anterior skeletal development Patterning of presomites in region II is required for proper tail development The Mesp2 promoter linked Lfng transgene lacks expression in the anterior PSM Future Directions Conclusions Chapter 4 Materials and Methods Works Cited ii
4 List of Tables Table Medians and Means of quantification Table PCR Primers iii
5 List of Figures Figure 1.1 Schematic of Presomitic Mesoderm... 9 Figure 1.2 Notch signaling Figure 1.3 Interactions in the posterior and anterior PSM Figure 1.4 The FCE1 deletion and corresponding phenotypes Figure 2.1 Lfng genotyping Figure 2.2 Anterior body defects Figure 2.3 Posterior body defects Figure 2.4 Anterior Specific transgene Figure 3.1 Transgene knocking out region II iv
6 Chapter 1 Introduction Overview of Somitogenesis Pairs of somites bud from the presomitic mesoderm (PSM) during somitogenesis. These somites later differentiate into the repeating structures of the back and tail. In mouse, a pair of somites forms every two hours; however, the time required is species specific. In chick embryos, somites bud every ninety minutes. The consistent time required for a pair of somites to bud has led many to believe that a molecular clock underlies somite formation. Notch signaling is a prime candidate for acting in the clock in somitogenesis. In addition, Notch has been shown to play an important role in the patterning of somites into their anterior and posterior regions. Analysis of one particular Notch pathway member, Lunatic fringe (Lfng), has supported conclusions that Notch signaling is important in somitogenesis. Primary vs. secondary body formation The mechanism for body development has been highly debated since 1925, Some believe that the posterior body, the tail, forms as a continuation of primary body formation, or the formation of the head and trunk. However, some also believe that the anterior body and posterior body form by two different mechanisms. In primary neurulation, or primary body formation, the anterior regions of the body are formed. The head and trunk differentiate and develop from tissues formed in early development. Regions of the vertebral column formed at this stage in develop are the cervical, thoracic, and lumbar vertebrae (Reviewed in Handrigan, et al., 2003). It is believed that primary and secondary body formation occurs through different mechanisms because different 1
7 genes are expressed during primary and secondary body formation. In primary development, gastrulation continues until the primitive streak is at the end of the PSM, but secondary development begins when gastrulation ceases and the tailbud forms. Secondary body formation seems to occur by a differently regulated mechanism than primary body formation. Secondary body formation in mouse begins around embryonic day 10.0 and ceases when somitogenesis terminates at embryonic day Structures formed during secondary body formation later differentiate into parts of the adult tail the sacral region and vertebrae of the tail. An important tissue in forming somites is the Tail Bud Mesenchyme (TBM), which is proposed to be undifferentiated mesenchyme that gives rise to somites. Without the TBM, somitogenesis ceases; in wildtype mouse embryos, the TBM is lost at E The TBM s role in tail length has been shown by grafting the TBM (Goldman, et. al, 2000). The TBM receives important signals from the Ventral Ectodermal Ridge (VER) which allows for somitogenesis to occur. The VER develops during secondary body formation and acts to signal for cell proliferation, promoting tail outgrowth. The VER appears to be a posterior body analog of the Apical Ectodermal Ridge of limb development and appear to have functional homology in their respective regions of development. Tails where the VER is removed have a normal segment appearance, but have truncated tails as a result of the cessation of somite development (Goldman, et. al, 2000). Somites are all of equal size and are equally patterned showing that the VER s role is only in tail elongation and not in patterning of somites. Further experiments showed that a VER implant on tails lacking a VER allowed somitogenesis to occur. The 2
8 VER s obvious role in tail development suggests that the PSM and the VER likely have some interaction. Two regions in the PSM: Clock vs. Patterning The tail develops from the PSM, which is divided into two regions. The posterior is termed region I and functions using a molecular clock used for cell allocation (Reviewed in Andrade, et. al, 2005). The oscillation in this particular region was suggested by finding oscillation of a gene, hairy1, in the PSM. Comparing the oscillation with embryos of the same somite number, that is, chicks less than 90 minutes apart in development showed that different localizations of hairy1 were found (Palmeirim, et. al, 1997), meaning expression of hairy1 oscillates in the chick PSM. The second region of the PSM, region II, functions to pattern each somite pair into rostral/caudal halves. Genes related to both of these regions are also members of the Notch signaling pathway. Therefore, it is reasonable to believe that Notch signaling plays an important role both in region I and region II of the PSM (Figure 1.1). Notch Pathway and somitogenesis Notch signaling functions in many important biological processes in development and cell differentiation. Notch signaling is one way cells can interact with one another. A signaling cell in this pathway displays a membrane bound ligand, often Delta, Jagged/Serrate-like. This ligand then binds to the Notch receptor found on a receiving cell. Following contact between a Notch ligand and the Notch receptor, the receptor undergoes a cleavage by Presenilin 1(Psen1), allowing the Notch Intracellular Domain (NICD) to enter the nucleus of the receiving cell and behave as a transcription factor. Notch can be modified by other members of the pathway such as Lunatic fringe, which 3
9 acts to glycosylate the Notch receptor, modulating its interactions with its ligands when expressed on the extracellular surface (Figure 1.2). Notch signaling has roles in both regions of the PSM (Reviewed in Weinmaster and Kintner, 2003). Mutations in Notch pathway members appear to have similar phenotypes. By mutating Hes7 (Chen, et. al, 2005), Notch signaling in the posterior PSM, or region I, is altered. This affects the molecular clock. The region II specific Mesp2 mutants only affect the patterning function of Notch (Morimoto, et. al, 2005). And lastly, mutating Lfng influences Notch activity both in region I and region II of the PSM (Zhang and Gridley, 1998; Evrard, et. al, 1998). Each mutant, though affecting different aspects of Notch signaling in the PSM, has a similar phenotype: a short stature accompanied by rib and vertebral defects. The similarity in phenotype leads to difficulties in determining the specific roles of each Notch member in somitogenesis. Regulation of Notch varies between region I and region II of the PSM. In region I, Lunatic fringe (Lfng) plays a key role in directing oscillatory Notch signaling during clock function. The action of LFNG, the glycosylating protein encoded by Lfng, serves to prevent communication between cells by placing sugars on the EGF repeats of the Notch receptor (reviewed in Shifley and Cole, 2007). This periodic inhibition caused by the addition of sugars tends to support to the oscillation both of active Notch signaling as well as Lfng transcript. Oscillation of the Lfng mrna plays a role in the ability to produce protein (Chen et. al, 2005) In the posterior PSM, it has been shown that oscillatory Notch activity is regulated by the interplay between Lfng and Hes7, a bhlh repressor (Chen, et. al, 2005). It has been suggested that though Notch can activate Lfng expression by its ICD, Hes7 s ability to repress Lfng expression overrides this particular 4
10 form of activation. Hes proteins encode transcription factors which behave both as activators and repressors of different genes. Though Hes7 acts to repress both its own transcription (Chen, et. al, 2005) as well as the transcription of Lfng, Hes7 actually activates Notch signaling (Reviewed in Kageyama, et. al, 2007). Hes7 transcription, like Lfng transcription, oscillates in the posterior PSM. The time required for Hes7 oscillation in mouse equals the time required for Lfng two hours. Hes7 expression is limited to the posterior PSM, but Hes1 is expressed in both regions of the PSM. Oscillation of these two transcription factors is in phase. Experiments relating to Hes7 null animals have suggested that Hes7 plays a critical role in the cell allocation clock (Figure 1.3a). Notch in region II of the PSM is likely under control of a different mechanism, as suggested by the difference in gene expressions. Interactions between Mesp2 and Notch also appear to influence patterning in the anterior PSM. Lfng expression is freed from the Notch signaling pathway and becomes under control of Mesp2 in region II. The loss of control by Notch, along with the gain of control by Mesp2 leads to the different type of expression found in the anterior PSM compared to that of the posterior PSM (Shifley and Cole, 2007). Notch signaling is not the only factor in Lfng expression. Mesp2 also has been shown to play roles in the expression of Lfng in the anterior PSM. Experiments have shown that Lfng expression is enhanced when Mesp2 is expressed (Morimoto, et. al, 2005). Using previous data, it is likely that this upregulation of Lfng in the rostral compartment leads to the downregulation of Notch only in that compartment. Some variety of change in regulation seems to play a role in this alteration in Lfng expression (Figure 1.3b). Mesp2 can bind to the Lfng promoter and affect its transcription in the 5
11 anterior PSM (Takahashi, et. al, 2007). Without Mesp2 expression in the anterior PSM, Lfng does not completely localize in the anterior (Takahashi, et. al, 2007). Lfng plays many roles in somitogenesis Lunatic fringe s (Lfng) roles in somitogenesis have been examined in great detail. When the Notch receptor is modified by the addition of the sugars added by Lfng, it is believed that the modified Notch receptor is rendered unable to interact with the signaling cell s ligands. The lack of communication between cells prevents the endocytosis of the Notch receptor and furthermore prevents cleavage by Psen1. A negative feedback loop between Notch and Lfng is one of the driving forces underlying somitogenesis (reviewed in Weinmaster and Kintner, 2003). Distinct patterns of Lfng expression are found in the two regions of the PSM. Expression oscillates every two hours in mouse embryos in the posterior PSM while it is ubiquitously expressed in region II of mouse embryos. The stable band of expression in region II is found in the rostral compartment of the presomite. It has been shown that the oscillation of Lfng in the posterior PSM is necessary for proper somitogenesis in mice (Serth, et. al, 2002). Mice with ubiquitously expressed Lfng were created and analyzed. It was found that the phenotypic defects are similar to those found in Lfng null animals. It has also been found that constitutive expression of Notch in chick embryos leads to an increase in Lfng expression (Dale, et. al, 2003). Animals defective in Lfng exhibit severe phenotypic consequences, showing the importance Lfng plays in the segmentation clock (Zhang and Gridley, 1998 and Evrard, et. al, 1998). Defects found in Lfng null animals include disordered vertebrae, resulting from the altered dynamic expression of Lfng, and truncated tails. It is unknown exactly 6
12 why Lfng null tails are severely truncated; and further study must be done. Not only do these animals exhibited truncated tails and disordered vertebrae, but they also show a variety of rib defects. Skeletal preps were done in order to visualize these defects. The loss of patterning in Lfng null animals leads to the hypothesis that patterning is required for secondary body formation. A novel allele that perturbs cyclic Lfng expression in the clock was created by removing a clock enhancer, Fringe Clock Element 1 (FCE1) (Cole, et al., 2002). In Lfng FCE mice, dynamic expression of Lfng is lost in the posterior PSM; however, stable, though slightly reduced, expression of Lfng is maintained in the anterior PSM where R/C somite patterning occurs (Figure 1.4). The difference in Lfng expression between the anterior and posterior PSM suggest different mechanisms control the anterior and posterior PSM. It has been shown that FCE1 mutants display phenotypes similar to those of Lfng null animals. Phenotypes in ΔFCE1/ΔFCE1 are similar to Lfng -/- in the anterior skeleton of the body. An interesting phenomenon occurs in the development of the spine in these mice; their anterior skeleton is very disordered, but upon reaching the posterior skeleton, the phenotype is rescued. The defects in segmentation of ΔFCE1 mutants are confined mostly to the anterior region of the body while those of the Lfng nulls range from the thoracic region through the tail (Shifley, et. al, 2008). Lfng expression in both region I and region II are required for proper development of both the anterior and posterior body. Oscillatory Lfng/Notch in the clock is important during primary somitogenesis while Lfng in region II during patterning is necessary during secondary somitogenesis. It is possible to begin analyzing what is happening with Notch1 in region II during secondary somitogenesis in order to produce these differences 7
13 in phenotype. To address this role, the dosage effects of the various transgenic mice will be examined. Next, the Lfng dosage will be manipulated by adding Lfng in an anteriorspecific manner. Lastly, potential mechanisms for regulation in region II will be analyzed. To analyze the effects of Lfng dosage on development, we first quantify the defects found amongst the various transgenic mice created, including a new genotype of FCE/-. This genotype appears to have its own unique phenotype associated with it. This quantification will be followed by the production of transgenic mice containing an anterior specific dose of Lfng driven by the Mesp2 promoter. It has been shown (Haraguchi, et. al, 2001) that the Mesp2 promoter can drive expression of LacZ in the anterior PSM. The anterior specific Lfng could serve to rescue the phenotype of some transgenic animals. If our hypothesis is correct, we would expect to see the rescue of tail development in Lfng -/- embryos. We believe rescue will occur because FCE/ FCE mice have some tail development. The sizes of somites will likely still vary since the clock will still be deleted. Creating a transgene containing a posterior-specific promoter and a Lfng coding sequence might lead to the restoration, to some extent, of somite size. This experiment would require a promoter that is dynamically expressed, as shown by the results of both Serth, et al. (2002), and Dale, et. al (2003). Oscillatory expression of Lfng is key in somite formation. 8
14 Figure 1.1 Schematic of Presomitic Mesoderm: The PSM is divided into two regions, cell allocation and rostral caudal (R/C) patterning. In region I cells are allocated while in region II, presomites are patterned into rostral and caudal halves. Somites bud in an anterior to posterior fashion, S0 through S-II represent presomites, while SI represents the most recently formed somite pair. 9
15 Figure 1.2 Notch signaling: This is a representation of the Notch Signaling pathway as it pertains to Lfng. The Notch receptor is expressed with the Notch Intracellular Domain (NICD) attached inside the cell. When a signal is received from a cell through a Delta, Serrate, or Jagged domain, the NICD is cleaved and enters the nucleus where transcription of target genes is affected. 10
16 Figure 1.3 Interactions in the posterior and anterior PSM these schematics show the gene interactions in the posterior (A) and anterior (B) PSM. Notch is capable of activating expression of Lfng and Hes7. Hes7 inhibits Lfng, and itself. Lastly, it is believed that Lfng inhibits Notch signaling, thus leading to the oscillatory expression of Lfng. (B) describes expression in the anterior PSM. Here, Mesp2 expression overrides Notch signaling and activates Lfng in the rostral compartment of the somite. 11
17 Figure 1.4 The FCE1 deletion and corresponding phenotypes (a) shows the targeted deletion of Fringe Clock Element 1 and the corresponding change in expression. (b) shows the dosage gradient produced by our various mice. Notice the decrease in loss of r/c patterning as we move from left to right. (c) shows corresponding phenotypes of adult mice. The body progressively shortens as the dosage of Lfng is lowered. The tail especially shortens. Photo s courtesy of Susan Cole and Emily Shifley 12
18 Chapter 2 Results Examining the requirements for anterior development We want to examine the role Lfng dosage plays in development of the skeleton. Since Lfng is expressed in different manners, we decided to see which region of expression is absolutely required for development of the axial skeleton. If development was solely dependent upon the clock, we would expect all mutant mice to have the same phenotype. However, if patterning is important, we would expect the FCE/ FCE and FCE/- embryos to have a different phenotype from the null and wild type embryos. To obtain our various genotypes, we cross Lfng +/- mice with other strains, such as Lfng FCE/-, or some other variety. The null allele must be kept in the heterozygous form, however, because null males tend to be sterile. We collected embryos 18.5 d.p.c from adult mice by non-survival Cesarean section, and then prepared them for examination as described in the materials and methods. After approximately one month, the end result was available for examination. Various axial skeleton defects were quantified starting with the anterior regions of the mice first. A comparison of the length of the anterior body was done by measuring the number of vertebrae present from the base of the skull to the pelvic girdle. In wild type embryos (n=25), we counted 26 ossification sites, or vertebrae, from the skull to the top of the pelvic girdle (Figure 2.2(e-h)). By removing the clock, or deleting FCE1, anterior vertebrae lost their well-maintained shape. Additionally, this removal of the clock found in our FCE/ FCE appears to impact the number of vertebrae contained in the anterior body. We found by direct counting that the total number of vertebrae in this region ranges from 16 to 21 (n=25), with an average of 18 13
19 vertebrae. This is a loss compared to the anterior body s normal length; loss of the clock leads to a loss of approximately 23% of the body length (6 missing/26 in wild type). Not only did these animals display a decrease in the anterior body length, but also the vertebrae lack consistent structure. Continuing our analysis of Lfng s role in axial skeleton development, we examined our FCE/- mice. The results were not quite as expected, with these mutant embryos showing a similar range of cervical to pelvic girdle vertebrae (17 to 20), and also sharing the average of 18 vertebrae per embryo (n=23). The expected result was to have further loss of anterior structures for a shorter anterior body. The expected results for these embryos would have been that they would have looked like the FCE/ FCE in their entire body shape. However, this finding suggests that dosage of Lfng in the patterning region influences development of the secondary body structures, as well as the primary body structures. Like the FCE/ FCE mice, anterior vertebrae of the FCE/- mice were always deformed. Our last genotype examined were Lfng -/-, those embryos lacking all Lfng. As expected, the complete loss of all Lfng played a clear role in the shortening of the anterior body; vertebral counts ranged from 13 to 17 vertebrae, with a median of 16 vertebrae (n=15). In addition to the much shorter stature of the Lfng -/- embryos, all vertebrae for these were poorly formed. These data suggest to us that the formation of vertebrae is dependent upon only the clock, and region II of PSM is insufficient to rescue anterior skeleton development. The vertebral column is not the only region of the anterior body affected by the varying levels of Lfng. We noticed that the ribcage is affected and thus determined three ways to quantify these defects. First, we counted the number of rib attachments on the sternum. Next, we examined the sternum of each ribcage for the number of misaligned 14
20 rib attachments (Figure 2.2(a-d; i)). The last area of rib quantification was simply to count the rib defects; the main classes of rib defects quantified were bifurcations, trifurcations, and rib fusions (Figure 2.2(a-h; j). Our wild type embryos were simply that: they had 7 attached ribs on each side of the sternum; they lacked misalignments on the sternum; they lacked any of the rib defects expected in our other mutant mice. Our next set of mice we examined were the FCE/ FCE mutants: their defects were quite evident. We found that some ribs never formed, but this lack of formation was not necessarily symmetric. For example, we counted 3 embryos that had fewer ribs on the right side of the sternum than the left. The range for right ribs was 5 to 7 and left ribs were 6 to 7. This rib asymmetry likely lead to the number of misalignments found in these embryos (median of 1, range= 0-4). Lastly, we examined the number of actual defects of ribs; these were most prominent in the dorsal ribcage. We found that all FCE/ FCE embryos exhibit at least some sort of rib abnormality, with the most common form of defect being a bifurcation. FCE / FCE embryos displayed about 3 rib defects per animal, but ranged from 2 to 5 defects. We next analyzed FCE/- embryos for defects in the ribcage region and found similar results to those obtained from analysis of the FCE/ FCE embryos. These embryos also showed asymmetry in number of ribs on each side of the sternum with the lower number most frequently found on the right side. Thus we found the median number of ribs on the left and right to both be 7, but the difference between FCE/ FCE and FCE/- was, however, the frequency of fewer than 7 ribs was higher. The range was the same as our FCE/ FCE embryos. The number of misalignments in our sample ranged from 1 to 3 with a median of 2 misalignments per embryo. We counted the number of rib abnormalities in the FCE/- animals, with a median of 4 rib 15
21 defects per embryo, and a range of 3 to 5. Our median suggests a slight increase in the number of rib defects between the FCE/ FCE and the FCE/- embryos, but this is rather insignificant as shown by the error. The Lfng -/- embryos were expected to have the most severe phenotype because they completely lack all Lfng in both regions of the PSM. We counted ribs on each side of the sternum and found that the left and right had medians of 7 and 6, respectively. The right had a low of 4 ribs while the left had a minimum of 5 attached ribs. Additionally, we noticed more rib misalignments, an average of 3.6, along with a low of at least two misalignments. We counted defects in these embryos and found that of our sample size, we had a median of 5 rib defects per animal; coupled with this, we had a higher range than in other genotypes (4 to 6). Quantifying the posterior defects in Lfng mutant mice We next turned our attention to the posterior skeleton, where development appears to occur by a different mechanism. We want to look in these structures to compare which level of Lfng expression leads to a loss of structures and which expression changes the organization of the posterior skeleton. To begin analysis of the posterior body, we counted the number of vertebrae from the pelvic girdle to the tip of the tail. In our wild type sample, this was the only real source of variation; posterior vertebrae totaled between 25 and 28 vertebrae (Figure 2.3(a-e)), which could be explained by the idea that tail length might be a polymorphic trait, or that some of the vertebrae at the tip of the tail were unable to be seen. The tails of these animals are best described as smooth; they lack rough bends, or kinks. We then moved on to quantify the FCE/ FCE posterior body. Our finding in these embryos was quite interesting: vertebral phenotype was rescued and tail length became mostly normal, with a median length of about 26 16
22 vertebrae and a range from 18 to 30. The number of deformed vertebrae in embryos with the homozygous deletion of FCE1 was surprisingly small: with between 0 and 2 vertebral defects in the posterior vertebral column. We found that some FCE/ FCE embryos had a tail kink. We counted a tail kink as a bend in the tail where the vertebra leads to a sharp change in direction. Our examination of Lfng FCE/- embryos showed that one less copy of Lfng in the patterning region lead to the loss of tail length (median length = 22 vertebrae), as well as the increase in tail kinks (median number of kinks = 2), compared to that of FCE/ FCE embryos. We also found that the number of vertebral defects extended further down the spine into the pelvic girdle, but subsequent vertebrae were nearly wild type. Our last sample, the Lfng -/- embryos, was examined for the same parameters (posterior body length and tail kinks). We found, however, that the body was so short, that the tail lacked length, and we were unable to count tail kinks. Thus, we classified their tails as truncated (Figure 2.3(a-d;f). The tail length for these embryos was shorter than those of the other genotypes, with a median length of 14 vertebrae. Our results, at this time, appear to support our hypothesis that Lfng dosage is important in the formation of the tail. Production of an anterior specific Lfng transgene It is known that Mesp2 is expressed only in the anterior region of a somite pair, along with Lfng. Because Mesp2 is expressed solely in the anterior PSM, we believed that when the transgene is activated for transcription, we will obtain Lfng in the anterior PSM only (Figure 2.3b) (Haraguchi, 2001). We would like to examine the effect this has on development, especially in Lfng -/- embryos. We constructed a transgene containing the Mesp2 promoter followed by the Lfng coding sequence, along with IRES and Bgeo in 17
23 order to test for expression (Figure 2.3a). The pieces of our transgene were assembled by a four-piece ligation; following assembly, we had mice injected with the transgene. The most evident way to address the issue of the importance of Lfng in secondary body formation is to breed our transgenic mice with Lfng -/- animals. These would give the most evident change in phenotype, as the tail would no longer be truncated. Tails obtained from offspring from the chimeric mouse were prepared as described in materials and methods. We analyzed the DNA with PCR to identity mice that were positive for our transgene. Our PCR showed that two transgenic mice were found (Figure 2.3c). We then used these founder mice to breed and produce more transgenic positive mice. We examined the expression of the transgene by RNA whole mount in situ hybridization on embryos (n=12) 10.5 d.p.c. and expected to find localized expression in the anterior region of the somites; our results, however, do not show expression of the transgene (Figure 2.3d) 18
24 Figure 2.1 Lfng genotyping Figures A-C show the chromosomes present in our various transgenic mice. 19
25 Figure 2.2 Anterior body defects (a-d) show the defects found on the ventral ribcage of these mice. The development of misalignments in the rib attachments is evident in those mice lacking oscillatory Lfng. (e-h) show dorsal rib defects. Notice the condensed cervical vertebrae in (f) compared to (e). Also, bifurcations and other rib defects are marked by arrows. Additionally, the vertebral disorder is marked in a red box, as an example. The numbers of attached ribs are shown in (i), while the total number of rib defects are shown in (j). (k) shows a comparison of the number of vertebrae to the number of defective vertebrae by genotype. Notice the progressive shortening of the anterior body. 20
26 Figure 2.3 Posterior body defects (a-d) are posterior skeletons of 18.5 d.p.c embryos. Notice the change in phenotype at the pelvic girdle. Also, (b) shows kinks in the tail of the FCE/ FCE mouse, which continue into the FCE/-. (d) shows the shorter, truncated tail evident in Lfng -/-, as well as the lack of change in disorder throughout the posterior body. A comparison of posterior body length and posterior body defects is shown in figure (e), with the FCE/ FCE being almost completely wild type. In (f), we counted the tail kinks and displayed them as a percent based on genotype. 21
27 Figure 2.4 Anterior Specific transgene The structure of our transgene is shown in (a), with the Mesp2 promoter linked to the Lfng gene. We included an IRES and Bgeo (LacZ + Neo) for verifying expression. (b) is the proposed effect of adding the transgene to a Lfng -/- mouse. Our initial founders yielded two positive transgenic mice (labeled). Embryos aged to 10.5 d.p.c. were examined by whole-mount in situ hybridization but we found not detectable expression in the anterior region of the somites. 22
28 Table 2.1 Medians and Means of quantification The above tables show the medians and means for the quantification of skeletal defects among the various genotypes. 23
29 Chapter 3 Discussion Clock function in somitogenesis is required for proper anterior skeletal development Our results indicate that a progressive decrease in Lfng leads to a shorter upper body. Removal of FCE1 leads to loss of oscillatory Lfng expression and produces defects in the anterior body. The anterior skeleton of the FCE/ FCE mice shows great disorder; FCE/ FCE mice lack all oscillatory Lfng. This suggests that the oscillatory expression of Lfng is required for proper body formation. However, this expression may not be sufficient to produce wild type appearance. This is because an examination of FCE/- animals displays a slightly more severe phenotypes in the anterior body (more rib defects and misalignments). Some other factor likely plays a role in this process though, because Lfng -/- produce somitic derivatives. The various phenotypes associated with our Lfng mutant mice suggest to us that the clock function, or region I expression of Lfng, is required for proper development of the anterior skeleton. Patterning of presomites in region II is required for proper tail development Upon examination of FCE/ FCE mice, we found an interesting phenomenon; formation of vertebrae at the lumbo-sacral junction is rescued and almost all subsequent vertebrae are wild type in appearance. Looking at the tail length, we found that it was essentially wild-type. Although we occasionally found tail kinks in these embryos, the observation of essentially wild-type embryos in the posterior body in mice with only patterning Lfng suggests that proper patterning is a requirement for formation of the posterior body. To examine this further, we examined the FCE/- mice, those that further reduced dosage of Lfng in the anterior PSM. We found that the lower dose has a slight, 24
30 but noticeable effect on development of the posterior skeleton. Most notably, the rescue found in the FCE/ FCE is essentially lost. Vertebrae become much more disordered and the tail becomes kinked in FCE/- embryos. The increase in tail kinks between FCE/ FCE and FCE/- suggests that tail kinks form as a result as a loss of Lfng in the anterior PSM. This is not as common in FCE/ FCE because they have two copies of Lfng, but because the FCE/- embryos lack an entire copy of Lfng, tail kinks are more common. Along with the noticeably shorter posterior region, we also noticed an increase in tail kinks and vertebral deformities in these mutant mice. A complete lack of both Lfng elements (clock and patterning) found in Lfng -/- embryos leads to poorly formed vertebrae along the entire axial skeleton of these mutant mice. Additionally, the truncation of the tail is a finding. The fact that the tail is truncated in these animals suggests that Lfng, in one way or another, is required for development of the tail. We therefore propose that exogenous expression of Lfng in the anterior PSM would lead to a rescued phenotype in null animals so they share a phenotype with the FCE/- embryos. The Mesp2 promoter linked Lfng transgene lacks expression in the anterior PSM The transgene we developed to directly assess the role of the patterning lacked expression in the PSM. One hypothesis to explain our result is that when injected into the mice, the transgene inserted in a region of a chromosome that is epigenetically repressed; an example of this could be insertion into the heterochromatin. This theory is supported by the fact that our transgene is transmitted down generations, as verified by PCR analysis of the offspring of our founders, but lacks RNA transcripts, determined through use of RNA whole mount in situ hybridization. 25
31 Another hypothesis to explain the lack of detectable expression is that the Mesp2 promoter is simply not robust enough to produce any product, although this promoter has been used in other transgenes (Haraguchi, 2001). In addition to this possibility, the Mesp2 promoter might require some additional factors not present in the promoter sequence we used. We could test this idea by transfecting cells with a modified transgene, one that has a reporter gene following the Mesp2 promoter, and looking for expression. If we found expression under wild type conditions, the previously mentioned hypothesis would be supported; however, if no expression occurs, additional factors would likely be needed. This experiment, however, would not directly reflect the way cells work in vivo. We could take a different approach to test the hypothesis that Lfng in the anterior PSM is required for proper development of the tail. We could look for genes that are expressed only in the anterior region of a presomite. To do this, we could do a microarray analysis on cells from the anterior PSM and cells from the posterior PSM. By comparing the results, we could determine which genes are expressed only in the anterior region of a somite. We could then produce a new transgene with the new promoter, followed by the Lfng coding sequence. Our hope would be to produce the anterior specific Lfng and rescue the skeletal phenotype. Additionally, we could use other regulatory sequences, such as the Lfng enhancer (Cole, 2002) or the EphA4 enhancer (Nakajima, 2006) to examine expression only at the anterior region of a presomite. Future Directions Because Lfng mutant animals show truncated tails, it is possible that some interaction between Lfng and the VER is present. The VER is a necessary component to proper tail formation and it is therefore reasonable to believe some interaction between 26
32 Lfng and the VER exists. Since Lfng null animals show truncated tails, and the VER signals for tail elongation, it is likely that Lfng interacts with the VER by promoting signaling. In the absence of Lfng, the VER would be unable to signal for tail elongation, thus resulting in the characteristic truncated tails of Lfng null mice. Since the anterior VER seems to be unable to influence tail outgrowth, it is expected that Lfng has no impact on this region, yet some interaction with either the middle or the posterior VER. While Hes7 s most important activity is located in the posterior PSM, the Mesp family of genes exclusively functions in the anterior PSM. Genes in this particular family serve as transcription factors; the two genes of interest in the anterior PSM are Mesp1 and Mesp2. Expression of Mesp2 is limited only to the rostral region of the maturing somite. Studies on the effect of Mesp2 on somitogenesis have been conducted by analyzing null mice. Mice lacking Mesp2 are unable to properly pattern somites; they lack rostralization. Its counterpart, Mesp1, seems to lack comparable importance in somitogenesis. Expression of Mesp1 and Mesp2 overlaps in the PSM; this suggests that redundancy in function might be present (Takahashi, 2005). Chimeric analysis of Mesp1 and Mesp2 double null animals is necessary because Mesp1/Mesp2 do not develop to term. The chimeric analysis of the double knockouts showed the inability of cells lacking both Mesp1 and Mesp2 to epithelialize. Mesp2 null animals appear to show at least some epithelialization which therefore suggests that Mesp1 is able to compensate for the lack of Mesp2 (Takahashi, 2005). It has also been shown that varying levels of Mesp1 and Mesp2 seems to impact the ability of the axial skeleton to develop (Morimoto, 2006). Mice lacking Mesp2 at a later developmental stage display fused pedicles, disordered vertebrae, and appear to lack rostral properties. Mesp2 plays roles in establishing the 27
33 rostral properties of each somite pair formed (Takahashi, 2003; Takahashi, 2007). Mice lacking Mesp2 only lack an aspect of patterning in somitogenesis, yet share a comparable phenotype to those lacking Hes7, a cell-allocation gene. To examine the role played by region II in anterior body development, a transgene expressing the anterior-specific, complementary Lfng coding region would be produced (Figure 3.1). The formation of the anterior-specific, antisense transcript would interfere with the wild-type expression of Lfng by producing a double stranded RNA molecule that would be destroyed by cellular mechanisms. This transgene would therefore remove only Lfng in the anterior PSM, but would allow for the expression of the oscillatory Lfng found in the posterior PSM. If our hypothesis is correct that only the oscillatory Lfng expression is required for proper development of the anterior body, it would be plausible to see a nearly wild-type phenotype. However, our data suggests that the loss of region II Lfng could lead to rostral-caudal patterning issues, but examination of these mice would answer the question of the requirements for somitogenesis in the anterior and the posterior PSM. We would expect to see tail truncations in transgenic mice due to the lack of Lfng in the patterning region. Additionally, defects would be present in the anterior skeleton due to the loss of patterning. We would find tail truncations because the mrna formed by our transgene would be able to bind, and thus sequester, any Lfng produced; the anti-sense Lfng would behave as a trans acting regulatory sequence. Tail truncations would be present because our transgene would make region II behave as if it is Lfng -/-, thus leading to a truncated tail. 28
34 Conclusions Through careful quantification of skeletons, we have determined that the dose of Lfng present in the PSM impacts the development of the axial skeleton. We also have found that the region I and region II impact development of different structures: region I expression promotes development of the anterior skeleton, while region II expression promotes development of the posterior skeleton. Additionally, we found that the amount of Lfng in the anterior PSM impacts the quality of the tail by comparing the phenotypes of FCE / FCE embryos to that of FCE /- embryos. We hope to gain further knowledge of the process of somitogenesis by knocking out only the R/C patterning region while maintaining oscillatory expression of Lfng. This experiment would show the true role of the clock. 29
35 Figure 3.1 Transgene knocking out region II The structure of the transgene mimics that of our earlier transgenic experiments (a). The likely effect of this transgene is shown in (b). We believe if would result in the loss of region II Lfng, but not oscillatory. 30
36 Chapter 4 Materials and Methods Genotyping of mice: Agarose gel electrophoresis was used to analyze all genotyping reactions. Tail DNA A salt-out technique was used to isolate tail DNA for all PCR reactions. Approximately 1 cm of tissue is removed from the specimen and placed in a 6 μl Proteinase K: 600 μl TENS overnight at 55 C overnight. The next day, saturated NaCl( μl) is added and mixed for 15 seconds. The mixture is then placed in a room temperature centrifuge at maximum speed for ten minutes. The resulting supernatant is then decanted into a new, sterile, 1.5-mL test tube. 600 μl of 95% ethanol is added and centrifuged as before. The supernatant is then removed and 70% ethanol is added in its place. The resulting solution is rocked at room temperature for no less than one hour. To concentrate the DNA, the tube is centrifuged for five minutes at maximum speed. The supernatant is removed and the sample is left to dry. Once the remaining ethanol has evaporated, 100 μl Tris (10 mmol, ph 7.5) is added. Samples are then stored at -20 C until needed for PCR. Yolk Sac DNA Yolk sac DNA comes from the membrane surrounding an embryo ranging from d.p.c. in age. A small amount is removed in order to determine the genotype of the embryo. It is placed in a 1.5-mL eppendorph test tube and spun down to concentrate the tissue. Next, a fresh 0.05M NaOH solution (100μl), prepared from 10N NaOH, is added and the test tube is placed on a heating block set to 95 C for thirty minutes. Next, 1/3 M Tris (ph 7.5) is added and the solution is mixed and 31
37 centrifuged. Finally, 90 μl of the upper region of the solution is placed in a separate, clean test tube and kept at -20 C until needed for further usage. PCR Genotyping reactions were done as follows: Lfng was genotyped using primers FNG322, FNG325, and PGK3 (Table 4.1). The FCE deletion requires SC284 and SC285 (Table 4.1). Reactions are run with 35 cycles with an annealing temperature of 55 C. Lfng null animals show a band at 200 bp while wild types have a band at 170 bp. FCE mutants displayed a negative band at 182 bp and a wild type band at 340 bps. For examination of transgenic mice, we used SC340 and SC 341, along with FNG 237 and FNG 239 as positive controls; transgenic animals display bands at about 250 bps. Skeleton Preparations: Mice aged at approximately 18.5 DPC are acquired from the mother by Cesarean section. After obtaining genotype information as above for Lfng as well as ΔFCE deletions, embryos are skinned and eviscerated and placed in 100% ethanol for four days. The embryos are removed from the 100% ethanol and then placed in acetone. The acetone is used to remove any remaining fat. After three days in acetone, embryos are rinsed with water then placed in the staining solution containing 2 volumes Alcian blue, 1 volume Alizarin red, 8 volumes Glacial acetic acid and 50 volumes 70% ethanol. Following staining, the embryos are transferred to a 1% KOH in 20% glycerol solution and placed in a 37 C water bath overnight. To prevent destruction of the specimens, they are then removed and remain in the 1% KOH/20% glycerol until the specimens are completely cleared. Following the clearing step, they are placed in a 2:2:1 Ethanol:Glycerol:Benzyl alcohol solution for analysis. Whole Mount RNA in situ hybridization 32
38 Embryos aged from 8.5 d.p.c. to 12.5 d.p.c were obtained from the mother via Caesarian section. Embryos were stored at -20 C in 100% methanol until needed; they are then rehydrated to 100% PBT. Bleaching of embryos using 6% hydrogen peroxide in PBT was the first step, followed by treatment with Proteinase K for fifteen minutes. Following Proteinase K treatment, 2μg/mL glycine in PBT was added. Next, embryos were fixed using 0.2% gluteraldehyde in 4% PFA in PBT for thirty minutes. Embryos were then incubated in Prehyb for one hour at 70 C with rocking. A combination of Prehyb and Dig-labeled probe (1μg/mL) was mixed and heated to 70 C. Following the Prehyb wash, the Prehyb and Dig-labeled probe were added to the embryos and incubated overnight at 70 C. Washes consisting of formamide and SCC (ph 4.5) were made and were used to wash the embryos for several hours. A series of washes using MABT were then prepared and the embryos were washed with these for several hours more. The solutions, in order, were MABT, MABT + Boehringer Blocking Reagent (BBR), MABT + BBR + 20% heat inactivated sheep serum, and MABT + BBR+ 20% heat inactivated sheep serum + anti- DIG antibody. The embryos remained in the final solution overnight at 4 C. The following day involved washing the embryos with a solution of MABT + 2 mm Levamoisole, first changing the wash every hour for five hours, and then switching the wash every twelve hours. The last day consisted of washing embryos in a solution of freshly prepared NTMT + 2mM Levamisole, then exposed to the same solution only with the addition of NBT and BCIP in the absence of light until staining became apparent. Once the desired staining became apparent, embryos were washed in PBT ph 5.5 and fixed in 4% 33
39 PFA/0.1% gluteraldehyde. Clearing of embryos was done by washing first with 1:1 Glycerol:PBT then followed by 4:1 Glycerol:PBT. Stained embryos were stored in this last solution at 4 C. 34
40 FNG GAGCACCAGGAGACAAGCC-3 FNG AGAGTTCCTGAAGCGAGAG-3 PGK3 5 -CTTGTGTAGCGCCAAGTGC-3 SC TTTGGTGGGAATGGATTAGC-3 SC CTGGTCCATTTGCTCTGAGG-3 SC CAGAATCCACACCTCTGCAA-3 SC ACCAGGAGACAAGCCAACAG-3 FNG CGACATTTTGCAGCACAG-3 FNG TTCACCGATGGAGACGAC-3 SC GCGACATGCTGGCTCTTCTA-3 SC CCAGGTTTTGACACTAGCACA-3 Table PCR Primers - List of PCR Primers and corresponding sequences used for genotyping and isolation of coding regions 35
41 Works Cited Andrade, R., Pascoal, S., and Palmeirim, I Thinking clockwise. Brain Research Reviews 49: Chen, J., Kang, L., and Zhang, N Negative Feedback Loop Formed by Lunatic Fringe and Hes7 Controls their Oscillatory Expression During Somitogenesis. Genesis 43, Cole, Susan E., John M. Levorse, Shirley M. Tilghman, and Thomas F. Vogt Clock Regulatory Elements Control Cyclic Expression of Lunatic fringe during Somitogenesis. Developmental Cell 3: Dale, J., Maroto, M., Dequeant, M., Malapert, P., McGrew, M., and Pourquie, O Periodic Notch inhibition by Lunatic Fringe underlies the chick segmentation clock. Nature 421, Evrard, Y.A., et al (1998). Lunatic fringe is an essential mediator of somite segmentation and patterning. Nature 394: Goldman, D., Martin, G., Tam, P Fate and function of the ventral ectodermal ridge during mouse tail development. Development 127, Haraguchi, S., Kitajima, S., Takagi, A., Takeda, H., Inoue, T., Saga, Y Transcriptional regulation of Mesp1 and Mesp2 genes: differential usage of enhancers during development. Mechanisms of Development 108: Kageyama, R., Masamizu, Y., and Niwa, Y Oscillator Mechanism of Notch Pathway in the Segmentation Clock. Developmental Dynamics 236, Nakajima, Y., Morimoto, M., Takahashi, Y., Koseki, H., Saga, Y Identification of Epha4 enhancer required for segmental expression and the regulation by Mesp2. Development 133(13), Morimoto, M., Takahashi, Y., Saga, Y The Mesp2 transcription factor establishes segmental borders by suppressing Notch activity. Nature 435, Morimoto, M., Kiso, M., Sasaki, N., Saga, Y Cooperative Mesp activity is required for normal somitogenesis along the anterior-posterior axis. Developmental Biology 300, Palmeirim, I., Henrique, D., Ish-Horowicz, D., and Pourquie, O Avian hairy gene expression identifies a molecular clock linked to vertebrate segmentation and somitogenesis. Cell 91:
2/23/09. Regional differentiation of mesoderm. Morphological changes at early postgastrulation. Segments organize the body plan during embryogenesis
 Regional differentiation of mesoderm Axial Paraxial Intermediate Somatic Splanchnic Chick embryo Morphological changes at early postgastrulation stages Segments organize the body plan during embryogenesis
Regional differentiation of mesoderm Axial Paraxial Intermediate Somatic Splanchnic Chick embryo Morphological changes at early postgastrulation stages Segments organize the body plan during embryogenesis
The oscillation of Notch activation, but not its boundary, is required for somite border formation and rostral-caudal patterning within a somite
 Access the Development most First recent posted version epress online at http://dev.biologists.org/lookup/doi/10.1242/dev.044545 on online 24 March publication 2010 as 10.1242/dev.044545 date March 2010
Access the Development most First recent posted version epress online at http://dev.biologists.org/lookup/doi/10.1242/dev.044545 on online 24 March publication 2010 as 10.1242/dev.044545 date March 2010
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Role of Organizer Chages in Late Frog Embryos
 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Notch Is a Critical Component of the Mouse Somitogenesis Oscillator and Is Essential for the Formation of the Somites
 Notch Is a Critical Component of the Mouse Somitogenesis Oscillator and Is Essential for the Formation of the Somites Zoltan Ferjentsik 1, Shinichi Hayashi 2., J. Kim Dale 1., Yasumasa Bessho 2, An Herreman
Notch Is a Critical Component of the Mouse Somitogenesis Oscillator and Is Essential for the Formation of the Somites Zoltan Ferjentsik 1, Shinichi Hayashi 2., J. Kim Dale 1., Yasumasa Bessho 2, An Herreman
THE MAKING OF THE SOMITE: MOLECULAR EVENTS IN VERTEBRATE SEGMENTATION. Yumiko Saga* and Hiroyuki Takeda
 THE MAKING OF THE SOMITE: MOLECULAR EVENTS IN VERTEBRATE SEGMENTATION Yumiko Saga* and Hiroyuki Takeda The reiterated structures of the vertebrate axial skeleton, spinal nervous system and body muscle
THE MAKING OF THE SOMITE: MOLECULAR EVENTS IN VERTEBRATE SEGMENTATION Yumiko Saga* and Hiroyuki Takeda The reiterated structures of the vertebrate axial skeleton, spinal nervous system and body muscle
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
The negative regulation of Mesp2 by mouse Ripply2 is required to establish the rostro-caudal patterning within a somite
 RESEARCH ARTICLE 1561 Development 134, 1561-1569 (2007) doi:10.1242/dev.000836 The negative regulation of by mouse Ripply2 is required to establish the rostro-caudal patterning within a somite Mitsuru
RESEARCH ARTICLE 1561 Development 134, 1561-1569 (2007) doi:10.1242/dev.000836 The negative regulation of by mouse Ripply2 is required to establish the rostro-caudal patterning within a somite Mitsuru
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
A systems approach to biology
 A systems approach to biology SB200 Lecture 7 7 October 2008 Jeremy Gunawardena jeremy@hms.harvard.edu Recap of Lecture 6 In phage lambda, cooperativity leads to bistability and hysteresis In HIV-1, sequestration
A systems approach to biology SB200 Lecture 7 7 October 2008 Jeremy Gunawardena jeremy@hms.harvard.edu Recap of Lecture 6 In phage lambda, cooperativity leads to bistability and hysteresis In HIV-1, sequestration
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
Biology 218, practise Exam 2, 2011
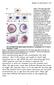 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Three different fusions led to three basic ideas: 1) If one fuses a cell in mitosis with a cell in any other stage of the cell cycle, the chromosomes
 Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
SIGNIFICANCE OF EMBRYOLOGY
 This lecture will discuss the following topics : Definition of Embryology Significance of Embryology Old and New Frontiers Introduction to Molecular Regulation and Signaling Descriptive terms in Embryology
This lecture will discuss the following topics : Definition of Embryology Significance of Embryology Old and New Frontiers Introduction to Molecular Regulation and Signaling Descriptive terms in Embryology
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
UNIVERSITY OF YORK BIOLOGY. Developmental Biology
 Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
Examination Candidate Number: UNIVERSITY OF YORK BSc Stage 2 Degree Examinations 2017-18 Department: BIOLOGY Title of Exam: Developmental Biology Desk Number: Time allowed: 1 hour and 30 minutes Total
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
Segment boundary formation in Drosophila embryos
 Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Segment boundary formation in Drosophila embryos Development 130, August 2003 Camilla W. Larsen, Elizabeth Hirst, Cyrille Alexandre and Jean Paul Vincent 1. Introduction: - Segment boundary formation:
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
ASpatio-TemporalModelofNotchSignallinginthe Zebrafish Segmentation Clock: Conditions for Synchronised Oscillatory Dynamics
 ASpatio-TemporalModelofNotchSignallinginthe Zebrafish Segmentation Clock: Conditions for Synchronised Oscillatory Dynamics Alan J. Terry 1 *, Marc Sturrock 1, J. Kim Dale 2, Miguel Maroto 2, Mark A. J.
ASpatio-TemporalModelofNotchSignallinginthe Zebrafish Segmentation Clock: Conditions for Synchronised Oscillatory Dynamics Alan J. Terry 1 *, Marc Sturrock 1, J. Kim Dale 2, Miguel Maroto 2, Mark A. J.
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Specification of vertebral identity is coupled to Notch signalling and the segmentation clock
 Research article 1221 Specification of vertebral identity is coupled to Notch signalling and the segmentation clock Ralf Cordes, Karin Schuster-Gossler, Katrin Serth and Achim Gossler* Institut für Molekularbiologie
Research article 1221 Specification of vertebral identity is coupled to Notch signalling and the segmentation clock Ralf Cordes, Karin Schuster-Gossler, Katrin Serth and Achim Gossler* Institut für Molekularbiologie
3/8/ Complex adaptations. 2. often a novel trait
 Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Chapter 10 Adaptation: from genes to traits p. 302 10.1 Cascades of Genes (p. 304) 1. Complex adaptations A. Coexpressed traits selected for a common function, 2. often a novel trait A. not inherited from
Q2 (4.6) Put the following in order from biggest to smallest: Gene DNA Cell Chromosome Nucleus. Q8 (Biology) (4.6)
 Q1 (4.6) What is variation? Q2 (4.6) Put the following in order from biggest to smallest: Gene DNA Cell Chromosome Nucleus Q3 (4.6) What are genes? Q4 (4.6) What sort of reproduction produces genetically
Q1 (4.6) What is variation? Q2 (4.6) Put the following in order from biggest to smallest: Gene DNA Cell Chromosome Nucleus Q3 (4.6) What are genes? Q4 (4.6) What sort of reproduction produces genetically
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
BIOLOGY 111. CHAPTER 5: Chromosomes and Inheritance
 BIOLOGY 111 CHAPTER 5: Chromosomes and Inheritance Chromosomes and Inheritance Learning Outcomes 5.1 Differentiate between sexual and asexual reproduction in terms of the genetic variation of the offspring.
BIOLOGY 111 CHAPTER 5: Chromosomes and Inheritance Chromosomes and Inheritance Learning Outcomes 5.1 Differentiate between sexual and asexual reproduction in terms of the genetic variation of the offspring.
Variation of Traits. genetic variation: the measure of the differences among individuals within a population
 Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
Genetic variability is the measure of the differences among individuals within a population. Because some traits are more suited to certain environments, creating particular niches and fits, we know that
1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms.
 Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Unit 4 Evaluation Question 1:
 Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Name: Unit 4 Evaluation Question 1: /7 points A naturally occurring dominant mutant in mice is the Doublefoot (Dbf) mutant. Below is an image of the bones from a wildtype (wt) and Doublefoot mutant mouse.
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
Drosophila Life Cycle
 Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Drosophila Life Cycle 1 Early Drosophila Cleavage Nuclei migrate to periphery after 10 nuclear divisions. Cellularization occurs when plasma membrane folds in to divide nuclei into cells. Drosophila Superficial
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Somites without a clock
 Dias, AS; de Almeida, I; Belmonte, JM; Glazier, JA; Stern, CD; (2014) Somites Without a Clock. Science 10.1126/science.1247575. ARTICLE Somites without a clock Ana S. Dias 1,, Irene de Almeida 1,, Julio
Dias, AS; de Almeida, I; Belmonte, JM; Glazier, JA; Stern, CD; (2014) Somites Without a Clock. Science 10.1126/science.1247575. ARTICLE Somites without a clock Ana S. Dias 1,, Irene de Almeida 1,, Julio
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Avian hairy Gene Expression Iden4fies a Molecular Clock Linked to Vertebrate Segmenta4on and Somitogenesis
 Avian hairy Gene Expression Iden4fies a Molecular Clock Linked to Vertebrate Segmenta4on and Somitogenesis Pourquie, et. al., 1997 Swetha Mummini, Victoria Fong What is a somite? Defini4on: Segmental block
Avian hairy Gene Expression Iden4fies a Molecular Clock Linked to Vertebrate Segmenta4on and Somitogenesis Pourquie, et. al., 1997 Swetha Mummini, Victoria Fong What is a somite? Defini4on: Segmental block
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11804 a Tailbud after cutting PSM after cutting b 3500 3000 2500 mean intensity 2000 1500 1000 ROI1 (TB) ROI2 (PSM) 500 0 0 1 2 3 4 5 6 7 8 9 time (h) Supplementary Fig.1 Lfng gene activity
doi:10.1038/nature11804 a Tailbud after cutting PSM after cutting b 3500 3000 2500 mean intensity 2000 1500 1000 ROI1 (TB) ROI2 (PSM) 500 0 0 1 2 3 4 5 6 7 8 9 time (h) Supplementary Fig.1 Lfng gene activity
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
The period of the somite segmentation clock is sensitive to Notch activity. Institute of Science and Technology, Ikoma, Nara , Japan
 The period of the somite segmentation clock is sensitive to Notch activity 1,7 Woong Kim, 1,7 Takaaki Matsui, 2 Masataka Yamao, 3 Makoto Ishibashi, 4 Kota Tamada, 4 Toru Takumi, 5 Kenji Kohno, 6 Shigeyuki
The period of the somite segmentation clock is sensitive to Notch activity 1,7 Woong Kim, 1,7 Takaaki Matsui, 2 Masataka Yamao, 3 Makoto Ishibashi, 4 Kota Tamada, 4 Toru Takumi, 5 Kenji Kohno, 6 Shigeyuki
The repression of Notch signaling occurs via the destabilization of mastermind-like 1 by Mesp2 and is essential for somitogenesis
 RESEARCH ARTICLE 55 Development 138, 55-64 (2011) doi:10.1242/dev.055533 2011. Published by The Company of Biologists Ltd The repression of Notch signaling occurs via the destabilization of mastermind-like
RESEARCH ARTICLE 55 Development 138, 55-64 (2011) doi:10.1242/dev.055533 2011. Published by The Company of Biologists Ltd The repression of Notch signaling occurs via the destabilization of mastermind-like
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Mesoderm Development
 Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Quiz rules: Spread out across available tables No phones, text books, or (lecture) notes on your desks No consultation with your colleagues No websites open other than the Quiz page No screen snap shots
Developmental processes Differential gene expression Introduction to determination The model organisms used to study developmental processes
 Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Date Title Topic(s) Learning Outcomes: Sept 28 Oct 3 1. What is developmental biology and why should we care? 2. What is so special about stem cells and gametes? Developmental processes Differential gene
Chromosome Chr Duplica Duplic t a ion Pixley
 Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Revisiting the involvement of signaling gradients in somitogenesis
 VIEWPOINT Revisiting the involvement of signaling gradients in somitogenesis Moises Mallo Instituto Gulbenkian de Ciencia, Oeiras, Portugal Keywords development; mesoderm; segmentation; signaling; somite
VIEWPOINT Revisiting the involvement of signaling gradients in somitogenesis Moises Mallo Instituto Gulbenkian de Ciencia, Oeiras, Portugal Keywords development; mesoderm; segmentation; signaling; somite
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Patterning embryos with oscillations: structure, function and dynamics of the vertebrate segmentation clock
 REVIEW 625 Development 139, 625-639 (2012) doi:10.1242/dev.063735 2012. Published by The Company of Biologists Ltd Patterning embryos with oscillations: structure, function and dynamics of the vertebrate
REVIEW 625 Development 139, 625-639 (2012) doi:10.1242/dev.063735 2012. Published by The Company of Biologists Ltd Patterning embryos with oscillations: structure, function and dynamics of the vertebrate
Initiation of translation in eukaryotic cells:connecting the head and tail
 Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Initiation of translation in eukaryotic cells:connecting the head and tail GCCRCCAUGG 1: Multiple initiation factors with distinct biochemical roles (linking, tethering, recruiting, and scanning) 2: 5
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Chapter 10 Development and Differentiation
 Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read
 Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read chapter 22 and chapter 10 [section on MATing type gene
Lecture 2: Read about the yeast MAT locus in Molecular Biology of the Gene. Watson et al. Chapter 10. Plus section on yeast as a model system Read chapter 22 and chapter 10 [section on MATing type gene
Homeotic Genes and Body Patterns
 Homeotic Genes and Body Patterns Every organism has a unique body pattern. Although specialized body structures, such as arms and legs, may be similar in makeup (both are made of muscle and bone), their
Homeotic Genes and Body Patterns Every organism has a unique body pattern. Although specialized body structures, such as arms and legs, may be similar in makeup (both are made of muscle and bone), their
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Biology 4361 Paraxial and Intermediate Mesoderm July 28, 2008 Paraxial and Intermediate Mesoderm Overview Development of major mesodermal lineages Somites: formation specification and differentiation Mesodermal
Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension
 Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension Nigel C. Harrison 1, Ruth Diez del Corral 2 *, Bakhtier Vasiev 1 * 1 Department of Mathematical
Coordination of Cell Differentiation and Migration in Mathematical Models of Caudal Embryonic Axis Extension Nigel C. Harrison 1, Ruth Diez del Corral 2 *, Bakhtier Vasiev 1 * 1 Department of Mathematical
Developmental Biology
 Developmental Biology 371 (2012) 235 245 Contents lists available at SciVerse ScienceDirect Developmental Biology journal homepage: www.elsevier.com/locate/developmentalbiology Signaling by FGF4 and FGF8
Developmental Biology 371 (2012) 235 245 Contents lists available at SciVerse ScienceDirect Developmental Biology journal homepage: www.elsevier.com/locate/developmentalbiology Signaling by FGF4 and FGF8
Biology I Level - 2nd Semester Final Review
 Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The role of FGF2 in craniofacial skeletogenesis
 The role of FGF2 in craniofacial skeletogenesis P. Ferretti, S. Sarkar, R. Moore, A. Petiot, C. J. Chan and A. Copp Summary E vidence that the major craniosynostosis syndromes are caused by mutations in
The role of FGF2 in craniofacial skeletogenesis P. Ferretti, S. Sarkar, R. Moore, A. Petiot, C. J. Chan and A. Copp Summary E vidence that the major craniosynostosis syndromes are caused by mutations in
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Jeopardy. Evolution Q $100 Q $100 Q $100 Q $100 Q $100 Q $200 Q $200 Q $200 Q $200 Q $200 Q $300 Q $300 Q $300 Q $300 Q $300
 Jeopardy Mutations Crosses & Punnett Sqs. Meiosis & Variability Evolution Photo, Cell Resp, Energy, Matter Q $100 Q $200 Q $300 Q $400 Q $500 Q $100 Q $100 Q $100 Q $100 Q $200 Q $200 Q $200 Q $200 Q $300
Jeopardy Mutations Crosses & Punnett Sqs. Meiosis & Variability Evolution Photo, Cell Resp, Energy, Matter Q $100 Q $200 Q $300 Q $400 Q $500 Q $100 Q $100 Q $100 Q $100 Q $200 Q $200 Q $200 Q $200 Q $300
The Making of the Fittest: Evolving Switches, Evolving Bodies
 INTRODUCTION MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH The types and amounts of proteins produced by a given cell in the body are very important and carefully regulated. Transcribing
INTRODUCTION MODELING THE REGULATORY SWITCHES OF THE PITX1 GENE IN STICKLEBACK FISH The types and amounts of proteins produced by a given cell in the body are very important and carefully regulated. Transcribing
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
13.4 Gene Regulation and Expression
 13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
13.4 Gene Regulation and Expression Lesson Objectives Describe gene regulation in prokaryotes. Explain how most eukaryotic genes are regulated. Relate gene regulation to development in multicellular organisms.
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
7.013 Problem Set
 7.013 Problem Set 5-2013 Question 1 During a summer hike you suddenly spot a huge grizzly bear. This emergency situation triggers a fight or flight response through a signaling pathway as shown below.
7.013 Problem Set 5-2013 Question 1 During a summer hike you suddenly spot a huge grizzly bear. This emergency situation triggers a fight or flight response through a signaling pathway as shown below.
CHAPTER 1 THE STRUCTURAL BIOLOGY OF THE FGF19 SUBFAMILY
 CHAPTER 1 THE STRUCTURAL BIOLOGY OF THE FGF19 SUBFAMILY Andrew Beenken and Moosa Mohammadi* Department of Pharmacology, New York University School of Medicine, New York, New York, USA. *Corresponding Author:
CHAPTER 1 THE STRUCTURAL BIOLOGY OF THE FGF19 SUBFAMILY Andrew Beenken and Moosa Mohammadi* Department of Pharmacology, New York University School of Medicine, New York, New York, USA. *Corresponding Author:
Evaluating Zebrafish as a Model for Craniosynostosis: Normal Cranial Vault and Suture Development
 Evaluating Zebrafish as a Model for Craniosynostosis: Normal Cranial Vault and Suture Development Ramy A. Shoela, MBBCh; Joanna P. Tomaszewski, MS; Jolanta M. Topczewska, PHD; Arun K. Gosain, MD Authors
Evaluating Zebrafish as a Model for Craniosynostosis: Normal Cranial Vault and Suture Development Ramy A. Shoela, MBBCh; Joanna P. Tomaszewski, MS; Jolanta M. Topczewska, PHD; Arun K. Gosain, MD Authors
