Geoglossomycetes cl. nov., Geoglossales ord. nov. and taxa above class rank in the Ascomycota Tree of Life
|
|
|
- Austin Asher Clarke
- 6 years ago
- Views:
Transcription
1 Persoonia 22, 2009: RESEARCH ARTICLE doi: / x Geoglossomycetes cl. nov., Geoglossales ord. nov. and taxa above class rank in the Ascomycota Tree of Life C.L. Schoch 1, Z. Wang 2, J.P. Townsend 2, J.W. Spatafora 3 Key words Bayesian inference hybrid classification maximum likelihood Abstract Featuring a high level of taxon sampling across Ascomycota, we evaluate a multi-gene phylogeny and propose a novel order and class in Ascomycota. We describe two new taxa, Geoglossomycetes and Geoglossales, to host three earth tongue genera: Geoglossum, Trichoglossum and Sarcoleotia as a lineage of Leotiomyceta. Correspondingly, we confirm that these genera are not closely related to the genera Neolecta, Mitrula, Cudonia, Microglossum, Thuemenidum, Spathularia and Bryoglossum, all of which have been previously placed within the Geoglossaceae. We also propose a non-hierarchical system for naming well-resolved nodes, such as Saccharomyceta, Dothideomyceta, and Sordariomyceta for supraordinal nodes, within the current phylogeny, acting as rankless taxa. As part of this revision, the continued use of Leotiomyceta, now as a rankless taxon, is proposed. Article info Received: 16 April 2009; Accepted: 11 May 2009; Published: 5 June INTRODUCTION The multi-gene sequence datasets generated by the research consortium Assembling the Fungal Tree of Life (AFTOL) have resulted in several multi-gene phylogenies incorporating comprehensive taxon sampling across Fungi (Lutzoni et al. 2004, Blackwell et al. 2006, James et al. 2006). AFTOL generated a data matrix spanning all currently accepted classes in the Ascomycota, the largest fungal phylum. The phylogenies produced by AFTOL prompted the proposal of a phylogenetic classification from phylum to ordinal level in fungi (Hibbett et al. 2007). Although the Botanical Code does not require the principle of priority in ranks above family (McNeill et al. 2006), this principle was nevertheless followed for all taxa. The following ranked taxa were defined: subkingdom, phylum (suffix -mycota, except for Microsporidia), subphylum (-mycotina), class (-mycetes), subclass (-mycetidae) and order (-ales). As in Hibbett et al. (2007), several phylogenetically well-supported nodes above the rank of order could not be accommodated in the current hierarchical classification system based on the International Code of Botanical Nomenclature. To remedy this deficiency, rankless (or unranked) taxa for unambiguously resolved nodes with strong statistical support was proposed (Hibbett & Donoghue 18). Hybrid classifications that include both rankless and Linnaean taxa have since been discussed elsewhere (Jørgensen 2002, Kuntner & Agnarsson 2006), and applied to diverse organisms from lichens (Stenroos et al. 2002) and plants (Sennblad & Bremer 2002, Pfeil & Crisp 2005) to spiders (Kuntner 2006). These studies all attempt to create a comprehensive code for phylogenetic nomenclature that retains the current Linnean hierarchical codes. In keeping with the practice of previous hybrid classifications, we propose to use names corresponding to clades of higher taxa that were resolved in this phylogeny as well as preceding studies. 1 National Center for Biological Information (GenBank), National Library of Medicine, National Institute of Health, 45 Center Drive, MSC 6510, Bethesda, Maryland , USA; corresponding author schoch2@ncbi.nlm.nih.gov 2 Department of Ecology and Evolutionary Biology, Yale University, 165 Prospect Street, New Haven, CT 06520, USA. 3 Department of Botany and Plant Pathology, Oregon State University, Corvallis, OR 97331, USA. The proposed informal, rankless names for well-supported clades above the class level in our phylogeny agrees with the principles of the Phylocode ( It is our hope that such names should function as rankless taxa, facilitating the naming of additional nodes/clades as they become resolved. Eventual codification will follow the example of Hibbett et al. (2007) by applying principles of type names and priority. A number of published manuscripts already provide background on other supraordinal relationships of Fungi; for more complete treatments of the various classes, see Blackwell et al. (2006). During the AFTOL project a data matrix was generated spanning all currently accepted classes in the Ascomycota, the largest fungal phylum. A multi-gene phylogeny was recently inferred from these data, demonstrating relevant patterns in biological and morphological character development as well as establishing several distinct lineages in Ascomycota (Schoch et al. 2009). Here we test whether the relationships reported in Schoch et al. (2009) remain valid by applying both maximum likelihood (ML) and Bayesian analyses on a more restricted but denser set of taxa, including expanded sampling in the Geoglossaceae. We will therefore address the taxonomic placement of a group of fungi with earth tongue morphologies that are shown to be unrelated to other known classes. This morphology is closely associated with the family Geoglossaceae (Corda 1838). With typical inoperculate asci and an exposed hymenium, Geoglossaceae has long been thought to be a member of Leotiomycetes, though the content of the family itself has experienced many changes (Nannfeldt 1942, Korf 1973, Spooner 17, Platt 2000, Wang et al. 2006a, b). It is currently listed with 48 species and 6 genera in the Dictionary of the Fungi (Kirk et al. 2008). Several analyses using molecular data supported a clade including three earth tongue genera, Geoglossum, Trichoglossum and Sarcoleotia (Fig. 1), and cast doubt upon their positions in Leotiomycetes (Platt 2000, Gernandt et al. 2001, Lutzoni et al. 2004, Sandnes 2006, Spatafora et al. 2006, Wang et al. 2006b). Here we present a comprehensive phylumwide phylogeny, including data from protein coding genes. We can confidently place the earth tongue family as separate from currently accepted classes in Ascomycota Nationaal Herbarium Nederland & Centraalbureau voor Schimmelcultures You are free to share - to copy, distribute and transmit the work, under the following conditions: Attribution: You must attribute the work in the manner specified by the author or licensor (but not in any way that suggests that they endorse you or your use of the work). Non-commercial: You may not use this work for commercial purposes. No derivative works: You may not alter, transform, or build upon this work. For any reuse or distribution, you must make clear to others the license terms of this work, which can be found at Any of the above conditions can be waived if you get permission from the copyright holder. Nothing in this license impairs or restricts the author s moral rights.
2 130 Persoonia Volume 22, Dothideomyceta Dothideomycetes Arthoniomycetes Eurotiomycetes Lecanoromycetes Lichinomycetes Sordariomycetes Laboulbeniomycetes A Leotiomycetes Geoglossomycetes Monilinialaxa Botryotinia fuckeliana 5380 Lambertella subrenispora 72 Mitrula elegans Mitrula brevispora Cudoniella clavus Lachnum virgineum 90 Bryoglossum gracile SordarioNeofabrea malicorticis myceta Dermea acerina Microglossum olivaceum Leotia lubrica Thuemenidium atropurpureum 1 Thuemenidium atropurpureum 2 Leotio52 myceta 84 Microglossum rufum 1 A Microglossum rufum 2 61 Cudonia circinans Spathularia velutipes 1 B Spathularia velutipes 2 Neobulgaria pura 58 Sarcoleotia globosa 1 Sarcoleotia globosa 2 C Sarcoleotia globosa 3 Trichoglossum hirsutum 1 PezizoD mycotina Trichoglossum hirsutum 2 Geoglossum umbratile Saccharo 88 Geoglossum nigritum E myceta Geoglossum glabrum Asco- B C D Orbiliomycetes Pezizomycetes mycota substitutions per site Saccharomycotina Saccharomycetes Taphrinomycotina Taphrinomycetes E Neolectomycetes Basidiomycota Schizosaccharomycetes Fig. 1 A most likely tree obtained by RAxML for Ascomycota. Subphyla, class and rankless taxa are indicated. Classes containing fungi designated as earth tongues are indicated in black. The tree was rooted with outgroup Rhizopus oryzae (not shown). Bootstrap values are shown in orange and Bayesian posterior probabilities in blue. Orange, bold branches are supported by more than 80 % bootstrap and 95 % posterior probability, respectively. The full phylogeny, without collapsed clades, are shown in Fig. 2. The inset figures illustrate morphological ascomal diversity in the earth tongues. The species are as follows: A. Trichoglossum hirsutum; B. Geoglossum nigritum; C. Microglossum rufum; D. Spathularia velutipes; E. Geoglossum nigritum. Photo credits: A: Zhuliang Yang; B, D, E: Kentaro Hosaka; C: Dan Luoma. MATERIALS AND METHODS Data were extracted from the complete data matrix obtained from the WASABI database ( incorporating representatives for all currently accepted classes, and maximizing the number of orders and available data. Following the approach of James et al. (2006) we performed a combined analysis, with both DNA and amino acid data, while allowing for missing data. This data was supplemented with additional ribosomal sequences from earth tongue genera obtained and deposited in GenBank from two previous studies (Wang et al. 2006a, b). To further minimise poorly aligned areas, 219 additional columns, which proved variable when viewed in BioEdit with a 40 % shade threshold, were excluded from the original AFTOL inclusion set. The refined dataset consisted of 161 taxa (including outgroups) and characters for six different loci: the nuclear small and large ribosomal subunits (nssu, nlsu), the mitochondrial small ribosomal subunit (mssu) and fragments from three proteins: transcription elongation factor 1 alpha (TEF1) and the largest and second largest subunits of
3 C.L. Schoch et al.: Geoglossomycetes 131 RNA polymerase II (RPB1, RPB2). A complete table with the published GenBank numbers is listed in Table 1. The phylogenetic analysis was run in RAxML v7.0.0 (Stamatakis 2006), partitioning by gene (six partitions) and estimating unique model parameters for each gene, as in Schoch et al. (2009). Models of evolution were evaluated as in Schoch et al. (2009) with the same models selected. For DNA sequences, this resulted in a general time reversible model (GTR) with a discrete gamma distribution composed of four rate classes plus an estimation of the proportion of invariable sites. The amino acid sequences were analysed with a RTREV model with similar accommodation of rate heterogeneity across sites and proportions of invariant sites. In addition, protein models for TEF1 and RPB2 incorporated a parameter to estimate amino acid frequencies. The tree shown in Fig. 1 was obtained by using an option in RAxML running a rapid bootstrap analysis and search for the best-scoring ML tree in one single run. This meant the GTRCAT model approximation was used, which does not produce likelihood values comparable to other programs. The full tree is shown here as Fig. 2 and was deposited in TreeBASE ( We also ran repetitions of RAxML under a gamma rate distribution option. The best scoring tree was included in TreeBASE. A second analysis was run using Bayesian inference of maximum likelihood in MrBayes v3.1.2 (Huelsenbeck & Ronquist 2001, Altekar et al. 2004) using models and parameters that were comparable to the maximum likelihood run. Data were similarly partitioned and amino acids were analysed, so that a mixture of models with fixed rate matrices for amino acid sequences could be evaluated. In all cases rate heterogeneity parameters were used by a discrete gamma distribution plus an estimation of the proportion of invariable sites. A metropolis coupled Markov Chain Monte Carlo analysis was run for 9 million generations sampling every 200th cycle, starting from a random tree and using 4 chains (three heated and one cold) under default settings. Two separate runs were confirmed to converge using Tracer v1.4.1 ( tracer/). The first sampled trees (2 million generations) were removed as burn in each run. A 50 % majority rule consensus tree of Bayesian likelihood trees from the two combined runs was subsequently constructed, and average branch lengths and posterior probabilities determined. The numbers of nodes shared with the most likely tree in Fig. 1 was determined and plotted on the branches. This tree was deposited in TreeBASE, along with the inclusive character set. RESULTS The phylogeny presented in Fig. 1 supports 15 classes (11 in Pezizomycotina, 1 in Saccharomycotina, 3 in Taphrinomycotina) with good statistical support (both ML bootstrap and Bayesian posterior probability) for 14. Phylogenies with all lineages in the analysed data matrix are included in Fig. 2. A run with repetitions of RAxML under a gamma rate distribution option resulted in a best scoring tree with a log likelihood of This tree shared the same supported nodes with the one presented in Fig. 1 but had changes in poorly supported nodes regarding placement of the Eurotiomycetes and Dothideomycetes. The two Bayesian runs produced trees with harmonic means of likelihood values of and , respectively, with similar topological differences in poorly supported nodes. As can be seen in Fig. 1, we continue to find low bootstrap and posterior probability support for Leotiomycetes as a monophyletic clade using a combined analysis of protein and nucleic acids. In our analysis, this includes Neobulgaria pura as the earliest diverging lineage. The node internal from this lineage is found in all ML bootstrap trees, suggesting that this taxon is unstable in our analyses. No conflicts were detected in Neobulgaria genes under a previous study and missing data did not affect important nodes (Schoch et al. 2009). A repeat run under maximum likelihood was done with Neobulgaria pura removed under the same settings but with only bootstrap repetitions. This trimmed dataset yielded a congruent phylogeny with increased bootstrap for Leotiomycetes (78 %; data not shown). The instability of the placement of Neobulgaria pura does not compromise any of the conclusions we present here and may be due to various reasons. Improved taxon sampling will likely help to resolve its placement in future analyses. We find support for numerous backbone nodes in Ascomycota, as did Schoch et al. (2009). Our phylum-wide sampling of Ascomycota classes in this study, combined with the results of a previous study (Schoch et al. 2009), facilitated addressing the placement of the previously problematic and unsampled lineages such as the Geoglossaceae in relation to all currently accepted Ascomycota classes. Taxonomy Given their unique ascomatal development, ultrastructure of ascus apical apparatus, mossy habitat, and our multilocus gene phylogeny, Geoglossomycetes cl. & ord. nov. is justified here as incertae sedis in Pezizomycotina and Leotiomyceta. Geoglossomycetes, Geoglossales Zheng Wang, C.L. Schoch & Spatafora, cl. & ord. nov. MycoBank MB513351, MB Ascomata solitaria vel gregaria, capitata, stipitata; stipe cylindricus, atrum, glabrum vel furfuraceus. Regio hymeniali capitata, clavata vel pileata, indistinctum ex stipite; hymenium atrum, continuatcum stipite ad praematuro incrementi grado. Asci clavati, inoperculati, octospori, poro parvo in iodo caerulescentes. Ascosporae elongatae, fuscae, pullae vel hyalinae, multiseptatae. Paraphyses filiformes, pullae vel hyalinae. Distributio generalis, terrestris, habitaile locus fere uliginoso et muscoso. Type genus. Geoglossum Pers., Neues Mag. Bot. 1: ; Geoglossaceae. Ascomata scattered to gregarious, capitate, stipitate; stipe cylindrical, black, smooth to furfuraceous. Ascigerous portion capitate, club-shaped to pileate, indistinguishable from stipe. Hymenium surface black, continues with stipe at early development stage. Asci clavate, inoperculate, thin-walled, J+, usually 8-spored. Ascospores elongate, dark-brown, blackish to hyaline, septate when mature. Paraphyses filiform, blackish to hyaline. Global distribution, terrestrial, habitat usually boggy and mossy. DISCUSSION In keeping with the phylogeny presented in Fig. 1, we endorse use of the -myceta suffix in order to circumscribe well-supported clades above class. The numbers of these clades are limited, and the use of such taxa will continue to become more practical as our biological knowledge base broadens. Use of this suffix will also allow for the continued use of Leotiomyceta, a taxon that has already been defined with a Latin diagnosis provided as a ranked superclass (Eriksson & Winka 17) and remains in use (Lumbsch et al. 2005, Wang et al. 2006a). We propose its continued use, but as a rankless taxon together with the newly proposed rankless taxa, Saccharomyceta, Dothideomyceta and Sordariomyceta. Since these taxa are not currently accepted under the Code (McNeill et al. 2006), we will refrain from formal designations. The relevant clades are discussed below with the informal designations indicated in single quotations.
4 Outgroup Rhizopus oryzae (not shown) 132 Persoonia Volume 22, Epichloë typhina Claviceps purpurea Elaphocordyceps capitata Elaphocordyceps ophioglossoides Cordyceps cardinalis Nectria cinnabarina Hypocrea lutea Bionectria cf. aureofulva Melanospora tiffanyae Melanosporales Coronophorales Bertia moriformis Gondwanamyces capensis Ceratocystis fimbriata Halosphaeria appendiculata Corollospora maritima Glomerella cingulata Verticillium dahliae Cryptosporella hypodermia Gnomonia gnomon Endothia gyrosa Diaporthe eres Calosphaeria pulchella Magnaporthe grisea Calosphaeriales Magnaporthales Gaeumannomyces medullaris Ophiostoma stenoceras Ophiostomatales Sordaria fimicola Sordariales Neurospora crassa Xylaria hypoxylon Xylariales Lindra thalassiae Lulworthia grandispora Stigmatomyces protrudens Hesperomyces virescens Pyxidiophora avernensis Botryotinia fuckeliana Monilinia laxa Lambertella subrenispora Mitrula brevispora Mitrula elegans Bryoglossum gracile Lachnum virgineum Cudoniella cf. clavus Dermea acerina Neofabraea malicorticis Microglossum rufum 2 Microglossum rufum 1 Thueminidium atropurpureum 1 Thueminidium atropurpureum 2 Leotia lubrica Microglossum olivaceum Spathularia velutipes 1 Spathularia velutipes 2 Cudonia circinans Neobulgaria pura Geoglossum glabrum Geoglossum nigritum Geoglossum umbratile Trichoglossum hirsutum 1 Trichoglossum hirsutum 2 Sarcoleotia globosa 2 Sarcoleotia globosa 3 Sarcoleotia globosa 1 Arthrobotrys elegans Orbilia vinosa l l Aleuria aurantia Scutellinia scutellata Pyronema domesticum Eleutherascus lectardii Sarcosoma latahense Gyromitra californica Verpa conica Caloscypha fulgens Ascobolus carbonarius Saccobolus dilutellus Peziza vesiculosa Sarcosphaera crassa Saccharomyces cerevisiae Candida glabrata Eremothecium gossypii Candida tropicalis Debaryomyces hansenii Taphrina wiesneri Taphrina deformans Taphrinales Saitoella complicata Schizosaccharomyces pombe Neolecta vitellina Neolecta irregularis Fomitopsis pinicola Grifola frondosa Calostoma cinnabarinum Calocera cornea Cryptococcus neoformans Geoglossomycetes, cl. nov. Geoglossales, ord. nov. Orbiliomycetes Orbiliales Taphrinomycetes Leotiomycetes Helotiales Rhytismatales Pezizomycetes Pezizales Hypocreales Microascales Diaporthales Schizosaccharomycetes, Schizosaccharomycetales Neolectomycetes Neolectales Lulworthiales Laboulbeniales Pyxidiophorales Saccharomycetes Saccharomycetales Dothideomyceta Leotiomyceta Sordariomyceta Pezizomycotina Saccharomycotina Taphrinomycotina 0.1 substitutions per site Chaetothyriales Ascomycota Saccharomyceta Sordariomycetes Pleospora herbarum Cochliobolus heterostrophus Allewia eureka Pyrenophora phaeocomes Leptosphaeria maculans Phoma herbarum Ascochyta pisi Sporormiella minima Westerdykella cylindrica Pleomassaria siparia Lepidosphaeria nicotiae Ulospora bilgramii Verruculina enalia Delitschia winteri Gloniopsis smilacis Hysterographium mori Hysterium angustatum 42 Mytilinidion australe 40 Mytilinidion rhenanum Lophium mytilinum Botryosphaeria tsugae Botryosphaeria dothidea Guignardia gaultheriae Guignardia bidwellii Tubeufia paludosa Helicomyces roseus Mycosphaerella punctiformis Mycosphaerella graminicola Mycosphaerella fijiensis Capnodium coffeae Scorias spongiosa Cladosporium cladosporioides Davidiella tassiana Lobaria scrobiculata Lobariella pallida Peltigera degenii Nephroma parile Coccocarpia erythroxyli Lecidea fuscoatra Hypogymnia physodes Canoparmelia caroliniana Pyxine subcinerea Heterodermia vulgaris Teloschistes exilis Diploschistes ocellatus Glomerobolus gelineus Trapelia placodioides Icmadophila ericetorum Pertusaria dactylina Ceramothyrium carniolicum Cyphellophora laciniata Exophiala salmonis Staurothele frustulenta Agonimia sp. Coccidioides immitis Ajellomyces capsulatum Aspergillus protuberus Caliciopsis orientalis Peltula umbilicata Peltula auriculata Lichinella iodopulchra Hysteriales Mytilinidiales Botryosphaeriales Dothiora cannabinae Dothidea sambuci Dothideales Sydowia polyspora Myriangium duriaei Myriangiales Elsinoë veneta Schismatomma decolorans Roccella fuciformis Roccellographa cretacea Peltigerales Lecanorales Capnodiales Teloschistales Ostropales Agyriales Pertusariales Onygenales Eupenicillium limosum Eurotiales Coryneliales Lichinomycetes Lichinales Pleosporales Arthoniomycetes Arthoniales Lecanoromycetes Basidiomycota Laboulbeniomycetes Verrucariales Fig. 2 A most likely tree obtained by RAxML for Ascomycota (as in Fig. 1). Phyla, subphyla, class, order and rankless taxa are indicated. Taxa designated as earth tongues are indicated in orange. The tree is displayed as two subtrees orange arrows indicate where the subtrees were joined. The tree was rooted with outgroup Rhizopus oryzae (not shown). Bootstrap values are shown in orange above nodes and Bayesian posterior probabilities in blue below. Numbers were removed for nodes with % bootstrap and % posterior probability. Eurotiomycetes Dothideomycetes
5 C.L. Schoch et al.: Geoglossomycetes 133 Subphylum Taphrinomycotina As in recent studies using large multi-gene datasets (Spatafora et al. 2006, Sugiyama et al. 2006, Liu et al. 2009, Schoch et al. 2009), we find ML bootstrap support here for the monophyly of the Taphrinomycotina. The addition of sequences from protein coding genes has been vital to the establishment of statistical support for this grouping. Recent work has shown that the short generation times characteristic of species in this group make phylogenetic analyses particularly susceptible to long branch attraction artefacts (Liu et al. 2009). The placement of Neolecta in this subclade is also confirmed here. The club-shaped apothecia of the members of Neolecta share superficial similarity with those of the Geoglossaceae. Neolecta was long thought to be included in the Geoglossaceae until molecular work proved otherwise (Landvik 16). In support of its placement in this early diverging group, Neolecta has several presumably ancestral features, such as simplified non-poricidal asci without croziers and the absence of paraphyses (Redhead 1979, Landvik et al. 2003). With additional sampling of both taxa and genes we find here moderate support for the monophyly of Taphrinomycotina, and thus demonstrate that the earliest diverging clade of the Ascomycota was dimorphic, with both filamentous and yeast growth forms. Nevertheless, it remains apparent that this part of the Ascomycota tree remains under sampled. This lack of adequate sampling is supported by the recent description of a clade labelled Soil Clone Group I (SCGI). SCGI is ubiquitous in soil and is only known from environmental sequence data (Porter et al. 2008). It appears possible that they form a novel early diverging lineage outside of Taphrinomycotina. Very little remains known about their ecology, morphology and general biology. Rankless taxon Saccharomyceta Saccharomyceta includes the two remaining subphyla of Ascomycota, Saccharomycotina and Pezizomycotina. Saccharomycotina comprises the true yeasts (e.g., Saccharomyces cerevisiae), although hyphal growth has been documented in some taxa (e.g., Eremothecium). The Pezizomycotina consists of the majority of filamentous, ascoma producing species, but numerous species are additionally capable of yeast and yeastlike growth phases. Thousands of species are only known to reproduce asexually. These two subphyla form a well-supported, monophyletic group that has been recovered in a large number of studies across a diversity of character and taxon sets. The recognition of Saccharomyceta highlights the shared common ancestry of these two taxa and the inaccurate characterisation of Saccharomycotina as a primitive or basal lineage of the Ascomycota. Rather, its small genome size (Dujon et al. 2004) and dominant yeast growth phase can be characterized as derived traits for this subphylum. Rankless taxon Leotiomyceta We apply Leotiomyceta as a rankless taxon containing the majority of fungi with a diversity of inoperculate asci (e.g., fissitunicate, poricidal, deliquescent). Leotiomyceta excludes the earliest diverging classes of Pezizomycotina, Pezizomycetes and Orbiliomycetes. It was first defined as a superclass (Eriksson & Winka 17). This definition has remained in use (Lumbsch et al. 2005, Spatafora et al. 2006). Included in this clade are the informal, rankless taxa Dothideomyceta, Sordariomyceta, as well as the classes Eurotiomycetes, Lecanoromycetes, Lichinomycetes, and a newly proposed class, Geoglossomycetes. The type genus of Geoglossaceae, Geoglossum was initially proposed by Persoon (1794). Persoon described it as clubshaped, with unitunicate, inoperculate asci, with the type species given as Geoglossum glabrum Pers. Trichoglossum have historically been classified in Geoglossaceae, and Sarcoleotia has historically been classified in the Helotiaceae (Leotiomycetes). These inoperculate Discomycetes produce terrestrial, stipitate, clavate ascomata, commonly referred to as earth tongues, which include Leotia, Microglossum, Cudonia, and Spathularia. In terms of ascomatal development, species of Geoglossum, Trichoglossum, and Sarcoleotia possess a hymenium that freely develops towards the base, while other earth tongue fungi feature a distinct ridge to their hymenium, implying a developmental stage during which the hymenium is enclosed (Schumacher & Sivertsen 17, Spooner 17, Wang et al. 2006b). An enclosed hymenium has been observed as well in several other lineages, such as Cyttaria, Erysiphales and Rhytismatales in the Leotiomycetes (Korf 13, Gargas et al. 15, Johnston 2001). Although the name earth tongue implies these fungi are terrestrial and have no direct association found with other organisms, Trichoglossum, Geoglossum and Sarcoleotia globosa have often been recorded in boggy habitats abundant with bryophytes (Seaver 1951, Dennis 1968, Schumacher & Sivertsen 17, Spooner 17, Jumpponen et al. 17, Zhuang 18). Ascus apical morphology is one of the major features in distinguishing higher ascomycetes, and operculate ascomycetes as members of Pezizales have an apical or subapical operculum which is thrown back at spore discharge while a definite plug is present in the thickened inoperculate ascus apex as in species of the Helotiales (Korf 1973). Ultrastructure of the ascus apical apparatus suggested no close relationship between Leotia lubrica and species of Geoglossum and Trichoglossum. A structure known as a tractus connects the uppermost spore to the apical wall and the spores to each other in Trichoglossum hirsutum, but is never found in other species of the Helotiales and is possibly homologous to structures in Sordariomycetes and Pezizomycetes (Verkley 14). Recent molecular phylogenetic analyses (Sandnes 2006, Wang et al. 2006a, b) confirmed that the earth tongue fungi are not monophyletic. At least two origins occurred in Leotiomycetes: in Leotia and allies in Helotiales, and in Cudonia and allies in Rhytismatales. Geoglossum, Trichoglossum. Sarcoleotia (Geoglossomycetes as we define it) represent a third, independent lineage of earth tongues, which we confirmed does not belong within the Leotiomycetes. DNA-only and combined model analyses produced conflicting placements of Geoglossaceae within Pezizomycotina. Previous analyses applying nucleotide sequences only placed the order as a sister group to the Lichinomycetes (Lutzoni et al. 2004, Spatafora et al. 2006), which includes a small number of lichenised species mainly associated with cyanobacteria (Reeb et al. 2004). Our sampling of Lichinomycetes includes two genera, Peltula and Lichinella that encompass at least some of the ascal diversity, i.e., rostrate and deliquescent, present in the class. In contrast, our combined amino acid and nucleotide model analyses resolved Geoglossaceae as an isolated, unique lineage of Leotiomyceta with no supported sister relationship, in agreement with Schoch et al. (2009). Different levels of missing data underlie these two conflicting topologies, and several phenomena can potentially explain this conflict, ranging from model misspecification to long-branch attraction. Regardless of these concerns, our conclusion that the Geoglossaceae is a monophyletic lineage, unallied with members of the Leotiomycetes and any of the other large fungal classes remains strongly supported. Eurotiomycetes and Lecanoromycetes are the two remaining classes in Leotiomyceta. Eurotiomycetes is arguably the most ecologically diverse class within Ascomycota including lichenised species, saprobes and pathogens of animals and plants. As currently defined, this class incorporates several distinct orders and three subclasses spanning virtually all known fun-
6 134 Persoonia Volume 22, 2009 Table 1 Taxa and sequences used in this study. AFTOL Class Order Voucher 1 Taxon nssu nlsu mssu RPB1 RPB2 TEF1 no Zygomycota outgroup GB Rhizopus oryzae AF AY AY Genome Genome Genome 438 Basidiomycota outgroup GEL 5359 Calocera cornea AY AY AY8570 AY AY Basidiomycota outgroup AW 136 Calostoma cinnabarinum AY AY AY AY AY Basidiomycota outgroup GB Cryptococcus neoformans Genome Genome XM_ XM_ Genome 770 Basidiomycota outgroup MB Fomitopsis pinicola AY AY FJ AY AY AY Basidiomycota outgroup DSH s.n. Grifola frondosa AY AY AY AY AY Arthoniomycetes Arthoniales Diederich Roccella fuciformis AY AY EU DQ DQ Arthoniomycetes Arthoniales BG Printzen11 Roccellographa cretacea DQ DQ FJ DQ DQ Arthoniomycetes Arthoniales DUKE Schismatomma decolorans AY AY AY DQ DQ DQ Dothideomycetes Botryosphaeriales CBS Botryosphaeria dothidea DQ6778 DQ FJ EU DQ DQ Dothideomycetes Botryosphaeriales CBS Botryosphaeria tsugae AF DQ DQ DQ Dothideomycetes Botryosphaeriales CBS Guignardia bidwellii DQ DQ DQ Dothideomycetes Botryosphaeriales CBS Guignardia gaultheriae DQ FJ DQ Dothideomycetes Capnodiales CBS Capnodium coffeae DQ DQ FJ DQ DQ DQ Dothideomycetes Capnodiales CBS Cladosporium cladosporioides DQ DQ FJ EU DQ DQ Dothideomycetes Capnodiales CBS 3.80 Davidiella tassiana DQ DQ DQ DQ Dothideomycetes Capnodiales OSC 622 Mycosphaerella fijiensis DQ DQ6780 FJ DQ Dothideomycetes Capnodiales CBS 2.38 Mycosphaerella graminicola DQ DQ DQ Dothideomycetes Capnodiales CBS Mycosphaerella punctiformis DQ DQ FJ DQ DQ4700 DQ Dothideomycetes Capnodiales CBS Scorias spongiosa DQ DQ FJ DQ DQ Dothideomycetes Dothideales DAOM Dothidea sambuci AY AY AY DQ DQ Dothideomycetes Dothideales CBS Dothiora cannabinae DQ4733 DQ4704 FJ DQ DQ DQ Dothideomycetes Dothideales CBS Sydowia polyspora DQ DQ FJ DQ DQ6778 Dothideomycetes Hysteriales CBS Gloniopsis smilacis FJ FJ FJ FJ Dothideomycetes Hysteriales EB 0324 Hysterium angustatum FJ FJ FJ FJ Dothideomycetes Hysteriales EB 0249 Hysterographium mori FJ FJ FJ Dothideomycetes Incertae sedis CBS Helicomyces roseus DQ DQ DQ6771 DQ Dothideomycetes Incertae sedis CBS Tubeufia paludosa DQ DQ DQ DQ Dothideomycetes Myriangiales CBS Elsinoë veneta DQ DQ FJ DQ Dothideomycetes Myriangiales CBS Myriangium duriaei AY DQ AY DQ DQ Dothideomycetes Mytilinidiales EB 0248 Lophium mytilinum FJ FJ FJ FJ Dothideomycetes Mytilinidiales CBS Mytilinidion australe FJ Dothideomycetes Mytilinidiales CBS Mytilinidion rhenanum FJ FJ FJ FJ Dothideomycetes Pleosporales DAOM Allewia eureka DQ6774 DQ DQ DQ Dothideomycetes Pleosporales CBS Ascochyta pisi var. pisi DQ DQ DQ DQ Dothideomycetes Pleosporales CBS Cochliobolus heterostrophus AY AY AY DQ DQ Dothideomycetes Pleosporales CBS Delitschia winteri DQ DQ FJ DQ DQ Dothideomycetes Pleosporales CBS Lepidosphaeria nicotiae DQ DQ DQ Dothideomycetes Pleosporales DAOM 2267 Leptosphaeria maculans DQ4703 DQ DQ DQ DQ Dothideomycetes Pleosporales CBS Phoma herbarum DQ DQ FJ DQ DQ Dothideomycetes Pleosporales CBS Pleomassaria siparia DQ DQ DQ DQ Dothideomycetes Pleosporales CBS Pleospora herbarum var. herbarum DQ DQ FJ DQ DQ DQ Dothideomycetes Pleosporales DAOM Pyrenophora phaeocomes DQ4595 DQ4596 FJ DQ DQ Dothideomycetes Pleosporales CBS Sporormiella minima DQ DQ FJ DQ DQ Dothideomycetes Pleosporales CBS 120 Ulospora bilgramii DQ DQ DQ DQ Dothideomycetes Pleosporales CBS Verruculina enalia DQ DQ DQ DQ Dothideomycetes Pleosporales CBS Westerdykella cylindrica AY AY AF DQ DQ4705 DQ Eurotiomycetes Chaetothyriales CBS Ceramothyrium carniolicum EF EF EF EF Eurotiomycetes Chaetothyriales CBS Cyphellophora laciniata EF EF Eurotiomycetes Chaetothyriales CBS Exophiala salmonis EF EF FJ EF EF EF Eurotiomycetes Coryneliales CBS Caliciopsis orientalis DQ DQ4707 FJ DQ DQ DQ Eurotiomycetes Eurotiales CBS Aspergillus protuberus FJ FJ FJ Eurotiomycetes Eurotiales CBS Eupenicillium limosum EF EF EF EF411070
7 C.L. Schoch et al.: Geoglossomycetes Eurotiomycetes Onygenales GB Ajellomyces capsulatum Genome Genome Genome Genome Genome 1084 Eurotiomycetes Onygenales TIGR Coccidioides immitis Genome Genome Genome Genome Genome 684 Eurotiomycetes Verrucariales NYBG Agonimia sp. DQ DQ DQ DQ DQ Eurotiomycetes Verrucariales DUKE Staurothele frustulenta DQ DQ8230 FJ DQ DQ Geoglossomycetes Geoglossales OSC Geoglossum glabrum AY AY Geoglossomycetes Geoglossales OSC 009 Geoglossum nigritum AY AY AY DQ DQ DQ Geoglossomycetes Geoglossales Mycorec1840 Geoglossum umbratile AY AY Geoglossomycetes Geoglossales HMAS Sarcoleotia globosa 1 AY78 AY78 Geoglossomycetes Geoglossales OSC Sarcoleotia globosa 2 AY Geoglossomycetes Geoglossales MBH Sarcoleotia globosa 3 AY Geoglossomycetes Geoglossales OSC 017 Trichoglossum hirsutum 1 AY AY AY DQ DQ DQ Geoglossomycetes Geoglossales OSC Trichoglossum hirsutum 2 AY AY Incertae sedis Incertae sedis IAM Saitoella complicata AY AY DQ AY DQ Laboulbeniomycetes Laboulbeniales GB Hesperomyces virescens AF2233 AF2235 Laboulbeniomycetes Laboulbeniales GB Stigmatomyces protrudens AF2232 AF Laboulbeniomycetes Pyxidiophorales CBS Pyxidiophora avernensis FJ FJ FJ FJ Lecanoromycetes Agyriales GB Trapelia placodioides AF AF AF DQ DQ DQ Lecanoromycetes Incertae sedis DUKE Lecidea fuscoatra DQ DQ DQ DQ DQ Lecanoromycetes Lecanorales DUKE Canoparmelia caroliniana AY AY AY DQ AY DQ Lecanoromycetes Lecanorales DUKE Hypogymnia physodes DQ DQ DQ DQ Lecanoromycetes Ostropales s.l. Lumbsch 5 Diploschistes ocellatus AF AY DQ DQ DQ Lecanoromycetes Ostropales s.l. JK 5548K Glomerobolus gelineus DQ DQ DQ DQ Lecanoromycetes Peltigerales DUKE Lobaria scrobiculata AY AY AY DQ DQ DQ Lecanoromycetes Peltigerales DUKE Lobariella pallida DQ DQ DQ DQ DQ DQ Lecanoromycetes Peltigerales DUKE Nephroma parile DQ DQ FJ Lecanoromycetes Peltigerales DUKE Peltigera degenii AY AY AY DQ AY DQ Lecanoromycetes Peltigerales DUKE Coccocarpia erythroxyli DQ DQ DQ DQ DQ DQ Lecanoromycetes Pertusariales DUKE Icmadophila ericetorum DQ DQ DQ6897 DQ DQ DQ Lecanoromycetes Pertusariales DUKE Pertusaria dactylina DQ DQ DQ DQ DQ DQ Lecanoromycetes Teloschistales DUKE Heterodermia vulgaris DQ DQ8837 DQ DQ DQ DQ Lecanoromycetes Teloschistales DUKE Pyxine subcinerea DQ DQ DQ9122 DQ DQ DQ Lecanoromycetes Teloschistales DUKE Teloschistes exilis AY AY FJ DQ DQ DQ Leotiomycetes Helotiales OSC 012 Botryotinia fuckeliana AY AY AY DQ DQ DQ Leotiomycetes Helotiales MBH Bryoglossum gracile AY AY Leotiomycetes Helotiales OSC 054 Cudoniella cf. clavus DQ4702 DQ FJ DQ DQ DQ Leotiomycetes Helotiales CBS Dermea acerina DQ DQ DQ DQ DQ DQ Leotiomycetes Helotiales OSC 002 Lachnum virgineum AY AY AY DQ DQ DQ Leotiomycetes Helotiales CBS Lambertella subrenispora DQ DQ DQ DQ DQ Leotiomycetes Helotiales OSC 001 Leotia lubrica AY AY AY DQ DQ DQ Leotiomycetes Helotiales FH-DSH Microglossum olivaceum AY AY Leotiomycetes Helotiales Ingo-Clark-Geo163 Microglossum rufum 1 DQ DQ Leotiomycetes Helotiales OSC 641 Microglossum rufum 2 DQ DQ4701 DQ DQ DQ Leotiomycetes Helotiales ZW Mitrula brevispora AY78 AY7893 Leotiomycetes Helotiales WZ-Geo47-Clark Mitrula elegans AY AY Leotiomycetes Helotiales OSC 063 Monilinia laxa AY AY AY FJ DQ DQ Leotiomycetes Helotiales CBS Neobulgaria pura FJ FJ FJ FJ Leotiomycetes Helotiales OSC 036 Neofabraea malicorticis AY AY AY DQ DQ DQ Leotiomycetes Helotiales 1803 Thueminidium atropurpureum 1 AY Leotiomycetes Helotiales Thueminidium atropurpureum 2 AY Leotiomycetes Rhytismatales DUKE Cudonia circinans AF AY AY AY Leotiomycetes Rhytismatales OSC 640 Spathularia velutipes 1 FJ7860 FJ7861 FJ7863 FJ7862 Leotiomycetes Rhytismatales ZW Geo58 Spathularia velutipes 2 AY AY Lichinomycetes Lichinales Schultz16319a Lichinella iodopulchra DQ DQ DQ Lichinomycetes Lichinales DUKE Peltula auriculata DQ DQ DQ DQ Lichinomycetes Lichinales DUKE Peltula umbilicata DQ DQ DQ2954 DQ DQ DQ Neolectomycetes Neolectales DAH-3 Neolecta irregularis DQ DQ4706 DQ Neolectomycetes Neolectales DAH-11 Neolecta vitellina DQ DQ4705 AAF19058
8 136 Persoonia Volume 22, 2009 Table 1 (cont.) AFTOL Class Order Voucher 1 Taxon nssu nlsu mssu RPB1 RPB2 TEF1 no Orbiliomycetes Orbiliales CBS Arthrobotrys elegans FJ FJ FJ FJ Orbiliomycetes Orbiliales CBS Orbilia vinosa DQ470 DQ DQ DQ Pezizomycetes Pezizales OSC 018 Aleuria aurantia AY5446 AY DQ DQ DQ Pezizomycetes Pezizales KH Ascobolus carbonarius AY AY FJ Pezizomycetes Pezizales OSC 062 Caloscypha fulgens DQ DQ2477 DQ DQ DQ Pezizomycetes Pezizales CBS Eleutherascus lectardii DQ DQ FJ DQ DQ DQ Pezizomycetes Pezizales OSC 068 Gyromitra californica AY AY AY DQ DQ DQ Pezizomycetes Pezizales TL-63 Peziza vesiculosa DQ4705 DQ DQ DQ4708 DQ Pezizomycetes Pezizales CBS Pyronema domesticum DQ DQ FJ DQ DQ DQ Pezizomycetes Pezizales CBS Saccobolus dilutellus FJ FJ FJ FJ FJ Pezizomycetes Pezizales CBS Sarcosoma latahense FJ FJ FJ FJ Pezizomycetes Pezizales OSC 049 Sarcosphaera crassa AY AY FJ Pezizomycetes Pezizales OSC 015 Scutellinia scutellata DQ DQ FJ DQ4735 DQ DQ Pezizomycetes Pezizales NRRL Verpa conica AY AY AY FJ Saccharomycetes Saccharomycetales GB Candida glabrata AY13 AY13 XM_ XM_ Genome 1269 Saccharomycetes Saccharomycetales GB Candida tropicalis M55527 Genome Genome Genome Genome 1077 Saccharomycetes Saccharomycetales GB Debaryomyces hansenii DHA AF4850 XM_4561 CR Genome 1072 Saccharomycetes Saccharomycetales GB Eremothecium gossypii AE AE AF NM_ AE Genome 1069 Saccharomycetes Saccharomycetales GB Saccharomyces cerevisiae SCYLR154C SCYLR154C AF X96876 SCYOR151C Genome 11 Schizosaccharomycetes Schizosaccharomycetales GB Schizosaccharomyces pombe X54866 Z19136 X54421 X56564 D13337 Genome 5086 Sordariomycetes Calosphaeriales CBS 1159 Calosphaeria pulchella AY AY FJ Sordariomycetes Coronophorales SMH4320 Bertia moriformis AY AY Sordariomycetes Diaporthales CBS Cryptosporella hypodermia DQ DQ DQ DQ Sordariomycetes Diaporthales CBS Diaporthe eres DQ AF FJ DQ DQ DQ Sordariomycetes Diaporthales CBS Endothia gyrosa DQ DQ DQ DQ4706 DQ Sordariomycetes Diaporthales CBS 1.53 Gnomonia gnomon DQ AF FJ DQ DQ4702 DQ Sordariomycetes Hypocreales GJS Bionectria cf. aureofulva DQ DQ FJ DQ DQ Sordariomycetes Hypocreales GAM Claviceps purpurea AF AF AY DQ AF Sordariomycetes Hypocreales OSC Cordyceps cardinalis AY AY EF DQ DQ DQ Sordariomycetes Hypocreales OSC Elaphocordyceps capitata AY AY FJ AY DQ AY Sordariomycetes Hypocreales OSC Elaphocordyceps ophioglossoides AY AY FJ AY DQ AY Sordariomycetes Hypocreales ATCC Epichloë typhina U32405 U17396 FJ DQ AF Sordariomycetes Hypocreales ATCC Hypocrea lutea AF AF FJ AY DQ AF Sordariomycetes Hypocreales CBS Nectria cinnabarina U32412 U00748 FJ AY DQ AF Sordariomycetes Incertae sedis FAU 553 Glomerella cingulata AF AF FJ AY DQ AF Sordariomycetes Incertae sedis ATCC Verticillium dahliae AY DQ FJ AY DQ AY Sordariomycetes Lulworthiales JK 5090A Lindra thalassiae DQ4704 DQ FJ DQ DQ DQ Sordariomycetes Lulworthiales JK 4686 Lulworthia grandispora DQ DQ FJ DQ DQ Sordariomycetes Magnaporthales JK 5528S Gaeumannomyces medullaris FJ FJ Sordariomycetes Magnaporthales Broad Magnaporthe grisea AB AB Genome Genome Genome Sordariomycetes Melanosporales ATCC Melanospora tiffanyae AY AY AY Sordariomycetes Microascales TCH C89 Ceratocystis fimbriata U32418 U17401 FJ Sordariomycetes Microascales 728a Corollospora maritima FJ FJ FJ FJ FJ Sordariomycetes Microascales CBS Gondwanamyces capensis FJ FJ FJ Sordariomycetes Microascales CBS Halosphaeria appendiculata U46872 U46885 FJ Sordariomycetes Ophiostomatales CBS Ophiostoma stenoceras DQ DQ FJ DQ DQ Sordariomycetes Sordariales Broad Neurospora crassa X04971 AF XM_ XM_ Genome 216 Sordariomycetes Sordariales CBSC Sordaria fimicola AY AY DQ Sordariomycetes Xylariales OSC 004 Xylaria hypoxylon AY5446 AY AY DQ DQ DQ Taphrinomycetes Taphrinales CBS Taphrina deformans DQ DQ FJ DQ DQ4707 DQ Taphrinomycetes Taphrinales IAM Taphrina wiesneri AY AY5482 AY DQ AY5482 DQ voucher GB = obtained from GenBank, or genome databases without clear voucher numbers.
9 C.L. Schoch et al.: Geoglossomycetes 137 gal ecological niches (Geiser et al. 2006). Lecanoromycetes contain the majority of the lichenised fungi (Miadlikowska et al. 2006). Earlier large-scale phylogenies (e.g. Lutzoni et al. 2004) have suggested a sister relationship between these two classes, but we find that such a relationship remains without strong statistical support (Fig. 1). Despite this, internal nodes are well supported enough to provide good support for the hypothesis that lichenisation evolved multiple times in the Ascomycota, with losses being rare (Gueidan et al. 2008, Schoch et al. 2009). The remaining classes are discussed in relation to their respective rankless taxa listed below. Rankless taxon Dothideomyceta This taxon is well supported, with ML bootstrap of 91 % and a moderate Bayesian posterior probability of %. It includes two classes of fungi which produce fissitunicate asci, Arthoniomycetes and Dothideomycetes. Arthoniomycetes consists of ± species of lichenised and lichenicolous fungi with fissitunicate asci and exposed hymenia (Grube 18, Ertz et al. 2009). Unlike other species with fissitunicate asci, these taxa have ascohymenial development, prompting their placement in a transitory group, or Zwischengruppe that is intermediate between ascohymenial and ascolocular development (Henssen & Jahns 1974). The class is resolved as sister to Dothideomycetes, consistent with recent studies (Lutzoni et al. 2004, Spatafora et al. 2006, Wang et al. 2006a). Dothideomycetes is a large class containing two subclasses, Dothideomycetidae and Pleosporomycetidae (Schoch et al. 2006). Our analysis contains members of all known orders in the class, including recent additions (Boehm et al. 2009). This broad representation yields increased resolution in the placement of an order previously labelled incertae sedis, Botryosphaeriales (Schoch et al. 2006). Placement of Botryosphaeriales within subclass Pleosporomycetidae is well supported, as is a close relationship with the unplaced family Tubeufiaceae (Fig. 2). Rankless taxon Sordariomyceta Sordariomyceta contains three classes, Leotiomycetes, Laboulbeniomycetes and Sordariomycetes. We find similar resolution for this clade as for the Dothideomyceta. These three classes are characterised by the production of unitunicate, poricidal asci, or derivatives of such asci (e.g., deliquescent asci). Leotiomycetes and Sordariomycetes include numerous fungi associated with plants as pathogens, endophytes and epiphytes. The sordariomycete phylogeny is comparatively well resolved with 15 orders and 3 subclasses named (Zhang et al. 2006, Kirk et al. 2008). In contrast, the leotiomycete classification still poorly matches its inferred phylogeny. A recent class-wide effort to assess morphological and ecological data in a phylogenetic context continued to find high levels of diversity unaccounted for in the current classification (Wang et al. 2009). In addition to the aforementioned two classes, Fig. 1 also supports the placement of the Laboulbeniomycetes reported in Schoch et al. (2009) as part of a monophyletic lineage. The relationship between the Sordariomycetes and Laboulbeniomycetes is also well supported but we will refrain from naming this node until sampling can be expanded for the Laboulbeniomycetes. The class Laboulbeniomycetes encompasses an enigmatic lineage of insect symbionts and mycoparasites that have long proved problematic with respect to placement in higher-level classification schemes. Laboulbeniomycetes comprises two orders, Laboulbeniales and Pyxidiophorales, that are united by an ascospore synapomorphy of a darkened holdfast region and by molecular data (Weir & Blackwell 2001, 2005). Members of Pyxidiophorales possess globose perithecia with a single layer of wall cells, and long perithecial necks that release their ascospores passively in droplets at the tips of their necks; this mechanism is repeatedly derived within Ascomycota for insect dispersal of ascospores (Blackwell 14). For this reason, they have been likened to other insect-dispersed perithecial ascomycetes (e.g., Ophiostomatales) that now are strongly supported as members of Sordariomycetes. Laboulbeniales includes ectoparasites of insects and displays morphological traits not found elsewhere in the Ascomycota. They form apomorphic ascomata produced by the division and enlargement of ascospores that are difficult to characterize in existing ascomatal terms. Laboulbeniales feature an ostiole, however, which is consistent with perithecia produced by hyphal growth. Determinate growth of the ascospore with a series of predictable cell divisions produces a thallus of a finite number of cells that is characteristic at the genus and species level (Tavares 1979). The analyses presented here strongly support Laboulbeniomycetes as sister to Sordariomycetes. This placement corresponds with the terminology originally applied to this group (Thaxter 1896). It is interesting to note that while species of Pyxidiophorales are endowed with a diverse group of anamorphs, members of Laboulbeniales are mainly known to reproduce sexually. Summary In conclusion, we propose two monotypic formal taxa and describe continued support for four informal rankless taxa. Important improvements in the resolution of deep nodes within the Ascomycota may be attributed to multi-gene sequence data produced by AFTOL and other projects during the last 5 years. The accelerating accumulation of genome-scale sequence data will continue to challenge and improve existing phylogenetic hypotheses. However, in order to direct limited resources towards under-sampled areas in the fungal phylogeny, an accurate, up-to-date classification is required. By placing three earth tongue genera in a separate newly described class, we underscore and communicate the genetic diversity that is found in the fungi producing these very convergent morphologies. Acknowledgements We are grateful to Kentaro Hosaka, Dan Luoma and Zhuliang Yang for the photographs used. We acknowledge funding provided by NSF through a grant (DEB ) to J.W.S. and C.L.S. (while at Oregon State). C.L.S. was also supported in part by the Intramural Research program of the National Institutes of Health. Z.W. was supported by the Yale University Anonymous Postdoctoral Fellowship. References Altekar G, Dwarkadas S, Huelsenbeck JP, Ronquist F Parallel Metropolis coupled Markov chain Monte Carlo for Bayesian phylogenetic inference. Bioinformatics 20: Blackwell M. 14. Minute mycological mysteries: the influence of Arthropods on the lives of Fungi. Mycologia 86: Blackwell M, Hibbett DS, Taylor JW, Spatafora JW Research Coordination Networks: a phylogeny for kingdom Fungi (Deep Hypha). Mycologia : Boehm EWA, Schoch CL, Spatafora JW On the evolution of the Hysteriaceae and Mytilinidiaceae (Pleosporomycetidae, Dothideomycetes, Ascomycota) using four nuclear genes. Mycological Research 113: Corda ACJ Abbildungen der Pilze und Schwaemme. Icones Fungorum, Hucusque Cognitorum 2: Dennis RWG British ascomycetes. Cramer Verlag, Lehre, Germany. Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S, et al Genome evolution in yeasts. Nature 430: Eriksson OE, Winka K. 17. Supraordinal taxa of Ascomycota. Myconet 1: Ertz D, Miadlikowska J, Lutzoni F, Dessein S, Raspe O, et al Towards a new classification of the Arthoniales (Ascomycota) based on a three-gene phylogeny focussing on the genus Opegrapha. Mycological Research 113:
Simplified ascus types
 Simplified ascus types unitunicate bitunicate operculum But numerous variants are recognized pore Ascospores +/- pigmentation (fungal melanin) aseptate, uniseptate or multiseptate +/- appendages +/- sheaths
Simplified ascus types unitunicate bitunicate operculum But numerous variants are recognized pore Ascospores +/- pigmentation (fungal melanin) aseptate, uniseptate or multiseptate +/- appendages +/- sheaths
Overview of Ascomycota
 Overview of Ascomycota Ascomycota ~ 6,350 Genera ~ 64,200 Species compared to Basidiomycota ~ 1,350 Genera ~31,500 Species Between 17,000-20,000 species (~ 30-40%) of Ascomycota are lichenized Many species
Overview of Ascomycota Ascomycota ~ 6,350 Genera ~ 64,200 Species compared to Basidiomycota ~ 1,350 Genera ~31,500 Species Between 17,000-20,000 species (~ 30-40%) of Ascomycota are lichenized Many species
Body plan evolution of ascomycetes, as inferred from an RNA polymerase II phylogeny
 Body plan evolution of ascomycetes, as inferred from an RNA polymerase II phylogeny Yajuan J. Liu and Benjamin D. Hall Departments of Biology and Genome Sciences, Box 355325, University of Washington,
Body plan evolution of ascomycetes, as inferred from an RNA polymerase II phylogeny Yajuan J. Liu and Benjamin D. Hall Departments of Biology and Genome Sciences, Box 355325, University of Washington,
A five-gene phylogeny of Pezizomycotina
 Mycologia, 98(6), 2006, pp. 1018 1028. # 2006 by The Mycological Society of America, Lawrence, KS 66044-8897 A five-gene phylogeny of Pezizomycotina Joseph W. Spatafora 1 Gi-Ho Sung Desiree Johnson Cedar
Mycologia, 98(6), 2006, pp. 1018 1028. # 2006 by The Mycological Society of America, Lawrence, KS 66044-8897 A five-gene phylogeny of Pezizomycotina Joseph W. Spatafora 1 Gi-Ho Sung Desiree Johnson Cedar
The Ascomycota Tree of Life: A Phylum-wide Phylogeny Clarifies the Origin and Evolution of Fundamental Reproductive and Ecological Traits
 Syst. Biol. 58(2):224 239, 2009 Copyright c Society of Systematic Biologists DOI:10.1093/sysbio/syp020 Advance Access publication on June 4, 2009 The Ascomycota Tree of Life: A Phylum-wide Phylogeny Clarifies
Syst. Biol. 58(2):224 239, 2009 Copyright c Society of Systematic Biologists DOI:10.1093/sysbio/syp020 Advance Access publication on June 4, 2009 The Ascomycota Tree of Life: A Phylum-wide Phylogeny Clarifies
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
National Science Foundation Assembling the Tree of Life AToL
 National Science Foundation Assembling the Tree of Life AToL PROPOSAL:: *1500 species of fungi for 8 loci ( 10 kb) in 4 years! 6 nuclear loci:nucssu rdna, nuclsu rdna, ITS rdna, RPB1, RPB2, EF-1α 2 mitochondrial
National Science Foundation Assembling the Tree of Life AToL PROPOSAL:: *1500 species of fungi for 8 loci ( 10 kb) in 4 years! 6 nuclear loci:nucssu rdna, nuclsu rdna, ITS rdna, RPB1, RPB2, EF-1α 2 mitochondrial
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Lecture 12. Ascomycota II
 Lecture 12 Ascomycota II - Saccharomycotina - Pezizomycotina 1 Lutzoni et al., 2004, American Journal of Botany Saccharomycotina - the true budding yeasts (--> Saccharomycetes, or the Saccharomycetales
Lecture 12 Ascomycota II - Saccharomycotina - Pezizomycotina 1 Lutzoni et al., 2004, American Journal of Botany Saccharomycotina - the true budding yeasts (--> Saccharomycetes, or the Saccharomycetales
Chapter 16: Reconstructing and Using Phylogenies
 Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Classification, Phylogeny yand Evolutionary History
 Classification, Phylogeny yand Evolutionary History The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize
Classification, Phylogeny yand Evolutionary History The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize
The practice of naming and classifying organisms is called taxonomy.
 Chapter 18 Key Idea: Biologists use taxonomic systems to organize their knowledge of organisms. These systems attempt to provide consistent ways to name and categorize organisms. The practice of naming
Chapter 18 Key Idea: Biologists use taxonomic systems to organize their knowledge of organisms. These systems attempt to provide consistent ways to name and categorize organisms. The practice of naming
Supporting Information
 Supporting Information Systematic analyses reveal uniqueness and origin of the CFEM domain in fungi Zhen-Na Zhang 1,2,, Qin-Yi Wu 1,, Gui-Zhi Zhang 2, Yue-Yan Zhu 1, Robert W. Murphy 3, Zhen Liu 3, * and
Supporting Information Systematic analyses reveal uniqueness and origin of the CFEM domain in fungi Zhen-Na Zhang 1,2,, Qin-Yi Wu 1,, Gui-Zhi Zhang 2, Yue-Yan Zhu 1, Robert W. Murphy 3, Zhen Liu 3, * and
Phylogenomics. Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University. Tuesday, January 29, 13
 Phylogenomics Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University How may we improve our inferences? How may we improve our inferences? Inferences Data How may we improve
Phylogenomics Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University How may we improve our inferences? How may we improve our inferences? Inferences Data How may we improve
Lecture 16. Ascomycota V
 Lecture 16 Ascomycota V - Lichenized fungi - Eurotiomycetes - Laboulbeniales Lutzoni et al., 2004, American Journal of Botany Lichen-forming Lichen-forming Sordariomycetes Nature 21:937-940. 2001 - Nuclear
Lecture 16 Ascomycota V - Lichenized fungi - Eurotiomycetes - Laboulbeniales Lutzoni et al., 2004, American Journal of Botany Lichen-forming Lichen-forming Sordariomycetes Nature 21:937-940. 2001 - Nuclear
Lecture 11 Friday, October 21, 2011
 Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Inclusion of Nothomitra in Geoglossomycetes
 Inclusion of Nothomitra in Geoglossomycetes Mycosphere Doi 10.5943/mycosphere/2/6/5/ Hustad VP 1,2*, Miller AN 2, Moingeon J-M 3 and Priou J-P 4 1 Department of Plant Biology, University of Illinois at
Inclusion of Nothomitra in Geoglossomycetes Mycosphere Doi 10.5943/mycosphere/2/6/5/ Hustad VP 1,2*, Miller AN 2, Moingeon J-M 3 and Priou J-P 4 1 Department of Plant Biology, University of Illinois at
Name: Class: Date: ID: A
 Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Class: _ Date: _ Ch 17 Practice test 1. A segment of DNA that stores genetic information is called a(n) a. amino acid. b. gene. c. protein. d. intron. 2. In which of the following processes does change
Microbial Taxonomy and the Evolution of Diversity
 19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
19 Microbial Taxonomy and the Evolution of Diversity Copyright McGraw-Hill Global Education Holdings, LLC. Permission required for reproduction or display. 1 Taxonomy Introduction to Microbial Taxonomy
Phylogenetic study of Diploschistes (lichen-forming Ascomycota: Ostropales: Graphidaceae), based on morphological, chemical, and molecular data
 Vol. 62 (2) April 2013 International Journal of Taxonomy, Phylogeny and Evolution Electronic Supplement to Phylogenetic study of Diploschistes (lichen-forming Ascomycota: Ostropales: Graphidaceae), based
Vol. 62 (2) April 2013 International Journal of Taxonomy, Phylogeny and Evolution Electronic Supplement to Phylogenetic study of Diploschistes (lichen-forming Ascomycota: Ostropales: Graphidaceae), based
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
The process by which the genetic structure of populations changes over time.
 Evolution The process by which the genetic structure of populations changes over time. Divergent evolution is the accumulation of differences between groups which can lead to the formation of new species.
Evolution The process by which the genetic structure of populations changes over time. Divergent evolution is the accumulation of differences between groups which can lead to the formation of new species.
CHAPTERS 24-25: Evidence for Evolution and Phylogeny
 CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
Fungi. Abstract. Fig. 1 A basidiomycete (Coprinopsis sp.) from the Eastern United States. Credit: M. E. Hood (Amherst College).
 Fungi Jaime E. Blair Department of Biology, 139 McGuire Life Sciences Building, Amherst College, Amherst, MA, 01002 USA. Current address: Department of Biology, Franklin and Marshall College, Lancaster,
Fungi Jaime E. Blair Department of Biology, 139 McGuire Life Sciences Building, Amherst College, Amherst, MA, 01002 USA. Current address: Department of Biology, Franklin and Marshall College, Lancaster,
Macroevolution Part I: Phylogenies
 Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
Kingdom Fungi. Learning Objectives. Introduction. Activity1: Zygomycota. Revised Fall 2017
 Kingdom Fungi Revised Fall 2017 ** You will require your text book Biological Science during this lab ** Learning Objectives Building on the learning objectives from your lab syllabus, you will be expected
Kingdom Fungi Revised Fall 2017 ** You will require your text book Biological Science during this lab ** Learning Objectives Building on the learning objectives from your lab syllabus, you will be expected
ESS 345 Ichthyology. Systematic Ichthyology Part II Not in Book
 ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
Family placement of Ascotaiwania and Ascolacicola based on DNA sequences from the large subunit rrna gene
 Fungal Diversity 2 (March 1999) Family placement of Ascotaiwania and Ascolacicola based on DNA sequences from the large subunit rrna gene v. Mala RanghooP, Kevin D. Hydel, E.C.Y. Liewl and J.W. Spatafora2
Fungal Diversity 2 (March 1999) Family placement of Ascotaiwania and Ascolacicola based on DNA sequences from the large subunit rrna gene v. Mala RanghooP, Kevin D. Hydel, E.C.Y. Liewl and J.W. Spatafora2
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley
 Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley B.D. Mishler Feb. 7, 2012. Morphological data IV -- ontogeny & structure of plants The last frontier
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley B.D. Mishler Feb. 7, 2012. Morphological data IV -- ontogeny & structure of plants The last frontier
A new species of Bertiella (Melanommataceae) from Brazil and a key to accepted species
 Mycosphere 8 (4): 392 396 (2017) www.mycosphere.org ISSN 2077 7019 Article Doi 10.5943/mycosphere/8/4/1 Copyright Guizhou Academy of Agricultural Sciences A new species of Bertiella (Melanommataceae) from
Mycosphere 8 (4): 392 396 (2017) www.mycosphere.org ISSN 2077 7019 Article Doi 10.5943/mycosphere/8/4/1 Copyright Guizhou Academy of Agricultural Sciences A new species of Bertiella (Melanommataceae) from
PHYLOGENY & THE TREE OF LIFE
 PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
Chapter 26 Phylogeny and the Tree of Life Chapter focus Shifting from the process of how evolution works to the pattern evolution produces over time. Phylogeny Phylon = tribe, geny = genesis or origin
The Tree of Life. Chapter 17
 The Tree of Life Chapter 17 1 17.1 Taxonomy The science of naming and classifying organisms 2000 years ago Aristotle Grouped plants and animals Based on structural similarities Greeks and Romans included
The Tree of Life Chapter 17 1 17.1 Taxonomy The science of naming and classifying organisms 2000 years ago Aristotle Grouped plants and animals Based on structural similarities Greeks and Romans included
Phylogenetic Analysis
 Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Phylogenetic Analysis Aristotle Through classification, one might discover the essence and purpose of species. Nelson & Platnick (1981) Systematics and Biogeography Carl Linnaeus Swedish botanist (1700s)
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
INTRODUCTION. Studies in Mycology 64:
 available online at www.studiesinmycology.org doi:10.3114/sim.2009.64.03 Studies in Mycology 64: 49 83. 2009. A molecular phylogenetic reappraisal of the Hysteriaceae, Mytilinidiaceae and Gloniaceae (Pleosporomycetidae,
available online at www.studiesinmycology.org doi:10.3114/sim.2009.64.03 Studies in Mycology 64: 49 83. 2009. A molecular phylogenetic reappraisal of the Hysteriaceae, Mytilinidiaceae and Gloniaceae (Pleosporomycetidae,
Using Trees for Classifications. Introduction
 Using Trees for Classifications The Phylogenetic Cibele Caio Principles and Practice of Phylogenetic Systematics, Spring 2009 Introduction The impusle to characterize and classify species Ancient Aristoteles
Using Trees for Classifications The Phylogenetic Cibele Caio Principles and Practice of Phylogenetic Systematics, Spring 2009 Introduction The impusle to characterize and classify species Ancient Aristoteles
9/19/2012. Chapter 17 Organizing Life s Diversity. Early Systems of Classification
 Section 1: The History of Classification Section 2: Modern Classification Section 3: Domains and Kingdoms Click on a lesson name to select. Early Systems of Classification Biologists use a system of classification
Section 1: The History of Classification Section 2: Modern Classification Section 3: Domains and Kingdoms Click on a lesson name to select. Early Systems of Classification Biologists use a system of classification
Advanced Mycology, Pawlowska
 Fall 2013 PLPA4490 Advanced Mycology 3 credits Lecture: Tuesday & Thursday 9:05 am 9:55 am 336 Plant Science Building Laboratory: Thursday 1:25 pm 4:25 pm 326 Plant Science Building Instructor: Teresa
Fall 2013 PLPA4490 Advanced Mycology 3 credits Lecture: Tuesday & Thursday 9:05 am 9:55 am 336 Plant Science Building Laboratory: Thursday 1:25 pm 4:25 pm 326 Plant Science Building Instructor: Teresa
An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny
 Mycologia, 98(6), 2006, pp. 1076 1087. # 2006 by The Mycological Society of America, Lawrence, KS 66044-8897 An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny Ning Zhang
Mycologia, 98(6), 2006, pp. 1076 1087. # 2006 by The Mycological Society of America, Lawrence, KS 66044-8897 An overview of the systematics of the Sordariomycetes based on a four-gene phylogeny Ning Zhang
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
New primers for promising single-copy genes in fungal phylogenetics and systematics
 Persoonia 23, 2009: 35 40 www.persoonia.org RESEARCH ARTICLE doi:10.3767/003158509x470602 New primers for promising single-copy genes in fungal phylogenetics and systematics I. Schmitt 1, A. Crespo 2,
Persoonia 23, 2009: 35 40 www.persoonia.org RESEARCH ARTICLE doi:10.3767/003158509x470602 New primers for promising single-copy genes in fungal phylogenetics and systematics I. Schmitt 1, A. Crespo 2,
Chapter 19: Taxonomy, Systematics, and Phylogeny
 Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Identification of Fungal DNA Barcode Targets and PCR Primers Based on Pfam Protein Families and Taxonomic Hierarchy
 30 The Open Applied Informatics Journal, 2011, 5, (Suppl 1-M5) 30-44 Open Access Identification of Fungal DNA Barcode Targets and PCR Primers Based on Pfam Protein Families and Taxonomic Hierarchy Christopher
30 The Open Applied Informatics Journal, 2011, 5, (Suppl 1-M5) 30-44 Open Access Identification of Fungal DNA Barcode Targets and PCR Primers Based on Pfam Protein Families and Taxonomic Hierarchy Christopher
Chapter 17A. Table of Contents. Section 1 Categories of Biological Classification. Section 2 How Biologists Classify Organisms
 Classification of Organisms Table of Contents Section 1 Categories of Biological Classification Section 1 Categories of Biological Classification Classification Section 1 Categories of Biological Classification
Classification of Organisms Table of Contents Section 1 Categories of Biological Classification Section 1 Categories of Biological Classification Classification Section 1 Categories of Biological Classification
How to read and make phylogenetic trees Zuzana Starostová
 How to read and make phylogenetic trees Zuzana Starostová How to make phylogenetic trees? Workflow: obtain DNA sequence quality check sequence alignment calculating genetic distances phylogeny estimation
How to read and make phylogenetic trees Zuzana Starostová How to make phylogenetic trees? Workflow: obtain DNA sequence quality check sequence alignment calculating genetic distances phylogeny estimation
infections beer Ascomycota wine Yeasts bread Dry yeast Yeast colonies
 infections beer Ascomycota wine Yeasts bread Dry yeast Yeast colonies 1. Fungi form a monophyletic group. Fungi -2011 Ascomycota No motile cells * Ascus = meiosporangium G. Wong Ascomycota Ascomycetous
infections beer Ascomycota wine Yeasts bread Dry yeast Yeast colonies 1. Fungi form a monophyletic group. Fungi -2011 Ascomycota No motile cells * Ascus = meiosporangium G. Wong Ascomycota Ascomycetous
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
The process by which the genetic structure of populations changes over time.
 Evolution The process by which the genetic structure of populations changes over time. Divergent evolution Goldfields and Ahinahina (silversword) a highly evolved member of the composite family. Evolution
Evolution The process by which the genetic structure of populations changes over time. Divergent evolution Goldfields and Ahinahina (silversword) a highly evolved member of the composite family. Evolution
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
Chapter 10. Classification and Phylogeny of Animals. Order in Diversity. Hierarchy of taxa. Table Linnaeus introduced binomial nomenclature
 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Chapter 10 Classification and Phylogeny of Animals Order in Diversity History Systematic zoologists have three
Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Chapter 10 Classification and Phylogeny of Animals Order in Diversity History Systematic zoologists have three
SPECIATION. REPRODUCTIVE BARRIERS PREZYGOTIC: Barriers that prevent fertilization. Habitat isolation Populations can t get together
 SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
Classification and Phylogeny
 Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny
 CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
CHAPTER 26 PHYLOGENY AND THE TREE OF LIFE Connecting Classification to Phylogeny To trace phylogeny or the evolutionary history of life, biologists use evidence from paleontology, molecular data, comparative
Outline. Classification of Living Things
 Outline Classification of Living Things Chapter 20 Mader: Biology 8th Ed. Taxonomy Binomial System Species Identification Classification Categories Phylogenetic Trees Tracing Phylogeny Cladistic Systematics
Outline Classification of Living Things Chapter 20 Mader: Biology 8th Ed. Taxonomy Binomial System Species Identification Classification Categories Phylogenetic Trees Tracing Phylogeny Cladistic Systematics
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
What is Phylogenetics
 What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
Classification and Phylogeny
 Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Phylogenetic inference
 Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Reconstructing the history of lineages
 Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
CHAPTER 10 Taxonomy and Phylogeny of Animals
 CHAPTER 10 Taxonomy and Phylogeny of Animals 10-1 10-2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Linnaeus and Taxonomy More than 1.5 million species of
CHAPTER 10 Taxonomy and Phylogeny of Animals 10-1 10-2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Linnaeus and Taxonomy More than 1.5 million species of
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis
 Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Hierarchies can be represented as trees:
 Diversity of Life Classification - an organized scheme for grouping organisms - a tool for communication - Hierarchical - a series of successive and inclusive rankings Domain - the highest rank - contains
Diversity of Life Classification - an organized scheme for grouping organisms - a tool for communication - Hierarchical - a series of successive and inclusive rankings Domain - the highest rank - contains
Using phylogenetics to estimate species divergence times... Basics and basic issues for Bayesian inference of divergence times (plus some digression)
 Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Biologists use a system of classification to organize information about the diversity of living things.
 Section 1: Biologists use a system of classification to organize information about the diversity of living things. K What I Know W What I Want to Find Out L What I Learned Essential Questions What are
Section 1: Biologists use a system of classification to organize information about the diversity of living things. K What I Know W What I Want to Find Out L What I Learned Essential Questions What are
Constructing Evolutionary/Phylogenetic Trees
 Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
!!!! FIG$S1!Rarefaction!curves!using!the!0%!RDP!Classifier!confidence!dataset!for!the!(A)!
 A 600 500 OK Control OK Warming C T Number of Genera 400 300 200 100 B 0 1.E+00 5.E+03 1.E+04 2.E+04 Number of Sequences 600 500 Control AK 0 to -20 AK -20 to -50 Warming 0-20 cm AK 0 to -20 20-50 cm AK
A 600 500 OK Control OK Warming C T Number of Genera 400 300 200 100 B 0 1.E+00 5.E+03 1.E+04 2.E+04 Number of Sequences 600 500 Control AK 0 to -20 AK -20 to -50 Warming 0-20 cm AK 0 to -20 20-50 cm AK
Consensus Methods. * You are only responsible for the first two
 Consensus Trees * consensus trees reconcile clades from different trees * consensus is a conservative estimate of phylogeny that emphasizes points of agreement * philosophy: agreement among data sets is
Consensus Trees * consensus trees reconcile clades from different trees * consensus is a conservative estimate of phylogeny that emphasizes points of agreement * philosophy: agreement among data sets is
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
How should we organize the diversity of animal life?
 How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
The Tree of Life. Phylogeny
 The Tree of Life Phylogeny Phylogenetics Phylogenetic trees illustrate the evolutionary relationships among groups of organisms, or among a family of related nucleic acid or protein sequences Each branch
The Tree of Life Phylogeny Phylogenetics Phylogenetic trees illustrate the evolutionary relationships among groups of organisms, or among a family of related nucleic acid or protein sequences Each branch
A. Incorrect! In the binomial naming convention the Kingdom is not part of the name.
 Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Microbiology Problem Drill 08: Classification of Microorganisms No. 1 of 10 1. In the binomial system of naming which term is always written in lowercase? (A) Kingdom (B) Domain (C) Genus (D) Specific
Verlag Ferdinand Berger & Söhne Ges.m.b.H., Horn, Austria, download unter Book Reviews
 Book Reviews Kiffer, E. & M. Morelet (2000). The deuteromycetes: mitosporic fungi, classification and generic keys. - Science Publishers, Inc., Enfield, NH 03748, USA. ISBN 1-57808-068-1; 273pp. Three
Book Reviews Kiffer, E. & M. Morelet (2000). The deuteromycetes: mitosporic fungi, classification and generic keys. - Science Publishers, Inc., Enfield, NH 03748, USA. ISBN 1-57808-068-1; 273pp. Three
Fungi are absorptive heterotrophs that secrete digestive enzymes and are major decomposers of dead organic material
 Fungi 1 2002 Prentice Hall, Inc The scarlet hood (Hygrocybe coccinea) Fungi are absorptive heterotrophs that secrete digestive enzymes and are major decomposers of dead organic material 2 Animals 3 Myxozoa
Fungi 1 2002 Prentice Hall, Inc The scarlet hood (Hygrocybe coccinea) Fungi are absorptive heterotrophs that secrete digestive enzymes and are major decomposers of dead organic material 2 Animals 3 Myxozoa
Anatomy of a species tree
 Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Chapter Chemical Uniqueness 1/23/2009. The Uses of Principles. Zoology: the Study of Animal Life. Fig. 1.1
 Fig. 1.1 Chapter 1 Life: Biological Principles and the Science of Zoology BIO 2402 General Zoology Copyright The McGraw Hill Companies, Inc. Permission required for reproduction or display. The Uses of
Fig. 1.1 Chapter 1 Life: Biological Principles and the Science of Zoology BIO 2402 General Zoology Copyright The McGraw Hill Companies, Inc. Permission required for reproduction or display. The Uses of
Lecture 6 Phylogenetic Inference
 Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
The problems of traditional and phylogenetic taxonomy of fungi
 The problems of traditional and phylogenetic taxonomy of fungi Mycosphere Vasilyeva LN 1* and Stephenson SL 2 1 Institute of Biology and Soil Science, Far East Branch of the Russian Academy of Sciences,Vladivostok
The problems of traditional and phylogenetic taxonomy of fungi Mycosphere Vasilyeva LN 1* and Stephenson SL 2 1 Institute of Biology and Soil Science, Far East Branch of the Russian Academy of Sciences,Vladivostok
Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food.
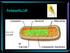 Prokaryotic Cell Eukaryotic Cell Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food. Heterotrophs must get energy by eating autotrophs or other heterotrophs.
Prokaryotic Cell Eukaryotic Cell Autotrophs capture the light energy from sunlight and convert it to chemical energy they use for food. Heterotrophs must get energy by eating autotrophs or other heterotrophs.
Concept Modern Taxonomy reflects evolutionary history.
 Concept 15.4 Modern Taxonomy reflects evolutionary history. What is Taxonomy: identification, naming, and classification of species. Common Names: can cause confusion - May refer to several species (ex.
Concept 15.4 Modern Taxonomy reflects evolutionary history. What is Taxonomy: identification, naming, and classification of species. Common Names: can cause confusion - May refer to several species (ex.
How Biological Diversity Evolves
 CHAPTER 14 How Biological Diversity Evolves PowerPoint Lectures for Essential Biology, Third Edition Neil Campbell, Jane Reece, and Eric Simon Essential Biology with Physiology, Second Edition Neil Campbell,
CHAPTER 14 How Biological Diversity Evolves PowerPoint Lectures for Essential Biology, Third Edition Neil Campbell, Jane Reece, and Eric Simon Essential Biology with Physiology, Second Edition Neil Campbell,
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Lecture 10. Basidiomycota V
 Lecture 10 Basidiomycota V - Agaricomycotina: --- Auriculariales, Dacrymycetales, Tremellales, Filobasidiales - Pucciniomycotina (rust fungi) - Ustilaginomycotina (smut fungi) Tulasnellales, Auriculariales,
Lecture 10 Basidiomycota V - Agaricomycotina: --- Auriculariales, Dacrymycetales, Tremellales, Filobasidiales - Pucciniomycotina (rust fungi) - Ustilaginomycotina (smut fungi) Tulasnellales, Auriculariales,
Pathogenic fungi genomics, evolution and epidemiology! John W. Taylor University of California Berkeley, USA
 Pathogenic fungi genomics, evolution and epidemiology! John W. Taylor University of California Berkeley, USA Adaptation! Divergence Divergence Adaptation Darwin 1859 Mendel 1866 1830 1850 1870 1890 1910
Pathogenic fungi genomics, evolution and epidemiology! John W. Taylor University of California Berkeley, USA Adaptation! Divergence Divergence Adaptation Darwin 1859 Mendel 1866 1830 1850 1870 1890 1910
MiGA: The Microbial Genome Atlas
 December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
December 12 th 2017 MiGA: The Microbial Genome Atlas Jim Cole Center for Microbial Ecology Dept. of Plant, Soil & Microbial Sciences Michigan State University East Lansing, Michigan U.S.A. Where I m From
Mixture Models in Phylogenetic Inference. Mark Pagel and Andrew Meade Reading University.
 Mixture Models in Phylogenetic Inference Mark Pagel and Andrew Meade Reading University m.pagel@rdg.ac.uk Mixture models in phylogenetic inference!some background statistics relevant to phylogenetic inference!mixture
Mixture Models in Phylogenetic Inference Mark Pagel and Andrew Meade Reading University m.pagel@rdg.ac.uk Mixture models in phylogenetic inference!some background statistics relevant to phylogenetic inference!mixture
Homework Assignment, Evolutionary Systems Biology, Spring Homework Part I: Phylogenetics:
 Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Chapter 26. Phylogeny and the Tree of Life. Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Pearson Education, Inc.
 Chapter 26 Phylogeny and the Tree of Life Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Investigating the Tree of Life Phylogeny is the evolutionary history of a species or group of
Chapter 26 Phylogeny and the Tree of Life Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Investigating the Tree of Life Phylogeny is the evolutionary history of a species or group of
Fungi evolved right on track
 Mycologia, 101(6), 2009, pp. 810 822. DOI: 10.3852/09-016 # 2009 by The Mycological Society of America, Lawrence, KS 66044-8897 Fungi evolved right on track Robert Lücking 1 Sabine Huhndorf Department
Mycologia, 101(6), 2009, pp. 810 822. DOI: 10.3852/09-016 # 2009 by The Mycological Society of America, Lawrence, KS 66044-8897 Fungi evolved right on track Robert Lücking 1 Sabine Huhndorf Department
Need for systematics. Applications of systematics. Linnaeus plus Darwin. Approaches in systematics. Principles of cladistics
 Topics Need for systematics Applications of systematics Linnaeus plus Darwin Approaches in systematics Principles of cladistics Systematics pp. 474-475. Systematics - Study of diversity and evolutionary
Topics Need for systematics Applications of systematics Linnaeus plus Darwin Approaches in systematics Principles of cladistics Systematics pp. 474-475. Systematics - Study of diversity and evolutionary
Announcements: 1. Labs meet this week 2. Lab manuals have been ordered 3. Some slides from each lecture will be on the web 4. Study questions will be
 Announcements: 1. Labs meet this week 2. Lab manuals have been ordered 3. Some slides from each lecture will be on the web 4. Study questions will be posted after each lecture Prokaryotes Eukaryotes Protozoa
Announcements: 1. Labs meet this week 2. Lab manuals have been ordered 3. Some slides from each lecture will be on the web 4. Study questions will be posted after each lecture Prokaryotes Eukaryotes Protozoa
AP Biology Exam #7 (PRACTICE) Subunit #7: Diversity of Life
 AP Biology Exam #7 (PRACTICE) Subunit #7: Diversity of Life Multiple Choice Questions: Choose the best answer then bubble your answer on your scantron sheet. 1. Armadillos and spiny anteaters are not related.
AP Biology Exam #7 (PRACTICE) Subunit #7: Diversity of Life Multiple Choice Questions: Choose the best answer then bubble your answer on your scantron sheet. 1. Armadillos and spiny anteaters are not related.
Using Phylogenomics to Predict Novel Fungal Pathogenicity Genes
 Using Phylogenomics to Predict Novel Fungal Pathogenicity Genes David DeCaprio, Ying Li, Hung Nguyen (sequenced Ascomycetes genomes courtesy of the Broad Institute) Phylogenomics Combining whole genome
Using Phylogenomics to Predict Novel Fungal Pathogenicity Genes David DeCaprio, Ying Li, Hung Nguyen (sequenced Ascomycetes genomes courtesy of the Broad Institute) Phylogenomics Combining whole genome
