BIOINFORMATICS ORIGINAL PAPER doi: /bioinformatics/btm017
|
|
|
- Maximillian Dixon
- 5 years ago
- Views:
Transcription
1 Vol. 23 no , pages BIOINFORMATICS ORIGINAL PAPER doi: /bioinformatics/btm017 Sequence analysis PROMALS: towards accurate multiple sequence alignments of distantly related proteins Jimin Pei 1, and Nick V. Grishin 1,2 1 Howard Hughes Medical Institute and 2 Department of Biochemistry, The University of Texas Southwestern Medical Center at Dallas, 6001 Forest Park Road, Dallas, TX , USA Received on December 4, 2006; revised on January 12, 2007; accepted on January 17, 2007 Advance Access publication January 31, 2007 Associate Editor: Alex Bateman ABSTRACT Motivation: Accurate multiple sequence alignments are essential in protein structure modeling, functional prediction and efficient planning of experiments. Although the alignment problem has attracted considerable attention, preparation of high-quality alignments for distantly related sequences remains a difficult task. Results: We developed PROMALS, a multiple alignment method that shows promising results for protein homologs with sequence identity below 10%, aligning close to half of the amino acid residues correctly on average. This is about three times more accurate than traditional pairwise sequence alignment methods. PROMALS algorithm derives its strength from several sources: (i) sequence database searches to retrieve additional homologs; (ii) accurate secondary structure prediction; (iii) a hidden Markov model that uses a novel combined scoring of amino acids and secondary structures; (iv) probabilistic consistency-based scoring applied to progressive alignment of profiles. Compared to the best alignment methods that do not use secondary structure prediction and database searches (e.g. MUMMALS, ProbCons and MAFFT), PROMALS is up to 30% more accurate, with improvement being most prominent for highly divergent homologs. Compared to SPEM and HHalign, which also employ database searches and secondary structure prediction, PROMALS shows an accuracy improvement of several percent. Availability: The PROMALS web server is available at: Contact: jpei@chop.swmed.edu Supplementary information: Supplementary data are available at Bioinformatics online. 1 INTRODUCTION Multiple sequence alignments have broad applications in sequence similarity searches, structure modeling and phylogenetic analysis (Altschul et al., 1997; Eddy, 1998; Ginalski and Rychlewski, 2003; Phillips et al., 2000). They also aid in experimental design by revealing conserved residues with potential functional importance. A variety of alignment methods that rely on different algorithms and scoring functions have been developed (Edgar and Batzoglou, 2006). A rigorous method that aligns all sequences simultaneously *To whom correspondence should be addressed. (Lipman et al., 1989) is computationally prohibitive for large sets of sequences. In contrast, a progressive method that aligns pairs of sequences and sequence groups along a tree is algorithmically simpler and much faster, requiring only N 1 steps of pairwise alignments for N sequences. However, in progressive methods, alignment errors made at each step are propagated to subsequent steps. Many progressive methods use a scoring function called sum-of-pairs, i.e. a sum of amino acid substitution scores for pairs of amino acids between two positions (Edgar and Batzoglou, 2006; Thompson et al., 1994). Such a scoring function yields reasonable alignment quality for closely related sequences (identity above 40%). However, alignment quality drops rapidly with decreasing sequence similarity (Thompson et al., 1999). Effective construction of multiple alignments with respect to accuracy and speed has been extensively researched in recent years. Refinement and consistency-based scoring are two major techniques to improve classical progressive methods. MUSCLE (Edgar, 2004) and MAFFT (Katoh et al., 2005) represent two recent methods that use extensive refinement to correct errors made in progressive steps. They both implement sum-of-pairs scores, which are easy to compute and offer the advantage of great speed. In T-COFFEE (Notredame et al., 2000), the scoring is derived by finding consistently aligned residue pairs in a library of pairwise alignments. Such consistency-based scoring functions can give better alignment quality than sum-of-pairs scores. Further improvement comes with a probabilistic treatment of consistency via pairwise hidden Markov models (HMMs), as first implemented in ProbCons (Do et al., 2005). MUMMALS (Pei and Grishin, 2006) builds on the success of probabilistic consistency by introducing HMMs with more states that capture local structural information. Consistency transformation requires operations on sequence triplets, and therefore is computationally intensive. By aligning similar sequences with general substitution matrices and aligning divergent sequence groups with profile-based consistency, PCMA (Pei et al., 2003) is able to achieve a balance between alignment accuracy and speed. Even with refinement and consistency-based scoring, current methods still have difficulty in obtaining high-quality alignments when sequence identity drops below 20%. As homologous proteins can have very low sequence similarity while maintaining similar structures and functions 802 ß The Author Published by Oxford University Press. All rights reserved. For Permissions, please journals.permissions@oxfordjournals.org
2 Accurate multiple sequence alignments of proteins (Murzin, 1998), aligning distantly related sequences is an important task. A recent trend in the multiple alignment field is to recruit various sources of sequence and structural information to improve alignment accuracy (Edgar and Batzoglou, 2006). Such sources include homologs detected in database searches (Katoh et al., 2005; Simossis and Heringa, 2005; Thompson et al., 2000), predicted secondary structure (Simossis and Heringa, 2005; Zhou and Zhou, 2005), and known 3D structures (O Sullivan et al., 2004). Since additional homologs improve the quality of sequence profiles, and structural features such as secondary structure are generally more conserved than sequences, their usage can lead to improved alignment quality. Here, we describe PROMALS, a multiple sequence alignment method that combines recent advances in computational approaches to tackle the difficult task of aligning divergent sequences. PROMALS improves probabilistic consistency-based scoring of profiles by utilizing predicted secondary structures and additional homologs found in database searches. To effectively combine these additional data, we developed and implemented a new hidden Markov model for profile-profile comparison, which scores both amino acid similarity and secondary structure similarity, and has local structure-dependent transition and emission probabilities. Like PCMA, PROMALS is made more computationally efficient by treating similar and divergent sequences with different alignment strategies. On several difficult data sets, we show that PROMALS gives the best alignment accuracy among leading methods such as SPEM, HHalign (Soding, 2005), MUMMALS, ProbCons and MAFFT. 2 METHODS 2.1 A hidden Markov model of profile profile alignment A classical pairwise HMM for aligning two sequences has three types of hidden states: a match state M emitting a residue pair, an X state emitting a residue in the first sequence and a Y state emitting a residue in the second sequence (Durbin et al., 1998). X and Y states correspond to insertions or deletions in the two sequences. Our hidden Markov model for aligning two alignments (having profile representations) has the same architecture as a pairwise sequence HMM. In our model, an M state emits a pair of positions instead of a pair of residues. For an X or Y state, a single position in the first alignment or in the second alignment is emitted, respectively. The emitted objects (observations) are amino acid frequency vectors and predicted secondary structure types. We adopt a representation of amino acid sequence profile similar to the ones in PSI-BLAST (Altschul et al., 1997) and COMPASS (Sadreyev and Grishin, 2003). Two profile components are estimated for a position in an alignment: (i) effective frequencies of amino acids, and (ii) target frequencies of amino acids. The effective frequencies serve as the emitted objects (observations) in a position for the hidden Markov model. They are estimated from the position-specific independent counts (PSIC) of amino acids (Pei and Grishin, 2001; Sunyaev et al., 1999), which is a sequence-weighting scheme that corrects for biased similarities between sequences. If an amino acid is not present in a position, it has an effective frequency of zero. The target frequencies serve as the hidden amino acid probabilistic generator for a position. The target frequencies are estimated from the effective frequencies, taking into account prior knowledge of amino acid substitution characteristics. The target frequency is a mixture (weighted average) between effective frequency and the pseudocount frequency (Altschul et al., 1997; Tatusov et al., 1994). Defined in this way, the target frequency of any amino acid, even if it is not present in a position, is always greater than zero. Details on derivation of the two profile components are in Supplementary Data. For an M state, the probability of emitting the observed amino acids for a position pair (i, j) is the product of two probabilities: (i) the probability of generating the effective frequencies of position i using the target frequencies of position j, and (ii) the probability of generating the effective frequencies of position j using the target frequencies of position i. For an X or Y state, the probability of emitting the observed amino acids in a position k is the probability of generating the effective frequencies of position k using the background amino acid frequencies in insertion regions. Besides amino acids, an M state also emits a pair of predicted secondary structures, and an X or Y state also emits a single predicted secondary structure. The emission probability in a hidden state ( M, X or Y ) is a weighted product of amino acid emission probability and secondary structure emission probability. The relative weights for the scoring terms of amino acids and predicted secondary structures have been optimized to increase the alignment accuracy of the training sequence pairs. Details on emission probability formulas, parameter estimation and the algorithm for aligning two profiles with optimal posterior probabilities of position matches are described in Supplementary Data. 2.2 PROMALS multiple sequence alignment procedure PROMALS (PROfile Multiple Alignment with predicted Local Structure) is a progressive method (Fig. 1). The alignment order is set by a tree built using a k-mer count method (Edgar, 2004). Like PCMA (Pei et al., 2003) and MUMMALS (Pei and Grishin, 2006), PROMALS has two alignment stages for easy and difficult alignments. In the first stage, highly similar sequences are progressively aligned in a fast way with a weighted sum-of-pairs measure of BLOSUM62 scores (Henikoff and Henikoff, 1992) (step 2 in Fig. 1). If two neighboring groups on the tree have an average sequence identity higher than a certain threshold (default: 60%), they are aligned in this fast way. The result of the first alignment stage is a set of sequences or pre-aligned groups that are relatively divergent from each other. In the second alignment stage, one representative sequence (the longest one) is selected from each pre-aligned group. For each representative, PSI-BLAST is used to search for homologs from sequence database UNIREF90 (Wu et al., 2006) with three iterations and an E-value cutoff of Hits with 520% identity to the query are removed and up to 300 hits are selected. The PSI-BLAST checkpoint file after three iterations is used to predict secondary structures by PSIPRED (Jones, 1999). For each pair of representatives, profiles are derived from the PSI-BLAST alignments and PSIPRED secondary structure prediction, and a matrix of posterior probabilities of matches between positions is obtained by forward and backward algorithms of the profile-profile HMM (see Supplementary Data for details). These matrices are used to calculate the probabilistic consistency scores as described in Do et al. (2005). The representatives are then aligned progressively according to the consistency-based scoring function, and the pre-aligned groups obtained in the first stage are merged to the multiple alignment of the representatives. Finally, gap placement is refined to make the gap patterns more realistic. For that, we define a core block as a set of consecutive positions with gap content less than 0.5 at each position. A highly gapped ( gappy ) region is defined as a set of consecutive positions with gap contents no less than 0.5 at each position. A gappy region is either bound by two adjacent core blocks, or is at the start or the end of the alignment. If there are l amino acid residues in a gappy segment, gap refinement introduces continuous gap characters in between the [l/2]th residue and the (l [l/2])th residue, with the exceptions for any gappy segment in N- or C-terminus, 803
3 J.Pei and N.V.Grishin 1. k-mer counting 2. Align similar sequences in a fast way 3. Select one sequence from each group N input sequences UPGMA tree N pre-aligned groups (N N) N representatives 4. Run PSI-BLAST and PSIPRED 7. Merge prealigned groups; refine gaps 6. Do progressive alignment based on consistency 5. Build profile-profile HMMs; consistency transformation Final alignment of N sequences Alignment of N representatives Probabilistic consistency objective function N profiles with predicted secondary structures Fig. 1. PROMALS multiple sequence alignment procedure. The gray arrows indicate the two most time-consuming steps: running PSI-BLAST and PSIPRED (step 4) and profile consistency transformation (step 5). where a single run of continuous gap characters is introduced at the sequence start or end. 2.3 Assessment of alignment methods The following methods were tested: SPEM (Zhou and Zhou, 2005), HHalign (Soding, 2005), MUMMALS (Pei and Grishin, 2006), ProbCons (version 1.10) (Do et al., 2005), MAFFT (version 5.667) (Katoh et al., 2005), MUSCLE (version 3.52) (Edgar, 2004) and ClustalW (version 1.83) (Thompson et al., 1994). For MAFFT, we report two alignment options ( -linsi and -ginsi ) that show the best results. HHalign is an enhanced version of HHsearch (Soding, 2005) that performs pairwise profile profile alignment with predicted secondary structures (J. Soding, personal communication). Several parameters (score shift, secondary structure weight, pseudocount weight) of HHalign were selected that gave optimal performance on SCOP domain pairs with identity 520%. For pairwise alignment tests, we used divergent SCOP superfamily domain pairs that were divided into three identity bins: below 10%, 10 15% and 15 20%. For multiple alignment tests, we added up to 24 homologs to each sequence in the testing cases of pairwise alignments. Details on construction of these testing data sets were given in our previous work (Pei and Grishin, 2006). Two large benchmark data sets compiled by other researchers were used as well. One is the SABmark database (version 1.65) (Van Walle et al., 2005), which contains two sets of multiple protein domains related at SCOP fold or superfamily level. The other is PREFAB database (version 4.0) (Edgar, 2004), which is based on structural alignments in FSSP database (Holm and Sander, 1998b) and homologous sequences from database searches. Reference-dependent alignment quality scores (Q-scores) were calculated using the built-in programs in SABmark and PREFAB packages. The Q-score is the number of correctly aligned residue pairs in the test alignment divided by the number of aligned residue pairs in the reference alignment. The value of the Q-score is between 0 and 1. Wilcoxon signed-ranks tests were performed to calculate the statistical significance of comparisons between alignment methods. In addition to Q-score, we applied reference-independent evaluation of alignment quality to SCOP domain pairs, as described in our previous work (Pei and Grishin, 2006). We calculated several scores reflecting structural similarity of two SCOP domains compared according to aligned residues in a test alignment: DALI Z-score (Holm and Sander, 1998a), GDT-TS score (Zemla et al., 1999), TM-score (Zhang and Skolnick, 2004), 3D-score (Rychlewski et al., 2003) and two LiveBench contact scores (Rychlewski et al., 2003). These scores were scaled by taking into account self-comparison scores, random scores and alignment coverage (scaled scores are no larger than 1 and usually above 0). We also calculated two referenceindependent sequence similarity scores: sequence identity and BLOSUM62 scores of aligned positions in a test alignment. These scores were also calculated for DaliLite (Holm and Sander, 1998a) structure-based alignments as a positive control. 3 RESULTS PROMALS is a progressive multiple alignment method based on probabilistic consistency of profile-profile comparison, with enhanced profile information from homologs detected by PSI-BLAST and secondary structures predicted by PSIPRED (Fig. 1). SPEM and HHalign are comparable methods as they also use these two sources of extra data. While PROMALS and SPEM can align two or more sequences, HHalign performs only pairwise alignments. The other tested methods (MUMMALS, ProbCons, MAFFT, MUSCLE and ClustalW) are stand-alone multiple sequence methods that do not resort to other data sources or programs. 3.1 Reference-dependent evaluation of methods Tests on weakly similar SCOP domain pairs We tested our profile-profile HMM on 1207 divergent SCOP domain pairs (Pei and Grishin, 2006) with 520% sequence identity (Table 1, first numbers in columns under SCOP ). The three methods that use extra data (PROMALS, SPEM and HHalign) produce substantially better results than stand-alone methods (MUMMALS, ProbCons, MAFFT, MUSCLE and ClustalW) that align a pair of sequences without using additional homologs or predicted secondary structures. For sequence 804
4 Accurate multiple sequence alignments of proteins Table 1. Reference-dependent evaluation of alignment methods Method SCOP a 0 10% (355) SCOP a 10 15% (432) SCOP a 15 20% (420) SABmark-twi (209) SABmark-sup (425) PREFAB c (1682) PROMALS 0.435/ / / SPEM 0.377/ / / HHalign b 0.406/ 0.567/ 0.730/ MUMMALS 0.151/ / / ProbCons 0.116/ / / MAFFT-linsi 0.116/ / / MAFFT-ginsi 0.116/ / / MUSCLE 0.139/ / / ClustalW 0.136/ / / Average Q-scores of three testing data sets of ASTRAL SCOP40 superfamily pairs, two SABmark data sets (twi twilight zone set, sup superfamily set) and the PREFAB 4.0 data set are shown. Q-score is the number of correctly aligned residue pairs in the test alignment divided by the total number of aligned residue pairs in the reference alignment. The number of alignments in each testing data set is shown in parentheses. Identity ranges are shown for the three SCOP data sets. The first three methods use extra data from PSI-BLAST and PSIPRED. The other five are stand-alone methods. The option of MUMMALS (modeling secondary structure and solvent accessibility) is set to produce the best results on these data sets. For each data set, PROMALS yields statistically higher accuracy (bold numbers) than any other method (P-value ) according to Wilcoxon signed rank test. a For tests on the SCOP data sets, there are two numbers in each cell separated by a slash. The first number is the average Q-score in pairwise alignment tests and the second number is the average Q-score in multiple alignment tests. b HHalign only performs pairwise profile profile alignments and does not construct multiple sequence alignments. Thus the values for SCOP multiple alignment tests and SABmark tests are not available. c For PREFAB 4.0 data set, the scores of PROMALS, HHalign and SPEM are based on pairwise profile profile alignments, while the scores for other methods are based on multiple alignments. pairs with identity below 10%, the average Q-score of PROMALS (0.431) is almost three times higher than that of MUMMALS (0.156). For alignments with identity ranges 10 15% and 15 20%, PROMALS also gives substantial accuracy increases over MUMMALS of and 0.176, respectively. PROMALS shows about 3 4% accuracy increases over SPEM and HHalign, suggesting that our profile-profile HMM utilizes homologs and predicted secondary structures in a better way. We also tested the methods (except HHalign, which is a pairwise alignment program) on data sets of multiple sequences constructed by adding up to 48 homologs to each SCOP domain pair (Table 1, second numbers in columns under SCOP ). With multiple sequences, PROMALS and SPEM both show slight improvement (1 2% for PROMALS and 2 3% for SPEM) over their pairwise profile profile alignments. PROMALS outperforms SPEM by 2% on multiple sequences. With added homologs, stand-alone methods all yield better accuracies than pairwise sequence alignments, among which MUMMALS is the best method. PROMALS outperforms MUMMALS by 0.13, 0.1, and 0.05 for data sets with identities 510%, 10 15% and 15 20%, respectively Tests on SABmark database SABmark database (version 1.65) has two multiple alignment benchmark sets. The twilight zone set contains 209 tests of SCOP (version 1.65) fold-level domains with very low similarity, and the superfamily set contains 425 tests of SCOP superfamily-level domains with low to intermediate similarity. PROMALS achieves the best results among all methods for both sets. Its accuracy is 6% and 4% higher than SPEM on twilight zone set and superfamily set, respectively. For the most difficult twilight zone set, PROMALS doubles the accuracy of the best stand-alone method (MUMMALS). Nevertheless, only 40% residues were correctly aligned on average by PROMALS for the twilight zone set, suggesting that homology modeling of extremely divergent domains remains a difficult problem with regard to alignment quality Tests on and PREFAB database PREFAB 4.0 database consists of 1682 alignments averaging 45.2 sequences per alignment. Each alignment consists of two sequences with known structures and their homologs found by PSI-BLAST database searches. The reference structural alignment in each test is based on the consensus of FSSP (Holm and Sander, 1998b) and CE (Shindyalov and Bourne, 1998) alignments. We have used the performances of pairwise profile profile alignments of PROMALS and SPEM as an indicator of their multiple alignment performances. The three methods that use additional data (PROMALS, SPEM and HHalign) give similar results, each with an average Q-score above Their accuracies are higher than those on the two SCOP data sets with identity515% and the two SABmark sets, suggesting that PREFAB 4.0 is an easier testing data set. PROMALS, SPEM and HHalign are more accurate than MUMMALS by 4 6%. PROMALS is statistically more accurate (P-value ) than SPEM and HHalign despite small differences in their average Q-scores. Results on PREFAB 4.0 confirm that alignment quality differences between methods become smaller on easier tests. 3.2 Reference-independent evaluation of methods On our data sets of 1207 SCOP domain pairs with identity below 20%, we evaluated alignment quality using referenceindependent scores that reflect the similarity between two structures compared according to aligned residue pairs in the test alignment (Pei and Grishin, 2006). These structural 805
5 J.Pei and N.V.Grishin Table 2. Reference-independent evaluation on 1207 representative SCOP40 domain pairs with identity 520% Method Structural similarity Sequence similarity DALI Z-score GDT-TS TM-score 3D-score LBcona LBconb Identity BLOSUM62 PROMALS a a a a a a SPEM HHalign MUMMALS ProbCons MAFFT-linsi MAFFT-ginsi MUSCLE ClustalW DaliLite b b The first three methods use extra data given by PSI-BLAST and PSIPRED. The last method (DaliLite) produces alignments based on comparison of known 3D structures. The other five are stand-alone methods. All sequence-based methods except HHalign construct multiple sequence alignments for target domain pairs with up to 48 homologs. HHalign constructs pairwise profile profile alignments. Scores are calculated for pairwise alignments of target domain pairs extracted from multiple sequence alignments. a PROMAL yields statistically higher structure-similarity scores (in bold) than other sequence alignment methods (P-value ) according to Wilcoxon signed rank test. b DaliLite structure-based sequence alignments have the lowest average sequence similarity scores (in bold). similarity scores are DALI Z-score, TM-score, GDT-TS score, 3D-score, and two LiveBench contact scores (Table 2). Consistent with reference-dependent evaluation, PROMALS produces significantly higher average structural similarity scores than other methods. Used as a positive control, structural alignment method DaliLite yields higher structural similarity scores than any sequence-based alignment method (Table 2). Interestingly, DaliLite alignments have the lowest reference-independent sequence similarity scores (sequence identity and BLOSUM62 scores). PROMALS also shows lower sequence similarity scores than several other sequencebased methods. These observations suggest that for distantly related sequences (sequence identity520%), sequence similarity scores, such as identity or BLOSUM62, may not correlate with alignment quality measured by 3D structural comparison, and maximization of these scores may not improve structural models based on sequence alignments. 3.3 Pairwise comparisons of alignment methods To gain further understanding of the differences between alignment methods, we compared their performance on individual domain pairs from the SCOP sets (identity 520%). Table 3 shows the number of pairs, for which one method performs better than another method by a relatively large margin of 0.1 or more (measured by scaled TM-score or Q-score, both scores are between 0 and 1). Although PROMALS clearly leads by a large margin, it does not offer the best alignment in each and every case. For example, PROMALS gives a TM-score increase of 0.1 or more over SPEM on 197 alignments, while producing significantly inferior alignments for 109 pairs. Even stand-alone methods (MUMMALS, ProbCons, MAFFT, MUSCLE and ClustalW) outperform PROMALS by a TM-score of 0.1 or more on a small number of pairs (5%, i.e out of 1207 alignments). These comparisons suggest that alignments constructed by different methods can vary much for divergent sequences, and a method with an overall inferior performance is capable of generating better alignments in some cases. Careful inspection of alignments produced by several programs could help improve alignment quality for divergent sequences. 4 DISCUSSION Judging by its performance, PROMALS is a definite advance compared to our previous alignment programs MUMMALS (Pei and Grishin, 2006). MUMMALS derives probabilistic consistency from pairwise HMMs with built-in local structural information (secondary structure and/or solvent accessibility), and shows slight but significant improvement (a few percent) over other stand-alone methods such as ProbCons (Do et al., 2005) and MAFFT (Katoh et al., 2005). However, since no additional homologs are used, the local structure prediction implicitly performed by MUMMALS is of low accuracy compared to advanced methods such as PSIPRED (Jones, 1999). In contrast, PROMALS incorporates database searches and more accurate secondary structure prediction, and derives probabilistic consistency from profile profile HMMs. Moreover, the HMM in PROMALS has a two-track structure (Karchin et al., 2003) that treats both amino acids and predicted secondary structures as emitted objects, while MUMMALS HMMs only emit amino acids. Owing to additional data sources and the advanced profile profile HMM, PROMALS shows significant improvement over MUMMALS and other stand-alone methods, especially for highly divergent sequences. The HMM in PROMALS adopts a numerical representation of sequence profile (see Supplementary Data for details) that successfully works in other profile-sequence or profile profile 806
6 Accurate multiple sequence alignments of proteins Table 3. Pairwise comparisons among alignment methods on 1207 SCOP domain pairs with identity 520% PROMALS SPEM HHalign MUMMALS ProbCons MAFFT-linsi MAFFT-ginsi MUSCLE ClustalW PROMALS 109/196 76/179 67/340 44/458 67/398 61/374 60/464 49/650 SPEM 199/81 140/ /281 71/389 98/324 99/326 82/400 43/574 HHalign 265/84 196/121 77/254 49/368 73/288 78/301 66/393 53/571 MUMMALS 685/ / /333 38/ /138 82/128 82/227 59/431 ProbCons 726/ / / /62 172/80 162/76 162/ /336 MAFFT-linsi 718/ / / / /188 85/98 93/196 60/387 MAFFT-ginsi 714/ / / / / /132 90/185 67/395 MUSCLE 783/ / / /83 302/ / /106 75/288 ClustalW 858/ / / /55 559/ /70 600/76 449/100 Each off-diagonal cell has two numbers separated by a slash. The first number is the number of pairs where the alignment score of the method listed to the left is inferior to that of the method listed above (in a column) by 0.1 or more. The second number is the number of pairs where the score of the method listed to the left is better than that of the method listed above by 0.1 or more. The alignment quality scores used for comparison in the lower triangle and the upper triangle are Q-scores and weighted and scaled TM-scores, respectively. These scores are calculated based on results of multiple sequence alignments (target domain pairs plus up to 48 added homologs), with the exception of HHalign alignments, which are pairwise profile profile alignments. Comparisons of PROMALS with other methods are highlighted in bold. alignment methods such as PSI-BLAST (Altschul et al., 1997) and COMPASS (Sadreyev and Grishin, 2003). A recent comprehensive study also supported the effectiveness of this profile profile scoring scheme (Wang and Dunbrack, 2004). To adequately use predicted secondary structures, we not only score them as emitted objects, but also use transition and emission probabilities that are dependent on predicted secondary structure types (Supplementary Data). Unlike HHalign, which treats each alignment as a classical profile HMM (Eddy, 1998), our HMM has a simpler structure similar to the classical 3-state pairwise HMM (Durbin et al., 1998). SPEM (Zhou and Zhou, 2005) does not use HMMs, but applies an empirical profile profile alignment method (SP 2 ) that identifies the optimal alignment path. In contrast, the HMM in PROMALS allows estimation of posterior probabilities of matches between positions. As a result, PROMALS has a probabilistic treatment of consistency similar to the one in ProbCons and MUMMALS, while simple consistency measures are used in SPEM, T-COFFEE (Notredame et al., 2000) and PCMA (Pei et al., 2003). PROMALS performs significantly better than SPEM and HHalign on difficult tests, suggesting the advantages of our profile profile comparison scheme. Since PROMALS relies on PSI-BLAST and PSIPRED to collect additional homologs and predicted secondary structures, the speed of PROMALS is considerably slower than that of stand-alone progressive methods. Our strategy for improving speed is to use different algorithms for easy and difficult alignments (Pei et al., 2003). By aligning highly similar sequences in a fast way, the number of sequences subject to the time-consuming steps (running PSI-BLAST, PSIPRED and consistency transformation) could be substantially reduced. For example, for 1207 SCOP domain pairs with up to 48 added homologs, the average number of sequences in an alignment is After PROMALS aligns similar sequences with identity above 60% in the first stage, only 24 sequences on average require database searches, secondary structure prediction, and consistency transformation. For these tests, the median CPU time of PROMALS is 30 min per alignment, as compared to 67 min for SPEM (on Redhat Enterprise Linux 3, AMD Opteron 2.0 GHz). The stand-alone methods (MUMMALS, PROBCONS, MAFFT, MUSCLE and ClustalW) are much faster, all with a median CPU time51 min. As in our previous work (Pei and Grishin, 2006), we demonstrated the effectiveness of reference-independent evaluation of alignment quality in this study. First, we observed a good correlation between reference-dependent and referenceindependent evaluations, suggesting that it may not be necessary to spend significant efforts on development of reference alignment databases. Second, reference-independent techniques solve the problem of reference alignment ambiguity, which becomes significant when similarity is low. Third, reference-independent evaluation helps answer general questions such as whether alignments can be further improved for sequences with low similarity, and whether such improvements will help structure modeling. For several structural similarity measures (GDT-TS, 3Dscore, TM-score, LB contact scores), the ratio between the average score of PROMALS sequencebased alignment and the average score of DaliLite structurebased alignment is 0.6 on domain pairs with 520% sequence identity (Table 2), suggesting that we are still 40% below what can be achieved with structures in hand. Notably, for these divergent sequences, DaliLite structural alignments have lower sequence similarity scores (identity and BLOSUM62 scores) than alignments produced by any sequence method, suggesting that scoring functions based only on amino acid sequence similarity may not be suitable for aligning divergent sequences for the purpose of homology modeling. This observation further justifies the use of alternative scoring schemes, such as the ones that recruit structural information. ACKNOWLEDGEMENTS We would like to thank Bong-Hyun Kim for the referenceindependent evaluation routine, and Johannes Soding for providing the HHalign program. We would like to thank Lisa Kinch, Ruslan Sadreyev and James Wrabl for critical reading 807
7 J.Pei and N.V.Grishin of the manuscript and helpful comments. This work was supported in part by NIH grant GM67165 to NVG. Conflict of Interest: none declared. REFERENCES Altschul,S.F. et al. (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res., 25, Do,C.B. et al. (2005) ProbCons: probabilistic consistency-based multiple sequence alignment. Genome Res., 15, Durbin,R. et al. (1998) Biological Sequence Analysis: Probabilistic Models of Proteins and Nucleic Acids. Cambridge University Press. Eddy,S.R. (1998) Profile hidden Markov models. Bioinformatics, 14, Edgar,R.C. (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res., 32, Edgar,R.C. and Batzoglou,S. (2006) Multiple sequence alignment. Curr. Opin. Struct. Biol, 16, Ginalski,K. and Rychlewski,L. (2003) Detection of reliable and unexpected protein fold predictions using 3D-Jury. Nucleic Acids Res., 31, Henikoff,S. and Henikoff,J.G. (1992) Amino acid substitution matrices from protein blocks. Proc. Natl. Acad. Sci. USA, 89, Holm,L. and Sander,C. (1998a) Dictionary of recurrent domains in protein structures. Proteins, 33, Holm,L. and Sander,C. (1998b) Touring protein fold space with Dali/FSSP. Nucleic Acids Res., 26, Jones,D.T. (1999) Protein secondary structure prediction based on positionspecific scoring matrices. J. Mol. Biol., 292, Karchin,R. et al. (2003) Hidden Markov models that use predicted local structure for fold recognition: alphabets of backbone geometry. Proteins, 51, Katoh,K. et al. (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res., 33, Lipman,D.J. et al. (1989) A tool for multiple sequence alignment. Proc. Natl. Acad. Sci. USA, 86, Murzin,A.G. (1998) How far divergent evolution goes in proteins. Curr. Opin. Struct. Biol., 8, Notredame,C. et al. (2000) T-Coffee: a novel method for fast and accurate multiple sequence alignment. J. Mol. Biol., 302, O Sullivan,O. et al. (2004) 3DCoffee: combining protein sequences and structures within multiple sequence alignments. J. Mol. Biol., 340, Pei,J. and Grishin,N.V. (2001) AL2CO: calculation of positional conservation in a protein sequence alignment. Bioinformatics, 17, Pei,J. and Grishin,N.V. (2006) MUMMALS: multiple sequence alignment improved by using hidden Markov models with local structural information. Nucleic Acids Res, 34, Pei,J. et al. (2003) PCMA: fast and accurate multiple sequence alignment based on profile consistency. Bioinformatics, 19, Phillips,A. et al. (2000) Multiple sequence alignment in phylogenetic analysis. Mol. Phylogenet. Evol., 16, Rychlewski,L. et al. (2003) LiveBench-6: large-scale automated evaluation of protein structure prediction servers. Proteins, 53 (Suppl. 6), Sadreyev,R. and Grishin,N. (2003) COMPASS: a tool for comparison of multiple protein alignments with assessment of statistical significance. J. Mol. Biol., 326, Shindyalov,I.N. and Bourne,P.E. (1998) Protein structure alignment by incremental combinatorial extension (CE) of the optimal path. Protein Eng., 11, Simossis,V.A. and Heringa,J. (2005) PRALINE: a multiple sequence alignment toolbox that integrates homology-extended and secondary structure information. Nucleic Acids Res., 33, W Soding,J. (2005) Protein homology detection by HMM-HMM comparison. Bioinformatics, 21, Sunyaev,S.R. et al. (1999) PSIC: profile extraction from sequence alignments with position-specific counts of independent observations. Protein Eng., 12, Tatusov,R.L. et al. (1994) Detection of conserved segments in proteins: iterative scanning of sequence databases with alignment blocks. Proc. Natl. Acad. Sci. USA, 91, Thompson,J.D. et al. (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res., 22, Thompson,J.D. et al. (1999) A comprehensive comparison of multiple sequence alignment programs. Nucleic Acids Res., 27, Thompson,J.D. et al. (2000) DbClustal: rapid and reliable global multiple alignments of protein sequences detected by database searches. Nucleic Acids Res., 28, Van Walle,I. et al. (2005) SABmark a benchmark for sequence alignment that covers the entire known fold space. Bioinformatics, 21, Wang,G. and Dunbrack,R.L., Jr. (2004) Scoring profile-to-profile sequence alignments. Protein Sci., 13, Wu,C.H. et al. (2006) The Universal Protein Resource (UniProt): an expanding universe of protein information. Nucleic Acids Res., 34, D Zemla,A. et al. (1999) Processing and analysis of CASP3 protein structure predictions. Proteins, (Suppl. 3), Zhang,Y. and Skolnick,J. (2004) Scoring function for automated assessment of protein structure template quality. Proteins, 57, Zhou,H. and Zhou,Y. (2005) SPEM: improving multiple sequence alignment with sequence profiles and predicted secondary structures. Bioinformatics, 21,
PROMALS3D: a tool for multiple protein sequence and structure alignments
 Nucleic Acids Research Advance Access published February 20, 2008 Nucleic Acids Research, 2008, 1 6 doi:10.1093/nar/gkn072 PROMALS3D: a tool for multiple protein sequence and structure alignments Jimin
Nucleic Acids Research Advance Access published February 20, 2008 Nucleic Acids Research, 2008, 1 6 doi:10.1093/nar/gkn072 PROMALS3D: a tool for multiple protein sequence and structure alignments Jimin
PROMALS3D web server for accurate multiple protein sequence and structure alignments
 W30 W34 Nucleic Acids Research, 2008, Vol. 36, Web Server issue Published online 24 May 2008 doi:10.1093/nar/gkn322 PROMALS3D web server for accurate multiple protein sequence and structure alignments
W30 W34 Nucleic Acids Research, 2008, Vol. 36, Web Server issue Published online 24 May 2008 doi:10.1093/nar/gkn322 PROMALS3D web server for accurate multiple protein sequence and structure alignments
proteins Refinement by shifting secondary structure elements improves sequence alignments
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Refinement by shifting secondary structure elements improves sequence alignments Jing Tong, 1,2 Jimin Pei, 3 Zbyszek Otwinowski, 1,2 and Nick V. Grishin 1,2,3
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Refinement by shifting secondary structure elements improves sequence alignments Jing Tong, 1,2 Jimin Pei, 3 Zbyszek Otwinowski, 1,2 and Nick V. Grishin 1,2,3
Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences
 Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences Yue Lu 1 and Sing-Hoi Sze 1,2 1 Department of Biochemistry & Biophysics 2 Department of Computer Science, Texas A&M University,
Multiple Sequence Alignment Based on Profile Alignment of Intermediate Sequences Yue Lu 1 and Sing-Hoi Sze 1,2 1 Department of Biochemistry & Biophysics 2 Department of Computer Science, Texas A&M University,
A profile-based protein sequence alignment algorithm for a domain clustering database
 A profile-based protein sequence alignment algorithm for a domain clustering database Lin Xu,2 Fa Zhang and Zhiyong Liu 3, Key Laboratory of Computer System and architecture, the Institute of Computing
A profile-based protein sequence alignment algorithm for a domain clustering database Lin Xu,2 Fa Zhang and Zhiyong Liu 3, Key Laboratory of Computer System and architecture, the Institute of Computing
proteins SHORT COMMUNICATION MALIDUP: A database of manually constructed structure alignments for duplicated domain pairs
 J_ID: Z7E Customer A_ID: 21783 Cadmus Art: PROT21783 Date: 25-SEPTEMBER-07 Stage: I Page: 1 proteins STRUCTURE O FUNCTION O BIOINFORMATICS SHORT COMMUNICATION MALIDUP: A database of manually constructed
J_ID: Z7E Customer A_ID: 21783 Cadmus Art: PROT21783 Date: 25-SEPTEMBER-07 Stage: I Page: 1 proteins STRUCTURE O FUNCTION O BIOINFORMATICS SHORT COMMUNICATION MALIDUP: A database of manually constructed
Multiple Sequence Alignment: A Critical Comparison of Four Popular Programs
 Multiple Sequence Alignment: A Critical Comparison of Four Popular Programs Shirley Sutton, Biochemistry 218 Final Project, March 14, 2008 Introduction For both the computational biologist and the research
Multiple Sequence Alignment: A Critical Comparison of Four Popular Programs Shirley Sutton, Biochemistry 218 Final Project, March 14, 2008 Introduction For both the computational biologist and the research
Week 10: Homology Modelling (II) - HHpred
 Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
Week 10: Homology Modelling (II) - HHpred Course: Tools for Structural Biology Fabian Glaser BKU - Technion 1 2 Identify and align related structures by sequence methods is not an easy task All comparative
frmsdalign: Protein Sequence Alignment Using Predicted Local Structure Information for Pairs with Low Sequence Identity
 1 frmsdalign: Protein Sequence Alignment Using Predicted Local Structure Information for Pairs with Low Sequence Identity HUZEFA RANGWALA and GEORGE KARYPIS Department of Computer Science and Engineering
1 frmsdalign: Protein Sequence Alignment Using Predicted Local Structure Information for Pairs with Low Sequence Identity HUZEFA RANGWALA and GEORGE KARYPIS Department of Computer Science and Engineering
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Statistical Machine Learning Methods for Bioinformatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD Department of Computer Science University of Missouri 2008 Free for Academic
Grouping of amino acids and recognition of protein structurally conserved regions by reduced alphabets of amino acids
 Science in China Series C: Life Sciences 2007 Science in China Press Springer-Verlag Grouping of amino acids and recognition of protein structurally conserved regions by reduced alphabets of amino acids
Science in China Series C: Life Sciences 2007 Science in China Press Springer-Verlag Grouping of amino acids and recognition of protein structurally conserved regions by reduced alphabets of amino acids
An Introduction to Sequence Similarity ( Homology ) Searching
 An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
An Introduction to Sequence Similarity ( Homology ) Searching Gary D. Stormo 1 UNIT 3.1 1 Washington University, School of Medicine, St. Louis, Missouri ABSTRACT Homologous sequences usually have the same,
Multiple sequence alignment
 Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Multiple sequence alignment Multiple sequence alignment: today s goals to define what a multiple sequence alignment is and how it is generated; to describe profile HMMs to introduce databases of multiple
Multiple Mapping Method: A Novel Approach to the Sequence-to-Structure Alignment Problem in Comparative Protein Structure Modeling
 63:644 661 (2006) Multiple Mapping Method: A Novel Approach to the Sequence-to-Structure Alignment Problem in Comparative Protein Structure Modeling Brajesh K. Rai and András Fiser* Department of Biochemistry
63:644 661 (2006) Multiple Mapping Method: A Novel Approach to the Sequence-to-Structure Alignment Problem in Comparative Protein Structure Modeling Brajesh K. Rai and András Fiser* Department of Biochemistry
Upcoming challenges for multiple sequence alignment methods in the high-throughput era
 BIOINFORMATICS REVIEW Vol. 25 no. 19 2009, pages 2455 2465 doi:10.1093/bioinformatics/btp452 Sequence analysis Upcoming challenges for multiple sequence alignment methods in the high-throughput era Carsten
BIOINFORMATICS REVIEW Vol. 25 no. 19 2009, pages 2455 2465 doi:10.1093/bioinformatics/btp452 Sequence analysis Upcoming challenges for multiple sequence alignment methods in the high-throughput era Carsten
Protein sequence alignment with family-specific amino acid similarity matrices
 TECHNICAL NOTE Open Access Protein sequence alignment with family-specific amino acid similarity matrices Igor B Kuznetsov Abstract Background: Alignment of amino acid sequences by means of dynamic programming
TECHNICAL NOTE Open Access Protein sequence alignment with family-specific amino acid similarity matrices Igor B Kuznetsov Abstract Background: Alignment of amino acid sequences by means of dynamic programming
HMM applications. Applications of HMMs. Gene finding with HMMs. Using the gene finder
 HMM applications Applications of HMMs Gene finding Pairwise alignment (pair HMMs) Characterizing protein families (profile HMMs) Predicting membrane proteins, and membrane protein topology Gene finding
HMM applications Applications of HMMs Gene finding Pairwise alignment (pair HMMs) Characterizing protein families (profile HMMs) Predicting membrane proteins, and membrane protein topology Gene finding
SUPPLEMENTARY INFORMATION
 Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
Supplementary information S1 (box). Supplementary Methods description. Prokaryotic Genome Database Archaeal and bacterial genome sequences were downloaded from the NCBI FTP site (ftp://ftp.ncbi.nlm.nih.gov/genomes/all/)
BIOINFORMATICS. RASCAL: rapid scanning and correction of multiple sequence alignments. J. D. Thompson, J. C. Thierry and O.
 BIOINFORMATICS Vol. 19 no. 9 2003, pages 1155 1161 DOI: 10.1093/bioinformatics/btg133 RASCAL: rapid scanning and correction of multiple sequence alignments J. D. Thompson, J. C. Thierry and O. Poch Laboratoire
BIOINFORMATICS Vol. 19 no. 9 2003, pages 1155 1161 DOI: 10.1093/bioinformatics/btg133 RASCAL: rapid scanning and correction of multiple sequence alignments J. D. Thompson, J. C. Thierry and O. Poch Laboratoire
Protein homology detection by HMM HMM comparison
 BIOINFORMATICS ORIGINAL PAPER Vol. 21 no. 7 2005, pages 951 960 doi:10.1093/bioinformatics/bti125 Sequence analysis Protein homology detection by HMM HMM comparison Johannes Söding Department of Protein
BIOINFORMATICS ORIGINAL PAPER Vol. 21 no. 7 2005, pages 951 960 doi:10.1093/bioinformatics/bti125 Sequence analysis Protein homology detection by HMM HMM comparison Johannes Söding Department of Protein
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences
 Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Statistical Machine Learning Methods for Biomedical Informatics II. Hidden Markov Model for Biological Sequences Jianlin Cheng, PhD William and Nancy Thompson Missouri Distinguished Professor Department
Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT
 22 Genome Informatics 16(1): 22-33 (2005) Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT Kazutaka Katohl kkatoh@kuicr.kyoto-u.ac.jp Takashi Miyata2" miyata@brh.co.jp Kei-ichi
22 Genome Informatics 16(1): 22-33 (2005) Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT Kazutaka Katohl kkatoh@kuicr.kyoto-u.ac.jp Takashi Miyata2" miyata@brh.co.jp Kei-ichi
BIOINFORMATICS ORIGINAL PAPER
 BIOINFORMATICS ORIGINAL PAPER Vol. 24 no. 19 2008, pages 2165 2171 doi:10.1093/bioinformatics/btn414 Sequence analysis Model-based prediction of sequence alignment quality Virpi Ahola 1,2,, Tero Aittokallio
BIOINFORMATICS ORIGINAL PAPER Vol. 24 no. 19 2008, pages 2165 2171 doi:10.1093/bioinformatics/btn414 Sequence analysis Model-based prediction of sequence alignment quality Virpi Ahola 1,2,, Tero Aittokallio
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
In-Depth Assessment of Local Sequence Alignment
 2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
2012 International Conference on Environment Science and Engieering IPCBEE vol.3 2(2012) (2012)IACSIT Press, Singapoore In-Depth Assessment of Local Sequence Alignment Atoosa Ghahremani and Mahmood A.
2 Dean C. Adams and Gavin J. P. Naylor the best three-dimensional ordination of the structure space is found through an eigen-decomposition (correspon
 A Comparison of Methods for Assessing the Structural Similarity of Proteins Dean C. Adams and Gavin J. P. Naylor? Dept. Zoology and Genetics, Iowa State University, Ames, IA 50011, U.S.A. 1 Introduction
A Comparison of Methods for Assessing the Structural Similarity of Proteins Dean C. Adams and Gavin J. P. Naylor? Dept. Zoology and Genetics, Iowa State University, Ames, IA 50011, U.S.A. 1 Introduction
The PRALINE online server: optimising progressive multiple alignment on the web
 Computational Biology and Chemistry 27 (2003) 511 519 Software Note The PRALINE online server: optimising progressive multiple alignment on the web V.A. Simossis a,b, J. Heringa a, a Bioinformatics Unit,
Computational Biology and Chemistry 27 (2003) 511 519 Software Note The PRALINE online server: optimising progressive multiple alignment on the web V.A. Simossis a,b, J. Heringa a, a Bioinformatics Unit,
THEORY. Based on sequence Length According to the length of sequence being compared it is of following two types
 Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Exp 11- THEORY Sequence Alignment is a process of aligning two sequences to achieve maximum levels of identity between them. This help to derive functional, structural and evolutionary relationships between
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
www.sciencemag.org/cgi/content/full/309/5742/1868/dc1 Supporting Online Material for Toward High-Resolution de Novo Structure Prediction for Small Proteins Philip Bradley, Kira M. S. Misura, David Baker*
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche
 Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Protein Structure Prediction II Lecturer: Serafim Batzoglou Scribe: Samy Hamdouche The molecular structure of a protein can be broken down hierarchically. The primary structure of a protein is simply its
Basic Local Alignment Search Tool
 Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
Basic Local Alignment Search Tool Alignments used to uncover homologies between sequences combined with phylogenetic studies o can determine orthologous and paralogous relationships Local Alignment uses
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT
 5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
5. MULTIPLE SEQUENCE ALIGNMENT BIOINFORMATICS COURSE MTAT.03.239 03.10.2012 ALIGNMENT Alignment is the task of locating equivalent regions of two or more sequences to maximize their similarity. Homology:
InDel 3-5. InDel 8-9. InDel 3-5. InDel 8-9. InDel InDel 8-9
 Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Lecture 5 Alignment I. Introduction. For sequence data, the process of generating an alignment establishes positional homologies; that is, alignment provides the identification of homologous phylogenetic
Homology Modeling. Roberto Lins EPFL - summer semester 2005
 Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Homology Modeling Roberto Lins EPFL - summer semester 2005 Disclaimer: course material is mainly taken from: P.E. Bourne & H Weissig, Structural Bioinformatics; C.A. Orengo, D.T. Jones & J.M. Thornton,
Neural Networks for Protein Structure Prediction Brown, JMB CS 466 Saurabh Sinha
 Neural Networks for Protein Structure Prediction Brown, JMB 1999 CS 466 Saurabh Sinha Outline Goal is to predict secondary structure of a protein from its sequence Artificial Neural Network used for this
Neural Networks for Protein Structure Prediction Brown, JMB 1999 CS 466 Saurabh Sinha Outline Goal is to predict secondary structure of a protein from its sequence Artificial Neural Network used for this
Sequence Database Search Techniques I: Blast and PatternHunter tools
 Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
Sequence Database Search Techniques I: Blast and PatternHunter tools Zhang Louxin National University of Singapore Outline. Database search 2. BLAST (and filtration technique) 3. PatternHunter (empowered
EECS730: Introduction to Bioinformatics
 EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
EECS730: Introduction to Bioinformatics Lecture 07: profile Hidden Markov Model http://bibiserv.techfak.uni-bielefeld.de/sadr2/databasesearch/hmmer/profilehmm.gif Slides adapted from Dr. Shaojie Zhang
CONCEPT OF SEQUENCE COMPARISON. Natapol Pornputtapong 18 January 2018
 CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
CONCEPT OF SEQUENCE COMPARISON Natapol Pornputtapong 18 January 2018 SEQUENCE ANALYSIS - A ROSETTA STONE OF LIFE Sequence analysis is the process of subjecting a DNA, RNA or peptide sequence to any of
BIOINFORMATICS. Consensus alignment for reliable framework prediction in homology modeling
 BIOINFORMATICS Vol. 19 no. 13 23, pages 1682 1691 DOI: 1.193/bioinformatics/btg211 Consensus alignment for reliable framework prediction in homology modeling J.C. Prasad 1, S.R. Comeau 1, S. Vajda 2 and
BIOINFORMATICS Vol. 19 no. 13 23, pages 1682 1691 DOI: 1.193/bioinformatics/btg211 Consensus alignment for reliable framework prediction in homology modeling J.C. Prasad 1, S.R. Comeau 1, S. Vajda 2 and
Supporting Text 1. Comparison of GRoSS sequence alignment to HMM-HMM and GPCRDB
 Structure-Based Sequence Alignment of the Transmembrane Domains of All Human GPCRs: Phylogenetic, Structural and Functional Implications, Cvicek et al. Supporting Text 1 Here we compare the GRoSS alignment
Structure-Based Sequence Alignment of the Transmembrane Domains of All Human GPCRs: Phylogenetic, Structural and Functional Implications, Cvicek et al. Supporting Text 1 Here we compare the GRoSS alignment
Sequence Alignment Techniques and Their Uses
 Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Sequence Alignment Techniques and Their Uses Sarah Fiorentino Since rapid sequencing technology and whole genomes sequencing, the amount of sequence information has grown exponentially. With all of this
Single alignment: Substitution Matrix. 16 march 2017
 Single alignment: Substitution Matrix 16 march 2017 BLOSUM Matrix BLOSUM Matrix [2] (Blocks Amino Acid Substitution Matrices ) It is based on the amino acids substitutions observed in ~2000 conserved block
Single alignment: Substitution Matrix 16 march 2017 BLOSUM Matrix BLOSUM Matrix [2] (Blocks Amino Acid Substitution Matrices ) It is based on the amino acids substitutions observed in ~2000 conserved block
Probalign: Multiple sequence alignment using partition function posterior probabilities
 Sequence Analysis Probalign: Multiple sequence alignment using partition function posterior probabilities Usman Roshan 1* and Dennis R. Livesay 2 1 Department of Computer Science, New Jersey Institute
Sequence Analysis Probalign: Multiple sequence alignment using partition function posterior probabilities Usman Roshan 1* and Dennis R. Livesay 2 1 Department of Computer Science, New Jersey Institute
Syllabus of BIOINF 528 (2017 Fall, Bioinformatics Program)
 Syllabus of BIOINF 528 (2017 Fall, Bioinformatics Program) Course Name: Structural Bioinformatics Course Description: Instructor: This course introduces fundamental concepts and methods for structural
Syllabus of BIOINF 528 (2017 Fall, Bioinformatics Program) Course Name: Structural Bioinformatics Course Description: Instructor: This course introduces fundamental concepts and methods for structural
Chapter 11 Multiple sequence alignment
 Chapter 11 Multiple sequence alignment Burkhard Morgenstern 1. INTRODUCTION Sequence alignment is of crucial importance for all aspects of biological sequence analysis. Virtually all methods of nucleic
Chapter 11 Multiple sequence alignment Burkhard Morgenstern 1. INTRODUCTION Sequence alignment is of crucial importance for all aspects of biological sequence analysis. Virtually all methods of nucleic
K-means-based Feature Learning for Protein Sequence Classification
 K-means-based Feature Learning for Protein Sequence Classification Paul Melman and Usman W. Roshan Department of Computer Science, NJIT Newark, NJ, 07102, USA pm462@njit.edu, usman.w.roshan@njit.edu Abstract
K-means-based Feature Learning for Protein Sequence Classification Paul Melman and Usman W. Roshan Department of Computer Science, NJIT Newark, NJ, 07102, USA pm462@njit.edu, usman.w.roshan@njit.edu Abstract
3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT
 3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT.03.239 25.09.2012 SEQUENCE ANALYSIS IS IMPORTANT FOR... Prediction of function Gene finding the process of identifying the regions of genomic DNA that encode
3. SEQUENCE ANALYSIS BIOINFORMATICS COURSE MTAT.03.239 25.09.2012 SEQUENCE ANALYSIS IS IMPORTANT FOR... Prediction of function Gene finding the process of identifying the regions of genomic DNA that encode
Tiffany Samaroo MB&B 452a December 8, Take Home Final. Topic 1
 Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Tiffany Samaroo MB&B 452a December 8, 2003 Take Home Final Topic 1 Prior to 1970, protein and DNA sequence alignment was limited to visual comparison. This was a very tedious process; even proteins with
Simultaneous Sequence Alignment and Tree Construction Using Hidden Markov Models. R.C. Edgar, K. Sjölander
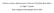 Simultaneous Sequence Alignment and Tree Construction Using Hidden Markov Models R.C. Edgar, K. Sjölander Pacific Symposium on Biocomputing 8:180-191(2003) SIMULTANEOUS SEQUENCE ALIGNMENT AND TREE CONSTRUCTION
Simultaneous Sequence Alignment and Tree Construction Using Hidden Markov Models R.C. Edgar, K. Sjölander Pacific Symposium on Biocomputing 8:180-191(2003) SIMULTANEOUS SEQUENCE ALIGNMENT AND TREE CONSTRUCTION
Detecting Distant Homologs Using Phylogenetic Tree-Based HMMs
 PROTEINS: Structure, Function, and Genetics 52:446 453 (2003) Detecting Distant Homologs Using Phylogenetic Tree-Based HMMs Bin Qian 1 and Richard A. Goldstein 1,2 * 1 Biophysics Research Division, University
PROTEINS: Structure, Function, and Genetics 52:446 453 (2003) Detecting Distant Homologs Using Phylogenetic Tree-Based HMMs Bin Qian 1 and Richard A. Goldstein 1,2 * 1 Biophysics Research Division, University
Received: 04 April 2006 Accepted: 25 July 2006
 BMC Bioinformatics BioMed Central Methodology article Improved alignment quality by combining evolutionary information, predicted secondary structure and self-organizing maps Tomas Ohlson 1, Varun Aggarwal
BMC Bioinformatics BioMed Central Methodology article Improved alignment quality by combining evolutionary information, predicted secondary structure and self-organizing maps Tomas Ohlson 1, Varun Aggarwal
Biologically significant sequence alignments using Boltzmann probabilities
 Biologically significant sequence alignments using Boltzmann probabilities P. Clote Department of Biology, Boston College Gasson Hall 416, Chestnut Hill MA 02467 clote@bc.edu May 7, 2003 Abstract In this
Biologically significant sequence alignments using Boltzmann probabilities P. Clote Department of Biology, Boston College Gasson Hall 416, Chestnut Hill MA 02467 clote@bc.edu May 7, 2003 Abstract In this
Sequence and Structure Alignment Z. Luthey-Schulten, UIUC Pittsburgh, 2006 VMD 1.8.5
 Sequence and Structure Alignment Z. Luthey-Schulten, UIUC Pittsburgh, 2006 VMD 1.8.5 Why Look at More Than One Sequence? 1. Multiple Sequence Alignment shows patterns of conservation 2. What and how many
Sequence and Structure Alignment Z. Luthey-Schulten, UIUC Pittsburgh, 2006 VMD 1.8.5 Why Look at More Than One Sequence? 1. Multiple Sequence Alignment shows patterns of conservation 2. What and how many
proteins Effect of using suboptimal alignments in template-based protein structure prediction Hao Chen 1 and Daisuke Kihara 1,2,3 * INTRODUCTION
 proteins STRUCTURE O FUNCTION O BIOINFORMATICS Effect of using suboptimal alignments in template-based protein structure prediction Hao Chen 1 and Daisuke Kihara 1,2,3 * 1 Department of Biological Sciences,
proteins STRUCTURE O FUNCTION O BIOINFORMATICS Effect of using suboptimal alignments in template-based protein structure prediction Hao Chen 1 and Daisuke Kihara 1,2,3 * 1 Department of Biological Sciences,
NRProF: Neural response based protein function prediction algorithm
 Title NRProF: Neural response based protein function prediction algorithm Author(s) Yalamanchili, HK; Wang, J; Xiao, QW Citation The 2011 IEEE International Conference on Systems Biology (ISB), Zhuhai,
Title NRProF: Neural response based protein function prediction algorithm Author(s) Yalamanchili, HK; Wang, J; Xiao, QW Citation The 2011 IEEE International Conference on Systems Biology (ISB), Zhuhai,
A New Similarity Measure among Protein Sequences
 A New Similarity Measure among Protein Sequences Kuen-Pin Wu, Hsin-Nan Lin, Ting-Yi Sung and Wen-Lian Hsu * Institute of Information Science Academia Sinica, Taipei 115, Taiwan Abstract Protein sequence
A New Similarity Measure among Protein Sequences Kuen-Pin Wu, Hsin-Nan Lin, Ting-Yi Sung and Wen-Lian Hsu * Institute of Information Science Academia Sinica, Taipei 115, Taiwan Abstract Protein sequence
SimShift: Identifying structural similarities from NMR chemical shifts
 BIOINFORMATICS ORIGINAL PAPER Vol. 22 no. 4 2006, pages 460 465 doi:10.1093/bioinformatics/bti805 Structural bioinformatics SimShift: Identifying structural similarities from NMR chemical shifts Simon
BIOINFORMATICS ORIGINAL PAPER Vol. 22 no. 4 2006, pages 460 465 doi:10.1093/bioinformatics/bti805 Structural bioinformatics SimShift: Identifying structural similarities from NMR chemical shifts Simon
Template-Based Modeling of Protein Structure
 Template-Based Modeling of Protein Structure David Constant Biochemistry 218 December 11, 2011 Introduction. Much can be learned about the biology of a protein from its structure. Simply put, structure
Template-Based Modeling of Protein Structure David Constant Biochemistry 218 December 11, 2011 Introduction. Much can be learned about the biology of a protein from its structure. Simply put, structure
PROTEIN FUNCTION PREDICTION WITH AMINO ACID SEQUENCE AND SECONDARY STRUCTURE ALIGNMENT SCORES
 PROTEIN FUNCTION PREDICTION WITH AMINO ACID SEQUENCE AND SECONDARY STRUCTURE ALIGNMENT SCORES Eser Aygün 1, Caner Kömürlü 2, Zafer Aydin 3 and Zehra Çataltepe 1 1 Computer Engineering Department and 2
PROTEIN FUNCTION PREDICTION WITH AMINO ACID SEQUENCE AND SECONDARY STRUCTURE ALIGNMENT SCORES Eser Aygün 1, Caner Kömürlü 2, Zafer Aydin 3 and Zehra Çataltepe 1 1 Computer Engineering Department and 2
HIDDEN MARKOV MODELS FOR REMOTE PROTEIN HOMOLOGY DETECTION
 From THE CENTER FOR GENOMICS AND BIOINFORMATICS Karolinska Institutet, Stockholm, Sweden HIDDEN MARKOV MODELS FOR REMOTE PROTEIN HOMOLOGY DETECTION Markus Wistrand Stockholm 2005 All previously published
From THE CENTER FOR GENOMICS AND BIOINFORMATICS Karolinska Institutet, Stockholm, Sweden HIDDEN MARKOV MODELS FOR REMOTE PROTEIN HOMOLOGY DETECTION Markus Wistrand Stockholm 2005 All previously published
Protein Structure Prediction using String Kernels. Technical Report
 Protein Structure Prediction using String Kernels Technical Report Department of Computer Science and Engineering University of Minnesota 4-192 EECS Building 200 Union Street SE Minneapolis, MN 55455-0159
Protein Structure Prediction using String Kernels Technical Report Department of Computer Science and Engineering University of Minnesota 4-192 EECS Building 200 Union Street SE Minneapolis, MN 55455-0159
Sequence Alignments. Dynamic programming approaches, scoring, and significance. Lucy Skrabanek ICB, WMC January 31, 2013
 Sequence Alignments Dynamic programming approaches, scoring, and significance Lucy Skrabanek ICB, WMC January 31, 213 Sequence alignment Compare two (or more) sequences to: Find regions of conservation
Sequence Alignments Dynamic programming approaches, scoring, and significance Lucy Skrabanek ICB, WMC January 31, 213 Sequence alignment Compare two (or more) sequences to: Find regions of conservation
Sara C. Madeira. Universidade da Beira Interior. (Thanks to Ana Teresa Freitas, IST for useful resources on this subject)
 Bioinformática Sequence Alignment Pairwise Sequence Alignment Universidade da Beira Interior (Thanks to Ana Teresa Freitas, IST for useful resources on this subject) 1 16/3/29 & 23/3/29 27/4/29 Outline
Bioinformática Sequence Alignment Pairwise Sequence Alignment Universidade da Beira Interior (Thanks to Ana Teresa Freitas, IST for useful resources on this subject) 1 16/3/29 & 23/3/29 27/4/29 Outline
COMPASS server for homology detection: improved statistical accuracy, speed and functionality
 Nucleic Acids Research Advance Access published May 12, 2009 Nucleic Acids Research, 2009, 1 5 doi:10.1093/nar/gkp360 COMPASS server for homology detection: improved statistical accuracy, speed and functionality
Nucleic Acids Research Advance Access published May 12, 2009 Nucleic Acids Research, 2009, 1 5 doi:10.1093/nar/gkp360 COMPASS server for homology detection: improved statistical accuracy, speed and functionality
Integrating multi-attribute similarity networks for robust representation of the protein space
 BIOINFORMATICS ORIGINAL PAPER Vol. 22 no. 13 2006, pages 1585 1592 doi:10.1093/bioinformatics/btl130 Structural bioinformatics Integrating multi-attribute similarity networks for robust representation
BIOINFORMATICS ORIGINAL PAPER Vol. 22 no. 13 2006, pages 1585 1592 doi:10.1093/bioinformatics/btl130 Structural bioinformatics Integrating multi-attribute similarity networks for robust representation
MUSCLE: multiple sequence alignment with high accuracy and high throughput
 Published online March 19, 2004 1792±1797 Nucleic Acids Research, 2004, Vol. 32, No. 5 DOI: 10.1093/nar/gkh340 MUSCLE: multiple sequence alignment with high accuracy and high throughput Robert C. Edgar*
Published online March 19, 2004 1792±1797 Nucleic Acids Research, 2004, Vol. 32, No. 5 DOI: 10.1093/nar/gkh340 MUSCLE: multiple sequence alignment with high accuracy and high throughput Robert C. Edgar*
SUPPLEMENTARY INFORMATION
 Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Supplementary information S3 (box) Methods Methods Genome weighting The currently available collection of archaeal and bacterial genomes has a highly biased distribution of isolates across taxa. For example,
Large-Scale Genomic Surveys
 Bioinformatics Subtopics Fold Recognition Secondary Structure Prediction Docking & Drug Design Protein Geometry Protein Flexibility Homology Modeling Sequence Alignment Structure Classification Gene Prediction
Bioinformatics Subtopics Fold Recognition Secondary Structure Prediction Docking & Drug Design Protein Geometry Protein Flexibility Homology Modeling Sequence Alignment Structure Classification Gene Prediction
Exercise 5. Sequence Profiles & BLAST
 Exercise 5 Sequence Profiles & BLAST 1 Substitution Matrix (BLOSUM62) Likelihood to substitute one amino acid with another Figure taken from https://en.wikipedia.org/wiki/blosum 2 Substitution Matrix (BLOSUM62)
Exercise 5 Sequence Profiles & BLAST 1 Substitution Matrix (BLOSUM62) Likelihood to substitute one amino acid with another Figure taken from https://en.wikipedia.org/wiki/blosum 2 Substitution Matrix (BLOSUM62)
Introduction to Bioinformatics
 Introduction to Bioinformatics Jianlin Cheng, PhD Department of Computer Science Informatics Institute 2011 Topics Introduction Biological Sequence Alignment and Database Search Analysis of gene expression
Introduction to Bioinformatics Jianlin Cheng, PhD Department of Computer Science Informatics Institute 2011 Topics Introduction Biological Sequence Alignment and Database Search Analysis of gene expression
Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics
 Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics Tandy Warnow Founder Professor of Engineering The University of Illinois at Urbana-Champaign http://tandy.cs.illinois.edu
Using Ensembles of Hidden Markov Models for Grand Challenges in Bioinformatics Tandy Warnow Founder Professor of Engineering The University of Illinois at Urbana-Champaign http://tandy.cs.illinois.edu
Homology Modeling (Comparative Structure Modeling) GBCB 5874: Problem Solving in GBCB
 Homology Modeling (Comparative Structure Modeling) Aims of Structural Genomics High-throughput 3D structure determination and analysis To determine or predict the 3D structures of all the proteins encoded
Homology Modeling (Comparative Structure Modeling) Aims of Structural Genomics High-throughput 3D structure determination and analysis To determine or predict the 3D structures of all the proteins encoded
A greedy, graph-based algorithm for the alignment of multiple homologous gene lists
 A greedy, graph-based algorithm for the alignment of multiple homologous gene lists Jan Fostier, Sebastian Proost, Bart Dhoedt, Yvan Saeys, Piet Demeester, Yves Van de Peer, and Klaas Vandepoele Bioinformatics
A greedy, graph-based algorithm for the alignment of multiple homologous gene lists Jan Fostier, Sebastian Proost, Bart Dhoedt, Yvan Saeys, Piet Demeester, Yves Van de Peer, and Klaas Vandepoele Bioinformatics
Detection of unrelated proteins in sequences multiple alignments by using predicted secondary structures
 BIOINFORMATICS Vol. 19 no. 4 2003, pages 506 512 DOI:.93/bioinformatics/btg016 Detection of unrelated proteins in sequences multiple alignments by using predicted secondary structures Mounir Errami, Christophe
BIOINFORMATICS Vol. 19 no. 4 2003, pages 506 512 DOI:.93/bioinformatics/btg016 Detection of unrelated proteins in sequences multiple alignments by using predicted secondary structures Mounir Errami, Christophe
Sequence Analysis '17- lecture 8. Multiple sequence alignment
 Sequence Analysis '17- lecture 8 Multiple sequence alignment Ex5 explanation How many random database search scores have e-values 10? (Answer: 10!) Why? e-value of x = m*p(s x), where m is the database
Sequence Analysis '17- lecture 8 Multiple sequence alignment Ex5 explanation How many random database search scores have e-values 10? (Answer: 10!) Why? e-value of x = m*p(s x), where m is the database
Copyright 2000 N. AYDIN. All rights reserved. 1
 Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Introduction to Bioinformatics Prof. Dr. Nizamettin AYDIN naydin@yildiz.edu.tr Multiple Sequence Alignment Outline Multiple sequence alignment introduction to msa methods of msa progressive global alignment
Sequence Alignment: A General Overview. COMP Fall 2010 Luay Nakhleh, Rice University
 Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Sequence Alignment: A General Overview COMP 571 - Fall 2010 Luay Nakhleh, Rice University Life through Evolution All living organisms are related to each other through evolution This means: any pair of
Identification of correct regions in protein models using structural, alignment, and consensus information
 Identification of correct regions in protein models using structural, alignment, and consensus information BJO RN WALLNER AND ARNE ELOFSSON Stockholm Bioinformatics Center, Stockholm University, SE-106
Identification of correct regions in protein models using structural, alignment, and consensus information BJO RN WALLNER AND ARNE ELOFSSON Stockholm Bioinformatics Center, Stockholm University, SE-106
SEQUENCE alignment is an underlying application in the
 194 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 8, NO. 1, JANUARY/FEBRUARY 2011 Pairwise Statistical Significance of Local Sequence Alignment Using Sequence-Specific and Position-Specific
194 IEEE/ACM TRANSACTIONS ON COMPUTATIONAL BIOLOGY AND BIOINFORMATICS, VOL. 8, NO. 1, JANUARY/FEBRUARY 2011 Pairwise Statistical Significance of Local Sequence Alignment Using Sequence-Specific and Position-Specific
Similarity searching summary (2)
 Similarity searching / sequence alignment summary Biol4230 Thurs, February 22, 2016 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 What have we covered? Homology excess similiarity but no excess similarity
Similarity searching / sequence alignment summary Biol4230 Thurs, February 22, 2016 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 What have we covered? Homology excess similiarity but no excess similarity
Page 1. References. Hidden Markov models and multiple sequence alignment. Markov chains. Probability review. Example. Markovian sequence
 Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
CMPS 3110: Bioinformatics. Tertiary Structure Prediction
 CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
CMPS 3110: Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the laws of physics! Conformation space is finite
CMPS 6630: Introduction to Computational Biology and Bioinformatics. Tertiary Structure Prediction
 CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
CMPS 6630: Introduction to Computational Biology and Bioinformatics Tertiary Structure Prediction Tertiary Structure Prediction Why Should Tertiary Structure Prediction Be Possible? Molecules obey the
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment
 Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Protein Structure Prediction
 Page 1 Protein Structure Prediction Russ B. Altman BMI 214 CS 274 Protein Folding is different from structure prediction --Folding is concerned with the process of taking the 3D shape, usually based on
Page 1 Protein Structure Prediction Russ B. Altman BMI 214 CS 274 Protein Folding is different from structure prediction --Folding is concerned with the process of taking the 3D shape, usually based on
Optimization of a New Score Function for the Detection of Remote Homologs
 PROTEINS: Structure, Function, and Genetics 41:498 503 (2000) Optimization of a New Score Function for the Detection of Remote Homologs Maricel Kann, 1 Bin Qian, 2 and Richard A. Goldstein 1,2 * 1 Department
PROTEINS: Structure, Function, and Genetics 41:498 503 (2000) Optimization of a New Score Function for the Detection of Remote Homologs Maricel Kann, 1 Bin Qian, 2 and Richard A. Goldstein 1,2 * 1 Department
Alignment principles and homology searching using (PSI-)BLAST. Jaap Heringa Centre for Integrative Bioinformatics VU (IBIVU)
 Alignment principles and homology searching using (PSI-)BLAST Jaap Heringa Centre for Integrative Bioinformatics VU (IBIVU) http://ibivu.cs.vu.nl Bioinformatics Nothing in Biology makes sense except in
Alignment principles and homology searching using (PSI-)BLAST Jaap Heringa Centre for Integrative Bioinformatics VU (IBIVU) http://ibivu.cs.vu.nl Bioinformatics Nothing in Biology makes sense except in
Pairwise sequence alignment and pair hidden Markov models
 Pairwise sequence alignment and pair hidden Markov models Martin C. Frith April 13, 2012 ntroduction Pairwise alignment and pair hidden Markov models (phmms) are basic textbook fare [2]. However, there
Pairwise sequence alignment and pair hidden Markov models Martin C. Frith April 13, 2012 ntroduction Pairwise alignment and pair hidden Markov models (phmms) are basic textbook fare [2]. However, there
Citation BMC Bioinformatics, 2016, v. 17 n. suppl. 8, p. 285:
 Title PnpProbs: A better multiple sequence alignment tool by better handling of guide trees Author(s) YE, Y; Lam, TW; Ting, HF Citation BMC Bioinformatics, 2016, v. 17 n. suppl. 8, p. 285:633-643 Issued
Title PnpProbs: A better multiple sequence alignment tool by better handling of guide trees Author(s) YE, Y; Lam, TW; Ting, HF Citation BMC Bioinformatics, 2016, v. 17 n. suppl. 8, p. 285:633-643 Issued
Lecture 7 Sequence analysis. Hidden Markov Models
 Lecture 7 Sequence analysis. Hidden Markov Models Nicolas Lartillot may 2012 Nicolas Lartillot (Universite de Montréal) BIN6009 may 2012 1 / 60 1 Motivation 2 Examples of Hidden Markov models 3 Hidden
Lecture 7 Sequence analysis. Hidden Markov Models Nicolas Lartillot may 2012 Nicolas Lartillot (Universite de Montréal) BIN6009 may 2012 1 / 60 1 Motivation 2 Examples of Hidden Markov models 3 Hidden
Computational Genomics and Molecular Biology, Fall
 Computational Genomics and Molecular Biology, Fall 2014 1 HMM Lecture Notes Dannie Durand and Rose Hoberman November 6th Introduction In the last few lectures, we have focused on three problems related
Computational Genomics and Molecular Biology, Fall 2014 1 HMM Lecture Notes Dannie Durand and Rose Hoberman November 6th Introduction In the last few lectures, we have focused on three problems related
Do Aligned Sequences Share the Same Fold?
 J. Mol. Biol. (1997) 273, 355±368 Do Aligned Sequences Share the Same Fold? Ruben A. Abagyan* and Serge Batalov The Skirball Institute of Biomolecular Medicine Biochemistry Department NYU Medical Center
J. Mol. Biol. (1997) 273, 355±368 Do Aligned Sequences Share the Same Fold? Ruben A. Abagyan* and Serge Batalov The Skirball Institute of Biomolecular Medicine Biochemistry Department NYU Medical Center
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Stephen Scott.
 1 / 21 sscott@cse.unl.edu 2 / 21 Introduction Designed to model (profile) a multiple alignment of a protein family (e.g., Fig. 5.1) Gives a probabilistic model of the proteins in the family Useful for
1 / 21 sscott@cse.unl.edu 2 / 21 Introduction Designed to model (profile) a multiple alignment of a protein family (e.g., Fig. 5.1) Gives a probabilistic model of the proteins in the family Useful for
P rotein structure alignment is a fundamental problem in computational structure biology and has been
 SUBJECT AREAS: PROTEIN ANALYSIS SOFTWARE STRUCTURAL BIOLOGY BIOINFORMATICS Received 15 November 2012 Accepted 25 February 2013 Published 14 March 2013 Correspondence and requests for materials should be
SUBJECT AREAS: PROTEIN ANALYSIS SOFTWARE STRUCTURAL BIOLOGY BIOINFORMATICS Received 15 November 2012 Accepted 25 February 2013 Published 14 March 2013 Correspondence and requests for materials should be
Some Problems from Enzyme Families
 Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
Some Problems from Enzyme Families Greg Butler Department of Computer Science Concordia University, Montreal www.cs.concordia.ca/~faculty/gregb gregb@cs.concordia.ca Abstract I will discuss some problems
A Tool for Structure Alignment of Molecules
 A Tool for Structure Alignment of Molecules Pei-Ken Chang, Chien-Cheng Chen and Ming Ouhyoung Department of Computer Science and Information Engineering, National Taiwan University {zick, ccchen}@cmlab.csie.ntu.edu.tw,
A Tool for Structure Alignment of Molecules Pei-Ken Chang, Chien-Cheng Chen and Ming Ouhyoung Department of Computer Science and Information Engineering, National Taiwan University {zick, ccchen}@cmlab.csie.ntu.edu.tw,
Subfamily HMMS in Functional Genomics. D. Brown, N. Krishnamurthy, J.M. Dale, W. Christopher, and K. Sjölander
 Subfamily HMMS in Functional Genomics D. Brown, N. Krishnamurthy, J.M. Dale, W. Christopher, and K. Sjölander Pacific Symposium on Biocomputing 10:322-333(2005) SUBFAMILY HMMS IN FUNCTIONAL GENOMICS DUNCAN
Subfamily HMMS in Functional Genomics D. Brown, N. Krishnamurthy, J.M. Dale, W. Christopher, and K. Sjölander Pacific Symposium on Biocomputing 10:322-333(2005) SUBFAMILY HMMS IN FUNCTIONAL GENOMICS DUNCAN
CISC 889 Bioinformatics (Spring 2004) Sequence pairwise alignment (I)
 CISC 889 Bioinformatics (Spring 2004) Sequence pairwise alignment (I) Contents Alignment algorithms Needleman-Wunsch (global alignment) Smith-Waterman (local alignment) Heuristic algorithms FASTA BLAST
CISC 889 Bioinformatics (Spring 2004) Sequence pairwise alignment (I) Contents Alignment algorithms Needleman-Wunsch (global alignment) Smith-Waterman (local alignment) Heuristic algorithms FASTA BLAST
Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics
 Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics Jianlin Cheng, PhD Department of Computer Science University of Missouri, Columbia
Statistical Machine Learning Methods for Bioinformatics IV. Neural Network & Deep Learning Applications in Bioinformatics Jianlin Cheng, PhD Department of Computer Science University of Missouri, Columbia
