Genetic Flexibility in the Convergent Evolution of Hermaphroditism in Caenorhabditis Nematodes
|
|
|
- Brendan Malone
- 5 years ago
- Views:
Transcription
1 Developmental Cell 10, , April, 2006 ª2006 Elsevier Inc. DOI /j.devcel Genetic Flexibility in the Convergent Evolution of Hermaphroditism in Caenorhabditis Nematodes Short Article Robin Cook Hill, 1 Carlos Egydio de Carvalho, 2,4 John Salogiannis, 1,5 Benjamin Schlager, 1,3,6 Dave Pilgrim, 2 and Eric S. Haag 1, * 1 Department of Biology University of Maryland College Park, Maryland Department of Biological Sciences University of Alberta Edmonton, Alberta T6G 2E9 Canada 3 Program in Molecular Biology Institute of Microbiology and Genetics University of Vienna Vienna A-1030 Austria Summary The self-fertile hermaphrodites of C. elegans and C. briggsae evolved from female ancestors by acquiring limited spermatogenesis. Initiation of C. elegans hermaphrodite spermatogenesis requires germline translational repression of the female-promoting gene tra-2, which allows derepression of the three male-promoting fem genes. Cessation of hermaphrodite spermatogenesis requires fem-3 translational repression. We show that C. briggsae requires neither fem-2 nor fem-3 for hermaphrodite development, and that XO Cb-fem-2/3 animals are transformed into hermaphrodites, not females as in C. elegans. Exhaustive screens for Cb-tra-2 suppressors identified another 75 fem-like mutants, but all are self-fertile hermaphrodites rather than females. Control of hermaphrodite spermatogenesis therefore acts downstream of the fem genes in C. briggsae. The outwardly similar hermaphrodites of C. elegans and C. briggsae thus achieve self-fertility via intervention at different points in the core sex determination pathway. These findings are consistent with convergent evolution of hermaphroditism, which is marked by considerable developmental genetic flexibility. Introduction In the nematode family Rhabditidae, which includes the model species Caenorhabditis elegans, self-fertile hermaphrodites have evolved from XX female ancestors several times (Fitch, 2002; Kiontke et al., 2004). In each case, the XX female sex gained a brief period of spermatogenesis that occurs before typical oogenesis. *Correspondence: ehaag@umd.edu 4 Present address: Harvard Medical School, New Research Building, Room 334, 77 Avenue Louis Pasteur, Boston, Massachusetts Present address: Department of Cell Biology, Harvard Medical School, 300 Longwood Avenue, Boston, Massachusetts Present address: Max-Planck-Institut für Entwicklungsbiologie, Spemannstr , Abteilung 4 Evolutionsbiologie, Tübingen, Germany. Recent phylogenetic studies have suggested the surprising possibility that the hermaphroditism of even the closely related C. elegans and C. briggsae may be convergent (Cho et al., 2004; Kiontke et al., 2004). Since spermatogenesis is the only anatomical difference between females and hermaphrodites, germline sex determination genes must have been the key targets of selection for self-fertility. As C. elegans hermaphrodite development is well understood, this offers an excellent opportunity to study the developmental genetics of parallel evolution. In this paper, we present what is, to our knowledge, the first genetic analysis of C. briggsae sex determination. In C. elegans, FEM-1, FEM-2, and FEM-3 form part of the core signal transduction pathway that mediates sex determination (Figure 1A) and act in the cytoplasm between the transmembrane receptor TRA-2 and the Gli-related transcription factor TRA-1 to promote male fate in XO animals (Doniach and Hodgkin, 1984; Hodgkin, 1986; Kimble et al., 1984). That all three fem genes are also required for hermaphrodite spermatogenesis suggests that they may have been instrumental for the evolution of self-fertility in the C. elegans lineage. FEM-1 is composed of ankyrin repeats (Spence et al., 1990), FEM-2 is a PP2C phosphatase (Chin-Sang and Spence, 1996; Pilgrim et al., 1995), and FEM-3 is a nematode-specific protein that forms a specific complex with FEM-2 (Ahringer et al., 1992; Chin-Sang and Spence, 1996). Nearly 20 years of research has shown that posttranscriptional controls regulate germline sexual fates in C. elegans hermaphrodites (Puoti et al., 1997). The translational repression of tra-2 by fog-2 and gld-1 is required to initiate fem-dependent hermaphrodite spermatogenesis (Clifford et al., 2000; Doniach, 1986; Goodwin et al., 1993; Schedl and Kimble, 1988), and similar repression of fem-3 by fbf-1/2 is required for its cessation (Ahringer and Kimble, 1991; Barton et al., 1987; Zhang et al., 1997). Nearly all C. elegans sex determination genes have orthologs in C. briggsae (Haag, 2005; Nayak et al., 2005), and Cb-tra-2 is translationally repressed in a manner consistent with it also being key for the initiation of hermaphrodite spermatogenesis (Jan et al., 1997). However, fog-2 is specific to C. elegans, and RNAi knockdown of Cb-gld-1 gives an unexpected Mog (masculinization of germline) phenotype (Clifford et al., 2000; Nayak et al., 2005). It is thus unclear at which point C. briggsae hermaphrodites regulate the core pathway to initiate XX spermatogenesis. In addition, Cb-fem- 2(RNAi) (Stothard et al., 2002) and Cb-fem-3(RNAi) (Haag et al., 2002) have no effect on hermaphrodites, but they partially feminize XO animals. In the case of Cb-fem-2, the XO germline is feminized, but the male soma was left intact (Stothard et al., 2002). For Cbfem-3, the soma is feminized, but the germline remained exclusively male (Haag et al., 2002). These results are intriguing (Stothard and Pilgrim, 2003), but the poor penetrance of RNAi in C. briggsae (Haag et al., 2002) limited their interpretation. We sought to rigorously test if spermatogenesis in C. briggsae hermaphrodites depends
2 Developmental Cell 532 Figure 1. Overview of fem Function, PP2C Evolution, and Isolation of Cb-fem-2 and Cb-fem-3 Deletions (A) The core sex determination pathway of C. elegans, which acts in both soma and germline. (B) Phylogenetic analysis of C. elegans and C. briggsae PP2C proteins. Aligned phosphatase domains from all predicted family members of each species were analyzed by the neighbor-joining method. Numbers indicate bootstrap support >50%. Parsimony analyses recovered the same orthologous pairs, but with different internal nodes. (C) Fully nested flanking primers were used to detect gene-specific deletions in pools of genomic DNA from mutagenized nematodes. Primer pairs ( WT only ) annealing within the deleted regions were subsequently designed to positively identify the wild-type chromosome. (D) Deletions were verified by single-worm PCR with flanking and WT-only primers. Each lane represents one of two reactions performed on random single worms of the indicated genotype, with replicates included to demonstrate homozygosity and assay reproducibility. In AF16 (wild-type) animals, the WT-only products are produced, and the flanking primers amplify the full-length product. In all Cb-fem-2(nm27) or Cb-fem-3(nm63) homozygotes, no WT-only product is produced, and flanking primers only generate the deletion-specific products. upon fem activity, and to clarify its roles in males, through analysis of strong loss-of-function mutations. Results and Discussion Isolation of Cb-fem-2 and Cb-fem-3 Deletions We screened for deletion mutations in C. briggsae fem genes with the same method (Figure 1C) used by the C. elegans Knockout Consortium (Edgley et al., 2002). The first screen produced the allele Cb-fem-2(nm27), a 1.6 kbp deletion that removes the entire phosphatase domain as well as part of the 3 0 UTR. As phosphatase activity is necessary for the sex determination function of C. elegans FEM-2 (Chin-Sang and Spence, 1996), nm27 is likely a null mutation. We used both the polymerase chain reaction (PCR) with primers inside and outside of the deleted region (Figure 1D) and Southern hybridization (A. Doty and E.S.H., unpublished data) to verify that a strain homozygous for Cb-fem-2(nm27) was generated. In addition, no Cb-fem-2 mrna expression is detected in mutant animals by high-sensitivity in situ hybridization (Figure 3D). This strain, CP36, is robustly self-fertile and morphologically indistinguishable from wild-type (Figure 2), demonstrating that Cb-FEM-2 is not required for hermaphrodite spermatogenesis. Another screen produced Cb-fem-3(nm63), a 1.1 kbp deletion (Figure 1C). This mutation removes DNA coding for 155 of the 409 total amino acids of Cb-FEM-3 (residues ). The deleted region codes for amino acids whose equivalents are essential for C. elegans FEM-3 function (Ahringer et al., 1992), as well as several short stretches of conservation between species (Haag et al., 2002). Homozygous Cb-fem-3(nm63) XX mutants are also self-fertile hermaphrodites (Figure 2F). It is formally possible that the nm63 mutation is not a null, as it maintains the correct reading frame of the remaining codons if splicing were unaffected. However, we show that nm63 has a strong loss-of-function phenotype in XO animals, indicating that Cb-fem-3, like Cb-fem-2, is not required for hermaphrodite spermatogenesis. The conservation of the FEM-2/FEM-3 interaction in C. briggsae (Stothard and Pilgrim, 2006) left the possibility that only one of them is necessary for hermaphrodite development. However, XX Cb-fem-2(nm27); Cb-fem- 3(nm63) worms are also self-fertile (Figure 2H and Table S1; see the Supplemental Data available with this article online). XO Cb-fem-2 and Cb-fem-3 Mutants Are Hermaphrodites The 11-member PP2C family has been stable since the divergence of C. elegans and C. briggsae (Figure 1B). An earlier study of a subset of the family (Stothard et al., 2002) indicated that fem-2 evolves faster than other family members. The current, complete analysis generally supports this, but it reveals that a second gene (CE07231/CBP03684) is comparably rapid. The large PP2C family suggested that the lack of Cb-fem- 2(nm27) XX feminization could be because Cb-FEM-2 is redundant with another phosphatase, and not
3 fem-independent Hermaphroditism in C. briggsae 533 Figure 2. Phenotypes of Cb-fem-2 and Cb-fem-3 Mutants (A) Wild-type XX adult hermaphrodites have two oocyte-producing gonad arms (the anterior visible here); two spermathecae with sperm flank the embryo-containing uterus and medial vulva. The gut is seen in the posterior, under which lies the posterior gonad arm. The tail is tapered. The scale bar = 100 mm. (B) XO wild-type males make sperm and have a modified tail used in copulation. (C) Cb-fem-2(nm27) XX animals are self-fertile hermaphrodites. The inset shows the uterus and posterior spermatheca of a virgin adult, with sperm (see also [I] [L]) and two embryos. (D) Cb-fem-2(nm27) XO animals are also self-fertile hermaphrodites with complete somatic feminization, including the tail (inset). Most are similar to wild-type hermaphrodites and have a completely female soma and two self-fertile gonad arms that produce viable embryos. (E) Abnormal XO hermaphrodites were seen at low frequency. This individual (shown slightly magnified relative to [A] [D]) has two spermathecae on the posterior gonad arm (asterisks) and none in the anterior. Sperm can be seen in a broad area of the anterior arm. (F) Cb-fem-3(nm63) XX mutants are also self-fertile hermaphrodites; an inset at higher magnification reveals the presence of oocytes, sperm, and an embryo of one gonad arm. (G) Cb-fem-3(nm63) XO mutants are transformed into hermaphrodites with completely feminized soma (the inset shows the tail at higher magnification). (H) XX Cb-fem-3(nm27); Cb-fem-3(nm63) double mutants (slightly magnified relative to [F] and [G]) are also self-fertile. (I L) fem-2(nm27) XX hermaphrodites produce normal sperm. (I) Proximal gonad of a virgin animal; the compact nuclei of sperm (J) are visualized with DAPI staining, and the sperm protein SPE-56 (K) is detected with monoclonal antibody staining. They colocalize ([L], merged image) in the spermatheca (indicated by arrowheads). Anterior is to the left, and ventral is down in all panels. Labels: e, embryo; g, gut; o, oocytes; s, sperm; v, vulva. essential for any aspect of sex determination. Similarly, the in-frame deletion of Cb-fem-3(nm63) left open the possibility that an internally truncated Cb-FEM-3 protein with sufficient activity is produced. To address these concerns, we examined the XO phenotype of Cb-fem- 2(nm27) by using three genetic assays. We then employed the two most definitive of these assays to similarly characterize Cb-fem-3(nm63). First, various crosses were scored for sex ratio changes indicative of somatic feminization (Table S1). Those that cannot produce Cb-fem-2(nm27) homozygotes had the 35% 40% male frequency characteristic of wild-type crosses (the ratio is not 1:1 due to rare selfing and errant males). However, crosses of Cb-fem-2(nm27) hermaphrodites
4 Developmental Cell 534 Figure 3. Expression of fem-2 and fem-3 mrna in the C. briggsae Germ Line (A) Hybridization of an antisense Cb-fem-2 cdna probe to extruded wild-type C. briggsae gonads. (B) Hybridization of an antisense Ce-fem-2 cdna probe with wild-type extruded C. elegans gonads. (C and D) (C) AF16 (wild-type) and (D) Cbfem-2(nm27) mutant germ lines processed with the concentration of proteinase K doubled, which increased sensitivity, but allowed only partially extruded germ lines to survive intact. (E G) (E) Hybridization of a Ce-fem-2 cdna sense control probe to wild-type C. elegans gonads. Partially dissected (F) C. briggsae hermaphrodite and (G) extruded gut and germ line stained with antisense Cb-fem-3 probe, showing strong staining throughout the body, including the gut and a loop of germline, but none at the distal tip of the germ line. (H) Cb-fem-3 sense control probe, which shows background staining only in the pharynx. (A 0 H 0 ) Fluorescence companion images of Hoechst stained DNA from the specimens shown in the panel above each. Proximal gonad ends are to the left in [A], [B], [E], [G], and [H] and are the bottom halves of the looped gonads in [D] and [D]. Labels: d, distal tip of germline; e, embryo; g, gut; o, mature oocyte in diakinesis; p, pharynx. with Cb-fem-2(nm27)/+ males produced roughly half as many male progeny, as expected if sexual transformation had occurred. A second test for XO transformation is to suppress male production in a high incidence of males (Him) strain. We serendipitously discovered that unmated hermaphrodites heterozygous for the X chromosomeintegrated green fluorescent protein (GFP) reporter transgene syis802 have small broods that produce 32.5% XO males, presumably due to transgene destabilization of meiotic pairing. In contrast, Cb-fem-2(nm27); syis802/+ hermaphrodites produce no males (Table S1). Similar Him suppression was seen in the progeny of Cbfem-3(nm63); syis802/+ animals. These results also suggest that male development cannot occur in Cb-fem-2 or Cb-fem-3 mutants, but feminized XO animals could not be positively identified. The most definitive test for putative XO feminization employs genetic markers for outcrossing and karyotype. We scored the progeny of crosses between Cbdpy(nm4) II; Cb-fem-2(nm27) III hermaphrodites and Cb-fem-2(nm27)/+ III; syis802[myo-2::gfp] X males. Half of the non-dpy, non-gfp offspring (XO cross progeny) are Cb-fem-2(nm27) homozygotes that also lack maternal Cb-fem-2 function and are thus potentially feminized, and half are Cb-fem-2(nm27)/+ and are expected to be male. We observed that 59% of non-dpy, non-gfp progeny (n = 90) were somatically feminized (Figure 2D), while the remainder were normal males. Interestingly, these XO Cb-fem-2(nm27) mutants are self-fertile hermaphrodites (Her), and not females as in C. elegans. They produce viable embryos, but they have small broods and minor defects in female somatic gonad development (e.g., Figure 2E). Cb-fem-2/+ XO animals show late-onset germline feminization similar to that seen with Cb-fem-2(RNAi) (Stothard et al., 2002) (data not shown). As detailed in the Supplemental Data, Cb-fem-2(nm27) exhibits neither the temperature sensitivity nor maternal rescue of Ce-fem-2 mutants (Hodgkin, 1986). The above-described experiments indicate that Cbfem-2 is essential for male somatic sex determination, and that it is therefore not generally redundant with other PP2c phosphatases. An identical strategy allowed for the isolation of Cb-fem-3(nm63) XO homozygotes: 59% of XO cross progeny (n = 88) were also self-fertile hermaphrodites (Figure 2G) with small broods and somatic gonad defects similar to those in the Cb-fem-2(nm27)
5 fem-independent Hermaphroditism in C. briggsae 535 Figure 4. Suppressors of Cb-tra-2(ts) Mutations Are Self-Fertile (A) At 15º, XX Cb-tra-2(nm9ts) animals are self-fertile hermaphrodites with a whip-like tail (arrowhead) and numerous embryos (e). (B) At 25º, they are transformed into Tra pseudomales with an imperfect male tail (arrowhead) and abundant sperm (s). (C) All suppressor mutations that restore a female soma to two such Cb-tra-2(ts) alleles are self-fertile. The different recovery rates for suppressors of the ed23ts and nm9ts alleles are due to methodological differences in the two screens (see Experimental Procedures). (D) Comparison of C. elegans and C. briggsae XX germline sex determination. Conserved components of the core sex determination pathway, which act in both males and hermaphrodites, are shown in black. In C. elegans (top), hermaphrodite spermatogenesis is initiated through the repression of tra-2 by the factors shown in green, and it is terminated by repression of fem-3 by the factors shown in red. A tra-1/fog-3-independent role for the Ce-fem genes and a possible spermpromoting role for tra-1 have been omitted for simplicity (Chen and Ellis, 2000; Hodgkin, 1987; Schedl et al., 1989). In C. briggsae (bottom), unknown factors acting downstream of the Cb-fem genes start and stop sperm production. The masculinizing activity that starts spermatogenesis is suggested here to repress Cb-tra-1 and/or stimulate Cb-fog-1/3, while the sperm-oocyte switch would depend upon either stimulating Cb-tra-1 and/or repressing Cb-fog-1/3. C. briggsae orthologs of xol-1 (Luz et al., 2003), tra-3, and the sdc genes (not shown) exist, but they have not been functionally characterized. Evidence for conservation of the remaining genes has recently been reviewed (Haag, 2005). XO mutants. In C. elegans, XO fem mutants are selfsterile because the fem genes act downstream of the tra-2-mediated initiation of spermatogenesis (Goodwin and Ellis, 2002). In contrast, C. briggsae XO fem-2 and fem-3 animals execute the sperm-then-oocytes pattern consistent with their transformed somas. This suggests that Cb-fem-2 and Cb-fem-3 act to repress the switch to oogenesis in males, even though they are not required for sperm production in hermaphrodites. Germline Expression of fem Genes Is Conserved Despite the differences in the germline phenotype of fem-2 and fem-3 mutants between C. elegans and C. briggsae, we find that adult hermaphrodites of the two species both produce germline fem-2 mrna at comparable levels, as judged by in situ hybridization (Figure 3). This expression is first detectable as oocytes exit pachytene and begin gametogenesis, and it is strongest in mature diakinesis oocytes. This is consistent with the maternal rescue of sexual transformation and the minor requirement for Ce-fem-2 in late embryogenesis (Piekny et al., 2000). Maternal Cb-fem-2 expression may have a similar embryonic function, but we have not observed embryogenesis defects like those reported for Ce-fem-2 mutants. Cb-fem-3 mrna is expressed in the germ line in a graded, proximal-biased manner similar to that seen in Cb-fem-2 (Figures 3F and 3G). While Cb-fem-2 staining is strong and consistently cytoplasmic in mature oocytes, Cb-fem-3 staining is often perinuclear. Cb-fem-3 staining is also observed in somatic tissues, including the gut. Germline expression of Ce-fem-3 is largely similar (A. Puoti, personal communication). Thus, as with fem-2, differences in fem-3 function cannot be explained by changes in germline transcription. However, Northern analysis of C. elegans mutants lacking germlines (Rosenquist and Kimble, 1988) indicates that somatic expression of Ce-fem-3 is much lower than for Cb-fem-3. Isolation of Cb-fem Genes as Cb-tra-2(ts) Suppressors Have other C. briggsae genes taken on a fem-like role of promoting spermatogenesis in the XX germline? In C. elegans, the first fem-3 mutations were isolated as genetic suppressors of tra-3 and tra-2 (Hodgkin, 1986). We undertook similar screens for mutants that suppress the somatic masculinization of two different temperature-sensitive alleles of Cb-tra-2 (C.E.d.C., M. Layton, J.S., E.S.H., and D.P., unpublished data; Figure 4). A total of 75 F2 suppressors were isolated from 760,000
6 Developmental Cell 536 haploid genomes screened. At least five failed to complement the Cb-fem-2(nm27) mutation, and the remaining alleles fell into at least two more complementation groups (C.E.d.C. and D.P., unpublished data). While the alleles varied in their ability to suppress the somatic and germline masculinization of the Cb-tra-2 alleles, all Cb-tra-2; sup double mutant strains can be maintained as self-fertile XX hermaphrodites at the restrictive temperature. No Cb-tra-2; sup XX females were isolated, although the large number of genomes screened and the isolation of multiple alleles of the suppressors suggest that the screen was close to saturation. Importantly, XX Cb-tra-2; Cb-fem-2 double mutants are self-fertile hermaphrodites (data not shown). This demonstrates that, despite the lack of an XX phenotype on its own, Cb-fem-2 acts downstream of tra-2 in both the soma and germline in C. briggsae, as in C. elegans. Conclusions and Prospects The hermaphrodites of C. elegans and C. briggsae both produce sperm around the L4-adult molt and then switch to oogenesis. Our results indicate that this outwardly similar phenotype is produced by intervention at different levels of the core sex determination pathway (Figure 4D). In C. elegans, the control of hermaphrodite spermatogenesis targets tra-2 ( sperm on ) and fem-3 ( sperm off ), and, consequently, the only loss-of-function mutations that convert XO males into hermaphrodites (the Her phenotype) affect her-1, which lies immediately upstream of tra-2. In contrast, we find that XO mutants lacking at least two of the Cb-fem genes are Her, and that no readily mutable C. briggsae fem-like gene (as defined by being downstream of Cb-tra-2) is required for normal XX spermatogenesis. The control of hermaphrodite spermatogenesis therefore operates downstream of the fem genes, implicating Cb-tra-1 and/or its targets. One likely Cb-tra-1 target, Cb-fog-3, is expressed during and is required for both male and hermaphrodite spermatogenesis (Chen et al., 2001). Since both its promoter and coding sequence can support the rescue of Ce-fog-3 germline feminization, this control may target Cb-tra-1. While it is formally possible that the last common ancestor of C. elegans and C. briggsae was hermaphroditic, our findings are most consistent with convergent evolution of selfing in Caenorhabditis, and they indicate the existence of flexibility in how it is achieved (True and Haag, 2001). Other studies of convergent evolution have often found the striking reuse of key genes (Colosimo et al., 2004; Cresko et al., 2004; Eizirik et al., 2003; Gompel and Carroll, 2003; Protas et al., 2006; Shapiro et al., 2004; Sucena et al., 2003; Yamamoto et al., 2004). However, a few cases of genetically distinct convergence have also been found (e.g., Hoekstra and Nachman, 2003; Wittkopp et al., 2003). Nematode hermaphroditism represents a new example of the latter, but it is distinguished by its basis in sex determination, its germline specificity, and the posttranscriptional regulation that accompanies them. Why might hermaphroditism evolve via distinct genetic changes? One possibility is that there are many routes to self-fertility; thus, independent use of any particular one is unlikely. Another is that differences in initial conditions greatly influence subsequent events. The closest known relative of C. briggsae is the gonochoristic (male/female) C. remanei (Cho et al., 2004; Kiontke et al., 2004). Germline sex determination in the gonochoristic C. remanei/c. briggsae common ancestor may have already diverged somewhat from that of the C. elegans lineage, such that evolution of hermaphroditism in C. briggsae necessarily required distinct modifications. These scenarios may be distinguished through further forward and reverse genetic characterization of C. briggsae and C. remanei sex determination, a task made much easier by the C. briggsae genomic sequence (Stein and others, 2003). Experimental Procedures Nematode Culture, Strains, and Genetics C. briggsae strains were cultured by using standard C. elegans conditions (Wood, 1988), with the use of 2.2% agar plates to discourage burrowing. All mutants were derived from the wild isolate AF16. The Cb-fem-2 deletion allele nm27 and the Cb-fem-3 deletion allele nm63 were isolated from different libraries of EMS-mutagenized worms (1.1 million haploid genomes each) by using standard C. elegans methods (Edgley et al., 2002) and without the poison primer modification (primer sequences are in Supplemental Data). Cbdpy(nm4), Cb-tra-2(nm9ts), and Cb-tra-2(ed23ts) were isolated in conventional F2 screens for recessive mutants that will be described elsewhere. Evidence that nm9ts and ed23ts are alleles of Cb-tra-2 includes tight linkage to a Cb-tra-2 polymorphism, RNAi phenocopy (Haag et al., 2002; Kuwabara, 1996), missense mutations in the Cbtra-2 open reading frame, and mutual noncomplementation. syis802, an X-integrated array consisting of a wild-type C. elegans daf-4 genomic fragment and the myo-2::gfp reporter construct, was kindly provided by T. Inouye and P. Sternberg (CalTech). All mutant alleles were outcrossed with wild-type AF16 males (and verified by PCR where necessary) at least four times prior to use in building strains. Cb-dpy(nm4)/+ II; Cb-fem-2(nm27)/+ III hermaphrodites were the progeny of a cross between Cb-dpy(nm4); Cb-fem-2(nm27) (strain CP48) and AF16 males. CP48 was produced as follows: AF16 males crossed with Cb-fem-2(nm27) III hermaphrodites produce Cb-fem- 2(nm27)/+ III males, which were then mated with dpy(nm4) II hermaphrodites. Many Dpy F2 were singled and PCR genotyped for Cb-fem-2 after they had laid most of their progeny. Cb-fem- 2(nm27)/+ III; syis802[myo-2::gfp] X males were the GFP+ male progeny of a cross between AF16 males and Cb-fem-2(nm27) III; syis802[myo-2::gfp] X hermaphrodites. These hermaphrodites were, in turn, produced from mating syis802[myo-2::gfp] X males (produced spontaneously by syis802[myo-2::gfp] X/+ hermaphrodites) with Cb-fem-2(nm27) III hermaphrodites and PCR genotyping GFP+ F2 for Cb-fem-2. Similar methods were used to build the analogous Cb-fem-3(nm63) strains. F2 suppressors of Cb-tra-2(nm9ts) and Cb-tra-2(ed23ts) were identified by plating the F1 progeny of EMS-mutagenized L4 hermaphrodite larvae at 15º (400 per 6 cm plate for ed23ts, 150 per 10 cm plate for nm9ts). F2 embryos were shifted to 25º and screened for rare mutants with normal female somas. Although the suppressors analyzed thus far are recessive, the collection may include dominant Cb-tra-1 alleles, similar to those found in C. elegans tra-3(sup) screens (Hodgkin, 1986). The ed23ts screen probably undercounted suppressors for two reasons: at the higher density, the youngest and sickly worms produce few progeny, and only a single F2 suppressor was retained per plate in both screens to ensure independence of mutations. A total of 74 of 75 mutants were verified as second-site mutations via outcrossing with AF16 and recovery of the Tra phenotype in the F2. Single-Worm PCR Single-worm PCR was carried out as described (Haag et al., 2002). For the assay of a single worm with two primer sets, PCR was first performed with the outer flanking primers, after which this outer reaction was diluted 1:250 for use in inner reactions with either the inner flanking or WT-only primers.
7 fem-independent Hermaphroditism in C. briggsae 537 Phylogenetic Analysis All C. elegans PP2C phosphatases, as predicted by PFAM profile searches, were obtained from WormBase ( After splice variants were removed, the set was used to perform BLAST searches on the predicted C. briggsae proteome (CB25 release), and all hits with E values below 0.01 were scrutinized for PP2C domains. This quickly established the largest possible set of C. briggsae family members. PP2c domains were then excised and aligned with Clustal W as implemented in Vector NTI (Invitrogen), and the alignment was validated by recovery of the conserved sequence blocks described by Pilgrim et al. (1995). Neighbor-joining and maximum parsimony analyses were performed with PAUP 4b10 (Swofford, 2002). In Situ Hybridization Single-stranded digoxygenin-labeled DNA probes were produced as described (Patel et al., 1992) from a 1.1 kbp Cb-fem-2 cdna clone covering the phosphatase domain, a complete 2.0 kbp Ce-fem-2 cdna plasmid, pdp#dh20 (D. Hansen and D.P., unpublished data), and the complete Cb-fem-3 coding sequence cdna in pjk769. Dissection of gonads, fixation, hybridization, and detection were generally performed as previously described (Jones et al., 1996), although the concentration of proteinase K was routinely increased to 100 mg/ml. At concentrations of 200 mg/ml, used for the experiments in Figures 3C and 3D, completely extruded gonads were fragmented. Staining of nuclei was with 0.5 mg/ml Hoechst in PBST. Immunostaining of Dissected Gonads Anti-SPE-56 monoclonal antisera were the gift of S. Strome. C. briggsae hermaphrodite gonads were dissected as described (Francis et al., 1995). Fixation was performed in ice-cold 100% methanol for 5 min. After three washes in PBST (0.1% Tween 20 in PBS), carcasses were blocked at 4ºC overnight with 1 mg/ml blocking solution (BSA in PBST). The primary antibody was diluted 1:50 in blocking solution. After 2 hr of incubation at room temperature, carcasses were washed three times in PBST and incubated with 1:500 dilutions (in PBST) of anti-mouse IgG conjugated to Alexa 488. Fluorescence microscopy was performed after one wash in PBST and a brief staining with 0.1 mg/ml 4 0,6-Diamidino-2-phenylindole dihydrochloride (DAPI). Supplemental Data Supplemental Data including Table S1 and primer sequences are available at 10/4/531/DC1/. Acknowledgments We thank M. Layton for isolation of Cb-dpy(nm4) and Cb-tra- 2(nm9ts); H. Smith, L. Mathies, P. Kroll-Conner, G. Seydoux, and M.H. Lee for technical advice; T. Inouye and P. Sternberg for syis802, and S. Strome for antisera. We also thank J. Rothman for helpful comments on the manuscript, and T. Schedl, S. Nayak, and A. Spence for communicating results prior to publication. This work was supported by the University of Maryland, an Oak Ridge- Associated Univeristies R.E. Powe, Jr. Faculty Enhancement Award, and National Science Foundation grant IBN to E.S.H., and by Natural Sciences and Engineering Research Council Canada grants to D.P. Received: October 28, 2005 Revised: January 25, 2006 Accepted: February 1, 2006 Published: April 3, 2006 References Ahringer, J., and Kimble, J. (1991). Control of the sperm-oocyte switch in Caenorhabditis elegans hermaphrodites by the fem untranslated region. Nature 349, Ahringer, J., Rosenquist, T., Lawson, D., and Kimble, J. (1992). The Caenorhabditis elegans sex determining gene fem-3 is regulated post-transcriptionally. EMBO J. 11, Barton, M.K., Schedl, T.B., and Kimble, J. (1987). Gain-of-function mutations of fem-3, a sex-determination gene in Caenorhabditis elegans. Genetics 115, Chen, P., and Ellis, R.E. (2000). TRA-1A regulates transcription of fog-3, which controls germ cell fate in C. elegans. Development 127, Chen, P., Cho, S., Jin, S., and Ellis, R. (2001). Specification of germ cell fates by FOG-3 has been conserved during nematode evolution. Genetics 158, Chin-Sang, I.D., and Spence, A.M. (1996). Caenorhabditis elegans sex-determining protein FEM-2 is a protein phosphatase that promotes male development and interacts directly with FEM-3. Genes Dev. 10, Cho, S., Jin, S.W., Cohen, A., and Ellis, R.E. (2004). A phylogeny of Caenorhabditis reveals frequent loss of introns during nematode evolution. Genome Res. 14, Clifford, R., Lee, M., Nayak, S., Ohmachi, M., Giorgini, F., and Schedl, T. (2000). FOG-2, a novel F-box-containing protein, associates with the GLD-1 RNA-binding protein and directs male sex determination in the C. elegans hermaphrodite germline. Development 127, Colosimo, P.F., Peichel, C.L., Nereng, K., Blackman, B.K., Shapiro, M.D., Schluter, D., and Kingsley, D.M. (2004). The genetic architecture of parallel armor plate reduction in threespine sticklebacks. PLoS Biol. 2, E109. Cresko, W.A., Amores, A., Wilson, C., Murphy, J., Currey, M., Phillips, P., Bell, M.A., Kimmel, C.B., and Postlethwait, J.H. (2004). Parallel genetic basis for repeated evolution of armor loss in Alaskan threespine stickleback populations. Proc. Natl. Acad. Sci. USA 101, Doniach, T. (1986). Activity of the sex-determining gene tra-2 is modulated to allow spermatogenesis in the C. elegans hermaphrodite. Genetics 114, Doniach, T., and Hodgkin, J. (1984). A sex-determining gene, fem-1, required for both male and hermaphrodite development in Caenorhabditis elegans. Dev. Biol. 106, Edgley, M., D Souza, A., Moulder, G., McKay, S., Shen, B., Gilchrist, E., Moerman, D., and Barstead, R. (2002). Improved detection of small deletions in complex pools of DNA. Nucleic Acids Res. 30, e52. Eizirik, E., Yuhki, N., Johnson, W.E., Menotti-Raymond, M., Hannah, S.S., and O Brien, S.J. (2003). Molecular genetics and evolution of melanism in the cat family. Curr. Biol. 13, Fitch, D. (2002). WormAtlas Chapter 2 of Appendix, Phylogeny ( Francis, R., Barton, M.K., Kimble, J., and Schedl, T. (1995). gld-1, a tumor suppressor gene required for oocyte development in Caenorhabditis elegans. Genetics 139, Gompel, N., and Carroll, S.B. (2003). Genetic mechanisms and constraints governing the evolution of correlated traits in drosophilid flies. Nature 424, Goodwin, E., and Ellis, R. (2002). Turning clustering loops: sex determination in Caenorhabditis elegans. Curr. Biol. 12, R111 R120. Goodwin, E.B., Okkema, P.G., Evans, T.C., and Kimble, J. (1993). Translational regulation of tra-2 by its 3 0 untranslated region controls sexual identity in C. elegans. Cell 75, Haag, E. (2005). The evolution of nematode sex determination: C. elegans as a reference point for comparative biology. In Wormbook ( Haag, E., Wang, S., and Kimble, J. (2002). Rapid coevolution of the nematode sex-determining genes fem-3 and tra-2. Curr. Biol. 12, Hodgkin, J. (1986). Sex determination in the nematode C. elegans: analysis of tra-3 suppressors and characterization of fem genes. Genetics 114, Hodgkin, J. (1987). A genetic analysis of the sex-determining gene, tra-1, in the nematode Caenorhabditis elegans. Genes Dev. 1,
8 Developmental Cell 538 Hoekstra, H.E., and Nachman, M.W. (2003). Different genes underlie adaptive melanism in different populations of rock pocket mice. Mol. Ecol. 12, Jan, E., Yoon, J.W., Walterhouse, D., Iannaccone, P., and Goodwin, E.B. (1997). Conservation of the C. elegans tra UTR translational control. EMBO J. 16, Jones, A.R., Francis, R., and Schedl, T. (1996). GLD-1, a cytoplasmic protein essential for oocyte differentiation, shows stage- and sexspecific expression during Caenorhabditis elegans germline development. Dev. Biol. 180, Kimble, J., Edgar, L., and Hirsh, D. (1984). Specification of male development in C. elegans: thefem genes. Dev. Biol. 105, Kiontke, K., Gavin, N.P., Raynes, Y., Roehrig, C., Piano, F., and Fitch, D.H. (2004). Caenorhabditis phylogeny predicts convergence of hermaphroditism and extensive intron loss. Proc. Natl. Acad. Sci. USA 101, Kuwabara, P.E. (1996). A novel regulatory mutation in the C. elegans sex determination gene tra-2 defines a candidate ligand/receptor interaction site. Development 122, Luz, J.G., Hassig, C.A., Pickle, C., Godzik, A., Meyer, B.J., and Wilson, I.A. (2003). XOL-1, primary determinant of sexual fate in C. elegans, is a GHMP kinase family member and a structural prototype for a class of developmental regulators. Genes Dev. 17, Nayak, S., Goree, J., and Schedl, T. (2005). fog-2 and the evolution of self-fertile hermaphroditism in Caenorhabditis. PLoS Biol. 3, e6. Patel, N.H., Ball, E.E., and Goodman, C.S. (1992). Changing role of even-skipped during the evolution of insect pattern formation. Nature 357, Piekny, A.J., Wissmann, A., and Mains, P.E. (2000). Embryonic morphogenesis in Caenorhabditis elegans integrates the activity of LET-502 Rho-binding kinase, MEL-11 myosin phosphatase, DAF-2 insulin receptor and FEM-2 PP2c phosphatase. Genetics 156, Pilgrim, D., McGregor, A., Jackle, P., Johnson, T., and Hansen, D. (1995). The C. elegans sex-determining gene fem-2 encodes a putative protein phosphatase. Mol. Biol. Cell 6, Protas, M.E., Hersey, C., Kochanek, D., Zhou, Y., Wilkens, H., Jeffery, W.R., Zon, L.I., Borowsky, R., and Tabin, C.J. (2006). Genetic analysis of cavefish reveals molecular convergence in the evolution of albinism. Nat. Genet. 38, Puoti, A., Gallegos, M., Zhang, B., Wickens, M.P., and Kimble, J. (1997). Controls of cell fate and pattern by 3 0 UTRs: the C. elegans sperm/oocyte decision. Cold Spring Harb. Symp. Quant. Biol. 62, Rosenquist, T., and Kimble, J. (1988). Molecular cloning and transcript analysis of fem-3, a sex determination gene in Caenorhabditis elegans. Genes Dev. 2, Schedl, T., and Kimble, J. (1988). fog-2, a germ-line-specific sex determination gene required for hermaphrodite spermatogenesis in Caenorhabditis elegans. Genetics 119, Schedl, T., Graham, P.L., Barton, M.K., and Kimble, J. (1989). Analysis of the role of tra-1 in germline sex determination in the nematode Caneorhabditis elegans. Genetics 123, Shapiro, M.D., Marks, M.E., Peichel, C.L., Blackman, B.K., Nereng, K.S., Jonsson, B., Schluter, D., and Kingsley, D.M. (2004). Genetic and developmental basis of evolutionary pelvic reduction in threespine sticklebacks. Nature 428, Spence, A., Coulson, A., and Hodgkin, J. (1990). The product of fem-1, a nematode sex-determining gene, contains a motif found in cell cycle control proteins and receptors for cell-cell interactions. Cell 60, Stein, L., Bao, Z., Blasiar, D., Blumenthal, T., Brent, M.R., Chen, N., Chinwalla, A., Clarke, L., Clee, C., Coghlan, A., et al. (2003). The genome sequence of Caenorhabditis briggsae: a platform for comparative genomics. PLoS Biol. 1, E45. Stothard, P., and Pilgrim, D. (2003). Sex determination gene and pathway evolution in nematodes. Bioessays 25, Stothard, P., and Pilgrim, D. (2006). Conspecific and interspecific interactions between the FEM-2 and FEM-3 sexdetermining proteins despite rapid sequence divergence. J. Mol. Evol., in press. Published online February 13, /s Stothard, P., Hansen, D., and Pilgrim, D. (2002). Evolution of the PP2C family in Caenorhabditis: rapid divergence of the sex-determining protein FEM-2. J. Mol. Evol. 54, Sucena, E., Delon, I., Jones, I., Payre, F., and Stern, D.L. (2003). Regulatory evolution of shavenbaby/ovo underlies multiple cases of morphological parallelism. Nature 424, Swofford, D. (2002). Phylogenetic Analysis Using Parsimony (and Other Methods) (Sunderland, MA: Sinauer). True, J., and Haag, E. (2001). Developmental system drift and flexibility in evolutionary trajectories. Evol. Dev. 3, Wittkopp, P.J., Williams, B.L., Selegue, J.E., and Carroll, S.B. (2003). Drosophila pigmentation evolution: divergent genotypes underlying convergent phenotypes. Proc. Natl. Acad. Sci. USA 100, Wood, W.B. (1988). The Nematode Caenorhabditis elegans (Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press). Yamamoto, Y., Stock, D.W., and Jeffery, W.R. (2004). Hedgehog signalling controls eye degeneration in blind cavefish. Nature 431, Zhang, B., Gallegos, M., Puoti, A., Durkin, A., Fields, S., Kimble, J., and Wickens, M.P. (1997). A conserved RNA binding protein that regulates sexual fates in the C. elegans hermaphrodite germ line. Nature 390,
Conspecific and Interspecific Interactions Between the FEM-2 and the FEM-3 Sex-Determining Proteins Despite Rapid Sequence Divergence
 J Mol Evol (2006) 62:281 291 DOI: 10.1007/s00239-005-0084-5 Conspecific and Interspecific Interactions Between the FEM-2 and the FEM-3 Sex-Determining Proteins Despite Rapid Sequence Divergence Paul Stothard,
J Mol Evol (2006) 62:281 291 DOI: 10.1007/s00239-005-0084-5 Conspecific and Interspecific Interactions Between the FEM-2 and the FEM-3 Sex-Determining Proteins Despite Rapid Sequence Divergence Paul Stothard,
Sex and the single worm: sex determination in the nematode C. elegans
 Mechanisms of Development 83 (1999) 3 15 Review article Sex and the single worm: sex determination in the nematode C. elegans Dave Hansen, Dave Pilgrim* Department of Biological Sciences, University of
Mechanisms of Development 83 (1999) 3 15 Review article Sex and the single worm: sex determination in the nematode C. elegans Dave Hansen, Dave Pilgrim* Department of Biological Sciences, University of
A complementation test would be done by crossing the haploid strains and scoring the phenotype in the diploids.
 Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Nature Biotechnology: doi: /nbt Supplementary Figure 1. Overexpression of YFP::GPR-1 in the germline.
 Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
Supplementary Figure 1 Overexpression of YFP::GPR-1 in the germline. The pie-1 promoter and 3 utr were used to express yfp::gpr-1 in the germline. Expression levels from the yfp::gpr-1(cai 1.0)-expressing
A predicted membrane protein, TRA-2A, directs hermaphrodite development in Caenorhabditis elegans
 Development 121, 2995-3004 (1995) Printed in Great Britain The Company of Biologists Limited 1995 2995 A predicted membrane protein,, directs hermaphrodite development in Caenorhabditis elegans Patricia
Development 121, 2995-3004 (1995) Printed in Great Britain The Company of Biologists Limited 1995 2995 A predicted membrane protein,, directs hermaphrodite development in Caenorhabditis elegans Patricia
Context-dependent function of a conserved translational regulatory module
 Development epress online publication date 7 March 2012 RESEARCH ARTICLE 1509 Development 139, 1509-1521 (2012) doi:10.1242/dev.070128 2012. Published by The Company of Biologists Ltd Context-dependent
Development epress online publication date 7 March 2012 RESEARCH ARTICLE 1509 Development 139, 1509-1521 (2012) doi:10.1242/dev.070128 2012. Published by The Company of Biologists Ltd Context-dependent
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/1178343/dc1 Supporting Online Material for Starvation Protects Germline Stem Cells and Extends Reproductive Longevity in C. elegans This PDF file includes: Giana Angelo
www.sciencemag.org/cgi/content/full/1178343/dc1 Supporting Online Material for Starvation Protects Germline Stem Cells and Extends Reproductive Longevity in C. elegans This PDF file includes: Giana Angelo
Evolution of the PP2C Family in Caenorhabditis: Rapid Divergence of the Sex-Determining Protein FEM-2
 J Mol Evol (2002) 54:267 282 DOI: 10.1007/s0023901-0008-y Springer-Verlag New York Inc. 2002 Evolution of the PP2C Family in Caenorhabditis: Rapid Divergence of the Sex-Determining Protein FEM-2 Paul Stothard,
J Mol Evol (2002) 54:267 282 DOI: 10.1007/s0023901-0008-y Springer-Verlag New York Inc. 2002 Evolution of the PP2C Family in Caenorhabditis: Rapid Divergence of the Sex-Determining Protein FEM-2 Paul Stothard,
Sex-determination gene and pathway evolution in nematodes
 Sex-determination gene and pathway evolution in nematodes Paul Stothard and Dave Pilgrim* Summary The pathway that controls sexual fate in the nematode Caenorhabditis elegans has been well characterized
Sex-determination gene and pathway evolution in nematodes Paul Stothard and Dave Pilgrim* Summary The pathway that controls sexual fate in the nematode Caenorhabditis elegans has been well characterized
UNLIKE many other developmental processes, such
 Copyright Ó 2008 by the Genetics Society of America DOI: 10.1534/genetics.107.073668 Comparative Genetics of Sex Determination: Masculinizing Mutations in Caenorhabditis briggsae Danielle F. Kelleher,*,1
Copyright Ó 2008 by the Genetics Society of America DOI: 10.1534/genetics.107.073668 Comparative Genetics of Sex Determination: Masculinizing Mutations in Caenorhabditis briggsae Danielle F. Kelleher,*,1
Genetic regulation of entry into meiosis in Caenorhabditis elegans
 Development 125, 1803-1813 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3823 1803 Genetic regulation of entry into meiosis in Caenorhabditis elegans Lisa C. Kadyk 1 and Judith
Development 125, 1803-1813 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3823 1803 Genetic regulation of entry into meiosis in Caenorhabditis elegans Lisa C. Kadyk 1 and Judith
Rapid Coevolution of the Nematode Sex-Determining Genes fem-3 and tra-2
 Current Biology, Vol. 12, 2035 2041, December 10, 2002, 2002 Elsevier Science Ltd. All rights reserved. PII S0960-9822(02)01333-7 Rapid Coevolution of the Nematode Sex-Determining Genes fem-3 and tra-2
Current Biology, Vol. 12, 2035 2041, December 10, 2002, 2002 Elsevier Science Ltd. All rights reserved. PII S0960-9822(02)01333-7 Rapid Coevolution of the Nematode Sex-Determining Genes fem-3 and tra-2
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Caenorhabditis elegans
 Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
The phenotype of this worm is wild type. When both genes are mutant: The phenotype of this worm is double mutant Dpy and Unc phenotype.
 Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams - Complementation There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Green Fluorescent Protein (GFP) Today s Nobel Prize in Chemistry
 In the news: High-throughput sequencing using Solexa/Illumina technology The copy number of each fetal chromosome can be determined by direct sequencing of DNA in cell-free plasma from pregnant women Confession:
In the news: High-throughput sequencing using Solexa/Illumina technology The copy number of each fetal chromosome can be determined by direct sequencing of DNA in cell-free plasma from pregnant women Confession:
Solutions to Problem Set 4
 Question 1 Solutions to 7.014 Problem Set 4 Because you have not read much scientific literature, you decide to study the genetics of garden peas. You have two pure breeding pea strains. One that is tall
Question 1 Solutions to 7.014 Problem Set 4 Because you have not read much scientific literature, you decide to study the genetics of garden peas. You have two pure breeding pea strains. One that is tall
Designer Genes C Test
 Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
Northern Regional: January 19 th, 2019 Designer Genes C Test Name(s): Team Name: School Name: Team Number: Rank: Score: Directions: You will have 50 minutes to complete the test. You may not write on the
2. Der Dissertation zugrunde liegende Publikationen und Manuskripte. 2.1 Fine scale mapping in the sex locus region of the honey bee (Apis mellifera)
 2. Der Dissertation zugrunde liegende Publikationen und Manuskripte 2.1 Fine scale mapping in the sex locus region of the honey bee (Apis mellifera) M. Hasselmann 1, M. K. Fondrk², R. E. Page Jr.² und
2. Der Dissertation zugrunde liegende Publikationen und Manuskripte 2.1 Fine scale mapping in the sex locus region of the honey bee (Apis mellifera) M. Hasselmann 1, M. K. Fondrk², R. E. Page Jr.² und
The Worm, Ceanorhabditis elegans
 1 1 Institute of Biology University of Iceland October, 2005 Lecture outline The problem of phenotype Dear Max Sidney Brenner A Nobel Prize in Medicine Genome sequence Some tools Gene structure Genomic
1 1 Institute of Biology University of Iceland October, 2005 Lecture outline The problem of phenotype Dear Max Sidney Brenner A Nobel Prize in Medicine Genome sequence Some tools Gene structure Genomic
Germline stem cell differentiation entails regional control of cell fate. regulator GLD-1 in Caenorhabditis elegans
 Genetics: Early Online, published on January 12, 216 as 1.1534/genetics.115.185678 Germline stem cell differentiation entails regional control of cell fate regulator GLD-1 in Caenorhabditis elegans John
Genetics: Early Online, published on January 12, 216 as 1.1534/genetics.115.185678 Germline stem cell differentiation entails regional control of cell fate regulator GLD-1 in Caenorhabditis elegans John
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Dose-dependent control of proliferation and sperm specification by FOG-1/CPEB
 First posted online on 6 July 2005 as 10.1242/dev.01921 Access the most recent version epress at http://dev.biologists.org/lookup/doi/10.1242/dev.01921 online publication date 6 July 2005 3471 Dose-dependent
First posted online on 6 July 2005 as 10.1242/dev.01921 Access the most recent version epress at http://dev.biologists.org/lookup/doi/10.1242/dev.01921 online publication date 6 July 2005 3471 Dose-dependent
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Upstream Elements Regulating mir-241 and mir-48 Abstract Introduction
 Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
CPEB proteins control two key steps in spermatogenesis in C. elegans
 CPEB proteins control two key steps in spermatogenesis in C. elegans Cameron Luitjens, 1 Maria Gallegos, 1,4,6 Brian Kraemer, 2,5,6 Judith Kimble, 2,3 and Marvin Wickens 2,7 1 Program in Cell and Molecular
CPEB proteins control two key steps in spermatogenesis in C. elegans Cameron Luitjens, 1 Maria Gallegos, 1,4,6 Brian Kraemer, 2,5,6 Judith Kimble, 2,3 and Marvin Wickens 2,7 1 Program in Cell and Molecular
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
PUF-8 negatively regulates RAS/MAPK signalling to promote differentiation of C. elegans germ cells
 DEVELOPMENT AND STEM CELLS RESEARCH ARTICLE 1645 Development 140, 1645-1654 (2013) doi:10.1242/dev.088013 2013. Published by The Company of Biologists Ltd PUF-8 negatively regulates RAS/MAPK signalling
DEVELOPMENT AND STEM CELLS RESEARCH ARTICLE 1645 Development 140, 1645-1654 (2013) doi:10.1242/dev.088013 2013. Published by The Company of Biologists Ltd PUF-8 negatively regulates RAS/MAPK signalling
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
SUPPLEMENTARY INFORMATION
 Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
Supplementary Discussion Rationale for using maternal ythdf2 -/- mutants as study subject To study the genetic basis of the embryonic developmental delay that we observed, we crossed fish with different
MANY animal tissues contain populations of prolif- ald and Greenwald 1995). Interaction between LAG-2
 Copyright 2004 by the Genetics Society of America DOI: 10.1534/genetics.104.029355 Caenorhabditis elegans atx-2 Promotes Germline Proliferation and the Oocyte Fate Eleanor M. Maine,*,1 Dave Hansen,,2 Deborah
Copyright 2004 by the Genetics Society of America DOI: 10.1534/genetics.104.029355 Caenorhabditis elegans atx-2 Promotes Germline Proliferation and the Oocyte Fate Eleanor M. Maine,*,1 Dave Hansen,,2 Deborah
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice Supplementary Figure 2.
 Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
Supplementary Figure 1. Markedly decreased numbers of marginal zone B cells in DOCK8 mutant mice. Percentage of marginal zone B cells in the spleen of wild-type mice (+/+), mice homozygous for cpm or pri
When one gene is wild type and the other mutant:
 Series 2: Cross Diagrams Linkage Analysis There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Series 2: Cross Diagrams Linkage Analysis There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome:
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Science Unit Learning Summary
 Learning Summary Inheritance, variation and evolution Content Sexual and asexual reproduction. Meiosis leads to non-identical cells being formed while mitosis leads to identical cells being formed. In
Learning Summary Inheritance, variation and evolution Content Sexual and asexual reproduction. Meiosis leads to non-identical cells being formed while mitosis leads to identical cells being formed. In
AP Biology Unit 6 Practice Test 1. A group of cells is assayed for DNA content immediately following mitosis and is found to have an average of 8
 AP Biology Unit 6 Practice Test Name: 1. A group of cells is assayed for DNA content immediately following mitosis and is found to have an average of 8 picograms of DNA per nucleus. How many picograms
AP Biology Unit 6 Practice Test Name: 1. A group of cells is assayed for DNA content immediately following mitosis and is found to have an average of 8 picograms of DNA per nucleus. How many picograms
C. ELEGANS STAR PROTEINS, GLD 1 AND ASD 2, REGULATE SPECIFIC RNA TARGETS TO CONTROL DEVELOPMENT
 CHAPTER C. ELEGANS STAR PROTEINS, GLD 1 AND ASD 2, REGULATE SPECIFIC RNA TARGETS TO CONTROL DEVELOPMENT Min Ho Lee* and Tim Schedl Abstract: A comprehensive understanding of the C. elegans STAR proteins
CHAPTER C. ELEGANS STAR PROTEINS, GLD 1 AND ASD 2, REGULATE SPECIFIC RNA TARGETS TO CONTROL DEVELOPMENT Min Ho Lee* and Tim Schedl Abstract: A comprehensive understanding of the C. elegans STAR proteins
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Genome-wide germline-enriched and sex-biased expression profiles in Caenorhabditis elegans
 311 Genome-wide germline-enriched and sex-biased expression profiles in Caenorhabditis elegans Valerie Reinke 1, *, Inigo San Gil 1, Samuel Ward 2 and Keith Kazmer 1 1 Department of Genetics, Yale University
311 Genome-wide germline-enriched and sex-biased expression profiles in Caenorhabditis elegans Valerie Reinke 1, *, Inigo San Gil 1, Samuel Ward 2 and Keith Kazmer 1 1 Department of Genetics, Yale University
SIGNIFICANCE OF EMBRYOLOGY
 This lecture will discuss the following topics : Definition of Embryology Significance of Embryology Old and New Frontiers Introduction to Molecular Regulation and Signaling Descriptive terms in Embryology
This lecture will discuss the following topics : Definition of Embryology Significance of Embryology Old and New Frontiers Introduction to Molecular Regulation and Signaling Descriptive terms in Embryology
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
MATHEMATICAL MODELS - Vol. III - Mathematical Modeling and the Human Genome - Hilary S. Booth MATHEMATICAL MODELING AND THE HUMAN GENOME
 MATHEMATICAL MODELING AND THE HUMAN GENOME Hilary S. Booth Australian National University, Australia Keywords: Human genome, DNA, bioinformatics, sequence analysis, evolution. Contents 1. Introduction:
MATHEMATICAL MODELING AND THE HUMAN GENOME Hilary S. Booth Australian National University, Australia Keywords: Human genome, DNA, bioinformatics, sequence analysis, evolution. Contents 1. Introduction:
Nature Genetics: doi: /ng Supplementary Figure 1. The phenotypes of PI , BR121, and Harosoy under short-day conditions.
 Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Supplementary Figure 1 The phenotypes of PI 159925, BR121, and Harosoy under short-day conditions. (a) Plant height. (b) Number of branches. (c) Average internode length. (d) Number of nodes. (e) Pods
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
ATX-2, the C. elegans ortholog of ataxin 2, functions in translational regulation in the germline
 Research article 4831 ATX-2, the C. elegans ortholog of ataxin 2, functions in translational regulation in the germline Rafal Ciosk 1,2, Michael DePalma 1,2 and James R. Priess 1,2,3, * 1 Division of Basic
Research article 4831 ATX-2, the C. elegans ortholog of ataxin 2, functions in translational regulation in the germline Rafal Ciosk 1,2, Michael DePalma 1,2 and James R. Priess 1,2,3, * 1 Division of Basic
Biology 218, practise Exam 2, 2011
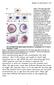 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms.
 Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Practicing Biology BIG IDEA 3.A 1. Draw, label and describe the structure of DNA and RNA including bonding mechanisms. 2. Using at least 2 well-known experiments, describe which features of DNA and RNA
Genetic Variation: The genetic substrate for natural selection. Horizontal Gene Transfer. General Principles 10/2/17.
 Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Genetic Variation: The genetic substrate for natural selection What about organisms that do not have sexual reproduction? Horizontal Gene Transfer Dr. Carol E. Lee, University of Wisconsin In prokaryotes:
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
The phenotype of this worm is wild type. When both genes are mutant: The phenotype of this worm is double mutant Dpy and Unc phenotype.
 Series 1: Cross Diagrams There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome: When both
Series 1: Cross Diagrams There are two alleles for each trait in a diploid organism In C. elegans gene symbols are ALWAYS italicized. To represent two different genes on the same chromosome: When both
A forkhead protein controls sexual identity of the C. elegans male somatic gonad
 Research article 1425 A forkhead protein controls sexual identity of the C. elegans male somatic gonad Weiru Chang 1,2, Christopher Tilmann 3,4, Kara Thoemke 1,2, Finn-Hugo Markussen 3,4, Laura D. Mathies
Research article 1425 A forkhead protein controls sexual identity of the C. elegans male somatic gonad Weiru Chang 1,2, Christopher Tilmann 3,4, Kara Thoemke 1,2, Finn-Hugo Markussen 3,4, Laura D. Mathies
Primary sex determination in the nematode C. elegans
 Development Supplement. 5-6 (987) Printed in Great Britain The Company of Biologists Limited 987 Primary sex determination in the nematode C. elegans JONATHAN HODGKIN MRC Laboratory of Molecular Biology,
Development Supplement. 5-6 (987) Printed in Great Britain The Company of Biologists Limited 987 Primary sex determination in the nematode C. elegans JONATHAN HODGKIN MRC Laboratory of Molecular Biology,
GLD-3, a Bicaudal-C Homolog that Inhibits FBF to Control Germline Sex Determination in C. elegans
 Developmental Cell, Vol. 3, 697 710, November, 2002, Copyright 2002 by Cell Press GLD-3, a Bicaudal-C Homolog that Inhibits FBF to Control Germline Sex Determination in C. elegans Christian R. Eckmann,
Developmental Cell, Vol. 3, 697 710, November, 2002, Copyright 2002 by Cell Press GLD-3, a Bicaudal-C Homolog that Inhibits FBF to Control Germline Sex Determination in C. elegans Christian R. Eckmann,
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Objective 3.01 (DNA, RNA and Protein Synthesis)
 Objective 3.01 (DNA, RNA and Protein Synthesis) DNA Structure o Discovered by Watson and Crick o Double-stranded o Shape is a double helix (twisted ladder) o Made of chains of nucleotides: o Has four types
Objective 3.01 (DNA, RNA and Protein Synthesis) DNA Structure o Discovered by Watson and Crick o Double-stranded o Shape is a double helix (twisted ladder) o Made of chains of nucleotides: o Has four types
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
(Write your name on every page. One point will be deducted for every page without your name!)
 POPULATION GENETICS AND MICROEVOLUTIONARY THEORY FINAL EXAMINATION (Write your name on every page. One point will be deducted for every page without your name!) 1. Briefly define (5 points each): a) Average
POPULATION GENETICS AND MICROEVOLUTIONARY THEORY FINAL EXAMINATION (Write your name on every page. One point will be deducted for every page without your name!) 1. Briefly define (5 points each): a) Average
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Q2 (4.6) Put the following in order from biggest to smallest: Gene DNA Cell Chromosome Nucleus. Q8 (Biology) (4.6)
 Q1 (4.6) What is variation? Q2 (4.6) Put the following in order from biggest to smallest: Gene DNA Cell Chromosome Nucleus Q3 (4.6) What are genes? Q4 (4.6) What sort of reproduction produces genetically
Q1 (4.6) What is variation? Q2 (4.6) Put the following in order from biggest to smallest: Gene DNA Cell Chromosome Nucleus Q3 (4.6) What are genes? Q4 (4.6) What sort of reproduction produces genetically
Genetics 275 Notes Week 7
 Cytoplasmic Inheritance Genetics 275 Notes Week 7 Criteriafor recognition of cytoplasmic inheritance: 1. Reciprocal crosses give different results -mainly due to the fact that the female parent contributes
Cytoplasmic Inheritance Genetics 275 Notes Week 7 Criteriafor recognition of cytoplasmic inheritance: 1. Reciprocal crosses give different results -mainly due to the fact that the female parent contributes
Chapter 13 Meiosis and Sexual Reproduction
 Biology 110 Sec. 11 J. Greg Doheny Chapter 13 Meiosis and Sexual Reproduction Quiz Questions: 1. What word do you use to describe a chromosome or gene allele that we inherit from our Mother? From our Father?
Biology 110 Sec. 11 J. Greg Doheny Chapter 13 Meiosis and Sexual Reproduction Quiz Questions: 1. What word do you use to describe a chromosome or gene allele that we inherit from our Mother? From our Father?
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Untitled Document. A. antibiotics B. cell structure C. DNA structure D. sterile procedures
 Name: Date: 1. The discovery of which of the following has most directly led to advances in the identification of suspects in criminal investigations and in the identification of genetic diseases? A. antibiotics
Name: Date: 1. The discovery of which of the following has most directly led to advances in the identification of suspects in criminal investigations and in the identification of genetic diseases? A. antibiotics
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
Aparticularly fruitful way to study the mechanisms saman 1989; Pannell 1997a,b), but little is known
 Copyright 2000 by the Genetics Society of America Regulatory Elements Required for Development of Caenorhabditis elegans Hermaphrodites Are Conserved in the tra-2 Homologue of C. remanei, a Male/Female
Copyright 2000 by the Genetics Society of America Regulatory Elements Required for Development of Caenorhabditis elegans Hermaphrodites Are Conserved in the tra-2 Homologue of C. remanei, a Male/Female
Bi Lecture 8 Genetic Pathways and Genetic Screens
 Bi190-2013 Lecture 8 Genetic Pathways and Genetic Screens WT A 2X:2A her-1 tra-1 1X:2A her-1 tra-1 Female body Male body Female body Male body her-1(lf) B 2X:2A her-1(lf) tra-1 1X:2A her-1(lf) tra-1 Female
Bi190-2013 Lecture 8 Genetic Pathways and Genetic Screens WT A 2X:2A her-1 tra-1 1X:2A her-1 tra-1 Female body Male body Female body Male body her-1(lf) B 2X:2A her-1(lf) tra-1 1X:2A her-1(lf) tra-1 Female
allosteric cis-acting DNA element coding strand dominant constitutive mutation coordinate regulation of genes denatured
 A B C D E F G H I J K L M N O P Q R S T U V W X Y Z AA BB CC DD EE FF GG HH II JJ KK LL MM NN OO PP QQ RR SS TT UU VV allosteric cis-acting DNA element coding strand codominant constitutive mutation coordinate
A B C D E F G H I J K L M N O P Q R S T U V W X Y Z AA BB CC DD EE FF GG HH II JJ KK LL MM NN OO PP QQ RR SS TT UU VV allosteric cis-acting DNA element coding strand codominant constitutive mutation coordinate
Functional exploration of the C. elegans genome using DNA microarrays
 Functional exploration of the C. elegans genome using DNA microarrays Valerie Reinke doi:10.1038/ng1039 Global changes in gene expression underlie developmental processes such as organogenesis, embryogenesis
Functional exploration of the C. elegans genome using DNA microarrays Valerie Reinke doi:10.1038/ng1039 Global changes in gene expression underlie developmental processes such as organogenesis, embryogenesis
MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition in C. Elegans
 Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition
Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition
GENETICS 603 Exam 3, Dec 2, 2011
 GENETICS 603 Exam 3, Dec 2, 2011 Name I. Chromosome IV in Drosophila is a very small chromosome with few genes. Fruitflies that are trisomic for chromosome IV have no apparent phenotypic abnormalities,
GENETICS 603 Exam 3, Dec 2, 2011 Name I. Chromosome IV in Drosophila is a very small chromosome with few genes. Fruitflies that are trisomic for chromosome IV have no apparent phenotypic abnormalities,
GENETICS UNIT VOCABULARY CHART. Word Definition Word Part Visual/Mnemonic Related Words 1. adenine Nitrogen base, pairs with thymine in DNA and uracil
 Word Definition Word Part Visual/Mnemonic Related Words 1. adenine Nitrogen base, pairs with thymine in DNA and uracil in RNA 2. allele One or more alternate forms of a gene Example: P = Dominant (purple);
Word Definition Word Part Visual/Mnemonic Related Words 1. adenine Nitrogen base, pairs with thymine in DNA and uracil in RNA 2. allele One or more alternate forms of a gene Example: P = Dominant (purple);
Cover Requirements: Name of Unit Colored picture representing something in the unit
 Name: Period: Cover Requirements: Name of Unit Colored picture representing something in the unit Biology B1 1 Target # Biology Unit B1 (Genetics & Meiosis) Learning Targets Genetics & Meiosis I can explain
Name: Period: Cover Requirements: Name of Unit Colored picture representing something in the unit Biology B1 1 Target # Biology Unit B1 (Genetics & Meiosis) Learning Targets Genetics & Meiosis I can explain
Full file at CHAPTER 2 Genetics
 CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
CHAPTER 2 Genetics MULTIPLE CHOICE 1. Chromosomes are a. small linear bodies. b. contained in cells. c. replicated during cell division. 2. A cross between true-breeding plants bearing yellow seeds produces
Genetically Engineering Yeast to Understand Molecular Modes of Speciation
 Genetically Engineering Yeast to Understand Molecular Modes of Speciation Mark Umbarger Biophysics 242 May 6, 2004 Abstract: An understanding of the molecular mechanisms of speciation (reproductive isolation)
Genetically Engineering Yeast to Understand Molecular Modes of Speciation Mark Umbarger Biophysics 242 May 6, 2004 Abstract: An understanding of the molecular mechanisms of speciation (reproductive isolation)
Analysis of Sex Reversal and TRA-2 Nucleotide Variation in Tropical and Temperate Clades of Caenorhabditis Briggsae
 Wright State University CORE Scholar Browse all Theses and Dissertations Theses and Dissertations 2007 Analysis of Sex Reversal and TRA-2 Nucleotide Variation in Tropical and Temperate Clades of Caenorhabditis
Wright State University CORE Scholar Browse all Theses and Dissertations Theses and Dissertations 2007 Analysis of Sex Reversal and TRA-2 Nucleotide Variation in Tropical and Temperate Clades of Caenorhabditis
Name Class Date. KEY CONCEPT Gametes have half the number of chromosomes that body cells have.
 Section 1: Chromosomes and Meiosis KEY CONCEPT Gametes have half the number of chromosomes that body cells have. VOCABULARY somatic cell autosome fertilization gamete sex chromosome diploid homologous
Section 1: Chromosomes and Meiosis KEY CONCEPT Gametes have half the number of chromosomes that body cells have. VOCABULARY somatic cell autosome fertilization gamete sex chromosome diploid homologous
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Biology I Level - 2nd Semester Final Review
 Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
Biology I Level - 2nd Semester Final Review The 2 nd Semester Final encompasses all material that was discussed during second semester. It s important that you review ALL notes and worksheets from the
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Sex-biased expression of the C. elegans X chromosome is a result of both X chromosome copy number and sex-specific gene regulation.
 Genetics: Early Online, published on June 9, 16 as 1.1534/genetics.116.1998 Sex-biased expression of the C. elegans X chromosome is a result of both X chromosome copy number and sex-specific gene regulation.
Genetics: Early Online, published on June 9, 16 as 1.1534/genetics.116.1998 Sex-biased expression of the C. elegans X chromosome is a result of both X chromosome copy number and sex-specific gene regulation.
GENETIC CONTROL OF CELL INTERACTIONS IN NEMATODE DEVELOPMENT
 Annu. Rev. Genet. 1991. 25. 411-36 Copyright 199l by Annual Reviews Inc. All rights reserved GENETIC CONTROL OF CELL INTERACTIONS IN NEMATODE DEVELOPMENT E. J. Lambie and Judith Kimble Laboratory of Molecular
Annu. Rev. Genet. 1991. 25. 411-36 Copyright 199l by Annual Reviews Inc. All rights reserved GENETIC CONTROL OF CELL INTERACTIONS IN NEMATODE DEVELOPMENT E. J. Lambie and Judith Kimble Laboratory of Molecular
1. Contains the sugar ribose instead of deoxyribose. 2. Single-stranded instead of double stranded. 3. Contains uracil in place of thymine.
 Protein Synthesis & Mutations RNA 1. Contains the sugar ribose instead of deoxyribose. 2. Single-stranded instead of double stranded. 3. Contains uracil in place of thymine. RNA Contains: 1. Adenine 2.
Protein Synthesis & Mutations RNA 1. Contains the sugar ribose instead of deoxyribose. 2. Single-stranded instead of double stranded. 3. Contains uracil in place of thymine. RNA Contains: 1. Adenine 2.
South Pacific Form Seven Certificate BIOLOGY. QUESTION and ANSWER BOOKLET
 / South Pacific Form Seven Certificate INSTRUCTIONS BIOLOGY 7 Write your Student Personal Identification Number (SPIN) in the space provided on the top right hand corner of this page. Answer ALL QUESTIONS.
/ South Pacific Form Seven Certificate INSTRUCTIONS BIOLOGY 7 Write your Student Personal Identification Number (SPIN) in the space provided on the top right hand corner of this page. Answer ALL QUESTIONS.
Reinforcement Unit 3 Resource Book. Meiosis and Mendel KEY CONCEPT Gametes have half the number of chromosomes that body cells have.
 6.1 CHROMOSOMES AND MEIOSIS KEY CONCEPT Gametes have half the number of chromosomes that body cells have. Your body is made of two basic cell types. One basic type are somatic cells, also called body cells,
6.1 CHROMOSOMES AND MEIOSIS KEY CONCEPT Gametes have half the number of chromosomes that body cells have. Your body is made of two basic cell types. One basic type are somatic cells, also called body cells,
doi: /nature09429
 SUPPLEMENTARY INFORMATION doi:10.1038/nature09429 C. e xnd-1 MSSEPIVANLDN------------------SMTTEPAPAAFPPISPMSFAFMNEEERAAVKFP--- C. b xnd-1 MN--DLAKN---------------------LLKHSQTTDSAPPISPLSFAFMTDEEKAAFKFPASE
SUPPLEMENTARY INFORMATION doi:10.1038/nature09429 C. e xnd-1 MSSEPIVANLDN------------------SMTTEPAPAAFPPISPMSFAFMNEEERAAVKFP--- C. b xnd-1 MN--DLAKN---------------------LLKHSQTTDSAPPISPLSFAFMTDEEKAAFKFPASE
Drosophila germline stem cells
 Development 125, 679-690 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3776 679 Nanos and Pumilio have critical roles in the development and function of Drosophila germline
Development 125, 679-690 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV3776 679 Nanos and Pumilio have critical roles in the development and function of Drosophila germline
Quiz Section 4 Molecular analysis of inheritance: An amphibian puzzle
 Genome 371, Autumn 2018 Quiz Section 4 Molecular analysis of inheritance: An amphibian puzzle Goals: To illustrate how molecular tools can be used to track inheritance. In this particular example, we will
Genome 371, Autumn 2018 Quiz Section 4 Molecular analysis of inheritance: An amphibian puzzle Goals: To illustrate how molecular tools can be used to track inheritance. In this particular example, we will
Supplementary Figure 1. Nature Genetics: doi: /ng.3848
 Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Supplementary Figure 1 Phenotypes and epigenetic properties of Fab2L flies. A- Phenotypic classification based on eye pigment levels in Fab2L male (orange bars) and female (yellow bars) flies (n>150).
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
BIOLOGY 111. CHAPTER 5: Chromosomes and Inheritance
 BIOLOGY 111 CHAPTER 5: Chromosomes and Inheritance Chromosomes and Inheritance Learning Outcomes 5.1 Differentiate between sexual and asexual reproduction in terms of the genetic variation of the offspring.
BIOLOGY 111 CHAPTER 5: Chromosomes and Inheritance Chromosomes and Inheritance Learning Outcomes 5.1 Differentiate between sexual and asexual reproduction in terms of the genetic variation of the offspring.
Chromosome Chr Duplica Duplic t a ion Pixley
 Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
Chromosome Duplication Pixley Figure 4-6 Molecular Biology of the Cell ( Garland Science 2008) Figure 4-72 Molecular Biology of the Cell ( Garland Science 2008) Interphase During mitosis (cell division),
