Functional Organization of the Transcriptome in Human Brain
|
|
|
- Ashlie Berry
- 5 years ago
- Views:
Transcription
1 Functional Organization of the Transcriptome in Human Brain Michael C. Oldham Laboratory of Daniel H. Geschwind, UCLA BIOCOMP 08, Las Vegas, NV July 15, 2008
2 Neurons Astrocytes Oligodendrocytes Microglia ~10 11 ~10 12
3 Microarray studies of the human brain face a number of challenges Cellular heterogeneity mrna extracted from tissue homogenates Sample quality Post-mortem tissue Sample size Typically, <10 individuals per group Focus on differential expression
4 What s wrong with differential expression?
5 Methodological overview: WGCNA
6 The data Network 1 (U133 cortex): 67 samples, 67 individuals Brain region Reference # samples # outliers # samples post QC BA46 Iwamoto et al.* ** BA9, BA11 Ryan et al. 42 (31 BA9, 11 BA11) 8 (5 BA9, 3 BA11) 21 (13 BA9, 8 BA11)** BA4, BA9 Hodges et al. 28 (16 BA4, 12 BA9) 2 (2 BA9) 26 (16 BA4, 10 BA9) Network 2 (U95 cortex): 42 samples, 32 individuals Brain region Reference # samples # outliers # samples post QC BrA (x2), ACC, PrV, Prf, BrRH, Prm Khaitovich et al (2 each region) FP, ait, IPL, MFG Caceres et al (2 FP) BA9 Enard et al (2 BA9) BA9 Khaitovich et al (BA9) BA10 Lu et al (BA10) BA10 Iwamoto et al.* (BA10) Network 3 (U133 caudate nucleus): 27 samples, 27 individuals Brain region Reference # samples # outliers # samples post QC Caudate nucleus Hodges et al Network 4 (U133 cerebellum): 24 samples, 24 individuals Brain region Reference # samples # outliers # samples post QC Cerebellum Hodges et al * Raw data obtained in collaboration with Dr. Kazuya Iwamoto and Dr. Tadafumi Kato at RIKEN. ** Following outlier removal, there were 11 pairs of samples from the same individuals in Iwamoto et al. and Ryan et al. One unique sample per individual was retained.
7 A: CTX The networks M1A M2 M3 M4A M5A M6A M7A M8A M9A M10A M11A M12A M13A M14A M15A M16A M17A M18A M19A B: CTX_95 M5B M20 M21 M17B M22 M23 M24 M4B M9B M16B M10B M15B M25 M8B M19B M26 M18B C: CN M27 M4C M11C M28 M29 M30 M31 M19C M9C M1C M32 M13C M18C M33 M34 M35 M36 M8C M16C M37 M5C M15C M38 D: CB M1D M12D M19D M39 M18D M40 M41 M6D M10D M42 M14D M11D M4D M7D M43 M44 M45 M16D M46 M9D M15D M47
8 Many modules are preserved across networks
9 Quantifying module membership with k me k me is the Pearson correlation between the expression level of a given probe set and a given eigengene, e.g.:
10 k me is significantly correlated across networks A B C
11
12
13
14 Conserved modules are enriched for markers of major cell classes Network Module Color Oligodendrocytes 1 Oligodendrocytes 2 Astrocytes 1 Astrocytes 2 Neurons 1 Neurons 2 CTX M9A turquoise 138 / 352 p = 8.1e / 60 p = 8.3e-43 NS NS NS NS CTX95 M9B turquoise 63 / 256 p = 4.3e / 55 p = 1.1e-27 NS NS NS NS CN M9C turquoise 93 / 352 p = 8.8e / 60 p = 2.1e-28 NS NS NS NS CB M9D turquoise 58 / 352 p = 5.6e / 60 p = 2.2e-35 NS NS NS NS CTX M15A brown NS NS 220 / 554 p = 2.0e / 32 p = 5.7e-14 NS NS CTX95 M15B brown NS 11 / 55 p = 3.2e / 430 p = 5.3e / 27 p = 7.2e-09 NS NS CN M15C brown NS NS 99 / 554 p = 2.5e / 32 p = 6.6e-14 NS NS CB M15D brown NS 9 / 60 p = 1.0e / 554 p = 6.1e / 32 p = 1.1e-17 NS NS CTX M16A blue NS NS NS NS 164 / 709 p = 8.4e / 50 p = 2.0e-03 CTX95 M16B blue NS NS NS NS 68 / 546 p = 4.0e-15 7 / 40 p = 2.4e-02 CN M16C blue NS NS NS NS 64 / 709 p = 1.7e / 50 p = 4.4e-06 CB M16D blue NS NS NS NS 50 / 709 p = 5.4e-05 7 / 50 p = 4.1e-02 Network Module Oligodendrocytes 3 Astrocytes 4 Synaptic proteins 5 Synaptic proteins 6 CTX M9A 128/458 p=5.5e-46 NS NS NS CTX_95 M9B 49/358 p=1.4e-16 4/18 p=4.6e-02 NS NS CN M9C 76/458 p=2.3e-37 NS NS NS CB M9D 43/458 p=1.0e-17 NS NS NS CTX M15A NS 15/22 p=2.6e-12 NS NS CTX_95 M15B NS 12/18 p=1.9e-10 NS NS CN M15C NS 9/22 p=1.5e-08 NS NS CB M15D NS 14/22 p=1.7e-17 NS NS CTX M16A NS NS 117/580 p=6.9e-16 23/60 p=1.6e-08 CTX_95 M16B NS NS 46/490 p=4.7e-06 10/55 p=1.6e-03 CN M16C NS NS 55/580 p=3.1e-13 13/60 p=4.1e-07 CB M16D NS NS 53/580 p=4.2e-09 15/60 p=4.0e-08 1 Cahoy, J.D. et al. J Neurosci 28: (2008) 3 Nielsen, J.A. et al. J Neurosci 26: (2006) 5 Genes2Cognition Consortium 2 Lein, E.S. et al. Nature 445: (2007) 4 Bachoo, R.M. et al. PNAS 101: (2004) 6 Morciano, M. et al. J Neurochem 95: (2005)
15 Thought experiment Amount Sample
16 The cortical transcriptome is organized into functional modules M1A M2 M3 M4A M5A M6A M7A M8A M9A M10A M11A M12A M13A M14A M15A M16A M17A M18A M19A Module Color Interpretation Characterization method P-value M1A palegreen Gender Correlation with gender status 4.8e-21 M2 midnightblue Ribosome Gene ontology 2.3e-64 M3 greenyellow? N/A N/A M4A purple Immune response: microglia Gene ontology 2.6e-56 M5A orange Immune response: microglia Gene ontology 9.8e-42 M6A darkolivegreen PVALB+ interneuron PVALB kme 5.3e-15 M7A honeydew Mitochondria Gene ontology 2.0e-16 M8A salmon? N/A N/A M9A turquoise Oligodendrocyte Enrichment analysis with cell-type markers 8.1e-71 M10A green Glutamatergic synapse Enrichment analysis with synaptic proteins 5.3e-18 M11A black? N/A N/A M12A tan Hypoxia Enrichment analysis with hypoxia genes 5.6e-13 M13A powderblue Neurogenesis Gene ontology 5.0e-03 M14A pink Glutamatergic synapse Enrichment analysis with synaptic proteins 1.6e-20 M15A brown Astrocyte Enrichment analysis with cell-type markers 2.0e-122 M16A blue Neuron Enrichment analysis with cell-type markers 8.4e-30 M17A tomato PVALB+ interneuron PVALB kme 2.0e-06 M18A yellow Metabolism Gene ontology 2.1e-07 M19A red Membrane/signal transduction Gene ontology 6.8e-08
17 Modules are organized into a functional meta-network Neurons Oligodendrocytes Astrocytes Meningeal cells Ribosomes Neurogenesis Hypoxia Glutamatergic synapses Mitochondria Gender Purkinje neurons Microglia PVALB+ interneurons
18 Applications 1) Context-specific annotation for genes expressed in the human brain ( guilt-by-association ) Rationale: genes with the strongest evidence of membership for the same module are likely to be driven by the same underlying factors
19
20
21 M6A: PVALB+ interneurons
22 Applications 1) Context-specific annotation for genes expressed in the human brain ( guilt-by-association ) 2) In silico comparisons of cellular specificity of gene expression across brain regions Rationale: genes with the most significant differences in membership for cell-type modules between brain regions imply differences in the cellular specificity and/or consistency of gene expression
23 Network 1 Network 2 Probe set Gene Module MM ntwk 1 MM ntwk 2 P-value Expr ntwk 1 Expr ntwk 2 P-value CN CB _s_at MON2 M e e-02 CN CB _s_at ATM /// LOC M e e-04 CN CB _at ST18 M e <2.2e-16 CTX CB _at ST18 M e <2.2e-16 CTX CB _at ZNF536 M e e-15 CTX CB _at HIP1 M e e-06 CTX CB _s_at NCAM1 M e <2.2e-16 CTX CB _at LRP2 M e e-10 CTX CB _s_at MTUS1 M e <2.2e-16 CTX CN _at CRTC3 M e <2.2e-16 CN CB _s_at LMO3 M e <2.2e-16 CN CB _at NKX2-2 M e e-03 CN CB _s_at LHFP M e <2.2e-16 CTX CB _s_at LMO3 M e <2.2e-16 CTX CB _at DAO M e <2.2e-16 CTX CB _at AGT M e e-09 CTX CB _at TIMP4 M e <2.2e-16 CTX CN _s_at MYO10 M e e-15 CTX CN _s_at TPD52L1 M e <2.2e-16 CTX CN _at ACSBG1 M e e-13 CN CB _at CX3CL1 M e <2.2e-16 CN CB _at DRD1 M e <2.2e-16 CTX CB _s_at RGS4 M e <2.2e-16 CTX CB _s_at MTUS1 M e <2.2e-16 CTX CB _s_at PCDHGC3 M e e-16 CTX CB _x_at CFLAR M e <2.2e-16 CTX CN _s_at GMFB M e e-11 CTX CN _s_at CA12 M e <2.2e-16 CTX CN _s_at PAK1 /// VDP M e e-10 CTX CN _s_at ENSA M e <2.2e-16
24 Applications 1) Context-specific annotation for genes expressed in the human brain ( guilt-by-association ) 2) In silico comparisons of cellular specificity of gene expression across brain regions 3) Cellular phenotype discovery Rationale: unsupervised analysis of gene coexpression patterns can identify novel distinctions among cell types within brain regions
25 Caudate nucleus M13C M15C Astrocytes 1 Astrocytes 2 Astrocytes 3 99 / 554 (p=2.5e-61) 14 / 32 (p=6.6e-14) 9 / 22 (p=1.5e-08) Astrocytes 1 Astrocytes 2 Astrocytes 3 45 / 554 (p=1.9e-24) 8 / 32 (p=6.6e-08) 7 / 22 (p=1.0e-07) 1 Cahoy, J.D. et al. J Neurosci 28: (2008) 2 Lein, E.S. et al. Nature 445: (2007) 3 Bachoo, R.M. et al. PNAS 101: (2004)
26 M13C (CN) identifies genes that are coexpressed in the SVZ
27 Conclusions The human brain transcriptome is organized into modules of coexpressed genes Many modules are reproducible across microarrays, individuals, and brain regions Several highly conserved modules are enriched for markers of major cell classes Core transcriptional programs for neurons, oligodendrocytes, astrocytes, and microglia Context-specific annotation for thousands of genes expressed in the human brain Potential to leverage consistency of k me for comparisons with other conditions of interest (e.g. disease)
28 Acknowledgements Dan Geschwind Steve Horvath Gena Konopka Peter Langfelder Collaborators: Dr. Kazuya Iwamoto Dr. Tadafumi Kato The Geschwind lab
Weighted gene co-expression analysis. Yuehua Cui June 7, 2013
 Weighted gene co-expression analysis Yuehua Cui June 7, 2013 Weighted gene co-expression network (WGCNA) A type of scale-free network: A scale-free network is a network whose degree distribution follows
Weighted gene co-expression analysis Yuehua Cui June 7, 2013 Weighted gene co-expression network (WGCNA) A type of scale-free network: A scale-free network is a network whose degree distribution follows
Neurochemistry 1. Nervous system is made of neurons & glia, as well as other cells. Santiago Ramon y Cajal Nobel Prize 1906
 Neurochemistry 1 Nervous system is made of neurons & glia, as well as other cells. Santiago Ramon y Cajal Nobel Prize 1906 How Many Neurons Do We Have? The human brain contains ~86 billion neurons and
Neurochemistry 1 Nervous system is made of neurons & glia, as well as other cells. Santiago Ramon y Cajal Nobel Prize 1906 How Many Neurons Do We Have? The human brain contains ~86 billion neurons and
A Geometric Interpretation of Gene Co-Expression Network Analysis. Steve Horvath, Jun Dong
 A Geometric Interpretation of Gene Co-Expression Network Analysis Steve Horvath, Jun Dong Outline Network and network concepts Approximately factorizable networks Gene Co-expression Network Eigengene Factorizability,
A Geometric Interpretation of Gene Co-Expression Network Analysis Steve Horvath, Jun Dong Outline Network and network concepts Approximately factorizable networks Gene Co-expression Network Eigengene Factorizability,
An Overview of Weighted Gene Co-Expression Network Analysis. Steve Horvath University of California, Los Angeles
 An Overview of Weighted Gene Co-Expression Network Analysis Steve Horvath University of California, Los Angeles Contents How to construct a weighted gene co-expression network? Why use soft thresholding?
An Overview of Weighted Gene Co-Expression Network Analysis Steve Horvath University of California, Los Angeles Contents How to construct a weighted gene co-expression network? Why use soft thresholding?
Cells. Steven McLoon Department of Neuroscience University of Minnesota
 Cells Steven McLoon Department of Neuroscience University of Minnesota 1 Microscopy Methods of histology: Treat the tissue with a preservative (e.g. formaldehyde). Dissect the region of interest. Embed
Cells Steven McLoon Department of Neuroscience University of Minnesota 1 Microscopy Methods of histology: Treat the tissue with a preservative (e.g. formaldehyde). Dissect the region of interest. Embed
Weighted Correlation Network Analysis and Systems Biologic Applications. Steve Horvath University of California, Los Angeles
 Weighted Correlation Network Analysis and Systems Biologic Applications Steve Horvath University of California, Los Angeles Contents Weighted correlation network analysis (WGCNA) Module preservation statistics
Weighted Correlation Network Analysis and Systems Biologic Applications Steve Horvath University of California, Los Angeles Contents Weighted correlation network analysis (WGCNA) Module preservation statistics
Intro and Homeostasis
 Intro and Homeostasis Physiology - how the body works. Homeostasis - staying the same. Functional Types of Neurons Sensory (afferent - coming in) neurons: Detects the changes in the body. Informations
Intro and Homeostasis Physiology - how the body works. Homeostasis - staying the same. Functional Types of Neurons Sensory (afferent - coming in) neurons: Detects the changes in the body. Informations
EXPLORING BRAIN TRANSCRIPTOMIC PATTERNS: A TOPOLOGICAL ANALYSIS USING SPATIAL EXPRESSION NETWORKS
 EXPLORING BRAIN TRANSCRIPTOMIC PATTERNS: A TOPOLOGICAL ANALYSIS USING SPATIAL EXPRESSION NETWORKS ZHANA KUNCHEVA Department of Mathematics Imperial College London, UK E-mail: z.kuncheva12@imperial.ac.uk
EXPLORING BRAIN TRANSCRIPTOMIC PATTERNS: A TOPOLOGICAL ANALYSIS USING SPATIAL EXPRESSION NETWORKS ZHANA KUNCHEVA Department of Mathematics Imperial College London, UK E-mail: z.kuncheva12@imperial.ac.uk
Cellular Neuroanatomy II The Prototypical Neuron: Neurites. Reading: BCP Chapter 2
 Cellular Neuroanatomy II The Prototypical Neuron: Neurites Reading: BCP Chapter 2 Major Internal Features of a Neuron The neuron is the functional unit of the nervous system. A typical neuron has a soma
Cellular Neuroanatomy II The Prototypical Neuron: Neurites Reading: BCP Chapter 2 Major Internal Features of a Neuron The neuron is the functional unit of the nervous system. A typical neuron has a soma
TRANSCRIPTOMICS. (or the analysis of the transcriptome) Mario Cáceres. Main objectives of genomics. Determine the entire DNA sequence of an organism
 TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
TRANSCRIPTOMICS (or the analysis of the transcriptome) Mario Cáceres Main objectives of genomics Determine the entire DNA sequence of an organism Identify and annotate the complete set of genes encoded
Module preservation statistics
 Module preservation statistics Module preservation is often an essential step in a network analysis Steve Horvath University of California, Los Angeles Construct a network Rationale: make use of interaction
Module preservation statistics Module preservation is often an essential step in a network analysis Steve Horvath University of California, Los Angeles Construct a network Rationale: make use of interaction
Dendrites - receives information from other neuron cells - input receivers.
 The Nerve Tissue Neuron - the nerve cell Dendrites - receives information from other neuron cells - input receivers. Cell body - includes usual parts of the organelles of a cell (nucleus, mitochondria)
The Nerve Tissue Neuron - the nerve cell Dendrites - receives information from other neuron cells - input receivers. Cell body - includes usual parts of the organelles of a cell (nucleus, mitochondria)
Ch 8: Neurons: Cellular and Network Properties, Part 1
 Developed by John Gallagher, MS, DVM Ch 8: Neurons: Cellular and Network Properties, Part 1 Objectives: Describe the Cells of the NS Explain the creation and propagation of an electrical signal in a nerve
Developed by John Gallagher, MS, DVM Ch 8: Neurons: Cellular and Network Properties, Part 1 Objectives: Describe the Cells of the NS Explain the creation and propagation of an electrical signal in a nerve
Physiology 2 nd year. Neuroscience Optional Lecture
 Academic year 2018/2019 Physiology 2 nd year Semester 1 Curricula Nervous system physiology Blood physiology Acid-base equilibrium Bibliography: Boron & Boulpaep Medical Physiology, 3 rd edition Physiology
Academic year 2018/2019 Physiology 2 nd year Semester 1 Curricula Nervous system physiology Blood physiology Acid-base equilibrium Bibliography: Boron & Boulpaep Medical Physiology, 3 rd edition Physiology
From Genes to Cognition to Social Behavior a DNA Microarray Research Proposal
 From Genes to Cognition to Social Behavior a DNA Microarray Research Proposal Jonathan Cachat & Melanie Huffman October 6, 2008 Abstract Cachat, Huffman 1 A difference in active gene expression in cerebrocortical
From Genes to Cognition to Social Behavior a DNA Microarray Research Proposal Jonathan Cachat & Melanie Huffman October 6, 2008 Abstract Cachat, Huffman 1 A difference in active gene expression in cerebrocortical
Singapore Institute for Neurotechnology & Memory Network Programme, National University of Singapore, Singapore
 SUPPLEMENTARY INFORMATION: Gene expression links functional networks across cortex and striatum Kevin M Anderson 1, Fenna M Krienen 2, Eun Young Choi 3, Jenna M Reinen 1, B T Thomas Yeo 4,5, Avram J Holmes
SUPPLEMENTARY INFORMATION: Gene expression links functional networks across cortex and striatum Kevin M Anderson 1, Fenna M Krienen 2, Eun Young Choi 3, Jenna M Reinen 1, B T Thomas Yeo 4,5, Avram J Holmes
WGCNA User Manual. (for version 1.0.x)
 (for version 1.0.x) WGCNA User Manual A systems biologic microarray analysis software for finding important genes and pathways. The WGCNA (weighted gene co-expression network analysis) software implements
(for version 1.0.x) WGCNA User Manual A systems biologic microarray analysis software for finding important genes and pathways. The WGCNA (weighted gene co-expression network analysis) software implements
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs
 Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Dynamic modular architecture of protein-protein interaction networks beyond the dichotomy of date and party hubs Xiao Chang 1,#, Tao Xu 2,#, Yun Li 3, Kai Wang 1,4,5,* 1 Zilkha Neurogenetic Institute,
Cellular neurophysiology
 Cellular neurophysiology Katalin Schlett Dept. Physiology and Neurobiology schlett.katalin@ttk.elte.hu tel: 8380 ext; room 6-411 recommended literature: From Molecules to Networks: An Introduction to Cellular
Cellular neurophysiology Katalin Schlett Dept. Physiology and Neurobiology schlett.katalin@ttk.elte.hu tel: 8380 ext; room 6-411 recommended literature: From Molecules to Networks: An Introduction to Cellular
Gene Ontology and Functional Enrichment. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein
 Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Gene Ontology and Functional Enrichment Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein The parsimony principle: A quick review Find the tree that requires the fewest
Assembling a Cellular User Manual for the Brain
 This Accepted Manuscript has not been copyedited and formatted. The final version may differ from this version. Progressions Assembling a Cellular User Manual for the Brain Steven A. Sloan and Ben A. Barres
This Accepted Manuscript has not been copyedited and formatted. The final version may differ from this version. Progressions Assembling a Cellular User Manual for the Brain Steven A. Sloan and Ben A. Barres
Cells were in by, who under a microscope He saw He called them Several worked to the which states:
 Cells were in by, who under a microscope He saw He called them Several worked to the which states: Cells Because, the before scientists could study them Cells are very tiny structures called Plant cells
Cells were in by, who under a microscope He saw He called them Several worked to the which states: Cells Because, the before scientists could study them Cells are very tiny structures called Plant cells
Clustering & microarray technology
 Clustering & microarray technology A large scale way to measure gene expression levels. Thanks to Kevin Wayne, Matt Hibbs, & SMD for a few of the slides 1 Why is expression important? Proteins Gene Expression
Clustering & microarray technology A large scale way to measure gene expression levels. Thanks to Kevin Wayne, Matt Hibbs, & SMD for a few of the slides 1 Why is expression important? Proteins Gene Expression
Introduction Principles of Signaling and Organization p. 3 Signaling in Simple Neuronal Circuits p. 4 Organization of the Retina p.
 Introduction Principles of Signaling and Organization p. 3 Signaling in Simple Neuronal Circuits p. 4 Organization of the Retina p. 5 Signaling in Nerve Cells p. 9 Cellular and Molecular Biology of Neurons
Introduction Principles of Signaling and Organization p. 3 Signaling in Simple Neuronal Circuits p. 4 Organization of the Retina p. 5 Signaling in Nerve Cells p. 9 Cellular and Molecular Biology of Neurons
NEURONS Excitable cells Therefore, have a RMP Synapse = chemical communication site between neurons, from pre-synaptic release to postsynaptic
 NEUROPHYSIOLOGY NOTES L1 WHAT IS NEUROPHYSIOLOGY? NEURONS Excitable cells Therefore, have a RMP Synapse = chemical communication site between neurons, from pre-synaptic release to postsynaptic receptor
NEUROPHYSIOLOGY NOTES L1 WHAT IS NEUROPHYSIOLOGY? NEURONS Excitable cells Therefore, have a RMP Synapse = chemical communication site between neurons, from pre-synaptic release to postsynaptic receptor
Eigengene Network Analysis of Human and Chimpanzee Microarray Data R Tutorial
 Eigengene Network Analysis of Human and Chimpanzee Microarray Data R Tutorial Peter Langfelder and Steve Horvath Correspondence: shorvath@mednet.ucla.edu, Peter.Langfelder@gmail.com This is a self contained
Eigengene Network Analysis of Human and Chimpanzee Microarray Data R Tutorial Peter Langfelder and Steve Horvath Correspondence: shorvath@mednet.ucla.edu, Peter.Langfelder@gmail.com This is a self contained
Lecture 3: A basic statistical concept
 Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Lecture 3: A basic statistical concept P value In statistical hypothesis testing, the p value is the probability of obtaining a result at least as extreme as the one that was actually observed, assuming
Cellular Neuroanatomy I The Prototypical Neuron: Soma. Reading: BCP Chapter 2
 Cellular Neuroanatomy I The Prototypical Neuron: Soma Reading: BCP Chapter 2 Functional Unit of the Nervous System The functional unit of the nervous system is the neuron. Neurons are cells specialized
Cellular Neuroanatomy I The Prototypical Neuron: Soma Reading: BCP Chapter 2 Functional Unit of the Nervous System The functional unit of the nervous system is the neuron. Neurons are cells specialized
Unit 2: The Structure and function of Organisms. Section 2: Inside Cells
 Unit 2: The Structure and function of Organisms Section 2: 42 Essential Question: Are all cells the same? - Vocabulary 1. 2. 3. 4. 5. 6. 7. 8. Eukaryotic Prokaryotic Organelle Plant Cell Animal Cell Chloroplast
Unit 2: The Structure and function of Organisms Section 2: 42 Essential Question: Are all cells the same? - Vocabulary 1. 2. 3. 4. 5. 6. 7. 8. Eukaryotic Prokaryotic Organelle Plant Cell Animal Cell Chloroplast
Neurons. The Molecular Basis of their Electrical Excitability
 Neurons The Molecular Basis of their Electrical Excitability Viva La Complexity! Consider, The human brain contains >10 11 neurons! Each neuron makes 10 3 (average) synaptic contacts on up to 10 3 other
Neurons The Molecular Basis of their Electrical Excitability Viva La Complexity! Consider, The human brain contains >10 11 neurons! Each neuron makes 10 3 (average) synaptic contacts on up to 10 3 other
Cells of the nervous system
 Cells of the nervous system There are approximately 100 billion neurons in the human brain There are about 100 times as many glial cells in the human brain Similar origin, different functions Other cells
Cells of the nervous system There are approximately 100 billion neurons in the human brain There are about 100 times as many glial cells in the human brain Similar origin, different functions Other cells
Human-Specific Transcriptional Networks in the Brain
 Human-Specific Transcriptional Networks in the Brain Genevieve Konopka, University of California Tara Friedrich, University of California Jeremy Davis-Turak, University of California Kellen Winden, University
Human-Specific Transcriptional Networks in the Brain Genevieve Konopka, University of California Tara Friedrich, University of California Jeremy Davis-Turak, University of California Kellen Winden, University
Housekeeping, 26 January 2009
 5 th & 6 th Lectures Mon 26 & Wed 28 Jan 2009 Vertebrate Physiology ECOL 437 (MCB/VetSci 437) Univ. of Arizona, spring 2009 Neurons Chapter 11 Kevin Bonine & Kevin Oh 1. Finish Solutes + Water 2. Neurons
5 th & 6 th Lectures Mon 26 & Wed 28 Jan 2009 Vertebrate Physiology ECOL 437 (MCB/VetSci 437) Univ. of Arizona, spring 2009 Neurons Chapter 11 Kevin Bonine & Kevin Oh 1. Finish Solutes + Water 2. Neurons
Neurons. 5 th & 6 th Lectures Mon 26 & Wed 28 Jan Finish Solutes + Water. 2. Neurons. Chapter 11
 5 th & 6 th Lectures Mon 26 & Wed 28 Jan 2009 Vertebrate Physiology ECOL 437 (MCB/VetSci 437) Univ. of Arizona, spring 2009 Neurons Chapter 11 Kevin Bonine & Kevin Oh 1. Finish Solutes + Water 2. Neurons
5 th & 6 th Lectures Mon 26 & Wed 28 Jan 2009 Vertebrate Physiology ECOL 437 (MCB/VetSci 437) Univ. of Arizona, spring 2009 Neurons Chapter 11 Kevin Bonine & Kevin Oh 1. Finish Solutes + Water 2. Neurons
A transcriptome meta-analysis identifies the response of plant to stresses. Etienne Delannoy, Rim Zaag, Guillem Rigaill, Marie-Laure Martin-Magniette
 A transcriptome meta-analysis identifies the response of plant to es Etienne Delannoy, Rim Zaag, Guillem Rigaill, Marie-Laure Martin-Magniette Biological context Multiple biotic and abiotic es impacting
A transcriptome meta-analysis identifies the response of plant to es Etienne Delannoy, Rim Zaag, Guillem Rigaill, Marie-Laure Martin-Magniette Biological context Multiple biotic and abiotic es impacting
1 Introduction. Abstract
 CBS 530 Assignment No 2 SHUBHRA GUPTA shubhg@asu.edu 993755974 Review of the papers: Construction and Analysis of a Human-Chimpanzee Comparative Clone Map and Intra- and Interspecific Variation in Primate
CBS 530 Assignment No 2 SHUBHRA GUPTA shubhg@asu.edu 993755974 Review of the papers: Construction and Analysis of a Human-Chimpanzee Comparative Clone Map and Intra- and Interspecific Variation in Primate
Ch 7. The Nervous System 7.1 & 7.2
 Ch 7 The Nervous System 7.1 & 7.2 SLOs Describe the different types of neurons and supporting cells, and identify their functions. Identify the myelin sheath and describe how it is formed in the CNS and
Ch 7 The Nervous System 7.1 & 7.2 SLOs Describe the different types of neurons and supporting cells, and identify their functions. Identify the myelin sheath and describe how it is formed in the CNS and
What are neurons for?
 5 th & 6 th Lectures Mon 26 & Wed 28 Jan 2009 Vertebrate Physiology ECOL 437 (MCB/VetSci 437) Univ. of Arizona, spring 2009 Kevin Bonine & Kevin Oh 1. Finish Solutes Water 2. Neurons Neurons Chapter 11
5 th & 6 th Lectures Mon 26 & Wed 28 Jan 2009 Vertebrate Physiology ECOL 437 (MCB/VetSci 437) Univ. of Arizona, spring 2009 Kevin Bonine & Kevin Oh 1. Finish Solutes Water 2. Neurons Neurons Chapter 11
Drosophila melanogaster and D. simulans, two fruit fly species that are nearly
 Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Comparative Genomics: Human versus chimpanzee 1. Introduction The chimpanzee is the closest living relative to humans. The two species are nearly identical in DNA sequence (>98% identity), yet vastly different
Human-Specific Transcriptional Networks in the Brain
 Article Human-Specific Transcriptional Networks in the Brain Genevieve Konopka, 1,6, * Tara Friedrich, 1 Jeremy Davis-Turak, 1 Kellen Winden, 1 Michael C. Oldham, 7 Fuying Gao, 1 Leslie Chen, 1 Guang-Zhong
Article Human-Specific Transcriptional Networks in the Brain Genevieve Konopka, 1,6, * Tara Friedrich, 1 Jeremy Davis-Turak, 1 Kellen Winden, 1 Michael C. Oldham, 7 Fuying Gao, 1 Leslie Chen, 1 Guang-Zhong
Gabriella Rustici EMBL-EBI
 Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
Towards a cellular phenotype ontology Gabriella Rustici EMBL-EBI gabry@ebi.ac.uk Systems Microscopy NoE at a glance The Systems Microscopy NoE (2011 2015) is a FP7 funded project, involving 15 research
Millifluidic culture improves human midbrain organoid vitality and differentiation
 Millifluidic culture improves human midbrain organoid vitality and differentiation Emanuel Berger University of Luxembourg 3D midbrain organoids for modelling Parkinson s disease Healthy control Midbrain
Millifluidic culture improves human midbrain organoid vitality and differentiation Emanuel Berger University of Luxembourg 3D midbrain organoids for modelling Parkinson s disease Healthy control Midbrain
The Nervous System. What did you learn at school today? Neurophysiology!
 The Nervous System What did you learn at school today? Neurophysiology! The Nervous System Controls heart rate, emotions, memories, consciousness, and much more. The most intricate and beautifully complex
The Nervous System What did you learn at school today? Neurophysiology! The Nervous System Controls heart rate, emotions, memories, consciousness, and much more. The most intricate and beautifully complex
Proteomics. 2 nd semester, Department of Biotechnology and Bioinformatics Laboratory of Nano-Biotechnology and Artificial Bioengineering
 Proteomics 2 nd semester, 2013 1 Text book Principles of Proteomics by R. M. Twyman, BIOS Scientific Publications Other Reference books 1) Proteomics by C. David O Connor and B. David Hames, Scion Publishing
Proteomics 2 nd semester, 2013 1 Text book Principles of Proteomics by R. M. Twyman, BIOS Scientific Publications Other Reference books 1) Proteomics by C. David O Connor and B. David Hames, Scion Publishing
Cells and Passive Transport Study Guide
 Cells and Passive Transport Study Guide Success Criteria: - Complete - If multiple choice, answer has explanations - Quality answers/best answer possible 1. List the 2 types of active transport and the
Cells and Passive Transport Study Guide Success Criteria: - Complete - If multiple choice, answer has explanations - Quality answers/best answer possible 1. List the 2 types of active transport and the
Geert Geeven. April 14, 2010
 iction of Gene Regulatory Interactions NDNS+ Workshop April 14, 2010 Today s talk - Outline Outline Biological Background Construction of Predictors The main aim of my project is to better understand the
iction of Gene Regulatory Interactions NDNS+ Workshop April 14, 2010 Today s talk - Outline Outline Biological Background Construction of Predictors The main aim of my project is to better understand the
Introduction to Bioinformatics
 CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
CSCI8980: Applied Machine Learning in Computational Biology Introduction to Bioinformatics Rui Kuang Department of Computer Science and Engineering University of Minnesota kuang@cs.umn.edu History of Bioinformatics
6.047 / Computational Biology: Genomes, Networks, Evolution Fall 2008
 MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
MIT OpenCourseWare http://ocw.mit.edu 6.047 / 6.878 Computational Biology: Genomes, Networks, Evolution Fall 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms.
Clustering and Network
 Clustering and Network Jing-Dong Jackie Han jdhan@picb.ac.cn http://www.picb.ac.cn/~jdhan Copy Right: Jing-Dong Jackie Han What is clustering? A way of grouping together data samples that are similar in
Clustering and Network Jing-Dong Jackie Han jdhan@picb.ac.cn http://www.picb.ac.cn/~jdhan Copy Right: Jing-Dong Jackie Han What is clustering? A way of grouping together data samples that are similar in
Nervous System Organization
 The Nervous System Chapter 44 Nervous System Organization All animals must be able to respond to environmental stimuli -Sensory receptors = Detect stimulus -Motor effectors = Respond to it -The nervous
The Nervous System Chapter 44 Nervous System Organization All animals must be able to respond to environmental stimuli -Sensory receptors = Detect stimulus -Motor effectors = Respond to it -The nervous
Monitoring neurite morphology and synapse formation in primary neurons for neurotoxicity assessments and drug screening
 APPLICATION NOTE ArrayScan High Content Platform Monitoring neurite morphology and synapse formation in primary neurons for neurotoxicity assessments and drug screening Suk J. Hong and Richik N. Ghosh
APPLICATION NOTE ArrayScan High Content Platform Monitoring neurite morphology and synapse formation in primary neurons for neurotoxicity assessments and drug screening Suk J. Hong and Richik N. Ghosh
The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector.
 The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector. Omar S. Akbari*, Igor Antoshechkin*, Henry Amrhein, Brian Williams, Race Diloreto, Jeremy
The Developmental Transcriptome of the Mosquito Aedes aegypti, an invasive species and major arbovirus vector. Omar S. Akbari*, Igor Antoshechkin*, Henry Amrhein, Brian Williams, Race Diloreto, Jeremy
Neurons and Nervous Systems
 34 Neurons and Nervous Systems Concept 34.1 Nervous Systems Consist of Neurons and Glia Nervous systems have two categories of cells: Neurons, or nerve cells, are excitable they generate and transmit electrical
34 Neurons and Nervous Systems Concept 34.1 Nervous Systems Consist of Neurons and Glia Nervous systems have two categories of cells: Neurons, or nerve cells, are excitable they generate and transmit electrical
Nervous Tissue. Neurons Neural communication Nervous Systems
 Nervous Tissue Neurons Neural communication Nervous Systems What is the function of nervous tissue? Maintain homeostasis & respond to stimuli Sense & transmit information rapidly, to specific cells and
Nervous Tissue Neurons Neural communication Nervous Systems What is the function of nervous tissue? Maintain homeostasis & respond to stimuli Sense & transmit information rapidly, to specific cells and
Neurons, Synapses, and Signaling
 LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 48 Neurons, Synapses, and Signaling
LECTURE PRESENTATIONS For CAMPBELL BIOLOGY, NINTH EDITION Jane B. Reece, Lisa A. Urry, Michael L. Cain, Steven A. Wasserman, Peter V. Minorsky, Robert B. Jackson Chapter 48 Neurons, Synapses, and Signaling
The Agrin Hypothesis (McMahan, 1990)
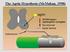 The Agrin Hypothesis (McMahan, 1990) Ectopic Expression of Agrin Reist et al., 1987 Kröger and Schröder, 2002 Agrin is a Component of b-amyloid Plaques in Alzheimer s Brains Donahue et al., (1999) PNAS
The Agrin Hypothesis (McMahan, 1990) Ectopic Expression of Agrin Reist et al., 1987 Kröger and Schröder, 2002 Agrin is a Component of b-amyloid Plaques in Alzheimer s Brains Donahue et al., (1999) PNAS
Protein function studies: history, current status and future trends
 19 3 2007 6 Chinese Bulletin of Life Sciences Vol. 19, No. 3 Jun., 2007 1004-0374(2007)03-0294-07 ( 100871) Q51A Protein function studies: history, current status and future trends MA Jing, GE Xi, CHANG
19 3 2007 6 Chinese Bulletin of Life Sciences Vol. 19, No. 3 Jun., 2007 1004-0374(2007)03-0294-07 ( 100871) Q51A Protein function studies: history, current status and future trends MA Jing, GE Xi, CHANG
Mixture models for analysing transcriptome and ChIP-chip data
 Mixture models for analysing transcriptome and ChIP-chip data Marie-Laure Martin-Magniette French National Institute for agricultural research (INRA) Unit of Applied Mathematics and Informatics at AgroParisTech,
Mixture models for analysing transcriptome and ChIP-chip data Marie-Laure Martin-Magniette French National Institute for agricultural research (INRA) Unit of Applied Mathematics and Informatics at AgroParisTech,
CJ LI LRRK2 MOUSE COMPARISON STUDY. Phenotyping Data Results
 CJ LI MOUSE COMPARISON STUDY Phenotyping Data Results NOTE FOR USE This comparison study was run as an opportunistic look at the phenotype of multiple CJ Li-generated mouse lines. The primary purpose of
CJ LI MOUSE COMPARISON STUDY Phenotyping Data Results NOTE FOR USE This comparison study was run as an opportunistic look at the phenotype of multiple CJ Li-generated mouse lines. The primary purpose of
Computational Genomics. Reconstructing dynamic regulatory networks in multiple species
 02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
02-710 Computational Genomics Reconstructing dynamic regulatory networks in multiple species Methods for reconstructing networks in cells CRH1 SLT2 SLR3 YPS3 YPS1 Amit et al Science 2009 Pe er et al Recomb
Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis
 Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Title Comparative RNA-seq analysis of transcriptome dynamics during petal development in Rosa chinensis Author list Yu Han 1, Huihua Wan 1, Tangren Cheng 1, Jia Wang 1, Weiru Yang 1, Huitang Pan 1* & Qixiang
Proteomics Systems Biology
 Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
Dr. Sanjeeva Srivastava IIT Bombay Proteomics Systems Biology IIT Bombay 2 1 DNA Genomics RNA Transcriptomics Global Cellular Protein Proteomics Global Cellular Metabolite Metabolomics Global Cellular
The Role of Network Science in Biology and Medicine. Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs
 The Role of Network Science in Biology and Medicine Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs Network Analysis Working Group 09.28.2017 Network-Enabled Wisdom (NEW) empirically
The Role of Network Science in Biology and Medicine Tiffany J. Callahan Computational Bioscience Program Hunter/Kahn Labs Network Analysis Working Group 09.28.2017 Network-Enabled Wisdom (NEW) empirically
9/11/18. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Multimodal Connectomics in Psychiatry: Bridging scales From Micro to Macro. Supplemental Information
 Multimodal Connectomics in Psychiatry: Bridging scales From Micro to Macro al Information Table S1. Overview of online resources for possible multiscale connectomics studies. Allen Brain Institute for
Multimodal Connectomics in Psychiatry: Bridging scales From Micro to Macro al Information Table S1. Overview of online resources for possible multiscale connectomics studies. Allen Brain Institute for
Fundamentals of Computational Neuroscience 2e
 Fundamentals of Computational Neuroscience 2e Thomas Trappenberg March 21, 2009 Chapter 9: Modular networks, motor control, and reinforcement learning Mixture of experts Expert 1 Input Expert 2 Integration
Fundamentals of Computational Neuroscience 2e Thomas Trappenberg March 21, 2009 Chapter 9: Modular networks, motor control, and reinforcement learning Mixture of experts Expert 1 Input Expert 2 Integration
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Bio334 Neurobiology 1 Lecture 2
 Evolution of Bio334 Neurobiology 1 Lecture 2 1 Questions from the last lecture Patients with Broca s area lesions do have trouble writing Golgi Cajal debate reticular theory of the brain vs. individual
Evolution of Bio334 Neurobiology 1 Lecture 2 1 Questions from the last lecture Patients with Broca s area lesions do have trouble writing Golgi Cajal debate reticular theory of the brain vs. individual
DEFINING THE PLAYERS IN HIGHER-ORDER NETWORKS: PREDICTIVE MODELING FOR REVERSE ENGINEERING FUNCTIONAL INFLUENCE NETWORKS
 DEFINING THE PLAYERS IN HIGHER-ORDER NETWORKS: PREDICTIVE MODELING FOR REVERSE ENGINEERING FUNCTIONAL INFLUENCE NETWORKS JASON E. MCDERMOTT 1, MICHELLE ARCHULETA 1, SUSAN L. STEVENS 2, MARY P. STENZEL-POORE
DEFINING THE PLAYERS IN HIGHER-ORDER NETWORKS: PREDICTIVE MODELING FOR REVERSE ENGINEERING FUNCTIONAL INFLUENCE NETWORKS JASON E. MCDERMOTT 1, MICHELLE ARCHULETA 1, SUSAN L. STEVENS 2, MARY P. STENZEL-POORE
MULTISCALE MODULARITY IN BRAIN SYSTEMS
 MULTISCALE MODULARITY IN BRAIN SYSTEMS Danielle S. Bassett University of California Santa Barbara Department of Physics The Brain: A Multiscale System Spatial Hierarchy: Temporal-Spatial Hierarchy: http://www.idac.tohoku.ac.jp/en/frontiers/column_070327/figi-i.gif
MULTISCALE MODULARITY IN BRAIN SYSTEMS Danielle S. Bassett University of California Santa Barbara Department of Physics The Brain: A Multiscale System Spatial Hierarchy: Temporal-Spatial Hierarchy: http://www.idac.tohoku.ac.jp/en/frontiers/column_070327/figi-i.gif
Bi 1x Spring 2014: LacI Titration
 Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
Bi 1x Spring 2014: LacI Titration 1 Overview In this experiment, you will measure the effect of various mutated LacI repressor ribosome binding sites in an E. coli cell by measuring the expression of a
9/2/17. Molecular and Cellular Biology. 3. The Cell From Genes to Proteins. key processes
 Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Molecular and Cellular Biology Animal Cell ((eukaryotic cell) -----> compare with prokaryotic cell) ENDOPLASMIC RETICULUM (ER) Rough ER Smooth ER Flagellum Nuclear envelope Nucleolus NUCLEUS Chromatin
Synchrony in Neural Systems: a very brief, biased, basic view
 Synchrony in Neural Systems: a very brief, biased, basic view Tim Lewis UC Davis NIMBIOS Workshop on Synchrony April 11, 2011 components of neuronal networks neurons synapses connectivity cell type - intrinsic
Synchrony in Neural Systems: a very brief, biased, basic view Tim Lewis UC Davis NIMBIOS Workshop on Synchrony April 11, 2011 components of neuronal networks neurons synapses connectivity cell type - intrinsic
Tracking whole-brain connectivity dynamics in the resting-state
 Tracking whole-brain connectivity dynamics in the resting-state Supplementary Table. Peak Coordinates of ICNs ICN regions BA t max Peak (mm) (continued) BA t max Peak (mm) X Y Z X Y Z Subcortical networks
Tracking whole-brain connectivity dynamics in the resting-state Supplementary Table. Peak Coordinates of ICNs ICN regions BA t max Peak (mm) (continued) BA t max Peak (mm) X Y Z X Y Z Subcortical networks
Bioinformatics. Transcriptome
 Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Bioinformatics Transcriptome Jacques.van.Helden@ulb.ac.be Université Libre de Bruxelles, Belgique Laboratoire de Bioinformatique des Génomes et des Réseaux (BiGRe) http://www.bigre.ulb.ac.be/ Bioinformatics
Computational Systems Biology
 Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
Computational Systems Biology Vasant Honavar Artificial Intelligence Research Laboratory Bioinformatics and Computational Biology Graduate Program Center for Computational Intelligence, Learning, & Discovery
CONJOINT 541. Translating a Transcriptome at Specific Times and Places. David Morris. Department of Biochemistry
 CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
CONJOINT 541 Translating a Transcriptome at Specific Times and Places David Morris Department of Biochemistry http://faculty.washington.edu/dmorris/ Lecture 1 The Biology and Experimental Analysis of mrna
Introduction to clustering methods for gene expression data analysis
 Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Introduction to clustering methods for gene expression data analysis Giorgio Valentini e-mail: valentini@dsi.unimi.it Outline Levels of analysis of DNA microarray data Clustering methods for functional
Information processing. Divisions of nervous system. Neuron structure and function Synapse. Neurons, synapses, and signaling 11/3/2017
 Neurons, synapses, and signaling Chapter 48 Information processing Divisions of nervous system Central nervous system (CNS) Brain and a nerve cord Integration center Peripheral nervous system (PNS) Nerves
Neurons, synapses, and signaling Chapter 48 Information processing Divisions of nervous system Central nervous system (CNS) Brain and a nerve cord Integration center Peripheral nervous system (PNS) Nerves
Pathway Association Analysis Trey Ideker UCSD
 Pathway Association Analysis Trey Ideker UCSD A working network map of the cell Network evolutionary comparison / cross-species alignment to identify conserved modules The Working Map Network-based classification
Pathway Association Analysis Trey Ideker UCSD A working network map of the cell Network evolutionary comparison / cross-species alignment to identify conserved modules The Working Map Network-based classification
Multilayer and dynamic networks
 OHBM 2016 Multilayer and dynamic networks Danielle S. Bassett University of Pennsylvania Department of Bioengineering Outline Statement of the problem Statement of a solution: Multilayer Modeling Utility
OHBM 2016 Multilayer and dynamic networks Danielle S. Bassett University of Pennsylvania Department of Bioengineering Outline Statement of the problem Statement of a solution: Multilayer Modeling Utility
1
 http://photos1.blogger.com/img/13/2627/640/screenhunter_047.jpg 1 The Nose Knows http://graphics.jsonline.com/graphics/owlive/img/mar05/sideways.one0308_big.jpg 2 http://www.stlmoviereviewweekly.com/sitebuilder/images/sideways-253x364.jpg
http://photos1.blogger.com/img/13/2627/640/screenhunter_047.jpg 1 The Nose Knows http://graphics.jsonline.com/graphics/owlive/img/mar05/sideways.one0308_big.jpg 2 http://www.stlmoviereviewweekly.com/sitebuilder/images/sideways-253x364.jpg
DOWNLOAD OR READ : THE NEURONAL CYTOSKELETON MOTOR PROTEINS AND ORGANELLE TRAFFICKING IN THE AXON PDF EBOOK EPUB MOBI
 DOWNLOAD OR READ : THE NEURONAL CYTOSKELETON MOTOR PROTEINS AND ORGANELLE TRAFFICKING IN THE AXON PDF EBOOK EPUB MOBI Page 1 Page 2 the neuronal cytoskeleton motor proteins and organelle trafficking in
DOWNLOAD OR READ : THE NEURONAL CYTOSKELETON MOTOR PROTEINS AND ORGANELLE TRAFFICKING IN THE AXON PDF EBOOK EPUB MOBI Page 1 Page 2 the neuronal cytoskeleton motor proteins and organelle trafficking in
Clustering. Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein. Some slides adapted from Jacques van Helden
 Clustering Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein Some slides adapted from Jacques van Helden Gene expression profiling A quick review Which molecular processes/functions
Clustering Genome 559: Introduction to Statistical and Computational Genomics Elhanan Borenstein Some slides adapted from Jacques van Helden Gene expression profiling A quick review Which molecular processes/functions
Fuzzy Clustering of Gene Expression Data
 Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Fuzzy Clustering of Gene Data Matthias E. Futschik and Nikola K. Kasabov Department of Information Science, University of Otago P.O. Box 56, Dunedin, New Zealand email: mfutschik@infoscience.otago.ac.nz,
Overview of clustering analysis. Yuehua Cui
 Overview of clustering analysis Yuehua Cui Email: cuiy@msu.edu http://www.stt.msu.edu/~cui A data set with clear cluster structure How would you design an algorithm for finding the three clusters in this
Overview of clustering analysis Yuehua Cui Email: cuiy@msu.edu http://www.stt.msu.edu/~cui A data set with clear cluster structure How would you design an algorithm for finding the three clusters in this
The majority of cells in the nervous system arise during the embryonic and early post
 Introduction Introduction The majority of cells in the nervous system arise during the embryonic and early post natal period. These cells are derived from population of neural stem cells first shown by
Introduction Introduction The majority of cells in the nervous system arise during the embryonic and early post natal period. These cells are derived from population of neural stem cells first shown by
Systems biology and complexity research
 Systems biology and complexity research Peter Schuster Institut für Theoretische Chemie, Universität Wien, Austria and The Santa Fe Institute, Santa Fe, New Mexico, USA Interdisciplinary Challenges for
Systems biology and complexity research Peter Schuster Institut für Theoretische Chemie, Universität Wien, Austria and The Santa Fe Institute, Santa Fe, New Mexico, USA Interdisciplinary Challenges for
Inferring disease-associated lncrnas using expression data and disease-associated protein coding genes
 Inferring disease-associated lncrnas using expression data and disease-associated protein coding genes Xiaoyong Pan Center for non-coding RNA in Technology and Health, Department of Clinical Veterinary
Inferring disease-associated lncrnas using expression data and disease-associated protein coding genes Xiaoyong Pan Center for non-coding RNA in Technology and Health, Department of Clinical Veterinary
Instituto de Acuiculturade Torre de la Sal (CSIC), Castellón, Spain. Laboratoire de Physiologie et Génomique des Poissons (INRA), Rennes, France
 Mitochondria as key responders of metabolic and hormonal disturbances in fish. Meta-analysis flowchart of transcriptomicdata in gilthead sea bream (Sparus aurata) Josep Calduch-Giner 1, YannEchasseriau
Mitochondria as key responders of metabolic and hormonal disturbances in fish. Meta-analysis flowchart of transcriptomicdata in gilthead sea bream (Sparus aurata) Josep Calduch-Giner 1, YannEchasseriau
The Role of G-Protein Coupled Estrogen Receptor (GPER) in Early Neurite Development. Kyle Pemberton
 The Role of G-Protein Coupled Estrogen Receptor (GPER) in Early Neurite Development Kyle Pemberton Acknowledgement Dr. Xu Lab Members Brittany Mersman Nicki Patel Pallavi Mhaskar Jason Cocjin Committee
The Role of G-Protein Coupled Estrogen Receptor (GPER) in Early Neurite Development Kyle Pemberton Acknowledgement Dr. Xu Lab Members Brittany Mersman Nicki Patel Pallavi Mhaskar Jason Cocjin Committee
Correlation Networks
 QuickTime decompressor and a are needed to see this picture. Correlation Networks Analysis of Biological Networks April 24, 2010 Correlation Networks - Analysis of Biological Networks 1 Review We have
QuickTime decompressor and a are needed to see this picture. Correlation Networks Analysis of Biological Networks April 24, 2010 Correlation Networks - Analysis of Biological Networks 1 Review We have
MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question.
 Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which body fluid compartment contains high levels of K +, large anions, and proteins?
Exam Name MULTIPLE CHOICE. Choose the one alternative that best completes the statement or answers the question. 1) Which body fluid compartment contains high levels of K +, large anions, and proteins?
Compartmental Modelling
 Modelling Neurons Computing and the Brain Compartmental Modelling Spring 2010 2 1 Equivalent Electrical Circuit A patch of membrane is equivalent to an electrical circuit This circuit can be described
Modelling Neurons Computing and the Brain Compartmental Modelling Spring 2010 2 1 Equivalent Electrical Circuit A patch of membrane is equivalent to an electrical circuit This circuit can be described
Comparative Network Analysis
 Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Comparative Network Analysis BMI/CS 776 www.biostat.wisc.edu/bmi776/ Spring 2016 Anthony Gitter gitter@biostat.wisc.edu These slides, excluding third-party material, are licensed under CC BY-NC 4.0 by
Extension of cortical synaptic development distinguishes humans from chimpanzees and macaques
 Research Extension of cortical synaptic development distinguishes humans from chimpanzees and macaques Xiling Liu, 1,8 Mehmet Somel, 1,2,8 Lin Tang, 1,3,8 Zheng Yan, 1 Xi Jiang, 1 Song Guo, 1 Yuan Yuan,
Research Extension of cortical synaptic development distinguishes humans from chimpanzees and macaques Xiling Liu, 1,8 Mehmet Somel, 1,2,8 Lin Tang, 1,3,8 Zheng Yan, 1 Xi Jiang, 1 Song Guo, 1 Yuan Yuan,
A. Visceral and somatic divisions. B. Sympathetic and parasympathetic divisions. C. Central and peripheral divisions
 Ch 8: Neurons: Cellular and Network Properties, Part 1 Review of the Nervous System Objectives: Describe the Cells of the NS Explain the creation and propagation of an electrical signal in a nerve cell
Ch 8: Neurons: Cellular and Network Properties, Part 1 Review of the Nervous System Objectives: Describe the Cells of the NS Explain the creation and propagation of an electrical signal in a nerve cell
Evolution in Action: The Power of Mutation in E. coli
 Evolution in Action: The Power of Mutation in E. coli by Merle K. Heidemann, College of Natural Science, Emeritus Peter J. T. White, Lyman Briggs College James J. Smith, Lyman Briggs College Michigan State
Evolution in Action: The Power of Mutation in E. coli by Merle K. Heidemann, College of Natural Science, Emeritus Peter J. T. White, Lyman Briggs College James J. Smith, Lyman Briggs College Michigan State
Biosciences in the 21st century
 Biosciences in the 21st century Lecture 1: Neurons, Synapses, and Signaling Dr. Michael Burger Outline: 1. Why neuroscience? 2. The neuron 3. Action potentials 4. Synapses 5. Organization of the nervous
Biosciences in the 21st century Lecture 1: Neurons, Synapses, and Signaling Dr. Michael Burger Outline: 1. Why neuroscience? 2. The neuron 3. Action potentials 4. Synapses 5. Organization of the nervous
Zhiguang Huo 1, Chi Song 2, George Tseng 3. July 30, 2018
 Bayesian latent hierarchical model for transcriptomic meta-analysis to detect biomarkers with clustered meta-patterns of differential expression signals BayesMP Zhiguang Huo 1, Chi Song 2, George Tseng
Bayesian latent hierarchical model for transcriptomic meta-analysis to detect biomarkers with clustered meta-patterns of differential expression signals BayesMP Zhiguang Huo 1, Chi Song 2, George Tseng
