BIOINFORMATICS ORIGINAL PAPER
|
|
|
- Elisabeth Cox
- 5 years ago
- Views:
Transcription
1 BIOINFORMATICS ORIGINAL PAPER Vol. 24 no , pages doi: /bioinformatics/btn414 Sequence analysis Model-based prediction of sequence alignment quality Virpi Ahola 1,2,, Tero Aittokallio 3, Mauno Vihinen 4,5 and Esa Uusipaikka 2 1 Biotechnology and Food Research, MTT Agrifood Research Finland, FI Jokioinen, 2 Department of Statistics, 3 Department of Mathematics, FI-20014, University of Turku, 4 Institute of Medical Technology, FI-33014, University of Tampere and 5 Tampere University Hospital, FI Tampere, Finland Received on May 20, 2008; revised on July 25, 2008; accepted on August 1, 2008 Advance Access publication August 4, 2008 Associate Editor: Burkhard Rost ABSTRACT Motivation: Multiple sequence alignment (MSA) is an essential prerequisite for many sequence analysis methods and valuable tool itself for describing relationships between protein sequences. Since the success of the sequence analysis is highly dependent on the reliability of alignments, measures for assessing the quality of alignments are highly requisite. Results: We present a statistical model-based alignment quality score. Unlike other quality scores, it does not require several parallel alignments for the same set of sequences or additional structural information. Our quality score is based on measuring the conservation level of reference alignments in Homstrad. Reference sequences were realigned with the Mafft, Muscle and Probcons alignment programs, and a sum-of-pairs (SP) score was used to measure the quality of the realignments. Statistical modelling of the SP score as a function of conservation level and other alignment characteristics makes it possible to predict the SP score for any global MSA. The predicted SP scores are highly correlated with the correct SP scores, when tested on the Homstrad and SABmark databases. The results are comparable to that of multiple overlap score (MOS) and better than those of normalized mean distance (NorMD) and normalized irmsd (NiRMSD) alignment quality criteria. Furthermore, the predicted SP score is able to detect alignments with badly aligned or unrelated sequences. Availability: The method is freely available at AlignmentQuality/ Contact: virpi.ahola@mtt.fi Supplementary information: Supplementary data are available at Bioinformatics online. 1 INTRODUCTION Biological nucleotide or amino acid sequence analysis is typically based on comparing a sequence of interest with homologous sequences. As more and more sequence data from different species is becoming available, comparative genomics using multiple sequence alignment (MSA) has increased its importance (Notredame, 2002). The quality of alignments is a focal limiting factor in sequence analysis (Landan and Graur, 2008). MSA programs are not always successful in constructing biologically plausible alignments, and the use of poor alignment results may lead, for instance, to incorrect To whom correspondence should be addressed. structural model and the error can cumulate in the subsequent structure predictions (Domingues et al., 2000). The alignment quality measures are important, e.g. for benchmarking the MSA programs and assessing the quality of a given alignment. The MSA programs have been validated and compared using reference sequence alignment databases such as Balibase, Homstrad, Prefab and SABmark (Bahr et al., 2001; Edgar, 2004; Mizuguchi et al., 1998; Walle et al., 2005). The benchmarking databases have usually been constructed using protein 3D structural alignments, and hence they are independent of sequence alignment methods (Edgar and Batzoglou, 2006). The quality of the MSA programs has typically been assessed by the sum-of-pairs (SP) or the column score (CS), that measure the proportion of correctly aligned residue pairs or alignment positions, respectively, in the alignment (Thompson et al., 1999). The lack of proper alignment validation methods, which could be used without reference alignments, is a disadvantage in benchmarking the MSA programs. The current approaches prohibit the analysis of any set of sequences and force to trust the structure-based reference alignment databases, although it is known that different structure alignment methods may produce different alignments (Goldsmith-Fischman and Honig, 2003; Lackner et al., 2000; Notredame, 2007). Assessing the quality of a single alignment is even more complicated than the validation of the MSA methods because no reference alignment is available. The only way is to use validation methods, which do not need a reference alignment. Traditionally, alignment quality measures have been formulated on the basis of column-based conservation measures, which are then summed over the entire length of the alignment. The NorMD is a normalized mean distance (MD) score that measures the mean distance between similarities of residues at each alignment position (Thompson et al., 2001). Other column-based methods, such as IC or AL2CO, use entropy or the sum of pairs measures (Hertz and Stormo, 1999; Pei and Grishin, 2001). Beiko et al. (2005) presented a word-oriented objective function (WOOF) for alignment validation. The method is not a column-based, but relies on scoring conserved amino acid regions between pairs of sequences. When the structures for at least two homologues are available, one can use the normalized irmsd score (NiRMSD), which measures the root mean squared distance (RMSD) of all possible residue pairs within a certain sphere of the radius between two sequences (Armougom et al., 2006). The drawback of the NiRMSD score is that it can only be used to compare relative accuracy of two alignments. Another possibility is to use structural similarity scores (Pei and The Author Published by Oxford University Press. All rights reserved. For Permissions, please journals.permissions@oxfordjournals.org 2165
2 V.Ahola et al. Grishin, 2006, 2007). The quality assessments of the structure-based measures are, however, based only on the sequences whose structure is available. Yet another approach is based on aligning the same set of sequences with several MSA programs and comparing the results (Morgenstern et al., 2003). Such consistency based scores assume that the residue pairs coinciding in most of the alignments are biologically correct (Gotoh, 1990; Vingron and Argos, 1991). The multiple overlap score (MOS) determines alignment quality as the proportion of similarly aligned residue pairs among all pairs of aligned residues (Lassmann and Sonnhammer, 2005b). Their score is essentially similar to alignment consistency of the overall residue evaluation (alcore) index in TCoffee alignment program (Notredame and Abergel, 2003). Landan and Graur (2007, 2008) realigned several suboptimal alignments and the same sequences in reversed residue order. Their reliability measure was obtained by calculating the proportion of alignments that reproduce the pairing of the residue pair at one alignment position and averaging the proportion over all the residue pairs and columns of the alignment. We present a model-based alignment quality scoring method, which combines the information available on the reference alignments with the conservation level and other observed properties of the MSA of interest. We used the Homstrad database as the source of reference information and aligned the reference sequences with Mafft (L-ins-I), Muscle and Probcons (Do et al., 2005; Edgar, 2004; Katoh et al., 2002, 2005). The divergence between the reference and realignments was evaluated by calculating the SP score, which was chosen as the representative alignment quality score (Thompson et al., 1999). The correct SP score for the Homstrad alignment was statistically modelled in terms of the conservation level by taking the identity, number of sequences and the alignment length into account. The conservation level of the alignment was calculated using our statistical conservation score (Ahola et al., 2006), which defines the proportion of conserved amino acids in the alignment. As a final outcome, the estimated model parameters can be used to predict the quality of any global MSA. We tested the novel quality score on the structure-based alignments in Homstrad and SABmark databases by comparing the predicted SP scores with the correct SP scores. Finally, we compared the performance of the quality score with previous scores, MOS, NiRMSD and NorMD, in the same benchmarking databases. 2 METHODS 2.1 Alignment quality assessment The Homstrad (Mizuguchi et al., 1998) database was used as the reference for constructing the alignment quality score. A set of training sequences in the Homstrad database were aligned using the Mafft with L-INS-i mode (Katoh et al., 2002, 2005), and Muscle (Edgar, 2004) and Probcons (Do et al., 2005) with default parameters (Fig. 1, stage 1). These programs were chosen, because they are fast and they are among the most accurate alignment programs (Ahola et al., 2006; Golubchik et al., 2007; Nuin et al., 2006). The SP scores, i.e. the proportion of correctly aligned residue pairs in an alignment, were calculated between the reference and realignments to measure the quality of the automatically generated MSAs (Thompson et al., 1999). Furthermore, the conservation-level ConsAA (Ahola et al., 2006) and other variables describing the alignment, such as the sequence identity, alignment length and the number of sequences in the alignment, were calculated for the Mafft, Muscle and Probcons alignments. Beta regression model was fitted for the SP score using the ConsAA and other descriptive variables as predictors. The predicted SP score value of a new alignment Fig. 1. Diagram for building and using the alignment quality model. Stage 1 illustrates the steps for calculating the beta regression parameter estimates and stage 2 their use to predict the SP score for an unknown alignment. could then be calculated by using the parameter values of the fitted model (Fig. 1, stage 2). The generalizability of the quality score was tested using unseen sequences of the SABmark and independent test sequences of the Homstrad databases. The predicted SP score values and their prediction intervals were calculated for the test alignments of the Homstrad and SABmark databases. The performance of the predicted SP scores was assessed by comparing them with the CS and correct SP scores in the Homstrad and with median f d and f m scores in the SABmark databases. Additionally, other alignment quality measures, MOS, NiRMSD and NorMD scores, were calculated for the Mafft, Muscle and Probcons alignments and compared with the correct alignment quality measures and the predicted SP scores. The ability of the quality scores to distinguish alignments with badly aligned or unrelated sequences from correct alignments was tested by adding random sequences to the reference alignments. 2.2 Benchmarking databases The Homstrad database provides homologous structure-based alignments for all known protein structures (Mizuguchi et al., 1998). The structures have been first clustered into homologous families and then the representative sequences of each family have been structure aligned and additional sequences have been added into the alignment by ClustalW. We used all the Homstrad families which contained at least two known structures, alignment had five or more sequences and was more than 50 residues long. Four database versions (Homs10, Homs20, Homs50 and Homs100) were build. The Homs10 includes alignments with 5 to 10 sequences, the Homs20 consists of alignments with 11 to 20 sequences, the Homs50 includes alignments with 20 to 50 sequences and the Homs100 the alignments with 50 to 100 sequences. In each of the rebuilt databases, each alignment includes all the representative sequences and, if needed, additional sequences. The additional sequences were randomly chosen and identical entries were excluded from the alignment. We used the combined set of four Homstrad datasets. The alignments were divided into training and test sets: two third (2390) of the alignments were included into the training and one third (1189) into the test set. The SP scores were calculated using the Bali_score program (Thompson et al., 1999). To study the effect of badly aligned or unrelated sequences on the quality scorings, we added 1 to 4 random sequences to each of the 284 test alignments of the Homs10 database and referred these datasets as Homs10 + R1, Homs10 + R2, Homs10 + R3 and Homs10 + R4. The SABmark database consists of two alignment sets (Walle et al., 2005). Superfamily set includes alignments with low to intermediate sequence identity ( 50%), while the alignments in the twilight zone set have very 2166
3 Prediction of sequence alignment quality low to low identity ( 25%). We used the combined set of those superfamily and twilight zone alignments which had at least five sequences of more than 50 residues (375). The results for the superfamily and twilight zone alignments are provided separately in the Supplementary Material. The SABmark database does not provide multiple reference alignments, but pairwise alignments of all sequences within a group are available. Therefore, since the SP scores could not be calculated for the SABmark database, we used instead the developer s viewpoint score f d, that describes which part of the structural alignment is correctly represented in the sequence alignment, and modeller s viewpoint score f m that measures the proportion of correctly aligned residues in the sequence alignment (Sauder et al., 2000). The f d and f m values were calculated for the pairwise alignments and the median values were used to estimate the SP score. Script supplied with the SABmark database was used to calculate the f d and f m scores. The pairwise alignments in which no residues were correctly aligned were excluded from the analysis similar to Lassmann and Sonnhammer (2005b). 2.3 Positional conservation score Let us assume that the amino acid frequencies n = (n 1,n 2,...,n 20 )inone position of an alignment follow a multinomial distribution with a parameter vector β =(β 1,β 2,...,β 20 ) being the true probabilities of the 20 amino acids. We denote the maximum likelihood (ML) estimates of the amino acid probabilities in one alignment position as b=(b 1,b 2,...,b 20 ) and in the whole alignment as β 0 =(β1 0,β0 2,...,β0 20 ). Here, the latter ones were used as background probabilities. For the calculation of the positional conservation score, we defined the maximum Z-score statistic (Ahola et al., 2006) as c T i maxz= max (b β0 ), i=1,2,...,20 c Ti 0 c i where c T i denotes the i-th row of the amino acid scoring matrix C. The denominator ij 0 = β0 i (δ ij βj 0), i,j =1,2,...,20 n is a covariance matrix of the multinomial distribution of the amino acid frequencies under the null hypothesis H 0 :β =β 0, δ ij is the Kronecker s delta function (δ jj =1 and δ ij =0 for all i =j), and n= 20 j=1 n j is the actual number of residues at the alignment position. The final positional conservation was determined by calculating the significance of the maxz score using an importance sampling (IS) method (Rubin, 1988) and correcting the p-values by controlling the false discovery rate (FDR). The aim was to test the null hypothesis H 0 :β j =βj 0 against the alternative H A :β j βj 0, where j =1,2,...20 and at least one inequality is proper. This means that the alignment positions where the amino acid frequencies diverge from the background distribution are considered as conserved. We used as the IS distribution a mixture of multinomial distribution for 20 amino acids with parameter values α =0.4 and ɛ =0.8. The detailed description of the IS distribution, the choice of the parameter values and the entire IS procedure have been presented in Ahola et al. (2006). The p-values were adjusted for making multiple test simultaneously by applying the step-up procedure by Benjamini and Yekutieli (2001). In the step-up procedure, we used 0.05 as the q-value. Further details of the step-up procedure have been presented in Ahola et al. (2006). 2.4 Whole alignment conservation score After determining which of the alignment positions were conserved, we defined a measure for conservation of the whole alignment (Ahola et al., 2006). Instead of using the proportion of conserved positions in the alignment, we used the proportion of conserved residues, because this measure takes gaps into account. The proportion of conserved residues, ConsAA, was obtained by dividing the number of residues in the conserved positions j J by the number of residues in the whole alignment j J ConsAA= n j mi=1. n i Here, J denotes the set of conserved positions, n i the number of residues at position i and m the length of the multiple sequence alignment. The ConsAA obtains values between 0 and 1; the higher the ConsAA value the higher is the conservation of the alignment. 2.5 Predicted SP score Beta regression model In this section, we present how the quality of the alignment can be assessed using the predicted SP score. The SP score is measured on the bounded unit interval [0,1], but otherwise the true distribution of the score is unknown. Let us assume that the SP score can have (almost) any shape between 0 and 1. The natural choice in this case is to assume that the SP score follows the beta distribution. The use of the normal distribution would not have been possible, since the values of the SP score are concentrated on the upper part of the [0,1] interval, and hence the SP score would have been very difficult to transform to follow the normal distribution. Moreover, the assumption of homogeneity of variances would be violated. Since the beta distribution is defined on the open unit interval (0,1), the SP score values were transformed to open unit scale according to the formula (McMillian and Creelman, 2005) SP = SP+ 1, if SP =0, 2N and SP = SP 1, if SP =1, 2N where N denotes the number of the observations. Now SP is assumed to follow a beta distribution f ( SP;ω,τ)= Ɣ(ω+τ) Ɣ(ω)Ɣ(τ) SP ω 1 (1 SP) τ 1, where SP (0,1), ω,τ >0 and Ɣ(.) is the gamma function. Both of the parameters ω and τ of the beta distribution are shape parameters. To work with more convenient parameters for the regression model, that is, location and precision parameters, let us reparameterize the model such that µ= ω ω+τ and φ =ω+τ, which yields ω =µφ and τ =φ µφ. This implies the mean and variance of SP to be E( SP)=µ and Var( SP)= µ(1 µ) 1+φ. Here, µ can be interpreted as location and φ as precision parameter, since decrease in φ results in increased variance values for fixed µ. The beta regression model for location of SP for alignment i is g(µ i )= k x ij θ j, j=0 where θ =θ 0,θ 1,...,θ k is a vector of unknown regression parameters and x i0,...,x ik are observed predictors. In our model, x i0 = grand mean, x i1 = ConsAA, x i2 = identity, x i3 = number of sequences and x i4 = alignment length. The link function g is chosen as logit link g(µ)=ln(µ/(1 µ)). Hence, the predicted values for the location of SP can be written as µ i = exp( k j=0 x ij θ j ) 1+exp( k j=0 x ij θ j ). (1) Similarly, the beta regression model for the precision of SP is h(φ i )= t w ij γ j, (2) j=0 where h is a link function, γ =γ 0,...,γ t are the unknown precision parameters and w i0,...,w it denote the predictors of the precision model. Although the predictors of the precision model could be different from that of the location model, in our model, we define w ij =x ij, where j =0,...t, and t =k. Because the precision parameters must always be positive, the log 2167
4 V.Ahola et al. function is a suitable choice for a link function. According to Smithson and Verkuilen (2006), we added a negative sign to the formula (2) to make the interpretation easier; the negative sign turns the interpretation of φ from precision to dispersion. Hence, the predicted dispersion values for SP can be obtained by k φ i =exp w ij γ j. j= Model estimation and prediction We fitted the same beta regression model to the Mafft, Muscle and Probcons aligned training set of the Homstrad database (Fig. 1, stage 1). The models for the location and dispersion are given by g(µ i ) = θ 0 +θ 1 ConsAA i +θ 2 identity i + θ 3 number of sequences i +θ 4 alignment length i (3) h(φ i ) = γ 0 γ 1 ConsAA i γ 2 identity i γ 3 number of sequences i γ 4 alignment length i, where identity is percentage identity divided by 100 and number of sequences and alignment length are scaled to the interval [0, 1]. The parameter estimates of the location and dispersion parameters and their standard errors are shown in Table 1. It should be noted that the combined set of four Homstrad datasets were not independent of each other, which should have been taken into account using the Homstrad dataset as a random effect in the model. This was, however, computationally too hard problem for the software available. We fitted the model using the Nlmixed procedure of the SAS software package (SAS R 9.1 for Windows). The Nlmixed procedure uses quasi- Newton optimization method for estimation of the model. After estimating the parameters of the model, the g(µ i ) was calculated by replacing the θ 0,θ 1,...,θ 4 in the formula (3) with the estimated parameter values (Table 1), and the predictors with the values calculated from the alignment of interest. The predicted SP scores were obtained using the formula (1). The predicted SP scores obtain values between 0 and 1; the higher the predicted value, the better is the quality of the alignment Prediction intervals Strictly speaking, the predicted SP score is a model-based estimate of the average SP score given ConsAA and other predictor values. To take into account the uncertainty of the predicted values, the prediction intervals can be calculated by the formula ŜP±t 1 α ˆσ, where ŜP denotes the predicted SP scores, t 1 α is the 1 α s quantile of the Student s t distribution and ˆσ is the standard error of ŜP. Parameter ˆσ can be calculated using the delta method (Cox, 1998). We calculated the 95% prediction intervals for the test alignments of the Homstrad and SABmark databases using SAS/Nlmixed procedure. 2.6 Other alignment quality scores The predicted SP scores were compared with other alignment quality scores, MOS, NiRMSD and NorMD scores. These particular scores were chosen because they represent different approaches in assessing the quality of MSAs and are known to produce good results (Armougom et al., 2006; Lassmann and Sonnhammer, 2005b; Thompson et al., 2001). The MOS is based on comparing several test alignments build for the same sequence set, and calculating the proportion of alignments where all pairs of residues are similarly aligned (Lassmann and Sonnhammer, 2005b). The MOS has been implemented into the MUMSA program package. Even if we were only interested in the MOS scores of the Mafft (L-INS-i), Muscle (default) and Probcons (default) alignments, for each MUMSA run, several parallel alignments from the same set of sequences were needed. We used seven alignment programs, six of which were the same as those used in the original paper of MUMSA (Lassmann and Sonnhammer, 2005b). We run Clustal, Kalign, Muscle, Mummals, Poa and Probcons with default parameters (Do et al., 2005; Edgar, 2004; Grasso and Lee, 2004; Lassmann and Sonnhammer, 2005a; Pei and Grishin, 2006; Thompson et al., 1997). Additionally, we run Muscle with two different parameter settings (-maxiters 1 and -maxiters 2), and Mafft with four different parameter settings [-Localpair, -Localpair -maxiterate 1000 (L-INS-i), -Globalpair, -Globalpair -maxiterate 1000] (Edgar, 2004; Katoh et al., 2002, 2005). The NiRMSD score is a structure-based alignment quality score, which does not need a reference alignment, but assumes that the structures of at least two of the aligned sequences are known (Armougom et al., 2006). For the NiRMSD runs, template files including the PDB codes of the sequences were created. For some of the alignments, the NiRMSD failed to find the structure of the two sequences from the PDB database, and the NiRMSD score could not be calculated. The NiRMSD is integrated into the T_coffee package (Notredame et al., 2000). The NorMD score is a conservation-based score. The NorMD measures the mean distance between similarities of residue pairs at each alignment column (Thompson et al., 2001). The column-wise scores were summed over the entire length of the alignment and normalized to obtain the final NorMD score. To calculate the NorMD score, we used the NorMD program with 1 and 0.1 as gap opening and gap extension penalties, respectively. Table 1. Beta regression parameter estimates with standard errors and significance levels Parameter Mafft Muscle Probcons Estimate Std Error p-value Estimate Std Error p-value Estimate Std Error p-value θ < <0.01 θ < < <0.01 θ < < <0.01 θ < < <0.01 θ < < <0.01 γ < < <0.01 γ < < <0.01 γ < γ γ < < The location and dispersion parameters are for the three alignment programs. 2168
5 Prediction of sequence alignment quality 2.7 Comparison of the quality scores The predicted SP, MOS, NiRMSD and NorMD scores were compared with the CS and correct SP scores in the independent test set of the Homstrad database and with the median f d and f m scores in the SABmark database. Additional comparisons were made between the tested quality scores. These comparisons were carried out using Spearman correlation coefficient (r s ), which measures the strength of the relationship between the two scores. The agreement of the predicted SP and MOS scores with the correct SP score values and between each other were studied with the mean squared error (MSE) addressing the question of how far the actual quality score values are from each other. 3 RESULTS 3.1 Evaluation of the predicted SP score We calculated the predicted SP scores for the Mafft, Muscle and Probcons alignments in the test set of the Homstrad database and in the whole SABmark database. First, the predicted SP scores were calculated for the alignments using the Mafft, Muscle and Probcons parameters, respectively. Second, the predicted SP scores were calculated for the same Mafft, Muscle and Probcons alignments, but using parameter set obtained by different alignment program. The purpose of this analysis was to study whether the parameter sets estimated by one algorithm could be used for assessing the quality of alignments obtained by another alignment algorithm. In the first analysis, the correlation coefficients indicate moderately strong relationships between the correct and predicted SP scores in the Mafft, Muscle and Probcons alignments (r = 0.645, and 0.663, respectively). The comparisons with the CS score showed similar results, the correlations being 5% lower than that for the correct SP score (Supplementary Table 1). The second analysis suggests that the quality assessment is independent from the chosen parameter set: the correlations between the correct and predicted SP scores differed at most 2.7 % (Supplementary Table 1). The results for the CS score had more variation between the parameter sets, the mean difference in correlation being 5.6 % between the parameter sets. The agreement between the correct and predicted SP scores was evaluated by calculating the MSE between the two scores. The low MSEs of the Mafft, Muscle and Probcons alignments (MSE = ) showed on average <1% mean squared difference between the two scores, suggesting that the predicted SP scores could be used instead of the correct SP scores. Similar conclusion can be drawn from the comparisons of the individual scorings of the two methods showing that 68%, 63% and 69% of the Mafft, Muscle and Probcons alignments are within the 5% range from the correct SP value and 87%, 84%, 86% within the 10% range from the correct value, respectively. It should be noted that most of the alignments, in which the predicted SP scores diverged significantly from the correct values, were difficult alignment cases involving, for example, several insertions of different length. In the SABmark dataset, the result shows that the predicted SP score is highly correlated with the correct quality scores in the Mafft, Muscle and Probcons alignments (r fd = 0.735, and 0.720, and r fm = 0.734, and 0.694, respectively) (Supplementary Fig. 1). When repeating the analysis using different parameter sets, the correlations differed not more than 1.5% (Supplementary Table 1). The results for the twilight zone alignments showed on average about 20% decrease in the quality scores (Supplementary Table 3). Although there was still a moderately high correlation between the correct and predicted quality scores, this result indicates that distantly related protein families should be more emphasized when performing the quality scoring. Interestingly, all the alignments obtained the highest correlations when the Muscle parameters were used. 3.2 Comparison of the quality scores To compare the performance of our quality score to those of the MOS, NiRMSD and NorMD scores, the correlation coefficients were calculated between the quality scores and the correct quality scores (Table 2). As the results were similar among the alignment programs, the following results are presented as the average correlation coefficients over the Mafft, Muscle and Probcons alignments. The only exception was the NorMD, for which the results were somewhat dependent on the alignment algorithm (Table 2). When analysing the Homstrad database, the MOS score showed especially high correlation with the correct SP score (r mean = 0.87). The correlation was somewhat decreased (r mean = 0.79) when compared with the CS score (Supplementary Table 2). These correlations were even higher than that of the predicted SP score. The relationships of the correct SP score for the NorMD and NiRMSD scores were substantially lower than that of the MOS and predicted SP scores (Table 2). The predicted SP and MOS scores were strongly related (r mean = 0.80), also the NorMD score showed moderate relationship between the both MOS and the predicted SP scores, although the result of the NorMD was dependent on the alignment programs. The NiRMSD showed negative correlation between all the other scores and was not related with the conservation-based scores NorMD and predicted SP. However, it was moderately correlated with the MOS. The MOS score obtained values within the bounded interval [0,1], while the NiRMSD and NorMD scores could obtain any real values. They could not be scaled because the maximum and minimum values Table 2. Correlation coefficients for the test sets of the Homstrad (upper triangle) and SABmark (lower triangle) databases Median f d Pred SP MOS NiRMSD NorMD Correct SP 0.64, 0.65, , 0.88, , 0.58, , 0.40, 0.64 Pred SP 0.73, 0.74, , 0.80, , 0.50, , 0.52, 0.73 MOS 0.82, 0.83, , 0.92, , 0.58, , 0.51, 0.74 NiRMSD 0.85, 0.82, , 0.86, , 0.91, , 0.26, 0.45 NorMD 0.70, 0.74, , 0.82, , 0.85, , 0.88 The three coefficients in each comparison show the correlations for the Mafft, Muscle and Probcons alignment programs. 2169
6 V.Ahola et al. are unknown. Hence, the agreement with the correct SP score can only be studied with the MOS and predicted SP scores. The MSE of the MOS from the correct SP (MSE mean = 0.004) and from the predicted SP scores (MSE mean = 0.005) showed only about 0.5% mean squared difference between the MOS and the correct and predicted SP scores, indicating very low divergence between all the three scores. Although the identity of the SABmark alignments is 50% in the alignments obtained from the superfamily dataset and 25% in the alignments from the twilight zone dataset, all the benchmarked programs performed rather well (Table 2, Supplementary Fig. 1). The MOS, NiRMSD and NorMD were all highly correlated with the correct quality scores f d and f m (r mean = 0.83, 0.85,0.74, respectively for both f d and f m ). When only twilight zone alignments were considered, the quality decreased on average 13%, 7%, 26% in the MOS, NiRMSD and NorMD scores, respectively (Supplementary Table 3). This indicates that the structure-based NiRMSD is well adapted for scoring distantly related sequences, while the scorings of the NorMD are strongly affected by the increased evolutionary distance. It should be noted that the MOS had very strong relationship with the predicted SP scores (r mean = 0.91). Also, the pairwise correlations between all the other tested alignment quality measures were relatively high (Table 2). 3.3 Sensitivity to adding random sequences The ability of the alignment quality measures to distinguish between correct alignments and alignments with badly aligned or unrelated sequences was studied in the Homs10+R1, Homs10+R2, Homs10+R3 and Homs10+R4 databases. The predicted SP score correctly reacts to the subtle appearances of random sequences and could be useful in detecting badly aligned or unrelated sequences from the alignment (Supplementary Table 4). The MOS score, on the contrary, seems to overcorrect the influence of unrelated or badly aligned sequences. For more detailed evaluation of the sensitivity of the predicted SP, MOS, Nirmsd and NorMD quality scores to random sequences, see Supplementary Material. 4 DISCUSSION AND CONCLUSIONS We introduced a statistical model-based score for the alignment quality assessment. Additionally, we presented a comparison of three alignment quality scores, MOS, NiRMSD and NorMD. The results indicate that the predicted SP scores have a moderate or strong relationship with the correct SP, CS, f d and f m scores in the Homstrad and SABmark test sets. This is true both in terms of correlation coefficient, measuring the strength of relationships between the correct and predicted scores, and of the MSE, measuring the agreement between the two scores. Thus, the predicted alignment quality scores can be used to estimate the correct SP scores. Importantly, the quality of our score decreases in the same proportion as the number of unrelated sequences increases. Our preliminary results showed that the predicted SP score may also be useful in identifying badly aligned or unrelated sequences from the alignment. When comparing the predicted SP scores with the other alignment quality measures, MOS, NiRMSD and NorMD, only the MOS slightly outperformed the predicted SP score. Although the MOS score obtained more accurate results in comparison with the correct alignment quality values, the MOS score is computationally laborious, since it demands approximately 12 alignments per each test alignment. Furthermore, the method is not robust to the number of alignments or the choice of alignment programs and their parameters (Lassmann and Sonnhammer, 2005b). It should be noted, that the consistency based scores cannot distinct between related and unrelated sequences (Vingron, 1996). This can be seen in the Homs10+R1 4 databases with added random sequences, for which the MOS score penalized random sequences even 27% more than which would have been necessary. The high correlation and low MSE between the predicted SP and MOS scores indicate that our simple quality score could be used instead of the MOS to measure the quality of the alignments. Although the NorMD and NiRMSD scores also had moderately strong relationships with the correct SP score, they cannot be directly used to estimate the SP score. This is because the range of their scoring values are not limited between zero and one, and the regression curves, SP = α 0 +α 1 NorMD(NiRMSD), do not lay in the y = x curve (Supplementary Fig. 1). After making a linear transformation, the NorMD and NiRMSD scores could be used to estimate the SP score, but not using the raw scores. The NorMD and NiRMSD scores do not measure the alignment quality in terms of sensitivity, as the SP score does. The NorMD measures the conservation of the sequence alignment, whereas the NiRMSD compares pairwise structural alignments. The advantage of the NiRMSD score is that the increased evolutionary distance between the sequences had no noticeable effect on the quality scorings (Supplementary Table 3). The drawback of the NiRMSD is that it can only be used when the structure is available for at least two of the aligned proteins. It determines the quality of the alignment based only on the sequences whose structure is known ignoring all the other sequences in the alignment. Moreover, since the NiRMSD can obtain any real values, only relative comparison between the other scoring methods is possible. The advantage of the predicted SP is that it can be applied to any sequence alignment and it does not require structural information or several alignments from the same set of sequences. The prediction intervals can be calculated for the SP score in order to measure the uncertainty of prediction. We also studied the robustness of the method to the choice of the alignment methods using Mafft (L-INS-i), Muscle and Probcons alignment programs to estimate the beta regression model and then cross-validating the quality scorings of the alignments by applying three different parameter sets. Different parameter values caused only minor differences to the quality scorings indicating that any parameter set can be used. The differences arising from the alignment methods, such as the Muscle provided slightly worse results than the other two methods, were in agreement with the earlier results of benchmarking the MSA programs (Ahola et al., 2006; Nuin et al., 2006). The use of the Homstrad database enabled us to incorporate information from the whole protein-fold space into the quality scoring method. However, the use of the reference alignments may also be a limitation, because of a lack of variety of sequence alignments in terms of sequence identities and numbers of sequences (Blackshields et al., 2006). The Homstrad database contains alignments with different numbers of sequences, but mostly with high sequence identity. Since the 3D structure of only some of the proteins is known and the rest of the sequences have been aligned with ClustalW, then alignments may not always be structurally correct in all details. The Homstrad also includes some 2170
7 Prediction of sequence alignment quality difficult alignment cases which are not suitable for modelling global alignment quality. The performance of the SP score could be further improved by applying complementary reference alignment databases. However, already these results indicate that the quality of any global MSA can be evaluated using the statistical prediction model of the SP score. ACKNOWLEDGEMENTS The authors thank Pentti Riikonen for support with the Linux cluster, the Academy of Finland, the Graduate School in Computational Biology, Bioinformatics and Biometry (ComBi) and the Medical Research Fund of Tampere University Hospital. Funding: Academy of Finland ( to V.A., to T.A.). Conflict of Interest: none declared. REFERENCES Ahola,V. et al. (2006) A statistical score for assessing the quality of multiple sequence alignments. BMC Bioinformatics, 7, 484. Armougom,F. et al. (2006) The irmsd: a local measure of sequence alignment accuracy using structural information. Bioinformatics, 22, e35 e39. Bahr,A. et al. (2001) Balibase (benchmark alignment database): enhancements for repeats, transmembrane sequences and circular permutations. Nucleic Acids Res., 29, Beiko,R.G. et al. (2005) A word-oriented approach to alignment validation. Bioinformatics, 21, Benjamini,Y. and Yekutieli,D. (2001) The control of the false discovery rate in multiple testing under dependency. Ann. Stat., 29, Blackshields,G. et al. (2006) Analysis and comparison of benchmarks for multiple sequence alignment. In Silico Biol., 6, Cox,C. (1998) Delta method. In Armitage,P. and Colton,T. (eds) Encyclopedia of Biostatistics. John Wiley & Sons, Inc., New York, pp Do,C.B. et al. (2005) Probcons: probabilistic consistency-based multiple sequence alignment. Genome Res., 15, Domingues,F.S. et al. (2000) Structure-based evaluation of sequence comparison and fold recognition alignment accuracy. J. Mol. Biol., 297, Edgar,R.C. (2004) Muscle: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res., 32, Edgar,R.C. and Batzoglou,S. (2006) Multiple sequence alignment. Curr. Opin. Struct. Biol., 16, Goldsmith-Fischman,S. and Honig,B. (2003) Structural genomics: computational methods for structure analysis. Protein Sci., 12, Golubchik,T. et al. (2007) Mind the gaps: evidence of bias in estimates of multiple sequence alignments. J. Mol. Evol., 24, Gotoh,O. (1990) Consistency of optimal sequence alignments. Bull. Math. Biol., 52, Grasso,C. and Lee,C. (2004) Combining partial order alignment and progressive multiple sequence alignment increases alignment speed and scalability to very large alignment problems. Bioinformatics, 20, Hertz,G.Z. and Stormo,G.D. (1999) Identifying DNA and protein patterns with statistically significant alignments of multiple sequences. Bioinformatics, 15, Katoh,K. et al. (2002) Mafft: a novel method for rapid multiple sequence alignment based on fast fourier transform. Nucleic Acids Res., 30, Katoh,K. et al. (2005) Mafft version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res., 33, Lackner,P. et al. (2000) Prosup: a refined tool for protein structure alignment. Protein Eng., 13, Landan,G. and Graur,D. (2007) Head or tails: a simple reliability check for multiple sequence alignments. Mol. Biol. Evol., 24, Landan,G. and Graur,D. (2008) Local reliability measures from sets of co-optimal multiple sequence alignments. Pac. Symp. Biocomput., 13, Lassmann,T. and Sonnhammer,E.L. (2005a) Kalign an accurate and fast multiple sequence alignment algorithm. BMC Bioinformatics, 6, 298. Lassmann,T. and Sonnhammer,E.L.L. (2005b) Automatic assessment of alignment quality. Nucleic Acids Res., 33, McMillian,N.A. and Creelman,C.D. (2005) Detection Theory; A User s Guide. Lawrence Erlebaum Associates, Mahwah, NJ. Mizuguchi,K. et al. (1998) Homstrad: a database of protein structure alignments for homologous families. Protein Sci., 7, Morgenstern,B. et al. (2003) AltAVisT: comparing alternative multiple sequence alignments. Bioinformatics, 19, Notredame,C. (2002) Recent progress in multiple sequence alignment: a survey. Pharmacogenomics, 3, Notredame,C. (2007) Recent evolutions of multiple sequence alignment algorithms. PLoS Comput. Biol., 3, e123. Notredame,C. and Abergel,C. (2003) Using multiple alignment methods to assess the quality of genomic data analysis. In Andrade,M.A. (ed.) Bioinformatics and Genomes: Current Perspectives. Horizon Scientific Press Ltd, Norwich, UK, pp Notredame,C. et al. (2000) T-coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol., 302, Nuin,P. et al. (2006) The accuracy of several multiple sequence alignment programs for proteins. BMC Bioinformatics, 7, 471. Pei,J. and Grishin,N.V. (2006) Mummals: multiple sequence alignment improved by using hidden Markov models with local structural information. Nucleic Acids Res., 34, Pei,J. and Grishin,N.V. (2007) Promals: towards accurate multiple sequence alignments of distantly related proteins. Bioinformatics, 23, Pei,J.M. and Grishin,N.V. (2001) Al2co: calculation of positional conservation in a protein sequence alignment. Bioinformatics, 17, Rubin,D.B. (1988) Using the sir algorithm to simulate posterior distributions. In Bernardo,M.H. et al. (eds) Bayesian Statistics 3. Oxford University Press, Oxford, UK, pp Sauder,J.M. et al. (2000) Large-scale comparison of protein sequence alignment algorithms with structure alignments. Proteins, 40, Smithson,M. and Verkuilen,J. (2006) A better lemon squeezer? Maximum-likelihood regression with beta-distributed dependent variables. Psychol. Methods, 11, Thompson,J.D. et al. (1997) The clustal_x windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res., 25, Thompson,J.D. et al. (1999) A comprehensive comparison of multiple sequence alignment programs. Nucleic Acids Res., 27, Thompson,J.D. et al. (2001) Towards a reliable objective function for multiple sequence alignments. J. Mol. Biol., 314, Vingron,M. (1996) Near-optimal sequence alignment. Curr. Opin. Struct. Biol., 6, Vingron,M. and Argos,P. (1991) Motif recognition and alignment for many sequences by comparison of dot-matrices. J. Mol. Biol., 218, Walle,I.V. et al. (2005) Sabmark a benchmark for sequence alignment that covers the entire known fold space. Bioinformatics, 21,
BIOINFORMATICS ORIGINAL PAPER doi: /bioinformatics/btm017
 Vol. 23 no. 7 2007, pages 802 808 BIOINFORMATICS ORIGINAL PAPER doi:10.1093/bioinformatics/btm017 Sequence analysis PROMALS: towards accurate multiple sequence alignments of distantly related proteins
Vol. 23 no. 7 2007, pages 802 808 BIOINFORMATICS ORIGINAL PAPER doi:10.1093/bioinformatics/btm017 Sequence analysis PROMALS: towards accurate multiple sequence alignments of distantly related proteins
PROMALS3D: a tool for multiple protein sequence and structure alignments
 Nucleic Acids Research Advance Access published February 20, 2008 Nucleic Acids Research, 2008, 1 6 doi:10.1093/nar/gkn072 PROMALS3D: a tool for multiple protein sequence and structure alignments Jimin
Nucleic Acids Research Advance Access published February 20, 2008 Nucleic Acids Research, 2008, 1 6 doi:10.1093/nar/gkn072 PROMALS3D: a tool for multiple protein sequence and structure alignments Jimin
Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences
 Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences Yue Lu 1 and Sing-Hoi Sze 1,2 1 Department of Biochemistry & Biophysics 2 Department of Computer Science, Texas A&M University,
Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences Yue Lu 1 and Sing-Hoi Sze 1,2 1 Department of Biochemistry & Biophysics 2 Department of Computer Science, Texas A&M University,
Multiple Sequence Alignment: A Critical Comparison of Four Popular Programs
 Multiple Sequence Alignment: A Critical Comparison of Four Popular Programs Shirley Sutton, Biochemistry 218 Final Project, March 14, 2008 Introduction For both the computational biologist and the research
Multiple Sequence Alignment: A Critical Comparison of Four Popular Programs Shirley Sutton, Biochemistry 218 Final Project, March 14, 2008 Introduction For both the computational biologist and the research
PROMALS3D web server for accurate multiple protein sequence and structure alignments
 W30 W34 Nucleic Acids Research, 2008, Vol. 36, Web Server issue Published online 24 May 2008 doi:10.1093/nar/gkn322 PROMALS3D web server for accurate multiple protein sequence and structure alignments
W30 W34 Nucleic Acids Research, 2008, Vol. 36, Web Server issue Published online 24 May 2008 doi:10.1093/nar/gkn322 PROMALS3D web server for accurate multiple protein sequence and structure alignments
Probalign: Multiple sequence alignment using partition function posterior probabilities
 Sequence Analysis Probalign: Multiple sequence alignment using partition function posterior probabilities Usman Roshan 1* and Dennis R. Livesay 2 1 Department of Computer Science, New Jersey Institute
Sequence Analysis Probalign: Multiple sequence alignment using partition function posterior probabilities Usman Roshan 1* and Dennis R. Livesay 2 1 Department of Computer Science, New Jersey Institute
Multiple sequence alignment
 Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
BIOINFORMATICS. RASCAL: rapid scanning and correction of multiple sequence alignments. J. D. Thompson, J. C. Thierry and O.
 BIOINFORMATICS Vol. 19 no. 9 2003, pages 1155 1161 DOI: 10.1093/bioinformatics/btg133 RASCAL: rapid scanning and correction of multiple sequence alignments J. D. Thompson, J. C. Thierry and O. Poch Laboratoire
BIOINFORMATICS Vol. 19 no. 9 2003, pages 1155 1161 DOI: 10.1093/bioinformatics/btg133 RASCAL: rapid scanning and correction of multiple sequence alignments J. D. Thompson, J. C. Thierry and O. Poch Laboratoire
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Protein sequence alignment with family-specific amino acid similarity matrices
 TECHNICAL NOTE Open Access Protein sequence alignment with family-specific amino acid similarity matrices Igor B Kuznetsov Abstract Background: Alignment of amino acid sequences by means of dynamic programming
TECHNICAL NOTE Open Access Protein sequence alignment with family-specific amino acid similarity matrices Igor B Kuznetsov Abstract Background: Alignment of amino acid sequences by means of dynamic programming
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Structure Comparison
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
CMPS 6630: Introduction to Computational Biology and Bioinformatics Structure Comparison Protein Structure Comparison Motivation Understand sequence and structure variability Understand Domain architecture
Some Problems from Enzyme Families
 Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Upcoming challenges for multiple sequence alignment methods in the high-throughput era
 BIOINFORMATICS REVIEW Vol. 25 no. 19 2009, pages 2455 2465 doi:10.1093/bioinformatics/btp452 Sequence analysis Upcoming challenges for multiple sequence alignment methods in the high-throughput era Carsten
BIOINFORMATICS REVIEW Vol. 25 no. 19 2009, pages 2455 2465 doi:10.1093/bioinformatics/btp452 Sequence analysis Upcoming challenges for multiple sequence alignment methods in the high-throughput era Carsten
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
InDel 3-5. InDel 8-9. InDel 3-5. InDel 8-9. InDel InDel 8-9
 Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT
 22 Genome Informatics 16(1): 22-33 (2005) Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT Kazutaka Katohl kkatoh@kuicr.kyoto-u.ac.jp Takashi Miyata2" miyata@brh.co.jp Kei-ichi
22 Genome Informatics 16(1): 22-33 (2005) Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT Kazutaka Katohl kkatoh@kuicr.kyoto-u.ac.jp Takashi Miyata2" miyata@brh.co.jp Kei-ichi
Week 10: Homology Modelling (II) - HHpred
 Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Chapter 11 Multiple sequence alignment
 Chapter 11 Multiple sequence alignment Burkhard Morgenstern 1. INTRODUCTION Sequence alignment is of crucial importance for all aspects of biological sequence analysis. Virtually all methods of nucleic
Chapter 11 Multiple sequence alignment Burkhard Morgenstern 1. INTRODUCTION Sequence alignment is of crucial importance for all aspects of biological sequence analysis. Virtually all methods of nucleic
Sequence Alignment: A General Overview. COMP Fall 2010 Luay Nakhleh, Rice University
 Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
A profile-based protein sequence alignment algorithm for a domain clustering database
 A profile-based protein sequence alignment algorithm for a domain clustering database Lin Xu,2 Fa Zhang and Zhiyong Liu 3, Key Laboratory of Computer System and architecture, the Institute of Computing
A profile-based protein sequence alignment algorithm for a domain clustering database Lin Xu,2 Fa Zhang and Zhiyong Liu 3, Key Laboratory of Computer System and architecture, the Institute of Computing
Sequence Alignment Techniques and Their Uses
 Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
An Introduction to Sequence Similarity ( Homology ) Searching
 An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
CONCEPT OF SEQUENCE COMPARISON. Natapol Pornputtapong 18 January 2018
 CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
The PRALINE online server: optimising progressive multiple alignment on the web
 Computational Biology and Chemistry 27 (2003) 511 519 Software Note The PRALINE online server: optimising progressive multiple alignment on the web V.A. Simossis a,b, J. Heringa a, a Bioinformatics Unit,
Computational Biology and Chemistry 27 (2003) 511 519 Software Note The PRALINE online server: optimising progressive multiple alignment on the web V.A. Simossis a,b, J. Heringa a, a Bioinformatics Unit,
Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling
 Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 009 Mark Craven craven@biostat.wisc.edu Sequence Motifs what is a sequence
Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 009 Mark Craven craven@biostat.wisc.edu Sequence Motifs what is a sequence
BLAST: Target frequencies and information content Dannie Durand
 Computational Genomics and Molecular Biology, Fall 2016 1 BLAST: Target frequencies and information content Dannie Durand BLAST has two components: a fast heuristic for searching for similar sequences
Computational Genomics and Molecular Biology, Fall 2016 1 BLAST: Target frequencies and information content Dannie Durand BLAST has two components: a fast heuristic for searching for similar sequences
Introduction to Bioinformatics Online Course: IBT
 Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Introduction to Bioinformatics Online Course: IBT Multiple Sequence Alignment Building Multiple Sequence Alignment Lec1 Building a Multiple Sequence Alignment Learning Outcomes 1- Understanding Why multiple
Mul$ple Sequence Alignment Methods. Tandy Warnow Departments of Bioengineering and Computer Science h?p://tandy.cs.illinois.edu
 Mul$ple Sequence Alignment Methods Tandy Warnow Departments of Bioengineering and Computer Science h?p://tandy.cs.illinois.edu Species Tree Orangutan Gorilla Chimpanzee Human From the Tree of the Life
Mul$ple Sequence Alignment Methods Tandy Warnow Departments of Bioengineering and Computer Science h?p://tandy.cs.illinois.edu Species Tree Orangutan Gorilla Chimpanzee Human From the Tree of the Life
Simultaneous Sequence Alignment and Tree Construction Using Hidden Markov Models. R.C. Edgar, K. Sjölander
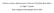 Simultaneous Sequence Alignment and Tree Construction Using Hidden Markov Models R.C. Edgar, K. Sjölander Pacific Symposium on Biocomputing 8:180-191(2003) SIMULTANEOUS SEQUENCE ALIGNMENT AND TREE CONSTRUCTION
Simultaneous Sequence Alignment and Tree Construction Using Hidden Markov Models R.C. Edgar, K. Sjölander Pacific Symposium on Biocomputing 8:180-191(2003) SIMULTANEOUS SEQUENCE ALIGNMENT AND TREE CONSTRUCTION
Copyright 2000 N. AYDIN. All rights reserved. 1
 Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Machine Learning for OR & FE
 Machine Learning for OR & FE Regression II: Regularization and Shrinkage Methods Martin Haugh Department of Industrial Engineering and Operations Research Columbia University Email: martin.b.haugh@gmail.com
Machine Learning for OR & FE Regression II: Regularization and Shrinkage Methods Martin Haugh Department of Industrial Engineering and Operations Research Columbia University Email: martin.b.haugh@gmail.com
Homology Modeling (Comparative Structure Modeling) GBCB 5874: Problem Solving in GBCB
 Homology Modeling (Comparative Structure Modeling) Aims of Structural Genomics High-throughput 3D structure determination and analysis To determine or predict the 3D structures of all the proteins encoded
Homology Modeling (Comparative Structure Modeling) Aims of Structural Genomics High-throughput 3D structure determination and analysis To determine or predict the 3D structures of all the proteins encoded
Machine Learning Linear Classification. Prof. Matteo Matteucci
 Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Machine Learning Linear Classification Prof. Matteo Matteucci Recall from the first lecture 2 X R p Regression Y R Continuous Output X R p Y {Ω 0, Ω 1,, Ω K } Classification Discrete Output X R p Y (X)
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche
 Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT
 5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
What is the expectation maximization algorithm?
 primer 2008 Nature Publishing Group http://www.nature.com/naturebiotechnology What is the expectation maximization algorithm? Chuong B Do & Serafim Batzoglou The expectation maximization algorithm arises
primer 2008 Nature Publishing Group http://www.nature.com/naturebiotechnology What is the expectation maximization algorithm? Chuong B Do & Serafim Batzoglou The expectation maximization algorithm arises
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
A greedy, graph-based algorithm for the alignment of multiple homologous gene lists
 A greedy, graph-based algorithm for the alignment of multiple homologous gene lists Jan Fostier, Sebastian Proost, Bart Dhoedt, Yvan Saeys, Piet Demeester, Yves Van de Peer, and Klaas Vandepoele Bioinformatics
A greedy, graph-based algorithm for the alignment of multiple homologous gene lists Jan Fostier, Sebastian Proost, Bart Dhoedt, Yvan Saeys, Piet Demeester, Yves Van de Peer, and Klaas Vandepoele Bioinformatics
Multiple Mapping Method: A Novel Approach to the Sequence-to-Structure Alignment Problem in Comparative Protein Structure Modeling
 63:644 661 (2006) Multiple Mapping Method: A Novel Approach to the Sequence-to-Structure Alignment Problem in Comparative Protein Structure Modeling Brajesh K. Rai and András Fiser* Department of Biochemistry
63:644 661 (2006) Multiple Mapping Method: A Novel Approach to the Sequence-to-Structure Alignment Problem in Comparative Protein Structure Modeling Brajesh K. Rai and András Fiser* Department of Biochemistry
proteins Refinement by shifting secondary structure elements improves sequence alignments
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Refinement by shifting secondary structure elements improves sequence alignments Jing Tong, 1,2 Jimin Pei, 3 Zbyszek Otwinowski, 1,2 and Nick V. Grishin 1,2,3
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Refinement by shifting secondary structure elements improves sequence alignments Jing Tong, 1,2 Jimin Pei, 3 Zbyszek Otwinowski, 1,2 and Nick V. Grishin 1,2,3
Gibbs Sampling Methods for Multiple Sequence Alignment
 Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
Local Alignment Statistics
 Local Alignment Statistics Stephen Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health Bethesda, MD Central Issues in Biological Sequence Comparison
Local Alignment Statistics Stephen Altschul National Center for Biotechnology Information National Library of Medicine National Institutes of Health Bethesda, MD Central Issues in Biological Sequence Comparison
Plausible Values for Latent Variables Using Mplus
 Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Plausible Values for Latent Variables Using Mplus Tihomir Asparouhov and Bengt Muthén August 21, 2010 1 1 Introduction Plausible values are imputed values for latent variables. All latent variables can
Giri Narasimhan. CAP 5510: Introduction to Bioinformatics. ECS 254; Phone: x3748
 CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/15/07 CAP5510 1 EM Algorithm Goal: Find θ, Z that maximize Pr
CAP 5510: Introduction to Bioinformatics Giri Narasimhan ECS 254; Phone: x3748 giri@cis.fiu.edu www.cis.fiu.edu/~giri/teach/bioinfs07.html 2/15/07 CAP5510 1 EM Algorithm Goal: Find θ, Z that maximize Pr
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
In-Depth Assessment of Local Sequence Alignment
 2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Supplementary text for the section Interactions conserved across species: can one select the conserved interactions?
 1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
Sequence analysis and comparison
 The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
The aim with sequence identification: Sequence analysis and comparison Marjolein Thunnissen Lund September 2012 Is there any known protein sequence that is homologous to mine? Are there any other species
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
proteins Estimating quality of template-based protein models by alignment stability Hao Chen 1 and Daisuke Kihara 1,2,3,4 * INTRODUCTION
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Estimating quality of template-based protein models by alignment stability Hao Chen 1 and Daisuke Kihara 1,2,3,4 * 1 Department of Biological Sciences, College
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Estimating quality of template-based protein models by alignment stability Hao Chen 1 and Daisuke Kihara 1,2,3,4 * 1 Department of Biological Sciences, College
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data
 DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
DEGseq: an R package for identifying differentially expressed genes from RNA-seq data Likun Wang Zhixing Feng i Wang iaowo Wang * and uegong Zhang * MOE Key Laboratory of Bioinformatics and Bioinformatics
Multiple Alignment. Anders Gorm Pedersen Molecular Evolution Group Center for Biological Sequence Analysis
 Multiple Alignment Anders Gorm Pedersen Molecular Evolution Group Center for Biological Sequence Analysis gorm@cbs.dtu.dk Refresher: pairwise alignments 43.2% identity; Global alignment score: 374 10 20
Multiple Alignment Anders Gorm Pedersen Molecular Evolution Group Center for Biological Sequence Analysis gorm@cbs.dtu.dk Refresher: pairwise alignments 43.2% identity; Global alignment score: 374 10 20
A New Similarity Measure among Protein Sequences
 A New Similarity Measure among Protein Sequences Kuen-Pin Wu, Hsin-Nan Lin, Ting-Yi Sung and Wen-Lian Hsu * Institute of Information Science Academia Sinica, Taipei 115, Taiwan Abstract Protein sequence
A New Similarity Measure among Protein Sequences Kuen-Pin Wu, Hsin-Nan Lin, Ting-Yi Sung and Wen-Lian Hsu * Institute of Information Science Academia Sinica, Taipei 115, Taiwan Abstract Protein sequence
Grouping of amino acids and recognition of protein structurally conserved regions by reduced alphabets of amino acids
 Science in China Series C: Life Sciences 2007 Science in China Press Springer-Verlag Grouping of amino acids and recognition of protein structurally conserved regions by reduced alphabets of amino acids
Science in China Series C: Life Sciences 2007 Science in China Press Springer-Verlag Grouping of amino acids and recognition of protein structurally conserved regions by reduced alphabets of amino acids
Sequence Bioinformatics. Multiple Sequence Alignment Waqas Nasir
 Sequence Bioinformatics Multiple Sequence Alignment Waqas Nasir 2010-11-12 Multiple Sequence Alignment One amino acid plays coy; a pair of homologous sequences whisper; many aligned sequences shout out
Sequence Bioinformatics Multiple Sequence Alignment Waqas Nasir 2010-11-12 Multiple Sequence Alignment One amino acid plays coy; a pair of homologous sequences whisper; many aligned sequences shout out
CAP 5510 Lecture 3 Protein Structures
 CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
CAP 5510 Lecture 3 Protein Structures Su-Shing Chen Bioinformatics CISE 8/19/2005 Su-Shing Chen, CISE 1 Protein Conformation 8/19/2005 Su-Shing Chen, CISE 2 Protein Conformational Structures Hydrophobicity
Chapter 5. Proteomics and the analysis of protein sequence Ⅱ
 Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Proteomics Chapter 5. Proteomics and the analysis of protein sequence Ⅱ 1 Pairwise similarity searching (1) Figure 5.5: manual alignment One of the amino acids in the top sequence has no equivalent and
Research Proposal. Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family.
 Research Proposal Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family. Name: Minjal Pancholi Howard University Washington, DC. June 19, 2009 Research
Research Proposal Title: Multiple Sequence Alignment used to investigate the co-evolving positions in OxyR Protein family. Name: Minjal Pancholi Howard University Washington, DC. June 19, 2009 Research
Modelling heterogeneous variance-covariance components in two-level multilevel models with application to school effects educational research
 Modelling heterogeneous variance-covariance components in two-level multilevel models with application to school effects educational research Research Methods Festival Oxford 9 th July 014 George Leckie
Modelling heterogeneous variance-covariance components in two-level multilevel models with application to school effects educational research Research Methods Festival Oxford 9 th July 014 George Leckie
Overview Multiple Sequence Alignment
 Overview Multiple Sequence Alignment Inge Jonassen Bioinformatics group Dept. of Informatics, UoB Inge.Jonassen@ii.uib.no Definition/examples Use of alignments The alignment problem scoring alignments
Overview Multiple Sequence Alignment Inge Jonassen Bioinformatics group Dept. of Informatics, UoB Inge.Jonassen@ii.uib.no Definition/examples Use of alignments The alignment problem scoring alignments
Quantifying sequence similarity
 Quantifying sequence similarity Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 16 th 2016 After this lecture, you can define homology, similarity, and identity
Quantifying sequence similarity Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, February 16 th 2016 After this lecture, you can define homology, similarity, and identity
Protein Bioinformatics. Rickard Sandberg Dept. of Cell and Molecular Biology Karolinska Institutet sandberg.cmb.ki.
 Protein Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology Karolinska Institutet rickard.sandberg@ki.se sandberg.cmb.ki.se Outline Protein features motifs patterns profiles signals 2 Protein
Protein Bioinformatics Rickard Sandberg Dept. of Cell and Molecular Biology Karolinska Institutet rickard.sandberg@ki.se sandberg.cmb.ki.se Outline Protein features motifs patterns profiles signals 2 Protein
Fall 2017 STAT 532 Homework Peter Hoff. 1. Let P be a probability measure on a collection of sets A.
 1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
1. Let P be a probability measure on a collection of sets A. (a) For each n N, let H n be a set in A such that H n H n+1. Show that P (H n ) monotonically converges to P ( k=1 H k) as n. (b) For each n
Pairwise & Multiple sequence alignments
 Pairwise & Multiple sequence alignments Urmila Kulkarni-Kale Bioinformatics Centre 411 007 urmila@bioinfo.ernet.in Basis for Sequence comparison Theory of evolution: gene sequences have evolved/derived
Pairwise & Multiple sequence alignments Urmila Kulkarni-Kale Bioinformatics Centre 411 007 urmila@bioinfo.ernet.in Basis for Sequence comparison Theory of evolution: gene sequences have evolved/derived
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Dimension Reduction Methods
 Dimension Reduction Methods And Bayesian Machine Learning Marek Petrik 2/28 Previously in Machine Learning How to choose the right features if we have (too) many options Methods: 1. Subset selection 2.
Dimension Reduction Methods And Bayesian Machine Learning Marek Petrik 2/28 Previously in Machine Learning How to choose the right features if we have (too) many options Methods: 1. Subset selection 2.
Citation BMC Bioinformatics, 2016, v. 17 n. suppl. 8, p. 285:
 Title PnpProbs: A better multiple sequence alignment tool by better handling of guide trees Author(s) YE, Y; Lam, TW; Ting, HF Citation BMC Bioinformatics, 2016, v. 17 n. suppl. 8, p. 285:633-643 Issued
Title PnpProbs: A better multiple sequence alignment tool by better handling of guide trees Author(s) YE, Y; Lam, TW; Ting, HF Citation BMC Bioinformatics, 2016, v. 17 n. suppl. 8, p. 285:633-643 Issued
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Low-Level Analysis of High- Density Oligonucleotide Microarray Data
 Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Low-Level Analysis of High- Density Oligonucleotide Microarray Data Ben Bolstad http://www.stat.berkeley.edu/~bolstad Biostatistics, University of California, Berkeley UC Berkeley Feb 23, 2004 Outline
Who Watches the Watchmen? An Appraisal of Benchmarks for Multiple Sequence Alignment
 Who Watches the Watchmen? An Appraisal of Benchmarks for Multiple Sequence Alignment Stefano Iantorno 1,2,*, Kevin Gori 3,*, Nick Goldman 3, Manuel Gil 4, Christophe Dessimoz 3, 1 Wellcome Trust Sanger
Who Watches the Watchmen? An Appraisal of Benchmarks for Multiple Sequence Alignment Stefano Iantorno 1,2,*, Kevin Gori 3,*, Nick Goldman 3, Manuel Gil 4, Christophe Dessimoz 3, 1 Wellcome Trust Sanger
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008
 Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM)
 Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
Bioinformatics II Probability and Statistics Universität Zürich and ETH Zürich Spring Semester 2009 Lecture 4: Evolutionary Models and Substitution Matrices (PAM and BLOSUM) Dr Fraser Daly adapted from
MACAU 2.0 User Manual
 MACAU 2.0 User Manual Shiquan Sun, Jiaqiang Zhu, and Xiang Zhou Department of Biostatistics, University of Michigan shiquans@umich.edu and xzhousph@umich.edu April 9, 2017 Copyright 2016 by Xiang Zhou
MACAU 2.0 User Manual Shiquan Sun, Jiaqiang Zhu, and Xiang Zhou Department of Biostatistics, University of Michigan shiquans@umich.edu and xzhousph@umich.edu April 9, 2017 Copyright 2016 by Xiang Zhou
Reducing storage requirements for biological sequence comparison
 Bioinformatics Advance Access published July 15, 2004 Bioinfor matics Oxford University Press 2004; all rights reserved. Reducing storage requirements for biological sequence comparison Michael Roberts,
Bioinformatics Advance Access published July 15, 2004 Bioinfor matics Oxford University Press 2004; all rights reserved. Reducing storage requirements for biological sequence comparison Michael Roberts,
2 Dean C. Adams and Gavin J. P. Naylor the best three-dimensional ordination of the structure space is found through an eigen-decomposition (correspon
 A Comparison of Methods for Assessing the Structural Similarity of Proteins Dean C. Adams and Gavin J. P. Naylor? Dept. Zoology and Genetics, Iowa State University, Ames, IA 50011, U.S.A. 1 Introduction
A Comparison of Methods for Assessing the Structural Similarity of Proteins Dean C. Adams and Gavin J. P. Naylor? Dept. Zoology and Genetics, Iowa State University, Ames, IA 50011, U.S.A. 1 Introduction
Subfamily HMMS in Functional Genomics. D. Brown, N. Krishnamurthy, J.M. Dale, W. Christopher, and K. Sjölander
 Subfamily HMMS in Functional Genomics D. Brown, N. Krishnamurthy, J.M. Dale, W. Christopher, and K. Sjölander Pacific Symposium on Biocomputing 10:322-333(2005) SUBFAMILY HMMS IN FUNCTIONAL GENOMICS DUNCAN
Subfamily HMMS in Functional Genomics D. Brown, N. Krishnamurthy, J.M. Dale, W. Christopher, and K. Sjölander Pacific Symposium on Biocomputing 10:322-333(2005) SUBFAMILY HMMS IN FUNCTIONAL GENOMICS DUNCAN
Dispersion modeling for RNAseq differential analysis
 Dispersion modeling for RNAseq differential analysis E. Bonafede 1, F. Picard 2, S. Robin 3, C. Viroli 1 ( 1 ) univ. Bologna, ( 3 ) CNRS/univ. Lyon I, ( 3 ) INRA/AgroParisTech, Paris IBC, Victoria, July
Dispersion modeling for RNAseq differential analysis E. Bonafede 1, F. Picard 2, S. Robin 3, C. Viroli 1 ( 1 ) univ. Bologna, ( 3 ) CNRS/univ. Lyon I, ( 3 ) INRA/AgroParisTech, Paris IBC, Victoria, July
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
Model Estimation Example
 Ronald H. Heck 1 EDEP 606: Multivariate Methods (S2013) April 7, 2013 Model Estimation Example As we have moved through the course this semester, we have encountered the concept of model estimation. Discussions
Ronald H. Heck 1 EDEP 606: Multivariate Methods (S2013) April 7, 2013 Model Estimation Example As we have moved through the course this semester, we have encountered the concept of model estimation. Discussions
Exercise 5. Sequence Profiles & BLAST
 Exercise 5 Sequence Profiles & BLAST 1 Substitution Matrix (BLOSUM62) Likelihood to substitute one amino acid with another Figure taken from https://en.wikipedia.org/wiki/blosum 2 Substitution Matrix (BLOSUM62)
Exercise 5 Sequence Profiles & BLAST 1 Substitution Matrix (BLOSUM62) Likelihood to substitute one amino acid with another Figure taken from https://en.wikipedia.org/wiki/blosum 2 Substitution Matrix (BLOSUM62)
Lecture 2, 5/12/2001: Local alignment the Smith-Waterman algorithm. Alignment scoring schemes and theory: substitution matrices and gap models
 Lecture 2, 5/12/2001: Local alignment the Smith-Waterman algorithm Alignment scoring schemes and theory: substitution matrices and gap models 1 Local sequence alignments Local sequence alignments are necessary
Lecture 2, 5/12/2001: Local alignment the Smith-Waterman algorithm Alignment scoring schemes and theory: substitution matrices and gap models 1 Local sequence alignments Local sequence alignments are necessary
Phylogenetics: Bayesian Phylogenetic Analysis. COMP Spring 2015 Luay Nakhleh, Rice University
 Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Similarity searching summary (2)
 Similarity searching / sequence alignment summary Biol4230 Thurs, February 22, 2016 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 What have we covered? Homology excess similiarity but no excess similarity
Similarity searching / sequence alignment summary Biol4230 Thurs, February 22, 2016 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 What have we covered? Homology excess similiarity but no excess similarity
TGDR: An Introduction
 TGDR: An Introduction Julian Wolfson Student Seminar March 28, 2007 1 Variable Selection 2 Penalization, Solution Paths and TGDR 3 Applying TGDR 4 Extensions 5 Final Thoughts Some motivating examples We
TGDR: An Introduction Julian Wolfson Student Seminar March 28, 2007 1 Variable Selection 2 Penalization, Solution Paths and TGDR 3 Applying TGDR 4 Extensions 5 Final Thoughts Some motivating examples We
Mixed- Model Analysis of Variance. Sohad Murrar & Markus Brauer. University of Wisconsin- Madison. Target Word Count: Actual Word Count: 2755
 Mixed- Model Analysis of Variance Sohad Murrar & Markus Brauer University of Wisconsin- Madison The SAGE Encyclopedia of Educational Research, Measurement and Evaluation Target Word Count: 3000 - Actual
Mixed- Model Analysis of Variance Sohad Murrar & Markus Brauer University of Wisconsin- Madison The SAGE Encyclopedia of Educational Research, Measurement and Evaluation Target Word Count: 3000 - Actual
STAT 461/561- Assignments, Year 2015
 STAT 461/561- Assignments, Year 2015 This is the second set of assignment problems. When you hand in any problem, include the problem itself and its number. pdf are welcome. If so, use large fonts and
STAT 461/561- Assignments, Year 2015 This is the second set of assignment problems. When you hand in any problem, include the problem itself and its number. pdf are welcome. If so, use large fonts and
Large-Scale Genomic Surveys
 Bioinformatics Subtopics Fold Recognition Secondary Structure Prediction Docking & Drug Design Protein Geometry Protein Flexibility Homology Modeling Sequence Alignment Structure Classification Gene Prediction
Bioinformatics Subtopics Fold Recognition Secondary Structure Prediction Docking & Drug Design Protein Geometry Protein Flexibility Homology Modeling Sequence Alignment Structure Classification Gene Prediction
Computational methods for predicting protein-protein interactions
 Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Computational methods for predicting protein-protein interactions Tomi Peltola T-61.6070 Special course in bioinformatics I 3.4.2008 Outline Biological background Protein-protein interactions Computational
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment
 Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
LOGISTIC REGRESSION Joseph M. Hilbe
 LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
LOGISTIC REGRESSION Joseph M. Hilbe Arizona State University Logistic regression is the most common method used to model binary response data. When the response is binary, it typically takes the form of
Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics
 Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics Tandy Warnow Founder Professor of Engineering The University of Illinois at Urbana-Champaign http://tandy.cs.illinois.edu
Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics Tandy Warnow Founder Professor of Engineering The University of Illinois at Urbana-Champaign http://tandy.cs.illinois.edu
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Sparse Linear Models (10/7/13)
 STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
STA56: Probabilistic machine learning Sparse Linear Models (0/7/) Lecturer: Barbara Engelhardt Scribes: Jiaji Huang, Xin Jiang, Albert Oh Sparsity Sparsity has been a hot topic in statistics and machine
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
frmsdalign: Protein Sequence Alignment Using Predicted Local Structure Information for Pairs with Low Sequence Identity
 1 frmsdalign: Protein Sequence Alignment Using Predicted Local Structure Information for Pairs with Low Sequence Identity HUZEFA RANGWALA and GEORGE KARYPIS Department of Computer Science and Engineering
1 frmsdalign: Protein Sequence Alignment Using Predicted Local Structure Information for Pairs with Low Sequence Identity HUZEFA RANGWALA and GEORGE KARYPIS Department of Computer Science and Engineering
LOCAL RELIABILITY MEASURES FROM SETS OF CO-OPTIMAL MULTIPLE SEQUENCE ALIGNMENTS
 LOCAL RELIABILITY MEASURES FROM SETS OF CO-OPTIMAL MULTIPLE SEQUENCE ALIGNMENTS GIDDY LANDAN DAN GRAUR Department of Biology & Biochemistry, University of Houston, Houston, TX 77204 The question of multiple
LOCAL RELIABILITY MEASURES FROM SETS OF CO-OPTIMAL MULTIPLE SEQUENCE ALIGNMENTS GIDDY LANDAN DAN GRAUR Department of Biology & Biochemistry, University of Houston, Houston, TX 77204 The question of multiple
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of omputer Science San José State University San José, alifornia, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Pairwise Sequence Alignment Homology
Algorithms in Bioinformatics Sami Khuri Department of omputer Science San José State University San José, alifornia, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Pairwise Sequence Alignment Homology
HAlign: Fast multiple similar DNA/RNA sequence alignment based on the centre star strategy
 Bioinformatics, 31(15), 2015, 2475 2481 doi: 10.1093/bioinformatics/btv177 Advance Access Publication Date: 25 March 2015 Original Paper Sequence analysis HAlign: Fast multiple similar DNA/RNA sequence
Bioinformatics, 31(15), 2015, 2475 2481 doi: 10.1093/bioinformatics/btv177 Advance Access Publication Date: 25 March 2015 Original Paper Sequence analysis HAlign: Fast multiple similar DNA/RNA sequence
