Small-Angle Scattering from Biomolecular Solutions
|
|
|
- Sydney Foster
- 5 years ago
- Views:
Transcription
1 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N Small-Angle Scattering from Biomolecular Solutions For students of HI Computational Structural Biology Dimitri I. Svergun, Ph.D. EMBL, Hamburg Outstation
2 Small-angle angle scattering experiment Detector Sample Log (Intensity) Monochromatic beam Wave vector k, k=2π/λ Radiation sources: X-ray generator (λ = nm) Synchrotron (λ = nm) Thermal neutrons (λ = nm) 2θ s=4π sinθ/λ, nm -1 Scattering vector s=k 1 -k, s=4π sinθ/λ k 1
3 Scattering from dilute macromolecular solutions (monodisperse( systems) I( s) = 4π D 0 sin sr p( r) dr sr The scattering is proportional to that of a single particle averaged over all orientations, which allows one to determine size, shape and internal structure of the particle at low (1-10 nm) resolution.
4 X-rays Neutrons X-rays: scattering factor increases with atomic number, no difference between H and D Neutrons: scattering factor is irregular, may be negative, huge difference between H and D Element H D C N O P S Au At. Weight N electrons bx,10-12 cm bn,10-12 cm
5 Solvent scattering and contrast I solution (s) I solvent (s) I particle (s) To obtain scattering from the particles, solvent scattering must be subtracted Contrast ρ = <ρ(r) - ρ s >, where ρ s is the scattering density of the solvent (is usually very small for biological samples)
6 Contrast variation Changing solvent density separates information about shape and internal structure Selective labelling permits to visualize specific structural fragments Stuhrmann, H.B. & Kirste, R.G. (1965) Z. Phys. Chem. 46, 247
7 X-rays Neutrons Addition of sucrose or salts RNA, 550 e/nm 3 Isotopic H/D substitution D-Protein, 130% D 2 O D-RNA, 120% D 2 O D 2 O, cm -2 60% sucrose, 430 e/nm 3 Protein, 410 e/nm 3 H-RNA, 70% D 2 O H-Protein, 40% D 2 O H 2 O, 344 e/nm 3 H 2 O, cm -2
8 Major applications of solution scattering Ab initio low resolution structure analysis Addition of missing fragments to high resolution models Rigid body refinement of complexes Quantitative characterization of mixtures Contrast variation on multicomponent particles
9 Major problem of scattering data analysis 3D search model M parameters Trial-and-error 1D scattering data Non-linear search Additional information is ALWAYS required to resolve or reduce ambiguity of interpretation
10 Ab initio initio methods Envelope function Stuhrmann, H. B. (1970) Z. Physik. Chem. N.F. 72, 177 Svergun, D.I. et al. (1996) Acta Crystallogr. A52, 419 Bead models Chacón, P. et al. (1998) Biophys. J. 74, 2760 Svergun, D.I. (1999) Biophys. J. 76, 2879 Dummy residues model Svergun, D.I., Petoukhov, M.V. & Koch, M.H.J. (2001) Biophys. J. 80, All the methods minimize Discrepancy[Data] + Penalty[Additional info]
11 Bead (dummy atoms) models Search volume The particle is represented as an ensemble of small densely packed volume elements (beads) in the search volume (e.g. a sphere with diameter D ) max The structure is described by phase assignments of each of these positions (e.g. for shape determination 1 = particle, 0 = solvent) A Monte-Carlo type search is employed to build a model fitting the scattering data Chacón,, P. et al. (1998) Biophys.. J. 74, Svergun, D.I. (1999) Biophys.. J. 76,
12 Ab initio program DAMMIN Using simulated annealing, finds a compact dummy atoms configuration X that fits the scattering data by minimizing f 2 ( X ) = χ [ Iexp( s), I( s, X )] + αp( X ) where χ is the discrepancy between the experimental and calculated curves, P(X) is the penalty to ensure compactness and connectivity, α>0 its weight. compact loose disconnected
13 Benchmarking ab initio methods Envelope Bead model Dummy residues log I, relative 2 Experimental data Envelope model Bead model Dummy residue model s, nm -1 Comparison with the crystal SASHA DAMMIN GASBOR structure of lysozyme
14 Study of nuclear exportins and importins Importin/exportin-mediated nuclear transport Examples of cargo recognition Targets for small-angle scattering study Two available crystal structures of bound importins built by stacks of HEAT repeats Snail-like importin β Z-like transportin
15 2 1 lg I, relative (1) (2) (3) (4) Importins ab initio lg I, relative 3 (1) (2) (3) (4) s, nm -1 Transportin- RanGTP: Z-like Free transportin: Z-like s, nm -1 Free importin β: Z-like Bound importin β: snail-like like lg I, relative lg I, relative 2 1 (1) (2) (3) (4) 3 (1) (2) (3) (4) s, nm s, nm -1 Fukuhara, N., Fernandez, E., Ebert, J., Conti, E. & Svergun, D. I. (2004) J. Biol. Chem. 279, 2176
16 lg I, relative 3 (1) (2) (3) (4) (5) 2 (6) (7) (8) (9) Exportins ab ab initio S.pombe Xpot Xpot+Ran+tRNA Yeast Los s, nm -1 lg I, relative 3 2 (1) (2) (3) (4) (5) (6) (7) s, nm -1 Yeast Cse1 Cse1+Ran+α S.pombe Crm1 Human Crm1 Fukuhara, N., Fernandez, E., Ebert, J., Conti, E. & Svergun, D. I. (2004) J. Biol. Chem. 279, 2176
17 Rigid body modelling Scattering amplitudes of the subunits are pre-computed and positional parameters are refined to fit the scattering from the complex Kozin & Svergun (2000). J. Appl. Cryst. 33, Konarev, Petoukhov & Svergun (2001). J. Appl. Cryst. 34, CRYSOL CRYSON MASSHA (Windows PC) ASSA (SUN/SGI/DEC)
18 Quaternary structure of glutamate synthase 3 2 lg I, relative β-subunit Catalytically active αβ holoenzyme of the iron-sulfur flavoprotein GltS, contains α subunit (162 kda) β subunit (52 kda) 1 GltS β-dimer 0-1 α subunit s, nm -1 α-tetramer αβ-tetramer M.V. Petoukhov, D.I. Svergun, P.V. Konarev, S. Ravasio, R.H.H. van den Heuvel, B. Curti & M.A. Vanoni (2003). J. Biol. Chem., 278, 29933
19 Scattering from a multiphase particle lg I, relative % D 2 O 40% D 2 O 55% D 2 O 75% D 2 O 100% D 2 O s, nm -1 I m ( s) = j ( ρ m j ) 2 I j ( s) + 2 j> k ρ m j ρ m i I jk ( s)
20 Ab initio initio multiphase modelling Start: random phase assignments within the search volume, no fit to the experimental data Finish: condensed multiphase model with minimum interfacial area fitting multiple data sets Program MONSA, Svergun, D.I. (1999) Biophys. J. 76, 2879
21 Contrast variation on hybrid ribosomes 0% D 2 O 40% D 2 O 70% D 2 O Protonated 70S ribosome, HH30+HH50 Hybrid 70S with 23S RNA deuterated, HH30+HD50
22 Scattering data from hybrid ribosomes Contrast variation by solvent exchange HH30+HH50 DD30+HH50 DH30+HH50 in 0, 35, 50, 75, 100% D 2 O HH30+DD50 in 0, 35, 50, 75% D 2 O DH30+DD50 and HH30+DH50 in 0, 40, 60, 100% D 2 O HH30 and HH50 in 0, 100% D 15 curves 4 curves 8 curves 0, 100% D 2 O 4 curves DD30 and DD50 in 0% D 2 O 2 curves Spin-dependent contrast variation data HH30+DD50, DD30+HH50, DH30+DH50 Polarization = 0 and 1 X-ray scattering curves from 70S, 30S and 50S 6 curves 3 curves TOTAL 42 curves
23 Search volume for the 70S ribosome Yellow pixels: cryo- EM model of Frank et al.. (1995) Red and blue circles: dummy atoms belonging to the 30S and 50S subunits, respectively Number of atoms M=7860 Packing radius r 0 =0.5 nm
24 Ribosomal data fitted Neutrons X-rays Svergun, D.I. & Nierhaus, K.H. (2000) J.Biol. Chem. 275,
25 A protein-rna map in the 70S ribosome E.coli
26 Solution crystal X-ray and neutron scattering map of protein-rna distribution in the 70S ribosome E. coli (left, resolution 3 nm) compared with later crystallographic models (right). 5 nm Top, 30S subunit from Th. thermophilus,, resolution 0.33 nm (Schluenzen,, F, et al, & Yonath,, A. (2000) Cell, 10, 615). Bottom, 50S subunit from H. marismortui,, resolution 0.24 nm (Ban, N., Nissen,, P., Hansen, J., Moore, P.B. & Steitz,, T.A. (2000) Science, 289,, 905).
27 Acknowledgments M.H.J. Koch, M.Malfois (EMBL, Hamburg Outstation), V.V. Volkov, M.B. Kozin, M.V.Petoukhov, P.V.Konarev, A.V.Sokolova (Institute of Crystallography, Moscow) hamburg.de/externalinfo/research/sax GltS: M.A.Vanoni (Milan University) Ribosome: K.H.Nierhaus (MPIMG, Berlin), H.B.Stuhrmann (GKSS, Geesthacht), J.Frank (Wadsworth Center, Albany), J. Skov Pedersen (Aarhus University) Importins/Exportins: N.Fukuhara (IBS, Grenoble), E.Conti (EMBL, Heidelberg), P.Timmins (ILL, Grenoble) Animation:
ID14-EH3. Adam Round
 Bio-SAXS @ ID14-EH3 Adam Round Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and
Bio-SAXS @ ID14-EH3 Adam Round Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and
SAXS/SANS data processing and overall parameters
 EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
Small-Angle Scattering Atomic Structure Based Modeling
 Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Modelling against small angle scattering data. Al Kikhney EMBL Hamburg, Germany
 Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Nuclear targeting by Nuclear Localization Signals (NLS) Richardson and Laskey (1988)
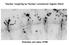 Nuclear targeting by Nuclear Localization Signals (NLS) Richardson and Laskey (1988) The nuclear import pathway of proteins containing a classical Nuclear Localization Signal (NLS) Uptake of NLS-containing
Nuclear targeting by Nuclear Localization Signals (NLS) Richardson and Laskey (1988) The nuclear import pathway of proteins containing a classical Nuclear Localization Signal (NLS) Uptake of NLS-containing
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands
 Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Introduction to Biological Small Angle Scattering
 Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
Data reduction and processing tutorial
 Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
SI Text S1 Solution Scattering Data Collection and Analysis. SI references
 SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013
 Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Small Angle X-Ray Solution Scattering of Biological Macromolecules
 Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
Joint use of small-angle X-ray and neutron scattering to study biological. European Molecular Biology Laboratory, Hamburg Outstation, Notkestraße 85,
 Joint use of small-angle X-ray and neutron scattering to study biological macromolecules in solution Maxim V. Petoukhov 1,2 and Dmitri I. Svergun 1,2 1 European Molecular Biology Laboratory, Hamburg Outstation,
Joint use of small-angle X-ray and neutron scattering to study biological macromolecules in solution Maxim V. Petoukhov 1,2 and Dmitri I. Svergun 1,2 1 European Molecular Biology Laboratory, Hamburg Outstation,
Introduction to biological small angle scattering
 Introduction to biological small angle scattering Frank Gabel (IBS/ILL) EMBO Practical Course (May 6th 013) F. Gabel (May 6th 013) EMBO Practical Course Length-scales and tools in structural biology small
Introduction to biological small angle scattering Frank Gabel (IBS/ILL) EMBO Practical Course (May 6th 013) F. Gabel (May 6th 013) EMBO Practical Course Length-scales and tools in structural biology small
Neutron Scattering (Basics).
 Neutron Scattering (Basics). Cy M. Jeffries Introduction One of the new frontiers of structural biology is the study of the interactome: There are approximately 30 000 structural genes in the human genome.
Neutron Scattering (Basics). Cy M. Jeffries Introduction One of the new frontiers of structural biology is the study of the interactome: There are approximately 30 000 structural genes in the human genome.
Studying conformational dynamics and molecular recognition using integrated structural biology in solution Michael Sattler
 Studying conformational dynamics and molecular recognition using integrated structural biology in solution Michael Sattler http://www.nmr.ch.tum.de http://www.helmholtz-muenchen.de/stb/ Outline Dynamics
Studying conformational dynamics and molecular recognition using integrated structural biology in solution Michael Sattler http://www.nmr.ch.tum.de http://www.helmholtz-muenchen.de/stb/ Outline Dynamics
Crystals, X-rays and Proteins
 Crystals, X-rays and Proteins Comprehensive Protein Crystallography Dennis Sherwood MA (Hons), MPhil, PhD Jon Cooper BA (Hons), PhD OXFORD UNIVERSITY PRESS Contents List of symbols xiv PART I FUNDAMENTALS
Crystals, X-rays and Proteins Comprehensive Protein Crystallography Dennis Sherwood MA (Hons), MPhil, PhD Jon Cooper BA (Hons), PhD OXFORD UNIVERSITY PRESS Contents List of symbols xiv PART I FUNDAMENTALS
Use of neutrons in biology and medicine. Jayne Lawrence Pharmaceutical Science Division King s College London London
 Use of neutrons in biology and medicine Jayne Lawrence Pharmaceutical Science Division King s College London London Useful reading Chapter 23 Neutron crystallography of proteins and Chapter 24 Molecular
Use of neutrons in biology and medicine Jayne Lawrence Pharmaceutical Science Division King s College London London Useful reading Chapter 23 Neutron crystallography of proteins and Chapter 24 Molecular
Overview - Macromolecular Crystallography
 Overview - Macromolecular Crystallography 1. Overexpression and crystallization 2. Crystal characterization and data collection 3. The diffraction experiment 4. Phase problem 1. MIR (Multiple Isomorphous
Overview - Macromolecular Crystallography 1. Overexpression and crystallization 2. Crystal characterization and data collection 3. The diffraction experiment 4. Phase problem 1. MIR (Multiple Isomorphous
Structure Analysis by Small-Angle X-Ray and Neutron Scattering
 Structure Analysis by Small-Angle X-Ray and Neutron Scattering L. A. Feigin and D. I. Svergun Institute of Crystallography Academy of Sciences of the USSR Moscow, USSR Edited by George W. Taylor Princeton
Structure Analysis by Small-Angle X-Ray and Neutron Scattering L. A. Feigin and D. I. Svergun Institute of Crystallography Academy of Sciences of the USSR Moscow, USSR Edited by George W. Taylor Princeton
Complementary use of SAXS and SANS. Jill Trewhella University of Sydney
 Complementary use of SAXS and SANS Jill Trewhella University of Sydney Conceptual diagram of the small-angle scattering experiment The conceptual experiment and theory is the same for X-rays and neutrons,
Complementary use of SAXS and SANS Jill Trewhella University of Sydney Conceptual diagram of the small-angle scattering experiment The conceptual experiment and theory is the same for X-rays and neutrons,
How to judge data quality
 SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
Small-angle scattering studies of biological macromolecules in solution
 INSTITUTE OF PHYSICS PUBLISHING Rep. Prog. Phys. 66 (2003) 1735 1782 REPORTS ON PROGRESS IN PHYSICS PII: S0034-4885(03)12688-7 Small-angle scattering studies of biological macromolecules in solution Dmitri
INSTITUTE OF PHYSICS PUBLISHING Rep. Prog. Phys. 66 (2003) 1735 1782 REPORTS ON PROGRESS IN PHYSICS PII: S0034-4885(03)12688-7 Small-angle scattering studies of biological macromolecules in solution Dmitri
Scattering Lecture. February 24, 2014
 Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
Scattering Lecture February 24, 2014 Structure Determination by Scattering Waves of radiation scattered by different objects interfere to give rise to an observable pattern! The wavelength needs to close
P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France
 Scattering of X-rays P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France SAXS measurement Sample SAXS measuring cell SAXS measurement Scattering experiment X-ray beam?? Detector SAXS measurement
Scattering of X-rays P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France SAXS measurement Sample SAXS measuring cell SAXS measurement Scattering experiment X-ray beam?? Detector SAXS measurement
Molecular shapes from small-angle X-ray scattering: extension of the theory to higher scattering angles
 Acta Crystallographica Section A Foundations of Crystallography ISSN 0108-7673 Editor: D. Schwarzenbach Molecular shapes from small-angle X-ray scattering: extension of the theory to higher scattering
Acta Crystallographica Section A Foundations of Crystallography ISSN 0108-7673 Editor: D. Schwarzenbach Molecular shapes from small-angle X-ray scattering: extension of the theory to higher scattering
SAXS Basics for BioSAXS. Robert P. Rambo Diamond Light Source B21
 SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
Advances in structure analysis using small-angle scattering in solution
 Advances in structure analysis using small-angle scattering in solution Dmitri I. Svergun 1,2) and Michel H. J. Koch 1) 1) EMBL, Hamburg Outstation, Notkestraße 85, D-22603 Hamburg, Germany 2) Institute
Advances in structure analysis using small-angle scattering in solution Dmitri I. Svergun 1,2) and Michel H. J. Koch 1) 1) EMBL, Hamburg Outstation, Notkestraße 85, D-22603 Hamburg, Germany 2) Institute
History of 3D Electron Microscopy and Helical Reconstruction
 T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
T H E U N I V E R S I T Y of T E X A S S C H O O L O F H E A L T H I N F O R M A T I O N S C I E N C E S A T H O U S T O N History of 3D Electron Microscopy and Helical Reconstruction For students of HI
Scattering of X-rays. EMBO Practical Course on Solution Scattering from Biological Macromolecules Hamburg 4 11 September 2001
 Scattering of X-rays Books on SAS - " The origins" (no recent edition) : Small Angle Scattering of X-rays A. Guinier and A. Fournet, (1955), in English, ed. Wiley, NY - Small-Angle X-ray Scattering O.
Scattering of X-rays Books on SAS - " The origins" (no recent edition) : Small Angle Scattering of X-rays A. Guinier and A. Fournet, (1955), in English, ed. Wiley, NY - Small-Angle X-ray Scattering O.
BM29 biosaxs data processing tutorial
 HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi
 Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Tools for Cryo-EM Map Fitting. Paul Emsley MRC Laboratory of Molecular Biology
 Tools for Cryo-EM Map Fitting Paul Emsley MRC Laboratory of Molecular Biology April 2017 Cryo-EM model-building typically need to move more atoms that one does for crystallography the maps are lower resolution
Tools for Cryo-EM Map Fitting Paul Emsley MRC Laboratory of Molecular Biology April 2017 Cryo-EM model-building typically need to move more atoms that one does for crystallography the maps are lower resolution
Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation, and crystallization
 HERCULES Specialized Course: Non-atomic resolution scattering in biology and soft matter Grenoble, September 14-19, 2014 Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation,
HERCULES Specialized Course: Non-atomic resolution scattering in biology and soft matter Grenoble, September 14-19, 2014 Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation,
Chapter 11 Protein shape and assembly studied with X ray solution scattering: Fundaments and practice
 Chapter 11 Protein shape and assembly studied with X ray solution scattering: Fundaments and practice Rubén M Buey, Pablo Chacón, José Manuel Andreu and José Fernando Díaz Abstract Small Angle X ray Scattering
Chapter 11 Protein shape and assembly studied with X ray solution scattering: Fundaments and practice Rubén M Buey, Pablo Chacón, José Manuel Andreu and José Fernando Díaz Abstract Small Angle X ray Scattering
Macromolecular X-ray Crystallography
 Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
Protein Structural Models for CHEM 641 Fall 07 Brian Bahnson Department of Chemistry & Biochemistry University of Delaware Macromolecular X-ray Crystallography Purified Protein X-ray Diffraction Data collection
BCM Protein crystallography - I. Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS
 BCM 6200 - Protein crystallography - I Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS What SAXS can do Small-angle X-ray scattering (SAXS) yields information on biological
BCM 6200 - Protein crystallography - I Crystal symmetry X-ray diffraction Protein crystallization X-ray sources SAXS What SAXS can do Small-angle X-ray scattering (SAXS) yields information on biological
Growing Biological crystals for Neutrons
 European Molecular Biology Laboratory Grenoble Outstation Growing Biological crystals for Neutrons Monika Budayova-Spano UJF EMBL CNRS / ILL Why neutron protein crystallography? Providing evidence on the
European Molecular Biology Laboratory Grenoble Outstation Growing Biological crystals for Neutrons Monika Budayova-Spano UJF EMBL CNRS / ILL Why neutron protein crystallography? Providing evidence on the
SAXS and SANS facilities and experimental practice. Clement Blanchet
 SAXS and SANS facilities and experimental practice Clement Blanchet SAS experiment Detector X-ray or neutron Beam Sample 2 s Buffer X-rays Roengten, 1895 Electromagnetic wave The electromagnetic spectrum
SAXS and SANS facilities and experimental practice Clement Blanchet SAS experiment Detector X-ray or neutron Beam Sample 2 s Buffer X-rays Roengten, 1895 Electromagnetic wave The electromagnetic spectrum
Small Angle Neutron Scattering in Different Fields of Research. Henrich Frielinghaus
 Small Angle Neutron Scattering in Different Fields of Research Henrich Frielinghaus Jülich Centre for Neutron Science Forschungszentrum Jülich GmbH Lichtenbergstrasse 1 85747 Garching (München) h.frielinghaus@fz-juelich.de
Small Angle Neutron Scattering in Different Fields of Research Henrich Frielinghaus Jülich Centre for Neutron Science Forschungszentrum Jülich GmbH Lichtenbergstrasse 1 85747 Garching (München) h.frielinghaus@fz-juelich.de
Handout 7 Reciprocal Space
 Handout 7 Reciprocal Space Useful concepts for the analysis of diffraction data http://homepages.utoledo.edu/clind/ Concepts versus reality Reflection from lattice planes is just a concept that helps us
Handout 7 Reciprocal Space Useful concepts for the analysis of diffraction data http://homepages.utoledo.edu/clind/ Concepts versus reality Reflection from lattice planes is just a concept that helps us
Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins. Michael Kachala EMBL-Hamburg, Germany
 Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins Michael Kachala EMBL-Hamburg, Germany 60 mkl >1 mg/ml Monocromatic X-ray beam Sample Mono- or polydisperse
Development of Novel Small- Angle X-ray Scattering Data Analysis Methods for Study of Flexible Proteins Michael Kachala EMBL-Hamburg, Germany 60 mkl >1 mg/ml Monocromatic X-ray beam Sample Mono- or polydisperse
Introduction to X-ray and neutron scattering
 UNESCO/IUPAC Postgraduate Course in Polymer Science Lecture: Introduction to X-ray and neutron scattering Zhigunov Alexander Institute of Macromolecular Chemistry ASCR, Heyrovsky sq., Prague -16 06 http://www.imc.cas.cz/unesco/index.html
UNESCO/IUPAC Postgraduate Course in Polymer Science Lecture: Introduction to X-ray and neutron scattering Zhigunov Alexander Institute of Macromolecular Chemistry ASCR, Heyrovsky sq., Prague -16 06 http://www.imc.cas.cz/unesco/index.html
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Crystal lattice Real Space. Reflections Reciprocal Space. I. Solving Phases II. Model Building for CHEM 645. Purified Protein. Build model.
 I. Solving Phases II. Model Building for CHEM 645 Purified Protein Solve Phase Build model and refine Crystal lattice Real Space Reflections Reciprocal Space ρ (x, y, z) pronounced rho F hkl 2 I F (h,
I. Solving Phases II. Model Building for CHEM 645 Purified Protein Solve Phase Build model and refine Crystal lattice Real Space Reflections Reciprocal Space ρ (x, y, z) pronounced rho F hkl 2 I F (h,
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA. President, Association of Women in Science, Bethesda Chapter STEM Consultant
 CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
CRYSTALLOGRAPHY AND STORYTELLING WITH DATA President, Association of Women in Science, Bethesda Chapter STEM Consultant MY STORY Passion for Science BS Biology Major MS Biotechnology & Project in Bioinformatics
Application examples of single particle 3D reconstruction. Ning Gao Tsinghua University
 Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Application examples of single particle 3D reconstruction Ning Gao Tsinghua University ninggao@tsinghua.edu.cn Electron Microscopes First electron microscope constructed by Ernst Ruska in 1930 s (1986
Determining Protein Structure BIBC 100
 Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Determining Protein Structure BIBC 100 Determining Protein Structure X-Ray Diffraction Interactions of x-rays with electrons in molecules in a crystal NMR- Nuclear Magnetic Resonance Interactions of magnetic
Protein Crystallography
 Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
Protein Crystallography Part II Tim Grüne Dept. of Structural Chemistry Prof. G. Sheldrick University of Göttingen http://shelx.uni-ac.gwdg.de tg@shelx.uni-ac.gwdg.de Overview The Reciprocal Lattice The
π p-elastic Scattering in the Resonance Region
 New Results on Spin Rotation Parameter A in the π p-elastic Scattering in the Resonance Region I.G. Alekseev Λ, P.E. Budkovsky Λ, V.P. Kanavets Λ, L.I. Koroleva Λ, B.V. Morozov Λ, V.M. Nesterov Λ, V.V.
New Results on Spin Rotation Parameter A in the π p-elastic Scattering in the Resonance Region I.G. Alekseev Λ, P.E. Budkovsky Λ, V.P. Kanavets Λ, L.I. Koroleva Λ, B.V. Morozov Λ, V.M. Nesterov Λ, V.V.
Supplementary materials. Crystal structure of the carboxyltransferase domain. of acetyl coenzyme A carboxylase. Department of Biological Sciences
 Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Three-Dimensional Electron Microscopy of Macromolecular Assemblies
 Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
Three-Dimensional Electron Microscopy of Macromolecular Assemblies Joachim Frank Wadsworth Center for Laboratories and Research State of New York Department of Health The Governor Nelson A. Rockefeller
BCMB / CHEM 8190 Biomolecular NMR GRADUATE COURSE OFFERING IN NUCLEAR MAGNETIC RESONANCE
 BCMB / CHEM 8190 Biomolecular NMR GRADUATE COURSE OFFERING IN NUCLEAR MAGNETIC RESONANCE "Biomolecular Nuclear Magnetic Resonance" is a course intended for all graduate students with an interest in applications
BCMB / CHEM 8190 Biomolecular NMR GRADUATE COURSE OFFERING IN NUCLEAR MAGNETIC RESONANCE "Biomolecular Nuclear Magnetic Resonance" is a course intended for all graduate students with an interest in applications
SHELXC/D/E. Andrea Thorn
 SHELXC/D/E Andrea Thorn What is experimental phasing? Experimental phasing is what you do if MR doesn t work. What is experimental phasing? Experimental phasing methods depend on intensity differences.
SHELXC/D/E Andrea Thorn What is experimental phasing? Experimental phasing is what you do if MR doesn t work. What is experimental phasing? Experimental phasing methods depend on intensity differences.
Approximation of the structure factor for nonspherical hard bodies using polydisperse spheres
 Journal of Applied Crystallography ISSN 21-8898 Approximation of the structure factor for nonspherical hard bodies using polydisperse spheres Steen Hansen J. Appl. Cryst. (213). 46, 18 116 Copyright c
Journal of Applied Crystallography ISSN 21-8898 Approximation of the structure factor for nonspherical hard bodies using polydisperse spheres Steen Hansen J. Appl. Cryst. (213). 46, 18 116 Copyright c
X-ray Crystallography
 2009/11/25 [ 1 ] X-ray Crystallography Andrew Torda, wintersemester 2009 / 2010 X-ray numerically most important more than 4/5 structures Goal a set of x, y, z coordinates different properties to NMR History
2009/11/25 [ 1 ] X-ray Crystallography Andrew Torda, wintersemester 2009 / 2010 X-ray numerically most important more than 4/5 structures Goal a set of x, y, z coordinates different properties to NMR History
Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes
 9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
9 The Nucleus Student Learning Outcomes: Nucleus distinguishes Eukaryotes from Prokaryotes Explain general structures of Nuclear Envelope, Nuclear Lamina, Nuclear Pore Complex Explain movement of proteins
The Small Angle X-ray Scattering Technique: An Overview
 The Small Angle X-ray Scattering Technique: An Overview Dr. Gianluca Croce, Ph.D DISTA - Univ. Piemonte Orientale Via T. Michel 11,15121 Alessandria (Italy) gianluca.croce@mfn.unipmn.it Dr. Gianluca Croce
The Small Angle X-ray Scattering Technique: An Overview Dr. Gianluca Croce, Ph.D DISTA - Univ. Piemonte Orientale Via T. Michel 11,15121 Alessandria (Italy) gianluca.croce@mfn.unipmn.it Dr. Gianluca Croce
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Schematic representation of relation between disorder and scattering
 Crystal lattice Reciprocal lattice FT Schematic representation of relation between disorder and scattering ρ = Δρ + Occupational disorder Diffuse scattering Bragg scattering ρ = Δρ + Positional
Crystal lattice Reciprocal lattice FT Schematic representation of relation between disorder and scattering ρ = Δρ + Occupational disorder Diffuse scattering Bragg scattering ρ = Δρ + Positional
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
Fundamentals of X-ray diffraction
 Fundamentals of X-ray diffraction Elena Willinger Lecture series: Modern Methods in Heterogeneous Catalysis Research Outline History of X-ray Sources of X-ray radiation Physics of X-ray scattering Fundamentals
Fundamentals of X-ray diffraction Elena Willinger Lecture series: Modern Methods in Heterogeneous Catalysis Research Outline History of X-ray Sources of X-ray radiation Physics of X-ray scattering Fundamentals
X-ray Crystallography. Kalyan Das
 X-ray Crystallography Kalyan Das Electromagnetic Spectrum NMR 10 um - 10 mm 700 to 10 4 nm 400 to 700 nm 10 to 400 nm 10-1 to 10 nm 10-4 to 10-1 nm X-ray radiation was discovered by Roentgen in 1895. X-rays
X-ray Crystallography Kalyan Das Electromagnetic Spectrum NMR 10 um - 10 mm 700 to 10 4 nm 400 to 700 nm 10 to 400 nm 10-1 to 10 nm 10-4 to 10-1 nm X-ray radiation was discovered by Roentgen in 1895. X-rays
Thesis and PhD themes
 Thesis and PhD themes DETERMINATION OF MASSES OF THE SUPER HEAVY ELEMENTS IN THE EXPERIMENTS ON SYNTHESIS OF 112 AND 114 ELEMENTS USING THE REACTIONS 48 CA+ 238 U AND 48 CA+ 242 PU Traditionally, in experiments
Thesis and PhD themes DETERMINATION OF MASSES OF THE SUPER HEAVY ELEMENTS IN THE EXPERIMENTS ON SYNTHESIS OF 112 AND 114 ELEMENTS USING THE REACTIONS 48 CA+ 238 U AND 48 CA+ 242 PU Traditionally, in experiments
CHARACTERIZATION OF WEAK INTRA- AND INTERMOLECULAR INTERACTIONS USING EXPERIMENTAL CHARGE DENSITY DISTRIBUTIONS
 CHARACTERIZATION OF WEAK INTRA- AND INTERMOLECULAR INTERACTIONS USING EXPERIMENTAL CHARGE DENSITY DISTRIBUTIONS Edwin D. Stevens and MaRyca Watts Department of Chemistry, University of New Orleans, New
CHARACTERIZATION OF WEAK INTRA- AND INTERMOLECULAR INTERACTIONS USING EXPERIMENTAL CHARGE DENSITY DISTRIBUTIONS Edwin D. Stevens and MaRyca Watts Department of Chemistry, University of New Orleans, New
Supporting Information for: Complexation of β-lactoglobulin Fibrils and Sulfated Polysaccharides
 Supporting Information for: Complexation of β-lactoglobulin Fibrils and Sulfated Polysaccharides Owen G Jones 1, Stephaandschin 1, Jozef Adamcik 1, Ludger Harnau 2, Sreenath Bolisetty 1, and Raffaele Mezzenga
Supporting Information for: Complexation of β-lactoglobulin Fibrils and Sulfated Polysaccharides Owen G Jones 1, Stephaandschin 1, Jozef Adamcik 1, Ludger Harnau 2, Sreenath Bolisetty 1, and Raffaele Mezzenga
V19 Topology of Protein Complexes on Graphs
 V19 Topology of Protein Complexes on Graphs (1) CombDock (2) Nuclear Pore Complex 19. Lecture WS 2013/14 Bioinformatics III 1 Prediction Structures of Protein Complexes from Connectivities CombDock: automated
V19 Topology of Protein Complexes on Graphs (1) CombDock (2) Nuclear Pore Complex 19. Lecture WS 2013/14 Bioinformatics III 1 Prediction Structures of Protein Complexes from Connectivities CombDock: automated
Supporting Information
 Submitted to Cryst. Growth Des. Version 1 of August 22, 2007 Supporting Information Engineering Hydrogen-Bonded Molecular Crystals Built from 1,3,5-Substituted Derivatives of Benzene: 6,6',6''-(1,3,5-Phenylene)tris-1,3,5-triazine-2,4-diamines
Submitted to Cryst. Growth Des. Version 1 of August 22, 2007 Supporting Information Engineering Hydrogen-Bonded Molecular Crystals Built from 1,3,5-Substituted Derivatives of Benzene: 6,6',6''-(1,3,5-Phenylene)tris-1,3,5-triazine-2,4-diamines
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
SUPPLEMENTARY INFORMATION doi:10.1038/nature11539 Supplementary Figure 1 Schematic representation of plant (A) and mammalian (B) P 2B -ATPase domain organization. Actuator (A-), nucleotide binding (N-),
Supporting Information
 Supporting Information Wilson et al. 10.1073/pnas.0804276105 Fig. S1. Sites of oxazolidinone resistance mutations in bacteria and archaea. (a) Secondary structure of the peptidyltransferase ring of the
Supporting Information Wilson et al. 10.1073/pnas.0804276105 Fig. S1. Sites of oxazolidinone resistance mutations in bacteria and archaea. (a) Secondary structure of the peptidyltransferase ring of the
Study of the Phase Composition of Fe 2 O 3 Nanoparticles
 WDS'9 Proceedings of Contributed Papers, Part III, 28 212, 29. ISBN 978-8-7378-13-3 MATFYZPRESS Study of the Phase Composition of Fe 2 O 3 Nanoparticles V. Valeš, J. Poltierová-Vejpravová, A. Mantlíková,
WDS'9 Proceedings of Contributed Papers, Part III, 28 212, 29. ISBN 978-8-7378-13-3 MATFYZPRESS Study of the Phase Composition of Fe 2 O 3 Nanoparticles V. Valeš, J. Poltierová-Vejpravová, A. Mantlíková,
Anomalous Small-Angle X-Ray Scattering of Core-shell Nanoparticles
 Anomalous Small-Angle X-Ray Scattering of Core-shell Nanoparticles Larissa Veiga University of Campinas Supervisor: Ulla Vainio DESY Summer Students Program 2008 September, 12 th 2008 1 1 Introduction
Anomalous Small-Angle X-Ray Scattering of Core-shell Nanoparticles Larissa Veiga University of Campinas Supervisor: Ulla Vainio DESY Summer Students Program 2008 September, 12 th 2008 1 1 Introduction
CS273: Algorithms for Structure Handout # 13 and Motion in Biology Stanford University Tuesday, 11 May 2003
 CS273: Algorithms for Structure Handout # 13 and Motion in Biology Stanford University Tuesday, 11 May 2003 Lecture #13: 11 May 2004 Topics: Protein Structure Determination Scribe: Minli Zhu We acknowledge
CS273: Algorithms for Structure Handout # 13 and Motion in Biology Stanford University Tuesday, 11 May 2003 Lecture #13: 11 May 2004 Topics: Protein Structure Determination Scribe: Minli Zhu We acknowledge
Christopher Pavlik Bioanalytical Chemistry March 2, 2011
 Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Web-based Auto-Rickshaw for validation of the X-ray experiment at the synchrotron beamline
 Web-based Auto-Rickshaw for validation of the X-ray experiment at the synchrotron beamline Auto-Rickshaw http://www.embl-hamburg.de/auto-rickshaw A platform for automated crystal structure determination
Web-based Auto-Rickshaw for validation of the X-ray experiment at the synchrotron beamline Auto-Rickshaw http://www.embl-hamburg.de/auto-rickshaw A platform for automated crystal structure determination
CCP4 Diamond 2014 SHELXC/D/E. Andrea Thorn
 CCP4 Diamond 2014 SHELXC/D/E Andrea Thorn SHELXC/D/E workflow SHELXC: α calculation, file preparation SHELXD: Marker atom search = substructure search SHELXE: density modification Maps and coordinate files
CCP4 Diamond 2014 SHELXC/D/E Andrea Thorn SHELXC/D/E workflow SHELXC: α calculation, file preparation SHELXD: Marker atom search = substructure search SHELXE: density modification Maps and coordinate files
arxiv: v3 [nucl-ex] 18 May 2018
![arxiv: v3 [nucl-ex] 18 May 2018 arxiv: v3 [nucl-ex] 18 May 2018](/thumbs/90/102861135.jpg) Observation of Pendellösung Fringes by Using Pulsed Neutrons Shigeyasu Itoh, Masaya Nakaji, Yuya Uchida, Masaaki Kitaguchi, and Hirohiko M. Shimizu Department of Physics, Nagoya University Furo-cho, Chikusa-ku,
Observation of Pendellösung Fringes by Using Pulsed Neutrons Shigeyasu Itoh, Masaya Nakaji, Yuya Uchida, Masaaki Kitaguchi, and Hirohiko M. Shimizu Department of Physics, Nagoya University Furo-cho, Chikusa-ku,
Structure Refinements of II-VI Semiconductor Nanoparticles based on PDF Measurements
 Structure Refinements of II-VI Semiconductor Nanoparticles based on PDF Measurements Reinhard B. Neder Institut für Physik der kondensierten Materie Lehrstuhl für Kristallographie und Strukturphysik Universität
Structure Refinements of II-VI Semiconductor Nanoparticles based on PDF Measurements Reinhard B. Neder Institut für Physik der kondensierten Materie Lehrstuhl für Kristallographie und Strukturphysik Universität
Structural characterization. Part 2
 Structural characterization Part Determining partial pair distribution functions X-ray absorption spectroscopy (XAS). Atoms of different elements have absorption edges at different energies. Structure
Structural characterization Part Determining partial pair distribution functions X-ray absorption spectroscopy (XAS). Atoms of different elements have absorption edges at different energies. Structure
Neutron Diffraction: a general overview
 RUG1 Neutron Diffraction: a general overview Graeme Blake Zernike Institute for Advanced Materials University of Groningen Outline Elastic scattering of neutrons from matter Comparison of neutron and X-ray
RUG1 Neutron Diffraction: a general overview Graeme Blake Zernike Institute for Advanced Materials University of Groningen Outline Elastic scattering of neutrons from matter Comparison of neutron and X-ray
Simulation for Proton Charge Radius (PRad) Experiment at Jefferson Lab1 Li Ye Mississippi State University For the PRad Collaboration The Proton Charg
 Simulation for Proton Charge Radius (PRad) Experiment at Jefferson Lab1 Li Ye Mississippi State University For the PRad Collaboration The Proton Charge Radius Puzzle refers to 7 σ discrepancy between the
Simulation for Proton Charge Radius (PRad) Experiment at Jefferson Lab1 Li Ye Mississippi State University For the PRad Collaboration The Proton Charge Radius Puzzle refers to 7 σ discrepancy between the
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
Nature Structural & Molecular Biology: doi: /nsmb Supplementary Figure 1
 Supplementary Figure 1 Cryo-EM structure and model of the C. thermophilum 90S preribosome. a, Gold standard FSC curve showing the average resolution of the 90S preribosome masked and unmasked (left). FSC
Supplementary Figure 1 Cryo-EM structure and model of the C. thermophilum 90S preribosome. a, Gold standard FSC curve showing the average resolution of the 90S preribosome masked and unmasked (left). FSC
Welcome to DESY. What is DESY and what kind of research is done here?
 Welcome to DESY. What is DESY and what kind of research is done here? Michael Grefe DESY Press and Public Relations (PR) What is DESY? > Deutsches Elektronen-Synchrotron (German electron synchrotron) DESY
Welcome to DESY. What is DESY and what kind of research is done here? Michael Grefe DESY Press and Public Relations (PR) What is DESY? > Deutsches Elektronen-Synchrotron (German electron synchrotron) DESY
Protein Crystallography. Mitchell Guss University of Sydney Australia
 Protein Crystallography Mitchell Guss University of Sydney Australia Outline of the talk Recap some basic crystallography and history Highlight the special requirements for protein (macromolecular) structure
Protein Crystallography Mitchell Guss University of Sydney Australia Outline of the talk Recap some basic crystallography and history Highlight the special requirements for protein (macromolecular) structure
Localization of Proteins and trna Molecules in the 70S Ribosome of the Escherichia coli Bacteria with Polarized Neutron Scatteringt
 1125 J. Appl. Cryst. (1997). 30, 1125-1131 Localization of Proteins and trna Molecules in the 70S Ribosome of the Escherichia coli Bacteria with Polarized Neutron Scatteringt R. WILLUMEIT, a* N. BURKHARDT,
1125 J. Appl. Cryst. (1997). 30, 1125-1131 Localization of Proteins and trna Molecules in the 70S Ribosome of the Escherichia coli Bacteria with Polarized Neutron Scatteringt R. WILLUMEIT, a* N. BURKHARDT,
Protein NMR. Bin Huang
 Protein NMR Bin Huang Introduction NMR and X-ray crystallography are the only two techniques for obtain three-dimentional structure information of protein in atomic level. NMR is the only technique for
Protein NMR Bin Huang Introduction NMR and X-ray crystallography are the only two techniques for obtain three-dimentional structure information of protein in atomic level. NMR is the only technique for
Fitting Function for Experimental Energy Ordered Spectra in Nuclear Continuum Studies
 Fitting Function for Experimental Energy Ordered Spectra in Nuclear Continuum Studies J.R. Pinzón, F. Cristancho January 17, 2012 Abstract We review the main features of the Hk-EOS method for the experimental
Fitting Function for Experimental Energy Ordered Spectra in Nuclear Continuum Studies J.R. Pinzón, F. Cristancho January 17, 2012 Abstract We review the main features of the Hk-EOS method for the experimental
Application of Bayesian analysis to indirect Fourier transformation in small-angle scattering
 Journal of Applied Crystallography ISSN 0021-8898 Editor: Gernot Kostorz Application of Bayesian analysis to indirect Fourier transformation in small-angle scattering Bente Vestergaard and Steen Hansen
Journal of Applied Crystallography ISSN 0021-8898 Editor: Gernot Kostorz Application of Bayesian analysis to indirect Fourier transformation in small-angle scattering Bente Vestergaard and Steen Hansen
Supplemental Information for:
 Supplemental Information for: New Insight into the Structure of RNA in Red clover necrotic mosaic virus and the Role of Divalent Cations Revealed by Small-Angle Neutron Scattering Stanton L. Martin a,
Supplemental Information for: New Insight into the Structure of RNA in Red clover necrotic mosaic virus and the Role of Divalent Cations Revealed by Small-Angle Neutron Scattering Stanton L. Martin a,
Fan, Hai-fu Institute of Physics, Chinese Academy of Sciences, Beijing , China
 Direct Methods in Crystallography Fan, Hai-fu Institute of Physics, Chinese Academy of Sciences, Beijing 100080, China An important branch of crystallography is the X-ray diffraction analysis of crystal
Direct Methods in Crystallography Fan, Hai-fu Institute of Physics, Chinese Academy of Sciences, Beijing 100080, China An important branch of crystallography is the X-ray diffraction analysis of crystal
From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available
 From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available Scripps EM course, November 14, 2007 What aspects of contemporary
From x-ray crystallography to electron microscopy and back -- how best to exploit the continuum of structure-determination methods now available Scripps EM course, November 14, 2007 What aspects of contemporary
Methoden Moderner Röntgenphysik II - Vorlesung im Haupt-/Masterstudiengang, Universität Hamburg, SoSe 2016, S. Roth
 > 31.05. : Small-Angle X-ray Scattering (SAXS) > 0.06. : Applications & A short excursion into Polymeric materials > 04.06. : Grazing incidence SAXS (GISAXS) Methoden Moderner Röntgenphysik II - Vorlesung
> 31.05. : Small-Angle X-ray Scattering (SAXS) > 0.06. : Applications & A short excursion into Polymeric materials > 04.06. : Grazing incidence SAXS (GISAXS) Methoden Moderner Röntgenphysik II - Vorlesung
shelxl: Refinement of Macromolecular Structures from Neutron Data
 ESS Neutron Protein Crystallography 2013 Aarhus, Denmark shelxl: Refinement of Macromolecular Structures from Neutron Data Tim Grüne University of Göttingen Dept. of Structural Chemistry http://shelx.uni-ac.gwdg.de
ESS Neutron Protein Crystallography 2013 Aarhus, Denmark shelxl: Refinement of Macromolecular Structures from Neutron Data Tim Grüne University of Göttingen Dept. of Structural Chemistry http://shelx.uni-ac.gwdg.de
Experimental phasing, Pattersons and SHELX Andrea Thorn
 Experimental phasing, Pattersons and SHELX Andrea Thorn What is experimental phasing? Experimental phasing is what you do if MR doesn t work. What is experimental phasing? Experimental phasing methods
Experimental phasing, Pattersons and SHELX Andrea Thorn What is experimental phasing? Experimental phasing is what you do if MR doesn t work. What is experimental phasing? Experimental phasing methods
Resolution: maximum limit of diffraction (asymmetric)
 Resolution: maximum limit of diffraction (asymmetric) crystal Y X-ray source 2θ X direct beam tan 2θ = Y X d = resolution 2d sinθ = λ detector 1 Unit Cell: two vectors in plane of image c* Observe: b*
Resolution: maximum limit of diffraction (asymmetric) crystal Y X-ray source 2θ X direct beam tan 2θ = Y X d = resolution 2d sinθ = λ detector 1 Unit Cell: two vectors in plane of image c* Observe: b*
X-ray non-resonant and resonant magnetic scattering Laurent C. Chapon, Diamond Light Source. European School on Magnetism L. C.
 X-ray non-resonant and resonant magnetic scattering Laurent C. Chapon, Diamond Light Source 1 The Diamond synchrotron 3 GeV, 300 ma Lienard-Wiechert potentials n.b: Use S.I units throughout. rq : position
X-ray non-resonant and resonant magnetic scattering Laurent C. Chapon, Diamond Light Source 1 The Diamond synchrotron 3 GeV, 300 ma Lienard-Wiechert potentials n.b: Use S.I units throughout. rq : position
Strong interplay between stripe spin fluctuations, nematicity and superconductivity in FeSe
 Strong interplay between stripe spin fluctuations, nematicity and superconductivity in FeSe Qisi Wang 1, Yao Shen 1, Bingying Pan 1, Yiqing ao 1, Mingwei Ma 2, Fang Zhou 2, P. Steens 3,. Schmalzl 4, T.
Strong interplay between stripe spin fluctuations, nematicity and superconductivity in FeSe Qisi Wang 1, Yao Shen 1, Bingying Pan 1, Yiqing ao 1, Mingwei Ma 2, Fang Zhou 2, P. Steens 3,. Schmalzl 4, T.
Copyright Mark Brandt, Ph.D A third method, cryogenic electron microscopy has seen increasing use over the past few years.
 Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Structure Determination and Sequence Analysis The vast majority of the experimentally determined three-dimensional protein structures have been solved by one of two methods: X-ray diffraction and Nuclear
Protein crystallography. Garry Taylor
 Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
Protein crystallography Garry Taylor X-ray Crystallography - the Basics Grow crystals Collect X-ray data Determine phases Calculate ρ-map Interpret map Refine coordinates Do the biology. Nitrogen at -180
New sample environment options at the neutron diffractometer BioDiff
 Mitglied der Helmholtz-Gemeinschaft New sample environment options at the neutron diffractometer BioDiff Cryostream - closed cycle cryostat - high pressure cell for powder diffraction March 22nd 2013 Tobias
Mitglied der Helmholtz-Gemeinschaft New sample environment options at the neutron diffractometer BioDiff Cryostream - closed cycle cryostat - high pressure cell for powder diffraction March 22nd 2013 Tobias
