Comparison of Multidimensional Scaling and Principal Component Analysis of Interspecific Variation in Bacteria*
|
|
|
- Alisha Stafford
- 6 years ago
- Views:
Transcription
1 ANNALS OF CLINICAL AND LABORATORY SCIENCE, Vol. 18., No. 6 Copyright 1988, Institute for Clinical Science, Inc. Comparison of Multidimensional Scaling and Principal Component Analysis of Interspecific Variation in Bacteria* DAVID A. LACHER, M.D., and EDWARD D. O DONNELL, Ph.D. Department of Pathology, Medical College of Ohio, Toledo, OH ABSTRACT Multidimensional scaling (MDS) and principal component analysis (PCA) were applied to bacterial taxonomy. The biochemical profiles of 42 isolates consisting of four species of Enterobacteriaceae were used. Both MDS and PCA use proximity measures such as the correlation coefficient or Euclidean distance to generate a spatial configuration (map) of points in multidimensional space where distances between points reflect the similarity among isolates. Multidimensional scaling and principal component analysis were able to discriminate organisms in two dimensions. The test components of the MDS and PCA factors (derived variables composed of linear combination of biochemical tests) were different for a two-dimensional solution. Introduction but these approaches assume statistically independent variables which do not usually occur.6,7 Discriminant analysis techniques have also been applied to bacterial taxonomy but may require reductions of large variable sets prior to analysis.3,17 Principal component analysis (PCA), a m ultivariate tech n iq u e which examines the correlations b e tween tests and reduces variable sets, has been applied to taxonomic studies of bacteria.1,4,10,11,14 Multidimensional scaling (MDS) can also be used to reduce variable sets and has been applied in the social sciences, but rarely to laboratory m edicine.12,13 Different populations of microorganisms can be represented by geometric models.9 A data set of p isolates analyzed by n tests can be visualized as a cloud of /88/ $01.20 Institute for Clinical Science, Inc. The system atic analysis of variation among bacterial species can help construct useful num eric taxonom ic schemes to classify and identify organisms. Bacterial isolates are commonly identified by a profile of biochemical tests. By exam ining th e correlation among these tests, statistical techniques can be useful in bacterial taxonomy. Many statistical techniques have been used to classify microorganisms. Bayesian and relative likelihood models can be applied to identify bacterial isolates, Address rep rin t requests to David A. Lacher, M.D., Departm ent of Pathology, Medical College of Ohio, C.S. #10008, Toledo, OH
2 456 M D S A N D PCA ANALYSIS OF BACTERIA p points in n-dimensional hyperellipsoid space. For efficiency, it is desirable to reduce the dimensional space (reduce the number of tests) while preserving the maximum amount o f information used to discriminate different bacterial groups. Geometrically, multidimensional scaling and principal component analysis seek a lower dimensional representation of the data set while retaining, as much as p o s s ib le, th e o rig in a l d ista n ce between points (the original underlying relationship o f data). Both M DS and PCA can map the p oin ts in higher dim ensional space onto lower dim ensional space. W hen moving to a lower dimensional representation, MDS and PCA generate factors which are linear combinations of variables that reflect features of the data. Tests which significantly influence a factor will be highly correlated and help to interpret the factor Ṫhere are several differences between MDS and PCA.8,12,16 Principal component analysis starts with a correlation matrix, while multidimensional scaling can start with an inter-subject distance matrix or a correlation matrix. The MDS m ethod is based on distances among points w hile PCA is based on angles among vectors. Also, principal component analysis is based on the general linear model, but multidimensional scaling has no such restrictive assumption. In addition, M DS may result in a lower dimensional solution than PCA. However, m ultidim ensional scaling cannot handle large data sets efficiently. A survey of the literature revealed no application of multidimensional scaling to bacterial taxonomy or identification. In this study, multidimensional scaling and previously used principal component analysis are applied to the analysis of variation among four closely related species of E n terobacteriaceae. Both MDS and PCA are compared for their ability to reduce the variable set (reduce dimensionality), to discriminate between species in various dim ensional space, and to explain the test components of the generated factors. Materials and Methods D a ta S e t Multidimensional scaling and principal component analysis were applied to a small test data set consisting of a biochemical profile of 42 selected isolates obtained from various anatom ic sites with the majority from urine cultures and endotracheal aspirates. The isolates consisted of four species of the Enterobacteriaceae family: Serratia marcescens (n = 8), E nterobacter aerogenes (n = 10), Klebsiella pneumoniae (n = 12), and Enterobacter cloacae (n = 12). These isolates were chosen because they are relatively closely related species within the family Enterobacteriaceae. There were no repeat isolates from any patient. The isolates were identified by biochemical tests incorporated into the gramnegative GNI card analyzed on the VITEK AMS instrument.* There are 30 wells in GNI card which contain 29 biochemical broths and one growth control broth. Test results were negative or postive and were coded as 0 and 1, respectively. Ten of 30 tests had uniformly positive or negative results for all isolates and were discarded since there was no discrimination among isolates. In addition, there were tests which correlated perfectly with other tests and this redundancy allowed the elimination of three tests. The 17 remaining biochem ical test results for the 42 isolates can be seen in table I. The biochem ical broths contained DP-300 (DP3 containing 2,4,4 trichloro-2 -hydroxydiphenylether), urea *McDonnell Douglas Health Systems Co., Hazelwood, MO
3 MDS AND PCA ANALYSIS OF BACTERIA 457 T ABLE I Biochemical Pattern Test* S p e c i e s DP3 URE I N O A R A A D O M A L C O U A R G A C E TDA LY S E S C P X B O N P O R N P L I L A C S e r r a t i a m a r c e s c e n s E n t e r o b a c t e r a e r o g e n s K l e b s i e l l a p n e u m o n i a e E n t e r o b a c t e r c l o a c a e * Test codes are 0 for a negative and 1 for a positive result. DP3- DP-300(DP3 containing 2,4,4' trichloro-2 '-hydroxydiphenylether) URE- Urea INO- Inositol AKA- L-arabinose ADO- Adonitol MAL- Malonate COU- p-coumaric ARG- Arginine ACE- Acetamide TDA- Tryptophan LYS- Lysine ESC- Esculin PXB- polymyxin B ONP- o-nitrophenyl-ß-d-galactopyranoside ORN- Ornithine PLI- Plant indican LAC- Lactose (U R E), in o sito l (IN O ), L -arabinose (ARA), a d o n ito l (A D O ), m alon ate (MAL), p-coum aric (COU), arginine (ARG), acetamide (ACE), tryptophan (TDA), lysine (LYS), esculin (ESC), polym yxin B (PXB), o-nitrophenyl-f$-d - galactopyranoside (ONP), ornithine (ORN), plant indican (PLI), and lactose (LAC). All isolates had com plete biochemical profiles. S t a t is t ic a l A n a l y sis M ultidimensional scaling was performed using the SAS ALSCAL program.18 The biochemical test data were
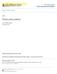
4 458 LACHER AND O DONNELL used to create a Euclidean distance between each pair of isolates using the following formula: du = -v/x (Xir Xjr)2 where i=l dy = Euclidean distance R = biochemical test number Xir = test result for the ith isolate for the rth test X,r = test result for the jth isolate for the rth test The range of the Euclidean distance was from zero (indicating isolates with identical biochemical patterns) to a maximum of the square of the number of biochemical tests (indicating a nonidentical p a tte r n ). T he E u c lid e a n d ista n c e betw een each isolate was used in the SAS ALSCAL program. Dim ensional scores produced by MDS for the isolates were plotted to assess the separation of the four species in various dimensional space (figures 1 and 2). Multiple linear regression, using the dimensional scores as dependent variables and biochemical tests as independent variables, was performed using the BMDP LR program5 to establish the standard regression coefficients for each dim ension (table II). These regression coefficients identify the tests which had significant positive and negative influences on each dimension. Also, stepwise linear regression identified those tests which contributed most significantly in predicting the dimension scores. x xx a a i Principal component analysis, a form of factor analysis, was performed on the biochemical data using the SAS FAC TOR program.15 Unrotated and orthogonal VARIMAX-rotated PCA were performed. Rotations were perform ed to simplify the interpretation of each factor. Factor scores were then plotted for each isolate (figures I and 3). Random bacterial isolates from a single population should produce normally (Gaussian) distributed PCA scores.4 The PCA factor pattern (table III) was examined to identify the tests which had salient loadings (strong influence) for each factor. Results and Discussion The biochemical profiles of isolates of S. marcescens, E. aerogenes, K. pneumoniae, and E. cloacae are seen in table I. Serratia m arcescens is identified if DP-300 or polymyxim B are positive or if ONPG or arabinose are negative. If inositol, adonitol or plant indican are negative or if arginine is positive, E. cloacae is identified. A positive reaction for acetam ide would indicate E. aerogenes. These tests are important in differentiatin g th e four sp ecies and w ou ld be expected to be influential test components of the factors in PCA or dimensions in MDS. Multidimensional scaling was applied to the biochemical pattern of the isolates. Young s S-stress measure (a goodness of fit function) was reduced significantly from a one-dimensional (0.2799) to a two-dimensional solution (0.2355), MDS -2-1 U i a X X X XX PCA F ig u r e 1. One-dimensional plot of multidimensional scaling (MDS) dimensional and principal component analysis (PCA) factor scores for S. marcescens ( ), E. aerogenes (A), K. pneumoniae (X) and E. cloacae ( ) isolates.
5 MDS AND PCA ANALYSIS O F BACTERIA T g 0'5 "', F igure 2. Twoin dimensional plot of multi- 5 dimensional scaling for S A marcescens ( ), E. aero- genes (A), K. pneumoniae (X), and E. cloacae ( ) o.s 4- x isolates DIMENSION 1 but the stress measure did not change significantly for a three-dim ensional solution (0.2348). This indicated that a two-dimensional MDS solution was optimal. Multiple linear regression, using the MDS dimensional coordinates as dependent variables and tests as independent variables, was performed to interpret the tw o-dim ensional solution (table II). Lysine, p-coumaric, and inositol had a strong positive influence (regression T A B L E II Two-dimensional Multidimensional Scaling Standard Regression Coefficients Tests D i m e n s i o n 1 D i m e n s i o n 2 D P ( D P 3 c o n t a i n i n g 2,4,4' t r i c hloro-2' -h y d r o x y d i p h e n y l e t h e r ) Urea I n o s i t o l L - a r a b i n o s e A d o n i t o l M a l o n a t e p - c o u m a r i c A r g i n i n e A c e t a m i d e T r y p t o p h a n Lysine E s c u l i n p o l y m y x i n B o -nitrophenyl- 3-D - g a l a c t o p y r a n o s i d e Orni thine P l a n t i n d i c a n L a c t o s e coefficients >0.10) and malonate, lactate and L-arabinose had a strong negative effect on dimension 1. For dimension 2, ornithine had a positive influence and urea, adonitol, lactate, esculin, and ONPG had a significant negative influence. Stepwise linear regression showed that L-arabinose and lysine best predicted the first dimensional scores. L- arabinose was useful in identifying S. marcescens, and lysine had the highest negative p ercentage for E. cloacae. Ornithine and esculin were the best predictors of dimension 2 scores. Ornithine helped differentiate K. pneumoniae and a negative esculin reaction indicated E. cloacae. Principal component analysis was also applied to the biochemical test reactions of the isolates. The first five factors had eigenvalues greater than one (considered significant factors) and explained 79.4 percent o f the variance. A scree plot (eigenvalue versus factor number) indicated that the first two factors (accounting for 54.2 percent of total variance) could adequately represent the information contained in the biochemical database. This was surprising because principal component analysis was developed to handle continuous, normally distributed variables, unlike the binary qualitative
6 460 LACHER AND O DONNELL F igure 3. T w o - dimensional plot of princip a l c o m p o n e n t fa c to r scores for S. marcescens ( ), E. aerogenes (A), K. pneum oniae (X) and E. cloacae ( ) isolates. FACTOR 1 biochemical test data analyzed in this study. Factoring data where variables are dichotomous may lead to spurious extra factors,8 but PC A efficiently reduced the biochemical data to two dimensions. The orthogonal VARIMAX rotation was applied to the principal component analysis since the unrotated PCA could not simply identify significant tests contributing to the two factor solution (table III). In general, tests with larger positive or negative coefficients should be the important tests which differentiate species. For factor one, L-arabinose, lactate, ONPG, and malonate had strong positive effects (factor loading > 0.50) and p- coumaric, polymix-b and DP-300 had salient negative loadings (less than 0.50). Inositol, esculin, lysine, adonitol and plant-indican had strong positive and ornithine had a strong negative influence on factor 2 scores. The communality estimates, the proportion of the variation explained by each test for the PCA solution, suggested that L-arabinose followed by p-coumaric and lysine were important tests in differentiating the four species. A positive p-coumaric T A B L E III Two-dimensional Varimax-rotated Factor Pattern Test Factor 1 Factor 2 DP-300(DP3 containing 2,4,4' trichloro-2 -hydroxydiphenylether) Urea Inositoi L-arabinose Adonitoi Malonate p-coumaric Arginine Acetamide Tryptophan Lusine Esculin polymyxin B o-nitrophenyl-ß-d -galactopyranoside Orni thine Plant indican Lactose
7 reaction was seen most often in S. marcescens. The influence of tests on the MDS dimensions and PCA factors were different. Multidimensional scaling and principal component analysis were compared for their ability to graphically separate S. m arcescens, E. aerogenes, E. cloacae and K. pneumoniae. The MDS and PCA scores were plotted for a one-dim ensional solution (figure 1). M ultidimensional scaling appeared to be the mirror image of the PCA one-dimensional solution and, hence, compared well to principal component analysis. Serratia marcesc e n s and E. clo a ca e w ere w e ll separated, but K. pneumoniae and E. aerogenes could not be fully differentiated for the one-dimensional solution. T he m u ltid im en sional scores and VARIMAX-rotated factor scores were then compared for a two-dimensional solution (figures 2 and 3). Both MDS and PCA separated the four sp ecies o f Enterobacteriaceae in two-dimensional space. E nterobacter aerogenes and K. pneumoniae appeared to be phylogenetically similar with respect to their biochemical test pattern. When compared to S. marcescens, E. cloacae was more closely related to E. aerogenes and K. pneumoniae. The relationships of these four species based on biochemical data was comparable to DNA relatedness groupings seen among Enterobacteriaceae species.2 Since the MDS plot could not be rotated to match the PCA plot, it appeared that the MDS and PCA twodimensional solutions were different. For MDS, a positive score for dimension 1 indicated S. marcescens. To get a positive score for dimension 1, lysine, p- coumaric and inositol should have a positive reaction and L-arabinose should have a negative result as indicated by the multiple linear regression analysis (vide supra). Serratia marcescens best fit this reaction pattern (table I). A negative score (less than 1.0) for dimension 2 MDS AND PCA ANALYSIS OF BACTERIA 461 indicated E. cloacae. Enterobacter aerogenes and K. pneum oniae w ere separated by the second MDS dimension. E n tero b a cter aerogen es had higher dimension 2 scores than K. pneumoniae. The interpretation of the tw o-dim ensional principal component analysis plot was different from the MDS graph. A negative factor score identified the S. marcescens group, but could not separate it from E. cloacae as the M DS dimension 1 did. The PCA dimension 2 helped separate K. pneumoniae, E. aero g en es and E. c lo a c a e w h ich had decreasing scores, respectively. Principal com ponent analysis and multidimensional scaling were applied to a profile of biochemical tests to reduce the dimensionality of the variable (test) set and to discriminate betw een four bacterial species. Both MDS and PCA could represent the data w ell in two dimensions but gave different interpretations of the dimensions. The analysis of a data set consisting of a large number of different isolates would increase the number of dimensions required to differentiate the organisms. In this study, the identities of the isolates were known a priori. Discriminate analysis of MDS or PCA scores of a training set of known isolates can be used to classify unknown isolates. Other pattern recognition techniques such as SIMCA, K-nearest neighbor and Bayesian analysis can discriminate bacterial isolates.1 Also, MDS or PCA followed by cluster analysis techniques can be used to group (a priori) unknown microorganisms.1,14 Multidimensional scaling and principal component analysis are used to reduce a large number of variables to a few significant variables in order to simplify data analysis. References I. Boyd, J. C., Lewis, J. W., Marr, J. J., Harper, A. M., and Kowalski, B. R.: Effect of atypical
8 462 LACHER AND O DONNELL antibiotic resistance on microorganism identification by pattern recognition. J. Clin. Microbiol. 8: , Brenner, D. J.: Family I. Enterobacteriaceae, Krieg, N. R., Holt J. G. Bergey s Manual of Systematic Bacteriology, volume 1. Baltimore, The Williams & Wilkins Co., 1984, pp DARLAND, G.: Discriminant analysis of antibiotic susceptibility as a means of bacterial identification. J. Clin. Microbiol. 2: , DARLAND, G.: Principal component analysis of infraspecific variation in bacteria. Appl. Microbiol. 30: , D ix o n, W. J., ed.: BMDP Statistical Software Manual. Berkley, CA, University of California Press, DYBOWSKI, W. and FRANKLIN, D. A.: Conditional probability and the identification of bacteria: A pilot study. J. Gen. Microbiol. 54: , F r i e d m a n, R. B. and M a c L o w ry, J.: Computer identification of bacteria on the basis of their antibiotic susceptibility patterns. Appl. Microbiol. 26: , GORSUCH, R. L.: Factor Analysis, 2nd ed. Hillsdale, NJ, Lawrence Earlbaum Associates, Inc., Gyllenberg, H. G.: A model for computer identification of micro-organisms. J. Gen. Microbiol. 39: , Hill, L. R., Silvestri, L. G., Ihm, P., Farchi, G., and Lenclani, P.: Automatic classification of staphylococci by principal component analysis and a gradient method. J. Bacteriol. 89: , Hornstein, M. J., Jupeau, A. M., Scavizzi, M. R., Phillippon, A. M., a n d Grimont, P. A. D.: In vitro susceptibilities of 126 clinical isolates of Yersinia enterocolitica to 21 p-lactam antibiotics. Amtimicrob. Agents Chem other. 27: , K r u s k a l, J. B. and W ISH, M.: Multidimensional Scaling. Beverly Hills, CA, Sage Publications, Inc., LaCHER, D. A.: In terp retatio n of laboratory results using multidimensional scaling and principal component analysis. Ann. Clin. Lab. Sci. 17: , Quadung, C. and Hospkins, J. W.: Evaluation of tests and grouping of cultures by a two-stage principal component method. Can. J. Microbiol. 13: , SAS User s Guide: Statistics, 5th ed Cary, NC, SAS Institute, Inc., Schiffman, S., Reynolds, M., and Young, F.: In tro d u ctio n to M ultidim ensional Scaling. Orlando, FL, Academic Press, Sielaff, B. H., Matsen, J. M., and McKie, J. E.: Novel approach to bacterial identification that uses the Autobac system. J. Clin. Microbiol. 25: , Young, F. W. and Lewyckyi, R.: The ALSCAL procedure, SUGI Supplemental Library User s Guide, 5th ed. Cary, NC, SAS Institute, Inc., 1986, pp
Interpretation of Laboratory Results Using M ultidimensional Scaling and Principal C om ponent Analysis*
 ANNALS OF CLINICAL AND LABORATORY SCIENCE, Vol. 17, No. 6 Copyright 1987, Institute for Clinical Science, Inc. Interpretation of Laboratory Results Using M ultidimensional Scaling and Principal C om ponent
ANNALS OF CLINICAL AND LABORATORY SCIENCE, Vol. 17, No. 6 Copyright 1987, Institute for Clinical Science, Inc. Interpretation of Laboratory Results Using M ultidimensional Scaling and Principal C om ponent
Clinical Laboratory Evaluation of the AutoMicrobic System
 JOURNAL OF CLINICAL MICROBIOLOGY, OCt. 1981, p. 370-375 0095-1 137/81/100370-06$02.00/0 Vol. 14, No. 4 Clinical Laboratory Evaluation of the AutoMicrobic System Enterobacteriaceae Biochemical Card JAMES
JOURNAL OF CLINICAL MICROBIOLOGY, OCt. 1981, p. 370-375 0095-1 137/81/100370-06$02.00/0 Vol. 14, No. 4 Clinical Laboratory Evaluation of the AutoMicrobic System Enterobacteriaceae Biochemical Card JAMES
Evaluation of the Modified Micro-ID System for Identification
 JOURNAL OF CLINICAL MICROBIOLOGY, Oct. 1979, p. 454-458 0095-1 137/79/10-0454/05$02.00/0 Vol. 10, No. 4 Evaluation of the Modified Micro-ID System for Identification of Enterobacteriaceae WILLIAM J. BUESCHING,'
JOURNAL OF CLINICAL MICROBIOLOGY, Oct. 1979, p. 454-458 0095-1 137/79/10-0454/05$02.00/0 Vol. 10, No. 4 Evaluation of the Modified Micro-ID System for Identification of Enterobacteriaceae WILLIAM J. BUESCHING,'
FACTOR ANALYSIS AND MULTIDIMENSIONAL SCALING
 FACTOR ANALYSIS AND MULTIDIMENSIONAL SCALING Vishwanath Mantha Department for Electrical and Computer Engineering Mississippi State University, Mississippi State, MS 39762 mantha@isip.msstate.edu ABSTRACT
FACTOR ANALYSIS AND MULTIDIMENSIONAL SCALING Vishwanath Mantha Department for Electrical and Computer Engineering Mississippi State University, Mississippi State, MS 39762 mantha@isip.msstate.edu ABSTRACT
Numerical Diagnostic Key for the Identification of Enterobacteriaceae
 APPLIED MICROBIOLOGY, Jan. 1972, p. 108-112 Copyright 0 1972 American Society for Microbiology Vol. 23, No. 1 Printed in U.SA. Numerical Diagnostic Key for the Identification of Enterobacteriaceae HERMAN
APPLIED MICROBIOLOGY, Jan. 1972, p. 108-112 Copyright 0 1972 American Society for Microbiology Vol. 23, No. 1 Printed in U.SA. Numerical Diagnostic Key for the Identification of Enterobacteriaceae HERMAN
ENTEROBACTER AEROGENES UNKNOWN BACTERIA FLOW CHART UNKNOWN LAB REPORT, MICROBIOLOGY ENTEROBACTER AEROGENES
 ENTEROBACTER AEROGENES UNKNOWN BACTERIA PDF UNKNOWN LAB REPORT, MICROBIOLOGY ENTEROBACTER AEROGENES IDENTIFICATION OF AN UNKNOWN BACTERIAL SPECIES OF 1 / 5 2 / 5 3 / 5 enterobacter aerogenes unknown bacteria
ENTEROBACTER AEROGENES UNKNOWN BACTERIA PDF UNKNOWN LAB REPORT, MICROBIOLOGY ENTEROBACTER AEROGENES IDENTIFICATION OF AN UNKNOWN BACTERIAL SPECIES OF 1 / 5 2 / 5 3 / 5 enterobacter aerogenes unknown bacteria
1 A factor can be considered to be an underlying latent variable: (a) on which people differ. (b) that is explained by unknown variables
 1 A factor can be considered to be an underlying latent variable: (a) on which people differ (b) that is explained by unknown variables (c) that cannot be defined (d) that is influenced by observed variables
1 A factor can be considered to be an underlying latent variable: (a) on which people differ (b) that is explained by unknown variables (c) that cannot be defined (d) that is influenced by observed variables
5. Discriminant analysis
 5. Discriminant analysis We continue from Bayes s rule presented in Section 3 on p. 85 (5.1) where c i is a class, x isap-dimensional vector (data case) and we use class conditional probability (density
5. Discriminant analysis We continue from Bayes s rule presented in Section 3 on p. 85 (5.1) where c i is a class, x isap-dimensional vector (data case) and we use class conditional probability (density
System in Comparison with the API 20E System
 JOURNAL OF CLINICAL MICROBIOLOGY, July 983, p. 2835 Vol. 8, No. 009537/83/0702808$02.00/0 Copyright C 983, American Society for Microbiology Evaluation of the Updated MS2 Bacterial Identification System
JOURNAL OF CLINICAL MICROBIOLOGY, July 983, p. 2835 Vol. 8, No. 009537/83/0702808$02.00/0 Copyright C 983, American Society for Microbiology Evaluation of the Updated MS2 Bacterial Identification System
Robot Image Credit: Viktoriya Sukhanova 123RF.com. Dimensionality Reduction
 Robot Image Credit: Viktoriya Sukhanova 13RF.com Dimensionality Reduction Feature Selection vs. Dimensionality Reduction Feature Selection (last time) Select a subset of features. When classifying novel
Robot Image Credit: Viktoriya Sukhanova 13RF.com Dimensionality Reduction Feature Selection vs. Dimensionality Reduction Feature Selection (last time) Select a subset of features. When classifying novel
Introduction to Machine Learning. PCA and Spectral Clustering. Introduction to Machine Learning, Slides: Eran Halperin
 1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
1 Introduction to Machine Learning PCA and Spectral Clustering Introduction to Machine Learning, 2013-14 Slides: Eran Halperin Singular Value Decomposition (SVD) The singular value decomposition (SVD)
Principal Component Analysis, A Powerful Scoring Technique
 Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
Principal Component Analysis, A Powerful Scoring Technique George C. J. Fernandez, University of Nevada - Reno, Reno NV 89557 ABSTRACT Data mining is a collection of analytical techniques to uncover new
A Model for Computer Identification of Micro-organisms
 J. gen, Microbial. (1965), 39, 401405 Printed.in Great Britain 401 A Model for Computer Identification of Micro-organisms BY H. G. GYLLENBERG Department of Microbiology, Ulziversity of Helsinki, Finland
J. gen, Microbial. (1965), 39, 401405 Printed.in Great Britain 401 A Model for Computer Identification of Micro-organisms BY H. G. GYLLENBERG Department of Microbiology, Ulziversity of Helsinki, Finland
7. Variable extraction and dimensionality reduction
 7. Variable extraction and dimensionality reduction The goal of the variable selection in the preceding chapter was to find least useful variables so that it would be possible to reduce the dimensionality
7. Variable extraction and dimensionality reduction The goal of the variable selection in the preceding chapter was to find least useful variables so that it would be possible to reduce the dimensionality
PCA & ICA. CE-717: Machine Learning Sharif University of Technology Spring Soleymani
 PCA & ICA CE-717: Machine Learning Sharif University of Technology Spring 2015 Soleymani Dimensionality Reduction: Feature Selection vs. Feature Extraction Feature selection Select a subset of a given
PCA & ICA CE-717: Machine Learning Sharif University of Technology Spring 2015 Soleymani Dimensionality Reduction: Feature Selection vs. Feature Extraction Feature selection Select a subset of a given
Effect of Methods of Platelet Resuspension on Stored Platelets
 ANNALS O F CLINICAL AND LABORATORY S C IEN C E, Vol. 14, No. 5 Copyright 1984, Institute for Clinical Science, Inc. Effect of Methods of Platelet Resuspension on Stored Platelets THOMAS KIRALY, M.A., S.B.B.
ANNALS O F CLINICAL AND LABORATORY S C IEN C E, Vol. 14, No. 5 Copyright 1984, Institute for Clinical Science, Inc. Effect of Methods of Platelet Resuspension on Stored Platelets THOMAS KIRALY, M.A., S.B.B.
identification system
 J Clin Pathol 1988;41:910-914 Evaluation of the Microbact-24E bacterial identification system JULIA M LING, Y-W HUT, G L FRENCH Department ofmicrobiology, The Chinese University of Hong Kong, The Prince
J Clin Pathol 1988;41:910-914 Evaluation of the Microbact-24E bacterial identification system JULIA M LING, Y-W HUT, G L FRENCH Department ofmicrobiology, The Chinese University of Hong Kong, The Prince
Chemometrics. Classification of Mycobacteria by HPLC and Pattern Recognition. Application Note. Abstract
 12-1214 Chemometrics Application Note Classification of Mycobacteria by HPLC and Pattern Recognition Abstract Mycobacteria include a number of respiratory and non-respiratory pathogens for humans, such
12-1214 Chemometrics Application Note Classification of Mycobacteria by HPLC and Pattern Recognition Abstract Mycobacteria include a number of respiratory and non-respiratory pathogens for humans, such
Linear Dimensionality Reduction
 Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Principal Component Analysis 3 Factor Analysis
Outline Hong Chang Institute of Computing Technology, Chinese Academy of Sciences Machine Learning Methods (Fall 2012) Outline Outline I 1 Introduction 2 Principal Component Analysis 3 Factor Analysis
Dimensionality Reduction for Exponential Family Data
 Dimensionality Reduction for Exponential Family Data Yoonkyung Lee* Department of Statistics The Ohio State University *joint work with Andrew Landgraf July 2-6, 2018 Computational Strategies for Large-Scale
Dimensionality Reduction for Exponential Family Data Yoonkyung Lee* Department of Statistics The Ohio State University *joint work with Andrew Landgraf July 2-6, 2018 Computational Strategies for Large-Scale
EKOLOGIE EN SYSTEMATIEK. T h is p a p e r n o t to be c i t e d w ith o u t p r i o r r e f e r e n c e to th e a u th o r. PRIMARY PRODUCTIVITY.
 EKOLOGIE EN SYSTEMATIEK Ç.I.P.S. MATHEMATICAL MODEL OF THE POLLUTION IN NORT H SEA. TECHNICAL REPORT 1971/O : B i o l. I T h is p a p e r n o t to be c i t e d w ith o u t p r i o r r e f e r e n c e to
EKOLOGIE EN SYSTEMATIEK Ç.I.P.S. MATHEMATICAL MODEL OF THE POLLUTION IN NORT H SEA. TECHNICAL REPORT 1971/O : B i o l. I T h is p a p e r n o t to be c i t e d w ith o u t p r i o r r e f e r e n c e to
Key words: Staphylococci, Classification, Antibiotic-susceptibility, Opportunistic infection
 Key words: Staphylococci, Classification, Antibiotic-susceptibility, Opportunistic infection Table 1. Species classification of staphylococcal isolates from clinical specimens Figures in parentheses indicate
Key words: Staphylococci, Classification, Antibiotic-susceptibility, Opportunistic infection Table 1. Species classification of staphylococcal isolates from clinical specimens Figures in parentheses indicate
Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA
 Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA Principle Components Analysis: Uses one group of variables (we will call this X) In
Principle Components Analysis (PCA) Relationship Between a Linear Combination of Variables and Axes Rotation for PCA Principle Components Analysis: Uses one group of variables (we will call this X) In
THE IDENTIFICATION OF TWO UNKNOWN BACTERIA AFUA WILLIAMS BIO 3302 TEST TUBE 3 PROF. N. HAQUE 5/14/18
 THE IDENTIFICATION OF TWO UNKNOWN BACTERIA AFUA WILLIAMS BIO 3302 TEST TUBE 3 PROF. N. HAQUE Introduction: The identification of bacteria is important in order for us to differentiate one microorganism
THE IDENTIFICATION OF TWO UNKNOWN BACTERIA AFUA WILLIAMS BIO 3302 TEST TUBE 3 PROF. N. HAQUE Introduction: The identification of bacteria is important in order for us to differentiate one microorganism
Drift Reduction For Metal-Oxide Sensor Arrays Using Canonical Correlation Regression And Partial Least Squares
 Drift Reduction For Metal-Oxide Sensor Arrays Using Canonical Correlation Regression And Partial Least Squares R Gutierrez-Osuna Computer Science Department, Wright State University, Dayton, OH 45435,
Drift Reduction For Metal-Oxide Sensor Arrays Using Canonical Correlation Regression And Partial Least Squares R Gutierrez-Osuna Computer Science Department, Wright State University, Dayton, OH 45435,
L11: Pattern recognition principles
 L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
L11: Pattern recognition principles Bayesian decision theory Statistical classifiers Dimensionality reduction Clustering This lecture is partly based on [Huang, Acero and Hon, 2001, ch. 4] Introduction
C o r p o r a t e l i f e i n A n c i e n t I n d i a e x p r e s s e d i t s e l f
 C H A P T E R I G E N E S I S A N D GROWTH OF G U IL D S C o r p o r a t e l i f e i n A n c i e n t I n d i a e x p r e s s e d i t s e l f i n a v a r i e t y o f f o r m s - s o c i a l, r e l i g i
C H A P T E R I G E N E S I S A N D GROWTH OF G U IL D S C o r p o r a t e l i f e i n A n c i e n t I n d i a e x p r e s s e d i t s e l f i n a v a r i e t y o f f o r m s - s o c i a l, r e l i g i
UCLA STAT 233 Statistical Methods in Biomedical Imaging
 UCLA STAT 233 Statistical Methods in Biomedical Imaging Instructor: Ivo Dinov, Asst. Prof. In Statistics and Neurology University of California, Los Angeles, Spring 2004 http://www.stat.ucla.edu/~dinov/
UCLA STAT 233 Statistical Methods in Biomedical Imaging Instructor: Ivo Dinov, Asst. Prof. In Statistics and Neurology University of California, Los Angeles, Spring 2004 http://www.stat.ucla.edu/~dinov/
Rapid Biochemical Characterization of Haemophilus Species
 JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 1980, p. 22-26 0095-1137/80/01-0022/05$02.00/0 Vol. 11, No. 1 Rapid Biochemical Characterization of Haemophilus Species by Using the Micro-ID STEPHEN C. EDBERG,*
JOURNAL OF CLINICAL MICROBIOLOGY, Jan. 1980, p. 22-26 0095-1137/80/01-0022/05$02.00/0 Vol. 11, No. 1 Rapid Biochemical Characterization of Haemophilus Species by Using the Micro-ID STEPHEN C. EDBERG,*
Machine Learning. B. Unsupervised Learning B.2 Dimensionality Reduction. Lars Schmidt-Thieme, Nicolas Schilling
 Machine Learning B. Unsupervised Learning B.2 Dimensionality Reduction Lars Schmidt-Thieme, Nicolas Schilling Information Systems and Machine Learning Lab (ISMLL) Institute for Computer Science University
Machine Learning B. Unsupervised Learning B.2 Dimensionality Reduction Lars Schmidt-Thieme, Nicolas Schilling Information Systems and Machine Learning Lab (ISMLL) Institute for Computer Science University
Unconstrained Ordination
 Unconstrained Ordination Sites Species A Species B Species C Species D Species E 1 0 (1) 5 (1) 1 (1) 10 (4) 10 (4) 2 2 (3) 8 (3) 4 (3) 12 (6) 20 (6) 3 8 (6) 20 (6) 10 (6) 1 (2) 3 (2) 4 4 (5) 11 (5) 8 (5)
Unconstrained Ordination Sites Species A Species B Species C Species D Species E 1 0 (1) 5 (1) 1 (1) 10 (4) 10 (4) 2 2 (3) 8 (3) 4 (3) 12 (6) 20 (6) 3 8 (6) 20 (6) 10 (6) 1 (2) 3 (2) 4 4 (5) 11 (5) 8 (5)
DIMENSION REDUCTION AND CLUSTER ANALYSIS
 DIMENSION REDUCTION AND CLUSTER ANALYSIS EECS 833, 6 March 2006 Geoff Bohling Assistant Scientist Kansas Geological Survey geoff@kgs.ku.edu 864-2093 Overheads and resources available at http://people.ku.edu/~gbohling/eecs833
DIMENSION REDUCTION AND CLUSTER ANALYSIS EECS 833, 6 March 2006 Geoff Bohling Assistant Scientist Kansas Geological Survey geoff@kgs.ku.edu 864-2093 Overheads and resources available at http://people.ku.edu/~gbohling/eecs833
A Cautionary Note on the Use of LISREL s Automatic Start Values in Confirmatory Factor Analysis Studies R. L. Brown University of Wisconsin
 A Cautionary Note on the Use of LISREL s Automatic Start Values in Confirmatory Factor Analysis Studies R. L. Brown University of Wisconsin The accuracy of parameter estimates provided by the major computer
A Cautionary Note on the Use of LISREL s Automatic Start Values in Confirmatory Factor Analysis Studies R. L. Brown University of Wisconsin The accuracy of parameter estimates provided by the major computer
ECE 521. Lecture 11 (not on midterm material) 13 February K-means clustering, Dimensionality reduction
 ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
ECE 521 Lecture 11 (not on midterm material) 13 February 2017 K-means clustering, Dimensionality reduction With thanks to Ruslan Salakhutdinov for an earlier version of the slides Overview K-means clustering
Evaluation of Mast-ID 15 system for identifying
 J Clin Pathol 7;4:6-73 Evaluation of Mast-ID 5 system for identifying Enterobacteriaceae, some Vibrionaceae, and Acinetobacter B HOLMES, C A DAWSON From the National Collection of Type Cultures, Central
J Clin Pathol 7;4:6-73 Evaluation of Mast-ID 5 system for identifying Enterobacteriaceae, some Vibrionaceae, and Acinetobacter B HOLMES, C A DAWSON From the National Collection of Type Cultures, Central
Statistical Machine Learning
 Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Statistical Machine Learning Christoph Lampert Spring Semester 2015/2016 // Lecture 12 1 / 36 Unsupervised Learning Dimensionality Reduction 2 / 36 Dimensionality Reduction Given: data X = {x 1,..., x
Table of Contents. Multivariate methods. Introduction II. Introduction I
 Table of Contents Introduction Antti Penttilä Department of Physics University of Helsinki Exactum summer school, 04 Construction of multinormal distribution Test of multinormality with 3 Interpretation
Table of Contents Introduction Antti Penttilä Department of Physics University of Helsinki Exactum summer school, 04 Construction of multinormal distribution Test of multinormality with 3 Interpretation
Principal Component Analysis
 I.T. Jolliffe Principal Component Analysis Second Edition With 28 Illustrations Springer Contents Preface to the Second Edition Preface to the First Edition Acknowledgments List of Figures List of Tables
I.T. Jolliffe Principal Component Analysis Second Edition With 28 Illustrations Springer Contents Preface to the Second Edition Preface to the First Edition Acknowledgments List of Figures List of Tables
Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size
 Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size Berkman Sahiner, a) Heang-Ping Chan, Nicholas Petrick, Robert F. Wagner, b) and Lubomir Hadjiiski
Feature selection and classifier performance in computer-aided diagnosis: The effect of finite sample size Berkman Sahiner, a) Heang-Ping Chan, Nicholas Petrick, Robert F. Wagner, b) and Lubomir Hadjiiski
Factor analysis. George Balabanis
 Factor analysis George Balabanis Key Concepts and Terms Deviation. A deviation is a value minus its mean: x - mean x Variance is a measure of how spread out a distribution is. It is computed as the average
Factor analysis George Balabanis Key Concepts and Terms Deviation. A deviation is a value minus its mean: x - mean x Variance is a measure of how spread out a distribution is. It is computed as the average
The Ability C ongress held at the Shoreham Hotel Decem ber 29 to 31, was a reco rd breaker for winter C ongresses.
 The Ability C ongress held at the Shoreham Hotel Decem ber 29 to 31, was a reco rd breaker for winter C ongresses. Attended by m ore than 3 00 people, all seem ed delighted, with the lectu res and sem
The Ability C ongress held at the Shoreham Hotel Decem ber 29 to 31, was a reco rd breaker for winter C ongresses. Attended by m ore than 3 00 people, all seem ed delighted, with the lectu res and sem
FREQUENCY DISTRIBUTION OF BACTERIA ISOLATED FROM DIFFERENT INDUSTRIAL EFFLUENTS
 28 DAFFODIL INTERNATIONAL UNIVERSITY JOURNAL OF SCIENCE AND TECHNOLOGY, VOLUME 7, ISSUE 1, JANUARY 2012 FREQUENCY DISTRIBUTION OF BACTERIA ISOLATED FROM DIFFERENT INDUSTRIAL EFFLUENTS Amna Ali 1 *and Fozia
28 DAFFODIL INTERNATIONAL UNIVERSITY JOURNAL OF SCIENCE AND TECHNOLOGY, VOLUME 7, ISSUE 1, JANUARY 2012 FREQUENCY DISTRIBUTION OF BACTERIA ISOLATED FROM DIFFERENT INDUSTRIAL EFFLUENTS Amna Ali 1 *and Fozia
Form and content. Iowa Research Online. University of Iowa. Ann A Rahim Khan University of Iowa. Theses and Dissertations
 University of Iowa Iowa Research Online Theses and Dissertations 1979 Form and content Ann A Rahim Khan University of Iowa Posted with permission of the author. This thesis is available at Iowa Research
University of Iowa Iowa Research Online Theses and Dissertations 1979 Form and content Ann A Rahim Khan University of Iowa Posted with permission of the author. This thesis is available at Iowa Research
Dimensionality Reduction Using PCA/LDA. Hongyu Li School of Software Engineering TongJi University Fall, 2014
 Dimensionality Reduction Using PCA/LDA Hongyu Li School of Software Engineering TongJi University Fall, 2014 Dimensionality Reduction One approach to deal with high dimensional data is by reducing their
Dimensionality Reduction Using PCA/LDA Hongyu Li School of Software Engineering TongJi University Fall, 2014 Dimensionality Reduction One approach to deal with high dimensional data is by reducing their
Machine Learning, Fall 2009: Midterm
 10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
10-601 Machine Learning, Fall 009: Midterm Monday, November nd hours 1. Personal info: Name: Andrew account: E-mail address:. You are permitted two pages of notes and a calculator. Please turn off all
gender mains treaming in Polis h practice
 gender mains treaming in Polis h practice B E R L IN, 1 9-2 1 T H A P R IL, 2 O O 7 Gender mains treaming at national level Parliament 25 % of women in S ejm (Lower Chamber) 16 % of women in S enat (Upper
gender mains treaming in Polis h practice B E R L IN, 1 9-2 1 T H A P R IL, 2 O O 7 Gender mains treaming at national level Parliament 25 % of women in S ejm (Lower Chamber) 16 % of women in S enat (Upper
Machine Learning. Principal Components Analysis. Le Song. CSE6740/CS7641/ISYE6740, Fall 2012
 Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
Machine Learning CSE6740/CS7641/ISYE6740, Fall 2012 Principal Components Analysis Le Song Lecture 22, Nov 13, 2012 Based on slides from Eric Xing, CMU Reading: Chap 12.1, CB book 1 2 Factor or Component
A L A BA M A L A W R E V IE W
 A L A BA M A L A W R E V IE W Volume 52 Fall 2000 Number 1 B E F O R E D I S A B I L I T Y C I V I L R I G HT S : C I V I L W A R P E N S I O N S A N D TH E P O L I T I C S O F D I S A B I L I T Y I N
A L A BA M A L A W R E V IE W Volume 52 Fall 2000 Number 1 B E F O R E D I S A B I L I T Y C I V I L R I G HT S : C I V I L W A R P E N S I O N S A N D TH E P O L I T I C S O F D I S A B I L I T Y I N
Functional pottery [slide]
![Functional pottery [slide] Functional pottery [slide]](/thumbs/89/99193754.jpg) Functional pottery [slide] by Frank Bevis Fabens A thesis submitted in partial fulfillment of the requirements for the degree of Master of Fine Arts Montana State University Copyright by Frank Bevis Fabens
Functional pottery [slide] by Frank Bevis Fabens A thesis submitted in partial fulfillment of the requirements for the degree of Master of Fine Arts Montana State University Copyright by Frank Bevis Fabens
Ch 10. Classification of Microorganisms
 Ch 10 Classification of Microorganisms Student Learning Outcomes Define taxonomy, taxon, and phylogeny. List the characteristics of the Bacteria, Archaea, and Eukarya domains. Differentiate among eukaryotic,
Ch 10 Classification of Microorganisms Student Learning Outcomes Define taxonomy, taxon, and phylogeny. List the characteristics of the Bacteria, Archaea, and Eukarya domains. Differentiate among eukaryotic,
PCA and LDA. Man-Wai MAK
 PCA and LDA Man-Wai MAK Dept. of Electronic and Information Engineering, The Hong Kong Polytechnic University enmwmak@polyu.edu.hk http://www.eie.polyu.edu.hk/ mwmak References: S.J.D. Prince,Computer
PCA and LDA Man-Wai MAK Dept. of Electronic and Information Engineering, The Hong Kong Polytechnic University enmwmak@polyu.edu.hk http://www.eie.polyu.edu.hk/ mwmak References: S.J.D. Prince,Computer
Applied Multivariate Analysis
 Department of Mathematics and Statistics, University of Vaasa, Finland Spring 2017 Dimension reduction Exploratory (EFA) Background While the motivation in PCA is to replace the original (correlated) variables
Department of Mathematics and Statistics, University of Vaasa, Finland Spring 2017 Dimension reduction Exploratory (EFA) Background While the motivation in PCA is to replace the original (correlated) variables
Introduction to Machine Learning
 10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
10-701 Introduction to Machine Learning PCA Slides based on 18-661 Fall 2018 PCA Raw data can be Complex, High-dimensional To understand a phenomenon we measure various related quantities If we knew what
Unsupervised Learning: K- Means & PCA
 Unsupervised Learning: K- Means & PCA Unsupervised Learning Supervised learning used labeled data pairs (x, y) to learn a func>on f : X Y But, what if we don t have labels? No labels = unsupervised learning
Unsupervised Learning: K- Means & PCA Unsupervised Learning Supervised learning used labeled data pairs (x, y) to learn a func>on f : X Y But, what if we don t have labels? No labels = unsupervised learning
Freeman (2005) - Graphic Techniques for Exploring Social Network Data
 Freeman (2005) - Graphic Techniques for Exploring Social Network Data The analysis of social network data has two main goals: 1. Identify cohesive groups 2. Identify social positions Moreno (1932) was
Freeman (2005) - Graphic Techniques for Exploring Social Network Data The analysis of social network data has two main goals: 1. Identify cohesive groups 2. Identify social positions Moreno (1932) was
PHONEME CLASSIFICATION OVER THE RECONSTRUCTED PHASE SPACE USING PRINCIPAL COMPONENT ANALYSIS
 PHONEME CLASSIFICATION OVER THE RECONSTRUCTED PHASE SPACE USING PRINCIPAL COMPONENT ANALYSIS Jinjin Ye jinjin.ye@mu.edu Michael T. Johnson mike.johnson@mu.edu Richard J. Povinelli richard.povinelli@mu.edu
PHONEME CLASSIFICATION OVER THE RECONSTRUCTED PHASE SPACE USING PRINCIPAL COMPONENT ANALYSIS Jinjin Ye jinjin.ye@mu.edu Michael T. Johnson mike.johnson@mu.edu Richard J. Povinelli richard.povinelli@mu.edu
Statistical Pattern Recognition
 Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
Statistical Pattern Recognition Feature Extraction Hamid R. Rabiee Jafar Muhammadi, Alireza Ghasemi, Payam Siyari Spring 2014 http://ce.sharif.edu/courses/92-93/2/ce725-2/ Agenda Dimensionality Reduction
176 Index. G Gradient, 4, 17, 22, 24, 42, 44, 45, 51, 52, 55, 56
 References Aljandali, A. (2014). Exchange rate forecasting: Regional applications to ASEAN, CACM, MERCOSUR and SADC countries. Unpublished PhD thesis, London Metropolitan University, London. Aljandali,
References Aljandali, A. (2014). Exchange rate forecasting: Regional applications to ASEAN, CACM, MERCOSUR and SADC countries. Unpublished PhD thesis, London Metropolitan University, London. Aljandali,
2/26/2017. This is similar to canonical correlation in some ways. PSY 512: Advanced Statistics for Psychological and Behavioral Research 2
 PSY 512: Advanced Statistics for Psychological and Behavioral Research 2 What is factor analysis? What are factors? Representing factors Graphs and equations Extracting factors Methods and criteria Interpreting
PSY 512: Advanced Statistics for Psychological and Behavioral Research 2 What is factor analysis? What are factors? Representing factors Graphs and equations Extracting factors Methods and criteria Interpreting
Lecture 7: Con3nuous Latent Variable Models
 CSC2515 Fall 2015 Introduc3on to Machine Learning Lecture 7: Con3nuous Latent Variable Models All lecture slides will be available as.pdf on the course website: http://www.cs.toronto.edu/~urtasun/courses/csc2515/
CSC2515 Fall 2015 Introduc3on to Machine Learning Lecture 7: Con3nuous Latent Variable Models All lecture slides will be available as.pdf on the course website: http://www.cs.toronto.edu/~urtasun/courses/csc2515/
Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization
 Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization Haiping Lu 1 K. N. Plataniotis 1 A. N. Venetsanopoulos 1,2 1 Department of Electrical & Computer Engineering,
Uncorrelated Multilinear Principal Component Analysis through Successive Variance Maximization Haiping Lu 1 K. N. Plataniotis 1 A. N. Venetsanopoulos 1,2 1 Department of Electrical & Computer Engineering,
How to Run the Analysis: To run a principal components factor analysis, from the menus choose: Analyze Dimension Reduction Factor...
 The principal components method of extraction begins by finding a linear combination of variables that accounts for as much variation in the original variables as possible. This method is most often used
The principal components method of extraction begins by finding a linear combination of variables that accounts for as much variation in the original variables as possible. This method is most often used
Canonical Correlation & Principle Components Analysis
 Canonical Correlation & Principle Components Analysis Aaron French Canonical Correlation Canonical Correlation is used to analyze correlation between two sets of variables when there is one set of IVs
Canonical Correlation & Principle Components Analysis Aaron French Canonical Correlation Canonical Correlation is used to analyze correlation between two sets of variables when there is one set of IVs
Taxonomy. Content. How to determine & classify a species. Phylogeny and evolution
 Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
Taxonomy Content Why Taxonomy? How to determine & classify a species Domains versus Kingdoms Phylogeny and evolution Why Taxonomy? Classification Arrangement in groups or taxa (taxon = group) Nomenclature
ECE 592 Topics in Data Science
 ECE 592 Topics in Data Science Final Fall 2017 December 11, 2017 Please remember to justify your answers carefully, and to staple your test sheet and answers together before submitting. Name: Student ID:
ECE 592 Topics in Data Science Final Fall 2017 December 11, 2017 Please remember to justify your answers carefully, and to staple your test sheet and answers together before submitting. Name: Student ID:
Phylogenetic Diversity of Coliform Isolates in USA. Phylogenetic Classification
 Phylogenetic Diversity of Coliform Isolates in USA Ya Zhang and Wen Tso Liu University of Illinois at Urbana Champaign Mark LeChevallier American Water Inc. Nov 2011 Phylogenetic Classification group organisms
Phylogenetic Diversity of Coliform Isolates in USA Ya Zhang and Wen Tso Liu University of Illinois at Urbana Champaign Mark LeChevallier American Water Inc. Nov 2011 Phylogenetic Classification group organisms
Chemometrics. 1. Find an important subset of the original variables.
 Chemistry 311 2003-01-13 1 Chemometrics Chemometrics: Mathematical, statistical, graphical or symbolic methods to improve the understanding of chemical information. or The science of relating measurements
Chemistry 311 2003-01-13 1 Chemometrics Chemometrics: Mathematical, statistical, graphical or symbolic methods to improve the understanding of chemical information. or The science of relating measurements
Lecture 24: Principal Component Analysis. Aykut Erdem May 2016 Hacettepe University
 Lecture 4: Principal Component Analysis Aykut Erdem May 016 Hacettepe University This week Motivation PCA algorithms Applications PCA shortcomings Autoencoders Kernel PCA PCA Applications Data Visualization
Lecture 4: Principal Component Analysis Aykut Erdem May 016 Hacettepe University This week Motivation PCA algorithms Applications PCA shortcomings Autoencoders Kernel PCA PCA Applications Data Visualization
Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations.
 Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Previously Focus was on solving matrix inversion problems Now we look at other properties of matrices Useful when A represents a transformations y = Ax Or A simply represents data Notion of eigenvectors,
Use precise language and domain-specific vocabulary to inform about or explain the topic. CCSS.ELA-LITERACY.WHST D
 Lesson eight What are characteristics of chemical reactions? Science Constructing Explanations, Engaging in Argument and Obtaining, Evaluating, and Communicating Information ENGLISH LANGUAGE ARTS Reading
Lesson eight What are characteristics of chemical reactions? Science Constructing Explanations, Engaging in Argument and Obtaining, Evaluating, and Communicating Information ENGLISH LANGUAGE ARTS Reading
Math for Machine Learning Open Doors to Data Science and Artificial Intelligence. Richard Han
 Math for Machine Learning Open Doors to Data Science and Artificial Intelligence Richard Han Copyright 05 Richard Han All rights reserved. CONTENTS PREFACE... - INTRODUCTION... LINEAR REGRESSION... 4 LINEAR
Math for Machine Learning Open Doors to Data Science and Artificial Intelligence Richard Han Copyright 05 Richard Han All rights reserved. CONTENTS PREFACE... - INTRODUCTION... LINEAR REGRESSION... 4 LINEAR
LSU Historical Dissertations and Theses
 Louisiana State University LSU Digital Commons LSU Historical Dissertations and Theses Graduate School 1976 Infestation of Root Nodules of Soybean by Larvae of the Bean Leaf Beetle, Cerotoma Trifurcata
Louisiana State University LSU Digital Commons LSU Historical Dissertations and Theses Graduate School 1976 Infestation of Root Nodules of Soybean by Larvae of the Bean Leaf Beetle, Cerotoma Trifurcata
Application of Indirect Race/ Ethnicity Data in Quality Metric Analyses
 Background The fifteen wholly-owned health plans under WellPoint, Inc. (WellPoint) historically did not collect data in regard to the race/ethnicity of it members. In order to overcome this lack of data
Background The fifteen wholly-owned health plans under WellPoint, Inc. (WellPoint) historically did not collect data in regard to the race/ethnicity of it members. In order to overcome this lack of data
I zm ir I nstiute of Technology CS Lecture Notes are based on the CS 101 notes at the University of I llinois at Urbana-Cham paign
 I zm ir I nstiute of Technology CS - 1 0 2 Lecture 1 Lecture Notes are based on the CS 101 notes at the University of I llinois at Urbana-Cham paign I zm ir I nstiute of Technology W hat w ill I learn
I zm ir I nstiute of Technology CS - 1 0 2 Lecture 1 Lecture Notes are based on the CS 101 notes at the University of I llinois at Urbana-Cham paign I zm ir I nstiute of Technology W hat w ill I learn
Simplifying Drug Discovery with JMP
 Simplifying Drug Discovery with JMP John A. Wass, Ph.D. Quantum Cat Consultants, Lake Forest, IL Cele Abad-Zapatero, Ph.D. Adjunct Professor, Center for Pharmaceutical Biotechnology, University of Illinois
Simplifying Drug Discovery with JMP John A. Wass, Ph.D. Quantum Cat Consultants, Lake Forest, IL Cele Abad-Zapatero, Ph.D. Adjunct Professor, Center for Pharmaceutical Biotechnology, University of Illinois
Gene Expression Data Classification With Kernel Principal Component Analysis
 Journal of Biomedicine and Biotechnology 25:2 25 55 59 DOI:.55/JBB.25.55 RESEARCH ARTICLE Gene Expression Data Classification With Kernel Principal Component Analysis Zhenqiu Liu, Dechang Chen, 2 and Halima
Journal of Biomedicine and Biotechnology 25:2 25 55 59 DOI:.55/JBB.25.55 RESEARCH ARTICLE Gene Expression Data Classification With Kernel Principal Component Analysis Zhenqiu Liu, Dechang Chen, 2 and Halima
Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation)
 Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation) PCA transforms the original input space into a lower dimensional space, by constructing dimensions that are linear combinations
Principal Component Analysis -- PCA (also called Karhunen-Loeve transformation) PCA transforms the original input space into a lower dimensional space, by constructing dimensions that are linear combinations
Introduction PCA classic Generative models Beyond and summary. PCA, ICA and beyond
 PCA, ICA and beyond Summer School on Manifold Learning in Image and Signal Analysis, August 17-21, 2009, Hven Technical University of Denmark (DTU) & University of Copenhagen (KU) August 18, 2009 Motivation
PCA, ICA and beyond Summer School on Manifold Learning in Image and Signal Analysis, August 17-21, 2009, Hven Technical University of Denmark (DTU) & University of Copenhagen (KU) August 18, 2009 Motivation
Comparison of Crystal Enteric/Nonfermenter System, API 20E System, and Vitek Automicrobic System for Identification of Gram-Negative Bacilli
 JOURNAL OF CLINICAL MICROBIOLOGY, Feb. 1995, p. 364 370 Vol. 33, No. 2 0095-1137/95/$04.00 0 Copyright 1995, American Society for Microbiology Comparison of Crystal Enteric/Nonfermenter System, API 20E
JOURNAL OF CLINICAL MICROBIOLOGY, Feb. 1995, p. 364 370 Vol. 33, No. 2 0095-1137/95/$04.00 0 Copyright 1995, American Society for Microbiology Comparison of Crystal Enteric/Nonfermenter System, API 20E
What is Principal Component Analysis?
 What is Principal Component Analysis? Principal component analysis (PCA) Reduce the dimensionality of a data set by finding a new set of variables, smaller than the original set of variables Retains most
What is Principal Component Analysis? Principal component analysis (PCA) Reduce the dimensionality of a data set by finding a new set of variables, smaller than the original set of variables Retains most
Experimental Design and Data Analysis for Biologists
 Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Experimental Design and Data Analysis for Biologists Gerry P. Quinn Monash University Michael J. Keough University of Melbourne CAMBRIDGE UNIVERSITY PRESS Contents Preface page xv I I Introduction 1 1.1
Machine Learning (Spring 2012) Principal Component Analysis
 1-71 Machine Learning (Spring 1) Principal Component Analysis Yang Xu This note is partly based on Chapter 1.1 in Chris Bishop s book on PRML and the lecture slides on PCA written by Carlos Guestrin in
1-71 Machine Learning (Spring 1) Principal Component Analysis Yang Xu This note is partly based on Chapter 1.1 in Chris Bishop s book on PRML and the lecture slides on PCA written by Carlos Guestrin in
Research Statement on Statistics Jun Zhang
 Research Statement on Statistics Jun Zhang (junzhang@galton.uchicago.edu) My interest on statistics generally includes machine learning and statistical genetics. My recent work focus on detection and interpretation
Research Statement on Statistics Jun Zhang (junzhang@galton.uchicago.edu) My interest on statistics generally includes machine learning and statistical genetics. My recent work focus on detection and interpretation
Principal Component Analysis & Factor Analysis. Psych 818 DeShon
 Principal Component Analysis & Factor Analysis Psych 818 DeShon Purpose Both are used to reduce the dimensionality of correlated measurements Can be used in a purely exploratory fashion to investigate
Principal Component Analysis & Factor Analysis Psych 818 DeShon Purpose Both are used to reduce the dimensionality of correlated measurements Can be used in a purely exploratory fashion to investigate
STA 414/2104: Lecture 8
 STA 414/2104: Lecture 8 6-7 March 2017: Continuous Latent Variable Models, Neural networks With thanks to Russ Salakhutdinov, Jimmy Ba and others Outline Continuous latent variable models Background PCA
STA 414/2104: Lecture 8 6-7 March 2017: Continuous Latent Variable Models, Neural networks With thanks to Russ Salakhutdinov, Jimmy Ba and others Outline Continuous latent variable models Background PCA
Factor Analysis (10/2/13)
 STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
STA561: Probabilistic machine learning Factor Analysis (10/2/13) Lecturer: Barbara Engelhardt Scribes: Li Zhu, Fan Li, Ni Guan Factor Analysis Factor analysis is related to the mixture models we have studied.
Structure in Data. A major objective in data analysis is to identify interesting features or structure in the data.
 Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Structure in Data A major objective in data analysis is to identify interesting features or structure in the data. The graphical methods are very useful in discovering structure. There are basically two
Advanced Introduction to Machine Learning CMU-10715
 Advanced Introduction to Machine Learning CMU-10715 Principal Component Analysis Barnabás Póczos Contents Motivation PCA algorithms Applications Some of these slides are taken from Karl Booksh Research
Advanced Introduction to Machine Learning CMU-10715 Principal Component Analysis Barnabás Póczos Contents Motivation PCA algorithms Applications Some of these slides are taken from Karl Booksh Research
Principles of factor analysis. Roger Watson
 Principles of factor analysis Roger Watson Factor analysis Factor analysis Factor analysis Factor analysis is a multivariate statistical method for reducing large numbers of variables to fewer underlying
Principles of factor analysis Roger Watson Factor analysis Factor analysis Factor analysis Factor analysis is a multivariate statistical method for reducing large numbers of variables to fewer underlying
Lecture: Face Recognition and Feature Reduction
 Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab Lecture 11-1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed
Lecture: Face Recognition and Feature Reduction Juan Carlos Niebles and Ranjay Krishna Stanford Vision and Learning Lab Lecture 11-1 Recap - Curse of dimensionality Assume 5000 points uniformly distributed
Computational Genomics
 Computational Genomics http://www.cs.cmu.edu/~02710 Introduction to probability, statistics and algorithms (brief) intro to probability Basic notations Random variable - referring to an element / event
Computational Genomics http://www.cs.cmu.edu/~02710 Introduction to probability, statistics and algorithms (brief) intro to probability Basic notations Random variable - referring to an element / event
Lecture 6: Methods for high-dimensional problems
 Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Lecture 6: Methods for high-dimensional problems Hector Corrada Bravo and Rafael A. Irizarry March, 2010 In this Section we will discuss methods where data lies on high-dimensional spaces. In particular,
Overview of clustering analysis. Yuehua Cui
 Overview of clustering analysis Yuehua Cui Email: cuiy@msu.edu http://www.stt.msu.edu/~cui A data set with clear cluster structure How would you design an algorithm for finding the three clusters in this
Overview of clustering analysis Yuehua Cui Email: cuiy@msu.edu http://www.stt.msu.edu/~cui A data set with clear cluster structure How would you design an algorithm for finding the three clusters in this
Factor Analysis (1) Factor Analysis
 Factor Analysis (1) Outlines: 1. Introduction of factor analysis 2. Principle component analysis 4. Factor rotation 5. Case Shan-Yu Chou 1 Factor Analysis Combines questions or variables to create new
Factor Analysis (1) Outlines: 1. Introduction of factor analysis 2. Principle component analysis 4. Factor rotation 5. Case Shan-Yu Chou 1 Factor Analysis Combines questions or variables to create new
Principal Components Analysis. Sargur Srihari University at Buffalo
 Principal Components Analysis Sargur Srihari University at Buffalo 1 Topics Projection Pursuit Methods Principal Components Examples of using PCA Graphical use of PCA Multidimensional Scaling Srihari 2
Principal Components Analysis Sargur Srihari University at Buffalo 1 Topics Projection Pursuit Methods Principal Components Examples of using PCA Graphical use of PCA Multidimensional Scaling Srihari 2
TECHNIQUE FOR RANKING POTENTIAL PREDICTOR LAYERS FOR USE IN REMOTE SENSING ANALYSIS. Andrew Lister, Mike Hoppus, and Rachel Riemam
 TECHNIQUE FOR RANKING POTENTIAL PREDICTOR LAYERS FOR USE IN REMOTE SENSING ANALYSIS Andrew Lister, Mike Hoppus, and Rachel Riemam ABSTRACT. Spatial modeling using GIS-based predictor layers often requires
TECHNIQUE FOR RANKING POTENTIAL PREDICTOR LAYERS FOR USE IN REMOTE SENSING ANALYSIS Andrew Lister, Mike Hoppus, and Rachel Riemam ABSTRACT. Spatial modeling using GIS-based predictor layers often requires
Ordination & PCA. Ordination. Ordination
 Ordination & PCA Introduction to Ordination Purpose & types Shepard diagrams Principal Components Analysis (PCA) Properties Computing eigenvalues Computing principal components Biplots Covariance vs. Correlation
Ordination & PCA Introduction to Ordination Purpose & types Shepard diagrams Principal Components Analysis (PCA) Properties Computing eigenvalues Computing principal components Biplots Covariance vs. Correlation
LONGITUDINAL ANALYSIS THROUGH MULTIPLE PROCESS STEPS, MEAN AND VARIABILITY MODULE OBJECTIVES
 LONGITUDINAL ANALYSIS THROUGH MULTIPLE PROCESS STEPS, MEAN AND VARIABILITY REPEATED MEASURES Tony Cooper SAS Inc. Tony.cooper@sas.com Doug Sanders 1 MODULE OBJECTIVES Introduce Manufacturing Repeated Measures
LONGITUDINAL ANALYSIS THROUGH MULTIPLE PROCESS STEPS, MEAN AND VARIABILITY REPEATED MEASURES Tony Cooper SAS Inc. Tony.cooper@sas.com Doug Sanders 1 MODULE OBJECTIVES Introduce Manufacturing Repeated Measures
Quantitative Understanding in Biology Principal Components Analysis
 Quantitative Understanding in Biology Principal Components Analysis Introduction Throughout this course we have seen examples of complex mathematical phenomena being represented as linear combinations
Quantitative Understanding in Biology Principal Components Analysis Introduction Throughout this course we have seen examples of complex mathematical phenomena being represented as linear combinations
Data Mining. Dimensionality reduction. Hamid Beigy. Sharif University of Technology. Fall 1395
 Data Mining Dimensionality reduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 42 Outline 1 Introduction 2 Feature selection
Data Mining Dimensionality reduction Hamid Beigy Sharif University of Technology Fall 1395 Hamid Beigy (Sharif University of Technology) Data Mining Fall 1395 1 / 42 Outline 1 Introduction 2 Feature selection
