The living cell is a society of molecules: molecules assembling and cooperating bring about life!
|
|
|
- Deirdre Reynolds
- 6 years ago
- Views:
Transcription
1 The Computational Microscope Computational microscope views the cell 100-1,000,000 processors The living cell is a society of molecules: molecules assembling and cooperating bring about life!
2 The Computational Microscope Computational microscope views the cell protein folding (10 4 atoms) photosynthetic chromatophore (10 8 atoms) fibrinogen (10 6 atoms) lipoprotein (10 5 atoms) 100-1,000,000 processors ribosome (10 6 atoms) vesicle formed by BAR domains (5x10 7 atoms)
3 The Computational Microscope View of blood clot elasticity atomic force microscope stretching fibrinogen computer simulation 100-1,000,000 processors
4 Mechanical Strength of a Blood Clot Collaborator: Bernard C. Lim (Mayo Clinic College of Medicine) measurement computation B. Lim, E. Lee, M. Sotomayor, and K. Schulten. Molecular basis of fibrin clot elasticity. Structure, 16: , A Blood Clot Red blood cells within a network of fibrin fibers, composed of polymerized fibrinogen molecules.
5 Mechanical Strength of a Blood Clot An even closer look B. Lim, E. Lee, M. Sotomayor, and K. Schulten. Molecular basis of fibrin clot elasticity. Structure, 16: , 2008.
6 Application to Ribosome X-ray crystallography High resolution (3-5Å) Crystal packing makes it difficult to determine functional state Cryo-EM Lower resolution (typically 8-12Å) Many functional states can be obtained with kirromycin Crystal structures of ribosome and ligands 30S and 50S from 2I2U/2I2V (Berk et al., 2006); L1 protuberance based on 1MZP (Nikulin et al., 2003); L1 protein using MODELLER (Sali and Blundell, 1993) with 1ZHO as template (Nevskaya et al., 2006); A-site finger using 1TWB (Tung and Sanbonmatsu, 2004) as template; trnas from Selmer et al., 2006; ternary complex from 1OB2 (P.Nissen,unpublished) Structures of the ribosome at different stages of the elongation cycle obtained by Cryo-EM (J. Frank. The dynamics of the Ribosome inferred from Cryo-EM, in Conformational Proteomics of Macromolecular Architectures, 2004)
7 Application to Ribosome X-ray crystallography High resolution (3-5Å) Crystal packing makes it difficult to determine functional state Cryo-EM Lower resolution (typically 8-12Å) Many functional states can be obtained map X-ray struct. with kirromycin Crystal structures of ribosome and ligands 30S and 50S from 2I2U/2I2V (Berk et al., 2006); L1 protuberance based on 1MZP (Nikulin et al., 2003); L1 protein using MODELLER (Sali and Blundell, 1993) with 1ZHO as template (Nevskaya et al., 2006); A-site finger using 1TWB (Tung and Sanbonmatsu, 2004) as template; trnas from Selmer et al., 2006; ternary complex from 1OB2 (P.Nissen,unpublished) Structures of the ribosome at different stages of the elongation cycle obtained by Cryo-EM (J. Frank. The dynamics of the Ribosome inferred from Cryo-EM, in Conformational Proteomics of Macromolecular Architectures, 2004)
8 Hybrid Microscopy with X-rays, Electrons, and Bits X-ray crystallography Computational Microscope Electron microscopy APS at Argonne L. Trabuco, E. Villa, K. Mitra, J. Frank, and K. Schulten. Flexible fitting of atomic structures into electron microscopy maps using molecular dynamics. Structure, 16: , NCSA supercomputer FEI microscope
9 The Computational Microscope Computational microscope recognizes atomic resolution picture of ribosome in action 100-1,000,000 processors 300,000 atoms structurally assigned
10 View of a Protein Being Born close view electron microscope structure computed
11 Current MDFF Applications Genetic decoding [1] Regulatory nascent chain [7] J. Frank (Columbia U.) R. Beckmann (U. Munich) Poliovirus J. Hogle (Harvard U.) Flagellar hook K. Namba (Osaka U.) B. pumilus cyanide dihydratase T. Sewell (U. Cape Town) Protein translocation [6,8] C. Akey (Boston U.) R. Beckmann (U. Munich) Ribosome ratcheting J. Frank (Columbia U.) T. Ha (UIUC) [1] Trabuco et al. Structure (2008) 16: [2] Villa et al. PNAS (2009) 106: [3] Sener et al. Chem Phys (2009) 357: [4] Trabuco et al. Methods (2009) 49: [5] Hsin et al. Biophys J (2009) 97: [6] Gumbart et al. Structure (2009) 17: [7] Seidelt et al. Science (2009) 326: [8] Becker et al. Science (2009) 326: Membrane curvature [3,5] N. Hunter (Sheffield U.)
12 The Computational Microscope Computational microscope recognizes atomic resolution picture of photosynthetic apparatus 100-1,000,000 processors
13 Molecular Dynamics Flexible Fitting (MDFF) Simulation In an MDFF simulation, RC-LH1-PufX dimer atoms are steered into high-density regions of the EM map 5 ns of MDFF, followed by a 29 ns of equilibration was performed The entire lipid patch became arched Curvature is anisotropic Lipid patch is twisted NIH Resource for Macromolecular Modeling and Bioinformatics Jen Hsin, James Gumbart, Leonardo G. Trabuco, Elizabeth Villa, Pu Qian, C. Neil Hunter, and Klaus Schulten. Protein-induced membrane curvature investigated through molecular dynamics flexible fitting.biophysical Journal, 97: , Beckman Institute, UIUC
14 LH2 Packing Induces Curvature Rps. spheroides (forms sphere) LH2s form spheres But, Rsp. rubrum (forms disks) LH2s are flat side view after 14 ns
15 NAMD leaders L. Kale J. Phillips S. Kumar (IBM) fibrinogen E. Lee B. Lim (Mayo) lipoprotein A. Shih S. Sligar (UIUC) Acknowledgements 10 µs folding P. Freddolino M. Gruebele (UIUC) BAR domain Y. Yin A. Arkhipov Funding: NIH, NSF Chromatophore J. Hsin D. Chandler JC. Gumbart J. Strumpfer M. Sener ribosome Elizabeth Villa L. Trabucco J. Gumbart J. Frank (Columbia U.) R. Beckman (Gene Center Munich) DOE - Incite Theoretical and Computational Biophysics Group Beckman Institute, UIUC
Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign
 Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign Molecular Modeling Flexible Fitting 3: Application to Ribosome Proofreading of codon recognition
Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign Molecular Modeling Flexible Fitting 3: Application to Ribosome Proofreading of codon recognition
NIH Center for Macromolecular Modeling and Bioinformatics Developer of VMD and NAMD. Beckman Institute
 NIH Center for Macromolecular Modeling and Bioinformatics Developer of VMD and NAMD 5 faculty members (2 physics, 1 chemistry, 1 biochemistry, 1 computer science); 8 developers; 1 system admin; 15 post
NIH Center for Macromolecular Modeling and Bioinformatics Developer of VMD and NAMD 5 faculty members (2 physics, 1 chemistry, 1 biochemistry, 1 computer science); 8 developers; 1 system admin; 15 post
Introduction to Protein Structures - Molecular Graphics Tool
 Lecture 1a Introduction to Protein Structures - Molecular Graphics Tool amino acid tyrosine LH2 energetics traficking Ubiquitin enzymatic control BPTI Highlights of the VMD Molecular Graphics Program >
Lecture 1a Introduction to Protein Structures - Molecular Graphics Tool amino acid tyrosine LH2 energetics traficking Ubiquitin enzymatic control BPTI Highlights of the VMD Molecular Graphics Program >
Molecular Dynamics Flexible Fitting
 Molecular Dynamics Flexible Fitting Ryan McGreevy Research Programmer University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Molecular Dynamics Flexible
Molecular Dynamics Flexible Fitting Ryan McGreevy Research Programmer University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Molecular Dynamics Flexible
NIH Center for Macromolecular Modeling and Bioinformatics Developer of VMD and NAMD. Beckman Institute
 NIH Center for Macromolecular Modeling and Bioinformatics Developer of VMD and NAMD 5 faculty members (2 physics, 1 chemistry, 1 biochemistry, 1 computer science); 8 developers; 1 system admin; 15 post
NIH Center for Macromolecular Modeling and Bioinformatics Developer of VMD and NAMD 5 faculty members (2 physics, 1 chemistry, 1 biochemistry, 1 computer science); 8 developers; 1 system admin; 15 post
VMD: Visual Molecular Dynamics. Our Microscope is Made of...
 VMD: Visual Molecular Dynamics Computational Microscope / Tool to Think amino acid tyrosine enzymatic control BPTI VMD tutorial traficking Ubiquitin case study http://www.ks.uiuc.edu/training/casestudies/
VMD: Visual Molecular Dynamics Computational Microscope / Tool to Think amino acid tyrosine enzymatic control BPTI VMD tutorial traficking Ubiquitin case study http://www.ks.uiuc.edu/training/casestudies/
Mechanical Proteins. Stretching imunoglobulin and fibronectin. domains of the muscle protein titin. Adhesion Proteins of the Immune System
 Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Don t forget to bring your MD tutorial. Potential Energy (hyper)surface
 Don t forget to bring your MD tutorial Lab session starts at 1pm You will have to finish an MD/SMD exercise on α-conotoxin in oxidized and reduced forms Potential Energy (hyper)surface What is Force? Energy
Don t forget to bring your MD tutorial Lab session starts at 1pm You will have to finish an MD/SMD exercise on α-conotoxin in oxidized and reduced forms Potential Energy (hyper)surface What is Force? Energy
Principles and Applications of Molecular Dynamics Simulations with NAMD
 Principles and Applications of Molecular Dynamics Simulations with NAMD Nov. 14, 2016 Computational Microscope NCSA supercomputer JC Gumbart Assistant Professor of Physics Georgia Institute of Technology
Principles and Applications of Molecular Dynamics Simulations with NAMD Nov. 14, 2016 Computational Microscope NCSA supercomputer JC Gumbart Assistant Professor of Physics Georgia Institute of Technology
Molecular Dynamics Flexible Fitting
 Molecular Dynamics Flexible Fitting Ryan McGreevy Research Programmer University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Molecular Dynamics Flexible
Molecular Dynamics Flexible Fitting Ryan McGreevy Research Programmer University of Illinois at Urbana-Champaign NIH Resource for Macromolecular Modeling and Bioinformatics Molecular Dynamics Flexible
Folding WT villin in silico Three folding simulations reach native state within 5-8 µs
 Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign Molecular Modeling Flexible Fitting 1: Introduction to Molecular Dynamics MD force field Equation
Modeling of Cryo-EM Maps Workshop Baylor College of Medicine Klaus Schulten, U. Illinois at Urbana-Champaign Molecular Modeling Flexible Fitting 1: Introduction to Molecular Dynamics MD force field Equation
Introduction to Protein Structures - Molecular Graphics Tool
 Lecture 1a Introduction to Protein Structures - Molecular Graphics Tool amino acid tyrosine LH2 energetics traficking Ubiquitin enzymatic control BPTI VMD - a Tool to Think Carl Woese Atomic, CG, Particle:
Lecture 1a Introduction to Protein Structures - Molecular Graphics Tool amino acid tyrosine LH2 energetics traficking Ubiquitin enzymatic control BPTI VMD - a Tool to Think Carl Woese Atomic, CG, Particle:
ParM filament images were extracted and from the electron micrographs and
 Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Supplemental methods Outline of the EM reconstruction: ParM filament images were extracted and from the electron micrographs and straightened. The digitized images were corrected for the phase of the Contrast
Steered Molecular Dynamics Introduction and Examples
 Steered Molecular Dynamics Introduction and Examples Klaus Schulten Justin Gullingsrud Hui Lu avidin Sergei Izrailev biotin Rosemary Braun Ferenc Molnar Dorina Kosztin Barry Isralewitz Why Steered Molecular
Steered Molecular Dynamics Introduction and Examples Klaus Schulten Justin Gullingsrud Hui Lu avidin Sergei Izrailev biotin Rosemary Braun Ferenc Molnar Dorina Kosztin Barry Isralewitz Why Steered Molecular
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein
 Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
GPU-Accelerated Analysis and Visualization of Large Structures Solved by Molecular Dynamics Flexible Fitting
 GPU-Accelerated Analysis and Visualization of Large Structures Solved by Molecular Dynamics Flexible Fitting John E. Stone, Ryan McGreevy, Barry Isralewitz, and Klaus Schulten Theoretical and Computational
GPU-Accelerated Analysis and Visualization of Large Structures Solved by Molecular Dynamics Flexible Fitting John E. Stone, Ryan McGreevy, Barry Isralewitz, and Klaus Schulten Theoretical and Computational
Mechanical Proteins. Stretching imunoglobulin and fibronectin. domains of the muscle protein titin. Adhesion Proteins of the Immune System
 Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
NSF Summer School UIUC 2003 Molecular Machines of the Living Cell : Photosynthetic. Unit from Light Absorption to ATP Synthesis Melih Sener
 Beckman Institute NSF Summer School UIUC 2003 Molecular Machines of the Living Cell : Photosynthetic Unit from Light Absorption to ATP Synthesis Melih Sener Thorsten Ritz Ioan Kosztin Ana Damjanovic Theoretical
Beckman Institute NSF Summer School UIUC 2003 Molecular Machines of the Living Cell : Photosynthetic Unit from Light Absorption to ATP Synthesis Melih Sener Thorsten Ritz Ioan Kosztin Ana Damjanovic Theoretical
The Molecular Dynamics Simulation Process
 The Molecular Dynamics Simulation Process For textbooks see: M.P. Allen and D.J. Tildesley. Computer Simulation of Liquids.Oxford University Press, New York, 1987. D. Frenkel and B. Smit. Understanding
The Molecular Dynamics Simulation Process For textbooks see: M.P. Allen and D.J. Tildesley. Computer Simulation of Liquids.Oxford University Press, New York, 1987. D. Frenkel and B. Smit. Understanding
The Computational Microscope
 The Computational Microscope Computational microscope views at atomic resolution... Rs SER RER C E M RER N GA L... how living cells maintain health and battle disease John Stone Our Microscope is Made
The Computational Microscope Computational microscope views at atomic resolution... Rs SER RER C E M RER N GA L... how living cells maintain health and battle disease John Stone Our Microscope is Made
Steered Molecular Dynamics Introduction and Examples
 Steered Molecular Dynamics Introduction and Examples Klaus Schulten Justin Gullingsrud Hui Lu avidin Sergei Izrailev biotin Rosemary Braun Ferenc Molnar Dorina Kosztin Barry Isralewitz Why Steered Molecular
Steered Molecular Dynamics Introduction and Examples Klaus Schulten Justin Gullingsrud Hui Lu avidin Sergei Izrailev biotin Rosemary Braun Ferenc Molnar Dorina Kosztin Barry Isralewitz Why Steered Molecular
Klaus Schulten Department of Physics and Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign
 Klaus Schulten Department of Physics and Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign GTC, San Jose Convention Center, CA Sept. 20 23, 2010 GPU and the Computational
Klaus Schulten Department of Physics and Theoretical and Computational Biophysics Group University of Illinois at Urbana-Champaign GTC, San Jose Convention Center, CA Sept. 20 23, 2010 GPU and the Computational
Flexible Fitting of Atomic Structures into Electron Microscopy Maps Using Molecular Dynamics
 Technical Advance Flexible Fitting of Atomic Structures into Electron Microscopy Maps Using Molecular Dynamics Leonardo G. Trabuco, 1,2,6 Elizabeth Villa, 1,2,6 Kakoli Mitra, 3,7 Joachim Frank, 3,4,8 and
Technical Advance Flexible Fitting of Atomic Structures into Electron Microscopy Maps Using Molecular Dynamics Leonardo G. Trabuco, 1,2,6 Elizabeth Villa, 1,2,6 Kakoli Mitra, 3,7 Joachim Frank, 3,4,8 and
protein synthesis and the ribosome
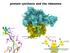 protein synthesis and the ribosome Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process RNA codes for proteins
protein synthesis and the ribosome Central dogma of biology DNA codes for DNA DNA codes for RNA RNA codes for proteins not surprisingly, many points for regulation of the process RNA codes for proteins
Application examples of single particle 3D reconstruction. Ning Gao Tsinghua University
 Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Part III - Bioinformatics Study of Aminoacyl trna Synthetases. VMD Multiseq Tutorial Web tools. Perth, Australia 2004 Computational Biology Workshop
 Part III - Bioinformatics Study of Aminoacyl trna Synthetases VMD Multiseq Tutorial Web tools Perth, Australia 2004 Computational Biology Workshop Multiple Sequence Alignments The aminoacyl-trna synthetases,
Part III - Bioinformatics Study of Aminoacyl trna Synthetases VMD Multiseq Tutorial Web tools Perth, Australia 2004 Computational Biology Workshop Multiple Sequence Alignments The aminoacyl-trna synthetases,
Our Microscope is Made of...
 The Computational Microscope Computational microscope views at atomic resolution... Rs E M RER SER N RER C GA L... how living cells maintain health and battle disease John%Stone Our Microscope is Made
The Computational Microscope Computational microscope views at atomic resolution... Rs E M RER SER N RER C GA L... how living cells maintain health and battle disease John%Stone Our Microscope is Made
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Our Microscope is Made of...
 The Computational Microscope Computational microscope views at atomic resolution... Rs E M RER SER N RER C GA L... how living cells maintain health and battle disease John%Stone Our Microscope is Made
The Computational Microscope Computational microscope views at atomic resolution... Rs E M RER SER N RER C GA L... how living cells maintain health and battle disease John%Stone Our Microscope is Made
Enhanced Sampling and Free Energy Applications in Biomolecular Modeling
 Enhanced Sampling and Free Energy Applications in Biomolecular Modeling Emad Tajkhorshid NIH Center for Macromolecular Modeling and Bioinformatics Beckman Institute for Advanced Science and Technology
Enhanced Sampling and Free Energy Applications in Biomolecular Modeling Emad Tajkhorshid NIH Center for Macromolecular Modeling and Bioinformatics Beckman Institute for Advanced Science and Technology
Measurement of Dynamic trna Movement and Visualizing Ribosome Biogenesis Applications of Time resolved Electron Cryo microscopy
 2011 TIGP CBMB Seminar Measurement of Dynamic trna Movement and Visualizing Ribosome Biogenesis Applications of Time resolved Electron Cryo microscopy Ribosome dynamics and trna movement by time resolved
2011 TIGP CBMB Seminar Measurement of Dynamic trna Movement and Visualizing Ribosome Biogenesis Applications of Time resolved Electron Cryo microscopy Ribosome dynamics and trna movement by time resolved
Molecular Dynamics Investigation of the ω-current in the Kv1.2 Voltage Sensor Domains
 Molecular Dynamics Investigation of the ω-current in the Kv1.2 Voltage Sensor Domains Fatemeh Khalili-Araghi, Emad Tajkhorshid, Benoît Roux, and Klaus Schulten Department of Physics, Department of Biochemistry,
Molecular Dynamics Investigation of the ω-current in the Kv1.2 Voltage Sensor Domains Fatemeh Khalili-Araghi, Emad Tajkhorshid, Benoît Roux, and Klaus Schulten Department of Physics, Department of Biochemistry,
Molecular Biology - Translation of RNA to make Protein *
 OpenStax-CNX module: m49485 1 Molecular Biology - Translation of RNA to make Protein * Jerey Mahr Based on Translation by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative
OpenStax-CNX module: m49485 1 Molecular Biology - Translation of RNA to make Protein * Jerey Mahr Based on Translation by OpenStax This work is produced by OpenStax-CNX and licensed under the Creative
Computational Modeling of Protein Dynamics. Yinghao Wu Department of Systems and Computational Biology Albert Einstein College of Medicine Fall 2014
 Computational Modeling of Protein Dynamics Yinghao Wu Department of Systems and Computational Biology Albert Einstein College of Medicine Fall 2014 Outline Background of protein dynamics Basic computational
Computational Modeling of Protein Dynamics Yinghao Wu Department of Systems and Computational Biology Albert Einstein College of Medicine Fall 2014 Outline Background of protein dynamics Basic computational
Our Microscope is Made of...
 The Computational Microscope Computational microscope views at atomic resolution... Rs E M RER SER N RER C GA L... how living cells maintain health and battle disease John%Stone Our Microscope is Made
The Computational Microscope Computational microscope views at atomic resolution... Rs E M RER SER N RER C GA L... how living cells maintain health and battle disease John%Stone Our Microscope is Made
Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass
 Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass University of California, Davis, USA http://www.cs.ucdavis.edu/~koehl/ims2017/ Biology = Multiscale. 10 6 m 10 3 m m mm µm nm Å ps
Tutorial 1 Geometry, Topology, and Biology Patrice Koehl and Joel Hass University of California, Davis, USA http://www.cs.ucdavis.edu/~koehl/ims2017/ Biology = Multiscale. 10 6 m 10 3 m m mm µm nm Å ps
Model building and validation for cryo-em maps at low resolution
 International Workshop of Advanced Image Processing of Cryo-Electron Microscopy 2015 Tsinghua University, Beijing, China June 3-7, 2015 Model building and validation for cryo-em maps at low resolution
International Workshop of Advanced Image Processing of Cryo-Electron Microscopy 2015 Tsinghua University, Beijing, China June 3-7, 2015 Model building and validation for cryo-em maps at low resolution
Reading the genetic code: The 3D version. Venki Ramakrishnan MRC Laboratory of Molecular Biology Cambridge, United Kingdom
 Reading the genetic code: The 3D version Venki Ramakrishnan MRC Laboratory of Molecular Biology Cambridge, United Kingdom DNA The Central Dogma The Central Dogma DNA mrna The Central Dogma DNA mrna Protein
Reading the genetic code: The 3D version Venki Ramakrishnan MRC Laboratory of Molecular Biology Cambridge, United Kingdom DNA The Central Dogma The Central Dogma DNA mrna The Central Dogma DNA mrna Protein
Overview & Applications. T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 04 June, 2015
 Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Overview & Applications T. Lezon Hands-on Workshop in Computational Biophysics Pittsburgh Supercomputing Center 4 June, 215 Simulations still take time Bakan et al. Bioinformatics 211. Coarse-grained Elastic
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data
 Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Computational Modeling of Protein Kinase A and Comparison with Nuclear Magnetic Resonance Data ABSTRACT Keyword Lei Shi 1 Advisor: Gianluigi Veglia 1,2 Department of Chemistry 1, & Biochemistry, Molecular
Joachim Frank Wadsworth Center Empire State Plaza P.O. Box 509 Albany, New York Tel: (518)
 This material is provided for educational use only. The information in these slides including all data, images and related materials are the property of : Joachim Frank Wadsworth Center Empire State Plaza
This material is provided for educational use only. The information in these slides including all data, images and related materials are the property of : Joachim Frank Wadsworth Center Empire State Plaza
Dissipative Particle Dynamics: Foundation, Evolution and Applications
 Dissipative Particle Dynamics: Foundation, Evolution and Applications Lecture 4: DPD in soft matter and polymeric applications George Em Karniadakis Division of Applied Mathematics, Brown University &
Dissipative Particle Dynamics: Foundation, Evolution and Applications Lecture 4: DPD in soft matter and polymeric applications George Em Karniadakis Division of Applied Mathematics, Brown University &
History of 3D Electron Microscopy and Helical Reconstruction
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
Nanotechnology? Source: National Science Foundation (NSF), USA
 2 2 Nanotechnology? Ability to work at the atomic, molecular and even sub-molecular levels in order to create and use material structures, devices and systems with new properties and functions Source:
2 2 Nanotechnology? Ability to work at the atomic, molecular and even sub-molecular levels in order to create and use material structures, devices and systems with new properties and functions Source:
Introduction to the Ribosome Overview of protein synthesis on the ribosome Prof. Anders Liljas
 Introduction to the Ribosome Molecular Biophysics Lund University 1 A B C D E F G H I J Genome Protein aa1 aa2 aa3 aa4 aa5 aa6 aa7 aa10 aa9 aa8 aa11 aa12 aa13 a a 14 How is a polypeptide synthesized? 2
Introduction to the Ribosome Molecular Biophysics Lund University 1 A B C D E F G H I J Genome Protein aa1 aa2 aa3 aa4 aa5 aa6 aa7 aa10 aa9 aa8 aa11 aa12 aa13 a a 14 How is a polypeptide synthesized? 2
What can be learnt from MD
 - 4.0 H-bond energy (kcal/mol) 0 What can be learnt from MD Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis ATPase, a molecular motor
- 4.0 H-bond energy (kcal/mol) 0 What can be learnt from MD Fibronectin III_1, a mechanical protein that glues cells together in wound healing and in preventing tumor metastasis ATPase, a molecular motor
Lesson Overview. Ribosomes and Protein Synthesis 13.2
 13.2 The Genetic Code The first step in decoding genetic messages is to transcribe a nucleotide base sequence from DNA to mrna. This transcribed information contains a code for making proteins. The Genetic
13.2 The Genetic Code The first step in decoding genetic messages is to transcribe a nucleotide base sequence from DNA to mrna. This transcribed information contains a code for making proteins. The Genetic
Videos. Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.
 Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Translation Translation Videos Bozeman, transcription and translation: https://youtu.be/h3b9arupxzg Crashcourse: Transcription and Translation - https://youtu.be/itsb2sqr-r0 Translation Translation The
Classification in Real Life (Precise Title to be Announced)
 Classification in Real Life (Precise Title to be Announced) Joachim Frank Howard Hughes Medical Institute Department of Biochemistry and Molecular Biophysics and Department of Biological Sciences Columbia
Classification in Real Life (Precise Title to be Announced) Joachim Frank Howard Hughes Medical Institute Department of Biochemistry and Molecular Biophysics and Department of Biological Sciences Columbia
Molecular Mechanisms of self-assembly and motion of the bacterial flagellum
 Molecular Mechanisms of self-assembly and motion of the bacterial flagellum Mexico-Japan Workshop on Pharmacobiology and Nanobiology Universidad Nacional Autonoma de México (UNAM) Mexico City, Mexico 2009.02.25-26
Molecular Mechanisms of self-assembly and motion of the bacterial flagellum Mexico-Japan Workshop on Pharmacobiology and Nanobiology Universidad Nacional Autonoma de México (UNAM) Mexico City, Mexico 2009.02.25-26
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Chapter
 Chapter 17 17.4-17.6 Molecular Components of Translation A cell interprets a genetic message and builds a polypeptide The message is a series of codons on mrna The interpreter is called transfer (trna)
Chapter 17 17.4-17.6 Molecular Components of Translation A cell interprets a genetic message and builds a polypeptide The message is a series of codons on mrna The interpreter is called transfer (trna)
MCB 110. "Molecular Biology: Macromolecular Synthesis and Cellular Function" Spring, 2018
 MCB 110 "Molecular Biology: Macromolecular Synthesis and Cellular Function" Spring, 2018 Faculty Instructors: Prof. Jeremy Thorner Prof. Qiang Zhou Prof. Eva Nogales GSIs:!!!! Ms. Samantha Fernandez Mr.
MCB 110 "Molecular Biology: Macromolecular Synthesis and Cellular Function" Spring, 2018 Faculty Instructors: Prof. Jeremy Thorner Prof. Qiang Zhou Prof. Eva Nogales GSIs:!!!! Ms. Samantha Fernandez Mr.
Translation Part 2 of Protein Synthesis
 Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Translation Part 2 of Protein Synthesis IN: How is transcription like making a jello mold? (be specific) What process does this diagram represent? A. Mutation B. Replication C.Transcription D.Translation
Diffusion in the cell
 Diffusion in the cell Single particle (random walk) Microscopic view Macroscopic view Measuring diffusion Diffusion occurs via Brownian motion (passive) Ex.: D = 100 μm 2 /s for typical protein in water
Diffusion in the cell Single particle (random walk) Microscopic view Macroscopic view Measuring diffusion Diffusion occurs via Brownian motion (passive) Ex.: D = 100 μm 2 /s for typical protein in water
Simple Models of Protein Folding
 Simple Models of Protein Folding Andrew Blanchard May 11, 2012 Abstract Lattice models with simplified interaction potentials have been used to analyze the ability of certain amino acid sequences to adopt
Simple Models of Protein Folding Andrew Blanchard May 11, 2012 Abstract Lattice models with simplified interaction potentials have been used to analyze the ability of certain amino acid sequences to adopt
Steered Molecular Dynamics Studies of Titin Domains
 Steered Molecular Dynamics Studies of Titin Domains Mu Gao Department of Physics and Beckman Institute University of Illinois at Urbana-Champaign Overview Biology Background AFM experiments Modeling and
Steered Molecular Dynamics Studies of Titin Domains Mu Gao Department of Physics and Beckman Institute University of Illinois at Urbana-Champaign Overview Biology Background AFM experiments Modeling and
Cellular Processes in Bacterial Cells
 Part II - Applications of MultiSeq Evolution of Translation: Dynamics of Recognition in RNA:Protein Complexes Part III Towards in silico Cells: Simulating processes in entire cells Zaida (Zan) Luthey-Schulten
Part II - Applications of MultiSeq Evolution of Translation: Dynamics of Recognition in RNA:Protein Complexes Part III Towards in silico Cells: Simulating processes in entire cells Zaida (Zan) Luthey-Schulten
BCH 4054 Spring 2001 Chapter 33 Lecture Notes
 BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
BCH 4054 Spring 2001 Chapter 33 Lecture Notes Slide 1 The chapter covers degradation of proteins as well. We will not have time to get into that subject. Chapter 33 Protein Synthesis Slide 2 Prokaryotic
Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy
 Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy Amit Singer Princeton University, Department of Mathematics and PACM July 23, 2014 Amit Singer (Princeton
Introduction to Three Dimensional Structure Determination of Macromolecules by Cryo-Electron Microscopy Amit Singer Princeton University, Department of Mathematics and PACM July 23, 2014 Amit Singer (Princeton
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
ATPase Synthase - A Molecular Double Motor
 ATPase Synthase - A Molecular Double Motor RC bc 1 bc 1 ATPase Photosynthesic Unit of Purple Bacteria Module that converts sun light into chemical energy (ATP) Light in hν H + Q/QH 2 /Q bc 1 ADP ATP out
ATPase Synthase - A Molecular Double Motor RC bc 1 bc 1 ATPase Photosynthesic Unit of Purple Bacteria Module that converts sun light into chemical energy (ATP) Light in hν H + Q/QH 2 /Q bc 1 ADP ATP out
A general overview of. Free energy methods. Comer et al. (2014) JPCB 119:1129. James C. (JC) Gumbart Georgia Institute of Technology, Atlanta
 A general overview of Free energy methods Comer et al. (2014) JPCB 119:1129. James C. (JC) Gumbart Georgia Institute of Technology, Atlanta Computational Biophysics Workshop Georgia Tech Nov. 16 2016 Outline
A general overview of Free energy methods Comer et al. (2014) JPCB 119:1129. James C. (JC) Gumbart Georgia Institute of Technology, Atlanta Computational Biophysics Workshop Georgia Tech Nov. 16 2016 Outline
Announcements. Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1.
 Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature12682 Supplementary Information 1 Theoretical models of flagellum growth on the outside of a bacterial cell A classic problem in polymer physics concerns the
SUPPLEMENTARY INFORMATION doi:10.1038/nature12682 Supplementary Information 1 Theoretical models of flagellum growth on the outside of a bacterial cell A classic problem in polymer physics concerns the
Electron microscopy in molecular cell biology II
 Electron microscopy in molecular cell biology II Cryo-EM and image processing Werner Kühlbrandt Max Planck Institute of Biophysics Sample preparation for cryo-em Preparation laboratory Specimen preparation
Electron microscopy in molecular cell biology II Cryo-EM and image processing Werner Kühlbrandt Max Planck Institute of Biophysics Sample preparation for cryo-em Preparation laboratory Specimen preparation
BACTERIAL PHYSIOLOGY SMALL GROUP. Monday, August 25, :00pm. Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D.
 BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
BACTERIAL PHYSIOLOGY SMALL GROUP Monday, August 25, 2014 1:00pm Faculty: Adam Driks, Ph.D. Alan Wolfe, Ph.D. Learning Goal To understand how bacterial physiology applies to the diagnosis and treatment
Protein Structures: Experiments and Modeling. Patrice Koehl
 Protein Structures: Experiments and Modeling Patrice Koehl Structural Bioinformatics: Proteins Proteins: Sources of Structure Information Proteins: Homology Modeling Proteins: Ab initio prediction Proteins:
Protein Structures: Experiments and Modeling Patrice Koehl Structural Bioinformatics: Proteins Proteins: Sources of Structure Information Proteins: Homology Modeling Proteins: Ab initio prediction Proteins:
DNA/RNA structure and packing
 DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
Supplemental Information
 1 Supplemental Information Head swivel on the ribosome facilitates translocation via intra-subunit trna hybrid sites Andreas H. Ratje, Justus Loerke, Aleksandra Mikolajka, Matthias Brünner,Peter W. Hildebrand,
1 Supplemental Information Head swivel on the ribosome facilitates translocation via intra-subunit trna hybrid sites Andreas H. Ratje, Justus Loerke, Aleksandra Mikolajka, Matthias Brünner,Peter W. Hildebrand,
Lecture CIMST Winter School. 1. What can you see by TEM?
 Lecture CIMST Winter School Cryo-electron microscopy and tomography of biological macromolecules 20.1.2011 9:00-9:45 in Y03G91 Dr. Takashi Ishikawa OFLB/005 Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch
Lecture CIMST Winter School Cryo-electron microscopy and tomography of biological macromolecules 20.1.2011 9:00-9:45 in Y03G91 Dr. Takashi Ishikawa OFLB/005 Tel: 056 310 4217 e-mail: takashi.ishikawa@psi.ch
2D surface confined nanoporous molecular networks at the liquid-solid interface: a scanning tunneling microscopy study
 2D surface confined nanoporous molecular networks at the liquid-solid interface: a scanning tunneling microscopy study Shengbin Lei Laboratory of Photochemistry and Spectroscopy and INPAC KULeuven Content
2D surface confined nanoporous molecular networks at the liquid-solid interface: a scanning tunneling microscopy study Shengbin Lei Laboratory of Photochemistry and Spectroscopy and INPAC KULeuven Content
Analyzing Ion channel Simulations
 Analyzing Ion channel Simulations (Neher and Sakmann, Scientific American 1992) Single channel current (Heurteaux et al, EMBO 2004) Computational Patch Clamp (Molecular Dynamics) Atoms move according to
Analyzing Ion channel Simulations (Neher and Sakmann, Scientific American 1992) Single channel current (Heurteaux et al, EMBO 2004) Computational Patch Clamp (Molecular Dynamics) Atoms move according to
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Fibrin Gelation During Blood Clotting
 Fibrin Gelation During Blood Clotting Aaron L. Fogelson Department of Mathematics University of Utah July 11, 2016 SIAM LS Conference Boston Acknowledgments Joint work with Jim Keener and Cheryl Zapata-Allegro,
Fibrin Gelation During Blood Clotting Aaron L. Fogelson Department of Mathematics University of Utah July 11, 2016 SIAM LS Conference Boston Acknowledgments Joint work with Jim Keener and Cheryl Zapata-Allegro,
Goals. Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions
 Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke 1,2 and Carlos Camacho 3 1 Bioengineering and Bioinformatics Summer Institute,
Structural Analysis of the EGR Family of Transcription Factors: Templates for Predicting Protein DNA Interactions Jamie Duke 1,2 and Carlos Camacho 3 1 Bioengineering and Bioinformatics Summer Institute,
Physical Biology of the Cell. Rob Phillips, Jane Kondev and Julie Theriot
 Physical Biology of the Cell Rob Phillips, Jane Kondev and Julie Theriot April 4, 2008 Contents 0.1 Preface................................ 14 I The Facts of Life 21 1 Why: Biology By the Numbers 23 1.1
Physical Biology of the Cell Rob Phillips, Jane Kondev and Julie Theriot April 4, 2008 Contents 0.1 Preface................................ 14 I The Facts of Life 21 1 Why: Biology By the Numbers 23 1.1
NAMD - An Exploring Tool
 NAMD - An Exploring Tool 20 years of computer science innovation and collaboration 1993: HP workstation cluster 1994: Writing NAMD in C++ 1998: Commodity Linux clusters 2002: Parallel on 3000 cores,10
NAMD - An Exploring Tool 20 years of computer science innovation and collaboration 1993: HP workstation cluster 1994: Writing NAMD in C++ 1998: Commodity Linux clusters 2002: Parallel on 3000 cores,10
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands
 Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Dynamic condensation of linker histone C-terminal domain regulates. chromatin structure
 Dynamic condensation of linker histone C-terminal domain regulates chromatin structure Supplementary Information Antoni Luque, Rosana Collepardo-Guevara, Sergei A. Grigoryev, and Tamar Schlick Refined
Dynamic condensation of linker histone C-terminal domain regulates chromatin structure Supplementary Information Antoni Luque, Rosana Collepardo-Guevara, Sergei A. Grigoryev, and Tamar Schlick Refined
Large Macromolecular Assemblies (LMAs)
 My Postdoctoral Research My Postdoctoral Research a reduced model My Postdoctoral Research Large Macromolecular Assemblies (LMAs) Large macromolecular assemblies (such as ribosomes, chaperonins, viruses,
My Postdoctoral Research My Postdoctoral Research a reduced model My Postdoctoral Research Large Macromolecular Assemblies (LMAs) Large macromolecular assemblies (such as ribosomes, chaperonins, viruses,
Computational Biology 1
 Computational Biology 1 Protein Function & nzyme inetics Guna Rajagopal, Bioinformatics Institute, guna@bii.a-star.edu.sg References : Molecular Biology of the Cell, 4 th d. Alberts et. al. Pg. 129 190
Computational Biology 1 Protein Function & nzyme inetics Guna Rajagopal, Bioinformatics Institute, guna@bii.a-star.edu.sg References : Molecular Biology of the Cell, 4 th d. Alberts et. al. Pg. 129 190
Evolution of Translation: Dynamics of Recognition in RNA:Protein Complexes
 Evolution of Translation: Dynamics of Recognition in RNA:Protein Complexes Zaida (Zan) Luthey-Schulten Dept. Chemistry, Beckman Institute, Biophysics, Institute of Genomics Biology, & Physics NIH Workshop
Evolution of Translation: Dynamics of Recognition in RNA:Protein Complexes Zaida (Zan) Luthey-Schulten Dept. Chemistry, Beckman Institute, Biophysics, Institute of Genomics Biology, & Physics NIH Workshop
From Gene to Protein
 From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
From Gene to Protein Gene Expression Process by which DNA directs the synthesis of a protein 2 stages transcription translation All organisms One gene one protein 1. Transcription of DNA Gene Composed
Anatoly B. Kolomeisky. Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS?
 Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Anatoly B. Kolomeisky Department of Chemistry CAN WE UNDERSTAND THE COMPLEX DYNAMICS OF MOTOR PROTEINS USING SIMPLE STOCHASTIC MODELS? Motor Proteins Enzymes that convert the chemical energy into mechanical
Supplementary Figures for Tong et al.: Structure and function of the intracellular region of the plexin-b1 transmembrane receptor
 Supplementary Figures for Tong et al.: Structure and function of the intracellular region of the plexin-b1 transmembrane receptor Figure S1. Plexin-B1 GAP segments are homologous to RasGAPs. Sequence alignment
Supplementary Figures for Tong et al.: Structure and function of the intracellular region of the plexin-b1 transmembrane receptor Figure S1. Plexin-B1 GAP segments are homologous to RasGAPs. Sequence alignment
NGF - twenty years a-growing
 NGF - twenty years a-growing A molecule vital to brain growth It is twenty years since the structure of nerve growth factor (NGF) was determined [ref. 1]. This molecule is more than 'quite interesting'
NGF - twenty years a-growing A molecule vital to brain growth It is twenty years since the structure of nerve growth factor (NGF) was determined [ref. 1]. This molecule is more than 'quite interesting'
Multiscale Structural Mechanics of Viruses: Stretching the Limits of Continuum Modeling
 Multiscale Structural Mechanics of Viruses: Stretching the Limits of Continuum Modeling William S. Klug Mechanical & Aerospace Engineering Department Acknowledgements: Melissa Gibbons, Lin Ma, Chuck Knobler,
Multiscale Structural Mechanics of Viruses: Stretching the Limits of Continuum Modeling William S. Klug Mechanical & Aerospace Engineering Department Acknowledgements: Melissa Gibbons, Lin Ma, Chuck Knobler,
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA. President, Association of Women in Science, Bethesda Chapter STEM Consultant
 CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar.
 Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Why Proteins Fold? (Parts of this presentation are based on work of Ashok Kolaskar) CS490B: Introduction to Bioinformatics Mar. 25, 2002 Molecular Dynamics: Introduction At physiological conditions, the
Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality
 JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 6 8 AUGUST 2003 Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality Sanghyun Park and Fatemeh Khalili-Araghi Beckman
JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 6 8 AUGUST 2003 Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality Sanghyun Park and Fatemeh Khalili-Araghi Beckman
Notes Chapter 4 Cell Reproduction. That cell divided and becomes two, two become, four become eight, and so on.
 Notes Chapter 4 Cell Reproduction 4.1 Cell Division and Mitosis Many organisms start as. That cell divided and becomes two, two become, four become eight, and so on. Many-celled organisms, including you,
Notes Chapter 4 Cell Reproduction 4.1 Cell Division and Mitosis Many organisms start as. That cell divided and becomes two, two become, four become eight, and so on. Many-celled organisms, including you,
BMB 170 Lecture 13 Nucleic Acids, Nov 7th
 BMB 170 Lecture 13 Nucleic Acids, Nov 7th Chloroplast Ribosome h"p://www.bangroup.ethz.ch/research/nomenclature-of-ribosomal-proteins.html Ban N, Beckmann R, Cate JH, Dinman JD, Dragon F, Ellis SR, Lafontaine
BMB 170 Lecture 13 Nucleic Acids, Nov 7th Chloroplast Ribosome h"p://www.bangroup.ethz.ch/research/nomenclature-of-ribosomal-proteins.html Ban N, Beckmann R, Cate JH, Dinman JD, Dragon F, Ellis SR, Lafontaine
Name: Class: Date: ID: A
 Class: Date: Ch 7 Review Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. Researchers use fluorescent labels and light microscopy to a. follow
Class: Date: Ch 7 Review Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. Researchers use fluorescent labels and light microscopy to a. follow
Multiple Choice Identify the letter of the choice that best completes the statement or answers the question.
 chapter 7 Test Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. Who was one of the first people to identify and see cork cells? a. Anton van
chapter 7 Test Multiple Choice Identify the letter of the choice that best completes the statement or answers the question. 1. Who was one of the first people to identify and see cork cells? a. Anton van
Chemistry Courses. Courses. Chemistry Courses 1
 Chemistry Courses 1 Chemistry Courses Courses CHEM 1105. Laboratory for CHEM 1305.. Prerequisite(s): (CHEM 1305 w/d or better) CHEM 1106. Laboratory for CHEM 1306.. Prerequisite(s): (CHEM 1306 w/d or better)
Chemistry Courses 1 Chemistry Courses Courses CHEM 1105. Laboratory for CHEM 1305.. Prerequisite(s): (CHEM 1305 w/d or better) CHEM 1106. Laboratory for CHEM 1306.. Prerequisite(s): (CHEM 1306 w/d or better)
Artificial Intelligence (AI) Common AI Methods. Training. Signals to Perceptrons. Artificial Neural Networks (ANN) Artificial Intelligence
 Artificial Intelligence (AI) Artificial Intelligence AI is an attempt to reproduce intelligent reasoning using machines * * H. M. Cartwright, Applications of Artificial Intelligence in Chemistry, 1993,
Artificial Intelligence (AI) Artificial Intelligence AI is an attempt to reproduce intelligent reasoning using machines * * H. M. Cartwright, Applications of Artificial Intelligence in Chemistry, 1993,
Large Macromolecular Assemblies (LMAs) Large Macromolecular Assemblies (LMAs) Large Macromolecular Assemblies (LMAs)
 Large Macromolecular Assemblies (LMAs) : an Automated Computational Tool for Identifying Conserved Domains in the Structures of Macromolecular Assemblies Large macromolecular assemblies (such as ribosomes,
Large Macromolecular Assemblies (LMAs) : an Automated Computational Tool for Identifying Conserved Domains in the Structures of Macromolecular Assemblies Large macromolecular assemblies (such as ribosomes,
