Reconstruction and Analysis of Metabolic Networks
|
|
|
- Egbert Watkins
- 5 years ago
- Views:
Transcription
1 1 Reconstruction and Analysis of Metabolic Networks
2 2 Outline What is a Reconstruction? Data Collection Interactions Between Network Components Special Considerations Applications
3 Genome-scale Metabolic Model 3 Genome Annotation - by homology, location Biochemical Data - protein characterized Physiological Data - indirect, pathway known Inferred Reactions - indirect, inferred from biomass requirements Quantitative Analysis - simulate cell behavior - drive experimental studies Reconstruction ORGANISM Genome Annotation Biochemistry Network Reconstruction Metabolic Model Cell Physiology Inferred Reactions Quantitative Analytical Methods New Predictions Emergent Properties
4 Genome-scale Metabolic Model 4 Model Development: an iterative process Reconstruction ORGANISM - Biochemical data -Revised ORF assignments Genome Annotation Biochemistry Cell Physiology Inferred Reactions Network Reconstruction Computational, Biochemical Investigation Metabolic Model New Predictions Emergent Properties Quantitative Analytical Methods
5 What is in a reconstruction? 5 Genome: Annotated genes Gene location Regulatory regions Wobble base pairs Biochemistry: Stereochemistry ph and pka (charge) Elemental balance Charge balance Multiple reactions/enzyme Multiple enzymes/reaction Transcription/translation: Gene to transcript to protein to reaction association Transcript half-lives trna abundances Ribosomal capacities Physiology: Flux data Knock-outs Balanced functions Overall phenotypic behavior Location of gene product compartmentalization
6 Defining Metabolic Reactions 6 ydbh hslj ldha 1st level: Primary metabolites LAC 2nd level: Neutral Formulas C 3 H 6 O 3 Charged Formulas C 3 H 5 O 3 1-3rd level: 4th level: 5th level: prokaryotes Lactate Dehydrogenase Metabolite Specificity PYR Metabolite Formulas C 3 H 4 O 3 C 3 H 3 O 3 1- Stoichiometry C 21 H 26 N 7 O 14 P 2 1 LAC + 1 NAD? 1 PYR + 1 NADH + 1 H Thermodynamic Considerations: Directionality Localization eukaryotes NAD C 21 H 26 N 7 O 14 P 2 1- Coenzymes 1 LAC + 1 NAD 1 PYR + 1 NADH + 1 H NADH C 21 H 27 N 7 O 14 P 2 C 21 H 27 N 7 O 14 P 2 1- [c]: cytoplasm [n]: nucleus [m]: mitochondria [e]: extracellular [g]: golgi aparatus [x]: peroxisome [p]: periplasm [v]: vacuole [h]: chloroplast [l]: lysosome [r]: endoplasmic reticulum 1 LAC [c] + 1 NAD [c] 1 PYR [c] + 1 NADH [c] + 1 H [c] STEPWISE INCORPORATION OF INFORMATION
7 Sources of Information 7 Genome Sequence & Annotation Physiological Data Databases OD 600 Growth Measurements Time (Hours) Available Literature Phylogenetic Data Bacteria Archaea Eukarya Localization Signal sequences...pllllpisgsalp...
8 Network Assembly and Representation 8 Reconstruction of Glycolytic Pathway Abbr. HEX1 PGI PFK FBA TPI GAPD PGK PGM ENO PYK Glycolytic Reactions [c]glc +atp g6p + adp [c]g6p f6p [c]atp + f6p adp + fdp + h [c]fdp dhap + g3p [c]dhap g3p [c]g3p + nad + pi 13dpg + h + nadh [c]13dpg + adp 3pg + atp [c]3pg 2pg [c]2pg h2o + pep [c]adp + h + pep atp + pyr Genes glk pgi pfka,pfkb fbaa,fbab tpia gapa,gapc_1,gapc_2 pgk gpma,gpmb eno pyka,pykf HEX1 PGI PFK FBA TPI GAPD PGK PGM ENO PYK atp glc adp g6p h f6p fdp dhap g3p nad pi dpg nadh pg pg pep h2o pyr PYK: IF pyka OR pykf ENO: IF eno GAPD: IF gapa OR (gapc_1 AND gapc_2)
9 Network Evaluation 9 Precursor Metabolite Formation Filling Network Gaps Incorporating Biomass Composition ATP Maintenance Calculation ATP (mmol/gdw/h) ATP m ATP biomass D (1/h) Physiological Data Comparison Knockout Data Comparison qco 2 (mmol/gdw/h) Experiment Model qo 2 (mmol/gdw/h) Experiment Model D (1/h) D (1/h) P/O Ratio Calculation 1.5 H + 3 H + Outer Membrane Inner Membrane I II III IV ATPase ETS 3 H +
10 10 Data Collection I. Genome Annotation II. Biochemistry III. Physiology
11 I. Genome Annotation ,238 protein sequences derived from whole genomes (expected to reach 1 million by 2005) 101,602 entries in Swiss Prot High-quality annotation requires substantial effort
12 Genome Annotation: how to 12 Open Reading Frame (ORF) Identification - Start & Stop codons, GLIMMER. Traditional Annotation Methods - Experimental (direct) - Sequence homology - Generally covers 40-70% of new genomes New Annotation Methods - Protein-protein interactions - Correlated mrna expression levels - Phylogenetic profile clustering - Protein fusion - Gene neighbors (operon clustering)
13 Genome Databases: TIGR 13 The Comprehensive Microbial Resource (CMR) 353 completed bacterial genomes 28 completed archaeal genomes Single-genome analysis: Genome overview, list by role category (eg amino acid biosynthesis) analysis methods, searches Multi-genome analysis also available
14 Genome Databases: TIGR 14
15 Genome Databases: NCBI 15 Microbial Genomes Resources 595 completed microbial genomes (47 archael) FTP sites for Protein Annotations (ptt files)
16 II. Biochemical Data: Reactions stoichiometry and reversibility 16 Gene: glk Enzyme: Glucokinase Reaction: ATP + D-Glucose > ADP + D-Glucose 6-phosphate E.C.: Human ß-cell glk
17 Trust the E.C. Nomenclature! 17 EC 1 Oxidoreductases EC 1.1 Acting on the CH-OH group of donors EC With NAD or NADP as acceptor EC With a cytochrome as acceptor EC With oxygen as acceptor EC With a disulfide as acceptor EC With a quinone or similar compound as acceptor EC With other acceptors EC 1.2 Acting on the aldehyde or oxo group of donors EC With NAD or NADP as acceptor EC With a cytochrome as acceptor EC With oxygen as acceptor EC With a disulfide as acceptor EC With an iron-sulfur protein acceptor EC With other acceptors EC 1.3 Acting on the CH-CH group of donors EC With NAD or NADP as acceptor EC With a cytochrome as acceptor EC With oxygen as acceptor EC With a quinone or related compound as acceptor EC With an iron-sulfur protein as acceptor Not widely available for other types of gene products (T.C. numbers are being developed) Kudos to enzymologists Make sure to balance elements when writing reaction
18 KEGG 18
19 How are these reconstructions 19 Reaction Entries different than KEGG? Not elementally or charge balanced Network Maps Don t show which compounds participate in the reactions Compound Entries Don t include details on ionization state Don t show localization info
20 Charge Determination on 20 Metabolite at neutral ph Identify compound & look up in KEGG Determine and identify ionizable group Determine acid and base forms Determine pka values based on the identifiable group (in the Table) If pka > ph, acid form dominant If pka < ph, base form dominant
21 21 Ionizable Groups 1 GROUP ACID BASE + H + pka Terminal Carboxyl -COOH COO - + H + ~4 Primary (Secondary, Tertiary) Amine -NH 3 + NH 2 + H + > 9 Thiol -SH S - + H + ~8.5 O - Phenol + H + ~10
22 22 Ionizable Groups 2 GROUP ACID BASE + H + pka Primary Alcohol -CH 2 OH CH 2 O - + H + ~15 Acetamide (Amide) + H + ~0 OH + Urea (Carbamide) OH + + H + ~1 Guanido Group H -N-C-NH 2 H -N-C-NH 2 + H + ~12 NH 2 + NH
23 23 Ionizable Groups 3 GROUP ACID BASE + H + pka Pyridine H+ + H + ~5 Pyrole + H + ~ -1 H 2 + Imidazole H+ + H + ~7 Pyrimidine H+ + H + ~0
24 24 Ionizable Groups 4 GROUP ACID BASE + H + pka Pyrrolidine (Like Sec Amine) H+ + H + ~10 Aniline H+ + H + ~5 -O Benzoic Acid + H + ~4
25 Ionizable Groups 5: Purines & Pyrimidines 25 Adenine Guanine Thymine Uracil Cytosine Net Charge = 0!
26 26 Example : Arginosuccinate Example: Argininosuccinate Primary Amine pka ~ 9 Guanido pka ~ 12 Neutral MF: C10H18N4O pka: 1.62, 2.70, 4.26, 9.58, >12 Net Charge: -1 Charged MF: C10H17N4O6
27 Biochemical Data: 27 Curation and Expansion of the Network H. pylori Glycolysis according to KEGG: Glucose G-6-P F-6-P FDP H. pylori Glycolysis according to Hoffman et al. (1996): Glucose G-6-P F-6-P FDP
28 28 Organism-specific Textbooks Great starting point Broad view of the organism s metabolism, biochemistry, physiology, uses, etc.
29 29 III. Physiological Information and Inferred Reactions: Filling in the Gaps based on indirect evidence
30 Filling in the Gaps an Example 30 Experiments determine which amino acids are taken up by H. pylori vs. which can be produced in vivo Missing steps of amino acid biosynthesis are added if necessary on the basis of this physiological evidence Amino Acid Requirements AA Reynolds Model Ala - - Arg - - Asn + + Asp + + Cys + + Gln + + Glu + + Gly + + His - - Ile - - Leu - - Lys + + Met - - Phe - - Pro + + Ser + + Thr + + Trp + + Tyr + + Val - - in vivo in silico
31 31 Inferred Reactions Some reactions are included based on indirect physiological evidence (by inference) Assumption: the cell must be able to produce all biomass components to grow Reactions are added if necessary Generally transporters, etc. Most tentative; should be examined more carefully
32 Reaction Confidence: Sources of 32 Evidence Increasing Confidence Biochemical Enzyme has been tested biochemically. Genetic Gene overexpression and purification, gene deletions. Sequence There is significant sequence similarity to another gene with known function. Physiological There is physiological data to support inclusion in the model. Modeling Reaction is included to improve simulation results. Kinetic Assay Overexpression 86% Homology Grows on Ascorbate
33 33 Gene to Reaction Connections
34 Escherichia coli Metabolism 34
35 From Genes to Reactions 35 Not all genes have a one-to-one relationship with their corresponding enzymes or reactions Many genes, one reaction: frdabcd Four subunits combine to form fumarate reductase enzyme, catalyzing FUM + FADH 2 SUCC + FAD One gene, many reactions: tkta One gene encodes transketolase I enzyme, catalyzing R5P + X5P T3P1 + S7P E4P + X5P T3P1 + F6P E. coli frdabcd
36 Integrating -omics Data 36 ORF gene Genomics ORF annotation Transcriptomics mrna levels protein reactions Proteomics protein levels Fluxomics flux measurements
37 Example of Isozymes 37 fructose-1,6-bisphosphate aldolase
38 A More Complex Example 38 Pyruvate Metabolism
39 39 Enolase 1 gene 1 reaction (342/904 Genes are one-to-one) Succinate Dehydrogenase 4 subunits 2 reactions (135/904 Genes interact as Subunits) Xylose ABC Transporter Protein complex (3 proteins) 1 reaction (153/904 Genes interact as PCs) Pyruvate Kinase 2 isozymes 1 reaction (149/931 Reactions have multiple isozymes)
40 Special Considerations 40
41 Biomass Composition 41 Indicates demands of the system (more detail in modeling section of class) Precursors may also be used for smaller networks Approximation of Biomass composition for less-characterized organisms (H. pylori, H. influenzae) Metabolite NADPH G6P F6P R5P E4P GA3P 3PG PEP PYR ACCOA OXA AKG SUCCOA Demand (mmol) ATP 41.3 NAD (trace)
42 42 Generally: Nutrient uptake Growth rate
43 43 Charge Considerations An underappreciated aspect of building reaction networks electrical charge should be conserved in all reactions Phosphofructokinase (from KEGG): ATP + F6P => ADP + FDP charges = -6 = -7 = -6 + H + + 1
44 44 Finding Compound Charges Consult diagram and look at each chemical group independently Determine if H s are attached or dissociated at cellular ph (attached if pka<ph) can find pka values F6P: Both H s dissociate at cellular ph, leaving a charge of -2
45 Balancing Reactions: An 45 Solution # 1: Example pyruvate decarboxylase pyr acald + co 2 C3H3O3 C2H4O CO2 ELEMENTALLY IMBALANCED (H and O) (-1) (0) (0) CHARGE IMBALANCED Solution # 2: h 2 o + pyr acald + co 2 + ½ o 2 + h + H2O C3H3O3 C2H4O CO2 O2 H ELEMENTALLY BALANCED (0) (-1) (0) (0) (0) (+1) CHARGE IMBALANCED Solution # 3: h + + pyr acald + co 2 H C3H3O3 C2H4O CO2 ELEMENTALLY BALANCED (+1) (-1) (0) (0) CHARGE BALANCED
46 Compartmentalization 46 extracellular space cytosol endoplasmic reticulum nucleus peroxisome mitochondria vacuole Golgi apparatus There are 8 compartments included in our yeast model May need to infer transport reactions between compartments H+, ATP, NADH, NADPH must be balanced within each compartment
47 47 Protein Localization Compartmentalization is a key part of network reconstruction Both a component (static localization) and state (dynamic localization) data type Techniques typically based on GFP-tagging of proteins or isolation of specific organelles Potential problems: Effect of the GFP tag on localization Usually human assistance is required in image analysis Condition dependence: e.g. mitochondrial localization agrees only in 30% of cases in yeast between three different data sets nucleus nucl. per. ER bud neck mitochon. lip. ves Huh et al. Nature 425:686 (2003)
48 48 Mitochondrial Isolation: Protein Identification 657 distinct proteins 498 (81%) functionally classified into 15 cellular processes. 153 unique enzymatic activities Glycolysis, TCA cycle, oxidative phosphorylation, urea cycle, fatty acid oxidation, lipid and heme biosynthesis. Taylor S. et al, Nature Biotech (21), 2003
49 Metabolic Databases 49
50 50 Genome-Scale Reconstructions
Constraint-Based Workshops
 Constraint-Based Workshops 2. Reconstruction Databases November 29 th, 2007 Defining Metabolic Reactions ydbh hslj ldha 1st level: Primary metabolites LAC 2nd level: Neutral Formulas C 3 H 6 O 3 Charged
Constraint-Based Workshops 2. Reconstruction Databases November 29 th, 2007 Defining Metabolic Reactions ydbh hslj ldha 1st level: Primary metabolites LAC 2nd level: Neutral Formulas C 3 H 6 O 3 Charged
Energy and Cellular Metabolism
 1 Chapter 4 About This Chapter Energy and Cellular Metabolism 2 Energy in biological systems Chemical reactions Enzymes Metabolism Figure 4.1 Energy transfer in the environment Table 4.1 Properties of
1 Chapter 4 About This Chapter Energy and Cellular Metabolism 2 Energy in biological systems Chemical reactions Enzymes Metabolism Figure 4.1 Energy transfer in the environment Table 4.1 Properties of
13. PG3 + ATP + NADH => GA3P + ADP + Pi + NAD +
 Additional file 2. 1.1 Stochiometric model for P. pastoris containing some additional reactions from the 13 C model (section 1.2) Methanol metabolism 1. Metoh => Form 2. Form => FOR + NADH 3. FOR + NAD
Additional file 2. 1.1 Stochiometric model for P. pastoris containing some additional reactions from the 13 C model (section 1.2) Methanol metabolism 1. Metoh => Form 2. Form => FOR + NADH 3. FOR + NAD
Oxidative Phosphorylation versus. Photophosphorylation
 Photosynthesis Oxidative Phosphorylation versus Photophosphorylation Oxidative Phosphorylation Electrons from the reduced cofactors NADH and FADH 2 are passed to proteins in the respiratory chain. In eukaryotes,
Photosynthesis Oxidative Phosphorylation versus Photophosphorylation Oxidative Phosphorylation Electrons from the reduced cofactors NADH and FADH 2 are passed to proteins in the respiratory chain. In eukaryotes,
Bringing Genomes to Life: The Use of Genome-Scale In Silico Models
 2 Bringing Genomes to Life: The Use of Genome-Scale In Silico Models Ines Thiele and Bernhard Ø. Palsson Summary Metabolic network reconstruction has become an established procedure that allows the integration
2 Bringing Genomes to Life: The Use of Genome-Scale In Silico Models Ines Thiele and Bernhard Ø. Palsson Summary Metabolic network reconstruction has become an established procedure that allows the integration
Winter School in Mathematical & Computational Biology
 1 Network reconstruction, topology and feasible solution space From component to systems biology Component biology Systems biology Component view Systems view Needed homeostasis Function S+E X E+P Reaction
1 Network reconstruction, topology and feasible solution space From component to systems biology Component biology Systems biology Component view Systems view Needed homeostasis Function S+E X E+P Reaction
Metabolic modelling. Metabolic networks, reconstruction and analysis. Esa Pitkänen Computational Methods for Systems Biology 1 December 2009
 Metabolic modelling Metabolic networks, reconstruction and analysis Esa Pitkänen Computational Methods for Systems Biology 1 December 2009 Department of Computer Science, University of Helsinki Metabolic
Metabolic modelling Metabolic networks, reconstruction and analysis Esa Pitkänen Computational Methods for Systems Biology 1 December 2009 Department of Computer Science, University of Helsinki Metabolic
BBS2710 Microbial Physiology. Module 5 - Energy and Metabolism
 BBS2710 Microbial Physiology Module 5 - Energy and Metabolism Topics Energy production - an overview Fermentation Aerobic respiration Alternative approaches to respiration Photosynthesis Summary Introduction
BBS2710 Microbial Physiology Module 5 - Energy and Metabolism Topics Energy production - an overview Fermentation Aerobic respiration Alternative approaches to respiration Photosynthesis Summary Introduction
Bio102 Problems Photosynthesis
 Bio102 Problems Photosynthesis 1. Why is it advantageous for chloroplasts to have a very large (in surface area) thylakoid membrane contained within the inner membrane? A. This limits the amount of stroma
Bio102 Problems Photosynthesis 1. Why is it advantageous for chloroplasts to have a very large (in surface area) thylakoid membrane contained within the inner membrane? A. This limits the amount of stroma
Bioinformatics: Network Analysis
 Bioinformatics: Network Analysis Flux Balance Analysis and Metabolic Control Analysis COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Flux Balance Analysis (FBA) Flux balance
Bioinformatics: Network Analysis Flux Balance Analysis and Metabolic Control Analysis COMP 572 (BIOS 572 / BIOE 564) - Fall 2013 Luay Nakhleh, Rice University 1 Flux Balance Analysis (FBA) Flux balance
2.A.2- Capture and Storage of Free Energy
 2.A.2- Capture and Storage of Free Energy Big Idea 2: Biological systems utilize free energy and molecular building blocks to grow, to reproduce, and to maintain dynamic homeostasis. EU 2.A- Growth, reproduction
2.A.2- Capture and Storage of Free Energy Big Idea 2: Biological systems utilize free energy and molecular building blocks to grow, to reproduce, and to maintain dynamic homeostasis. EU 2.A- Growth, reproduction
2. In regards to the fluid mosaic model, which of the following is TRUE?
 General Biology: Exam I Sample Questions 1. How many electrons are required to fill the valence shell of a neutral atom with an atomic number of 24? a. 0 the atom is inert b. 1 c. 2 d. 4 e. 6 2. In regards
General Biology: Exam I Sample Questions 1. How many electrons are required to fill the valence shell of a neutral atom with an atomic number of 24? a. 0 the atom is inert b. 1 c. 2 d. 4 e. 6 2. In regards
Predicting rice (Oryza sativa) metabolism
 Predicting rice (Oryza sativa) metabolism Sudip Kundu Department of Biophysics, Molecular Biology & Bioinformatics, University of Calcutta, WB, India. skbmbg@caluniv.ac.in Collaborators: Mark G Poolman
Predicting rice (Oryza sativa) metabolism Sudip Kundu Department of Biophysics, Molecular Biology & Bioinformatics, University of Calcutta, WB, India. skbmbg@caluniv.ac.in Collaborators: Mark G Poolman
Lecture 10. Proton Gradient-dependent ATP Synthesis. Oxidative. Photo-Phosphorylation
 Lecture 10 Proton Gradient-dependent ATP Synthesis Oxidative Phosphorylation Photo-Phosphorylation Model of the Electron Transport Chain (ETC) Glycerol-3-P Shuttle Outer Mitochondrial Membrane G3P DHAP
Lecture 10 Proton Gradient-dependent ATP Synthesis Oxidative Phosphorylation Photo-Phosphorylation Model of the Electron Transport Chain (ETC) Glycerol-3-P Shuttle Outer Mitochondrial Membrane G3P DHAP
Systems II. Metabolic Cycles
 Systems II Metabolic Cycles Overview Metabolism is central to cellular functioning Maintenance of mass and energy requirements for the cell Breakdown of toxic compounds Consists of a number of major pathways
Systems II Metabolic Cycles Overview Metabolism is central to cellular functioning Maintenance of mass and energy requirements for the cell Breakdown of toxic compounds Consists of a number of major pathways
2. Cellular and Molecular Biology
 2. Cellular and Molecular Biology 2.1 Cell Structure 2.2 Transport Across Cell Membranes 2.3 Cellular Metabolism 2.4 DNA Replication 2.5 Cell Division 2.6 Biosynthesis 2.1 Cell Structure What is a cell?
2. Cellular and Molecular Biology 2.1 Cell Structure 2.2 Transport Across Cell Membranes 2.3 Cellular Metabolism 2.4 DNA Replication 2.5 Cell Division 2.6 Biosynthesis 2.1 Cell Structure What is a cell?
Objective: You will be able to justify the claim that organisms share many conserved core processes and features.
 Objective: You will be able to justify the claim that organisms share many conserved core processes and features. Do Now: Read Enduring Understanding B Essential knowledge: Organisms share many conserved
Objective: You will be able to justify the claim that organisms share many conserved core processes and features. Do Now: Read Enduring Understanding B Essential knowledge: Organisms share many conserved
Lecture Series 9 Cellular Pathways That Harvest Chemical Energy
 Lecture Series 9 Cellular Pathways That Harvest Chemical Energy Reading Assignments Review Chapter 3 Energy, Catalysis, & Biosynthesis Read Chapter 13 How Cells obtain Energy from Food Read Chapter 14
Lecture Series 9 Cellular Pathways That Harvest Chemical Energy Reading Assignments Review Chapter 3 Energy, Catalysis, & Biosynthesis Read Chapter 13 How Cells obtain Energy from Food Read Chapter 14
Giving you the energy you need!
 Giving you the energy you need! Use your dominant hand Open and close the pin (with your thumb and forefinger) as many times as you can for 20 seconds while holding the other fingers straight out! Repeat
Giving you the energy you need! Use your dominant hand Open and close the pin (with your thumb and forefinger) as many times as you can for 20 seconds while holding the other fingers straight out! Repeat
Metabolism. Fermentation vs. Respiration. End products of fermentations are waste products and not fully.
 Outline: Metabolism Part I: Fermentations Part II: Respiration Part III: Metabolic Diversity Learning objectives are: Learn about respiratory metabolism, ATP generation by respiration linked (oxidative)
Outline: Metabolism Part I: Fermentations Part II: Respiration Part III: Metabolic Diversity Learning objectives are: Learn about respiratory metabolism, ATP generation by respiration linked (oxidative)
Quiz answers. Allele. BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA)
 BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA) http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Quiz answers Kinase: An enzyme
BIO 5099: Molecular Biology for Computer Scientists (et al) Lecture 17: The Quiz (and back to Eukaryotic DNA) http://compbio.uchsc.edu/hunter/bio5099 Larry.Hunter@uchsc.edu Quiz answers Kinase: An enzyme
Description of the algorithm for computing elementary flux modes
 Description of the algorithm for computing elementary flux modes by Stefan Schuster, Thomas Dandekar,, and David Fell Department of Bioinformatics, Max Delbrück Centre for Molecular Medicine D-9 Berlin-Buch,
Description of the algorithm for computing elementary flux modes by Stefan Schuster, Thomas Dandekar,, and David Fell Department of Bioinformatics, Max Delbrück Centre for Molecular Medicine D-9 Berlin-Buch,
Principles of Cellular Biology
 Principles of Cellular Biology آشنایی با مبانی اولیه سلول Biologists are interested in objects ranging in size from small molecules to the tallest trees: Cell Basic building blocks of life Understanding
Principles of Cellular Biology آشنایی با مبانی اولیه سلول Biologists are interested in objects ranging in size from small molecules to the tallest trees: Cell Basic building blocks of life Understanding
V19 Metabolic Networks - Overview
 V19 Metabolic Networks - Overview There exist different levels of computational methods for describing metabolic networks: - stoichiometry/kinetics of classical biochemical pathways (glycolysis, TCA cycle,...
V19 Metabolic Networks - Overview There exist different levels of computational methods for describing metabolic networks: - stoichiometry/kinetics of classical biochemical pathways (glycolysis, TCA cycle,...
CHEM 3653 Exam # 1 (03/07/13)
 1. Using phylogeny all living organisms can be divided into the following domains: A. Bacteria, Eukarya, and Vertebrate B. Archaea and Eukarya C. Bacteria, Eukarya, and Archaea D. Eukarya and Bacteria
1. Using phylogeny all living organisms can be divided into the following domains: A. Bacteria, Eukarya, and Vertebrate B. Archaea and Eukarya C. Bacteria, Eukarya, and Archaea D. Eukarya and Bacteria
Stoichiometric analysis of metabolic networks
 1 SGN-6156 Computational systems biology II Antti Larjo antti.larjo@tut.fi Computational Systems Biology group 30.4.2008 30.4.2008 2 Contents Networks in cells Metabolism Metabolic networks and pathways
1 SGN-6156 Computational systems biology II Antti Larjo antti.larjo@tut.fi Computational Systems Biology group 30.4.2008 30.4.2008 2 Contents Networks in cells Metabolism Metabolic networks and pathways
TCA Cycle. Voet Biochemistry 3e John Wiley & Sons, Inc.
 TCA Cycle Voet Biochemistry 3e Voet Biochemistry 3e The Electron Transport System (ETS) and Oxidative Phosphorylation (OxPhos) We have seen that glycolysis, the linking step, and TCA generate a large number
TCA Cycle Voet Biochemistry 3e Voet Biochemistry 3e The Electron Transport System (ETS) and Oxidative Phosphorylation (OxPhos) We have seen that glycolysis, the linking step, and TCA generate a large number
Chapter 7: Metabolic Networks
 Chapter 7: Metabolic Networks 7.1 Introduction Prof. Yechiam Yemini (YY) Computer Science epartment Columbia University Introduction Metabolic flux analysis Applications Overview 2 1 Introduction 3 Metabolism:
Chapter 7: Metabolic Networks 7.1 Introduction Prof. Yechiam Yemini (YY) Computer Science epartment Columbia University Introduction Metabolic flux analysis Applications Overview 2 1 Introduction 3 Metabolism:
2015 AP Biology PRETEST Unit 3: Cellular Energetics Week of October
 Name: Class: _ Date: _ 2015 AP Biology PRETEST Unit 3: Cellular Energetics Week of 19-23 October Multiple Choice Identify the choice that best completes the statement or answers the question. 1) Which
Name: Class: _ Date: _ 2015 AP Biology PRETEST Unit 3: Cellular Energetics Week of 19-23 October Multiple Choice Identify the choice that best completes the statement or answers the question. 1) Which
Chapter 4: Amino Acids
 Chapter 4: Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. lipid polysaccharide enzyme 1940s 1980s. Lipids membrane 1960s. Polysaccharide Are energy metabolites and many of
Chapter 4: Amino Acids All peptides and polypeptides are polymers of alpha-amino acids. lipid polysaccharide enzyme 1940s 1980s. Lipids membrane 1960s. Polysaccharide Are energy metabolites and many of
Advanced Topics in RNA and DNA. DNA Microarrays Aptamers
 Quiz 1 Advanced Topics in RNA and DNA DNA Microarrays Aptamers 2 Quantifying mrna levels to asses protein expression 3 The DNA Microarray Experiment 4 Application of DNA Microarrays 5 Some applications
Quiz 1 Advanced Topics in RNA and DNA DNA Microarrays Aptamers 2 Quantifying mrna levels to asses protein expression 3 The DNA Microarray Experiment 4 Application of DNA Microarrays 5 Some applications
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 AP Exam Chapters 9 and 10; Photosynthesis and Respiration Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Carbon dioxide (CO2) is released
AP Exam Chapters 9 and 10; Photosynthesis and Respiration Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Carbon dioxide (CO2) is released
Translation. A ribosome, mrna, and trna.
 Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
Translation The basic processes of translation are conserved among prokaryotes and eukaryotes. Prokaryotic Translation A ribosome, mrna, and trna. In the initiation of translation in prokaryotes, the Shine-Dalgarno
V14 extreme pathways
 V14 extreme pathways A torch is directed at an open door and shines into a dark room... What area is lighted? Instead of marking all lighted points individually, it would be sufficient to characterize
V14 extreme pathways A torch is directed at an open door and shines into a dark room... What area is lighted? Instead of marking all lighted points individually, it would be sufficient to characterize
Cellular Respiration: Harvesting Chemical Energy. 9.1 Catabolic pathways yield energy by oxidizing organic fuels
 Cellular Respiration: Harvesting Chemical Energy 9.1 Catabolic pathways yield energy by oxidizing organic fuels 9.2 Glycolysis harvests chemical energy by oxidizing glucose to pyruvate 9.3 The citric acid
Cellular Respiration: Harvesting Chemical Energy 9.1 Catabolic pathways yield energy by oxidizing organic fuels 9.2 Glycolysis harvests chemical energy by oxidizing glucose to pyruvate 9.3 The citric acid
C. Incorrect! Catalysts themselves are not altered or consumed during the reaction.
 Human Physiology - Problem Drill 04: Enzymes and Energy Question No. 1 of 10 Instructions: (1) Read the problem and answer choices carefully, (2) Work the problems on paper as needed, (3) Pick the answer,
Human Physiology - Problem Drill 04: Enzymes and Energy Question No. 1 of 10 Instructions: (1) Read the problem and answer choices carefully, (2) Work the problems on paper as needed, (3) Pick the answer,
Introduction to the Ribosome Overview of protein synthesis on the ribosome Prof. Anders Liljas
 Introduction to the Ribosome Molecular Biophysics Lund University 1 A B C D E F G H I J Genome Protein aa1 aa2 aa3 aa4 aa5 aa6 aa7 aa10 aa9 aa8 aa11 aa12 aa13 a a 14 How is a polypeptide synthesized? 2
Introduction to the Ribosome Molecular Biophysics Lund University 1 A B C D E F G H I J Genome Protein aa1 aa2 aa3 aa4 aa5 aa6 aa7 aa10 aa9 aa8 aa11 aa12 aa13 a a 14 How is a polypeptide synthesized? 2
BIOCHEMISTRY. František Vácha. JKU, Linz.
 BIOCHEMISTRY František Vácha http://www.prf.jcu.cz/~vacha/ JKU, Linz Recommended reading: D.L. Nelson, M.M. Cox Lehninger Principles of Biochemistry D.J. Voet, J.G. Voet, C.W. Pratt Principles of Biochemistry
BIOCHEMISTRY František Vácha http://www.prf.jcu.cz/~vacha/ JKU, Linz Recommended reading: D.L. Nelson, M.M. Cox Lehninger Principles of Biochemistry D.J. Voet, J.G. Voet, C.W. Pratt Principles of Biochemistry
Table 1. ODEs used in the model based on mass balances.
 Table. ODEs used in the model based on mass balances. ODE s _ = _ 6 = + _ 6 = + + = +! 3 = +2!!$ = +!$ %&' ( = _ + %&' ) *+ = + _, - +2 ),$.(/0 = +,!%!-.(023 = + - $!. 56..(023_ = +!. 56. 2,3 8.3(20 =
Table. ODEs used in the model based on mass balances. ODE s _ = _ 6 = + _ 6 = + + = +! 3 = +2!!$ = +!$ %&' ( = _ + %&' ) *+ = + _, - +2 ),$.(/0 = +,!%!-.(023 = + - $!. 56..(023_ = +!. 56. 2,3 8.3(20 =
Biochemistry: A Review and Introduction
 Biochemistry: A Review and Introduction CHAPTER 1 Chem 40/ Chem 35/ Fundamentals of 1 Outline: I. Essence of Biochemistry II. Essential Elements for Living Systems III. Classes of Organic Compounds IV.
Biochemistry: A Review and Introduction CHAPTER 1 Chem 40/ Chem 35/ Fundamentals of 1 Outline: I. Essence of Biochemistry II. Essential Elements for Living Systems III. Classes of Organic Compounds IV.
CHMI 2227 EL. Biochemistry I. Test January Prof : Eric R. Gauthier, Ph.D.
 CHMI 2227 EL Biochemistry I Test 1 26 January 2007 Prof : Eric R. Gauthier, Ph.D. Guidelines: 1) Duration: 55 min 2) 14 questions, on 7 pages. For 70 marks (5 marks per question). Worth 15 % of the final
CHMI 2227 EL Biochemistry I Test 1 26 January 2007 Prof : Eric R. Gauthier, Ph.D. Guidelines: 1) Duration: 55 min 2) 14 questions, on 7 pages. For 70 marks (5 marks per question). Worth 15 % of the final
Electron Transport Chain (Respiratory Chain) - exercise - Vladimíra Kvasnicová
 Electron Transport Chain (Respiratory Chain) - exercise - Vladimíra Kvasnicová Respiratory chain (RCH) a) is found in all cells b) is located in a mitochondrion c) includes enzymes integrated in the inner
Electron Transport Chain (Respiratory Chain) - exercise - Vladimíra Kvasnicová Respiratory chain (RCH) a) is found in all cells b) is located in a mitochondrion c) includes enzymes integrated in the inner
Change to Office Hours this Friday and next Monday. Tomorrow (Abel): 8:30 10:30 am. Monday (Katrina): Cancelled (05/04)
 Change to Office Hours this Friday and next Monday Tomorrow (Abel): 8:30 10:30 am Monday (Katrina): Cancelled (05/04) Lecture 10 Proton Gradient-dependent ATP Synthesis Oxidative Phosphorylation Photo-Phosphorylation
Change to Office Hours this Friday and next Monday Tomorrow (Abel): 8:30 10:30 am Monday (Katrina): Cancelled (05/04) Lecture 10 Proton Gradient-dependent ATP Synthesis Oxidative Phosphorylation Photo-Phosphorylation
Big Idea 1: The process of evolution drives the diversity and unity of life. Sunday, August 28, 16
 Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.B: Organisms are linked by lines of descent from common ancestry. Essential knowledge 1.B.1: Organisms
Big Idea 1: The process of evolution drives the diversity and unity of life. Enduring understanding 1.B: Organisms are linked by lines of descent from common ancestry. Essential knowledge 1.B.1: Organisms
Components of a functional cell. Boundary-membrane Cytoplasm: Cytosol (soluble components) & particulates DNA-information Ribosomes-protein synthesis
 Cell (Outline) - Components of a functional cell - Major Events in the History of Earth: abiotic and biotic phases; anaerobic and aerobic atmosphere - Prokaryotic cells impact on the biosphere - Origin
Cell (Outline) - Components of a functional cell - Major Events in the History of Earth: abiotic and biotic phases; anaerobic and aerobic atmosphere - Prokaryotic cells impact on the biosphere - Origin
Gene Regulatory Network Identification
 Gene Regulatory Network Identification Korkut Uygun and Yinlun Huang Department of Chemical Engineering and Materials Science Wayne State University AIChE Annual National Meeting San Francisco November
Gene Regulatory Network Identification Korkut Uygun and Yinlun Huang Department of Chemical Engineering and Materials Science Wayne State University AIChE Annual National Meeting San Francisco November
Biological Process Term Enrichment
 Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Biological Process Term Enrichment cellular protein localization cellular macromolecule localization intracellular protein transport intracellular transport generation of precursor metabolites and energy
Biology Reading Assignment: Chapter 9 in textbook
 Biology 205 5.10.06 Reading Assignment: Chapter 9 in textbook HTTP://WUNMR.WUSTL.EDU/EDUDEV/LABTUTORIALS/CYTOCHROMES/CYTOCHROMES.HTML What does a cell need to do? propagate itself (and its genetic program)
Biology 205 5.10.06 Reading Assignment: Chapter 9 in textbook HTTP://WUNMR.WUSTL.EDU/EDUDEV/LABTUTORIALS/CYTOCHROMES/CYTOCHROMES.HTML What does a cell need to do? propagate itself (and its genetic program)
Photosynthesis and Cellular Respiration
 Photosynthesis and Cellular Respiration Photosynthesis and Cellular Respiration All cellular activities require energy. Directly or indirectly nearly all energy for life comes from the sun. Autotrophs:
Photosynthesis and Cellular Respiration Photosynthesis and Cellular Respiration All cellular activities require energy. Directly or indirectly nearly all energy for life comes from the sun. Autotrophs:
All organisms require a constant expenditure of energy to maintain the living state - "LIFE".
 CELLULAR RESPIRATION All organisms require a constant expenditure of energy to maintain the living state - "LIFE". Where does the energy come from and how is it made available for life? With rare exception,
CELLULAR RESPIRATION All organisms require a constant expenditure of energy to maintain the living state - "LIFE". Where does the energy come from and how is it made available for life? With rare exception,
Division Ave. High School AP Biology
 Overview 10 reactions u convert () to pyruvate (3C) u produces: 4 & NADH u consumes: u net: & NADH C-C-C-C-C-C fructose-1,6bp P-C-C-C-C-C-C-P DHAP P-C-C-C G3P C-C-C-P H P i P i pyruvate C-C-C 4 4 NAD +
Overview 10 reactions u convert () to pyruvate (3C) u produces: 4 & NADH u consumes: u net: & NADH C-C-C-C-C-C fructose-1,6bp P-C-C-C-C-C-C-P DHAP P-C-C-C G3P C-C-C-P H P i P i pyruvate C-C-C 4 4 NAD +
Chapter 15 part 2. Biochemistry I Introduction to Metabolism Bioenergetics: Thermodynamics in Biochemistry. ATP 4- + H 2 O ADP 3- + P i + H +
 Biochemistry I Introduction to Metabolism Bioenergetics: Thermodynamics in Biochemistry ATP 4- + 2 ADP 3- + P i 2- + + Chapter 15 part 2 Dr. Ray 1 Energy flow in biological systems: Energy Transformations
Biochemistry I Introduction to Metabolism Bioenergetics: Thermodynamics in Biochemistry ATP 4- + 2 ADP 3- + P i 2- + + Chapter 15 part 2 Dr. Ray 1 Energy flow in biological systems: Energy Transformations
9/11/18. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Division Ave. High School AP Biology
 Tour of the Cell 1 Types of cells Prokaryote bacteria cells - no organelles - organelles Eukaryote animal cells Eukaryote plant cells Why organelles? Specialized structures u specialized functions cilia
Tour of the Cell 1 Types of cells Prokaryote bacteria cells - no organelles - organelles Eukaryote animal cells Eukaryote plant cells Why organelles? Specialized structures u specialized functions cilia
The Discovery of Cells
 The Discovery of Cells Microscope observations! General Cell & Organelle Discovery 1600s Observations made by scientists using more powerful microscopes in the 1800s led to the formation of the cell theory.
The Discovery of Cells Microscope observations! General Cell & Organelle Discovery 1600s Observations made by scientists using more powerful microscopes in the 1800s led to the formation of the cell theory.
Courtesy of Elsevier. Used with permission.
 Chemistry 5.07 2013 Problem Set 5 Answers Problem 1 Succinate dehydrogenase (SDH) is a heterotetramer enzyme complex that catalyzes the oxidation of succinate to fumarate with concomitant reduction of
Chemistry 5.07 2013 Problem Set 5 Answers Problem 1 Succinate dehydrogenase (SDH) is a heterotetramer enzyme complex that catalyzes the oxidation of succinate to fumarate with concomitant reduction of
Chemistry of Life Cells & Bioprocesses CRT Review
 Chemistry of Life Cells & Bioprocesses CRT Review Chapter 2: The Chemistry of Life macromolecules - The four types of macromolecules are carbohydrates, lipids, nucleic acids, and proteins Types of Macromolecules
Chemistry of Life Cells & Bioprocesses CRT Review Chapter 2: The Chemistry of Life macromolecules - The four types of macromolecules are carbohydrates, lipids, nucleic acids, and proteins Types of Macromolecules
Biochemistry 3300 Problems (and Solutions) Metabolism I
 (1) Provide a reasonable systematic name for an enzyme that catalyzes the following reaction: fructose + ATP > fructose-1 phosphate + ADP (2) The IUBMB has a developed a set of rules for classifying enzymes
(1) Provide a reasonable systematic name for an enzyme that catalyzes the following reaction: fructose + ATP > fructose-1 phosphate + ADP (2) The IUBMB has a developed a set of rules for classifying enzymes
Biochemical Pathways
 Biochemical Pathways Living organisms can be divided into two large groups according to the chemical form in which they obtain carbon from the environment. Autotrophs can use carbon dioxide from the atmosphere
Biochemical Pathways Living organisms can be divided into two large groups according to the chemical form in which they obtain carbon from the environment. Autotrophs can use carbon dioxide from the atmosphere
Chapter 1. DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d
 Chapter 1 1. Matching Questions DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d 2. Matching Questions : Unbranched polymer that, when folded into its three-dimensional shape,
Chapter 1 1. Matching Questions DNA is made from the building blocks adenine, guanine, cytosine, and. Answer: d 2. Matching Questions : Unbranched polymer that, when folded into its three-dimensional shape,
Review Questions - Lecture 5: Metabolism, Part 1
 Review Questions - Lecture 5: Metabolism, Part 1 Questions: 1. What is metabolism? 2. What does it mean to say that a cell has emergent properties? 3. Define metabolic pathway. 4. What is the difference
Review Questions - Lecture 5: Metabolism, Part 1 Questions: 1. What is metabolism? 2. What does it mean to say that a cell has emergent properties? 3. Define metabolic pathway. 4. What is the difference
9/2/17. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Stamford Public Schools Science Department District Midterm Examination REVIEW
 Stamford Public Schools Science Department District Midterm Examination REVIEW 2015-2016 Honors Biology Student Name: School/Teacher: Date: SPS Honors Biology Midterm Review, January 2016 Page 1 Dear Biology
Stamford Public Schools Science Department District Midterm Examination REVIEW 2015-2016 Honors Biology Student Name: School/Teacher: Date: SPS Honors Biology Midterm Review, January 2016 Page 1 Dear Biology
Genome Annotation. Bioinformatics and Computational Biology. Genome sequencing Assembly. Gene prediction. Protein targeting.
 Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Genome Annotation Bioinformatics and Computational Biology Genome Annotation Frank Oliver Glöckner 1 Genome Analysis Roadmap Genome sequencing Assembly Gene prediction Protein targeting trna prediction
Cell Organelles. a review of structure and function
 Cell Organelles a review of structure and function TEKS and Student Expectations (SE s) B.4 Science concepts. The student knows that cells are the basic structures of all living things with specialized
Cell Organelles a review of structure and function TEKS and Student Expectations (SE s) B.4 Science concepts. The student knows that cells are the basic structures of all living things with specialized
Introduction to Systems Biology. James Gomes KSBS, IITD
 Introduction to Systems Biology James Gomes KSBS, IITD Inner Life of a Cell Cells Exhibit a High Degree of Complexity What are the different types of cells? Is Life an emergent property? What is Systems
Introduction to Systems Biology James Gomes KSBS, IITD Inner Life of a Cell Cells Exhibit a High Degree of Complexity What are the different types of cells? Is Life an emergent property? What is Systems
Systems Biology Lecture 1 history, introduction and definitions. Pawan Dhar
 Systems Biology Lecture 1 history, introduction and definitions Pawan Dhar Historical context 1900 1950 2000 Dominant approach Physiology Molecular biology Focus of study Paradigmatic discovery Functioning
Systems Biology Lecture 1 history, introduction and definitions Pawan Dhar Historical context 1900 1950 2000 Dominant approach Physiology Molecular biology Focus of study Paradigmatic discovery Functioning
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Outer Glycolysis mitochondrial membrane Glucose ATP
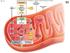 Fig. 7.5 uter Glycolysis mitochondrial membrane Glucose Intermembrane space xidation Mitochondrial matrix Acetyl-oA Krebs FAD e NAD + FAD Inner mitochondrial membrane e Electron e Transport hain hemiosmosis
Fig. 7.5 uter Glycolysis mitochondrial membrane Glucose Intermembrane space xidation Mitochondrial matrix Acetyl-oA Krebs FAD e NAD + FAD Inner mitochondrial membrane e Electron e Transport hain hemiosmosis
Ch. 6 & 7 Photosynthesis & Cellular Respiration
 Ch. 6 & 7 Photosynthesis & Cellular Respiration 6.1 Energy Reactions The Cycle of Energy Sun CO 2 H 2 O Photosynthesis (energy stored) Cellular Respiration (energy released) O 2 Glucose Obtaining Energy
Ch. 6 & 7 Photosynthesis & Cellular Respiration 6.1 Energy Reactions The Cycle of Energy Sun CO 2 H 2 O Photosynthesis (energy stored) Cellular Respiration (energy released) O 2 Glucose Obtaining Energy
Cellular Respiration Stage 4: Electron Transport Chain
 Cellular Respiration Stage 4: Electron Transport Chain 2006-2007 Cellular respiration What s the point? The point is to make ATP! ATP 2006-2007 ATP accounting so far Glycolysis 2 ATP Kreb s cycle 2 ATP
Cellular Respiration Stage 4: Electron Transport Chain 2006-2007 Cellular respiration What s the point? The point is to make ATP! ATP 2006-2007 ATP accounting so far Glycolysis 2 ATP Kreb s cycle 2 ATP
MITOCW watch?v=0xajihttcns
 MITOCW watch?v=0xajihttcns The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high quality educational resources for free. To
MITOCW watch?v=0xajihttcns The following content is provided under a Creative Commons license. Your support will help MIT OpenCourseWare continue to offer high quality educational resources for free. To
Biomolecules. Energetics in biology. Biomolecules inside the cell
 Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Biomolecules Energetics in biology Biomolecules inside the cell Energetics in biology The production of energy, its storage, and its use are central to the economy of the cell. Energy may be defined as
Final Chem 4511/6501 Spring 2011 May 5, 2011 b Name
 Key 1) [10 points] In RNA, G commonly forms a wobble pair with U. a) Draw a G-U wobble base pair, include riboses and 5 phosphates. b) Label the major groove and the minor groove. c) Label the atoms of
Key 1) [10 points] In RNA, G commonly forms a wobble pair with U. a) Draw a G-U wobble base pair, include riboses and 5 phosphates. b) Label the major groove and the minor groove. c) Label the atoms of
20. Electron Transport and Oxidative Phosphorylation
 20. Electron Transport and Oxidative Phosphorylation 20.1 What Role Does Electron Transport Play in Metabolism? Electron transport - Role of oxygen in metabolism as final acceptor of electrons - In inner
20. Electron Transport and Oxidative Phosphorylation 20.1 What Role Does Electron Transport Play in Metabolism? Electron transport - Role of oxygen in metabolism as final acceptor of electrons - In inner
Unit 3: Cell Energy Guided Notes
 Enzymes Unit 3: Cell Energy Guided Notes 1 We get energy from the food we eat by breaking apart the chemical bonds where food is stored. energy is in the bonds, energy is the energy we use to do things.
Enzymes Unit 3: Cell Energy Guided Notes 1 We get energy from the food we eat by breaking apart the chemical bonds where food is stored. energy is in the bonds, energy is the energy we use to do things.
Genome-wide multilevel spatial interactome model of rice
 Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
Sino-German Workshop on Multiscale Spatial Computational Systems Biology, Beijing, Oct 8-12, 2015 Genome-wide multilevel spatial interactome model of rice Ming CHEN ( 陈铭 ) mchen@zju.edu.cn College of Life
Importance of Protein sorting. A clue from plastid development
 Importance of Protein sorting Cell organization depend on sorting proteins to their right destination. Cell functions depend on sorting proteins to their right destination. Examples: A. Energy production
Importance of Protein sorting Cell organization depend on sorting proteins to their right destination. Cell functions depend on sorting proteins to their right destination. Examples: A. Energy production
Photosynthesis and Cellular Respiration Practice Test Name
 Photosynthesis and Cellular Respiration Practice Test Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which H+ has just passed through the
Photosynthesis and Cellular Respiration Practice Test Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which H+ has just passed through the
Cellular Energetics. Photosynthesis, Cellular Respiration and Fermentation
 Cellular Energetics Photosynthesis, Cellular Respiration and Fermentation TEKS B.4 Science concepts. The student knows that cells are the basic structures of all living things with specialized parts that
Cellular Energetics Photosynthesis, Cellular Respiration and Fermentation TEKS B.4 Science concepts. The student knows that cells are the basic structures of all living things with specialized parts that
Chapter 8 Metabolism: Energy, Enzymes, and Regulation
 Chapter 8 Metabolism: Energy, Enzymes, and Regulation Energy: Capacity to do work or cause a particular change. Thus, all physical and chemical processes are the result of the application or movement of
Chapter 8 Metabolism: Energy, Enzymes, and Regulation Energy: Capacity to do work or cause a particular change. Thus, all physical and chemical processes are the result of the application or movement of
Honors Biology Fall Final Exam Study Guide
 Honors Biology Fall Final Exam Study Guide Helpful Information: Exam has 100 multiple choice questions. Be ready with pencils and a four-function calculator on the day of the test. Review ALL vocabulary,
Honors Biology Fall Final Exam Study Guide Helpful Information: Exam has 100 multiple choice questions. Be ready with pencils and a four-function calculator on the day of the test. Review ALL vocabulary,
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1)
 MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) 1) Which of the following statements about the atom A) It has 12 neutrons in its nucleus. B) It
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) 1) Which of the following statements about the atom A) It has 12 neutrons in its nucleus. B) It
Cells & Cell Organelles. Doing Life s Work
 Cells & Cell Organelles Doing Life s Work Types of cells bacteria cells Prokaryote Eukaryotes animal cells plant cells Cell size comparison Animal cell Bacterial cell most bacteria 1-10 microns eukaryotic
Cells & Cell Organelles Doing Life s Work Types of cells bacteria cells Prokaryote Eukaryotes animal cells plant cells Cell size comparison Animal cell Bacterial cell most bacteria 1-10 microns eukaryotic
Biomolecules: lecture 9
 Biomolecules: lecture 9 - understanding further why amino acids are the building block for proteins - understanding the chemical properties amino acids bring to proteins - realizing that many proteins
Biomolecules: lecture 9 - understanding further why amino acids are the building block for proteins - understanding the chemical properties amino acids bring to proteins - realizing that many proteins
MOLECULAR ACTIVITIES OF PLANT CELLS
 MOLECULAR ACTIVITIES OF PLANT CELLS An introduction to plant biochemistry JOHN W. ANDERSON BAgrSc, PhD Reader, Botany Department, School of Biological Sciences, La Trobe University, Bundoora, Victoria,
MOLECULAR ACTIVITIES OF PLANT CELLS An introduction to plant biochemistry JOHN W. ANDERSON BAgrSc, PhD Reader, Botany Department, School of Biological Sciences, La Trobe University, Bundoora, Victoria,
Cellular Energy. How Organisms Obtain Energy Section 2: Photosynthesis Section 3: Cellular Respiration. Click on a lesson name to select.
 Section 1: How Organisms Obtain Energy Section 2: Photosynthesis Section 3: Cellular Respiration Click on a lesson name to select. Section 1 How Organisms Obtain Energy Transformation of Energy Energy
Section 1: How Organisms Obtain Energy Section 2: Photosynthesis Section 3: Cellular Respiration Click on a lesson name to select. Section 1 How Organisms Obtain Energy Transformation of Energy Energy
7.014 Quiz I Handout
 7.014 Quiz I andout Quiz I announcements: Quiz I: Friday, February 27 12:05 12:55 Walker Gym, rd floor (room 5040) **This will be a closed book exam** Quiz Review Session: Wednesday, February 25 7:00 9:00
7.014 Quiz I andout Quiz I announcements: Quiz I: Friday, February 27 12:05 12:55 Walker Gym, rd floor (room 5040) **This will be a closed book exam** Quiz Review Session: Wednesday, February 25 7:00 9:00
Lecture 7 Cell Biolog y ٢٢٢ ١
 Lecture 7 ١ Mitochondria ٢ Mitochondria Mitochondria are the energy factories of the cells. The energy currency for the work that animals must do is the energy-rich molecule adenosine triphosphate (ATP).
Lecture 7 ١ Mitochondria ٢ Mitochondria Mitochondria are the energy factories of the cells. The energy currency for the work that animals must do is the energy-rich molecule adenosine triphosphate (ATP).
Introduction to molecular biology. Mitesh Shrestha
 Introduction to molecular biology Mitesh Shrestha Molecular biology: definition Molecular biology is the study of molecular underpinnings of the process of replication, transcription and translation of
Introduction to molecular biology Mitesh Shrestha Molecular biology: definition Molecular biology is the study of molecular underpinnings of the process of replication, transcription and translation of
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Figure 2.1
 Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Figure 2.1 1) Which compound in Figure 2.1 is an ester? 1) A) a b c d e Answer: D 2) A scientist
Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. Figure 2.1 1) Which compound in Figure 2.1 is an ester? 1) A) a b c d e Answer: D 2) A scientist
5. The cells in the liver that detoxify poison substances contain lots of a. smooth ER b. rough ER c. Golgi apparatus d. lysosomes e.
 Chapter 7 practice 1. What scientist originally came up with the term "cell"? a. von Leeuwenhoek d. Watson b. Hooke e. Virchow c. van der Waals 2. When you wish to look at the coat of a virus on the surface
Chapter 7 practice 1. What scientist originally came up with the term "cell"? a. von Leeuwenhoek d. Watson b. Hooke e. Virchow c. van der Waals 2. When you wish to look at the coat of a virus on the surface
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Transformation of Energy! Energy is the ability to do work.! Thermodynamics is the study of the flow and transformation of energy in the universe.
 Section 1 How Organisms Obtain Energy Transformation of Energy! Energy is the ability to do work.! Thermodynamics is the study of the flow and transformation of energy in the universe. Section 1 How Organisms
Section 1 How Organisms Obtain Energy Transformation of Energy! Energy is the ability to do work.! Thermodynamics is the study of the flow and transformation of energy in the universe. Section 1 How Organisms
METABOLISM CHAPTER 04 BIO 211: ANATOMY & PHYSIOLOGY I. Dr. Lawrence G. Altman Some illustrations are courtesy of McGraw-Hill.
 BIO 211: ANATOMY & PHYSIOLOGY I CHAPTER 04 1 Please wait 20 seconds before starting slide show. Mouse click or Arrow keys to navigate. Hit ESCAPE Key to exit. CELLULAR METABOLISM Dr. Lawrence G. Altman
BIO 211: ANATOMY & PHYSIOLOGY I CHAPTER 04 1 Please wait 20 seconds before starting slide show. Mouse click or Arrow keys to navigate. Hit ESCAPE Key to exit. CELLULAR METABOLISM Dr. Lawrence G. Altman
CELL METABOLISM OVERVIEW Keep the big picture in mind as we discuss the particulars!
 BIO 211: ANATOMY & PHYSIOLOGY I CHAPTER 04 CELLULAR METABOLISM 1 Please wait 20 seconds before starting slide show. Mouse click or Arrow keys to navigate. Hit ESCAPE Key to exit. Dr. Lawrence G. Altman
BIO 211: ANATOMY & PHYSIOLOGY I CHAPTER 04 CELLULAR METABOLISM 1 Please wait 20 seconds before starting slide show. Mouse click or Arrow keys to navigate. Hit ESCAPE Key to exit. Dr. Lawrence G. Altman
Gene regulation II Biochemistry 302. February 27, 2006
 Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Gene regulation II Biochemistry 302 February 27, 2006 Molecular basis of inhibition of RNAP by Lac repressor 35 promoter site 10 promoter site CRP/DNA complex 60 Lewis, M. et al. (1996) Science 271:1247
Lectures by Kathleen Fitzpatrick
 Chapter 10 Chemotrophic Energy Metabolism: Aerobic Respiration Lectures by Kathleen Fitzpatrick Simon Fraser University Figure 10-1 Figure 10-6 Conversion of pyruvate The conversion of pyruvate to acetyl
Chapter 10 Chemotrophic Energy Metabolism: Aerobic Respiration Lectures by Kathleen Fitzpatrick Simon Fraser University Figure 10-1 Figure 10-6 Conversion of pyruvate The conversion of pyruvate to acetyl
Bio 1B Lecture Outline (please print and bring along) Fall, 2007
 Bio 1B Lecture Outline (please print and bring along) Fall, 2007 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #5 -- Molecular genetics and molecular evolution
Bio 1B Lecture Outline (please print and bring along) Fall, 2007 B.D. Mishler, Dept. of Integrative Biology 2-6810, bmishler@berkeley.edu Evolution lecture #5 -- Molecular genetics and molecular evolution
Structural Analysis of Expanding Metabolic Networks
 Genome Informatics 15(1): 35 45 (24) 35 Structural Analysis of Expanding Metabolic Networks Oliver Ebenhöh oliver.ebenhoeh@rz.hu-berlin.de Reinhart Heinrich reinhart.heinrich@rz.hu-berlin.de Thomas Handorf
Genome Informatics 15(1): 35 45 (24) 35 Structural Analysis of Expanding Metabolic Networks Oliver Ebenhöh oliver.ebenhoeh@rz.hu-berlin.de Reinhart Heinrich reinhart.heinrich@rz.hu-berlin.de Thomas Handorf
PHOTOSYNTHESIS: A BRIEF STORY!!!!!
 PHOTOSYNTHESIS: A BRIEF STORY!!!!! This is one of the most important biochemical processes in plants and is amongst the most expensive biochemical processes in plant in terms of investment. Photosynthesis
PHOTOSYNTHESIS: A BRIEF STORY!!!!! This is one of the most important biochemical processes in plants and is amongst the most expensive biochemical processes in plant in terms of investment. Photosynthesis
