What is the expectation maximization algorithm?
|
|
|
- Britton Lloyd
- 5 years ago
- Views:
Transcription
1 primer 2008 Nature Publishing Group What is the expectation maximization algorithm? Chuong B Do & Serafim Batzoglou The expectation maximization algorithm arises in many computational biology applications that involve probabilistic models. What is it good for, and how does it work? Probabilistic models, such as hidden Markov models or Bayesian networks, are commonly used to model biological data. Much of their popularity can be attributed to the existence of efficient and robust procedures for learning parameters from observations. Often, however, the only data available for training a probabilistic model are incomplete. Missing values can occur, for example, in medical diagnosis, where patient histories generally include results from a limited battery of tests. Alternatively, in gene expression clustering, incomplete data arise from the intentional omission of gene-to-cluster assignments in the probabilistic model. The expectation maximization algorithm enables parameter estimation in probabilistic models with incomplete data. A coin-flipping experiment As an example, consider a simple coin-flipping experiment in which we are given a pair of coins A and B of unknown biases, and, respectively (that is, on any given flip, coin A will land on heads with probability and tails with probability 1 and similarly for coin B). Our goal is to estimate θ = (, ) by repeating the following procedure five times: randomly choose one of the two coins (with equal probability), and perform ten independent coin tosses with the selected coin. Thus, the entire procedure involves a total of 50 coin tosses (Fig. 1a). During our experiment, suppose that we keep track of two vectors x = (x 1, x 2,, x 5 ) and Chuong B. Do and Serafim Batzoglou are in the Computer Science Department, Stanford University, 318 Campus Drive, Stanford, California , USA. chuong@cs.stanford.edu z = (z 1, z 2,, z 5 ), where x i {0,1,,10} is the number of heads observed during the ith set of tosses, and z i {A,B} is the identity of the coin used during the ith set of tosses. Parameter estimation in this setting is known as the complete data case in that the values of all relevant random variables in our model (that is, the result of each coin flip and the type of coin used for each flip) are known. Here, a simple way to estimate and is to return the observed proportions of heads for each coin: and # of heads using coin A θ Αˆ = total # of flips using coin A # of heads using coin B θ Βˆ = total # of flips using coin B (1) This intuitive guess is, in fact, known in the statistical literature as maximum likelihood estimation (roughly speaking, the maximum likelihood method assesses the quality of a statistical model based on the probability it assigns to the observed data). If logp(x,z;θ) is the logarithm of the joint probability (or loglikelihood) of obtaining any particular vector of observed head counts x and coin types z, then the formulas in (1) solve for the parameters θˆ = ( θ ˆ ˆ A, ) that maximize logp(x,z;θ). Now consider a more challenging variant of the parameter estimation problem in which we are given the recorded head counts x but not the identities z of the coins used for each set of tosses. We refer to z as hidden variables or latent factors. Parameter estimation in this new setting is known as the incomplete data case. This time, computing proportions of heads for each coin is no longer possible, because we don t know the coin used for each set of tosses. However, if we had some way of completing the data (in our case, guessing correctly which coin was used in each of the five sets), then we could reduce parameter estimation for this problem with incomplete data to maximum likelihood estimation with complete data. One iterative scheme for obtaining completions could work as follows: starting from some initial parameters, (t) θˆ = ( θ ˆ (t) ˆ (t) Α,θ Β ), determine for each of the five sets whether coin A or coin B was more likely to have generated the observed flips (using the current parameter estimates). Then, assume these completions (that is, guessed coin assignments) to be correct, and apply the regular maximum likelihood estimation procedure to get θˆ(t+1). Finally, repeat these two steps until convergence. As the estimated model improves, so too will the quality of the resulting completions. The expectation maximization algorithm is a refinement on this basic idea. Rather than picking the single most likely completion of the missing coin assignments on each iteration, the expectation maximization algorithm computes probabilities for each possible completion of the missing data, using the current parameters θˆ(t). These probabilities are used to create a weighted training set consisting of all possible completions of the data. Finally, a modified version of maximum likelihood estimation that deals with weighted training examples provides new parameter estimates, θˆ(t+1). By using weighted training examples rather than choosing the single best completion, the expectation maximization algorithm accounts for the confidence of the model in each completion of the data (Fig. 1b). In summary, the expectation maximization algorithm alternates between the steps nature biotechnology volume 26 number 8 august
2 primer 2008 Nature Publishing Group of guessing a probability distribution over completions of missing data given the current model (known as the E-step) and then reestimating the model parameters using these completions (known as the M-step). The name E-step comes from the fact that one does not usually need to form the probability distribution over completions explicitly, but rather need only compute expected sufficient statistics over these completions. Similarly, the name M-step comes from the fact that model reestimation can be thought of as maximization of the expected log-likelihood of the data. Introduced as early as 1950 by Ceppellini et al. 1 in the context of gene frequency estimation, the expectation maximization algorithm a b Maximum likelihood H T T T HHT H T H HHHHT HHHHH H T H H H HHT H H H T H T T T H H T T T HHH T HHH T H 5 sets, 10 tosses per set Expectation maximization 1 E-step H T T T H H T H T H H H H H T H H H H H H T H H H H H T H H H T H T T T H H T T T H H H T H H H T H ˆ = 0.60 (0) ˆ = 0.50 (0) x 0.80 x 0.73 x 0.35 x 0.65 x (1) Coin A 9 H, 1 T 8 H, 2 T 7 H, 3 T 24 H, 6 T was analyzed more generally by Hartley 2 and by Baum et al. 3 in the context of hidden Markov models, where it is commonly known as the Baum-Welch algorithm. The standard reference on the expectation maximization algorithm and its convergence is Dempster et al 4. Mathematical foundations How does the expectation maximization algorithm work? More importantly, why is it even necessary? The expectation maximization algorithm is a natural generalization of maximum likelihood estimation to the incomplete data case. In particular, expectation maximization attempts to find the parameters θˆ that maximize the Coin B 5 H, 5 T 4 H, 6 T 9 H, 11 T ˆ ˆ (1) 0.55 x 0.20 x 0.27 x 0.65 x 0.35x ˆ 24 = = ˆ 9 = = Coin A 2.2 H, 2.2 T 7.2 H, 0.8 T 5.9 H, 1.5 T 1.4 H, 2.1 T 4.5 H, 1.9 T 21.3 H, 8.6 T 3 M-step 4 Coin B 2.8 H, 2.8 T 1.8 H, 0.2 T 2.1 H, 0.5 T 2.6 H, 3.9 T 2.5 H, 1.1 T 11.7 H, 8.4 T ˆ 0.80 (10) ˆ 0.52 (10) Figure 1 Parameter estimation for complete and incomplete data. (a) Maximum likelihood estimation. For each set of ten tosses, the maximum likelihood procedure accumulates the counts of heads and tails for coins A and B separately. These counts are then used to estimate the coin biases. (b) Expectation maximization. 1. EM starts with an initial guess of the parameters. 2. In the E-step, a probability distribution over possible completions is computed using the current parameters. The counts shown in the table are the expected numbers of heads and tails according to this distribution. 3. In the M-step, new parameters are determined using the current completions. 4. After several repetitions of the E-step and M-step, the algorithm converges. log probability logp(x;θ) of the observed data. Generally speaking, the optimization problem addressed by the expectation maximization algorithm is more difficult than the optimization used in maximum likelihood estimation. In the complete data case, the objective function logp(x,z;θ) has a single global optimum, which can often be found in closed form (e.g., equation 1). In contrast, in the incomplete data case the function logp(x;θ) has multiple local maxima and no closed form solution. To deal with this, the expectation maximization algorithm reduces the difficult task of optimizing logp(x;θ) into a sequence of simpler optimization subproblems, whose objective functions have unique global maxima that can often be computed in closed form. These subproblems are chosen in a way that guarantees their corresponding solutions θˆ(1),θˆ(2), and will converge to a local optimum of logp(x;θ). More specifically, the expectation maximization algorithm alternates between two phases. During the E-step, expectation maximization chooses a function g t that lower bounds logp(x;θ) everywhere, and for which g θˆ(t) t ( )=logp(x; θˆ(t) ). During the M-step, the expectation maximization algorithm moves to a new parameter set θˆ(t+1) that maximizes g t. As the value of the lower-bound g t matches the objective function at θˆ(t), it follows that logp(x; θˆ(t) )= g θˆ (t) t ( ) g t ( θˆ(t+1) ) = logp(x; θˆ(t+1) ) s o the objective function monotonically increases during each iteration of expectation maximization! A graphical illustration of this argument is provided in Supplementary Figure 1 online, and a concise mathematical derivation of the expectation maximization algorithm is given in Supplementary Note 1 online. As with most optimization methods for nonconcave functions, the expectation maximization algorithm comes with guarantees only of convergence to a local maximum of the objective function (except in degenerate cases). Running the procedure using multiple initial starting parameters is often helpful; similarly, initializing parameters in a way that breaks symmetry in models is also important. With this limited set of tricks, the expectation maximization algorithm provides a simple and robust tool for parameter estimation in models with incomplete data. In theory, other numerical optimization techniques, such as gradient descent or Newton-Raphson, could be used instead of expectation maximization; in practice, however, expectation maximization has the advantage of being simple, robust and easy to implement. Applications Many probabilistic models in computational biology include latent variables. In some 898 volume 26 number 8 august 2008 nature biotechnology
3 primer 2008 Nature Publishing Group cases, these latent variables are present due to missing or corrupted data; in most applications of expectation maximization to computational biology, however, the latent factors are intentionally included, and parameter learning itself provides a mechanism for knowledge discovery. For instance, in gene expression clustering 5, we are given microarray gene expression measurements for thousands of genes under varying conditions, and our goal is to group the observed expression vectors into distinct clusters of related genes. One approach is to model the vector of expression measurements for each gene as being sampled from a multivariate Gaussian distribution (a generalization of a standard Gaussian distribution to multiple correlated variables) associated with that gene s cluster. In this case, the observed data x correspond to microarray measurements, the unobserved latent factors z are the assignments of genes to clusters, and the parameters θ include the means and covariance matrices of the multivariate Gaussian distributions representing the expression patterns for each cluster. For parameter learning, the expectation maximization algorithm alternates between computing probabilities for assignments of each gene to each cluster (E-step) and updating the cluster means and covariances based on the set of genes predominantly belonging to that cluster (M-step). This can be thought of as a soft variant of the popular k-means clustering algorithm, in which one alternates between hard assignments of genes to clusters and reestimation of cluster means based on their assigned genes. In motif finding 6, we are given a set of unaligned DNA sequences and asked to identify a pattern of length W that is present (though possibly with minor variations) in every sequence from the set. To apply the expectation maximization algorithm, we model the instance of the motif in each sequence as having each letter sampled independently from a position-specific distribution over letters, and the remaining letters in each sequence as coming from some fixed background distribution. The observed data x consist of the letters of sequences, the unobserved latent factors z include the starting position of the motif in each sequence and the parameters θ describe the position-specific letter frequencies for the motif. Here, the expectation maximization algorithm involves computing the probability distribution over motif start positions for each sequence (E-step) and updating the motif letter frequencies based on the expected letter counts for each position in the motif (M-step). In the haplotype inference problem 7, we are given the unphased genotypes of individuals from some population, where each unphased genotype consists of unordered pairs of single-nucleotide polymorphisms (SNPs) taken from homologous chromosomes of the individual. Contiguous blocks of SNPs inherited from a single chromosome are called haplotypes. Assuming for simplicity that each individual s genotype is a combination of two haplotypes (one maternal and one paternal), the goal of haplotype inference is to determine a small set of haplotypes that best explain all of the unphased genotypes observed in the population. Here, the observed data x are the known unphased genotypes for each individual, the unobserved latent factors z are putative assignments of unphased genotypes to pairs of haplotypes and the parameters θ describe the frequencies of each haplotype in the population. The expectation maximization algorithm alternates between using the current haplotype frequencies to estimate probability distributions over phasing assignments for each unphased genotype (E-step) and using the expected phasing assignments to refine estimates of haplotype frequencies (M-step). Other problems in which the expectation maximization algorithm plays a prominent role include learning profiles of protein domains 8 and RNA families 9, discovery of transcriptional modules 10, tests of linkage disequilibrium 11, protein identification 12 and medical imaging 13. In each case, expectation maximization provides a simple, easy-to-implement and efficient tool for learning parameters of a model; once these parameters are known, we can use probabilistic inference to ask interesting queries about the model. For example, what cluster does a particular gene most likely belong to? What is the most likely starting location of a motif in a particular sequence? What are the most likely haplotype blocks making up the genotype of a specific individual? By providing a straightforward mechanism for parameter learning in all of these models, expectation maximization provides a mechanism for building and training rich probabilistic models for biological applications. Note: Supplementary information is available on the Nature Biotechnology website. ACKNOWLEDGMENTS C.B.D. is supported in part by an National Science Foundation (NSF) Graduate Fellowship. S.B. wishes to acknowledge support by the NSF CAREER Award. We thank four anonymous referees for helpful suggestions. 1. Ceppellini, R., Siniscalco, M. & Smith, C.A. Ann. Hum. Genet. 20, (1955). 2. Hartley, H. Biometrics 14, (1958). 3. Baum, L.E., Petrie, T., Soules, G. & Weiss, N. Ann. Math. Stat. 41, (1970). 4. Dempster, A.P., Laird, N.M. & Rubin, D.B. J. R. Stat. Soc. Ser. B 39, 1 38 (1977). 5. D haeseleer, P. Nat. Biotechnol. 23, (2005). 6. Lawrence, C.E. & Reilly, A.A. Proteins 7, (1990). 7. Excoffier, L. & Slatkin, M. Mol. Biol. Evol. 12, (1995). 8. Krogh, A., Brown, M., Mian, I.S., Sjölander, K. & Haussler, D. J. Mol. Biol. 235, (1994). 9. Eddy, S.R. & Durbin, R. Nucleic Acids Res. 22, (1994). 10. Segal, E., Yelensky, R. & Koller, D. Bioinformatics 19, i273 i282 (2003). 11. Slatkin, M. & Excoffier, L. Heredity 76, (1996). 12. Nesvizhskii, A.I., Keller, A., Kolker, E. & Aebersold, R. Anal. Chem. 75, (2003). 13. De Pierro, A.R. IEEE Trans. Med. Imaging 14, (1995). nature biotechnology volume 26 number 8 august
4 Supplementary Figure 1 Convergence of the EM algorithm. Starting from initial parameters, the E-step of the EM algorithm constructs a function that lower-bounds the objective function. In the M-step, is computed as the maximum of. In the next E-step, a new lower-bound is constructed; maximization of in the next M-step gives, etc.
5 Supplementary Note 1 Here, we provide a short derivation of the EM algorithm based on the idea of bound maximization. A more general (though somewhat subtle) argument, which leads to a number of variants of the EM algorithm, was described in: Neal RM and Hinton GE A view of the EM algorithm that justifies incremental, sparse, and other variants. In MI Jordan, ed., Learning in Graphical Models, Kluwer Academic Publishers, Mathematically, the EM algorithm derives from the fact that for any probability distribution, (3) where the inequality is tight (i.e. holds with equality) whenever. The derivation of (3) relies on a tool from mathematical analysis known as Jensen s inequality for concave functions. 1 Now, consider the update rule,, where (4) Applying the tightness conditions of equation (3),. However, observe that by definition of the update rule. Furthermore, since (3) guarantees that is a lower bound on for any parameter vector. Therefore, the update rule results in monotonic improvement of the maximum likelihood objective for incomplete data. To see the connection between (4) and the description of the EM algorithm given in the text, consider the following equivalent update rule: (5) Here, equivalence follows from the fact that the objective function of (5) differs from by a constant offset which does not depend on. In this final form, we see that the EM update rule effectively maximizes the log-likelihood of a dataset expanded to contain all possible completions 1 In brief, Jensen s inequality states that a concave function (e.g., ) of the expectation of a random variable (e.g., ) is never less than expected value of that concave function applied to the random variable (i.e., ); furthermore, equality holds if and only if the random variable is constant with probability 1 (i.e., ). Equation (3) follows from Jensen s inequality, using the random variable.
6 of the unobserved variables, where each completion is weighted by the posterior probability,.
Expectation-Maximization (EM) algorithm
 I529: Machine Learning in Bioinformatics (Spring 2017) Expectation-Maximization (EM) algorithm Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Contents Introduce
I529: Machine Learning in Bioinformatics (Spring 2017) Expectation-Maximization (EM) algorithm Yuzhen Ye School of Informatics and Computing Indiana University, Bloomington Spring 2017 Contents Introduce
The Expectation Maximization Algorithm
 The Expectation Maximization Algorithm Frank Dellaert College of Computing, Georgia Institute of Technology Technical Report number GIT-GVU-- February Abstract This note represents my attempt at explaining
The Expectation Maximization Algorithm Frank Dellaert College of Computing, Georgia Institute of Technology Technical Report number GIT-GVU-- February Abstract This note represents my attempt at explaining
O 3 O 4 O 5. q 3. q 4. Transition
 Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Hidden Markov Models Hidden Markov models (HMM) were developed in the early part of the 1970 s and at that time mostly applied in the area of computerized speech recognition. They are first described in
Lecture 6: April 19, 2002
 EE596 Pat. Recog. II: Introduction to Graphical Models Spring 2002 Lecturer: Jeff Bilmes Lecture 6: April 19, 2002 University of Washington Dept. of Electrical Engineering Scribe: Huaning Niu,Özgür Çetin
EE596 Pat. Recog. II: Introduction to Graphical Models Spring 2002 Lecturer: Jeff Bilmes Lecture 6: April 19, 2002 University of Washington Dept. of Electrical Engineering Scribe: Huaning Niu,Özgür Çetin
Mixture Models and EM
 Mixture Models and EM Goal: Introduction to probabilistic mixture models and the expectationmaximization (EM) algorithm. Motivation: simultaneous fitting of multiple model instances unsupervised clustering
Mixture Models and EM Goal: Introduction to probabilistic mixture models and the expectationmaximization (EM) algorithm. Motivation: simultaneous fitting of multiple model instances unsupervised clustering
Estimating Gaussian Mixture Densities with EM A Tutorial
 Estimating Gaussian Mixture Densities with EM A Tutorial Carlo Tomasi Due University Expectation Maximization (EM) [4, 3, 6] is a numerical algorithm for the maximization of functions of several variables
Estimating Gaussian Mixture Densities with EM A Tutorial Carlo Tomasi Due University Expectation Maximization (EM) [4, 3, 6] is a numerical algorithm for the maximization of functions of several variables
The E-M Algorithm in Genetics. Biostatistics 666 Lecture 8
 The E-M Algorithm in Genetics Biostatistics 666 Lecture 8 Maximum Likelihood Estimation of Allele Frequencies Find parameter estimates which make observed data most likely General approach, as long as
The E-M Algorithm in Genetics Biostatistics 666 Lecture 8 Maximum Likelihood Estimation of Allele Frequencies Find parameter estimates which make observed data most likely General approach, as long as
Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling
 Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 009 Mark Craven craven@biostat.wisc.edu Sequence Motifs what is a sequence
Learning Sequence Motif Models Using Expectation Maximization (EM) and Gibbs Sampling BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 009 Mark Craven craven@biostat.wisc.edu Sequence Motifs what is a sequence
Weighted Finite-State Transducers in Computational Biology
 Weighted Finite-State Transducers in Computational Biology Mehryar Mohri Courant Institute of Mathematical Sciences mohri@cims.nyu.edu Joint work with Corinna Cortes (Google Research). 1 This Tutorial
Weighted Finite-State Transducers in Computational Biology Mehryar Mohri Courant Institute of Mathematical Sciences mohri@cims.nyu.edu Joint work with Corinna Cortes (Google Research). 1 This Tutorial
Introduction to Machine Learning. Maximum Likelihood and Bayesian Inference. Lecturers: Eran Halperin, Lior Wolf
 1 Introduction to Machine Learning Maximum Likelihood and Bayesian Inference Lecturers: Eran Halperin, Lior Wolf 2014-15 We know that X ~ B(n,p), but we do not know p. We get a random sample from X, a
1 Introduction to Machine Learning Maximum Likelihood and Bayesian Inference Lecturers: Eran Halperin, Lior Wolf 2014-15 We know that X ~ B(n,p), but we do not know p. We get a random sample from X, a
Page 1. References. Hidden Markov models and multiple sequence alignment. Markov chains. Probability review. Example. Markovian sequence
 Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Page Hidden Markov models and multiple sequence alignment Russ B Altman BMI 4 CS 74 Some slides borrowed from Scott C Schmidler (BMI graduate student) References Bioinformatics Classic: Krogh et al (994)
Use of hidden Markov models for QTL mapping
 Use of hidden Markov models for QTL mapping Karl W Broman Department of Biostatistics, Johns Hopkins University December 5, 2006 An important aspect of the QTL mapping problem is the treatment of missing
Use of hidden Markov models for QTL mapping Karl W Broman Department of Biostatistics, Johns Hopkins University December 5, 2006 An important aspect of the QTL mapping problem is the treatment of missing
The Expectation-Maximization Algorithm
 1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
1/29 EM & Latent Variable Models Gaussian Mixture Models EM Theory The Expectation-Maximization Algorithm Mihaela van der Schaar Department of Engineering Science University of Oxford MLE for Latent Variable
A Gentle Tutorial of the EM Algorithm and its Application to Parameter Estimation for Gaussian Mixture and Hidden Markov Models
 A Gentle Tutorial of the EM Algorithm and its Application to Parameter Estimation for Gaussian Mixture and Hidden Markov Models Jeff A. Bilmes (bilmes@cs.berkeley.edu) International Computer Science Institute
A Gentle Tutorial of the EM Algorithm and its Application to Parameter Estimation for Gaussian Mixture and Hidden Markov Models Jeff A. Bilmes (bilmes@cs.berkeley.edu) International Computer Science Institute
Parametric Unsupervised Learning Expectation Maximization (EM) Lecture 20.a
 Parametric Unsupervised Learning Expectation Maximization (EM) Lecture 20.a Some slides are due to Christopher Bishop Limitations of K-means Hard assignments of data points to clusters small shift of a
Parametric Unsupervised Learning Expectation Maximization (EM) Lecture 20.a Some slides are due to Christopher Bishop Limitations of K-means Hard assignments of data points to clusters small shift of a
Expectation maximization tutorial
 Expectation maximization tutorial Octavian Ganea November 18, 2016 1/1 Today Expectation - maximization algorithm Topic modelling 2/1 ML & MAP Observed data: X = {x 1, x 2... x N } 3/1 ML & MAP Observed
Expectation maximization tutorial Octavian Ganea November 18, 2016 1/1 Today Expectation - maximization algorithm Topic modelling 2/1 ML & MAP Observed data: X = {x 1, x 2... x N } 3/1 ML & MAP Observed
Using Expectation-Maximization for Reinforcement Learning
 NOTE Communicated by Andrew Barto and Michael Jordan Using Expectation-Maximization for Reinforcement Learning Peter Dayan Department of Brain and Cognitive Sciences, Center for Biological and Computational
NOTE Communicated by Andrew Barto and Michael Jordan Using Expectation-Maximization for Reinforcement Learning Peter Dayan Department of Brain and Cognitive Sciences, Center for Biological and Computational
Hidden Markov Models for biological sequence analysis
 Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Hidden Markov Models for biological sequence analysis Master in Bioinformatics UPF 2017-2018 http://comprna.upf.edu/courses/master_agb/ Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA
Learning gene regulatory networks Statistical methods for haplotype inference Part I
 Learning gene regulatory networks Statistical methods for haplotype inference Part I Input: Measurement of mrn levels of all genes from microarray or rna sequencing Samples (e.g. 200 patients with lung
Learning gene regulatory networks Statistical methods for haplotype inference Part I Input: Measurement of mrn levels of all genes from microarray or rna sequencing Samples (e.g. 200 patients with lung
Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
An Introduction to Bioinformatics Algorithms Hidden Markov Models
 Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models Outline 1. CG-Islands 2. The Fair Bet Casino 3. Hidden Markov Model 4. Decoding Algorithm 5. Forward-Backward Algorithm 6. Profile HMMs 7. HMM Parameter Estimation 8. Viterbi Training
Hidden Markov Models for biological sequence analysis I
 Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Hidden Markov Models for biological sequence analysis I Master in Bioinformatics UPF 2014-2015 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Example: CpG Islands
Computer Vision Group Prof. Daniel Cremers. 6. Mixture Models and Expectation-Maximization
 Prof. Daniel Cremers 6. Mixture Models and Expectation-Maximization Motivation Often the introduction of latent (unobserved) random variables into a model can help to express complex (marginal) distributions
Prof. Daniel Cremers 6. Mixture Models and Expectation-Maximization Motivation Often the introduction of latent (unobserved) random variables into a model can help to express complex (marginal) distributions
ECE521 week 3: 23/26 January 2017
 ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
ECE521 week 3: 23/26 January 2017 Outline Probabilistic interpretation of linear regression - Maximum likelihood estimation (MLE) - Maximum a posteriori (MAP) estimation Bias-variance trade-off Linear
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation. Volker Tresp Summer 2014
 Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2014 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2014 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Expectation Maximization
 Expectation Maximization Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr 1 /
Expectation Maximization Seungjin Choi Department of Computer Science and Engineering Pohang University of Science and Technology 77 Cheongam-ro, Nam-gu, Pohang 37673, Korea seungjin@postech.ac.kr 1 /
Speech Recognition Lecture 8: Expectation-Maximization Algorithm, Hidden Markov Models.
 Speech Recognition Lecture 8: Expectation-Maximization Algorithm, Hidden Markov Models. Mehryar Mohri Courant Institute and Google Research mohri@cims.nyu.com This Lecture Expectation-Maximization (EM)
Speech Recognition Lecture 8: Expectation-Maximization Algorithm, Hidden Markov Models. Mehryar Mohri Courant Institute and Google Research mohri@cims.nyu.com This Lecture Expectation-Maximization (EM)
EM algorithm. Rather than jumping into the details of the particular EM algorithm, we ll look at a simpler example to get the idea of how it works
 EM algorithm The example in the book for doing the EM algorithm is rather difficult, and was not available in software at the time that the authors wrote the book, but they implemented a SAS macro to implement
EM algorithm The example in the book for doing the EM algorithm is rather difficult, and was not available in software at the time that the authors wrote the book, but they implemented a SAS macro to implement
U-Likelihood and U-Updating Algorithms: Statistical Inference in Latent Variable Models
 U-Likelihood and U-Updating Algorithms: Statistical Inference in Latent Variable Models Jaemo Sung 1, Sung-Yang Bang 1, Seungjin Choi 1, and Zoubin Ghahramani 2 1 Department of Computer Science, POSTECH,
U-Likelihood and U-Updating Algorithms: Statistical Inference in Latent Variable Models Jaemo Sung 1, Sung-Yang Bang 1, Seungjin Choi 1, and Zoubin Ghahramani 2 1 Department of Computer Science, POSTECH,
Gaussian Mixture Models
 Gaussian Mixture Models Pradeep Ravikumar Co-instructor: Manuela Veloso Machine Learning 10-701 Some slides courtesy of Eric Xing, Carlos Guestrin (One) bad case for K- means Clusters may overlap Some
Gaussian Mixture Models Pradeep Ravikumar Co-instructor: Manuela Veloso Machine Learning 10-701 Some slides courtesy of Eric Xing, Carlos Guestrin (One) bad case for K- means Clusters may overlap Some
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 11 Project
Computational Genomics. Systems biology. Putting it together: Data integration using graphical models
 02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
02-710 Computational Genomics Systems biology Putting it together: Data integration using graphical models High throughput data So far in this class we discussed several different types of high throughput
Hidden Markov Models
 Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
Andrea Passerini passerini@disi.unitn.it Statistical relational learning The aim Modeling temporal sequences Model signals which vary over time (e.g. speech) Two alternatives: deterministic models directly
Gibbs Sampling Methods for Multiple Sequence Alignment
 Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
Gibbs Sampling Methods for Multiple Sequence Alignment Scott C. Schmidler 1 Jun S. Liu 2 1 Section on Medical Informatics and 2 Department of Statistics Stanford University 11/17/99 1 Outline Statistical
MACHINE LEARNING INTRODUCTION: STRING CLASSIFICATION
 MACHINE LEARNING INTRODUCTION: STRING CLASSIFICATION THOMAS MAILUND Machine learning means different things to different people, and there is no general agreed upon core set of algorithms that must be
MACHINE LEARNING INTRODUCTION: STRING CLASSIFICATION THOMAS MAILUND Machine learning means different things to different people, and there is no general agreed upon core set of algorithms that must be
1.5 MM and EM Algorithms
 72 CHAPTER 1. SEQUENCE MODELS 1.5 MM and EM Algorithms The MM algorithm [1] is an iterative algorithm that can be used to minimize or maximize a function. We focus on using it to maximize the log likelihood
72 CHAPTER 1. SEQUENCE MODELS 1.5 MM and EM Algorithms The MM algorithm [1] is an iterative algorithm that can be used to minimize or maximize a function. We focus on using it to maximize the log likelihood
Based on slides by Richard Zemel
 CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 3: Directed Graphical Models and Latent Variables Based on slides by Richard Zemel Learning outcomes What aspects of a model can we
CSC 412/2506 Winter 2018 Probabilistic Learning and Reasoning Lecture 3: Directed Graphical Models and Latent Variables Based on slides by Richard Zemel Learning outcomes What aspects of a model can we
Machine Learning for Signal Processing Expectation Maximization Mixture Models. Bhiksha Raj 27 Oct /
 Machine Learning for Signal rocessing Expectation Maximization Mixture Models Bhiksha Raj 27 Oct 2016 11755/18797 1 Learning Distributions for Data roblem: Given a collection of examples from some data,
Machine Learning for Signal rocessing Expectation Maximization Mixture Models Bhiksha Raj 27 Oct 2016 11755/18797 1 Learning Distributions for Data roblem: Given a collection of examples from some data,
STATS 306B: Unsupervised Learning Spring Lecture 3 April 7th
 STATS 306B: Unsupervised Learning Spring 2014 Lecture 3 April 7th Lecturer: Lester Mackey Scribe: Jordan Bryan, Dangna Li 3.1 Recap: Gaussian Mixture Modeling In the last lecture, we discussed the Gaussian
STATS 306B: Unsupervised Learning Spring 2014 Lecture 3 April 7th Lecturer: Lester Mackey Scribe: Jordan Bryan, Dangna Li 3.1 Recap: Gaussian Mixture Modeling In the last lecture, we discussed the Gaussian
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008
 Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
Lecture 8 Learning Sequence Motif Models Using Expectation Maximization (EM) Colin Dewey February 14, 2008 1 Sequence Motifs what is a sequence motif? a sequence pattern of biological significance typically
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project
 Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Performance Comparison of K-Means and Expectation Maximization with Gaussian Mixture Models for Clustering EE6540 Final Project Devin Cornell & Sushruth Sastry May 2015 1 Abstract In this article, we explore
Mixture Models and Expectation-Maximization
 Mixture Models and Expectation-Maximiation David M. Blei March 9, 2012 EM for mixtures of multinomials The graphical model for a mixture of multinomials π d x dn N D θ k K How should we fit the parameters?
Mixture Models and Expectation-Maximiation David M. Blei March 9, 2012 EM for mixtures of multinomials The graphical model for a mixture of multinomials π d x dn N D θ k K How should we fit the parameters?
Latent Variable Models
 Latent Variable Models Stefano Ermon, Aditya Grover Stanford University Lecture 5 Stefano Ermon, Aditya Grover (AI Lab) Deep Generative Models Lecture 5 1 / 31 Recap of last lecture 1 Autoregressive models:
Latent Variable Models Stefano Ermon, Aditya Grover Stanford University Lecture 5 Stefano Ermon, Aditya Grover (AI Lab) Deep Generative Models Lecture 5 1 / 31 Recap of last lecture 1 Autoregressive models:
Convergence Rate of Expectation-Maximization
 Convergence Rate of Expectation-Maximiation Raunak Kumar University of British Columbia Mark Schmidt University of British Columbia Abstract raunakkumar17@outlookcom schmidtm@csubcca Expectation-maximiation
Convergence Rate of Expectation-Maximiation Raunak Kumar University of British Columbia Mark Schmidt University of British Columbia Abstract raunakkumar17@outlookcom schmidtm@csubcca Expectation-maximiation
ECE521 Tutorial 11. Topic Review. ECE521 Winter Credits to Alireza Makhzani, Alex Schwing, Rich Zemel and TAs for slides. ECE521 Tutorial 11 / 4
 ECE52 Tutorial Topic Review ECE52 Winter 206 Credits to Alireza Makhzani, Alex Schwing, Rich Zemel and TAs for slides ECE52 Tutorial ECE52 Winter 206 Credits to Alireza / 4 Outline K-means, PCA 2 Bayesian
ECE52 Tutorial Topic Review ECE52 Winter 206 Credits to Alireza Makhzani, Alex Schwing, Rich Zemel and TAs for slides ECE52 Tutorial ECE52 Winter 206 Credits to Alireza / 4 Outline K-means, PCA 2 Bayesian
CSC321 Lecture 18: Learning Probabilistic Models
 CSC321 Lecture 18: Learning Probabilistic Models Roger Grosse Roger Grosse CSC321 Lecture 18: Learning Probabilistic Models 1 / 25 Overview So far in this course: mainly supervised learning Language modeling
CSC321 Lecture 18: Learning Probabilistic Models Roger Grosse Roger Grosse CSC321 Lecture 18: Learning Probabilistic Models 1 / 25 Overview So far in this course: mainly supervised learning Language modeling
Expectation Maximization Mixture Models HMMs
 11-755 Machine Learning for Signal rocessing Expectation Maximization Mixture Models HMMs Class 9. 21 Sep 2010 1 Learning Distributions for Data roblem: Given a collection of examples from some data, estimate
11-755 Machine Learning for Signal rocessing Expectation Maximization Mixture Models HMMs Class 9. 21 Sep 2010 1 Learning Distributions for Data roblem: Given a collection of examples from some data, estimate
Mixture Models & EM. Nicholas Ruozzi University of Texas at Dallas. based on the slides of Vibhav Gogate
 Mixture Models & EM icholas Ruozzi University of Texas at Dallas based on the slides of Vibhav Gogate Previously We looed at -means and hierarchical clustering as mechanisms for unsupervised learning -means
Mixture Models & EM icholas Ruozzi University of Texas at Dallas based on the slides of Vibhav Gogate Previously We looed at -means and hierarchical clustering as mechanisms for unsupervised learning -means
Expectation Maximization and Mixtures of Gaussians
 Statistical Machine Learning Notes 10 Expectation Maximiation and Mixtures of Gaussians Instructor: Justin Domke Contents 1 Introduction 1 2 Preliminary: Jensen s Inequality 2 3 Expectation Maximiation
Statistical Machine Learning Notes 10 Expectation Maximiation and Mixtures of Gaussians Instructor: Justin Domke Contents 1 Introduction 1 2 Preliminary: Jensen s Inequality 2 3 Expectation Maximiation
PACKAGE LMest FOR LATENT MARKOV ANALYSIS
 PACKAGE LMest FOR LATENT MARKOV ANALYSIS OF LONGITUDINAL CATEGORICAL DATA Francesco Bartolucci 1, Silvia Pandofi 1, and Fulvia Pennoni 2 1 Department of Economics, University of Perugia (e-mail: francesco.bartolucci@unipg.it,
PACKAGE LMest FOR LATENT MARKOV ANALYSIS OF LONGITUDINAL CATEGORICAL DATA Francesco Bartolucci 1, Silvia Pandofi 1, and Fulvia Pennoni 2 1 Department of Economics, University of Perugia (e-mail: francesco.bartolucci@unipg.it,
Nonparametric Bayesian Methods (Gaussian Processes)
 [70240413 Statistical Machine Learning, Spring, 2015] Nonparametric Bayesian Methods (Gaussian Processes) Jun Zhu dcszj@mail.tsinghua.edu.cn http://bigml.cs.tsinghua.edu.cn/~jun State Key Lab of Intelligent
[70240413 Statistical Machine Learning, Spring, 2015] Nonparametric Bayesian Methods (Gaussian Processes) Jun Zhu dcszj@mail.tsinghua.edu.cn http://bigml.cs.tsinghua.edu.cn/~jun State Key Lab of Intelligent
But if z is conditioned on, we need to model it:
 Partially Unobserved Variables Lecture 8: Unsupervised Learning & EM Algorithm Sam Roweis October 28, 2003 Certain variables q in our models may be unobserved, either at training time or at test time or
Partially Unobserved Variables Lecture 8: Unsupervised Learning & EM Algorithm Sam Roweis October 28, 2003 Certain variables q in our models may be unobserved, either at training time or at test time or
Mixture Models & EM. Nicholas Ruozzi University of Texas at Dallas. based on the slides of Vibhav Gogate
 Mixture Models & EM icholas Ruozzi University of Texas at Dallas based on the slides of Vibhav Gogate Previously We looed at -means and hierarchical clustering as mechanisms for unsupervised learning -means
Mixture Models & EM icholas Ruozzi University of Texas at Dallas based on the slides of Vibhav Gogate Previously We looed at -means and hierarchical clustering as mechanisms for unsupervised learning -means
The Variational Gaussian Approximation Revisited
 The Variational Gaussian Approximation Revisited Manfred Opper Cédric Archambeau March 16, 2009 Abstract The variational approximation of posterior distributions by multivariate Gaussians has been much
The Variational Gaussian Approximation Revisited Manfred Opper Cédric Archambeau March 16, 2009 Abstract The variational approximation of posterior distributions by multivariate Gaussians has been much
CS221 / Autumn 2017 / Liang & Ermon. Lecture 15: Bayesian networks III
 CS221 / Autumn 2017 / Liang & Ermon Lecture 15: Bayesian networks III cs221.stanford.edu/q Question Which is computationally more expensive for Bayesian networks? probabilistic inference given the parameters
CS221 / Autumn 2017 / Liang & Ermon Lecture 15: Bayesian networks III cs221.stanford.edu/q Question Which is computationally more expensive for Bayesian networks? probabilistic inference given the parameters
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm
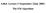 6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
6.864: Lecture 5 (September 22nd, 2005) The EM Algorithm Overview The EM algorithm in general form The EM algorithm for hidden markov models (brute force) The EM algorithm for hidden markov models (dynamic
Inferring Transcriptional Regulatory Networks from Gene Expression Data II
 Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Inferring Transcriptional Regulatory Networks from Gene Expression Data II Lectures 9 Oct 26, 2011 CSE 527 Computational Biology, Fall 2011 Instructor: Su-In Lee TA: Christopher Miles Monday & Wednesday
Estimating the parameters of hidden binomial trials by the EM algorithm
 Hacettepe Journal of Mathematics and Statistics Volume 43 (5) (2014), 885 890 Estimating the parameters of hidden binomial trials by the EM algorithm Degang Zhu Received 02 : 09 : 2013 : Accepted 02 :
Hacettepe Journal of Mathematics and Statistics Volume 43 (5) (2014), 885 890 Estimating the parameters of hidden binomial trials by the EM algorithm Degang Zhu Received 02 : 09 : 2013 : Accepted 02 :
Multiscale Systems Engineering Research Group
 Hidden Markov Model Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of echnology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Learning Objectives o familiarize the hidden
Hidden Markov Model Prof. Yan Wang Woodruff School of Mechanical Engineering Georgia Institute of echnology Atlanta, GA 30332, U.S.A. yan.wang@me.gatech.edu Learning Objectives o familiarize the hidden
Mixtures of Gaussians. Sargur Srihari
 Mixtures of Gaussians Sargur srihari@cedar.buffalo.edu 1 9. Mixture Models and EM 0. Mixture Models Overview 1. K-Means Clustering 2. Mixtures of Gaussians 3. An Alternative View of EM 4. The EM Algorithm
Mixtures of Gaussians Sargur srihari@cedar.buffalo.edu 1 9. Mixture Models and EM 0. Mixture Models Overview 1. K-Means Clustering 2. Mixtures of Gaussians 3. An Alternative View of EM 4. The EM Algorithm
Clustering K-means. Machine Learning CSE546. Sham Kakade University of Washington. November 15, Review: PCA Start: unsupervised learning
 Clustering K-means Machine Learning CSE546 Sham Kakade University of Washington November 15, 2016 1 Announcements: Project Milestones due date passed. HW3 due on Monday It ll be collaborative HW2 grades
Clustering K-means Machine Learning CSE546 Sham Kakade University of Washington November 15, 2016 1 Announcements: Project Milestones due date passed. HW3 due on Monday It ll be collaborative HW2 grades
Technical Details about the Expectation Maximization (EM) Algorithm
 Technical Details about the Expectation Maximization (EM Algorithm Dawen Liang Columbia University dliang@ee.columbia.edu February 25, 2015 1 Introduction Maximum Lielihood Estimation (MLE is widely used
Technical Details about the Expectation Maximization (EM Algorithm Dawen Liang Columbia University dliang@ee.columbia.edu February 25, 2015 1 Introduction Maximum Lielihood Estimation (MLE is widely used
Unsupervised Learning
 2018 EE448, Big Data Mining, Lecture 7 Unsupervised Learning Weinan Zhang Shanghai Jiao Tong University http://wnzhang.net http://wnzhang.net/teaching/ee448/index.html ML Problem Setting First build and
2018 EE448, Big Data Mining, Lecture 7 Unsupervised Learning Weinan Zhang Shanghai Jiao Tong University http://wnzhang.net http://wnzhang.net/teaching/ee448/index.html ML Problem Setting First build and
Bayesian Methods for Machine Learning
 Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Bayesian Methods for Machine Learning CS 584: Big Data Analytics Material adapted from Radford Neal s tutorial (http://ftp.cs.utoronto.ca/pub/radford/bayes-tut.pdf), Zoubin Ghahramni (http://hunch.net/~coms-4771/zoubin_ghahramani_bayesian_learning.pdf),
Brief Introduction of Machine Learning Techniques for Content Analysis
 1 Brief Introduction of Machine Learning Techniques for Content Analysis Wei-Ta Chu 2008/11/20 Outline 2 Overview Gaussian Mixture Model (GMM) Hidden Markov Model (HMM) Support Vector Machine (SVM) Overview
1 Brief Introduction of Machine Learning Techniques for Content Analysis Wei-Ta Chu 2008/11/20 Outline 2 Overview Gaussian Mixture Model (GMM) Hidden Markov Model (HMM) Support Vector Machine (SVM) Overview
CSC2535: Computation in Neural Networks Lecture 7: Variational Bayesian Learning & Model Selection
 CSC2535: Computation in Neural Networks Lecture 7: Variational Bayesian Learning & Model Selection (non-examinable material) Matthew J. Beal February 27, 2004 www.variational-bayes.org Bayesian Model Selection
CSC2535: Computation in Neural Networks Lecture 7: Variational Bayesian Learning & Model Selection (non-examinable material) Matthew J. Beal February 27, 2004 www.variational-bayes.org Bayesian Model Selection
EM-algorithm for motif discovery
 EM-algorithm for motif discovery Xiaohui Xie University of California, Irvine EM-algorithm for motif discovery p.1/19 Position weight matrix Position weight matrix representation of a motif with width
EM-algorithm for motif discovery Xiaohui Xie University of California, Irvine EM-algorithm for motif discovery p.1/19 Position weight matrix Position weight matrix representation of a motif with width
Clustering K-means. Clustering images. Machine Learning CSE546 Carlos Guestrin University of Washington. November 4, 2014.
 Clustering K-means Machine Learning CSE546 Carlos Guestrin University of Washington November 4, 2014 1 Clustering images Set of Images [Goldberger et al.] 2 1 K-means Randomly initialize k centers µ (0)
Clustering K-means Machine Learning CSE546 Carlos Guestrin University of Washington November 4, 2014 1 Clustering images Set of Images [Goldberger et al.] 2 1 K-means Randomly initialize k centers µ (0)
Latent Variable models for GWAs
 Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Latent Variable models for GWAs Oliver Stegle Machine Learning and Computational Biology Research Group Max-Planck-Institutes Tübingen, Germany September 2011 O. Stegle Latent variable models for GWAs
Biostat 2065 Analysis of Incomplete Data
 Biostat 2065 Analysis of Incomplete Data Gong Tang Dept of Biostatistics University of Pittsburgh October 20, 2005 1. Large-sample inference based on ML Let θ is the MLE, then the large-sample theory implies
Biostat 2065 Analysis of Incomplete Data Gong Tang Dept of Biostatistics University of Pittsburgh October 20, 2005 1. Large-sample inference based on ML Let θ is the MLE, then the large-sample theory implies
Homework 2 Solutions Kernel SVM and Perceptron
 Homework 2 Solutions Kernel SVM and Perceptron CMU 1-71: Machine Learning (Fall 21) https://piazza.com/cmu/fall21/17115781/home OUT: Sept 25, 21 DUE: Oct 8, 11:59 PM Problem 1: SVM decision boundaries
Homework 2 Solutions Kernel SVM and Perceptron CMU 1-71: Machine Learning (Fall 21) https://piazza.com/cmu/fall21/17115781/home OUT: Sept 25, 21 DUE: Oct 8, 11:59 PM Problem 1: SVM decision boundaries
p L yi z n m x N n xi
 y i z n x n N x i Overview Directed and undirected graphs Conditional independence Exact inference Latent variables and EM Variational inference Books statistical perspective Graphical Models, S. Lauritzen
y i z n x n N x i Overview Directed and undirected graphs Conditional independence Exact inference Latent variables and EM Variational inference Books statistical perspective Graphical Models, S. Lauritzen
Linear Models for Regression CS534
 Linear Models for Regression CS534 Example Regression Problems Predict housing price based on House size, lot size, Location, # of rooms Predict stock price based on Price history of the past month Predict
Linear Models for Regression CS534 Example Regression Problems Predict housing price based on House size, lot size, Location, # of rooms Predict stock price based on Price history of the past month Predict
Lecture 3: Latent Variables Models and Learning with the EM Algorithm. Sam Roweis. Tuesday July25, 2006 Machine Learning Summer School, Taiwan
 Lecture 3: Latent Variables Models and Learning with the EM Algorithm Sam Roweis Tuesday July25, 2006 Machine Learning Summer School, Taiwan Latent Variable Models What to do when a variable z is always
Lecture 3: Latent Variables Models and Learning with the EM Algorithm Sam Roweis Tuesday July25, 2006 Machine Learning Summer School, Taiwan Latent Variable Models What to do when a variable z is always
Unsupervised Learning with Permuted Data
 Unsupervised Learning with Permuted Data Sergey Kirshner skirshne@ics.uci.edu Sridevi Parise sparise@ics.uci.edu Padhraic Smyth smyth@ics.uci.edu School of Information and Computer Science, University
Unsupervised Learning with Permuted Data Sergey Kirshner skirshne@ics.uci.edu Sridevi Parise sparise@ics.uci.edu Padhraic Smyth smyth@ics.uci.edu School of Information and Computer Science, University
Learning in Bayesian Networks
 Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Learning in Bayesian Networks Florian Markowetz Max-Planck-Institute for Molecular Genetics Computational Molecular Biology Berlin Berlin: 20.06.2002 1 Overview 1. Bayesian Networks Stochastic Networks
Hidden Markov Models. Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from:
 Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Hidden Markov Models Ivan Gesteira Costa Filho IZKF Research Group Bioinformatics RWTH Aachen Adapted from: www.ioalgorithms.info Outline CG-islands The Fair Bet Casino Hidden Markov Model Decoding Algorithm
Deriving the EM-Based Update Rules in VARSAT. University of Toronto Technical Report CSRG-580
 Deriving the EM-Based Update Rules in VARSAT University of Toronto Technical Report CSRG-580 Eric I. Hsu Department of Computer Science University of Toronto eihsu@cs.toronto.edu Abstract Here we show
Deriving the EM-Based Update Rules in VARSAT University of Toronto Technical Report CSRG-580 Eric I. Hsu Department of Computer Science University of Toronto eihsu@cs.toronto.edu Abstract Here we show
Lecture: Mixture Models for Microbiome data
 Lecture: Mixture Models for Microbiome data Lecture 3: Mixture Models for Microbiome data Outline: - - Sequencing thought experiment Mixture Models (tangent) - (esp. Negative Binomial) - Differential abundance
Lecture: Mixture Models for Microbiome data Lecture 3: Mixture Models for Microbiome data Outline: - - Sequencing thought experiment Mixture Models (tangent) - (esp. Negative Binomial) - Differential abundance
Language as a Stochastic Process
 CS769 Spring 2010 Advanced Natural Language Processing Language as a Stochastic Process Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu 1 Basic Statistics for NLP Pick an arbitrary letter x at random from any
CS769 Spring 2010 Advanced Natural Language Processing Language as a Stochastic Process Lecturer: Xiaojin Zhu jerryzhu@cs.wisc.edu 1 Basic Statistics for NLP Pick an arbitrary letter x at random from any
MIXTURE OF EXPERTS ARCHITECTURES FOR NEURAL NETWORKS AS A SPECIAL CASE OF CONDITIONAL EXPECTATION FORMULA
 MIXTURE OF EXPERTS ARCHITECTURES FOR NEURAL NETWORKS AS A SPECIAL CASE OF CONDITIONAL EXPECTATION FORMULA Jiří Grim Department of Pattern Recognition Institute of Information Theory and Automation Academy
MIXTURE OF EXPERTS ARCHITECTURES FOR NEURAL NETWORKS AS A SPECIAL CASE OF CONDITIONAL EXPECTATION FORMULA Jiří Grim Department of Pattern Recognition Institute of Information Theory and Automation Academy
Introduction: MLE, MAP, Bayesian reasoning (28/8/13)
 STA561: Probabilistic machine learning Introduction: MLE, MAP, Bayesian reasoning (28/8/13) Lecturer: Barbara Engelhardt Scribes: K. Ulrich, J. Subramanian, N. Raval, J. O Hollaren 1 Classifiers In this
STA561: Probabilistic machine learning Introduction: MLE, MAP, Bayesian reasoning (28/8/13) Lecturer: Barbara Engelhardt Scribes: K. Ulrich, J. Subramanian, N. Raval, J. O Hollaren 1 Classifiers In this
STA 4273H: Statistical Machine Learning
 STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
STA 4273H: Statistical Machine Learning Russ Salakhutdinov Department of Statistics! rsalakhu@utstat.toronto.edu! http://www.utstat.utoronto.ca/~rsalakhu/ Sidney Smith Hall, Room 6002 Lecture 7 Approximate
University of Cambridge. MPhil in Computer Speech Text & Internet Technology. Module: Speech Processing II. Lecture 2: Hidden Markov Models I
 University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
University of Cambridge MPhil in Computer Speech Text & Internet Technology Module: Speech Processing II Lecture 2: Hidden Markov Models I o o o o o 1 2 3 4 T 1 b 2 () a 12 2 a 3 a 4 5 34 a 23 b () b ()
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation. Volker Tresp Summer 2016
 Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2016 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Bayesian Networks: Construction, Inference, Learning and Causal Interpretation Volker Tresp Summer 2016 1 Introduction So far we were mostly concerned with supervised learning: we predicted one or several
Programming Assignment 4: Image Completion using Mixture of Bernoullis
 Programming Assignment 4: Image Completion using Mixture of Bernoullis Deadline: Tuesday, April 4, at 11:59pm TA: Renie Liao (csc321ta@cs.toronto.edu) Submission: You must submit two files through MarkUs
Programming Assignment 4: Image Completion using Mixture of Bernoullis Deadline: Tuesday, April 4, at 11:59pm TA: Renie Liao (csc321ta@cs.toronto.edu) Submission: You must submit two files through MarkUs
Predicting the Treatment Status
 Predicting the Treatment Status Nikolay Doudchenko 1 Introduction Many studies in social sciences deal with treatment effect models. 1 Usually there is a treatment variable which determines whether a particular
Predicting the Treatment Status Nikolay Doudchenko 1 Introduction Many studies in social sciences deal with treatment effect models. 1 Usually there is a treatment variable which determines whether a particular
The knee-jerk mapping
 arxiv:math/0606068v [math.pr] 2 Jun 2006 The knee-jerk mapping Peter G. Doyle Jim Reeds Version dated 5 October 998 GNU FDL Abstract We claim to give the definitive theory of what we call the kneejerk
arxiv:math/0606068v [math.pr] 2 Jun 2006 The knee-jerk mapping Peter G. Doyle Jim Reeds Version dated 5 October 998 GNU FDL Abstract We claim to give the definitive theory of what we call the kneejerk
10-701/15-781, Machine Learning: Homework 4
 10-701/15-781, Machine Learning: Homewor 4 Aarti Singh Carnegie Mellon University ˆ The assignment is due at 10:30 am beginning of class on Mon, Nov 15, 2010. ˆ Separate you answers into five parts, one
10-701/15-781, Machine Learning: Homewor 4 Aarti Singh Carnegie Mellon University ˆ The assignment is due at 10:30 am beginning of class on Mon, Nov 15, 2010. ˆ Separate you answers into five parts, one
Introduction to Machine Learning. Maximum Likelihood and Bayesian Inference. Lecturers: Eran Halperin, Yishay Mansour, Lior Wolf
 1 Introduction to Machine Learning Maximum Likelihood and Bayesian Inference Lecturers: Eran Halperin, Yishay Mansour, Lior Wolf 2013-14 We know that X ~ B(n,p), but we do not know p. We get a random sample
1 Introduction to Machine Learning Maximum Likelihood and Bayesian Inference Lecturers: Eran Halperin, Yishay Mansour, Lior Wolf 2013-14 We know that X ~ B(n,p), but we do not know p. We get a random sample
Chapter 08: Direct Maximum Likelihood/MAP Estimation and Incomplete Data Problems
 LEARNING AND INFERENCE IN GRAPHICAL MODELS Chapter 08: Direct Maximum Likelihood/MAP Estimation and Incomplete Data Problems Dr. Martin Lauer University of Freiburg Machine Learning Lab Karlsruhe Institute
LEARNING AND INFERENCE IN GRAPHICAL MODELS Chapter 08: Direct Maximum Likelihood/MAP Estimation and Incomplete Data Problems Dr. Martin Lauer University of Freiburg Machine Learning Lab Karlsruhe Institute
STA 414/2104: Machine Learning
 STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
STA 414/2104: Machine Learning Russ Salakhutdinov Department of Computer Science! Department of Statistics! rsalakhu@cs.toronto.edu! http://www.cs.toronto.edu/~rsalakhu/ Lecture 9 Sequential Data So far
EM Algorithm II. September 11, 2018
 EM Algorithm II September 11, 2018 Review EM 1/27 (Y obs, Y mis ) f (y obs, y mis θ), we observe Y obs but not Y mis Complete-data log likelihood: l C (θ Y obs, Y mis ) = log { f (Y obs, Y mis θ) Observed-data
EM Algorithm II September 11, 2018 Review EM 1/27 (Y obs, Y mis ) f (y obs, y mis θ), we observe Y obs but not Y mis Complete-data log likelihood: l C (θ Y obs, Y mis ) = log { f (Y obs, Y mis θ) Observed-data
Bayesian Models in Machine Learning
 Bayesian Models in Machine Learning Lukáš Burget Escuela de Ciencias Informáticas 2017 Buenos Aires, July 24-29 2017 Frequentist vs. Bayesian Frequentist point of view: Probability is the frequency of
Bayesian Models in Machine Learning Lukáš Burget Escuela de Ciencias Informáticas 2017 Buenos Aires, July 24-29 2017 Frequentist vs. Bayesian Frequentist point of view: Probability is the frequency of
Discovering molecular pathways from protein interaction and ge
 Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
Discovering molecular pathways from protein interaction and gene expression data 9-4-2008 Aim To have a mechanism for inferring pathways from gene expression and protein interaction data. Motivation Why
PARAMETER CONVERGENCE FOR EM AND MM ALGORITHMS
 Statistica Sinica 15(2005), 831-840 PARAMETER CONVERGENCE FOR EM AND MM ALGORITHMS Florin Vaida University of California at San Diego Abstract: It is well known that the likelihood sequence of the EM algorithm
Statistica Sinica 15(2005), 831-840 PARAMETER CONVERGENCE FOR EM AND MM ALGORITHMS Florin Vaida University of California at San Diego Abstract: It is well known that the likelihood sequence of the EM algorithm
Gaussian processes. Chuong B. Do (updated by Honglak Lee) November 22, 2008
 Gaussian processes Chuong B Do (updated by Honglak Lee) November 22, 2008 Many of the classical machine learning algorithms that we talked about during the first half of this course fit the following pattern:
Gaussian processes Chuong B Do (updated by Honglak Lee) November 22, 2008 Many of the classical machine learning algorithms that we talked about during the first half of this course fit the following pattern:
Machine Learning. Gaussian Mixture Models. Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall
 Machine Learning Gaussian Mixture Models Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall 2012 1 The Generative Model POV We think of the data as being generated from some process. We assume
Machine Learning Gaussian Mixture Models Zhiyao Duan & Bryan Pardo, Machine Learning: EECS 349 Fall 2012 1 The Generative Model POV We think of the data as being generated from some process. We assume
The connection of dropout and Bayesian statistics
 The connection of dropout and Bayesian statistics Interpretation of dropout as approximate Bayesian modelling of NN http://mlg.eng.cam.ac.uk/yarin/thesis/thesis.pdf Dropout Geoffrey Hinton Google, University
The connection of dropout and Bayesian statistics Interpretation of dropout as approximate Bayesian modelling of NN http://mlg.eng.cam.ac.uk/yarin/thesis/thesis.pdf Dropout Geoffrey Hinton Google, University
