arm, we estimate the time to initial suppression using the nonparametric Kaplan Meier
|
|
|
- Asher Mitchell
- 5 years ago
- Views:
Transcription
1 Appendix 1: Technical details Let patients be indexed from 1 to 761 and index the treatment arms. is the time from randomization to first viral suppression, is the time from randomization to viral rebound, and is the time from randomization to censoring at loss to follow up, day 1012, or death. Define min, and min,. is an indicator that and is an indicator that. is the number of patients in arm. For each arm, we estimate the time to initial suppression using the nonparametric Kaplan Meier product limit estimator of the survivor function:, 1, /,, where indexes the distinct suppression event times,, is the number of suppression events at time,, and, is the number of patients at risk of initial suppression at time,. Similarly, we estimate the time from randomization to viral rebound as, 1, /,, where indexes the distinct rebound event times,, is the number of viral rebound events at time, and, is the number of patients at risk for viral rebound at time,. The probability that a patient with exposure has a suppressed viral load at time is the difference of these two curves:. Rather than choose a time point at which to estimate this probability, we can summarize this probability over followup interval [0, ] as the mean time spent in a state of suppression, estimated as. The mean time in a state of suppression can be compared between exposure groups as d, which can be calculated as the Riemann sum Δ. 1
2 where The variance of DS can be estimated as 1, :. 1 1, Where is an indicator for individual of being at risk of suppression at time u, : and is an indicator of being at risk of rebound at time u, :. 2
3 Appendix 2: SAS code ***************************************************************************** Code for viral suppression manuscript illustrating approach outlined by Gouskova 2014 start with dataset d with the following variables: t1: time from randomization to suppression d1: indicator of suppression at time t1 t2: time from randomization to rebound d2: indicator of rebound at t2 x: exposure indicator ****************************************************************************; *Estimate S(Suppression); proc phreg data=d noprint; model t1*d1(0)=; strata x; baseline out=sup(keep= x t1 s1) survival=s1; data sup; set sup; t=t1; *Estimate S(Rebound); proc phreg data=d noprint; model t2*d2(0)=; strata x; baseline out=reb(keep= x t2 s2) survival=s2; data reb; set reb; t=t2; *Make G(t) = S(Reb,t)-S(Sup,t); proc sort data=sup; by x t; proc sort data=reb; by x t; data times; do x=0 to 1; do t=0 to 1012; end; end; data g; merge sup reb times; by x t; keep x t s1 s2; data g; set g; by x t; retain holdr1 holdr2; if first.x then do; holdr1=0; holdr2=0; end; if s1=. then s1=holdr1; if s2=. then s2=holdr2; delta=s2-s1; holdr1=s1; holdr2=s2; data three; set g; if x=0; delta0=delta; keep t delta0; data four; set g; if x=1; delta1=delta; keep t delta1; data deltag; merge three four; by t; retain sum0 sum1 0; if first.b then do; sum0=0; sum1=0; end; sum0=sum0+delta0; sum1=sum1+delta1; if t=0 then do; g0=0; g1=0; end; if t>0 then do; g0=(sum0/t)*1012; g1=(sum1/t)*1012;end; delta=g1-g0; if t=1012 then proc print data=deltag noobs; var g0 g1 delta; title "Mean time in suppression and difference for x=0 and =1"; *Variance; proc phreg data=d atrisk noprint; model t1*d1(0)=; strata x; 3
4 output out=ars(keep=x t1 ys) num_left=ys ; baseline out=sup(keep=x t1 s) survival=s; proc sort data=ars; by x t1; data sup; set sup; by x t1; if last.t1;* get rid of duplicate record at t=0; data ars; set ars; by x t1; if first.t1; data shell; do x=0 to 1; do t1=1 to 1012;t2=t1; end; end; data sup2; merge sup ars; by x t1; retain s2; if s ne. then s2=s; else s2=s2; drop s; rename s2=s; data sup2; set sup2; by x t1; retain chm1 0 sm1 1 tm1 0 nsu; if first.x then do; nsu=0; chm1=0; sm1=1; end; if s>0 then ch=-log(s); nsu=round((ch-chm1)*ys,1);*above recreates y because SAS does not sst=s; t=t1; chm1=ch; sm1=s; keep x t nsu ys sst ; proc phreg data=d atrisk noprint; model t2*d2(0)=; strata x; output out=arr(keep=x t2 yr) num_left=yr ; baseline out=reb(keep=x t2 s) survival=s; proc sort data=arr; by x t2; data reb; set reb; by x t2; if last.t2;* get rid of duplicate record at t=0; data arr; set arr; by x t1; if first.t2; data shell; do x=0 to 1; do t1=1 to 1012;t2=t1; t=t1; end; end; data reb2; merge reb arr; by x t2; retain s2; if s ne. then s2=s; else s2=s2; drop s; rename s2=s; data reb2; set reb2; by x t2; retain chm1 0 sm1 1 tm1 0 nru; if first.x then do; nru=0; chm1=0; sm1=1; end; if s>0 then ch=-log(s); nru=round((ch-chm1)*yr,1);* recreates nru because SAS does not ssr=s; t=t2; chm1=ch; sm1=s; keep x t ssr nru yr ; data ltp4; merge reb2 sup2 shell; by x t; retain yr2 nru2 ys2 nsu2 ssr2 sst2; if yr ne. then yr2=yr; else yr2=yr2; 4
5 if nru ne. then nru2=nru; else nru2=0; if ys ne. then ys2=ys; else ys2=ys2; if nsu ne. then nsu2=nsu; else nsu2=0; if sst ne. then sst2=sst; else sst2=sst2; if ssr ne. then ssr2=ssr; else ssr2=ssr2; if last.t then drop yr nru ys nsu sst ssr t1 t2; rename yr2=yr nru2=nru ys2=ys nsu2=nsu ssr2=ssr sst2=sst; data e; set d; do t=0 to 1012; end; proc sort data=e; by x t; data lte; merge e(in=ine) ltp4 ; by x t ; if ine; proc sort data=lte; by x id t; data lte; set lte; if t<=t1 then ysui=1; else ysui=0; if t<=t2 then yrui=1; else yrui=0; if t1=t and d1=1 then nsi=1; else nsi=0; if t2=t and d2=1 then nri=1; else nri=0; if t<=1012 then *count number in each arm; proc sql; create table counts as select x, count(distinct(id)) as nx from lte group by x;quit; data lte2; merge counts lte; by x; proc sort data=lte2; by x id; *get components of Pi(t); data lte2; set lte2; by x id; q1u=(1/ys)*nsi; q2u=(ysui/ys**2)*nsu; q3u=(1/yr)*nri; q4u=(yrui/yr**2)*nru; *sum components of Pi(t) over time (until time t) and calculate Pi(t); data lte3; set lte2; by x id; retain q1t q2t q3t q4t; if first.id then do; q1t=0; q2t=0; q3t=0; q4t=0; end; q1t=q1t+q1u; q2t=q2t+q2u; q3t=q3t+q3u; q4t=q4t+q4u; pit=nx*sst*(q1t-q2t)-(nx*ssr*(q3t-q4t)); data lte4; set lte3; by x id; retain q6i; if first.id then q6i=0; q6i=pit+q6i; if t>0 then p=(q6i); if last.id then data lte5; set lte4; by x; retain q8; if first.x then q8=0; q7=p**2; q8=q7+q8; 5
6 if last.x then data lte6; set lte5 ; by x; retain v0 n0; if x=0 then do; v0=(1/nx**2)*q8; n0=nx;end; if x=1 then do; v1=(1/nx**2)*q8; n1=nx;end; if x=1 then keep v0 v1 n0 n1; data lte7; set lte6; se1=sqrt(v1); se0=sqrt(v0); sediff=sqrt(v0+v1); ods listing; *get estimate and 95% CI; data est; merge reb2 sup2 shell ;by x t; data est; set est;by x; retain sr st;if first.x then do; sr=1; st=1; end; if ssr ne. then sr=ssr; if sst ne. then st=sst; data estx1; set est; if x=1 ; gt1=sr-st; data estx0; set est; if x=0 ; gt0=sr-st; data estx; merge estx1 estx0 end=eof; by t; if last.t; retain sum0 0 sum1 0 g0 g1; sum0=sum(gt0,sum0); sum1=sum(gt1,sum1); if t=0 then do; g0=0; g1=0; end; else do; g0=sum0/(t)*1012; g1=sum1/(t)*1012; end; if eof then data estx; merge estx(keep=g0 g1) lte7(keep=sediff se1 se0); gtdiff=g1-g0; lcl=gtdiff-1.96*sediff; ucl=gtdiff+1.96*sediff; proc print data=estx; var g1 g0 gtdiff lcl ucl; title "Mean days suppressed for 3- and 4-drug arms with different and 95% CI"; 6
Multiple imputation to account for measurement error in marginal structural models
 Multiple imputation to account for measurement error in marginal structural models Supplementary material A. Standard marginal structural model We estimate the parameters of the marginal structural model
Multiple imputation to account for measurement error in marginal structural models Supplementary material A. Standard marginal structural model We estimate the parameters of the marginal structural model
Survival Analysis with Time- Dependent Covariates: A Practical Example. October 28, 2016 SAS Health Users Group Maria Eberg
 Survival Analysis with Time- Dependent Covariates: A Practical Example October 28, 2016 SAS Health Users Group Maria Eberg Outline Why use time-dependent covariates? Things to consider in definition of
Survival Analysis with Time- Dependent Covariates: A Practical Example October 28, 2016 SAS Health Users Group Maria Eberg Outline Why use time-dependent covariates? Things to consider in definition of
Advantages of Mixed-effects Regression Models (MRM; aka multilevel, hierarchical linear, linear mixed models) 1. MRM explicitly models individual
 Advantages of Mixed-effects Regression Models (MRM; aka multilevel, hierarchical linear, linear mixed models) 1. MRM explicitly models individual change across time 2. MRM more flexible in terms of repeated
Advantages of Mixed-effects Regression Models (MRM; aka multilevel, hierarchical linear, linear mixed models) 1. MRM explicitly models individual change across time 2. MRM more flexible in terms of repeated
Extensions of Cox Model for Non-Proportional Hazards Purpose
 PhUSE 2013 Paper SP07 Extensions of Cox Model for Non-Proportional Hazards Purpose Jadwiga Borucka, PAREXEL, Warsaw, Poland ABSTRACT Cox proportional hazard model is one of the most common methods used
PhUSE 2013 Paper SP07 Extensions of Cox Model for Non-Proportional Hazards Purpose Jadwiga Borucka, PAREXEL, Warsaw, Poland ABSTRACT Cox proportional hazard model is one of the most common methods used
Extensions of Cox Model for Non-Proportional Hazards Purpose
 PhUSE Annual Conference 2013 Paper SP07 Extensions of Cox Model for Non-Proportional Hazards Purpose Author: Jadwiga Borucka PAREXEL, Warsaw, Poland Brussels 13 th - 16 th October 2013 Presentation Plan
PhUSE Annual Conference 2013 Paper SP07 Extensions of Cox Model for Non-Proportional Hazards Purpose Author: Jadwiga Borucka PAREXEL, Warsaw, Poland Brussels 13 th - 16 th October 2013 Presentation Plan
Statistics 262: Intermediate Biostatistics Non-parametric Survival Analysis
 Statistics 262: Intermediate Biostatistics Non-parametric Survival Analysis Jonathan Taylor & Kristin Cobb Statistics 262: Intermediate Biostatistics p.1/?? Overview of today s class Kaplan-Meier Curve
Statistics 262: Intermediate Biostatistics Non-parametric Survival Analysis Jonathan Taylor & Kristin Cobb Statistics 262: Intermediate Biostatistics p.1/?? Overview of today s class Kaplan-Meier Curve
Let us use the term failure time to indicate the time of the event of interest in either a survival analysis or reliability analysis.
 10.2 Product-Limit (Kaplan-Meier) Method Let us use the term failure time to indicate the time of the event of interest in either a survival analysis or reliability analysis. Let T be a continuous random
10.2 Product-Limit (Kaplan-Meier) Method Let us use the term failure time to indicate the time of the event of interest in either a survival analysis or reliability analysis. Let T be a continuous random
A Survival Analysis of GMO vs Non-GMO Corn Hybrid Persistence Using Simulated Time Dependent Covariates in SAS
 Western Kentucky University From the SelectedWorks of Matt Bogard 2012 A Survival Analysis of GMO vs Non-GMO Corn Hybrid Persistence Using Simulated Time Dependent Covariates in SAS Matt Bogard, Western
Western Kentucky University From the SelectedWorks of Matt Bogard 2012 A Survival Analysis of GMO vs Non-GMO Corn Hybrid Persistence Using Simulated Time Dependent Covariates in SAS Matt Bogard, Western
Nonparametric Model Construction
 Nonparametric Model Construction Chapters 4 and 12 Stat 477 - Loss Models Chapters 4 and 12 (Stat 477) Nonparametric Model Construction Brian Hartman - BYU 1 / 28 Types of data Types of data For non-life
Nonparametric Model Construction Chapters 4 and 12 Stat 477 - Loss Models Chapters 4 and 12 (Stat 477) Nonparametric Model Construction Brian Hartman - BYU 1 / 28 Types of data Types of data For non-life
Analysis of Time-to-Event Data: Chapter 2 - Nonparametric estimation of functions of survival time
 Analysis of Time-to-Event Data: Chapter 2 - Nonparametric estimation of functions of survival time Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term
Analysis of Time-to-Event Data: Chapter 2 - Nonparametric estimation of functions of survival time Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term
Survival Regression Models
 Survival Regression Models David M. Rocke May 18, 2017 David M. Rocke Survival Regression Models May 18, 2017 1 / 32 Background on the Proportional Hazards Model The exponential distribution has constant
Survival Regression Models David M. Rocke May 18, 2017 David M. Rocke Survival Regression Models May 18, 2017 1 / 32 Background on the Proportional Hazards Model The exponential distribution has constant
9 Estimating the Underlying Survival Distribution for a
 9 Estimating the Underlying Survival Distribution for a Proportional Hazards Model So far the focus has been on the regression parameters in the proportional hazards model. These parameters describe the
9 Estimating the Underlying Survival Distribution for a Proportional Hazards Model So far the focus has been on the regression parameters in the proportional hazards model. These parameters describe the
ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES. Cox s regression analysis Time dependent explanatory variables
 ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES Cox s regression analysis Time dependent explanatory variables Henrik Ravn Bandim Health Project, Statens Serum Institut 4 November 2011 1 / 53
ADVANCED STATISTICAL ANALYSIS OF EPIDEMIOLOGICAL STUDIES Cox s regression analysis Time dependent explanatory variables Henrik Ravn Bandim Health Project, Statens Serum Institut 4 November 2011 1 / 53
MAS3301 / MAS8311 Biostatistics Part II: Survival
 MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
MAS3301 / MAS8311 Biostatistics Part II: Survival M. Farrow School of Mathematics and Statistics Newcastle University Semester 2, 2009-10 1 13 The Cox proportional hazards model 13.1 Introduction In the
Estimation for Modified Data
 Definition. Estimation for Modified Data 1. Empirical distribution for complete individual data (section 11.) An observation X is truncated from below ( left truncated) at d if when it is at or below d
Definition. Estimation for Modified Data 1. Empirical distribution for complete individual data (section 11.) An observation X is truncated from below ( left truncated) at d if when it is at or below d
Missing Data in Longitudinal Studies: Mixed-effects Pattern-Mixture and Selection Models
 Missing Data in Longitudinal Studies: Mixed-effects Pattern-Mixture and Selection Models Hedeker D & Gibbons RD (1997). Application of random-effects pattern-mixture models for missing data in longitudinal
Missing Data in Longitudinal Studies: Mixed-effects Pattern-Mixture and Selection Models Hedeker D & Gibbons RD (1997). Application of random-effects pattern-mixture models for missing data in longitudinal
USING THE SAS SYSTEM TO DEVELOP RISK PREDICTION MODELS FOR PATIENT RE-ADMISSION REDUCTION IN VAMCS. Issac Shams
 USING THE SAS SYSTEM TO DEVELOP RISK PREDICTION MODELS FOR PATIENT RE-ADMISSION REDUCTION IN VAMCS Issac Shams Veteran Engineering Resource Center (VERC)-VA-CASE Healthcare Systems Engineering Research
USING THE SAS SYSTEM TO DEVELOP RISK PREDICTION MODELS FOR PATIENT RE-ADMISSION REDUCTION IN VAMCS Issac Shams Veteran Engineering Resource Center (VERC)-VA-CASE Healthcare Systems Engineering Research
STAT Sample Problem: General Asymptotic Results
 STAT331 1-Sample Problem: General Asymptotic Results In this unit we will consider the 1-sample problem and prove the consistency and asymptotic normality of the Nelson-Aalen estimator of the cumulative
STAT331 1-Sample Problem: General Asymptotic Results In this unit we will consider the 1-sample problem and prove the consistency and asymptotic normality of the Nelson-Aalen estimator of the cumulative
Philosophy and Features of the mstate package
 Introduction Mathematical theory Practice Discussion Philosophy and Features of the mstate package Liesbeth de Wreede, Hein Putter Department of Medical Statistics and Bioinformatics Leiden University
Introduction Mathematical theory Practice Discussion Philosophy and Features of the mstate package Liesbeth de Wreede, Hein Putter Department of Medical Statistics and Bioinformatics Leiden University
Testing Indirect Effects for Lower Level Mediation Models in SAS PROC MIXED
 Testing Indirect Effects for Lower Level Mediation Models in SAS PROC MIXED Here we provide syntax for fitting the lower-level mediation model using the MIXED procedure in SAS as well as a sas macro, IndTest.sas
Testing Indirect Effects for Lower Level Mediation Models in SAS PROC MIXED Here we provide syntax for fitting the lower-level mediation model using the MIXED procedure in SAS as well as a sas macro, IndTest.sas
Multi-state models: prediction
 Department of Medical Statistics and Bioinformatics Leiden University Medical Center Course on advanced survival analysis, Copenhagen Outline Prediction Theory Aalen-Johansen Computational aspects Applications
Department of Medical Statistics and Bioinformatics Leiden University Medical Center Course on advanced survival analysis, Copenhagen Outline Prediction Theory Aalen-Johansen Computational aspects Applications
Chapter 4 Regression Models
 23.August 2010 Chapter 4 Regression Models The target variable T denotes failure time We let x = (x (1),..., x (m) ) represent a vector of available covariates. Also called regression variables, regressors,
23.August 2010 Chapter 4 Regression Models The target variable T denotes failure time We let x = (x (1),..., x (m) ) represent a vector of available covariates. Also called regression variables, regressors,
IP WEIGHTING AND MARGINAL STRUCTURAL MODELS (CHAPTER 12) BIOS IPW and MSM
 IP WEIGHTING AND MARGINAL STRUCTURAL MODELS (CHAPTER 12) BIOS 776 1 12 IPW and MSM IP weighting and marginal structural models ( 12) Outline 12.1 The causal question 12.2 Estimating IP weights via modeling
IP WEIGHTING AND MARGINAL STRUCTURAL MODELS (CHAPTER 12) BIOS 776 1 12 IPW and MSM IP weighting and marginal structural models ( 12) Outline 12.1 The causal question 12.2 Estimating IP weights via modeling
Part III. Hypothesis Testing. III.1. Log-rank Test for Right-censored Failure Time Data
 1 Part III. Hypothesis Testing III.1. Log-rank Test for Right-censored Failure Time Data Consider a survival study consisting of n independent subjects from p different populations with survival functions
1 Part III. Hypothesis Testing III.1. Log-rank Test for Right-censored Failure Time Data Consider a survival study consisting of n independent subjects from p different populations with survival functions
Exercises. (a) Prove that m(t) =
 Exercises 1. Lack of memory. Verify that the exponential distribution has the lack of memory property, that is, if T is exponentially distributed with parameter λ > then so is T t given that T > t for
Exercises 1. Lack of memory. Verify that the exponential distribution has the lack of memory property, that is, if T is exponentially distributed with parameter λ > then so is T t given that T > t for
Survival Analysis. Stat 526. April 13, 2018
 Survival Analysis Stat 526 April 13, 2018 1 Functions of Survival Time Let T be the survival time for a subject Then P [T < 0] = 0 and T is a continuous random variable The Survival function is defined
Survival Analysis Stat 526 April 13, 2018 1 Functions of Survival Time Let T be the survival time for a subject Then P [T < 0] = 0 and T is a continuous random variable The Survival function is defined
Multivariate Survival Analysis
 Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
β j = coefficient of x j in the model; β = ( β1, β2,
 Regression Modeling of Survival Time Data Why regression models? Groups similar except for the treatment under study use the nonparametric methods discussed earlier. Groups differ in variables (covariates)
Regression Modeling of Survival Time Data Why regression models? Groups similar except for the treatment under study use the nonparametric methods discussed earlier. Groups differ in variables (covariates)
4. Comparison of Two (K) Samples
 4. Comparison of Two (K) Samples K=2 Problem: compare the survival distributions between two groups. E: comparing treatments on patients with a particular disease. Z: Treatment indicator, i.e. Z = 1 for
4. Comparison of Two (K) Samples K=2 Problem: compare the survival distributions between two groups. E: comparing treatments on patients with a particular disease. Z: Treatment indicator, i.e. Z = 1 for
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials
 Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Assessing the effect of a partly unobserved, exogenous, binary time-dependent covariate on -APPENDIX-
 Assessing the effect of a partly unobserved, exogenous, binary time-dependent covariate on survival probabilities using generalised pseudo-values Ulrike Pötschger,2, Harald Heinzl 2, Maria Grazia Valsecchi
Assessing the effect of a partly unobserved, exogenous, binary time-dependent covariate on survival probabilities using generalised pseudo-values Ulrike Pötschger,2, Harald Heinzl 2, Maria Grazia Valsecchi
PRINCIPAL COMPONENTS ANALYSIS (PCA)
 PRINCIPAL COMPONENTS ANALYSIS (PCA) Introduction PCA is considered an exploratory technique that can be used to gain a better understanding of the interrelationships between variables. PCA is performed
PRINCIPAL COMPONENTS ANALYSIS (PCA) Introduction PCA is considered an exploratory technique that can be used to gain a better understanding of the interrelationships between variables. PCA is performed
Package threg. August 10, 2015
 Package threg August 10, 2015 Title Threshold Regression Version 1.0.3 Date 2015-08-10 Author Tao Xiao Maintainer Tao Xiao Depends R (>= 2.10), survival, Formula Fit a threshold regression
Package threg August 10, 2015 Title Threshold Regression Version 1.0.3 Date 2015-08-10 Author Tao Xiao Maintainer Tao Xiao Depends R (>= 2.10), survival, Formula Fit a threshold regression
UNIVERSITY OF CALIFORNIA, SAN DIEGO
 UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
Kaplan-Meier in SAS. filename foo url "http://math.unm.edu/~james/small.txt"; data small; infile foo firstobs=2; input time censor; run;
 Kaplan-Meier in SAS filename foo url "http://math.unm.edu/~james/small.txt"; data small; infile foo firstobs=2; input time censor; run; proc print data=small; run; proc lifetest data=small plots=survival;
Kaplan-Meier in SAS filename foo url "http://math.unm.edu/~james/small.txt"; data small; infile foo firstobs=2; input time censor; run; proc print data=small; run; proc lifetest data=small plots=survival;
Product-limit estimators of the survival function with left or right censored data
 Product-limit estimators of the survival function with left or right censored data 1 CREST-ENSAI Campus de Ker-Lann Rue Blaise Pascal - BP 37203 35172 Bruz cedex, France (e-mail: patilea@ensai.fr) 2 Institut
Product-limit estimators of the survival function with left or right censored data 1 CREST-ENSAI Campus de Ker-Lann Rue Blaise Pascal - BP 37203 35172 Bruz cedex, France (e-mail: patilea@ensai.fr) 2 Institut
Mixed Models Lecture Notes By Dr. Hanford page 199 More Statistics& SAS Tutorial at
 Mixed Models Lecture Notes By Dr. Hanford page 199 Variance Balance Cross-Over Designs Variance balance cross-over designs are designs where all treatment contrasts have the same precision and all carry-over
Mixed Models Lecture Notes By Dr. Hanford page 199 Variance Balance Cross-Over Designs Variance balance cross-over designs are designs where all treatment contrasts have the same precision and all carry-over
Beyond GLM and likelihood
 Stat 6620: Applied Linear Models Department of Statistics Western Michigan University Statistics curriculum Core knowledge (modeling and estimation) Math stat 1 (probability, distributions, convergence
Stat 6620: Applied Linear Models Department of Statistics Western Michigan University Statistics curriculum Core knowledge (modeling and estimation) Math stat 1 (probability, distributions, convergence
In contrast, parametric techniques (fitting exponential or Weibull, for example) are more focussed, can handle general covariates, but require
 Chapter 5 modelling Semi parametric We have considered parametric and nonparametric techniques for comparing survival distributions between different treatment groups. Nonparametric techniques, such as
Chapter 5 modelling Semi parametric We have considered parametric and nonparametric techniques for comparing survival distributions between different treatment groups. Nonparametric techniques, such as
Chapter 4 Fall Notations: t 1 < t 2 < < t D, D unique death times. d j = # deaths at t j = n. Y j = # at risk /alive at t j = n
 Bios 323: Applied Survival Analysis Qingxia (Cindy) Chen Chapter 4 Fall 2012 4.2 Estimators of the survival and cumulative hazard functions for RC data Suppose X is a continuous random failure time with
Bios 323: Applied Survival Analysis Qingxia (Cindy) Chen Chapter 4 Fall 2012 4.2 Estimators of the survival and cumulative hazard functions for RC data Suppose X is a continuous random failure time with
Point and Interval Estimation II Bios 662
 Point and Interval Estimation II Bios 662 Michael G. Hudgens, Ph.D. mhudgens@bios.unc.edu http://www.bios.unc.edu/ mhudgens 2006-09-13 17:17 BIOS 662 1 Point and Interval Estimation II Nonparametric CI
Point and Interval Estimation II Bios 662 Michael G. Hudgens, Ph.D. mhudgens@bios.unc.edu http://www.bios.unc.edu/ mhudgens 2006-09-13 17:17 BIOS 662 1 Point and Interval Estimation II Nonparametric CI
18.465, further revised November 27, 2012 Survival analysis and the Kaplan Meier estimator
 18.465, further revised November 27, 2012 Survival analysis and the Kaplan Meier estimator 1. Definitions Ordinarily, an unknown distribution function F is estimated by an empirical distribution function
18.465, further revised November 27, 2012 Survival analysis and the Kaplan Meier estimator 1. Definitions Ordinarily, an unknown distribution function F is estimated by an empirical distribution function
Some Relatively Simple Ways to Get Good Answers
 OBSERVATIONAL DATA ANALYSIS COMPETITION: HETEROGENEOUS RESPONSE CHALLENGE Some Relatively Simple Ways to Get Good Answers Xiaochun Li, Ph.D. Associate Professor Division of Biostatistics Indiana University
OBSERVATIONAL DATA ANALYSIS COMPETITION: HETEROGENEOUS RESPONSE CHALLENGE Some Relatively Simple Ways to Get Good Answers Xiaochun Li, Ph.D. Associate Professor Division of Biostatistics Indiana University
Comparison of Two Population Means
 Comparison of Two Population Means Esra Akdeniz March 15, 2015 Independent versus Dependent (paired) Samples We have independent samples if we perform an experiment in two unrelated populations. We have
Comparison of Two Population Means Esra Akdeniz March 15, 2015 Independent versus Dependent (paired) Samples We have independent samples if we perform an experiment in two unrelated populations. We have
A Regression Model For Recurrent Events With Distribution Free Correlation Structure
 A Regression Model For Recurrent Events With Distribution Free Correlation Structure J. Pénichoux(1), A. Latouche(2), T. Moreau(1) (1) INSERM U780 (2) Université de Versailles, EA2506 ISCB - 2009 - Prague
A Regression Model For Recurrent Events With Distribution Free Correlation Structure J. Pénichoux(1), A. Latouche(2), T. Moreau(1) (1) INSERM U780 (2) Université de Versailles, EA2506 ISCB - 2009 - Prague
A SAS Macro to Find the Best Fitted Model for Repeated Measures Using PROC MIXED
 Paper AD18 A SAS Macro to Find the Best Fitted Model for Repeated Measures Using PROC MIXED Jingjing Wang, U.S. Army Institute of Surgical Research, Ft. Sam Houston, TX ABSTRACT Specifying an appropriate
Paper AD18 A SAS Macro to Find the Best Fitted Model for Repeated Measures Using PROC MIXED Jingjing Wang, U.S. Army Institute of Surgical Research, Ft. Sam Houston, TX ABSTRACT Specifying an appropriate
Constrained estimation for binary and survival data
 Constrained estimation for binary and survival data Jeremy M. G. Taylor Yong Seok Park John D. Kalbfleisch Biostatistics, University of Michigan May, 2010 () Constrained estimation May, 2010 1 / 43 Outline
Constrained estimation for binary and survival data Jeremy M. G. Taylor Yong Seok Park John D. Kalbfleisch Biostatistics, University of Michigan May, 2010 () Constrained estimation May, 2010 1 / 43 Outline
for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park
 Threshold Regression for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park MLTLEE@UMD.EDU Outline The proportional hazards (PH) model is widely used in analyzing time-to-event data.
Threshold Regression for Time-to-event Data Mei-Ling Ting Lee University of Maryland, College Park MLTLEE@UMD.EDU Outline The proportional hazards (PH) model is widely used in analyzing time-to-event data.
REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520
 REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520 Department of Statistics North Carolina State University Presented by: Butch Tsiatis, Department of Statistics, NCSU
REGRESSION ANALYSIS FOR TIME-TO-EVENT DATA THE PROPORTIONAL HAZARDS (COX) MODEL ST520 Department of Statistics North Carolina State University Presented by: Butch Tsiatis, Department of Statistics, NCSU
Statistical Methods III Statistics 212. Problem Set 2 - Answer Key
 Statistical Methods III Statistics 212 Problem Set 2 - Answer Key 1. (Analysis to be turned in and discussed on Tuesday, April 24th) The data for this problem are taken from long-term followup of 1423
Statistical Methods III Statistics 212 Problem Set 2 - Answer Key 1. (Analysis to be turned in and discussed on Tuesday, April 24th) The data for this problem are taken from long-term followup of 1423
Adjusting Analyses of Survey Results using a Predicted Probability of Response Rob Gately, INNOVUS: Epidemiology, Waltham, MA
 PharmaSUG2011 Paper HS07 Adjusting Analyses of Survey Results using a Predicted Probability of Response Rob Gately, INNOVUS: Epidemiology, Waltham, MA ABSTRACT When conducting surveys, an issue that is
PharmaSUG2011 Paper HS07 Adjusting Analyses of Survey Results using a Predicted Probability of Response Rob Gately, INNOVUS: Epidemiology, Waltham, MA ABSTRACT When conducting surveys, an issue that is
Estimating Causal Effects of Organ Transplantation Treatment Regimes
 Estimating Causal Effects of Organ Transplantation Treatment Regimes David M. Vock, Jeffrey A. Verdoliva Boatman Division of Biostatistics University of Minnesota July 31, 2018 1 / 27 Hot off the Press
Estimating Causal Effects of Organ Transplantation Treatment Regimes David M. Vock, Jeffrey A. Verdoliva Boatman Division of Biostatistics University of Minnesota July 31, 2018 1 / 27 Hot off the Press
Mapping Participants to the Closest Medical Center
 SESUG Paper RV-192-2017 Mapping Participants to the Closest Medical Center David Franklin, QuintilesIMS, Cambridge, MA ABSTRACT How far are patients from Clinics? That was the question which was asked
SESUG Paper RV-192-2017 Mapping Participants to the Closest Medical Center David Franklin, QuintilesIMS, Cambridge, MA ABSTRACT How far are patients from Clinics? That was the question which was asked
S The Over-Reliance on the Central Limit Theorem
 S04-2008 The Over-Reliance on the Central Limit Theorem Abstract The objective is to demonstrate the theoretical and practical implication of the central limit theorem. The theorem states that as n approaches
S04-2008 The Over-Reliance on the Central Limit Theorem Abstract The objective is to demonstrate the theoretical and practical implication of the central limit theorem. The theorem states that as n approaches
CIMAT Taller de Modelos de Capture y Recaptura Known Fate Survival Analysis
 CIMAT Taller de Modelos de Capture y Recaptura 2010 Known Fate urvival Analysis B D BALANCE MODEL implest population model N = λ t+ 1 N t Deeper understanding of dynamics can be gained by identifying variation
CIMAT Taller de Modelos de Capture y Recaptura 2010 Known Fate urvival Analysis B D BALANCE MODEL implest population model N = λ t+ 1 N t Deeper understanding of dynamics can be gained by identifying variation
Chapter 7: Hypothesis testing
 Chapter 7: Hypothesis testing Hypothesis testing is typically done based on the cumulative hazard function. Here we ll use the Nelson-Aalen estimate of the cumulative hazard. The survival function is used
Chapter 7: Hypothesis testing Hypothesis testing is typically done based on the cumulative hazard function. Here we ll use the Nelson-Aalen estimate of the cumulative hazard. The survival function is used
Logistic regression model for survival time analysis using time-varying coefficients
 Logistic regression model for survival time analysis using time-varying coefficients Accepted in American Journal of Mathematical and Management Sciences, 2016 Kenichi SATOH ksatoh@hiroshima-u.ac.jp Research
Logistic regression model for survival time analysis using time-varying coefficients Accepted in American Journal of Mathematical and Management Sciences, 2016 Kenichi SATOH ksatoh@hiroshima-u.ac.jp Research
Time Series Forecasting Methods
 Time Series Forecasting Methods Nate Derby Statis Pro Data Analytics Seattle, WA, USA Edmonton SAS Users Group, 11/13/09 Nate Derby Time Series Forecasting Methods 1 / 43 Outline Introduction 1 Introduction
Time Series Forecasting Methods Nate Derby Statis Pro Data Analytics Seattle, WA, USA Edmonton SAS Users Group, 11/13/09 Nate Derby Time Series Forecasting Methods 1 / 43 Outline Introduction 1 Introduction
Sample size re-estimation in clinical trials. Dealing with those unknowns. Chris Jennison. University of Kyoto, January 2018
 Sample Size Re-estimation in Clinical Trials: Dealing with those unknowns Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj University of Kyoto,
Sample Size Re-estimation in Clinical Trials: Dealing with those unknowns Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj University of Kyoto,
SAS Analysis Examples Replication C11
 SAS Analysis Examples Replication C11 * SAS Analysis Examples Replication for ASDA 2nd Edition * Berglund April 2017 * Chapter 11 ; libname d "P:\ASDA 2\Data sets\hrs 2012\HRS 2006_2012 Longitudinal File\"
SAS Analysis Examples Replication C11 * SAS Analysis Examples Replication for ASDA 2nd Edition * Berglund April 2017 * Chapter 11 ; libname d "P:\ASDA 2\Data sets\hrs 2012\HRS 2006_2012 Longitudinal File\"
Results of a simulation of modeling and nonparametric methodology for count data (from patient diary) in drug studies
 Results of a simulation of modeling and nonparametric methodology for count data (from patient diary) in drug studies Erhard Quebe-Fehling Workshop Stat. Methoden für korrelierte Daten Bochum 24-Nov-06
Results of a simulation of modeling and nonparametric methodology for count data (from patient diary) in drug studies Erhard Quebe-Fehling Workshop Stat. Methoden für korrelierte Daten Bochum 24-Nov-06
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Power and Sample Size Bios 662
 Power and Sample Size Bios 662 Michael G. Hudgens, Ph.D. mhudgens@bios.unc.edu http://www.bios.unc.edu/ mhudgens 2008-10-31 14:06 BIOS 662 1 Power and Sample Size Outline Introduction One sample: continuous
Power and Sample Size Bios 662 Michael G. Hudgens, Ph.D. mhudgens@bios.unc.edu http://www.bios.unc.edu/ mhudgens 2008-10-31 14:06 BIOS 662 1 Power and Sample Size Outline Introduction One sample: continuous
Statistics 262: Intermediate Biostatistics Regression & Survival Analysis
 Statistics 262: Intermediate Biostatistics Regression & Survival Analysis Jonathan Taylor & Kristin Cobb Statistics 262: Intermediate Biostatistics p.1/?? Introduction This course is an applied course,
Statistics 262: Intermediate Biostatistics Regression & Survival Analysis Jonathan Taylor & Kristin Cobb Statistics 262: Intermediate Biostatistics p.1/?? Introduction This course is an applied course,
Survival Analysis. 732G34 Statistisk analys av komplexa data. Krzysztof Bartoszek
 Survival Analysis 732G34 Statistisk analys av komplexa data Krzysztof Bartoszek (krzysztof.bartoszek@liu.se) 10, 11 I 2018 Department of Computer and Information Science Linköping University Survival analysis
Survival Analysis 732G34 Statistisk analys av komplexa data Krzysztof Bartoszek (krzysztof.bartoszek@liu.se) 10, 11 I 2018 Department of Computer and Information Science Linköping University Survival analysis
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL
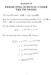 Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
Lecture 6 PREDICTING SURVIVAL UNDER THE PH MODEL The Cox PH model: λ(t Z) = λ 0 (t) exp(β Z). How do we estimate the survival probability, S z (t) = S(t Z) = P (T > t Z), for an individual with covariates
Introduction to Statistical Analysis
 Introduction to Statistical Analysis Changyu Shen Richard A. and Susan F. Smith Center for Outcomes Research in Cardiology Beth Israel Deaconess Medical Center Harvard Medical School Objectives Descriptive
Introduction to Statistical Analysis Changyu Shen Richard A. and Susan F. Smith Center for Outcomes Research in Cardiology Beth Israel Deaconess Medical Center Harvard Medical School Objectives Descriptive
Lecture 4 - Survival Models
 Lecture 4 - Survival Models Survival Models Definition and Hazards Kaplan Meier Proportional Hazards Model Estimation of Survival in R GLM Extensions: Survival Models Survival Models are a common and incredibly
Lecture 4 - Survival Models Survival Models Definition and Hazards Kaplan Meier Proportional Hazards Model Estimation of Survival in R GLM Extensions: Survival Models Survival Models are a common and incredibly
Lecture 9. Statistics Survival Analysis. Presented February 23, Dan Gillen Department of Statistics University of California, Irvine
 Statistics 255 - Survival Analysis Presented February 23, 2016 Dan Gillen Department of Statistics University of California, Irvine 9.1 Survival analysis involves subjects moving through time Hazard may
Statistics 255 - Survival Analysis Presented February 23, 2016 Dan Gillen Department of Statistics University of California, Irvine 9.1 Survival analysis involves subjects moving through time Hazard may
Models for Ordinal Response Data
 Models for Ordinal Response Data Robin High Department of Biostatistics Center for Public Health University of Nebraska Medical Center Omaha, Nebraska Recommendations Analyze numerical data with a statistical
Models for Ordinal Response Data Robin High Department of Biostatistics Center for Public Health University of Nebraska Medical Center Omaha, Nebraska Recommendations Analyze numerical data with a statistical
Summarizing and Displaying Measurement Data/Understanding and Comparing Distributions
 Summarizing and Displaying Measurement Data/Understanding and Comparing Distributions Histograms, Mean, Median, Five-Number Summary and Boxplots, Standard Deviation Thought Questions 1. If you were to
Summarizing and Displaying Measurement Data/Understanding and Comparing Distributions Histograms, Mean, Median, Five-Number Summary and Boxplots, Standard Deviation Thought Questions 1. If you were to
^l,2-, 3. f 2_ ' >v., 4. b/-e. of- fee. -^3-5- I70(fa = )&*>$)
 & ^l,2-, 3 f 2_ ' s\ >v., 4 b/-e of- fee O = )&*>$) -^3-5- I70(fa \J QQQ ~E - V-?' = f rff ^ if -j- 41 -Ofc -Ofc v OH OH ST/VT H SoJU^, 55-3 2. ^ T_ 2, _ ^ * ^3W^(M7 2. X 2.! Z. d-f- S-5- /V Z 33?> 5YJ
& ^l,2-, 3 f 2_ ' s\ >v., 4 b/-e of- fee O = )&*>$) -^3-5- I70(fa \J QQQ ~E - V-?' = f rff ^ if -j- 41 -Ofc -Ofc v OH OH ST/VT H SoJU^, 55-3 2. ^ T_ 2, _ ^ * ^3W^(M7 2. X 2.! Z. d-f- S-5- /V Z 33?> 5YJ
Power and Sample Size Calculations with the Additive Hazards Model
 Journal of Data Science 10(2012), 143-155 Power and Sample Size Calculations with the Additive Hazards Model Ling Chen, Chengjie Xiong, J. Philip Miller and Feng Gao Washington University School of Medicine
Journal of Data Science 10(2012), 143-155 Power and Sample Size Calculations with the Additive Hazards Model Ling Chen, Chengjie Xiong, J. Philip Miller and Feng Gao Washington University School of Medicine
Statistical Methods for Alzheimer s Disease Studies
 Statistical Methods for Alzheimer s Disease Studies Rebecca A. Betensky, Ph.D. Department of Biostatistics, Harvard T.H. Chan School of Public Health July 19, 2016 1/37 OUTLINE 1 Statistical collaborations
Statistical Methods for Alzheimer s Disease Studies Rebecca A. Betensky, Ph.D. Department of Biostatistics, Harvard T.H. Chan School of Public Health July 19, 2016 1/37 OUTLINE 1 Statistical collaborations
Survival analysis in R
 Survival analysis in R Niels Richard Hansen This note describes a few elementary aspects of practical analysis of survival data in R. For further information we refer to the book Introductory Statistics
Survival analysis in R Niels Richard Hansen This note describes a few elementary aspects of practical analysis of survival data in R. For further information we refer to the book Introductory Statistics
PASS Sample Size Software. Poisson Regression
 Chapter 870 Introduction Poisson regression is used when the dependent variable is a count. Following the results of Signorini (99), this procedure calculates power and sample size for testing the hypothesis
Chapter 870 Introduction Poisson regression is used when the dependent variable is a count. Following the results of Signorini (99), this procedure calculates power and sample size for testing the hypothesis
Machine Learning. Module 3-4: Regression and Survival Analysis Day 2, Asst. Prof. Dr. Santitham Prom-on
 Machine Learning Module 3-4: Regression and Survival Analysis Day 2, 9.00 16.00 Asst. Prof. Dr. Santitham Prom-on Department of Computer Engineering, Faculty of Engineering King Mongkut s University of
Machine Learning Module 3-4: Regression and Survival Analysis Day 2, 9.00 16.00 Asst. Prof. Dr. Santitham Prom-on Department of Computer Engineering, Faculty of Engineering King Mongkut s University of
Model Building Chap 5 p251
 Model Building Chap 5 p251 Models with one qualitative variable, 5.7 p277 Example 4 Colours : Blue, Green, Lemon Yellow and white Row Blue Green Lemon Insects trapped 1 0 0 1 45 2 0 0 1 59 3 0 0 1 48 4
Model Building Chap 5 p251 Models with one qualitative variable, 5.7 p277 Example 4 Colours : Blue, Green, Lemon Yellow and white Row Blue Green Lemon Insects trapped 1 0 0 1 45 2 0 0 1 59 3 0 0 1 48 4
Pattern Structures for Risk Group Identification
 Pattern Structures for Risk Group Identification Natalia V. Korepanova and Sergei O. Kuznetsov National Research University Higher School of Economics, Moscow, Russia, nkorepanova@hse.ru, skuznetsov@hse.ru
Pattern Structures for Risk Group Identification Natalia V. Korepanova and Sergei O. Kuznetsov National Research University Higher School of Economics, Moscow, Russia, nkorepanova@hse.ru, skuznetsov@hse.ru
Analysis of Survival Data Using Cox Model (Continuous Type)
 Australian Journal of Basic and Alied Sciences, 7(0): 60-607, 03 ISSN 99-878 Analysis of Survival Data Using Cox Model (Continuous Type) Khawla Mustafa Sadiq Department of Mathematics, Education College,
Australian Journal of Basic and Alied Sciences, 7(0): 60-607, 03 ISSN 99-878 Analysis of Survival Data Using Cox Model (Continuous Type) Khawla Mustafa Sadiq Department of Mathematics, Education College,
Definitions and examples Simple estimation and testing Regression models Goodness of fit for the Cox model. Recap of Part 1. Per Kragh Andersen
 Recap of Part 1 Per Kragh Andersen Section of Biostatistics, University of Copenhagen DSBS Course Survival Analysis in Clinical Trials January 2018 1 / 65 Overview Definitions and examples Simple estimation
Recap of Part 1 Per Kragh Andersen Section of Biostatistics, University of Copenhagen DSBS Course Survival Analysis in Clinical Trials January 2018 1 / 65 Overview Definitions and examples Simple estimation
Interpolation {x,y} Data with Suavity. Peter K. Ott Forest Analysis & Inventory Branch BC Ministry of FLNRO Victoria, BC
 Interpolation {x,y} Data with Suavity Peter K. Ott Forest Analysis & Inventory Branch BC Ministry of FLNRO Victoria, BC Peter.Ott@gov.bc.ca 1 The Goal Given a set of points: x i, y i, i = 1,2,, n find
Interpolation {x,y} Data with Suavity Peter K. Ott Forest Analysis & Inventory Branch BC Ministry of FLNRO Victoria, BC Peter.Ott@gov.bc.ca 1 The Goal Given a set of points: x i, y i, i = 1,2,, n find
3003 Cure. F. P. Treasure
 3003 Cure F. P. reasure November 8, 2000 Peter reasure / November 8, 2000/ Cure / 3003 1 Cure A Simple Cure Model he Concept of Cure A cure model is a survival model where a fraction of the population
3003 Cure F. P. reasure November 8, 2000 Peter reasure / November 8, 2000/ Cure / 3003 1 Cure A Simple Cure Model he Concept of Cure A cure model is a survival model where a fraction of the population
MAS361. MAS361 1 Turn Over SCHOOL OF MATHEMATICS AND STATISTICS. Medical Statistics
 t r r t r r t r t s s MAS361 SCHOOL OF MATHEMATICS AND STATISTICS Medical Statistics Autumn Semester 2015 16 2 hours t s 2 r t t 1 t t r t t r s t rs t2 r t s q st s r t r t r 2 t st s rs q st s rr2 q
t r r t r r t r t s s MAS361 SCHOOL OF MATHEMATICS AND STATISTICS Medical Statistics Autumn Semester 2015 16 2 hours t s 2 r t t 1 t t r t t r s t rs t2 r t s q st s r t r t r 2 t st s rs q st s rr2 q
Supply Forecasting via Survival Curve Estimation
 Supply Forecasting via Survival Curve Estimation Gary D. Knott, Ph.D. Civilized Software Inc. 12109 Heritage Park Circle Silver Spring, MD 20906 USA email:csi@civilized.com URL:www.civilized.com Tel: (301)
Supply Forecasting via Survival Curve Estimation Gary D. Knott, Ph.D. Civilized Software Inc. 12109 Heritage Park Circle Silver Spring, MD 20906 USA email:csi@civilized.com URL:www.civilized.com Tel: (301)
Analysis of Incomplete Non-Normal Longitudinal Lipid Data
 Analysis of Incomplete Non-Normal Longitudinal Lipid Data Jiajun Liu*, Devan V. Mehrotra, Xiaoming Li, and Kaifeng Lu 2 Merck Research Laboratories, PA/NJ 2 Forrest Laboratories, NY *jiajun_liu@merck.com
Analysis of Incomplete Non-Normal Longitudinal Lipid Data Jiajun Liu*, Devan V. Mehrotra, Xiaoming Li, and Kaifeng Lu 2 Merck Research Laboratories, PA/NJ 2 Forrest Laboratories, NY *jiajun_liu@merck.com
Statistical Inference and Methods
 Department of Mathematics Imperial College London d.stephens@imperial.ac.uk http://stats.ma.ic.ac.uk/ das01/ 31st January 2006 Part VI Session 6: Filtering and Time to Event Data Session 6: Filtering and
Department of Mathematics Imperial College London d.stephens@imperial.ac.uk http://stats.ma.ic.ac.uk/ das01/ 31st January 2006 Part VI Session 6: Filtering and Time to Event Data Session 6: Filtering and
Survival models and health sequences
 Survival models and health sequences Walter Dempsey University of Michigan July 27, 2015 Survival Data Problem Description Survival data is commonplace in medical studies, consisting of failure time information
Survival models and health sequences Walter Dempsey University of Michigan July 27, 2015 Survival Data Problem Description Survival data is commonplace in medical studies, consisting of failure time information
Multimodal Deep Learning for Predicting Survival from Breast Cancer
 Multimodal Deep Learning for Predicting Survival from Breast Cancer Heather Couture Deep Learning Journal Club Nov. 16, 2016 Outline Background on tumor histology & genetic data Background on survival
Multimodal Deep Learning for Predicting Survival from Breast Cancer Heather Couture Deep Learning Journal Club Nov. 16, 2016 Outline Background on tumor histology & genetic data Background on survival
1 The problem of survival analysis
 1 The problem of survival analysis Survival analysis concerns analyzing the time to the occurrence of an event. For instance, we have a dataset in which the times are 1, 5, 9, 20, and 22. Perhaps those
1 The problem of survival analysis Survival analysis concerns analyzing the time to the occurrence of an event. For instance, we have a dataset in which the times are 1, 5, 9, 20, and 22. Perhaps those
Part III Measures of Classification Accuracy for the Prediction of Survival Times
 Part III Measures of Classification Accuracy for the Prediction of Survival Times Patrick J Heagerty PhD Department of Biostatistics University of Washington 102 ISCB 2010 Session Three Outline Examples
Part III Measures of Classification Accuracy for the Prediction of Survival Times Patrick J Heagerty PhD Department of Biostatistics University of Washington 102 ISCB 2010 Session Three Outline Examples
6 Single Sample Methods for a Location Parameter
 6 Single Sample Methods for a Location Parameter If there are serious departures from parametric test assumptions (e.g., normality or symmetry), nonparametric tests on a measure of central tendency (usually
6 Single Sample Methods for a Location Parameter If there are serious departures from parametric test assumptions (e.g., normality or symmetry), nonparametric tests on a measure of central tendency (usually
Model Selection Procedures
 Model Selection Procedures Statistics 135 Autumn 2005 Copyright c 2005 by Mark E. Irwin Model Selection Procedures Consider a regression setting with K potential predictor variables and you wish to explore
Model Selection Procedures Statistics 135 Autumn 2005 Copyright c 2005 by Mark E. Irwin Model Selection Procedures Consider a regression setting with K potential predictor variables and you wish to explore
SAS/STAT 13.1 User s Guide. The PHREG Procedure
 SAS/STAT 13.1 User s Guide The PHREG Procedure This document is an individual chapter from SAS/STAT 13.1 User s Guide. The correct bibliographic citation for the complete manual is as follows: SAS Institute
SAS/STAT 13.1 User s Guide The PHREG Procedure This document is an individual chapter from SAS/STAT 13.1 User s Guide. The correct bibliographic citation for the complete manual is as follows: SAS Institute
STA 303H1F: Two-way Analysis of Variance Practice Problems
 STA 303H1F: Two-way Analysis of Variance Practice Problems 1. In the Pygmalion example from lecture, why are the average scores of the platoon used as the response variable, rather than the scores of the
STA 303H1F: Two-way Analysis of Variance Practice Problems 1. In the Pygmalion example from lecture, why are the average scores of the platoon used as the response variable, rather than the scores of the
ST745: Survival Analysis: Cox-PH!
 ST745: Survival Analysis: Cox-PH! Eric B. Laber Department of Statistics, North Carolina State University April 20, 2015 Rien n est plus dangereux qu une idee, quand on n a qu une idee. (Nothing is more
ST745: Survival Analysis: Cox-PH! Eric B. Laber Department of Statistics, North Carolina State University April 20, 2015 Rien n est plus dangereux qu une idee, quand on n a qu une idee. (Nothing is more
Description Syntax for predict Menu for predict Options for predict Remarks and examples Methods and formulas References Also see
 Title stata.com stcrreg postestimation Postestimation tools for stcrreg Description Syntax for predict Menu for predict Options for predict Remarks and examples Methods and formulas References Also see
Title stata.com stcrreg postestimation Postestimation tools for stcrreg Description Syntax for predict Menu for predict Options for predict Remarks and examples Methods and formulas References Also see
Paired comparisons. We assume that
 To compare to methods, A and B, one can collect a sample of n pairs of observations. Pair i provides two measurements, Y Ai and Y Bi, one for each method: If we want to compare a reaction of patients to
To compare to methods, A and B, one can collect a sample of n pairs of observations. Pair i provides two measurements, Y Ai and Y Bi, one for each method: If we want to compare a reaction of patients to
Analysis of Time-to-Event Data: Chapter 6 - Regression diagnostics
 Analysis of Time-to-Event Data: Chapter 6 - Regression diagnostics Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/25 Residuals for the
Analysis of Time-to-Event Data: Chapter 6 - Regression diagnostics Steffen Unkel Department of Medical Statistics University Medical Center Göttingen, Germany Winter term 2018/19 1/25 Residuals for the
