MINISYMPOSIUM LOGICAL MODELLING OF (MULTI)CELLULAR NETWORKS
|
|
|
- Carol Dean
- 5 years ago
- Views:
Transcription
1 Friday, July 27th, 11:00 MINISYMPOSIUM LOGICAL MODELLING OF (MULTI)CELLULAR NETWORKS Organizer Pedro T. Monteiro INESC-ID/IST - Univ. de Lisboa Lisboa, PT Pedro.Tiago.Monteiro@tecnico.ulisboa.pt Co-organizer Claudine Chaouiya Instituto Gulbenkian de Ciência Oeiras, PT chaouiya@igc.gulbenkian.pt Minisymposium Keywords: Boolean regulatory networks, Logical models, regulatory circuits, discrete dynamics Boolean and multilevel logical approaches are increasingly used to study the behaviours of biological regulatory networks, which govern essential cellular processes. Moreover, modellers can rely on a broad array of model definitions, simulation methods, computational algorithms and software tools. This mini-symposium aims at introducing these modelling approaches as well as discussing recent advances on both formal aspects and applications. It is organized in connection with the Consortium for Logical Modelling and Tools (CoLoMoTo, which has been launched to promote the logical modelling framework and to provide scientists with dedicated standards, repositories and tools. First, an overview of the field will be presented, covering different model classes, current challenges and application ranges. Four speakers will then present their recent achievements. These will first concern the assessment of the dynamics relying on topological features namely regulatory circuits of the modelled regulatory networks. Appropriate abstractions of large asynchronous dynamics will be then discussed, as well as the concept of model sets. Finally, definition of multicellular logical models by specifying cell-cell communication rules will be illustrated through the dorsalventral axis specification during sea urchin embryogenesis. Acknowledgements: We received supported from Fundação Calouste Gulbenkian and from national funds through Fundação para a Ciência e a Tecnologia (FCT) with reference UID/CEC/50021/2013 and FCT project PTDC/EEI-CTP/2914/2014.
2 Friday, July 27th, 11:00 LOGICAL MODELLING OF CELLULAR NETWORKS, INTRODUCTION AND METHODOLOGICAL CHALLENGES Claudine Chaouiya Instituto Gulbenkian de Ciência Rua da Quinta Grande 6, Oeiras, PT Keywords: Regulatory networks, Logical modelling, Discrete dynamics. This introductory talk will provide the basics on the logical modelling formalism, covering some distinct variants. Logical models define discrete dynamics whose properties of interest will be introduced, including their counterparts for the biological networks under study. Distinctive features of signalling and regulatory networks amenable to such a qualitative approach will be discussed. For large regulatory networks, model definition and analysis are often hampered due to diverse sources of complexity. We will conclude by discussing how to tackle these issues, through methods that have been developed relying on diverse mathematical and computational approaches. Acknowledgements: CC acknowledges financial support from Fundação Calouste Gulbenkian.
3 Friday, July 27th, 11:20 LOCAL NEGATIVE CIRCUITS AND CYCLIC ATTRACTORS: A BOOLEAN SATISFIABILITY APPROACH Elisa Tonello Elisa.Tonello@nottingham.ac.uk School of Mathematical Sciences, University of Nottingham Nottingham, NG7 2RD, UK Keywords: Boolean networks, Negative regulatory circuits, Cyclic attractors, Boolean satisfiability problem. Negative circuits in Boolean networks are necessary for the system to display sustained oscillations. In dimension greater than 6, the negative circuits are not necessarily local. In this work we investigate the existence of Boolean maps with desired properties using Boolean satisfiability problems. We translate the absence of local negative circuits into an expression on n2 n variables. We then implement a necessary condition for the existence of a cyclic attractor. We verify with a satisfiability solver that, for networks of dimension less or equal to 5, a local negative circuit is required for sustained oscillations, and find new counterexamples for n 6.
4 Friday, July 27th, 11:40 BOOLEAN DYNAMICS OF COMPOUND REGULATORY CIRCUITS Elisabeth Remy Institut de Mathématiques de Luminy, Marseille cedex 9, FR Keywords: Boolean networks, Regulatory circuits, Synchronous/asynchronous dynamics, Asymptotic behaviours. In biological regulatory networks represented in terms of signed, directed graphs, topological motifs such as circuits are known to play key dynamical roles. After reviewing established results on the roles of simple motifs, we present recent results on the impact of the addition of a short-cut in a regulatory circuit. More precisely, based on Boolean formalisation of regulatory graphs, we provide complete descriptions of the discrete dynamics of particular motifs, under the synchronous and asynchronous updating schemes. These motifs are made of a circuit of arbitrary length, combining positive and negative interactions in any sequence, and are including a short-cut, and hence a smaller embedded circuit. The asymptotical behavior of such motifs strongly depends on the coherence of the signs of the two involved circuits. Such results on the functionality of motifs allow to improve the dynamical analysis of large regulatory biological networks.
5 Friday, July 27th, 12:00 APPROXIMATING THE BEHAVIOR OF BOOLEAN NETWORKS Heike Siebert DFG Research Center Matheon, Freie Universität Berlin Arnimallee 6, Berlin, DE Keywords: Boolean networks, Asynchronous dynamics, Invariant sets, Boolean model sets. One of the key advantages of Boolean models is their low complexity that allows for a more comprehensive analysis of the state space comparative to, e.g., ordinary differential equation (ODE) models. However, scalability is still an issue, in particular, when considering asynchronous update schemes that are often better suited for capturing the behaviour of biological systems and more comparable to ODE models. To tackle this issue, instead of considering the full state transition graph the model dynamics can be analysed on a coarser level. Rather than computing one trajectory step by step, sets of trajectories can be described by sequences of invariant sets finally leading to an attractor. In this talk, we will discuss some applications of this idea both in the context of single model analysis as well as for model sets.
6 Friday, July 27th, 12:20 LOGICAL MODELLING OF THE REGULATORY NETWORK GOVERNING DORSAL-VENTRAL AXIS SPECIFICATION IN THE SEA URCHIN P. LIVIDUS Denis Thieffry Institut de Biologie de l Ecole Normale Supérieure (IBENS) 46 Rue d Ulm, F Paris, FR Keywords: Developmental patterning, Logical modelling, Multicellular system. Located at the basis of the deuterostome branch, echinoderms occupy a unique position to study the regulatory networks governing embryo morphogenesis. The dorsal-ventral (D-V) axis specification in the sea urchin Paracentrotus lividus is controlled by various transcription factors, including two TGF-βs: Nodal and BMP2/4. However, the signalling network downstream of these key morphogens is not yet fully understood. To identify Nodal and BMP2/4 target genes, we have performed a systematic functional analysis using RNA sequencing and in situ hybridization screens. The analysis of these data enables to delineate novel interactions. To gain further insights into this developmental process, we have developed a predictive dynamical model of the corresponding signalling/regulatory network. More specifically, using a logical modelling framework, we account for the specification of three main ectodermal regions along the D-V axis (ventral, ciliary and dorsal ectoderm) in terms of specific marker gene expression patterns. In our model analysis, we first focused on the computation of stable states and on their reachability in single representative cells, depending on signalling inputs. Next, taking advantage of the software EpiLog ( we have simulated grids of cells connected through signalling molecules. These model simulations correctly reproduces the wild-type pattern, as well as various reported embryo mutant phenotypes, including double Nodal mrna injections.
Synchronous state transition graph
 Heike Siebert, FU Berlin, Molecular Networks WS10/11 2-1 Synchronous state transition graph (0, 2) (1, 2) vertex set X (state space) edges (x,f(x)) every state has only one successor attractors are fixed
Heike Siebert, FU Berlin, Molecular Networks WS10/11 2-1 Synchronous state transition graph (0, 2) (1, 2) vertex set X (state space) edges (x,f(x)) every state has only one successor attractors are fixed
arxiv: v1 [cs.dm] 13 Nov 2014
![arxiv: v1 [cs.dm] 13 Nov 2014 arxiv: v1 [cs.dm] 13 Nov 2014](/thumbs/94/118221044.jpg) 1 arxiv:1411.3539v1 [cs.dm] 13 Nov 2014 Quantification of reachable attractors in asynchronous discrete dynamics Nuno D. Mendes 1,3,4,, Pedro T. Monteiro 1,3,, Jorge Carneiro 1, Elisabeth Remy 2, Claudine
1 arxiv:1411.3539v1 [cs.dm] 13 Nov 2014 Quantification of reachable attractors in asynchronous discrete dynamics Nuno D. Mendes 1,3,4,, Pedro T. Monteiro 1,3,, Jorge Carneiro 1, Elisabeth Remy 2, Claudine
Networks in systems biology
 Networks in systems biology Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2017 M. Macauley (Clemson) Networks in systems
Networks in systems biology Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2017 M. Macauley (Clemson) Networks in systems
Simulation of Gene Regulatory Networks
 Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Simulation of Gene Regulatory Networks Overview I have been assisting Professor Jacques Cohen at Brandeis University to explore and compare the the many available representations and interpretations of
Logical modelling of cellular decisions
 Logical modelling of cellular decisions Denis Thieffry (thieffry@ens.fr) Contents Introduction Logical modelling T-helper cell differentiation MAPK network Porquerolles, June 25th, 2013 Cell proliferation,
Logical modelling of cellular decisions Denis Thieffry (thieffry@ens.fr) Contents Introduction Logical modelling T-helper cell differentiation MAPK network Porquerolles, June 25th, 2013 Cell proliferation,
Logical modelling: Inferring structure from dynamics
 Logical modelling: Inferring structure from dynamics Ling Sun, Therese Lorenz and Alexander Bockmayr Freie Universität Berlin Speaker: Ling Sun 08/09/2018 Ling Sun, Therese Lorenz and Alexander Bockmayr
Logical modelling: Inferring structure from dynamics Ling Sun, Therese Lorenz and Alexander Bockmayr Freie Universität Berlin Speaker: Ling Sun 08/09/2018 Ling Sun, Therese Lorenz and Alexander Bockmayr
arxiv: v1 [q-bio.mn] 2 Aug 2018
![arxiv: v1 [q-bio.mn] 2 Aug 2018 arxiv: v1 [q-bio.mn] 2 Aug 2018](/thumbs/83/87254517.jpg) arxiv:1808.00775v1 [q-bio.mn] 2 Aug 2018 Representing Model Ensembles as Boolean Functions Robert Schwieger, Heike Siebert Department of Mathematics, Freie Universität Berlin, Germany Abstract August 3,
arxiv:1808.00775v1 [q-bio.mn] 2 Aug 2018 Representing Model Ensembles as Boolean Functions Robert Schwieger, Heike Siebert Department of Mathematics, Freie Universität Berlin, Germany Abstract August 3,
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Biological networks CS449 BIOINFORMATICS
 CS449 BIOINFORMATICS Biological networks Programming today is a race between software engineers striving to build bigger and better idiot-proof programs, and the Universe trying to produce bigger and better
CS449 BIOINFORMATICS Biological networks Programming today is a race between software engineers striving to build bigger and better idiot-proof programs, and the Universe trying to produce bigger and better
Logic-Based Modeling in Systems Biology
 Logic-Based Modeling in Systems Biology Alexander Bockmayr LPNMR 09, Potsdam, 16 September 2009 DFG Research Center Matheon Mathematics for key technologies Outline A.Bockmayr, FU Berlin/Matheon 2 I. Systems
Logic-Based Modeling in Systems Biology Alexander Bockmayr LPNMR 09, Potsdam, 16 September 2009 DFG Research Center Matheon Mathematics for key technologies Outline A.Bockmayr, FU Berlin/Matheon 2 I. Systems
Modelling the cell cycle regulatory network
 Chapter 3 Modelling the cell cycle regulatory network 3.1 Dynamical modelling Dynamical properties of a system arise from the interaction of its components. In the case of the cell division cycle, cellular
Chapter 3 Modelling the cell cycle regulatory network 3.1 Dynamical modelling Dynamical properties of a system arise from the interaction of its components. In the case of the cell division cycle, cellular
Positive circuits and d-dimensional spatial differentiation: Application to the formation of sense organs in Drosophila
 Positive circuits and d-dimensional spatial differentiation: Application to the formation of sense organs in Drosophila Anne Crumière Institut de Mathématiques de Luminy Campus de Luminy, Case 97, 3288
Positive circuits and d-dimensional spatial differentiation: Application to the formation of sense organs in Drosophila Anne Crumière Institut de Mathématiques de Luminy Campus de Luminy, Case 97, 3288
Level 3 Proposals/Qualitative Models
 Proposal title Qualitative Models (qual) Proposal authors a Duncan Berenguier TAGC INSERM U928 and IML CNRS UMR 6206, Luminy, 163 av. de Luminy 13288 Marseille, France Claudine Chaouiya IGC Rua da Quinta
Proposal title Qualitative Models (qual) Proposal authors a Duncan Berenguier TAGC INSERM U928 and IML CNRS UMR 6206, Luminy, 163 av. de Luminy 13288 Marseille, France Claudine Chaouiya IGC Rua da Quinta
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
GINsim: A software suite for the qualitative modelling, simulation and analysis of regulatory networks
 BioSystems 84 (2006) 91 100 GINsim: A software suite for the qualitative modelling, simulation and analysis of regulatory networks A. Gonzalez Gonzalez a,b, A. Naldi a,l.sánchez b, D. Thieffry a, C. Chaouiya
BioSystems 84 (2006) 91 100 GINsim: A software suite for the qualitative modelling, simulation and analysis of regulatory networks A. Gonzalez Gonzalez a,b, A. Naldi a,l.sánchez b, D. Thieffry a, C. Chaouiya
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Boolean models of gene regulatory networks. Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016
 Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Boolean models of gene regulatory networks Matthew Macauley Math 4500: Mathematical Modeling Clemson University Spring 2016 Gene expression Gene expression is a process that takes gene info and creates
Positive and negative cycles in Boolean networks
 Positive and negative cycles in Boolean networks In the memory of René Thomas Adrien Richard April 2, 2018 Abstract We review and discuss some results about the influence of positive and negative feedback
Positive and negative cycles in Boolean networks In the memory of René Thomas Adrien Richard April 2, 2018 Abstract We review and discuss some results about the influence of positive and negative feedback
Classification of Random Boolean Networks
 Classification of Random Boolean Networks Carlos Gershenson, School of Cognitive and Computer Sciences University of Sussex Brighton, BN1 9QN, U. K. C.Gershenson@sussex.ac.uk http://www.cogs.sussex.ac.uk/users/carlos
Classification of Random Boolean Networks Carlos Gershenson, School of Cognitive and Computer Sciences University of Sussex Brighton, BN1 9QN, U. K. C.Gershenson@sussex.ac.uk http://www.cogs.sussex.ac.uk/users/carlos
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Random Boolean Networks
 Random Boolean Networks Boolean network definition The first Boolean networks were proposed by Stuart A. Kauffman in 1969, as random models of genetic regulatory networks (Kauffman 1969, 1993). A Random
Random Boolean Networks Boolean network definition The first Boolean networks were proposed by Stuart A. Kauffman in 1969, as random models of genetic regulatory networks (Kauffman 1969, 1993). A Random
Stability Depends on Positive Autoregulation in Boolean Gene Regulatory Networks
 Stability Depends on Positive Autoregulation in Boolean Gene Regulatory Networks Ricardo Pinho 1,2 *, Victor Garcia 3, Manuel Irimia 4 *, Marcus W. Feldman 1 1 Department of Biology, Stanford University,
Stability Depends on Positive Autoregulation in Boolean Gene Regulatory Networks Ricardo Pinho 1,2 *, Victor Garcia 3, Manuel Irimia 4 *, Marcus W. Feldman 1 1 Department of Biology, Stanford University,
FUNDAMENTALS of SYSTEMS BIOLOGY From Synthetic Circuits to Whole-cell Models
 FUNDAMENTALS of SYSTEMS BIOLOGY From Synthetic Circuits to Whole-cell Models Markus W. Covert Stanford University 0 CRC Press Taylor & Francis Group Boca Raton London New York Contents /... Preface, xi
FUNDAMENTALS of SYSTEMS BIOLOGY From Synthetic Circuits to Whole-cell Models Markus W. Covert Stanford University 0 CRC Press Taylor & Francis Group Boca Raton London New York Contents /... Preface, xi
Course plan Academic Year Qualification MSc on Bioinformatics for Health Sciences. Subject name: Computational Systems Biology Code: 30180
 Course plan 201-201 Academic Year Qualification MSc on Bioinformatics for Health Sciences 1. Description of the subject Subject name: Code: 30180 Total credits: 5 Workload: 125 hours Year: 1st Term: 3
Course plan 201-201 Academic Year Qualification MSc on Bioinformatics for Health Sciences 1. Description of the subject Subject name: Code: 30180 Total credits: 5 Workload: 125 hours Year: 1st Term: 3
SCOTCAT Credits: 20 SCQF Level 7 Semester 1 Academic year: 2018/ am, Practical classes one per week pm Mon, Tue, or Wed
 Biology (BL) modules BL1101 Biology 1 SCOTCAT Credits: 20 SCQF Level 7 Semester 1 10.00 am; Practical classes one per week 2.00-5.00 pm Mon, Tue, or Wed This module is an introduction to molecular and
Biology (BL) modules BL1101 Biology 1 SCOTCAT Credits: 20 SCQF Level 7 Semester 1 10.00 am; Practical classes one per week 2.00-5.00 pm Mon, Tue, or Wed This module is an introduction to molecular and
Qualitative Petri Net Modelling of Genetic Networks
 Qualitative Petri Net Modelling of Genetic Networks Claudine Chaouiya 1, Elisabeth Remy 2, and Denis Thieffry 1 1 Institut de biologie du Développement de Marseille Luminy UMR 6216, Case 907 - Luminy,
Qualitative Petri Net Modelling of Genetic Networks Claudine Chaouiya 1, Elisabeth Remy 2, and Denis Thieffry 1 1 Institut de biologie du Développement de Marseille Luminy UMR 6216, Case 907 - Luminy,
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
Cybergenetics: Control theory for living cells
 Department of Biosystems Science and Engineering, ETH-Zürich Cybergenetics: Control theory for living cells Corentin Briat Joint work with Ankit Gupta and Mustafa Khammash Introduction Overview Cybergenetics:
Department of Biosystems Science and Engineering, ETH-Zürich Cybergenetics: Control theory for living cells Corentin Briat Joint work with Ankit Gupta and Mustafa Khammash Introduction Overview Cybergenetics:
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Towards a framework for multi-level modelling in Computational Biology
 Towards a framework for multi-level modelling in Computational Biology Sara Montagna sara.montagna@unibo.it Alma Mater Studiorum Università di Bologna PhD in Electronics, Computer Science and Telecommunications
Towards a framework for multi-level modelling in Computational Biology Sara Montagna sara.montagna@unibo.it Alma Mater Studiorum Università di Bologna PhD in Electronics, Computer Science and Telecommunications
Cooperative development of logical modelling standards and tools with CoLoMoTo
 Bioinformatics Advance Access published January 25, 2015 Cooperative development of logical modelling standards and tools with CoLoMoTo Aurélien Naldi 1, Pedro T. Monteiro 2,3, Christoph Müssel 4, the
Bioinformatics Advance Access published January 25, 2015 Cooperative development of logical modelling standards and tools with CoLoMoTo Aurélien Naldi 1, Pedro T. Monteiro 2,3, Christoph Müssel 4, the
New Computational Methods for Systems Biology
 New Computational Methods for Systems Biology François Fages, Sylvain Soliman The French National Institute for Research in Computer Science and Control INRIA Paris-Rocquencourt Constraint Programming
New Computational Methods for Systems Biology François Fages, Sylvain Soliman The French National Institute for Research in Computer Science and Control INRIA Paris-Rocquencourt Constraint Programming
networks in molecular biology Wolfgang Huber
 networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
networks in molecular biology Wolfgang Huber networks in molecular biology Regulatory networks: components = gene products interactions = regulation of transcription, translation, phosphorylation... Metabolic
Introduction to Bioinformatics
 Systems biology Introduction to Bioinformatics Systems biology: modeling biological p Study of whole biological systems p Wholeness : Organization of dynamic interactions Different behaviour of the individual
Systems biology Introduction to Bioinformatics Systems biology: modeling biological p Study of whole biological systems p Wholeness : Organization of dynamic interactions Different behaviour of the individual
Attractor computation using interconnected Boolean networks: testing growth rate models in E. Coli
 Attractor computation using interconnected Boolean networks: testing growth rate models in E. Coli Madalena Chaves, Alfonso Carta To cite this version: Madalena Chaves, Alfonso Carta. Attractor computation
Attractor computation using interconnected Boolean networks: testing growth rate models in E. Coli Madalena Chaves, Alfonso Carta To cite this version: Madalena Chaves, Alfonso Carta. Attractor computation
Accepted Manuscript. Boolean Modeling of Biological Regulatory Networks: A Methodology Tutorial. Assieh Saadatpour, Réka Albert
 Accepted Manuscript Boolean Modeling of Biological Regulatory Networks: A Methodology Tutorial Assieh Saadatpour, Réka Albert PII: S1046-2023(12)00277-0 DOI: http://dx.doi.org/10.1016/j.ymeth.2012.10.012
Accepted Manuscript Boolean Modeling of Biological Regulatory Networks: A Methodology Tutorial Assieh Saadatpour, Réka Albert PII: S1046-2023(12)00277-0 DOI: http://dx.doi.org/10.1016/j.ymeth.2012.10.012
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
Basins of Attraction, Commitment Sets and Phenotypes of Boolean Networks arxiv: v1 [math.ds] 26 Jul 2018
![Basins of Attraction, Commitment Sets and Phenotypes of Boolean Networks arxiv: v1 [math.ds] 26 Jul 2018 Basins of Attraction, Commitment Sets and Phenotypes of Boolean Networks arxiv: v1 [math.ds] 26 Jul 2018](/thumbs/90/103239239.jpg) Basins of Attraction, Commitment Sets and Phenotypes of Boolean Networks arxiv:1807.10103v1 [math.ds] 26 Jul 2018 Hannes Klarner, Frederike Heinitz, Sarah Nee, Heike Siebert Department of Mathematics,
Basins of Attraction, Commitment Sets and Phenotypes of Boolean Networks arxiv:1807.10103v1 [math.ds] 26 Jul 2018 Hannes Klarner, Frederike Heinitz, Sarah Nee, Heike Siebert Department of Mathematics,
Classification of Random Boolean Networks
 in Artificial Life VIII, Standish, Abbass, Bedau (eds)(mit Press) 2002. pp 1 8 1 Classification of Random Boolean Networks Carlos Gershenson, School of Cognitive and Computer Sciences University of Sussex
in Artificial Life VIII, Standish, Abbass, Bedau (eds)(mit Press) 2002. pp 1 8 1 Classification of Random Boolean Networks Carlos Gershenson, School of Cognitive and Computer Sciences University of Sussex
Basic Synthetic Biology circuits
 Basic Synthetic Biology circuits Note: these practices were obtained from the Computer Modelling Practicals lecture by Vincent Rouilly and Geoff Baldwin at Imperial College s course of Introduction to
Basic Synthetic Biology circuits Note: these practices were obtained from the Computer Modelling Practicals lecture by Vincent Rouilly and Geoff Baldwin at Imperial College s course of Introduction to
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
Hybrid Model of gene regulatory networks, the case of the lac-operon
 Hybrid Model of gene regulatory networks, the case of the lac-operon Laurent Tournier and Etienne Farcot LMC-IMAG, 51 rue des Mathématiques, 38041 Grenoble Cedex 9, France Laurent.Tournier@imag.fr, Etienne.Farcot@imag.fr
Hybrid Model of gene regulatory networks, the case of the lac-operon Laurent Tournier and Etienne Farcot LMC-IMAG, 51 rue des Mathématiques, 38041 Grenoble Cedex 9, France Laurent.Tournier@imag.fr, Etienne.Farcot@imag.fr
Early Development in Invertebrates
 Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Linear Models and Empirical Bayes Methods for Microarrays
 Methods for Microarrays by Gordon Smyth Alex Sánchez and Carme Ruíz de Villa Department d Estadística Universitat de Barcelona 16-12-2004 Outline 1 An introductory example Paper overview 2 3 Lönnsted and
Methods for Microarrays by Gordon Smyth Alex Sánchez and Carme Ruíz de Villa Department d Estadística Universitat de Barcelona 16-12-2004 Outline 1 An introductory example Paper overview 2 3 Lönnsted and
Synthesis of Biological Models from Mutation Experiments
 Synthesis of Biological Models from Mutation Experiments Ali Sinan Köksal, Saurabh Srivastava, Rastislav Bodík, UC Berkeley Evan Pu, MIT Jasmin Fisher, Microsoft Research Cambridge Nir Piterman, University
Synthesis of Biological Models from Mutation Experiments Ali Sinan Köksal, Saurabh Srivastava, Rastislav Bodík, UC Berkeley Evan Pu, MIT Jasmin Fisher, Microsoft Research Cambridge Nir Piterman, University
Biology 218, practise Exam 2, 2011
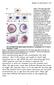 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Mathematical Biology - Lecture 1 - general formulation
 Mathematical Biology - Lecture 1 - general formulation course description Learning Outcomes This course is aimed to be accessible both to masters students of biology who have a good understanding of the
Mathematical Biology - Lecture 1 - general formulation course description Learning Outcomes This course is aimed to be accessible both to masters students of biology who have a good understanding of the
A&S 320: Mathematical Modeling in Biology
 A&S 320: Mathematical Modeling in Biology David Murrugarra Department of Mathematics, University of Kentucky http://www.ms.uky.edu/~dmu228/as320/ These slides were modified from Matthew Macauley s lecture
A&S 320: Mathematical Modeling in Biology David Murrugarra Department of Mathematics, University of Kentucky http://www.ms.uky.edu/~dmu228/as320/ These slides were modified from Matthew Macauley s lecture
Petri net models. tokens placed on places define the state of the Petri net
 Petri nets Petri net models Named after Carl Adam Petri who, in the early sixties, proposed a graphical and mathematical formalism suitable for the modeling and analysis of concurrent, asynchronous distributed
Petri nets Petri net models Named after Carl Adam Petri who, in the early sixties, proposed a graphical and mathematical formalism suitable for the modeling and analysis of concurrent, asynchronous distributed
Francisco M. Couto Mário J. Silva Pedro Coutinho
 Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Francisco M. Couto Mário J. Silva Pedro Coutinho DI FCUL TR 03 29 Departamento de Informática Faculdade de Ciências da Universidade de Lisboa Campo Grande, 1749 016 Lisboa Portugal Technical reports are
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites
 Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Evolutionary analysis of the well characterized endo16 promoter reveals substantial variation within functional sites Paper by: James P. Balhoff and Gregory A. Wray Presentation by: Stephanie Lucas Reviewed
Introduction to Systems Biology
 Introduction to Systems Biology References: Watson s Molecular Biology of the Gene, Chapter 22 Alberts Molecular Biology of the Cell, Chapter 7 Yousof Gheisari ygheisari@med.mui.ac.ir Why is this picture
Introduction to Systems Biology References: Watson s Molecular Biology of the Gene, Chapter 22 Alberts Molecular Biology of the Cell, Chapter 7 Yousof Gheisari ygheisari@med.mui.ac.ir Why is this picture
biological networks Claudine Chaouiya SBML Extention L3F meeting August
 Campus de Luminy - Marseille - France Petri nets and qualitative modelling of biological networks Claudine Chaouiya chaouiya@igc.gulbenkian.pt chaouiya@tagc.univ-mrs.fr SML Extention L3F meeting 1-13 ugust
Campus de Luminy - Marseille - France Petri nets and qualitative modelling of biological networks Claudine Chaouiya chaouiya@igc.gulbenkian.pt chaouiya@tagc.univ-mrs.fr SML Extention L3F meeting 1-13 ugust
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Discrete and Indiscrete Models of Biological Networks
 Discrete and Indiscrete Models of Biological Networks Winfried Just Ohio University November 17, 2010 Who are we? What are we doing here? Who are we? What are we doing here? A population of interacting
Discrete and Indiscrete Models of Biological Networks Winfried Just Ohio University November 17, 2010 Who are we? What are we doing here? Who are we? What are we doing here? A population of interacting
Logical modelling of genetic regulatory networks Contents
 Logical modelling of genetic regulatory networks Contents Boolean modelling of gene networks Multilevel logical modelling Regulatory circuits Application to Drosophila segmentation Biological regulatory
Logical modelling of genetic regulatory networks Contents Boolean modelling of gene networks Multilevel logical modelling Regulatory circuits Application to Drosophila segmentation Biological regulatory
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Complex Systems Theory
 Complex Systems Theory 1988 Some approaches to the study of complex systems are outlined. They are encompassed by an emerging field of science concerned with the general analysis of complexity. Throughout
Complex Systems Theory 1988 Some approaches to the study of complex systems are outlined. They are encompassed by an emerging field of science concerned with the general analysis of complexity. Throughout
7.32/7.81J/8.591J: Systems Biology. Fall Exam #1
 7.32/7.81J/8.591J: Systems Biology Fall 2013 Exam #1 Instructions 1) Please do not open exam until instructed to do so. 2) This exam is closed- book and closed- notes. 3) Please do all problems. 4) Use
7.32/7.81J/8.591J: Systems Biology Fall 2013 Exam #1 Instructions 1) Please do not open exam until instructed to do so. 2) This exam is closed- book and closed- notes. 3) Please do all problems. 4) Use
ACTA PHYSICA DEBRECINA XLVI, 47 (2012) MODELLING GENE REGULATION WITH BOOLEAN NETWORKS. Abstract
 ACTA PHYSICA DEBRECINA XLVI, 47 (2012) MODELLING GENE REGULATION WITH BOOLEAN NETWORKS E. Fenyvesi 1, G. Palla 2 1 University of Debrecen, Department of Experimental Physics, 4032 Debrecen, Egyetem 1,
ACTA PHYSICA DEBRECINA XLVI, 47 (2012) MODELLING GENE REGULATION WITH BOOLEAN NETWORKS E. Fenyvesi 1, G. Palla 2 1 University of Debrecen, Department of Experimental Physics, 4032 Debrecen, Egyetem 1,
Rui Dilão NonLinear Dynamics Group, IST
 1st Conference on Computational Interdisciplinary Sciences (CCIS 2010) 23-27 August 2010, INPE, São José dos Campos, Brasil Modeling, Simulating and Calibrating Genetic Regulatory Networks: An Application
1st Conference on Computational Interdisciplinary Sciences (CCIS 2010) 23-27 August 2010, INPE, São José dos Campos, Brasil Modeling, Simulating and Calibrating Genetic Regulatory Networks: An Application
Three different fusions led to three basic ideas: 1) If one fuses a cell in mitosis with a cell in any other stage of the cell cycle, the chromosomes
 Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
Section Notes The cell division cycle presents an interesting system to study because growth and division must be carefully coordinated. For many cells it is important that it reaches the correct size
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Chaotic Dynamics in an Electronic Model of a Genetic Network
 Journal of Statistical Physics, Vol. 121, Nos. 5/6, December 2005 ( 2005) DOI: 10.1007/s10955-005-7009-y Chaotic Dynamics in an Electronic Model of a Genetic Network Leon Glass, 1 Theodore J. Perkins,
Journal of Statistical Physics, Vol. 121, Nos. 5/6, December 2005 ( 2005) DOI: 10.1007/s10955-005-7009-y Chaotic Dynamics in an Electronic Model of a Genetic Network Leon Glass, 1 Theodore J. Perkins,
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
A New Kind of Language for Complex Engineering Systems:
 NKS 2004 A New Kind of Language for Complex Engineering Systems: Case Study: NASA s Apollo Program Benjamin Koo Edward F. Crawley Engineering Systems Division Department of Aeronautical and Astronautical
NKS 2004 A New Kind of Language for Complex Engineering Systems: Case Study: NASA s Apollo Program Benjamin Koo Edward F. Crawley Engineering Systems Division Department of Aeronautical and Astronautical
Motivation. Evolution has rediscovered several times multicellularity as a way to build complex living systems
 Cellular Systems 1 Motivation Evolution has rediscovered several times multicellularity as a way to build complex living systems Multicellular systems are composed by many copies of a unique fundamental
Cellular Systems 1 Motivation Evolution has rediscovered several times multicellularity as a way to build complex living systems Multicellular systems are composed by many copies of a unique fundamental
DYNAMIC MODELING OF BIOLOGICAL AND PHYSICAL SYSTEMS
 The Pennsylvania State University The Graduate School Eberly College of Science DYNAMIC MODELING OF BIOLOGICAL AND PHYSICAL SYSTEMS A Dissertation in Mathematics by Assieh Saadatpour Moghaddam 2012 Assieh
The Pennsylvania State University The Graduate School Eberly College of Science DYNAMIC MODELING OF BIOLOGICAL AND PHYSICAL SYSTEMS A Dissertation in Mathematics by Assieh Saadatpour Moghaddam 2012 Assieh
Dynamical Systems in Biology
 Dynamical Systems in Biology Hal Smith A R I Z O N A S T A T E U N I V E R S I T Y H.L. Smith (ASU) Dynamical Systems in Biology ASU, July 5, 2012 1 / 31 Outline 1 What s special about dynamical systems
Dynamical Systems in Biology Hal Smith A R I Z O N A S T A T E U N I V E R S I T Y H.L. Smith (ASU) Dynamical Systems in Biology ASU, July 5, 2012 1 / 31 Outline 1 What s special about dynamical systems
1. Supplemental figures and legends
 1. Supplemental figures and legends Figure S1. Phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands. AD, phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands at 72h. A, sea urchin
1. Supplemental figures and legends Figure S1. Phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands. AD, phenotypes of the inactivation of the BMP2/4 and BMP5/8 ligands at 72h. A, sea urchin
Constraint and Contingency in Multifunctional Gene Regulatory Circuits
 in Multifunctional Gene Regulatory Circuits Joshua L. Payne 1,2, Andreas Wagner 1,2,3,4 * 1 University of Zurich, Institute of Evolutionary Biology and Environmental Studies, Zurich, Switzerland, 2 Swiss
in Multifunctional Gene Regulatory Circuits Joshua L. Payne 1,2, Andreas Wagner 1,2,3,4 * 1 University of Zurich, Institute of Evolutionary Biology and Environmental Studies, Zurich, Switzerland, 2 Swiss
Chapter 1. Scientific Process and Themes of Biology
 Chapter 1 Scientific Process and Themes of Biology What is Science? u Scientific knowledge is acquired using a rigorous process u Science is an organized way of gathering and analyzing evidence about the
Chapter 1 Scientific Process and Themes of Biology What is Science? u Scientific knowledge is acquired using a rigorous process u Science is an organized way of gathering and analyzing evidence about the
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Network Biology: Understanding the cell s functional organization. Albert-László Barabási Zoltán N. Oltvai
 Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Network Biology: Understanding the cell s functional organization Albert-László Barabási Zoltán N. Oltvai Outline: Evolutionary origin of scale-free networks Motifs, modules and hierarchical networks Network
Predictive Modeling of Signaling Crosstalk... Model execution and model checking can be used to test a biological hypothesis
 Model execution and model checking can be used to test a biological hypothesis The model is an explanation of a biological mechanism and its agreement with experimental results is used to either validate
Model execution and model checking can be used to test a biological hypothesis The model is an explanation of a biological mechanism and its agreement with experimental results is used to either validate
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Optimal State Estimation for Boolean Dynamical Systems using a Boolean Kalman Smoother
 Optimal State Estimation for Boolean Dynamical Systems using a Boolean Kalman Smoother Mahdi Imani and Ulisses Braga-Neto Department of Electrical and Computer Engineering Texas A&M University College
Optimal State Estimation for Boolean Dynamical Systems using a Boolean Kalman Smoother Mahdi Imani and Ulisses Braga-Neto Department of Electrical and Computer Engineering Texas A&M University College
Correlation Networks
 QuickTime decompressor and a are needed to see this picture. Correlation Networks Analysis of Biological Networks April 24, 2010 Correlation Networks - Analysis of Biological Networks 1 Review We have
QuickTime decompressor and a are needed to see this picture. Correlation Networks Analysis of Biological Networks April 24, 2010 Correlation Networks - Analysis of Biological Networks 1 Review We have
TUNING-RULES FOR FRACTIONAL PID CONTROLLERS
 TUNING-RULES FOR FRACTIONAL PID CONTROLLERS Duarte Valério, José Sá da Costa Technical Univ. of Lisbon Instituto Superior Técnico Department of Mechanical Engineering GCAR Av. Rovisco Pais, 49- Lisboa,
TUNING-RULES FOR FRACTIONAL PID CONTROLLERS Duarte Valério, José Sá da Costa Technical Univ. of Lisbon Instituto Superior Técnico Department of Mechanical Engineering GCAR Av. Rovisco Pais, 49- Lisboa,
A Simple Protein Synthesis Model
 A Simple Protein Synthesis Model James K. Peterson Department of Biological Sciences and Department of Mathematical Sciences Clemson University September 3, 213 Outline A Simple Protein Synthesis Model
A Simple Protein Synthesis Model James K. Peterson Department of Biological Sciences and Department of Mathematical Sciences Clemson University September 3, 213 Outline A Simple Protein Synthesis Model
Stochastic simulations
 Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
An introduction to SYSTEMS BIOLOGY
 An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
An introduction to SYSTEMS BIOLOGY Paolo Tieri CNR Consiglio Nazionale delle Ricerche, Rome, Italy 10 February 2015 Universidade Federal de Minas Gerais, Belo Horizonte, Brasil Course outline Day 1: intro
Computational Modelling in Systems and Synthetic Biology
 Computational Modelling in Systems and Synthetic Biology Fran Romero Dpt Computer Science and Artificial Intelligence University of Seville fran@us.es www.cs.us.es/~fran Models are Formal Statements of
Computational Modelling in Systems and Synthetic Biology Fran Romero Dpt Computer Science and Artificial Intelligence University of Seville fran@us.es www.cs.us.es/~fran Models are Formal Statements of
Sorting Network Development Using Cellular Automata
 Sorting Network Development Using Cellular Automata Michal Bidlo, Zdenek Vasicek, and Karel Slany Brno University of Technology, Faculty of Information Technology Božetěchova 2, 61266 Brno, Czech republic
Sorting Network Development Using Cellular Automata Michal Bidlo, Zdenek Vasicek, and Karel Slany Brno University of Technology, Faculty of Information Technology Božetěchova 2, 61266 Brno, Czech republic
Rigorous numerical computation of polynomial differential equations over unbounded domains
 Rigorous numerical computation of polynomial differential equations over unbounded domains Olivier Bournez 1, Daniel S. Graça 2,3, Amaury Pouly 1,2 1 Ecole Polytechnique, LIX, 91128 Palaiseau Cedex, France
Rigorous numerical computation of polynomial differential equations over unbounded domains Olivier Bournez 1, Daniel S. Graça 2,3, Amaury Pouly 1,2 1 Ecole Polytechnique, LIX, 91128 Palaiseau Cedex, France
BINARY COUNTING WITH CHEMICAL REACTIONS
 BINARY COUNTING WITH CHEMICAL REACTIONS ALEKSANDRA KHARAM, HUA JIANG, MARC RIEDEL, and KESHAB PARHI Electrical and Computer Engineering, University of Minnesota Minneapolis, MN 55455 http://cctbio.ece.umn.edu
BINARY COUNTING WITH CHEMICAL REACTIONS ALEKSANDRA KHARAM, HUA JIANG, MARC RIEDEL, and KESHAB PARHI Electrical and Computer Engineering, University of Minnesota Minneapolis, MN 55455 http://cctbio.ece.umn.edu
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Preface. Motivation and Objectives
 Preface Motivation and Objectives In control theory, complex models of physical processes, such as systems of differential or difference equations, are usually checked against simple specifications, such
Preface Motivation and Objectives In control theory, complex models of physical processes, such as systems of differential or difference equations, are usually checked against simple specifications, such
BILD7: Problem Set. 2. What did Chargaff discover and why was this important?
 BILD7: Problem Set 1. What is the general structure of DNA? 2. What did Chargaff discover and why was this important? 3. What was the major contribution of Rosalind Franklin? 4. How did solving the structure
BILD7: Problem Set 1. What is the general structure of DNA? 2. What did Chargaff discover and why was this important? 3. What was the major contribution of Rosalind Franklin? 4. How did solving the structure
SPA for quantitative analysis: Lecture 6 Modelling Biological Processes
 1/ 223 SPA for quantitative analysis: Lecture 6 Modelling Biological Processes Jane Hillston LFCS, School of Informatics The University of Edinburgh Scotland 7th March 2013 Outline 2/ 223 1 Introduction
1/ 223 SPA for quantitative analysis: Lecture 6 Modelling Biological Processes Jane Hillston LFCS, School of Informatics The University of Edinburgh Scotland 7th March 2013 Outline 2/ 223 1 Introduction
Analog Electronics Mimic Genetic Biochemical Reactions in Living Cells
 Analog Electronics Mimic Genetic Biochemical Reactions in Living Cells Dr. Ramez Daniel Laboratory of Synthetic Biology & Bioelectronics (LSB 2 ) Biomedical Engineering, Technion May 9, 2016 Cytomorphic
Analog Electronics Mimic Genetic Biochemical Reactions in Living Cells Dr. Ramez Daniel Laboratory of Synthetic Biology & Bioelectronics (LSB 2 ) Biomedical Engineering, Technion May 9, 2016 Cytomorphic
On the convergence of Boolean automata networks without negative cycles
 On the convergence of Boolean automata networks without negative cycles Tarek Melliti, Damien Regnault, Adrien Richard, and Sylvain Sené,3 Laboratoire IBISC, EA456, Université d Évry Val-d Essonne, 9000
On the convergence of Boolean automata networks without negative cycles Tarek Melliti, Damien Regnault, Adrien Richard, and Sylvain Sené,3 Laboratoire IBISC, EA456, Université d Évry Val-d Essonne, 9000
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
Coexistence of Dynamics for Two- Dimensional Cellular Automata
 Coexistence of Dynamics for Two- Dimensional Cellular Automata Ricardo Severino Department of Mathematics and Applications University of Minho Campus de Gualtar - 4710-057 Braga, Portugal Maria Joana Soares
Coexistence of Dynamics for Two- Dimensional Cellular Automata Ricardo Severino Department of Mathematics and Applications University of Minho Campus de Gualtar - 4710-057 Braga, Portugal Maria Joana Soares
Lecture 9: June 21, 2007
 Analysis of Gene Expression Data Spring Semester, 2007 Lecture 9: June 21, 2007 Lecturer: Ron Shamir Scribe: Maria Natanzon and Yevgenia Koren 1 9.1 Genetic networks 9.1.1 Preface An ultimate goal of a
Analysis of Gene Expression Data Spring Semester, 2007 Lecture 9: June 21, 2007 Lecturer: Ron Shamir Scribe: Maria Natanzon and Yevgenia Koren 1 9.1 Genetic networks 9.1.1 Preface An ultimate goal of a
BIOINFORMATICS CS4742 BIOLOGICAL NETWORKS
 CS4742 BIOINFORMATICS BIOLOGICAL NETWORKS Programming today is a race between software engineers striving to build bigger and better idiot-proof programs, and the Universe trying to produce bigger and
CS4742 BIOINFORMATICS BIOLOGICAL NETWORKS Programming today is a race between software engineers striving to build bigger and better idiot-proof programs, and the Universe trying to produce bigger and
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
www.sciencesignaling.org/cgi/content/full/7/345/ra91/dc1 Supplementary Materials for TGF-β induced epithelial-to-mesenchymal transition proceeds through stepwise activation of multiple feedback loops Jingyu
