Computational and physical models of RNA structure
|
|
|
- Kerry Quinn
- 5 years ago
- Views:
Transcription
1 omputational and physical models of RN structure Ralf Bundschuh Ohio State niversity July 25, 2007 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
2 Outline of all lectures Outline Part I : Statistical physics of RN Part II : Quantitative modeling of force-extension experiments Part III : RN folding kinetics Part IV : microrn target prediction Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
3 Outline of part I 1 Boltzmann partition function 2 Molten RN 3 Molten-native transition 4 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
4 Outline of part I Boltzmann partition function 1 Boltzmann partition function Secondary structure Partition function Recursion equation 2 Molten RN 3 Molten-native transition 4 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
5 Boltzmann partition function Secondary structure Definition of RN secondary structure Definition n RN secondary structure on an RN sequence of length N is a set S of base pairs (i, j) with 1 i < j N that fulfill the following conditions: Each base is involved in at most one base pair No pseudoknots, i.e., if (i, j) and (k, l) are base pairs with i < k, either i < k < l < j or i < j < k < l Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
6 Boltzmann partition function Diagrammatic representation Secondary structure n RN secondary structure can be represented by an arch diagram Every base is end point to at most one arch Two arches never cross One to one correspondance between arch diagrams that fulfil the above conditions and secondary structures Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
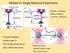
7 Energy models Boltzmann partition function Partition function Each structure S has a certain (free) energy E[S] Different models possible ll base pairs are equal E[S] = ε 0 S (ε 0 < 0) Energy associated with base pairs E[S] = (i,j) S e(i, j) Turner parameters (nearest neighbor model): energy associated with loops Base pair stacks Hairpin loops Interior loops Bulges Multi-loops multiloop bulge hairpin loop stacking loop interior loop Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
8 Partition function Boltzmann partition function Partition function Definition The partition function of an RN molecule with energy function E[S] is given by Z S e E[S] RT R = cal mol K Temperature T in Kelvin RT 0.6 kcal mol Ensemble free energy F = RT ln Z Thermodynamics completely specified if partition function is known Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
9 Boltzmann partition function alculating partition functions Recursion equation Definition Let Z i,j be the partition function for the RN molecule starting at base i and ending at base j. Denote this quantity by i j. Observation For the base pairing energy model the Z i,j obey the recursion equation = + i j i j 1 j Σ k i k 1 k k+1 j 1 j j 1 Z i,j = Z i,j 1 + Z i,k 1 e e(k,j) RT Zk+1,j 1 k=i alculate from shortest to longest substrands O(N 3 ) algorithm for arbitrary sequence Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
10 Outline of part I Molten RN 1 Boltzmann partition function 2 Molten RN Energy model z transform Properties 3 Molten-native transition 4 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
11 Molten RN Energetics in molten phase Energy model Definition In the molten phase of RN every base can pair with every other base equally well, i.e., e(i, j) = ε 0. Properties pplies to repetitive sequences:,, pplies to arbitrary sequences at temperatures close to denaturation on a coarse-grained level Minimum energy is always N 2 ε 0 Structural entropy plays a major role Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
12 Molten RN Molten phase partition function Energy model Reminder = + i j i j 1 j Σ k i k 1 k k+1 j 1 j Simplification j 1 Z i,j = Z i,j 1 + Z i,k 1 e e(k,j) RT Zk+1,j 1 k=i In the molten phase the partition function depends only on the length j i of the substrand, not on i and j individually: Z i,j (j i + 2) onsequence N 1 (N + 1) = (N) + q (k)(n k) with q e ε 0 RT k=1 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
13 z transform Molten RN z transform Definition For any series Q(N) the z transform Q is defined as Q(z) Q(N)z N N=1 Properties Function of the complex variable z nalytic outside a radius of convergence Discrete version of Fourier transform Back transform: Q(N) = 1 Q(z)z N 1 dz onvolution property: 2πi N 1 k=1 Q(k)W (N k) = Q(z) Ŵ (z) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
14 Molten RN z transform z transform for molten RN Ĝ(z) can be calculated: N 1 (N + 1) = (N) + q (k)(n k) k=1 N 1 (N + 1)z N = (N)z N + qz N (N + 1)z N = Ĝ(z) + qĝ 2 (z) N=1 zĝ(z) (1) = Ĝ(z) + qĝ 2 (z) zĝ(z) 1 = Ĝ(z) + qĝ 2 (z) k=1 Ĝ(z) = 1 [ ] z 1 (z 1) 2q 2 4q (k)(n k) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
15 Back transform I Molten RN z transform Integral expression for (N) (N) = 1 Ĝ(z)z N 1 dz 2πi [ 1 = z 1 4πqi = 1 4πqi (z 1) 2 4q (z 1) 2 4q z N 1 dz ] z N 1 dz Singularity structure Branch cut from 1 2 q to z q Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
16 Back transform II Molten RN z transform Reminder (N) = 1 4πqi (z 1) 2 4q z N 1 dz Leading behavior For large N only the singularity with largest real part contributes (N) z0 N = (1 + 2 q) N Prefactor µc (N) N 3/2 (1 + 2 q) N µ 0 (µ c µ) α e µn dµ Γ(1 + α)n (1+α) e µcn Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
17 Molten RN Properties Properties of molten RN ( ) q 1/2 (N) N 3/2 (1 + 2 q) N 4πq 3/2 N 3/2 characteristic behavior due to entropy an be observed in pairing probability: (k)(n k) 3/2 (N k) 3/2 Pr{1 and k paired} = q k (N + 1) N 3/2 k 3/2 Free energy is F = RT ln (N) RTN ln(1 + 2 q) For q 1: F RTN ln(2 q) = RT 2 N ln(q) = N ε 0 2 For q = 1: (N) =number of secondary structures N 3/2 3 N. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
18 Outline of part I Molten-native transition 1 Boltzmann partition function 2 Molten RN 3 Molten-native transition Model Solution Results 4 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
19 Molten-native transition Model Model for molten-native transition Observation Structural RNs have to fold into a specific native structure there must be something in the sequence that prefers this structure Model se perfect hairpin as native structure 1 N/2 1 1 N N ssign binding energy ε 1 to all native base pairs ssign binding energy ε 0 to all other base pairs N Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
20 Molten-native transition Solution Partition function Definition Let Z(N; q, q) be the partition function of the ō model for the molten-native transition with 2N 2 bases, q = e ε 0/RT, and q = e ε 1/RT Definition Let W (N; q) Z(N; q, q = 0) be the partition function of the ō model for 2N 2 bases in which native contacts are disallowed. Observation Z(N; q, q) =(p.f. with 0 native contacts) + (p.f. with 1 native contacts) + (p.f. with 2 native contacts) +... = Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
21 Molten-native transition Solution Individual terms 0 native contacts (p.f. with 0 native contacts) = = W (N; q) z-transform: Ŵ (z) 1 native contact (p.f. with 1 native contacts) = z-transform: qŵ 2 (z) = q N 1 k=1 W (k; q)w (N k; q) 2 native contacts (p.f. with 2 native contacts) = = q 2 1 k 1 <k 2 <N W (k 1; q)w (k 2 k 1 ; q)w (N k 2 ; q) z-transform: q 2 Ŵ 3 (z) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
22 Putting it all together Molten-native transition Solution Summing up Ẑ(z; q, q) = Ŵ (z) + qŵ 2 (z) + q 2 Ŵ 3 (z) +... = Ŵ (z) 1 qŵ (z) What is Ŵ (z)? For q = q we have Z(N; q, q = q) = (2N 1; q) Ẑ(z; q, q) = Ê(z; q) N=1 (2N 1; q)z N Ŵ (z) = Ê(z) 1 + qê(z) Ẑ(z; q, q) = Ê(z) 1 ( q q)ê(z) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
23 Molten-native transition Solution Behavior of Ê(z) Expression for Ê(z) Ê(z) = N=1 (2N 1; q)z N can be calculated similarly to Ĝ(z). Ê(z) = 1 2q 1 [ ] (z 1) 4qz 2 4q + (z + 1) 2 4q. Properties Vanishes as z Square root branch cut at z 0 = q Finite limit Ê(1 + 2 q) at branch cut E(z ) 0 E(z) Other branch cuts have smaller real part z 0 z Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
24 Molten-native transition Solution Singularity structure of Ẑ(z) andidate singularities Ẑ(z; q, q) = Ê(z) 1 ( q q)ê(z) E(z ) 0 E(z) 0.1 Square root branch cut at z 0 = q 0.05 ~ 1 q q Pole at z 1 ( q) given by Ê(z 1) = 1/( q q) z Dominant singularity Square root branch cut at z 0 if 1/( q q) > Ê(z 0) Pole at z 1 ( q) if 1/( q q) < Ê(z 0) Phase transition z 0 z (q) ~ 1 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
25 Molten-native transition Results Properties of molten-native transition haracterization of phase transition ritical bias q c = q + 1/Ê(z 0) If q < q c regular molten behavior Z N 3/2 (1 + 2 q) N native base pairs do not play any role If q > q c native behavior with Z N 0 z 1 ( q) N finite fraction of native base pairs but still many bubbles For q q c we get Z N 0 q N only native base pairs onclusion It takes a finite amount of sequence bias to enforce a native structure. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
26 Outline of part I Summary 1 Boltzmann partition function 2 Molten RN 3 Molten-native transition 4 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
27 Summary Summary of part I The partition function of RN can be calculated in polynomial time symptotic behavior for homogeneous RN can be calculated by analytical methods The partition function of molten RN has a characteristic N 3/2 behavior It takes a finite amount of sequence bias to enforce a native structure. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
28 Outline of part II 5 Force-extension experiments 6 Quantitative modeling 7 Results for simple force-extension experiments 8 Nanopores 9 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
29 Outline of part II Force-extension experiments 5 Force-extension experiments Motivation Experimental setup 6 Quantitative modeling 7 Results for simple force-extension experiments 8 Nanopores 9 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
30 Force-extension experiments Motivation Experimental methods to determine RN structure Experimental methods X-ray crystallography Nuclear Magnetic Resonance Biochemical evidence: protection essays orrelated mutation analysis Problem RN is very floppy two RN molecules rarely have the same three-dimensional structure Idea Look at one RN molecule at a time. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
31 Experimental setup Force-extension experiments Experimental setup ttach ends of RN molecule to two beads Keep beads at fixed distance R with optical tweezers Measure force f on beads as function of distance R r Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
32 Measurement Force-extension experiments Experimental setup Subject P5ab hairpin of Tetrahymena thermophila group I intron Data (Liphardt, Onoa, Smith, Tinoco, and Bustamante, Science, 2001) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
33 Outline of part II Quantitative modeling 5 Force-extension experiments 6 Quantitative modeling Force from free energy Secondary structures Backbone Putting it back together 7 Results for simple force-extension experiments 8 Nanopores 9 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
34 alculating the force Quantitative modeling Force from free energy r Basic physics energy = force distance Force from free energy Need free energy F (r) at fixed end to end distance r force f = F (r) r Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
35 Partition function I Quantitative modeling Force from free energy r Free energy from partition function F (r) = RT ln Z(r) need partition function Z(r) at fixed distance r Partition function Z(r) = secondary structures S polymer configurations P with distance r and structure S e E[S]+E[P] RT Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
36 Partition function II Quantitative modeling Force from free energy r Z(r) = = = secondary structures S secondary structures S N m=0 secondary structures S with m exterior bases polymer configurations P with distance r and structure S polymer configurations P of the exterior bases with distance r and structure S polymer configurations P of the exterior bases with distance r and structure S e E[S]+E[P] RT e E[S]+E[P] RT e E[S]+E[P] RT Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
37 Partition function III Quantitative modeling Force from free energy Z(r) = = = N m=0 secondary structures S with m exterior bases N m=0 secondary structures S with m exterior bases N m=0 secondary structures S with m exterior bases r polymer configurations P of the exterior bases with distance r and structure S e E[S] RT e E[S] RT polymer configurations P of m exterior bases with distance r e E[S]+E[P] RT e E[P] RT polymer configurations P of ssrn with m bases with distance r e E[P] RT Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
38 Partition function IV Quantitative modeling Force from free energy N Z(r) = Q(m)W (m, r) m=0 with and Q(m) W (m, r) secondary structures S with m exterior bases e E[S] RT polymer configurations P of ssrn with m bases with distance r e E[P] RT Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
39 Recursion equation Quantitative modeling Secondary structures Reminder i j = i j 1 j + k i k 1 k k+1 j 1 j j 1 Z i,j = Z i,j 1 + k=i Σ Z i,k 1 e e(k,j) RT Zk+1,j 1 Definition Let Q j (m) be the partition function for the first j bases with exactly m exterior bases ( j). eneralization = Σ j 1 j 1 j k 1 k 1 k k+1 j 1 j j 1 Q j (m) = Q j 1 (m 1) + k=1 Q k 1 (m)e e(k,j) RT Zk+1,j 1 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
40 Quantitative modeling Properties of recursion equation Secondary structures Properties: = Σ j 1 j 1 j k 1 k 1 k k+1 j 1 j j 1 Q j (m) = Q j 1 (m 1) + O(N 3 ) complexity Q(m) = Q N (m) k=1 an be easily generalized to Turner parameters Q k 1 (m)e e(k,j) RT Zk+1,j 1 an be calculated by modifying the Vienna package (Hofacker, Fontana, Stadler, Bonhoeffer, Tacker, and Schuster, Monatshefte f. hemie, 1994) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
41 Quantitative modeling Backbone Polymer physics Need W (m, r) = polymer configurations P of ssrn with m bases with distance r e E[P] RT Model Elastic freely jointed chain Persistence length 1.9nm/base distance 0.7nm Partition function alculation of W (m, r) is standard polymer physics. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
42 RNpull Quantitative modeling Putting it back together Putting it together Q(m), W (m, r) N Z(r) = Q(m)W (m, r) m=0 F (r) = RT ln Z(r) F (r) f (r) = r Web server Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
43 Results for simple force-extension experiments Outline of part II 5 Force-extension experiments 6 Quantitative modeling 7 Results for simple force-extension experiments Hairpin real molecule Why the force-extension curve is smooth? 8 Nanopores 9 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
44 Hairpin Results for simple force-extension experiments Hairpin pply to P5ab hairpin of Tetrahymena thermophila group I intron Quantitative agreement! Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
45 Results for simple force-extension experiments The full group I intron real molecule The group I intron of Tetrahymena thermophila roup I intron contains pseudo-knot! Quantitative modeling ignores pseudo-knot known inactive conformation (Pan and Woodson, J. Mol. Biol., 1998) P5c P5a P5b P6b P5 P4 P6a P1 5 P2.1 P2 P9.1a P9 P9.1 P7 lt P3 3 P8 P9.2 omputational result No sign of secondary structure! <F> [pn] <R> [nm] Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
46 Results for simple force-extension experiments What s happening? Why the force-extension curve is smooth? Intermediate structure Look at intermediate structure (here r = 100nm) Like Socks on the @ Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
47 Results for simple force-extension experiments What s happening? Why the force-extension curve is smooth? ompensation effect Extension r is increased One of the socks disappears The other socks take up the slag No rapid change in force as sock disappears smooth force-extension curve Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
48 Outline of part II Nanopores 5 Force-extension experiments 6 Quantitative modeling 7 Results for simple force-extension experiments 8 Nanopores Introduction Force-extension experiments 9 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
49 Nanopores Nanopores Introduction What is a nanopore? nanopore is a little hole is so small that only single-stranded but no double-stranded RN can pass through it can be placed between two chambers such that there is only one nanopore connecting the chambers Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
50 Nanopores Nanopores Introduction Types of nanopores natural ion channel (α-hemolysin) (Meller, J. Phys. ond. Mat., 2003) solid state pore (Storm et al., Nature Mat., 2003) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
51 Nanopores Force-extension experiments ombining nanopores with force-extension experiments Suggestion ombine a nanopore with a force-extension setup "cis" "trans" n nucleotides f r Prediction The force will rise at every structural element The structural elements will break in their order along the sequence Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
52 Nanopores Force-extension experiments Simulation approach "cis" "trans" n nucleotides f Simulation Fix r(t) to be linear (set by experiment) Degree of freedom: number n of base in the pore t each time n can increase: calculable gain in mechanical energy on the right potentially loss in binding energy calculable from E[S] decrease: calculable loss in mechanical energy on the right Monte arlo simulation (see also part III) r Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
53 Simulation result Nanopores Force-extension experiments pply to group I intron of Tetrahymena thermophila 50 <f> [pn] n= P9.1 P9 P7 lt P3 P9.2 P8 P6 P4-P5a P5bc <R> [nm] P2.1 P2 P1 P5 P5a P1 P4 P5c 5 P6a P5b P6b P2.1 P2 P9.1a P9.1 P9 P9.2 3 P7 lt P3 P8 Signature of every structural element an extract sequence position of stalling sites Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
54 Structure reconstruction I Nanopores Force-extension experiments Repeat pulling in opposite direction <f> [pn] <f> [pn] n= n= <R> [nm]...<...>...<...<...>...<... 5 (((((((((((...))...)))))))))...((((((((((...)))))))))).(((((((( ccccccccc...ccccccccc...aaaaaaaaaa...aaaaaaaaaa.gggggg.....>...>...<..<...<...<... (((.((...)).))))).))))))...((((((...((((((...(((.(((((...gggggg...jjjj...kkkkkk...iiii......>...<...>...>...>..>...>. (((..(((((((((...)))))))))..(((...)))...)))...).)))))))...))))))....llll...llll...iiii...kkkkkk....>..>.<...<...<...>...>...<.....)).))))((...((((...((((((((...))))))))..))))...))...(.(((((..((((....jjjj...dddddddd...dddddddd...hhhhh......<..<...>...>.<...<...>...<...<....(((((((...)))))))...))))))))))((((.(((((...)))))(((((((...(((...eeeeeee...eeeeeee...hhhhh...ffff...ffffbbbbbbb......>...>...>...>... 3 (...))))...))))))).((((((((.((((((...)))))).))))))))..))...))....bbbbbbb... Match stalling sites by sequence comparison reconstruct structure Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
55 Nanopores Force-extension experiments Structure reconstruction II 3 P9 P9.1 P9.2 P9.0 P9.1a P8 P2 P2.1 5 P1 P7 lt P3 P5b P5a P5 P5c P6b P6a P4 3 P9 P9.1 P9.2 P9.0 P9.1a P8 P2 P2.1 5 P1 P7 P3 P5b P5a P5 P5c P6b P6a P4 Overall structure reconstructed correctly an distinguish different structures on the same sequence Even pseudoknot reconstructed Just needs to be implemented experimentally... Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
56 Outline of part II Summary 5 Force-extension experiments 6 Quantitative modeling 7 Results for simple force-extension experiments 8 Nanopores 9 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
57 Summary of part II Summary Quantitative description of single-molecule experiments possible Force-extension curves do not reveal secondary structure information due to compensation effects Single-molecule experiments reveal structure information with the help of a nanopore Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
58 Outline of part III 10 Motivation 11 Molecular Dynamics 12 Monte arlo simulation 13 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
59 Outline of part III Motivation 10 Motivation What is kinetics? Why care about kinetics? 11 Molecular Dynamics 12 Monte arlo simulation 13 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
60 Kinetics introduction Motivation What is kinetics? Thermodynamics What structure(s) does an RN molecule take on? Kinetics How long does it take an RN molecule to reach a certain structure? long which pathway does an RN molecule get to a certain structure? Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
61 Motivation Why is kinetics important? Why care about kinetics? Why is kinetics a problem? Take, say, a short RN with 50 bases Has roughly = structures Let s be generous and say that it can explore a structure per picosecond it takes 500,000,000s 18yr to explore all structures! Processes that need to happen in time: rrn have to fold riboswitches have to get from one configuration to another terminator hairpins have to form in time for termination RN viruses have to be folded for packaging splicing might depend on secondary structure Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
62 Outline of part III Molecular Dynamics 10 Motivation 11 Molecular Dynamics eneral principle Pros and cons 12 Monte arlo simulation 13 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
63 Molecular Dynamics What is Molecular Dynamics? eneral principle Model an RN molecule atom by atom dd water and salt atom by atom or as an effective medium hoose a force field Integrate Newton s equations Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
64 Molecular Dynamics Pros and cons dvantages and disadvantages of Molecular Dynamics dvantages First principle based modeling Full three dimensional structure included Disadvantages Need many atoms ( 30 atoms per base) an only simulate very short sequences (10 base pairs) an only simulate very short times (at most µs) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
65 Molecular Dynamics What can be done with MD? Pros and cons Structural dynamics of very small building blocks: Sorin et al., RN simulations: probing hairpin unfolding and the dynamics of a NR tetraloop, J. Mol. Biol Sarzynska et al., Effects of base substitutions in an RN hairpin from molecular dynamics and free energy simulations, Biophys J Hart et al., Molecular dynamics simulations and free energy calculations of base flipping in dsrn, RN Mazier et al., Molecular dynamics simulation for probing the flexibility of the 35 nucleotide SL1 sequence kissing complex from HIV-1Lai genomic RN, J Biomol Struct Dyn Nystrom et al.,molecular dynamics study of intrinsic stability in six RN terminal loop motifs, J Biomol Struct Dyn Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
66 Outline of part III Monte arlo simulation 10 Motivation 11 Molecular Dynamics 12 Monte arlo simulation Method Base pair level approach Helix level approach 13 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
67 Monte arlo simulation What is Monte arlo dynamics? Method Description of a system State space Energy for each state llowed transitions from state to state Procedure hoose random initial state S 0 Repeat as often as necessary for i = 1, 2, 3,...: Enumerate all states into which state S i 1 is allowed to transition. Pick one of these states T randomly alculate E E[S] E[T ] If E 0 let S i T If E < 0 pick a random number r If r < e E RT let S i T, otherwise let S i S i 1 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
68 Monte arlo simulation Method Properties of Monte arlo dynamics Boltzmann distribution Transitions have to be chosen such that every state can be reached from every other state The sequence S 0, S 1,... visits each state with a frequency proportional to its Boltzmann probability p = e E[S] RT /Z It avoids calculating a partition function Monte arlo and real time In general, Monte arlo dynamics has nothing to do with real dynamics of system Monte arlo dynamics is a good representation of real dynamics if: The allowed transitions correspond to true elementary steps of real dynamics There is a good physical reason for all downhill transitions occuring all with the same rate k 0 In this case, the real time is t = i/k 0 for the ith step Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
69 Numerical tricks Monte arlo simulation Method illespie sampling Instead of accepting or rejecting a move, choose a move in every step but increase time by a larger chunk if moves are unfavorable. (illespie, J. Phys. hem. 1977) State clustering If algorithm revisits the same state in a local minimum over and over again, precalculate the distribution and residence time in local valley by diagonalizing a reasonably sized matrix and directly choose a move that leads out of the valley. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
70 The Vienna approach Monte arlo simulation Base pair level approach Specification States: all secondary structures Energies: calculated by Turner model Transitions: Opening of a base pair losing of a base pair Repairing of a base Flamm et al., RN folding at elementary step resolution, RN 2000 kinfold program in Vienna package Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
71 Properties Monte arlo simulation Base pair level approach Validity Every state can be reached from every other Transitions are reasonable elementary steps losing of a base is an activated process itself with a roughly constant free energy barrier Time scales Time for closing a base pair experimentally determined to be 1µs Monte arlo time is real time in µs Time for closing a hairpin of length 4: e E[S] RT µs e µs 1ms Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
72 Monte arlo simulation Isambert and Siggia approach Helix level approach Specification Precompute all possible helices of minimum length 3 States: all possible combinations of non-overlapping helices (Note: this includes pseudo-knots!) Energy: Turner stacking energies for stacks stick and string model for loop entropies Transitions: formation and removal of a complete helix Isambert and Siggia, Modeling RN folding paths with pseudoknots: pplication to hepatitis delta virus ribozyme, PNS 2000 Xayaphoummine et al., Kinefold web server for RN/DN folding path and structure prediction including pseudoknots and knots, Nucleic cids Research 2005 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
73 Results I Monte arlo simulation Helix level approach Pseudoknots Test on 7 relatively short RN molecules with experimentally known pseudoknots For 5 out of the 7 correct pseudo-knotted is found For 1 the pseudo-knotted structure is only marginally higher than minimum energy structure For 1 specific Mg binding is suspected Isambert and Siggia, PNS 2000 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
74 Results II Monte arlo simulation Helix level approach Kinetics Folding of HDV ribozyme 1/3 of the time folds within a fraction of a second 2/3 of the time trapped for up to a minute Molecule has to be functional after 4s to self-cleave! If folding co-transcriptional only 10% of molecules get trapped Isambert and Siggia, PNS 2000 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
75 Outline of part III Summary 10 Motivation 11 Molecular Dynamics 12 Monte arlo simulation 13 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
76 Summary of part III Summary Dynamics of RN folding is important for many biological processes Molecular dynamics is useful to study fast structural changes of small substructures Monte arlo simulations can describe folding of realistically sized molecules Monte arlo simulations can describe folding including pseudo-knots The elementary time scale for closing of a base pair on top of an existing step is on the order of 1µs Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
77 Outline of part IV 14 Introduction to microrns 15 First generation target prediction 16 urrent target prediction methods 17 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
78 Outline of part IV Introduction to microrns 14 Introduction to microrns What are microrns Biogenesis Functions omputational problems 15 First generation target prediction 16 urrent target prediction methods 17 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
79 Introduction to microrns What is a micro RN? What are microrns Definition microrn is an approximately 22 nucleotide long RN that regulates expression of other genes Properties microrn are post-transcriptional regulators microrn themselves are temporally and spatially regulated There are most likely hundreds of microrn in higher eukaryotes Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
80 Introduction to microrns How are microrn made? Biogenesis microrn is made in four steps: 1 Transcription of the primary RN (pri-mirn) by RN polymerase (in the nucleus) 2 Processing into hairpin-like RNs called pre-mirn (in the nucleus) 3 leavage into the mature microrn by Dicer (in the cytoplasm) 4 Incorporation into the RN-induced silencing complex (RIS) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
81 Introduction to microrns What do microrn do? Functions In plants microrns in RIS complexes bind to exactly complementary regions of the mrns of target genes The mrn of the target gene is cut and degraded In animals microrns in RIS complexes bind to partially complementary regions of the mrns of target genes The mrn of the target gene is prevented from translation Processes microrns are known to be involved in Development Immune system ancer Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
82 Introduction to microrns omputational problems omputational problems ene finding problem iven a genome sequence, predict the microrn sequences of the organism (in plants and animals). Target prediction problem iven a microrn sequence, predict the genes that are regulated by that microrn (in animals). Here, only talk about the target prediction problem Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
83 Outline of part IV First generation target prediction 14 Introduction to microrns 15 First generation target prediction pproaches lgorithm Results 16 urrent target prediction methods 17 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
84 First generation target prediction pproaches for Drosophila pproaches Many algorithms proposed essentially simultaneously For Drosophila: Stark et al., Identification of Drosophila microrn targets, PLoS Biollogy (2003) Enright et al., MicroRN targets in Drosophila, enome Biology (2003) Rajewsky and Socci, omputational identification of microrn targets, Developmental Biology (2004) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
85 First generation target prediction pproaches for mammals pproaches For mammals: Lewis et al., Prediction of mammalian microrn targets, ell (2003) John et al., Human MicroRN targets, PLoS Biology (2004) Kiriakidou et al., combined computational-experimental approach predicts human microrn targets, enes & Development (2004) ll approaches rather similar, but resulting predictions have only small overlap Here: Rajewsky and Socci Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
86 First generation target prediction lgorithm Features used in target prediction Observations: 1 MicroRN target sites are in 3 TRs. 2 lthough microrns are not exactly complementary to their targets they always contain a nucleus of length 6-8 bases with exact complementarity. 3 Even in the not exactly complementary regions, there is still a lot of stabilizing base pairing. 4 MicroRN target relationships tend to be conserved across species. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
87 First generation target prediction Nucleus identification lgorithm sing observation 2: there is a nucleus ssign to every base pairing stretch a score by the formula score = w # base pairs + w # b.p. + w # b.p. se training sets of true targets and of true non-targets to choose w, w, and w such that they maximize the distance between expected scores of targets and non-targets w = 5, w = 2, w = 0 Fix a p-value threshold Score many random mrn sequences for a given microrn sequence and determine the score cutoff that belongs to the p-value cutoff Score all 3 TR regions of interest and keep only those for which the score exceeds the threshold. Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
88 sing other features First generation target prediction lgorithm sing observation 3: other bases pair as well ut 40 base window around identified nucleus se MFOLD to predict binding energy of microrn-mrn hybrid Keep only candidate targets with binding energy below 17.4kcal/mol sing observation 4: targets are conserved Keep only candidates for which a target is predicted in D. melanogaster as well as in D. pseudoobscura Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
89 First generation target prediction Results Results Test1 Search D. melanogaster hid transcript (not used in training) for bantam sites: 4 of the 5 experimentally known sites found, no false positive Test2 Prediction of target sites for 74 D. melanogaster microrns in 31 body patterning genes: found 39 putative target sites, some of them make biological sense Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
90 Outline of part IV urrent target prediction methods 14 Introduction to microrns 15 First generation target prediction 16 urrent target prediction methods dditional features Performance 17 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
91 urrent target prediction methods dditional features dditional features used for prediction Position of nucleus in microrn (5 end) onservation of nucleus in large multiple sequence alignments bsence of conservation outside nucleus n immediately upstream of the nucleus ombinatorics of target sites for multiple microrns Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
92 urrent target prediction methods Performance comparison Performance ssay Evaluate prediction method on 133 experimentally tested targets. (Stark, Brennecke, Bushati, Russell, ohen, ell 2005) Results Best methods: TargetScanS (Stark et al.) PicTar (rün et al.) Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
93 Outline of part IV Summary 14 Introduction to microrns 15 First generation target prediction 16 urrent target prediction methods 17 Summary Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
94 Summary of part IV Summary microrns are a relatively new but important class of regulatory RNs There are many programs for microrn target prediction The main features used for target prediction are stability of a nucleus, stability of the full hybrid, and conservation between species Best current algorithms (TargetScanS and PicTar) have about 90% accuracy and 70% 80% sensitivity Reviews: Issac Bentwich, Prediction of microrns and their targets, FEBS Letters 2005 Nikolaus Rajewksy, microrn target predictions in animals, Nature enetics 2006 Ralf Bundschuh (Ohio State niversity) Modelling RN structure July 25, / 98
Quantitative modeling of RNA single-molecule experiments. Ralf Bundschuh Department of Physics, Ohio State University
 Quantitative modeling of RN single-molecule experiments Ralf Bundschuh Department of Physics, Ohio State niversity ollaborators: lrich erland, LM München Terence Hwa, San Diego Outline: Single-molecule
Quantitative modeling of RN single-molecule experiments Ralf Bundschuh Department of Physics, Ohio State niversity ollaborators: lrich erland, LM München Terence Hwa, San Diego Outline: Single-molecule
Statistical Physics of RNA folding
 Statistical Physics of RN folding Ralf Bundschuh ollaborators: Terry Hwa, SD, lrich erland, niversität zu Köln, Robijn Bruinsma, L Ohio State niversity March 18, 2008 Ralf Bundschuh (Ohio State niversity)
Statistical Physics of RN folding Ralf Bundschuh ollaborators: Terry Hwa, SD, lrich erland, niversität zu Köln, Robijn Bruinsma, L Ohio State niversity March 18, 2008 Ralf Bundschuh (Ohio State niversity)
proteins are the basic building blocks and active players in the cell, and
 12 RN Secondary Structure Sources for this lecture: R. Durbin, S. Eddy,. Krogh und. Mitchison, Biological sequence analysis, ambridge, 1998 J. Setubal & J. Meidanis, Introduction to computational molecular
12 RN Secondary Structure Sources for this lecture: R. Durbin, S. Eddy,. Krogh und. Mitchison, Biological sequence analysis, ambridge, 1998 J. Setubal & J. Meidanis, Introduction to computational molecular
Mechanically probing the folding pathway of single RNA molecules
 Mechanically probing the folding pathway of single RN molecules lrich erland, Ralf Bundschuh, and Terence Hwa Department of Physics, niversity of alifornia at San Diego, La Jolla, alifornia 9293-319 and
Mechanically probing the folding pathway of single RN molecules lrich erland, Ralf Bundschuh, and Terence Hwa Department of Physics, niversity of alifornia at San Diego, La Jolla, alifornia 9293-319 and
98 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 6, 2006
 98 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 6, 2006 8.3.1 Simple energy minimization Maximizing the number of base pairs as described above does not lead to good structure predictions.
98 Algorithms in Bioinformatics I, WS 06, ZBIT, D. Huson, December 6, 2006 8.3.1 Simple energy minimization Maximizing the number of base pairs as described above does not lead to good structure predictions.
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri RNA Structure Prediction Secondary
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri RNA Structure Prediction Secondary
RNA Abstract Shape Analysis
 ourse: iegerich RN bstract nalysis omplete shape iegerich enter of Biotechnology Bielefeld niversity robert@techfak.ni-bielefeld.de ourse on omputational RN Biology, Tübingen, March 2006 iegerich ourse:
ourse: iegerich RN bstract nalysis omplete shape iegerich enter of Biotechnology Bielefeld niversity robert@techfak.ni-bielefeld.de ourse on omputational RN Biology, Tübingen, March 2006 iegerich ourse:
A Novel Statistical Model for the Secondary Structure of RNA
 ISBN 978-1-8466-93-3 Proceedings of the 5th International ongress on Mathematical Biology (IMB11) Vol. 3 Nanjing, P. R. hina, June 3-5, 11 Novel Statistical Model for the Secondary Structure of RN Liu
ISBN 978-1-8466-93-3 Proceedings of the 5th International ongress on Mathematical Biology (IMB11) Vol. 3 Nanjing, P. R. hina, June 3-5, 11 Novel Statistical Model for the Secondary Structure of RN Liu
DNA/RNA Structure Prediction
 C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Master Course DNA/Protein Structurefunction Analysis and Prediction Lecture 12 DNA/RNA Structure Prediction Epigenectics Epigenomics:
C E N T R E F O R I N T E G R A T I V E B I O I N F O R M A T I C S V U Master Course DNA/Protein Structurefunction Analysis and Prediction Lecture 12 DNA/RNA Structure Prediction Epigenectics Epigenomics:
Combinatorial approaches to RNA folding Part II: Energy minimization via dynamic programming
 ombinatorial approaches to RNA folding Part II: Energy minimization via dynamic programming Matthew Macauley Department of Mathematical Sciences lemson niversity http://www.math.clemson.edu/~macaule/ Math
ombinatorial approaches to RNA folding Part II: Energy minimization via dynamic programming Matthew Macauley Department of Mathematical Sciences lemson niversity http://www.math.clemson.edu/~macaule/ Math
RNA Secondary Structure Prediction
 RN Secondary Structure Prediction Perry Hooker S 531: dvanced lgorithms Prof. Mike Rosulek University of Montana December 10, 2010 Introduction Ribonucleic acid (RN) is a macromolecule that is essential
RN Secondary Structure Prediction Perry Hooker S 531: dvanced lgorithms Prof. Mike Rosulek University of Montana December 10, 2010 Introduction Ribonucleic acid (RN) is a macromolecule that is essential
RNA-Strukturvorhersage Strukturelle Bioinformatik WS16/17
 RNA-Strukturvorhersage Strukturelle Bioinformatik WS16/17 Dr. Stefan Simm, 01.11.2016 simm@bio.uni-frankfurt.de RNA secondary structures a. hairpin loop b. stem c. bulge loop d. interior loop e. multi
RNA-Strukturvorhersage Strukturelle Bioinformatik WS16/17 Dr. Stefan Simm, 01.11.2016 simm@bio.uni-frankfurt.de RNA secondary structures a. hairpin loop b. stem c. bulge loop d. interior loop e. multi
GCD3033:Cell Biology. Transcription
 Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Transcription Transcription: DNA to RNA A) production of complementary strand of DNA B) RNA types C) transcription start/stop signals D) Initiation of eukaryotic gene expression E) transcription factors
Elucidation of the RNA-folding mechanism at the level of both
 RNA hairpin-folding kinetics Wenbing Zhang and Shi-Jie Chen* Department of Physics and Astronomy and Department of Biochemistry, University of Missouri, Columbia, MO 65211 Edited by Peter G. Wolynes, University
RNA hairpin-folding kinetics Wenbing Zhang and Shi-Jie Chen* Department of Physics and Astronomy and Department of Biochemistry, University of Missouri, Columbia, MO 65211 Edited by Peter G. Wolynes, University
Predicting RNA Secondary Structure
 7.91 / 7.36 / BE.490 Lecture #6 Mar. 11, 2004 Predicting RNA Secondary Structure Chris Burge Review of Markov Models & DNA Evolution CpG Island HMM The Viterbi Algorithm Real World HMMs Markov Models for
7.91 / 7.36 / BE.490 Lecture #6 Mar. 11, 2004 Predicting RNA Secondary Structure Chris Burge Review of Markov Models & DNA Evolution CpG Island HMM The Viterbi Algorithm Real World HMMs Markov Models for
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION
 THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
Conserved RNA Structures. Ivo L. Hofacker. Institut for Theoretical Chemistry, University Vienna.
 onserved RN Structures Ivo L. Hofacker Institut for Theoretical hemistry, University Vienna http://www.tbi.univie.ac.at/~ivo/ Bled, January 2002 Energy Directed Folding Predict structures from sequence
onserved RN Structures Ivo L. Hofacker Institut for Theoretical hemistry, University Vienna http://www.tbi.univie.ac.at/~ivo/ Bled, January 2002 Energy Directed Folding Predict structures from sequence
Biology Chemistry & Physics of Biomolecules. Examination #1. Proteins Module. September 29, Answer Key
 Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Biology 5357 Chemistry & Physics of Biomolecules Examination #1 Proteins Module September 29, 2017 Answer Key Question 1 (A) (5 points) Structure (b) is more common, as it contains the shorter connection
Bi 8 Lecture 11. Quantitative aspects of transcription factor binding and gene regulatory circuit design. Ellen Rothenberg 9 February 2016
 Bi 8 Lecture 11 Quantitative aspects of transcription factor binding and gene regulatory circuit design Ellen Rothenberg 9 February 2016 Major take-home messages from λ phage system that apply to many
Bi 8 Lecture 11 Quantitative aspects of transcription factor binding and gene regulatory circuit design Ellen Rothenberg 9 February 2016 Major take-home messages from λ phage system that apply to many
The Double Helix. CSE 417: Algorithms and Computational Complexity! The Central Dogma of Molecular Biology! DNA! RNA! Protein! Protein!
 The Double Helix SE 417: lgorithms and omputational omplexity! Winter 29! W. L. Ruzzo! Dynamic Programming, II" RN Folding! http://www.rcsb.org/pdb/explore.do?structureid=1t! Los lamos Science The entral
The Double Helix SE 417: lgorithms and omputational omplexity! Winter 29! W. L. Ruzzo! Dynamic Programming, II" RN Folding! http://www.rcsb.org/pdb/explore.do?structureid=1t! Los lamos Science The entral
RNA Basics. RNA bases A,C,G,U Canonical Base Pairs A-U G-C G-U. Bases can only pair with one other base. wobble pairing. 23 Hydrogen Bonds more stable
 RNA STRUCTURE RNA Basics RNA bases A,C,G,U Canonical Base Pairs A-U G-C G-U wobble pairing Bases can only pair with one other base. 23 Hydrogen Bonds more stable RNA Basics transfer RNA (trna) messenger
RNA STRUCTURE RNA Basics RNA bases A,C,G,U Canonical Base Pairs A-U G-C G-U wobble pairing Bases can only pair with one other base. 23 Hydrogen Bonds more stable RNA Basics transfer RNA (trna) messenger
 Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Lecture 7: Simple genetic circuits I
 Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Lecture 7: Simple genetic circuits I Paul C Bressloff (Fall 2018) 7.1 Transcription and translation In Fig. 20 we show the two main stages in the expression of a single gene according to the central dogma.
Combinatorial approaches to RNA folding Part I: Basics
 Combinatorial approaches to RNA folding Part I: Basics Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2015 M. Macauley (Clemson)
Combinatorial approaches to RNA folding Part I: Basics Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2015 M. Macauley (Clemson)
2012 Univ Aguilera Lecture. Introduction to Molecular and Cell Biology
 2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
2012 Univ. 1301 Aguilera Lecture Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the
Computational RNA Secondary Structure Design:
 omputational RN Secondary Structure Design: Empirical omplexity and Improved Methods by Rosalía guirre-hernández B.Sc., niversidad Nacional utónoma de México, 996 M. Eng., niversidad Nacional utónoma de
omputational RN Secondary Structure Design: Empirical omplexity and Improved Methods by Rosalía guirre-hernández B.Sc., niversidad Nacional utónoma de México, 996 M. Eng., niversidad Nacional utónoma de
Chapter 6- An Introduction to Metabolism*
 Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Chapter 6- An Introduction to Metabolism* *Lecture notes are to be used as a study guide only and do not represent the comprehensive information you will need to know for the exams. The Energy of Life
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
EVALUATION OF RNA SECONDARY STRUCTURE MOTIFS USING REGRESSION ANALYSIS
 EVLTION OF RN SEONDRY STRTRE MOTIFS SIN RERESSION NLYSIS Mohammad nwar School of Information Technology and Engineering, niversity of Ottawa e-mail: manwar@site.uottawa.ca bstract Recent experimental evidences
EVLTION OF RN SEONDRY STRTRE MOTIFS SIN RERESSION NLYSIS Mohammad nwar School of Information Technology and Engineering, niversity of Ottawa e-mail: manwar@site.uottawa.ca bstract Recent experimental evidences
DANNY BARASH ABSTRACT
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 11, Number 6, 2004 Mary Ann Liebert, Inc. Pp. 1169 1174 Spectral Decomposition for the Search and Analysis of RNA Secondary Structure DANNY BARASH ABSTRACT Scales
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 11, Number 6, 2004 Mary Ann Liebert, Inc. Pp. 1169 1174 Spectral Decomposition for the Search and Analysis of RNA Secondary Structure DANNY BARASH ABSTRACT Scales
Master equation approach to finding the rate-limiting steps in biopolymer folding
 JOURNAL OF CHEMICAL PHYSICS VOLUME 118, NUMBER 7 15 FEBRUARY 2003 Master equation approach to finding the rate-limiting steps in biopolymer folding Wenbing Zhang and Shi-Jie Chen a) Department of Physics
JOURNAL OF CHEMICAL PHYSICS VOLUME 118, NUMBER 7 15 FEBRUARY 2003 Master equation approach to finding the rate-limiting steps in biopolymer folding Wenbing Zhang and Shi-Jie Chen a) Department of Physics
RNA Secondary Structures: A Tractable Model of Biopolymer Folding
 R Secondary Structures: Tractable Model of Biopolymer Folding Ivo L. Hofacker Institut für Theoretische hemie der niversität Wien, Währingerstr. 17, 1090 Wien, ustria E-mail: ivo@tbi.univie.ac.at R secondary
R Secondary Structures: Tractable Model of Biopolymer Folding Ivo L. Hofacker Institut für Theoretische hemie der niversität Wien, Währingerstr. 17, 1090 Wien, ustria E-mail: ivo@tbi.univie.ac.at R secondary
Introduction to Molecular and Cell Biology
 Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Introduction to Molecular and Cell Biology Molecular biology seeks to understand the physical and chemical basis of life. and helps us answer the following? What is the molecular basis of disease? What
Protein folding. α-helix. Lecture 21. An α-helix is a simple helix having on average 10 residues (3 turns of the helix)
 Computat onal Biology Lecture 21 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Computat onal Biology Lecture 21 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
2.4 DNA structure. S(l) bl + c log l + d, with c 1.8k B. (2.72)
 2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
Computational Approaches for determination of Most Probable RNA Secondary Structure Using Different Thermodynamics Parameters
 Computational Approaches for determination of Most Probable RNA Secondary Structure Using Different Thermodynamics Parameters 1 Binod Kumar, Assistant Professor, Computer Sc. Dept, ISTAR, Vallabh Vidyanagar,
Computational Approaches for determination of Most Probable RNA Secondary Structure Using Different Thermodynamics Parameters 1 Binod Kumar, Assistant Professor, Computer Sc. Dept, ISTAR, Vallabh Vidyanagar,
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
A graph kernel approach to the identification and characterisation of structured non-coding RNAs using multiple sequence alignment information
 graph kernel approach to the identification and characterisation of structured noncoding RNs using multiple sequence alignment information Mariam lshaikh lbert Ludwigs niversity Freiburg, Department of
graph kernel approach to the identification and characterisation of structured noncoding RNs using multiple sequence alignment information Mariam lshaikh lbert Ludwigs niversity Freiburg, Department of
Lab III: Computational Biology and RNA Structure Prediction. Biochemistry 208 David Mathews Department of Biochemistry & Biophysics
 Lab III: Computational Biology and RNA Structure Prediction Biochemistry 208 David Mathews Department of Biochemistry & Biophysics Contact Info: David_Mathews@urmc.rochester.edu Phone: x51734 Office: 3-8816
Lab III: Computational Biology and RNA Structure Prediction Biochemistry 208 David Mathews Department of Biochemistry & Biophysics Contact Info: David_Mathews@urmc.rochester.edu Phone: x51734 Office: 3-8816
S(l) bl + c log l + d, with c 1.8k B. (2.71)
 2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
BIOINFORMATICS. Prediction of RNA secondary structure based on helical regions distribution
 BIOINFORMATICS Prediction of RNA secondary structure based on helical regions distribution Abstract Motivation: RNAs play an important role in many biological processes and knowing their structure is important
BIOINFORMATICS Prediction of RNA secondary structure based on helical regions distribution Abstract Motivation: RNAs play an important role in many biological processes and knowing their structure is important
Impact Of The Energy Model On The Complexity Of RNA Folding With Pseudoknots
 Impact Of The Energy Model On The omplexity Of RN Folding With Pseudoknots Saad Sheikh, Rolf Backofen Yann Ponty, niversity of Florida, ainesville, S lbert Ludwigs niversity, Freiburg, ermany LIX, NRS/Ecole
Impact Of The Energy Model On The omplexity Of RN Folding With Pseudoknots Saad Sheikh, Rolf Backofen Yann Ponty, niversity of Florida, ainesville, S lbert Ludwigs niversity, Freiburg, ermany LIX, NRS/Ecole
CSEP 590A Summer Tonight MLE. FYI, re HW #2: Hemoglobin History. Lecture 4 MLE, EM, RE, Expression. Maximum Likelihood Estimators
 CSEP 59A Summer 26 Lecture 4 MLE, EM, RE, Expression FYI, re HW #2: Hemoglobin History 1 Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
CSEP 59A Summer 26 Lecture 4 MLE, EM, RE, Expression FYI, re HW #2: Hemoglobin History 1 Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
CSEP 590A Summer Lecture 4 MLE, EM, RE, Expression
 CSEP 590A Summer 2006 Lecture 4 MLE, EM, RE, Expression 1 FYI, re HW #2: Hemoglobin History Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
CSEP 590A Summer 2006 Lecture 4 MLE, EM, RE, Expression 1 FYI, re HW #2: Hemoglobin History Alberts et al., 3rd ed.,pg389 2 Tonight MLE: Maximum Likelihood Estimators EM: the Expectation Maximization Algorithm
Introduction to" Protein Structure
 Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Introduction to" Protein Structure Function, evolution & experimental methods Thomas Blicher, Center for Biological Sequence Analysis Learning Objectives Outline the basic levels of protein structure.
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Predicting free energy landscapes for complexes of double-stranded chain molecules
 JOURNAL OF CHEMICAL PHYSICS VOLUME 114, NUMBER 9 1 MARCH 2001 Predicting free energy landscapes for complexes of double-stranded chain molecules Wenbing Zhang and Shi-Jie Chen a) Department of Physics
JOURNAL OF CHEMICAL PHYSICS VOLUME 114, NUMBER 9 1 MARCH 2001 Predicting free energy landscapes for complexes of double-stranded chain molecules Wenbing Zhang and Shi-Jie Chen a) Department of Physics
Multiple Choice Review- Eukaryotic Gene Expression
 Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Multiple Choice Review- Eukaryotic Gene Expression 1. Which of the following is the Central Dogma of cell biology? a. DNA Nucleic Acid Protein Amino Acid b. Prokaryote Bacteria - Eukaryote c. Atom Molecule
Grand Plan. RNA very basic structure 3D structure Secondary structure / predictions The RNA world
 Grand Plan RNA very basic structure 3D structure Secondary structure / predictions The RNA world very quick Andrew Torda, April 2017 Andrew Torda 10/04/2017 [ 1 ] Roles of molecules RNA DNA proteins genetic
Grand Plan RNA very basic structure 3D structure Secondary structure / predictions The RNA world very quick Andrew Torda, April 2017 Andrew Torda 10/04/2017 [ 1 ] Roles of molecules RNA DNA proteins genetic
A Method for Aligning RNA Secondary Structures
 Method for ligning RN Secondary Structures Jason T. L. Wang New Jersey Institute of Technology J Liu, JTL Wang, J Hu and B Tian, BM Bioinformatics, 2005 1 Outline Introduction Structural alignment of RN
Method for ligning RN Secondary Structures Jason T. L. Wang New Jersey Institute of Technology J Liu, JTL Wang, J Hu and B Tian, BM Bioinformatics, 2005 1 Outline Introduction Structural alignment of RN
Bio nformatics. Lecture 23. Saad Mneimneh
 Bio nformatics Lecture 23 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Bio nformatics Lecture 23 Protein folding The goal is to determine the three-dimensional structure of a protein based on its amino acid sequence Assumption: amino acid sequence completely and uniquely
Secondary structure formation of homopolymeric single-stranded nucleic acids including force and loop entropy: implications for DNA hybridization
 EPJ manuscript No. (will be inserted by the editor) Secondary structure formation of homopolymeric single-stranded nucleic acids including force and loop entropy: implications for DNA hybridization Thomas
EPJ manuscript No. (will be inserted by the editor) Secondary structure formation of homopolymeric single-stranded nucleic acids including force and loop entropy: implications for DNA hybridization Thomas
Sparse RNA Folding Revisited: Space-Efficient Minimum Free Energy Prediction
 Sparse RNA Folding Revisited: Space-Efficient Minimum Free Energy Prediction Sebastian Will 1 and Hosna Jabbari 2 1 Bioinformatics/IZBI, University Leipzig, swill@csail.mit.edu 2 Ingenuity Lab, National
Sparse RNA Folding Revisited: Space-Efficient Minimum Free Energy Prediction Sebastian Will 1 and Hosna Jabbari 2 1 Bioinformatics/IZBI, University Leipzig, swill@csail.mit.edu 2 Ingenuity Lab, National
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
Short Announcements. 1 st Quiz today: 15 minutes. Homework 3: Due next Wednesday.
 Short Announcements 1 st Quiz today: 15 minutes Homework 3: Due next Wednesday. Next Lecture, on Visualizing Molecular Dynamics (VMD) by Klaus Schulten Today s Lecture: Protein Folding, Misfolding, Aggregation
Short Announcements 1 st Quiz today: 15 minutes Homework 3: Due next Wednesday. Next Lecture, on Visualizing Molecular Dynamics (VMD) by Klaus Schulten Today s Lecture: Protein Folding, Misfolding, Aggregation
RNA Protein Interaction
 Introduction RN binding in detail structural analysis Examples RN Protein Interaction xel Wintsche February 16, 2009 xel Wintsche RN Protein Interaction Introduction RN binding in detail structural analysis
Introduction RN binding in detail structural analysis Examples RN Protein Interaction xel Wintsche February 16, 2009 xel Wintsche RN Protein Interaction Introduction RN binding in detail structural analysis
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Lecture 11: Protein Folding & Stability
 Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Structure - Function Protein Folding: What we know Lecture 11: Protein Folding & Stability 1). Amino acid sequence dictates structure. 2). The native structure represents the lowest energy state for a
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall Protein Folding: What we know. Protein Folding
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2003 Structure - Function Protein Folding: What we know 1). Amino acid sequence dictates structure. 2). The native structure represents
arxiv:cond-mat/ v1 [cond-mat.soft] 25 Jul 2002
![arxiv:cond-mat/ v1 [cond-mat.soft] 25 Jul 2002 arxiv:cond-mat/ v1 [cond-mat.soft] 25 Jul 2002](/thumbs/80/82151644.jpg) Slow nucleic acid unzipping kinetics from sequence-defined barriers S. Cocco, J.F. Marko, R. Monasson CNRS Laboratoire de Dynamique des Fluides Complexes, rue de l Université, 67 Strasbourg, France; Department
Slow nucleic acid unzipping kinetics from sequence-defined barriers S. Cocco, J.F. Marko, R. Monasson CNRS Laboratoire de Dynamique des Fluides Complexes, rue de l Université, 67 Strasbourg, France; Department
Many proteins spontaneously refold into native form in vitro with high fidelity and high speed.
 Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Macromolecular Processes 20. Protein Folding Composed of 50 500 amino acids linked in 1D sequence by the polypeptide backbone The amino acid physical and chemical properties of the 20 amino acids dictate
Structural biomathematics: an overview of molecular simulations and protein structure prediction
 : an overview of molecular simulations and protein structure prediction Figure: Parc de Recerca Biomèdica de Barcelona (PRBB). Contents 1 A Glance at Structural Biology 2 3 1 A Glance at Structural Biology
: an overview of molecular simulations and protein structure prediction Figure: Parc de Recerca Biomèdica de Barcelona (PRBB). Contents 1 A Glance at Structural Biology 2 3 1 A Glance at Structural Biology
Olle Inganäs: Polymers structure and dynamics. Polymer physics
 Polymer physics Polymers are macromolecules formed by many identical monomers, connected through covalent bonds, to make a linear chain of mers a polymer. The length of the chain specifies the weight of
Polymer physics Polymers are macromolecules formed by many identical monomers, connected through covalent bonds, to make a linear chain of mers a polymer. The length of the chain specifies the weight of
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Lecture 12. DNA/RNA Structure Prediction. Epigenectics Epigenomics: Gene Expression
 C N F O N G A V B O N F O M A C S V U Master Course DNA/Protein Structurefunction Analysis and Prediction Lecture 12 DNA/NA Structure Prediction pigenectics pigenomics: Gene xpression ranscription factors
C N F O N G A V B O N F O M A C S V U Master Course DNA/Protein Structurefunction Analysis and Prediction Lecture 12 DNA/NA Structure Prediction pigenectics pigenomics: Gene xpression ranscription factors
BME 5742 Biosystems Modeling and Control
 BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
BME 5742 Biosystems Modeling and Control Lecture 24 Unregulated Gene Expression Model Dr. Zvi Roth (FAU) 1 The genetic material inside a cell, encoded in its DNA, governs the response of a cell to various
Origin of the Electrophoretic Force on DNA in Nanopores. Biological and Soft Systems - Cavendish Laboratory
 Origin of the Electrophoretic Force on DNA in Nanopores Ulrich F. Keyser Biological and Soft Systems - Cavendish Laboratory Acknowledgements Delft Cees Dekker, Nynke H. Dekker, Serge G. Lemay R. Smeets,
Origin of the Electrophoretic Force on DNA in Nanopores Ulrich F. Keyser Biological and Soft Systems - Cavendish Laboratory Acknowledgements Delft Cees Dekker, Nynke H. Dekker, Serge G. Lemay R. Smeets,
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 8: RNA folding
 1/16, Lecture 8: RNA folding Hamidreza Chitsaz Colorado State University chitsaz@cs.colostate.edu Fall 2018 September 13, 2018 2/16, Nearest Neighbor Model 3/16, McCaskill s Algorithm for MFE Structure
1/16, Lecture 8: RNA folding Hamidreza Chitsaz Colorado State University chitsaz@cs.colostate.edu Fall 2018 September 13, 2018 2/16, Nearest Neighbor Model 3/16, McCaskill s Algorithm for MFE Structure
F. Piazza Center for Molecular Biophysics and University of Orléans, France. Selected topic in Physical Biology. Lecture 1
 Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
Zhou Pei-Yuan Centre for Applied Mathematics, Tsinghua University November 2013 F. Piazza Center for Molecular Biophysics and University of Orléans, France Selected topic in Physical Biology Lecture 1
The Multistrand Simulator: Stochastic Simulation of the Kinetics of Multiple Interacting DNA Strands
 The Multistrand Simulator: Stochastic Simulation of the Kinetics of Multiple Interacting DNA Strands Joseph Schaeffer, Caltech (slides by John Reif) Entropy: Is a measure of disorder or randomness of a
The Multistrand Simulator: Stochastic Simulation of the Kinetics of Multiple Interacting DNA Strands Joseph Schaeffer, Caltech (slides by John Reif) Entropy: Is a measure of disorder or randomness of a
Lecture 8: RNA folding
 1/16, Lecture 8: RNA folding Hamidreza Chitsaz Colorado State University chitsaz@cs.colostate.edu Spring 2018 February 15, 2018 2/16, Nearest Neighbor Model 3/16, McCaskill s Algorithm for MFE Structure
1/16, Lecture 8: RNA folding Hamidreza Chitsaz Colorado State University chitsaz@cs.colostate.edu Spring 2018 February 15, 2018 2/16, Nearest Neighbor Model 3/16, McCaskill s Algorithm for MFE Structure
Predicting Protein Functions and Domain Interactions from Protein Interactions
 Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Predicting Protein Functions and Domain Interactions from Protein Interactions Fengzhu Sun, PhD Center for Computational and Experimental Genomics University of Southern California Outline High-throughput
Molecular Mechanics, Dynamics & Docking
 Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Molecular Mechanics, Dynamics & Docking Lawrence Hunter, Ph.D. Director, Computational Bioscience Program University of Colorado School of Medicine Larry.Hunter@uchsc.edu http://compbio.uchsc.edu/hunter
Thermodynamics. Entropy and its Applications. Lecture 11. NC State University
 Thermodynamics Entropy and its Applications Lecture 11 NC State University System and surroundings Up to this point we have considered the system, but we have not concerned ourselves with the relationship
Thermodynamics Entropy and its Applications Lecture 11 NC State University System and surroundings Up to this point we have considered the system, but we have not concerned ourselves with the relationship
Prediction of Conserved and Consensus RNA Structures
 Prediction of onserved and onsensus RN Structures DISSERTTION zur Erlangung des akademischen rades Doctor rerum naturalium Vorgelegt der Fakultät für Naturwissenschaften und Mathematik der niversität Wien
Prediction of onserved and onsensus RN Structures DISSERTTION zur Erlangung des akademischen rades Doctor rerum naturalium Vorgelegt der Fakultät für Naturwissenschaften und Mathematik der niversität Wien
Free energy, electrostatics, and the hydrophobic effect
 Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
Protein Physics 2016 Lecture 3, January 26 Free energy, electrostatics, and the hydrophobic effect Magnus Andersson magnus.andersson@scilifelab.se Theoretical & Computational Biophysics Recap Protein structure
10-810: Advanced Algorithms and Models for Computational Biology. microrna and Whole Genome Comparison
 10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
10-810: Advanced Algorithms and Models for Computational Biology microrna and Whole Genome Comparison Central Dogma: 90s Transcription factors DNA transcription mrna translation Proteins Central Dogma:
Measurements of interaction forces in (biological) model systems
 Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
The RNA World, Fitness Landscapes, and all that
 The RN World, Fitness Landscapes, and all that Peter F. Stadler Bioinformatics roup, Dept. of omputer Science & Interdisciplinary enter for Bioinformatics, niversity of Leipzig RNomics roup, Fraunhofer
The RN World, Fitness Landscapes, and all that Peter F. Stadler Bioinformatics roup, Dept. of omputer Science & Interdisciplinary enter for Bioinformatics, niversity of Leipzig RNomics roup, Fraunhofer
Models for Single Molecule Experiments
 Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
Free energy recovery in single molecule experiments
 Supplementary Material Free energy recovery in single molecule experiments Single molecule force measurements (experimental setup shown in Fig. S1) can be used to determine free-energy differences between
Supplementary Material Free energy recovery in single molecule experiments Single molecule force measurements (experimental setup shown in Fig. S1) can be used to determine free-energy differences between
Protein Folding Prof. Eugene Shakhnovich
 Protein Folding Eugene Shakhnovich Department of Chemistry and Chemical Biology Harvard University 1 Proteins are folded on various scales As of now we know hundreds of thousands of sequences (Swissprot)
Protein Folding Eugene Shakhnovich Department of Chemistry and Chemical Biology Harvard University 1 Proteins are folded on various scales As of now we know hundreds of thousands of sequences (Swissprot)
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water?
 Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Can a continuum solvent model reproduce the free energy landscape of a β-hairpin folding in water? Ruhong Zhou 1 and Bruce J. Berne 2 1 IBM Thomas J. Watson Research Center; and 2 Department of Chemistry,
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Chapter 9 DNA recognition by eukaryotic transcription factors
 Chapter 9 DNA recognition by eukaryotic transcription factors TRANSCRIPTION 101 Eukaryotic RNA polymerases RNA polymerase RNA polymerase I RNA polymerase II RNA polymerase III RNA polymerase IV Function
Chapter 9 DNA recognition by eukaryotic transcription factors TRANSCRIPTION 101 Eukaryotic RNA polymerases RNA polymerase RNA polymerase I RNA polymerase II RNA polymerase III RNA polymerase IV Function
Molecular Interactions F14NMI. Lecture 4: worked answers to practice questions
 Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
Molecular Interactions F14NMI Lecture 4: worked answers to practice questions http://comp.chem.nottingham.ac.uk/teaching/f14nmi jonathan.hirst@nottingham.ac.uk (1) (a) Describe the Monte Carlo algorithm
CONTRAfold: RNA Secondary Structure Prediction without Physics-Based Models
 Supplementary Material for CONTRAfold: RNA Secondary Structure Prediction without Physics-Based Models Chuong B Do, Daniel A Woods, and Serafim Batzoglou Stanford University, Stanford, CA 94305, USA, {chuongdo,danwoods,serafim}@csstanfordedu,
Supplementary Material for CONTRAfold: RNA Secondary Structure Prediction without Physics-Based Models Chuong B Do, Daniel A Woods, and Serafim Batzoglou Stanford University, Stanford, CA 94305, USA, {chuongdo,danwoods,serafim}@csstanfordedu,
The protein folding problem consists of two parts:
 Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Energetics and kinetics of protein folding The protein folding problem consists of two parts: 1)Creating a stable, well-defined structure that is significantly more stable than all other possible structures.
Searching for Noncoding RNA
 Searching for Noncoding RN Larry Ruzzo omputer Science & Engineering enome Sciences niversity of Washington http://www.cs.washington.edu/homes/ruzzo Bio 2006, Seattle, 8/4/2006 1 Outline Noncoding RN Why
Searching for Noncoding RN Larry Ruzzo omputer Science & Engineering enome Sciences niversity of Washington http://www.cs.washington.edu/homes/ruzzo Bio 2006, Seattle, 8/4/2006 1 Outline Noncoding RN Why
Copyright (c) 2007 IEEE. Personal use of this material is permitted. However, permission to use this material for any other purposes must be obtained
 Copyright (c) 2007 IEEE. Personal use of this material is permitted. However, permission to use this material for any other purposes must be obtained from the IEEE by sending an email to pubs-permissions@ieee.org.
Copyright (c) 2007 IEEE. Personal use of this material is permitted. However, permission to use this material for any other purposes must be obtained from the IEEE by sending an email to pubs-permissions@ieee.org.
Protein Folding & Stability. Lecture 11: Margaret A. Daugherty. Fall How do we go from an unfolded polypeptide chain to a
 Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Lecture 11: Protein Folding & Stability Margaret A. Daugherty Fall 2004 How do we go from an unfolded polypeptide chain to a compact folded protein? (Folding of thioredoxin, F. Richards) Structure - Function
Proteins polymer molecules, folded in complex structures. Konstantin Popov Department of Biochemistry and Biophysics
 Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Chapter 1. A Method to Predict the 3D Structure of an RNA Scaffold. Xiaojun Xu and Shi-Jie Chen. Abstract. 1 Introduction
 Chapter 1 Abstract The ever increasing discoveries of noncoding RNA functions draw a strong demand for RNA structure determination from the sequence. In recently years, computational studies for RNA structures,
Chapter 1 Abstract The ever increasing discoveries of noncoding RNA functions draw a strong demand for RNA structure determination from the sequence. In recently years, computational studies for RNA structures,
DNA/RNA structure and packing
 DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
RNA folding at elementary step resolution
 RNA (2000), 6:325 338+ Cambridge University Press+ Printed in the USA+ Copyright 2000 RNA Society+ RNA folding at elementary step resolution CHRISTOPH FLAMM, 1 WALTER FONTANA, 2,3 IVO L. HOFACKER, 1 and
RNA (2000), 6:325 338+ Cambridge University Press+ Printed in the USA+ Copyright 2000 RNA Society+ RNA folding at elementary step resolution CHRISTOPH FLAMM, 1 WALTER FONTANA, 2,3 IVO L. HOFACKER, 1 and
Biochemistry in Singulo:
 Biochemistry in Singulo: When Less Means More Carlos Bustamante University of California, Berkeley The Ensemble Approach The study of chemical transforma9ons has been dominated by the ensemble method:
Biochemistry in Singulo: When Less Means More Carlos Bustamante University of California, Berkeley The Ensemble Approach The study of chemical transforma9ons has been dominated by the ensemble method:
Biophysical Model Building
 Biophysical Model Building Step 1: Come up with a hypothesis about how a system works How many binding sites? Is there cooperativity? Step 2: Translate the qualitative hypotheses into an observable mathematical
Biophysical Model Building Step 1: Come up with a hypothesis about how a system works How many binding sites? Is there cooperativity? Step 2: Translate the qualitative hypotheses into an observable mathematical
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
