research papers Online ion-exchange chromatography for small-angle X-ray scattering
|
|
|
- Abel Dickerson
- 5 years ago
- Views:
Transcription
1 Online ion-exchange chromatography for small-angle X-ray scattering ISSN Stephanie Hutin, a Martha Brennich, b * Benoit Maillot b,c and Adam Round d,e Received 20 November 2015 Accepted 9 August 2016 a Institut de Biologie Structurale, 71 Avenue des Martyrs, Grenoble, France, b European Synchrotron Radiation Facilty, 71 Avenue des Martyrs, Grenoble, France, c Integrated Structural Biology Department, Institut de Génétique et de Biologie Moléculaire et Cellulaire, BP 10142, Illkirch, France, d EMBL Grenoble Outstation, 71 Avenue des Martyrs, Grenoble, France, and e EPSAM, Keele University, Keele, England. *Correspondence brennich@esrf.fr Edited by S. Wakatsuki, Stanford University, USA These authors contributed equally to this work. Current address: Single Particles, Clusters and Biomolecules and Serial Femtosecond Crystallography (SPB/SFX), European XFEL, Schenefeld, Germany. Keywords: BioSAXS; ion-exchange chromatography; online purification. Supporting information: this article has supporting information at journals.iucr.org/d Biological small-angle X-ray scattering (BioSAXS) is a powerful technique to determine the solution structure, particle size, shape and surface-to-volume ratio of macromolecules. However, a drawback is that the sample needs to be monodisperse. To ensure this, size-exclusion chromatography (SEC) has been implemented on many BioSAXS beamlines. Here, the integration of ionexchange chromatography (IEC) using both continuous linear and step gradients on a beamline is described. Background subtraction for continuous gradients by shifting a reference measurement and two different approaches for step gradients, which are based on interpolating between two background measurements, are discussed. The results presented here serve as a proof of principle for online IEC and subsequent data treatment. 1. Introduction Biological small-angle X-ray scattering (BioSAXS) can reveal solution structures in terms of the average particle size and shape of biological macromolecules, as well as information on the surface-to-volume ratio (Graewert & Svergun, 2013; Kikhney & Svergun, 2015; Putnam et al., 2007; Jacques & Trewhella, 2010). This method is an accurate, mostly nondestructive approach, which requires little sample preparation compared with that required for other structural biology techniques, such as crystallography. However, for accurate interpretation the sample is required to be monodisperse; this requirement is often a problem as biological macromolecules can be susceptible to aggregation. Recently, the combination of online size-exclusion chromatography (SEC) and SAXS has been implemented directly on beamlines in order to overcome this obstacle, to ensure data quality and to make this technique more accessible for increasingly difficult samples (Lambright et al., 2013; Round et al., 2013; David & Pérez, 2009; Graewert et al., 2015; Mathew et al., 2004; Watanabe & Inoko, 2009; Acerbo et al., 2015; Grant et al., 2011). Additional techniques, such as time-resolved (TR) SAXS experiments using desalting columns for quick buffer exchanges (Jensen et al., 2010) and differential ultracentrifugation, have been successfully coupled with SAXS (Hynson et al., 2015). However, even when using SEC SAXS, samples can still be difficult to measure, for example when different species do not separate. This can be owing to their size (the difference in molecular mass should be at least 10%), owing to the limited resolution range of the SEC column or owing to the physical properties of the sample such as hydrophobic surfaces, flexibility or lack of stability, e.g. separation of complexes into Acta Cryst. (2016). D72,
2 individual protomers. In these cases, data collection, analysis and interpretation can be difficult. Another common problem is the quantity of sample needed. Many proteins are difficult to express or purify and the required quantity (about 100 ml at 3 mg ml 1 ) of monodisperse sample cannot be obtained. For SEC SAXS measurements, the high degree of dilution on the column [up to a factor of ten depending on the column type and sample (Watanabe & Inoko, 2009; Kirby et al., 2013)] requires even more concentrated samples (5 mg ml 1 or above), although depending on the column less volume (down to 10 ml) might be sufficient. Even if the samples are soluble at these concentrations, they are not always stable and can often aggregate. An alternative purification method is ion-exchange chromatography (IEC). IEC separates ionized molecules based on their total net surface charge, which changes gradually with the ph and/or the salt concentration of the buffer (Karlsson & Hirsh, 2011). This technique enables the separation of molecules of similar size, which can be difficult to separate by other techniques. In general, if the net surface charge of a protein is higher (positive or negative) than that of the IEC resin (anion or cation exchange, respectively), the protein will bind to it. When using a linear salt gradient to increase the charge in the mobile phase, a specific protein will be eluted at a specific salt concentration. Given that different oligomeric states and aggregates differ in their surface area, and hence their surfacecharge distribution, it is also possible to separate oligomeric species (Kluters et al., 2015). IEC has the advantage of providing moderate resolution and, when working with large volumes of dilute samples, concentrating the sample, since the eluate concentration is determined by the capacity of the column and the binding affinity of protein to the column, and not by the initial concentration. Therefore, it is possible to store and transport samples at low concentrations prior to the IEC experiment. Additionally, many IEC columns perform well at flow rates in the millilitre per minute range, which limits the risk of collecting data from radiation-damaged material. However, IEC does require some optimization regarding the ph and the salt gradient (Yigzaw et al., 2009), which can and should be performed prior to any SAXS-coupled data collection. Here, we present a proof of principle for an alternative to SEC SAXS measurements using ion-exchange chromatography (Selkirk, 2004) online with the SAXS experiment. This has been implemented and tested on the BioSAXS beamline BM29 at the European Synchrotron Radiation Facility (ESRF) in Grenoble, France (Pernot et al., 2013). 2. Materials and methods 2.1. Purification of BSA on an ion-exchange column For the preparative offline IEC experiment, 100 ml of a 77 mg ml 1 BSA (lyophilized powder, essentially globulinfree; Sigma) solution in buffer A [20 mm Tris ph 7, 25 mm NaCl, 5% glycerol, 1 mm dithiothreitol (DTT)] was prepared and injected onto a Uno Q-1R (Bio-Rad) column on a Biologic system (Bio-Rad) equilibrated with buffer A. The protein concentration was determined by measuring the absorption at 280 nm with a NanoDrop (NanoDrop 1000, Thermo Fisher) using a mass extinction coefficient of 6.7 for a 1% (10 mg ml 1 ) BSA solution. The high concentration enables easy injection onto the column using injection loops. A salt gradient was made by mixing buffer A with buffer B (20 mm Tris HCl ph 7, 1 M NaCl, 5% glycerol, 1 mm DTT). The flow rate for the offline experiment was 2 ml min 1. For IEC SAXS experiments, only 50 ml was injected using the autosampler of the HPLC system (SIL-20ACXR) and the flow rate was 1 ml min 1. Peak fractions from the preparative run (20 ml per 1500 ml fraction size) were analysed on a 12% SDS PAGE stained with InstantBlue (Expedeon). PageRuler Plus Prestained Protein Ladder (Thermo Fisher) was used as a molecularweight marker D cloning, expression and purification A fragment of the helicase primase D5R representing the D5N and helicase domains, D , was cloned, expressed and purified as described in Hutin et al. (2016). The construct was cloned into the pproex HTb vector (Life Technologies) using the primers 5 0 -GCGCCATGGGTAATAAACTGTTT- AATATTGCAC-3 0 and 5 0 -ATGCAAGCTTTTACGGAGA TGAAATATCCTCTATGA-3 0 and expressed in Escherichia coli BL21 (DE3) Star cells (Novagen). The bacterial pellet was resuspended in lysis buffer (50 mm Tris HCl ph 7, 150 mm NaCl, 5 mm MgCl 2,10mM-mercaptoethanol, 10% glycerol) with complete protease-inhibitor cocktail (Roche) and 1 ml benzonase per 10 ml and lysed by sonication. The supernatant was loaded onto a nickel-affinity column (HIS-Select; Sigma), which was washed with 10 column volumes (CV) of lysis buffer, 10 CV washing buffer (50 mm Tris HCl ph 7, 1 M NaCl, 10 mm -mercaptoethanol, 10% glycerol) and 10 CV imidazole wash (50 mm Tris HCl ph 7, 150 mm NaCl, 10 mm -mercaptoethanol, 10% glycerol, 30 mm imidazole). D was eluted in 20 mm Tris HCl ph 7, 150 mm NaCl, 10 mm -mercaptoethanol, 10% glycerol, 200 mm imidazole and the buffer was exchanged back to the lysis buffer on an Econo-Pac 10DG Desalting column. His-TEV cleavage was performed at room temperature overnight and the cleaved protein was passed over a second nickel column before injection onto a Superose 6 column (GE Healthcare) equilibrated with gel-filtration buffer (20 mm Tris HCl ph 7, 150 mm NaCl, 10% glycerol, 1 mm DTT). The eluted peak fractions of D were then combined and diluted to 25 mm NaCl and 5% glycerol by keeping the buffer concentration equal. 30 ml of the sample, containing about 5 mg of protein, were then loaded onto a Uno Q-1R column (Bio-Rad; buffer A, 20mM Tris ph 7, 25 mm NaCl, 5% glycerol, 1 mm DTT; buffer B, 20mM Tris ph 7, 1 M NaCl, 5% glycerol, 1mM DTT). The protein concentration was estimated by measuring the absorption at 280 nm with a NanoDrop (NanoDrop 1000, Thermo Fisher) using a mass extinction Acta Cryst. (2016). D72, Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering 1091
3 Table 1 Parameters of SAXS data acquisition and analysis. Data-collection parameters Instrument BM29, ESRF Wavelength (Å) 0.99 q-range (Å 1 ) Sample-to-detector distance (m) Exposure time per frame (s) 1 Concentration range n.a. Temperature (K) 293 Detector PILATUS 1M (Dectris) Flux (photons s 1 ) Beam size (mm) Structural parameters for BSA, linear gradient I 0 (from Guinier) (arbitrary units) R g (from Guinier) (Å) q min R g q max R g used for Guinier Theoretical R g (from CRYSOL) (Å) Porod volume V p (from SCÅTTER) (Å 3 ) (116 5) 10 3 Molecular mass M r (from V p ) (kda) 68.2 Calculated monomeric M r from sequence (kda) 66.5 Structural parameters for BSA, step gradient I 0 (from Guinier) (arbitrary units) R g (from Guinier) (Å) q min R g q max R g used for Guinier Theoretical R g (from CRYSOL) (Å) Porod volume V p (from SCÅTTER) (Å 3 ) (116 5) 10 3 Molecular mass M r (from V p ) (kda) 68.2 Calculated monomeric M r from sequence (kda) 66.5 Structural parameters for D , step gradient I 0 (from Guinier) (arbitrary units) R g (from Guinier) (Å) q min R g q max R g used for Guinier I 0 [from P(r)] R g [from P(r)] (Å) D max [from P(r)] (Å) 120 q-range used for P(r) (Å 1 ) Porod volume V p (from SCÅTTER) (Å 3 ) (577 5) 10 3 Molecular mass M r (from V p ) (kda) 338 Calculated monomeric M r from sequence (kda) 321 coefficient of 8.38 for a 1% (10 mg ml 1 )D solution. For offline IEC the flow rate used was 2 ml min 1, which was reduced to 1 ml min 1 in the online experiments. Glycerol was added to all buffers as a co-solvent to enhance the stability of D in aqueous solution in order to prevent protein aggregation. As for BSA, fractions from the preparative run were analysed by SDS PAGE IEC SAXS data collection SAXS data were collected on BM29 at ESRF (Pernot et al., 2013) using a PILATUS 1M detector (Dectris) at a distance of m from the 1.8 mm diameter flowthrough capillary. The scattering of pure water was used to calibrate the intensity to absolute units (Orthaber et al., 2000). The intensities were scaled such that the forward scattering corresponds directly to the concentration (in mg ml 1 ) times the molar mass (in kda) of idealized proteins, i.e. 1 a.u. = cm 1, unless explicitly stated otherwise. Data collection was performed continuously throughout the chromatography run at a frame rate of 1 Hz. The X-ray energy was 12.5 kevand the accessible q-range was nm 1. The incoming flux at the sample position was of the order of photons s 1 in mm. A summary of the acquisition parameters is given in Table 1. All images were automatically azimuthally averaged with pyfai (Ashiotis et al., 2015; Kieffer & Wright, 2013; Kieffer & Karkoulis, 2013). Online purification was performed with a high-pressure liquid-chromatography (HPLC) system (Shimadzu, France) consisting of an inline degasser (DGU-20A5R), a binary pump (LC-20ADXR), a valve for buffer selection and gradients, an auto-sampler (SIL-20ACXR), a UV Vis array spectrophotometer (SPD-M20A) and a conductimeter (CDD- 10AVP). The HPLC system was directly coupled to the flowthrough capillary of the SAXS exposure unit (Round et al., 2015). The flow rate for all online experiments was 1 ml min 1, resulting in a mean passage time of material through the X-ray beam of 0.1 s. Initial evaluation of the data quality was performed using the automatic SEC SAXS processing pipeline available at the BM29 beamline (Brennich et al., 2016; De Maria Antolinos et al., 2015) Background subtraction for BSA on a linear gradient In order to subtract an appropriate background, the method developed by Hynson and coworkers for differential ultracentrifugation (Hynson et al., 2015) was applied. Similarly to the salt gradient in IEC SAXS used here, the background signal in differential ultracentrifugation-coupled SAXS changes owing to a sucrose gradient. The backgroundcorrection method of Hynson and coworkers identifies the appropriate background subtraction as that which provides a stable SAXS signal throughout the peak. The stability of the signal can be assessed by comparing the ratio of scattering in a low-q region to that in a mid-q region: for a stable signal this ratio is constant, whereas for an unstable signal it changes throughout the peak. Incorrect background correction results in a systematic change in the scattering signal depending on the protein concentration. The choice of the regions depends to some extent on the protein of interest and in particular on its size. However, we found that the regions used by Hynson and coworkers ( and nm 1 for low-q and mid-q regions, respectively) suited well: the low-q region reflects the correct overall size of the protein, whereas most of the characteristic features of BSA are present in the mid-q region (see, for example, Fig. 5d). In Hynson et al. (2015) the offset between the individual sample and buffer runs is first determined by matching the high-q region of the scattering profile. This shift is then fine tuned by assessing the variability of the low-q to mid-q ratio for a fine grid of interpolated buffers. In IEC SAXS experiments, the overall change as well as the difference between individual frames in the background signal throughout the gradient and the offset between the buffer and sample measurements are much smaller than in the sucrose gradient used in the ultracentrifugation experiment [compare the bottom part of Fig. 1(b) in this paper with Fig. 2(a) in Hynson et al. (2015)]. As a consequence, the approach can be slightly simplified: the optimal shift between the recorded frames can be determined directly without the 1092 Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering Acta Cryst. (2016). D72,
4 need for interpolation. To achieve this, we shift the buffer run with respect to the sample run in steps of five frames and subtract frame-wise. The low-q to mid-q ratio is then calculated for each frame and the variability in the region of interest is assessed. The perfect shift would result in a parallel line in the ratio versus frame-number plot throughout the region, whereas systematic under-subtraction results in a concave shape and over-subtraction in a convex shape. Using this approach, we can determine the best shift with an error of ten frames. This might seem less accurate in comparison with the subframe precision of Hynson and coworkers, but the difference between individual buffer frames in an IEC gradient is much lower than in differential ultracentrifugation [a CORMAP test (Franke et al., 2015) on 20 frames in the region of interest showed no significant deviation]. This implies that the effect of these ten frames on the subtraction is small SAXS data analysis To compare the scattering from different frames, the ratio of scattering in the low-q region ( nm 1 ) to the high-q region ( nm 1 ) was calculated in the same manner as in Hynson et al. (2015). For easier comparison between different buffer subtractions, the result was normalized to give the same mean over the region of interest. Radii of gyration were calculated using AUTORG from the ATSAS package (Petoukhov et al., 2007), P(r) functions were calculated using GNOM (Svergun, 1992) and Porod volumes were estimated using the Volume interface of SCÅTTER (available at The estimation of molecular mass based on the Porod analysis was performed by dividing the Porod volume by 1.7 nm 3 kda 1 (Petoukhov et al., 2007). All other manipulations were performed in the Spyder interface for Python 3.4 using the NumPy and SciPy packages (Jones et al., 2001). The scripts used are available at maaeli/iec. As the background scattering is dependent on the buffer composition, the gradient or stepwise buffer changes during elution must be accounted for in the subtraction process. We present three methods to find the optimal background subtractions, which are explained for the individual cases in x3. Known BSA crystal structures were compared with the experimental data using CRYSOL (Svergun et al., 1995). Default fitting parameters were used, which allow the hydration shell to be adjusted to optimize the fit. Figure 1 Linear gradient IEC SAXS performed on BSA. (a) Standard ionexchange chromatogram of BSA on the Uno Q-1R column, highlighting the locations of the three peaks. Orange line, percentage of buffer B; red line, resulting conductivity; violet line, absorbance at 280 nm. (b) IEC SAXS chromatograms of BSA using a linear salt gradient. Top panel, UV absorbance (violet) and total scattering intensity (green). Middle and bottom panels, chromatograms of sample (red) and buffer (black) runs at q =1nm 1 (middle) and 4 nm 1 (bottom). (c) Difference between the scattering intensities measured at q =4nm 1 in the sample and buffer runs (blue symbols). The dotted orange line illustrates the salt gradient. (d) Comparison of the scattering ratio ( nm 1 versus nm 1 ) for background correction using a shift of 100 frames (cyan), 130 frames (blue) and 160 frames (green) between the sample and buffer runs. The dotted black line serves as a visual aid for a constant line. 3. Results and discussion 3.1. IEC SAXS of bovine serum albumin using a linear salt gradient The first test case was bovine serum albumin (BSA), using a standard linear salt gradient for elution and a blank run of the same gradient for buffer subtraction. We chose BSA as it is known to form higher oligomers and aggregates in solution (Folta-Stogniew & Williams, 1999), allowing us to assess the ability of the method to reduce sample polydispersity. We performed a standard BSA run on the ion-exchange column using an offline FLPC system, evaluated the chromatogram and observed several peaks in the first third of the chromatogram (Fig. 1a). Their approximate buffer B percentages were determined to be 10, 12 and 16%, corresponding to NaCl concentrations of 122.5, 142 and 181 mm, respectively. SDS PAGE of different fractions throughout the peak confirmed that the first two peaks correspond to BSA, while the third peak is a higher molecular-mass contaminant (Fig. 2a). Acta Cryst. (2016). D72, Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering 1093
5 We then ran the same sample online on the BM29 beamline at ESRF, as described in x2, and continuously acquired SAXS data from the eluent at a rate of 1 Hz. In order to improve the resolution of the peaks and to ensure a sufficiently high sampling rate, the flow rate for the online experiments was reduced to 1 ml min 1. The UV absorption signal at 280 nm and the total scattering elution profiles show the expected three peaks (Fig. 1b, top), whereas the scattering at higher angles increases linearly (Fig. 1b, bottom). The matching background was found to be slightly shifted with respect to the buffer run (compare the bottom graph in Fig. 1b with Fig. 1c). It is therefore necessary to subtract not the directly corresponding frames but slightly shifted ones. Possible reasons for this shift are a nonperfect synchronization between the ion-exchange run and the SAXS data acquisition or the co-elution of ions bound to the column. To identify the optimal shift, as described in x2, we applied the approach developed by Hynson et al. (2015) for differential ultracentrifugation coupled to SAXS. For each shift the ratio of scattering at low q ( nm 1 ) versus mid-to-high q ( nm 1 ) is calculated for each frame. If the sample scatters in the same way throughout the peak, this value should not change and a constant value over the peak indicates a suitable background subtraction. Fig. 1(d) shows how the ratio changes in the region of interest for different values of the shift. For the first two peaks the ideal shift was found to be frames, which corresponds to about 2.2 ml. In addition, the ratio was constant over both peaks, indicating that one species elutes in a double peak. However, for the third peak no shift giving a flat line was found, indicating that the scattering from the sample is not constant throughout the peak and that the peak therefore represents more than one species. The SDS PAGE from the offline run shows that in this peak BSA is contaminated by a larger protein species (Fig. 2a). The forward scattering and the radius of gyration obtained using AUTORG (Petoukhov et al., 2007) were calculated from subtracted curves and the mass was estimated via the correlated volume approach (Rambo & Tainer, 2013). Fig. 3(a) confirms that, as expected for a single species, both the mass and the radius of gyration are constant throughout the initial double peak. Based on the forward scattering intensity (Mylonas & Svergun, 2007), and assuming monomeric BSA (66.5 kda), the maximum protein concentration throughout the peak was estimated at 1.75 mg ml 1, which is well below the regime in which significant structure-factor contributions to the signal would become relevant (Skou et al., 2014; Zhang et al., 2007). An additional comparison of individual frames throughout the double peak shows that the SAXS signal is identical throughout the peak, with the lowest Figure 2 (a) 10% SDS PAGE of fractions of BSA along the salt gradient collected after the Uno Q-R1 column run. Pure BSA is indicated in teal and the contaminant is indicated in cyan. The protein ladder is labelled in kda. (b) Fractions from a Uno Q-R1 column run of D along the salt gradient. The red box frames the D protein, while the magenta box indicates the contaminant. Figure 3 Background analysis of linear gradient IEC SAXS on BSA. (a) Forward scattering (black), radius of gyration (green) and mass (red) based on the correlated volume for the background-corrected curves in the region of interest. The dotted lines correspond to the expected radius of gyration (dark green) and molecular mass (dark red) of BSA. The grey area indicates which frames were used for subsequent averaging; the arrows indicate the positions of the individual curves shown in (b) and (c). (b) Individual scattering curves and (c) Guinier plots at different positions throughout the chromatogram (the curves are offset for clarity). The colour code and positions in the chromatogram are as indicated in (a). (d) Averaged scattering curve from the region indicated in grey in (a). (e) Guinier fit (light green line) of the scattering curve shown in (d). Open symbols represent points that were not used for fitting. The lower panel shows the residuals. ( f) Differences in scattering between the curve found by shifting the buffer run by 130 frames and a clearly under-subtracted curve (120 frames shift, orange) and over-subtracted curve (160 frames shift, blue) Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering Acta Cryst. (2016). D72,
6 adjusted p-value in a CORMAP test (Franke et al., 2015) being 0.16 (Figs. 3b and 3c). This suggests that the two subpeaks do not represent different conformations of BSA. 169 frames were averaged in the region of interest shown (grey) in Fig. 3(a). The resulting curve (Fig. 3d) gives a radius of gyration of nm (Fig. 3e) and a Porod volume of 116 5nm 3, corresponding to a molecular weight of about 68 kda (Petoukhov et al., 2007; see Table 1 for further details). Both of these values correspond well to the expected size (2.77 nm; PDB entry 3v03; Majorek et al., 2012) and molecular weight (66.5 kda) of monomeric BSA. Additional comparison to the monomeric crystal structure (PDB entry 3v03; Majorek et al., 2012) shows a similarly good match as the SEC SAXS data for BSA ( 2 = 1.82; Fig. 5d and Supplementary Fig. S1b). To estimate to what extent an incorrect shift affects these results, the corresponding averages for shifts of 120 and 140 frames, respectively, were calculated. The absolute differences from the 130-frame shift are shown in Fig. 3(f). In both cases, above 1nm 1 the curve is shifted by a small constant, which many modelling algorithms take into account (Knight & Hub, 2015; Petoukhov et al., 2007; Svergun et al., 1995). Below 1 nm 1 differences in the relative contribution of capillary scattering result in a q-dependent difference, which could in principle affect modelling. However, these differences contribute less than 0.2% to the scattering signal and thus their influence is negligible. the two buffers recorded before and after the peak. As the scattering of BSA at high angles is rather low and the increase in signal above q =4.5nm 1 does not follow the protein concentration (Fig. 4a), one can assume that the changes in signal in this region are only owing to the difference in background. In order to limit the effect of the rather high noise level in this region, the increase is modelled with a 3.2. IEC SAXS of bovine serum albumin using a stepwise salt gradient The linear gradient IEC works well for many proteins, but for samples where the elution peak is broad or is not well separated from other peaks, manual selection of appropriate steps in the gradient can ensure a purer and higher peak on faster timescales. For these cases, we describe an elution system in which the salt concentration is increased in predefined steps. The advantage of this approach is that by choosing a sufficiently long step length, it is possible to use buffer measurements from the same chromatography run for background correction, reducing the risk of nonmatching buffers owing to slow drifts. On the downside, even if the change in the buffer mixing ratio at the pumps is instantaneous, various effects, such as Taylor dispersion of flow in the capillary (see, for example, Wunderlich et al., 2014), co-elution of small molecules from the columns and the creation of new interaction sites for salt on the column and the eluted protein, result in non-instantaneous gradients at the measurement position. To show the validity of this approach, we again used BSA with the salt concentrations determined above from the offline IEC run. The elution steps were chosen in such a way that the first two subpeaks of the BSA elute in the same step. In this case, BSA elutes as a single peak (Fig. 4a). As expected, the background scattering at high angles does not increase instantaneously, but saturates slowly after a steep initial increase. In order to subtract an appropriate background from each measured frame, it is necessary to interpolate between Figure 4 Stepwise-gradient IEC SAXS performed on BSA. (a) IEC SAXS chromatograms of BSA using a stepwise salt gradient. Top panel, UV absorbance (violet) and total scattering intensity (green). Middle and bottom panels, chromatograms at 1 and 4 nm 1, respectively. (b) The buffer signal (mean scattering above 4.5 nm 1, blue) increases slowly during the elution of the peak (total scattering intensity, green) and can be modelled by an exponential decay (red) between the buffer before the peak (I) and after the peak (II). (c) Comparison of the scattering ratio ( nm 1 versus nm 1 ) for background correction using the buffer before the peak (I, cyan), after the peak (II, green) and the modelled buffer (blue). (d) Forward scattering (black), radius of gyration (green) and mass (red) based on the correlated volume for the background-corrected curves in the peak region. The grey area indicates the frames that were used for subsequent averaging; the arrows indicate the positions of the individual curves presented in (e) and ( f). (e) Individual scattering curves and ( f) Guinier plots at different positions throughout the chromatogram (curves are offset for clarity). The colour code and the positions in the chromatogram are as indicated in (d). Acta Cryst. (2016). D72, Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering 1095
7 continuous function instead of using the high-q region of each frame to interpolate the buffer individually. A least-parameter model for an asymmetric, saturating increase as observed here is a single exponential decay from the buffer before (region I) to the buffer after (region II) the peak, i.e. afittoii (II I)exp[ (N N 0 )/N 0 ], where N is the frame number, N 0 is a constant offset and 1/N 0 is the rate of increase. To obtain the correct values for N 0 and N 0, the mean of the data was fitted above 4.5 nm 1 (Fig. 4b). The best fit is obtained with N 0 = and N 0 =17 3, giving an 2 of These parameters match the starting point of the increase (frame 1290) and the estimated half-life of 12 2 frames well and allow us to model the entire q-range. This procedure allows the background to be subtracted individually for each frame and a constant scattering from the sample throughout the peak to be confirmed by assessing the stability of the ratio of scattering at low q versus mid/high q,as discussed above (Figs. 4c and 4d). Individual comparison of frames confirmed this constant signal (Figs. 4e and 4f). Based on the forward scattering, the maximum concentration in this case was only 0.35 mg ml 1. After averaging (Fig. 5a), a scattering curve which corresponds to monomeric BSA ( 2 = 3.63; Fig. 5d) was found. The radius of gyration is nm (Fig. 5b) and the Porod volume is 116 5nm 3, matching previous results (also see Table 1). This shows that important structural parameters can be determined using either linear or stepwise gradients. In order to estimate the mis-subtraction of the buffer and its effect on the conclusions that can be drawn from the data, one needs to determine corresponding over-subtracted and undersubtracted curves. We decided to over-subtract by directly subtracting buffer II and to under-subtract by subtracting the mean of buffer I and buffer II. This choice might be more extreme than strictly necessary, but provides an upper limit of the effect of the mis-subtraction. The differences in this case (Fig. 5c) are much more pronounced than for a linear gradient case (Fig. 3f). This means that the background subtraction for the linear case is more accurate. Above 1 nm 1, buffer mismatch causes a constant shift in the signal. Below 1 nm 1, the differences are larger but do not impact the structural parameters determined from this region, with the radius of gyration remaining at nm and the Porod volume increasing within its error to 120 5nm 3 only for the subtraction of buffer II and remaining unchanged for the other case. This implies that while interpretations of domain arrangements might be affected by buffer mismatches, the observed overall shape (size, anisotropy,...) of the protein can still be determined despite the larger inaccuracy in the background subtraction. Direct comparison of the two scattering curves of BSA obtained using the two different IEC SAXS approaches (Fig. 5d) shows two very similar curves. Owing to the lower protein concentration at the peak, the curve resulting from the step-gradient method is noisier. However, from about 2.5 nm 1 onwards the signal determined using the linear gradient method decreases more steeply. In particular, above 3.5 nm 1 the signal determined using the step gradient turns upward. This increase also results in a clear deviation from the predicted signal and is responsible for the higher 2. Figure 5 Stepwise-gradient IEC SAXS analysis of averaged BSA data. (a) Scattering curve based on averaging of the region indicated in grey in Fig. 4(d). (b) Guinier fit (light blue line) of the scattering curve shown in (a). Open symbols represent points that were not used for fitting. The lower panel shows the residuals. (c) Differences in scattering between the curve found by interpolating the buffer and a clearly under-subtracted curve (average of buffers from regions I and II, orange) and oversubtracted curve (buffer from region II, blue). (d) Fits to the monomeric crystal structure of BSA (PDB entry 3v03; Majorek et al., 2012) eluted with a linear gradient (green) and a stepwise gradient (blue) IEC SAXS of the helicase primase protein D using a stepwise salt gradient To demonstrate the applicability of this approach to a novel sample, we selected D , the D5N and helicase domains of the Vaccinia virus helicase primase D5. The protein fragment forms a hexamer ( kda) that is required for its activity (Hutin et al., 2016). After two nickel columns and a size-exclusion chromatography step, a contaminant still persisted and required an additional purification step via ionexchange chromatography (Fig. 2b), followed by an additional size-exclusion chromatography step. Each step takes time and about 30 50% of the material is lost in total. To optimize the data-collection strategy it would be advantageous to measure the hexamer directly from the Uno Q-1R column. In the standard offline IEC experiment using a linear gradient, three not completely separated peaks at 11, 14 and 16% high-salt buffer were observed (Fig. 6a). When using a stepwise gradient a sharp peak is obtained (Fig. 6b) at 12% salt buffer (142 mm NaCl) and small addi Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering Acta Cryst. (2016). D72,
8 3.4. Additional considerations One of the major difficulties in online SEC SAXS is the effect of radiation damage on the observed signal. The continuously changing signal makes assessment of radiationresearch papers tional peaks in the subsequent steps. This indicates that D elutes completely at 142 mm NaCl and the peaks are better separated than in the linear-gradient elution procedure (Fig. 2a). The scattering at high q (4 nm 1 ) slightly overshoots the scattering level expected from the buffer run (Fig. 6c), probably owing to the co-elution of small ions. To check that the sample composition does not change throughout the peak, the low-salt buffer from each frame was subtracted and their radii of gyration were compared. This operation is not affected by the small change of background throughout the peak (data not shown). Based on the forward scattering from this preliminary processing, the maximum protein concentration is estimated to be 3.4 mg ml 1, corresponding to a 20-fold increase from the injected concentration of 0.17 mg ml 1. Owing to the irregular changes in the background signal that were observed in these data, the approaches to modelling the change in background signal on a frame-by-frame basis that were applied to the previous examples are not applicable. To overcome this issue, first the frames in the peak were averaged in order to improve the signal-to-noise ratio in the high-q region, and a matching buffer from a salt-concentration series measured directly before the ion-exchange run was then chosen by comparing the average scattering in the q-range between 4.25 and 4.75 nm 1. The resulting curve (Fig. 7a) has a radius of gyration of 4.7 nm (Fig. 7b) and a Porod volume of 577 5nm 3, corresponding to a mass of 338 kda (Petoukhov et al., 2007), matching the hexamer mass of 320 kda well (see also Table 1). The P(r) function (Fig. 7c) shows one peak that is slightly shifted to larger distances. This shape, in addition to the clear minima, hints at a mostly spherical shape with a cavity and matches the expected molecular shape well (Hutin et al., 2016). As the steps were chosen more finely than in the previous experiment with BSA, it is easier to estimate the degree of buffer mismatch in the D experiment. The extent to which it influences the interpretation of the results can be estimated by selecting buffers from a higher and a lower salt step of the buffer run (Fig. 7d). In both cases the resulting curve is distinguished from the reported curve by a small constant above q =0.8nm 1. Below q =0.8nm 1 the deviations are no longer constant but are below 0.1% of the signal, and therefore do not influence the interpretation. Figure 6 IEC SAXS of D (a) Ion-exchange chromatogram of D on the Uno Q-1R column with the three peaks indicated. Orange line, percentage of buffer B; red line, the resulting conductivity; violet line, absorbance at 280 nm. (b) IEC SAXS chromatograms of D using a stepwise salt gradient. Top panel, UV absorbance (violet) and total scattering intensity (green). Middle and bottom panels, chromatograms of sample (red) and buffer (black) runs at q =1nm 1 (middle) and 4 nm 1 (bottom). (c) Enlargement of the lower panel in (b): in the protein run (red) the signal at high q overshoots in comparison to the buffer run. Figure 7 (a) Average sample (violet) and buffer (black) curves for the D protein, the resulting subtraction (red) and the fit for calculating the P(r) function (blue). (b) Guinier fit (black line) of the scattering curve shown in (a). Open symbols represent points that were not used for fitting. The lower panel shows the residuals. (c) Pair distance distribution function of the average curve, showing a single peak. (d) Differences in scattering between the curve based on the best-matching buffer and a clearly undersubtracted curve (preceding buffer step, orange) and over-subtracted curve (subsequent buffer step, blue). Acta Cryst. (2016). D72, Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering 1097
9 induced changes to the SAXS signal difficult. In addition, radiation-damaged material often displays a tendency to adhere to the surfaces of the sample environment (Brookes et al., 2013; Jeffries et al., 2015). IEC SAXS can be performed at relatively high flow rates (1 ml min 1 for the studies in this publication); consequently, the average dwelling time of material in the X-ray beam is shorter than for standard SEC SAXS experiments and the contribution of damaged material to the signal is reduced. Furthermore, the higher flow rate reduces the risk of damaged material spoiling the sample environment by adhering to the surface (Epstein, 1997). 4. Conclusions For many macromolecules in solution, ion-exchange chromatography is a required step and is the best-suited purification method, as it can often separate similarly sized proteins. However, the IEC step is usually followed by another purification step, or dialysis, to ensure optimal buffer subtraction. Here, we demonstrate for the first time the application of ionexchange chromatography directly prior to SAXS measurements. The linear gradient method presented here is analogous to a standard ion-exchange protocol and does not inherently require additional offline tests. In addition, background correction can be achieved by correct alignment of a buffer run without any need for interpolation. As each frame can be corrected individually, it is possible to confirm a stable signal throughout the peak. However, owing to the continuity of the gradient, sharp and well separated peaks are beneficial. By carefully selecting the salt concentration of the steps in the step-elution method, this method allows the separation of the peaks to be improved, as no limit exists on the number of substeps. However, background subtraction requires more caution, as the discontinuity of the elution gradient results in the discontinuous co-elution of small ions. In our first example (BSA), it was still possible to correct the background individually for each frame by interpolating the buffer signal. In our second example (D ) it was not possible to interpolate the background throughout the peak, and the correction by framewise comparisons was not applicable. Nevertheless, the correct background for the average signal can be found by measuring a variety of mixing ratios prior to the experiment and carefully choosing the matching one. Despite this difficulty, we are confident that it can be applied to a wide variety of biological macromolecules. Online ion-exchange chromatography adds another important biochemical purification method to the repertoire of purification methods which can be coupled with SAXS (Round et al., 2013; David & Pérez, 2009; Graewert et al., 2015; Jensen et al., 2010; Hynson et al., 2015; Mathew et al., 2004; Watanabe & Inoko, 2009). Proteins of similar apparent molecular weight, but different net surface charges, can now be separated online using the IEC SAXS method. While background subtraction is slightly less straightforward than for SEC SAXS, the variation in the background is smaller than for differential ultracentrifugation coupled with SAXS owing to the high sugar content of the latter (from 15 to 35% sucrose; Hynson et al., 2015). The background-correction method here for D , based on averages, can be performed using any software package for SAXS data analysis, such as PRIMUS (Konarev et al., 2003) or SCÅTTER, assuming that the possible buffer range has been well determined experimentally. For IEC, buffer conditions, such as salt concentrations or ph, are primarily determined by the biochemical characteristics of the protein of interest, such as its isoelectric point and solubility (Yigzaw et al., 2009). However, one should aim to minimize the amounts of all additives in order to maximize the X-ray contrast between the protein and the buffer in a SAXS experiment. As with any SAS data, IEC SAXS results need to be carefully validated. Special care should be paid to sample monodispersity, which cannot be directly assessed by IEC SAXS, and correct background subtraction, especially in the case of flexible proteins (Jacques et al., 2012; Jacques & Trewhella, 2010). It is important to estimate how missubtraction would affect the conclusions drawn from the data. For the results shown in this paper, we have shown that deviations from the ideal background subtraction are small enough to allow reliable conclusions. In conclusion, IEC SAXS is a useful technique that allows accurate data collection with fewer preparation steps and minimizing the loss of time and sample. Proof of principle of a simple elution method is demonstrated in this paper with the option to optimize species separation by using steps in the salt gradient. Validation of three different approaches to achieve optimal background subtractions is also given. A suitable combination of elution method and background subtraction can be chosen to suit the sample of interest best and to provide necessary information for data validation (as demonstrated), so that the resulting data can be used with confidence for subsequent analysis and modelling using the standard tools available within the scientific community. The online SEC system installed at BM29 can routinely perform these experiments and is available for user access on request. Acknowledgements We thank the ESRF for SAXS beam time and support. We further wish to acknowledge Wim Burmeister from Université Grenoble Alpes for fruitful discussions and financial support. This work was supported by an ANR grant (REPLIPOX, ANR-13-BSV8-0014). We would like to thank Petra Pernot and Matthew Bowler for proofreading the manuscript. References Acerbo, A. S., Cook, M. J. & Gillilan, R. E. (2015). J. Synchrotron Rad. 22, Ashiotis, G., Deschildre, A., Nawaz, Z., Wright, J. P., Karkoulis, D., Picca, F. E. & Kieffer, J. (2015). J. Appl. Cryst. 48, Brennich, M. E., Kieffer, J., Bonamis, G., De Maria Antolinos, A., Hutin, S., Pernot, P. & Round, A. (2016). J. Appl. Cryst. 49, Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering Acta Cryst. (2016). D72,
10 Brookes, E., Pérez, J., Cardinali, B., Profumo, A., Vachette, P. & Rocco, M. (2013). J. Appl. Cryst. 46, David, G. & Pérez, J. (2009). J. Appl. Cryst. 42, De Maria Antolinos, A., Pernot, P., Brennich, M. E., Kieffer, J., Bowler, M. W., Delageniere, S., Ohlsson, S., Malbet Monaco, S., Ashton, A., Franke, D., Svergun, D., McSweeney, S., Gordon, E. & Round, A. (2015). Acta Cryst. D71, Epstein, N. (1997). Exp. Therm. Fluid Sci. 14, Folta-Stogniew, E. & Williams, K. (1999). J. Biomol. Tech. 10, Franke, D., Jeffries, C. M. & Svergun, D. I. (2015). Nature Methods, 12, Graewert, M. A., Franke, D., Jeffries, C. M., Blanchet, C. E., Ruskule, D., Kuhle, K., Flieger, A., Schäfer, B., Tartsch, B., Meijers, R. & Svergun, D. I. (2015). Sci. Rep. 5, Graewert, M. A. & Svergun, D. I. (2013). Curr. Opin. Struct. Biol. 23, Grant, T. D., Luft, J. R., Wolfley, J. R., Tsuruta, H., Martel, A., Montelione, G. T. & Snell, E. H. (2011). Biopolymers, 95, Hutin, S., Ling, W. L., Round, A., Effantin, G., Reich, S., Iseni, F., Tarbouriech, N., Schoehn, G. & Burmeister, W. P. (2016). J. Virol. 90, Hynson, R. M. G., Duff, A. P., Kirby, N., Mudie, S. & Lee, L. K. (2015). J. Appl. Cryst. 48, Jacques, D. A., Guss, J. M., Svergun, D. I. & Trewhella, J. (2012). Acta Cryst. D68, Jacques, D. A. & Trewhella, J. (2010). Protein Sci. 19, Jeffries, C. M., Graewert, M. A., Svergun, D. I. & Blanchet, C. E. (2015). J. Synchrotron Rad. 22, Jensen, M. H., Toft, K. N., David, G., Havelund, S., Pérez, J. & Vestergaard, B. (2010). J. Synchrotron Rad. 17, Jones, E., Oliphant, T. E. & Peterson, P. (2001). SciPy: Open Source Scientific Tools for Python. Karlsson, E. & Hirsh, I. (2011). Methods Biochem. Anal. 54, Kieffer, J. & Karkoulis, D. (2013). J. Phys. Conf. Ser. 425, Kieffer, J. & Wright, J. (2013). Powder Diffr. 28, S339 S350. Kikhney, A. G. & Svergun, D. I. (2015). FEBS Lett. 589, Kirby, N., Mudie, S., Hawley, A., Mertens, H. D. T., Cowieson, N., Samardzic Boban, V., Felzmann, U., Mudie, N., Dwyer, J., Beckham, S., Wilce, M., Huang, J., Sinclair, J. & Moll, A. (2013). Trans. Am. Crystallogr. Assoc. 44, 3. documents/2013 Transactions/3-Kirby.pdf. Kluters, S., Wittkopp, F., Jöhnck, M. & Frech, C. (2015). J. Sep. Sci. 39, Knight, C. J. & Hub, J. S. (2015). Nucleic Acids Res. 43, W225 W230. Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. (2003). J. Appl. Cryst. 36, Lambright, D., Malaby, A. W., Kathuria, S. V., Nobrega, R. P., Bilsel, O., Matthews, C. R., Muthurajan, U., Luger, K., Chopra, R. & Irving, T. C. (2013). Trans. Am. Crystallogr. Assoc. 44, 1. Transactions/ 1-Chakravarthy.pdf. Majorek, K. A., Porebski, P. J., Dayal, A., Zimmerman, M. D., Jablonska, K., Stewart, A. J., Chruszcz, M. & Minor, W. (2012). Mol. Immunol. 52, Mathew, E., Mirza, A. & Menhart, N. (2004). J. Synchrotron Rad. 11, Mylonas, E. & Svergun, D. I. (2007). J. Appl. Cryst. 40, s245 s249. Orthaber, D., Bergmann, A. & Glatter, O. (2000). J. Appl. Cryst. 33, Pernot, P. et al. (2013). J. Synchrotron Rad. 20, Petoukhov, M. V., Konarev, P. V., Kikhney, A. G. & Svergun, D. I. (2007). J. Appl. Cryst. 40, s223 s228. Putnam, C. D., Hammel, M., Hura, G. L. & Tainer, J. A. (2007). Q. Rev. Biophys. 40, Rambo, R. P. & Tainer, J. A. (2013). Nature (London), 496, Round, A., Brown, E., Marcellin, R., Kapp, U., Westfall, C. S., Jez, J. M. & Zubieta, C. (2013). Acta Cryst. D69, Round, A., Felisaz, F., Fodinger, L., Gobbo, A., Huet, J., Villard, C., Blanchet, C. E., Pernot, P., McSweeney, S., Roessle, M., Svergun, D. I. & Cipriani, F. (2015). Acta Cryst. D71, Selkirk, C. (2004). Methods Mol. Biol. 244, Skou, S., Gillilan, R. E. & Ando, N. (2014). Nature Protoc. 9, Svergun, D. I. (1992). J. Appl. Cryst. 25, Svergun, D., Barberato, C. & Koch, M. H. J. (1995). J. Appl. Cryst. 28, Watanabe, Y. & Inoko, Y. (2009). J. Chromatogr. A, 1216, Wunderlich, B., Nettels, D. & Schuler, B. (2014). Lab Chip, 14, Yigzaw, Y., Hinckley, P., Hewig, A. & Vedantham, G. (2009). Curr. Pharm. Biotechnol. 10, Zhang, F., Skoda, M. W., Jacobs, R. M., Martin, R. A., Martin, C. M. & Schreiber, F. (2007). J. Phys. Chem. B, 111, Acta Cryst. (2016). D72, Hutin et al. Online ion-exchange chromatography for small-angle X-ray scattering 1099
ID14-EH3. Adam Round
 Bio-SAXS @ ID14-EH3 Adam Round Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and
Bio-SAXS @ ID14-EH3 Adam Round Contents What can be obtained from Bio-SAXS Measurable parameters Modelling strategies How to collect data at Bio-SAXS Procedure Data collection tests Data Verification and
Data reduction and processing tutorial
 Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
Data reduction and processing tutorial Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group EMBL BioSAXS beamline X33, 2012 Optics Vacuum cell Completely redesigned 2005-2012
SAXS and SANS facilities and experimental practice. Clement Blanchet
 SAXS and SANS facilities and experimental practice Clement Blanchet SAS experiment Detector X-ray or neutron Beam Sample 2 s Buffer X-rays Roengten, 1895 Electromagnetic wave The electromagnetic spectrum
SAXS and SANS facilities and experimental practice Clement Blanchet SAS experiment Detector X-ray or neutron Beam Sample 2 s Buffer X-rays Roengten, 1895 Electromagnetic wave The electromagnetic spectrum
BM29 biosaxs data processing tutorial
 HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
HERCULES 2014 BM29 biosaxs data processing tutorial Page 2 OUTLINE Sample Changer Primary Data Processing Model Validation HPLC-SAXS Primary Data Processing Model Validation Ab Initio Model Software in
Introduction to Biological Small Angle Scattering
 Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
Introduction to Biological Small Angle Scattering Tom Grant, Ph.D. Staff Scientist BioXFEL Science and Technology Center Hauptman-Woodward Institute Buffalo, New York, USA tgrant@hwi.buffalo.edu SAXS Literature
How to judge data quality
 SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
SSRL Workshop: Small-Angle X-ray Scattering and Diffraction Studies, March 28-30, 2016 How to judge data quality Tsutomu Matsui SSRL Lab / Dept. of Chemistry Stanford University Subject of this session
SI Text S1 Solution Scattering Data Collection and Analysis. SI references
 SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
SI Text S1 Solution Scattering Data Collection and Analysis. The X-ray photon energy was set to 8 kev. The PILATUS hybrid pixel array detector (RIGAKU) was positioned at a distance of 606 mm from the sample.
Acta Cryst. (2017). D73, , doi: /s
 Supporting information Volume 73 (2017) Supporting information for article: 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update
Supporting information Volume 73 (2017) Supporting information for article: 2017 publication guidelines for structural modelling of small-angle scattering data from biomolecules in solution: an update
SAXS/SANS data processing and overall parameters
 EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
EMBO Global Exchange Lecture Course 30 November 2012 Hyderabad India SAXS/SANS data processing and overall parameters Petr V. Konarev European Molecular Biology Laboratory, Hamburg Outstation BioSAXS group
Small Angle X-Ray Solution Scattering of Biological Macromolecules
 Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
Small Angle X-Ray Solution Scattering of Biological Macromolecules Emre Brookes UltraScan Workshop 15 June 2014 Overview Experimental method Sample preparation Experimental data analysis Experimental method
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands
 Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Characterizing Biological Macromolecules by SAXS Detlef Beckers, Jörg Bolze, Bram Schierbeek, PANalytical B.V., Almelo, The Netherlands This document was presented at PPXRD - Pharmaceutical Powder X-ray
Small-Angle Scattering Atomic Structure Based Modeling
 Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Small-Angle Scattering Atomic Structure Based Modeling Alejandro Panjkovich EMBL Hamburg 07.12.2017 A. Panjkovich (EMBL) BioSAS atomic modeling 07.12.2017 1 / 49 From the forest to the particle accelerator
Modelling against small angle scattering data. Al Kikhney EMBL Hamburg, Germany
 Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Modelling against small angle scattering data Al Kikhney EMBL Hamburg, Germany Validation of atomic models CRYSOL Rigid body modelling SASREF BUNCH CORAL Oligomeric mixtures OLIGOMER Flexible systems EOM
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013
 Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Biological Small Angle X-ray Scattering (SAXS) Dec 2, 2013 Structural Biology Shape Dynamic Light Scattering Electron Microscopy Small Angle X-ray Scattering Cryo-Electron Microscopy Wide Angle X- ray
Interaction of Gold Nanoparticle with Proteins
 Chapter 7 Interaction of Gold Nanoparticle with Proteins 7.1. Introduction The interfacing of nanoparticle with biomolecules such as protein is useful for applications ranging from nano-biotechnology (molecular
Chapter 7 Interaction of Gold Nanoparticle with Proteins 7.1. Introduction The interfacing of nanoparticle with biomolecules such as protein is useful for applications ranging from nano-biotechnology (molecular
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1.
 Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Table S1. Overview of used PDZK1 constructs and their binding affinities to peptides. Related to figure 1. PDZK1 constru cts Amino acids MW [kda] KD [μm] PEPT2-CT- FITC KD [μm] NHE3-CT- FITC KD [μm] PDZK1-CT-
Supplementary materials. Crystal structure of the carboxyltransferase domain. of acetyl coenzyme A carboxylase. Department of Biological Sciences
 Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
Supplementary materials Crystal structure of the carboxyltransferase domain of acetyl coenzyme A carboxylase Hailong Zhang, Zhiru Yang, 1 Yang Shen, 1 Liang Tong Department of Biological Sciences Columbia
radiation damage Development of tools to automate quantitative analysis of radiation damage in SAXS experiments 1. Introduction
 Development of tools to automate quantitative analysis of radiation damage in SAXS experiments ISSN 1600-5775 Jonathan C. Brooks-Bartlett, a Rebecca A. Batters, a Charles S. Bury, a Edward D. Lowe, a Helen
Development of tools to automate quantitative analysis of radiation damage in SAXS experiments ISSN 1600-5775 Jonathan C. Brooks-Bartlett, a Rebecca A. Batters, a Charles S. Bury, a Edward D. Lowe, a Helen
Protein separation and characterization
 Address:800 S Wineville Avenue, Ontario, CA 91761,USA Website:www.aladdin-e.com Email USA: tech@aladdin-e.com Email EU: eutech@aladdin-e.com Email Asia Pacific: cntech@aladdin-e.com Protein separation
Address:800 S Wineville Avenue, Ontario, CA 91761,USA Website:www.aladdin-e.com Email USA: tech@aladdin-e.com Email EU: eutech@aladdin-e.com Email Asia Pacific: cntech@aladdin-e.com Protein separation
Sample preparation and characterization around SAXS
 Sample preparation and characterization around SAXS Experimental verification and validation? Rob Meijers EMBL Hamburg Garbage in? The right stuff Molecular weight Oligomerization state Monodispersity
Sample preparation and characterization around SAXS Experimental verification and validation? Rob Meijers EMBL Hamburg Garbage in? The right stuff Molecular weight Oligomerization state Monodispersity
ProPac WCX-10 Columns
 ProPac WCX-10 Columns Guidance for column use Tips to maximize column lifetime ProPac WCX-10 Column Tips and Tricks This guide provides essential information and invaluable guidelines for mobile phases,
ProPac WCX-10 Columns Guidance for column use Tips to maximize column lifetime ProPac WCX-10 Column Tips and Tricks This guide provides essential information and invaluable guidelines for mobile phases,
Experimental Design and Data Collection
 A. Checks to run to make sure the instrument is in good working condition B. Decide which experiment type is most appropriate, equilibrium or velocity C. Buffer considerations D. Speed selection and length
A. Checks to run to make sure the instrument is in good working condition B. Decide which experiment type is most appropriate, equilibrium or velocity C. Buffer considerations D. Speed selection and length
Supporting Information
 Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2013. Supporting Information for Adv. Mater., DOI: 10.1002/adma.201301472 Reconfigurable Infrared Camouflage Coatings from a Cephalopod
Copyright WILEY-VCH Verlag GmbH & Co. KGaA, 69469 Weinheim, Germany, 2013. Supporting Information for Adv. Mater., DOI: 10.1002/adma.201301472 Reconfigurable Infrared Camouflage Coatings from a Cephalopod
Introduction to biological small angle scattering
 Introduction to biological small angle scattering Frank Gabel (IBS/ILL) EMBO Practical Course (May 6th 013) F. Gabel (May 6th 013) EMBO Practical Course Length-scales and tools in structural biology small
Introduction to biological small angle scattering Frank Gabel (IBS/ILL) EMBO Practical Course (May 6th 013) F. Gabel (May 6th 013) EMBO Practical Course Length-scales and tools in structural biology small
Tips & Tricks GPC/SEC: Protein Analysis with Size-Exclusion Chromatography
 Tips & Tricks GPC/SEC: Protein Analysis with Size-Exclusion Chromatography Daniela Held and Thorsten Hofe, PSS Polymer Standards Service GmbH, Mainz, Germany Gel permeation chromatography/size-exclusion
Tips & Tricks GPC/SEC: Protein Analysis with Size-Exclusion Chromatography Daniela Held and Thorsten Hofe, PSS Polymer Standards Service GmbH, Mainz, Germany Gel permeation chromatography/size-exclusion
Advantages of Agilent AdvanceBio SEC Columns for Biopharmaceutical Analysis
 Advantages of Agilent AdvanceBio SEC Columns for Biopharmaceutical Analysis Comparing Columns from Different Vendors to Improve Data Quality Technical Overview Introduction Size exclusion chromatography
Advantages of Agilent AdvanceBio SEC Columns for Biopharmaceutical Analysis Comparing Columns from Different Vendors to Improve Data Quality Technical Overview Introduction Size exclusion chromatography
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Isolation & Purification of Proteoglycans (PGs) and Glycosaminoglycans (GAGs) PEG Trainee Lecture July 23, 2012
 Isolation & Purification of Proteoglycans (PGs) and Glycosaminoglycans (GAGs) PEG Trainee Lecture July 23, 2012 Most Common Extraction Procedure for PGs 4 M Guanidine-HCl Detergents such as 2% CHAPS or
Isolation & Purification of Proteoglycans (PGs) and Glycosaminoglycans (GAGs) PEG Trainee Lecture July 23, 2012 Most Common Extraction Procedure for PGs 4 M Guanidine-HCl Detergents such as 2% CHAPS or
Dr. Christoph Johann Wyatt Technology Europe GmbH Copyright Wyatt Technology Europe GmbH All Rights reserved 1
 Dr. Christoph Johann Wyatt Technology Europe GmbH 2010 Copyright Wyatt Technology Europe GmbH All Rights reserved 1 Introduction Overview The Nature of Scattered Light: Intensity of scattered light Angular
Dr. Christoph Johann Wyatt Technology Europe GmbH 2010 Copyright Wyatt Technology Europe GmbH All Rights reserved 1 Introduction Overview The Nature of Scattered Light: Intensity of scattered light Angular
SAXS Basics for BioSAXS. Robert P. Rambo Diamond Light Source B21
 SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
SAXS Basics for BioSAXS Robert P. Rambo Diamond Light Source B21 Scattering and Size SAXS 0.11 nm (1.1 Å) smaller than C-C bond 6000x range 690 nm (6900 Å) 3.5x larger than E.coli 10x larger than a ribosome
Degree of labeling (DOL)
 Degree of labeling (DOL) A written explanation for the determination of the degree of labeling when using NuLink reagents. The DOL is the average number of reagent molecules that have been covalently attached
Degree of labeling (DOL) A written explanation for the determination of the degree of labeling when using NuLink reagents. The DOL is the average number of reagent molecules that have been covalently attached
Quantification Strategies Using the High Sensitivity Protein 250 Assay for the Agilent 2100 Bioanalyzer. Technical Note. Abstract
 Quantification Strategies Using the High Sensitivity Protein 25 Assay for the Agilent 21 Bioanalyzer Technical Note.. 1. 1. 1..1.1 1. 1. 1... Abstract The Agilent High Sensitivity Protein 25 assay for
Quantification Strategies Using the High Sensitivity Protein 25 Assay for the Agilent 21 Bioanalyzer Technical Note.. 1. 1. 1..1.1 1. 1. 1... Abstract The Agilent High Sensitivity Protein 25 assay for
Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System
 Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System Keywords: Hydrodynamic radius, diffusion coefficient, sizing, Taylor dispersion analysis, protein, antibodies,
Precision and accuracy of protein size determination using the ActiPix TDA200 Nano-Sizing System Keywords: Hydrodynamic radius, diffusion coefficient, sizing, Taylor dispersion analysis, protein, antibodies,
Polymer analysis by GPC-SEC. Technical Note. Introduction
 Polymer analysis by GPC-SEC Technical Note Introduction Gel Permeation Chromatography (GPC), also referred to as Size Exclusion Chromatography (SEC) is a mode of liquid chromatography in which the components
Polymer analysis by GPC-SEC Technical Note Introduction Gel Permeation Chromatography (GPC), also referred to as Size Exclusion Chromatography (SEC) is a mode of liquid chromatography in which the components
Liquid storage: Holds the solvent which is going to act as the mobile phase. Pump: Pushes the solvent through to the column at high pressure.
 High performance liquid chromatography (HPLC) is a much more sensitive and useful technique than paper and thin layer chromatography. The instrument used for HPLC is called a high performance liquid chromatograph.
High performance liquid chromatography (HPLC) is a much more sensitive and useful technique than paper and thin layer chromatography. The instrument used for HPLC is called a high performance liquid chromatograph.
Nature Structural and Molecular Biology: doi: /nsmb Supplementary Figure 1. Definition and assessment of ciap1 constructs.
 Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Supplementary Figure 1 Definition and assessment of ciap1 constructs. (a) ciap1 constructs used in this study are shown as primary structure schematics with domains colored as in the main text. Mutations
Electronic Supplementary Information (ESI) Synthesis of gold nanoparticles in a biocompatible fluid from sputtering deposition onto castor oil
 Electronic Supplementary Information (ESI) Synthesis of gold nanoparticles in a biocompatible fluid from sputtering deposition onto castor oil Heberton Wender, a Luciane F. de Oliveira, b Adriano F. Feil,
Electronic Supplementary Information (ESI) Synthesis of gold nanoparticles in a biocompatible fluid from sputtering deposition onto castor oil Heberton Wender, a Luciane F. de Oliveira, b Adriano F. Feil,
Supplementary Figure 1 Pairing alignments, turns and extensions within the structure of the ribozyme-product complex. (a) The alignment of the G27
 Supplementary Figure 1 Pairing alignments, turns and extensions within the structure of the ribozyme-product complex. (a) The alignment of the G27 A40 non-canonical pair stacked over the A41 (G1-C26) three-base
Supplementary Figure 1 Pairing alignments, turns and extensions within the structure of the ribozyme-product complex. (a) The alignment of the G27 A40 non-canonical pair stacked over the A41 (G1-C26) three-base
Bis sulfone Reagents. Figure 1.
 Bis sulfone Reagents An intact IgG molecule has four accessible inter chain disulfide bonds that can be reduced to form eight free cysteine thiols, which can serve as sites for conjugation. The reaction
Bis sulfone Reagents An intact IgG molecule has four accessible inter chain disulfide bonds that can be reduced to form eight free cysteine thiols, which can serve as sites for conjugation. The reaction
Small-Angle X-ray Scattering (SAXS)/X-ray Absorption Near Edge Spectroscopy (XANES).
 S1 Small-Angle X-ray Scattering (SAXS)/X-ray Absorption Near Edge Spectroscopy (XANES). The combined SAXS/XANES measurements were carried out at the µspot beamline at BESSY II (Berlin, Germany). The beamline
S1 Small-Angle X-ray Scattering (SAXS)/X-ray Absorption Near Edge Spectroscopy (XANES). The combined SAXS/XANES measurements were carried out at the µspot beamline at BESSY II (Berlin, Germany). The beamline
Instantaneous and Quantitative Functionalization of Gold Nanoparticles with Thiolated DNA Using a ph-assisted and Surfactant-Free Route
 Supporting Information Instantaneous and Quantitative Functionalization of Gold Nanoparticles with Thiolated DNA Using a ph-assisted and Surfactant-Free Route Xu Zhang,, Mark R. Servos and Juewen Liu *
Supporting Information Instantaneous and Quantitative Functionalization of Gold Nanoparticles with Thiolated DNA Using a ph-assisted and Surfactant-Free Route Xu Zhang,, Mark R. Servos and Juewen Liu *
LC Technical Information
 LC Technical Information Method Transfer to Accucore.6 μm Columns Containing solid core particles, which are engineered to a diameter of.6μm and a very narrow particle size distribution; Accucore HPLC
LC Technical Information Method Transfer to Accucore.6 μm Columns Containing solid core particles, which are engineered to a diameter of.6μm and a very narrow particle size distribution; Accucore HPLC
Static and dynamic light scattering. Cy Jeffries EMBL Hamburg
 Static and dynamic light scattering. Cy Jeffries EMBL Hamburg Introduction. The electromagnetic spectrum. visible 10-16 10-10 10-8 10-4 10-2 10 4 (l m) g-rays X-rays UV IR micro wave Long radio waves 400
Static and dynamic light scattering. Cy Jeffries EMBL Hamburg Introduction. The electromagnetic spectrum. visible 10-16 10-10 10-8 10-4 10-2 10 4 (l m) g-rays X-rays UV IR micro wave Long radio waves 400
Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS)
 nature methods Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS) Greg L Hura, Angeli L Menon, Michal Hammel, Robert P Rambo, Farris L Poole II, Susan E Tsutakawa,
nature methods Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS) Greg L Hura, Angeli L Menon, Michal Hammel, Robert P Rambo, Farris L Poole II, Susan E Tsutakawa,
MEASURING PROTEIN AGGREGATION WITH THE VISCOTEK SEC-MALS 20
 MEASURING PROTEIN AGGREGATION WITH THE VISCOTEK SEC-MALS 20 Introduction Protein aggregation is recognised as a major issue in the biopharmaceutical industry. Proteins have a tendency to aggregate over
MEASURING PROTEIN AGGREGATION WITH THE VISCOTEK SEC-MALS 20 Introduction Protein aggregation is recognised as a major issue in the biopharmaceutical industry. Proteins have a tendency to aggregate over
Supplementary figure 1. Comparison of unbound ogm-csf and ogm-csf as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine
 Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
Supplementary figure 1. Comparison of unbound and as captured in the GIF:GM-CSF complex. Alignment of two copies of unbound ovine GM-CSF (slate) with bound GM-CSF in the GIF:GM-CSF complex (GIF: green,
SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW
 SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW Paula Hong, John Van Antwerp, and Patricia McConville Waters Corporation, Milford, MA, USA Historically UV detection has been
SEAMLESS INTEGRATION OF MASS DETECTION INTO THE UV CHROMATOGRAPHIC WORKFLOW Paula Hong, John Van Antwerp, and Patricia McConville Waters Corporation, Milford, MA, USA Historically UV detection has been
Size Exclusion Chromatography in the Presence of an Anionic Surfactant
 Size Exclusion Chromatography in the Presence of an Anionic Surfactant Intact Protein Profiling Application Note Author Andy Coffey Agilent Technologies, Inc Abstract Sodium dodecyl sulfate (SDS, or SLS)
Size Exclusion Chromatography in the Presence of an Anionic Surfactant Intact Protein Profiling Application Note Author Andy Coffey Agilent Technologies, Inc Abstract Sodium dodecyl sulfate (SDS, or SLS)
Crystal Structure of Fibroblast Growth Factor 9 (FGF9) Reveals Regions. Implicated in Dimerization and Autoinhibition
 JBC Papers in Press. Published on November 1, 2000 as Manuscript M006502200 Crystal Structure of Fibroblast Growth Factor 9 (FGF9) Reveals Regions Implicated in Dimerization and Autoinhibition 1 Copyright
JBC Papers in Press. Published on November 1, 2000 as Manuscript M006502200 Crystal Structure of Fibroblast Growth Factor 9 (FGF9) Reveals Regions Implicated in Dimerization and Autoinhibition 1 Copyright
Fajun Zhang, Roland Roth, Marcell Wolf, Felix Roosen-Runge, Maximilian W. A. Skoda, Robert M. J. Jacobs, Michael Stzuckie and Frank Schreiber
 Soft Matter, 2012, 8, 1313 Fajun Zhang, Roland Roth, Marcell Wolf, Felix Roosen-Runge, Maximilian W. A. Skoda, Robert M. J. Jacobs, Michael Stzuckie and Frank Schreiber Universität Tübingen, Institut für
Soft Matter, 2012, 8, 1313 Fajun Zhang, Roland Roth, Marcell Wolf, Felix Roosen-Runge, Maximilian W. A. Skoda, Robert M. J. Jacobs, Michael Stzuckie and Frank Schreiber Universität Tübingen, Institut für
TA Instruments Application Note
 Q ( µj) TA Instruments Application Note Isothermal Titration Calorimetry (ITC) with Reduced Cell Volumes: A Comparison of the TA Instruments Nano ITC-Low Volume with the GE Healthcare Auto-iTC 200. Colette
Q ( µj) TA Instruments Application Note Isothermal Titration Calorimetry (ITC) with Reduced Cell Volumes: A Comparison of the TA Instruments Nano ITC-Low Volume with the GE Healthcare Auto-iTC 200. Colette
Protein Structure Analysis and Verification. Course S Basics for Biosystems of the Cell exercise work. Maija Nevala, BIO, 67485U 16.1.
 Protein Structure Analysis and Verification Course S-114.2500 Basics for Biosystems of the Cell exercise work Maija Nevala, BIO, 67485U 16.1.2008 1. Preface When faced with an unknown protein, scientists
Protein Structure Analysis and Verification Course S-114.2500 Basics for Biosystems of the Cell exercise work Maija Nevala, BIO, 67485U 16.1.2008 1. Preface When faced with an unknown protein, scientists
Comparison of Polymer Separation by Size Exclusion Chromatography and Asymmetric Flow Field Flow Fractionation
 Comparison of Polymer Separation by Size Exclusion Chromatography and Asymmetric Flow Field Flow Fractionation Stepan Podzimek, 1 Christoph Johann 2 1 SYNPO / University of Pardubice, Czech Republic, stepan.podzimek@synpo.cz
Comparison of Polymer Separation by Size Exclusion Chromatography and Asymmetric Flow Field Flow Fractionation Stepan Podzimek, 1 Christoph Johann 2 1 SYNPO / University of Pardubice, Czech Republic, stepan.podzimek@synpo.cz
Supplemental Information. Structural and Mechanistic Paradigm. of Leptin Receptor Activation Revealed
 Structure, Volume 22 Supplemental Information Structural and Mechanistic Paradigm of Leptin Receptor Activation Revealed by Complexes with Wild-Type and Antagonist Leptins Kedar Moharana, Lennart Zabeau,
Structure, Volume 22 Supplemental Information Structural and Mechanistic Paradigm of Leptin Receptor Activation Revealed by Complexes with Wild-Type and Antagonist Leptins Kedar Moharana, Lennart Zabeau,
Non-Interfering Protein Assay Kit Cat. No
 Visit our interactive pathways at /pathways User Protocol 488250 Rev. 5 January 2006 RFH Page 1 of 1 Non-Interfering Protein Assay Kit Cat. No. 488250 Note that this user protocol is not lot-specific and
Visit our interactive pathways at /pathways User Protocol 488250 Rev. 5 January 2006 RFH Page 1 of 1 Non-Interfering Protein Assay Kit Cat. No. 488250 Note that this user protocol is not lot-specific and
Liquid Chromatography
 Liquid Chromatography 1. Introduction and Column Packing Material 2. Retention Mechanisms in Liquid Chromatography 3. Method Development 4. Column Preparation 5. General Instrumental aspects 6. Detectors
Liquid Chromatography 1. Introduction and Column Packing Material 2. Retention Mechanisms in Liquid Chromatography 3. Method Development 4. Column Preparation 5. General Instrumental aspects 6. Detectors
Soil Cation Analysis Using High-Performance Capillary Zone Electrophoresis Last Modified: October 20, 2006
 Soil Cation Analysis Using High-Performance Capillary Zone Electrophoresis Last Modified: October 20, 2006 Introduction: Capillary electrophoresis (CE) is a relatively new, but rapidly growing separation
Soil Cation Analysis Using High-Performance Capillary Zone Electrophoresis Last Modified: October 20, 2006 Introduction: Capillary electrophoresis (CE) is a relatively new, but rapidly growing separation
Hints for Strong Ion Exchange Resins
 Hints for Strong Ion Exchange Resins Chromatography Application Note AN98 Abstract Ion exchange columns are a powerful means of isolating and purifying compounds, but their use is limited due to lack of
Hints for Strong Ion Exchange Resins Chromatography Application Note AN98 Abstract Ion exchange columns are a powerful means of isolating and purifying compounds, but their use is limited due to lack of
HPLC Columns. HILICpak VT-50 2D MANUAL. Shodex HPLC Columns Europe, Middle East, Africa, Russia
 HPLC Columns MANUAL HILICpak VT-50 2D Shodex HPLC Columns Europe, Middle East, Africa, Russia For technical support please use contact details shown below: SHOWA DENKO EUROPE GmbH Shodex Business Konrad-Zuse-Platz
HPLC Columns MANUAL HILICpak VT-50 2D Shodex HPLC Columns Europe, Middle East, Africa, Russia For technical support please use contact details shown below: SHOWA DENKO EUROPE GmbH Shodex Business Konrad-Zuse-Platz
Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation, and crystallization
 HERCULES Specialized Course: Non-atomic resolution scattering in biology and soft matter Grenoble, September 14-19, 2014 Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation,
HERCULES Specialized Course: Non-atomic resolution scattering in biology and soft matter Grenoble, September 14-19, 2014 Proteins in solution: charge-tuning, cluster formation, liquid-liquid phase separation,
Supporting Online Material. On-Chip Dielectrophoretic Co-Assembly of Live Cells and. Particles into Responsive Biomaterials
 Supporting Online Material On-Chip Dielectrophoretic Co-Assembly of Live Cells and Particles into esponsive Biomaterials Shalini Gupta, ossitza G. Alargova, Peter K. Kilpatrick and Orlin D. Velev* Description
Supporting Online Material On-Chip Dielectrophoretic Co-Assembly of Live Cells and Particles into esponsive Biomaterials Shalini Gupta, ossitza G. Alargova, Peter K. Kilpatrick and Orlin D. Velev* Description
Analytical determination of testosterone in human serum using an Agilent Ultivo Triple Quadrupole LC/MS
 Application Note Clinical Research Analytical determination of testosterone in human serum using an Agilent Ultivo Triple Quadrupole LC/MS Authors Yanan Yang 1, Victor Mandragon 2, and Peter Stone 1 1
Application Note Clinical Research Analytical determination of testosterone in human serum using an Agilent Ultivo Triple Quadrupole LC/MS Authors Yanan Yang 1, Victor Mandragon 2, and Peter Stone 1 1
Assay Transfer from HPLC to UPLC for Higher Analysis Throughput
 MAY 2005 SEPARATION SCIENCE REDEFINED 31 Assay Transfer from HPLC to UPLC for Higher Analysis Throughput A typical HPLC assay was transferred and optimized for a Waters ACQUITY UPLC system to achieve both
MAY 2005 SEPARATION SCIENCE REDEFINED 31 Assay Transfer from HPLC to UPLC for Higher Analysis Throughput A typical HPLC assay was transferred and optimized for a Waters ACQUITY UPLC system to achieve both
The Theory of HPLC. Quantitative and Qualitative HPLC
 The Theory of HPLC Quantitative and Qualitative HPLC i Wherever you see this symbol, it is important to access the on-line course as there is interactive material that cannot be fully shown in this reference
The Theory of HPLC Quantitative and Qualitative HPLC i Wherever you see this symbol, it is important to access the on-line course as there is interactive material that cannot be fully shown in this reference
ANALYSIS OF LOW DENSITY PARTICLES USING DIFFERENTIAL CENTRIFUGAL SEDIMENTATION
 ANALYSIS OF LOW DENSITY PARTICLES USING DIFFERENTIAL CENTRIFUGAL SEDIMENTATION Conventional Centrifugal Methods Centrifugal sedimentation of particles suspended in a fluid is a well known method (1, 2)
ANALYSIS OF LOW DENSITY PARTICLES USING DIFFERENTIAL CENTRIFUGAL SEDIMENTATION Conventional Centrifugal Methods Centrifugal sedimentation of particles suspended in a fluid is a well known method (1, 2)
Prep 150 LC System: Considerations for Analytical to Preparative Scaling
 Andrew Aubin and Jo-Ann Jablonski Waters Corporation, Milford, MA, USA APPLICATION BENEFITS The Prep 150 LC System is an affordable, highly reliable system for preparative chromatography and is suitable
Andrew Aubin and Jo-Ann Jablonski Waters Corporation, Milford, MA, USA APPLICATION BENEFITS The Prep 150 LC System is an affordable, highly reliable system for preparative chromatography and is suitable
Quantifying the Affinities and Kinetics of Protein Interactions Using Silicon Nanowire Biosensors
 SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.82 Quantifying the Affinities and Kinetics of Protein Interactions Using Silicon Nanowire Biosensors Xuexin Duan, Yue Li, Nitin K. Rajan, David A. Routenberg,
SUPPLEMENTARY INFORMATION DOI: 10.1038/NNANO.2012.82 Quantifying the Affinities and Kinetics of Protein Interactions Using Silicon Nanowire Biosensors Xuexin Duan, Yue Li, Nitin K. Rajan, David A. Routenberg,
Protein assay of SpectroArt 200
 Technical Bulletin 14 SpectroArt 200 12/01/2008 Protein assay of SpectroArt 200 MATERIAL BSA: Albumin, bovine serum (Sigma) PBS: BupH TM Phosphate Buffered Saline packs (PIERCE) Bradford assay: Bio-Rad
Technical Bulletin 14 SpectroArt 200 12/01/2008 Protein assay of SpectroArt 200 MATERIAL BSA: Albumin, bovine serum (Sigma) PBS: BupH TM Phosphate Buffered Saline packs (PIERCE) Bradford assay: Bio-Rad
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R)
 Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
Supplementary Figure 1 Crystal contacts in COP apo structure (PDB code 3S0R) Shown in cyan and green are two adjacent tetramers from the crystallographic lattice of COP, forming the only unique inter-tetramer
specified quantity of a solvent at a given temperature. To deconvolute the value from the
 S.1 Calculations of Dilution Enthalpy and Enthalpic Interaction Coefficients. When a solute is dissolved in a solvent a solution is formed. During dissolution of a solute in any solvent, heat is either
S.1 Calculations of Dilution Enthalpy and Enthalpic Interaction Coefficients. When a solute is dissolved in a solvent a solution is formed. During dissolution of a solute in any solvent, heat is either
Zetasizer Nano ZSP: A Perfect Tool For Life Science Applications
 Zetasizer Nano ZSP: A Perfect Tool For Life Science Applications Dr Mike Kaszuba Technical Support Manager E-mail: michael.kaszuba@malvern.com Contents Zetasizer Nano ZSP Software Enhancements Protein
Zetasizer Nano ZSP: A Perfect Tool For Life Science Applications Dr Mike Kaszuba Technical Support Manager E-mail: michael.kaszuba@malvern.com Contents Zetasizer Nano ZSP Software Enhancements Protein
Biochemistry. Biochemical Techniques HPLC
 Description of Module Subject Name Paper Name 12 Module Name/Title 13 1. Objectives 1.1. To understand the basic concept and principle of 1.2. To understand the components and techniques of 1.3. To know
Description of Module Subject Name Paper Name 12 Module Name/Title 13 1. Objectives 1.1. To understand the basic concept and principle of 1.2. To understand the components and techniques of 1.3. To know
Supporting Information
 Supporting Information Structural Analysis of the Binding of Type I, I 1/2, and II Inhibitors to Eph Tyrosine Kinases Jing Dong, *1 Hongtao Zhao, 1 Ting Zhou, 1 Dimitrios Spiliotopoulos, 1 Chitra Rajendran,
Supporting Information Structural Analysis of the Binding of Type I, I 1/2, and II Inhibitors to Eph Tyrosine Kinases Jing Dong, *1 Hongtao Zhao, 1 Ting Zhou, 1 Dimitrios Spiliotopoulos, 1 Chitra Rajendran,
Enzyme Experiments Using THz Light at the SRS and J-Lab
 Enzyme Experiments Using THz Light at the SRS and J-Lab Hannan Fersi Manchester Interdisciplinary Biocentre The University of Manchester May 16, 2007 Feasibility Study Can high power THz radiation be used
Enzyme Experiments Using THz Light at the SRS and J-Lab Hannan Fersi Manchester Interdisciplinary Biocentre The University of Manchester May 16, 2007 Feasibility Study Can high power THz radiation be used
Supplemental Information for:
 Supplemental Information for: New Insight into the Structure of RNA in Red clover necrotic mosaic virus and the Role of Divalent Cations Revealed by Small-Angle Neutron Scattering Stanton L. Martin a,
Supplemental Information for: New Insight into the Structure of RNA in Red clover necrotic mosaic virus and the Role of Divalent Cations Revealed by Small-Angle Neutron Scattering Stanton L. Martin a,
Supporting Information
 Supporting Information Wiley-VCH 2008 69451 Weinheim, Germany Electronic Supporting Information A Highly Selective Luminescence Switch-on Probe to Histidine/Histidine-rich proteins and Its Application
Supporting Information Wiley-VCH 2008 69451 Weinheim, Germany Electronic Supporting Information A Highly Selective Luminescence Switch-on Probe to Histidine/Histidine-rich proteins and Its Application
Supporting Information
 Supporting Information Role of Molecular Weight Distribution on Charge Transport in Semiconducting Polymers Scott Himmelberger, Koen Vandewal, Zhuping Fei, Martin Heeney, Alberto Salleo* Mobility Model
Supporting Information Role of Molecular Weight Distribution on Charge Transport in Semiconducting Polymers Scott Himmelberger, Koen Vandewal, Zhuping Fei, Martin Heeney, Alberto Salleo* Mobility Model
Anion Analysis of Ophthalmic Irrigation Solutions with the Shimadzu Prominence IC System Craig Young, PhD, Sr. Tech Support Specialist
 High Performance Liquid Chromatography SSI HPLC 5 Anion Analysis of Ophthalmic Irrigation Solutions with the Shimadzu Prominence IC System Craig Young, PhD, Sr. Tech Support Specialist Introduction Ophthalmic
High Performance Liquid Chromatography SSI HPLC 5 Anion Analysis of Ophthalmic Irrigation Solutions with the Shimadzu Prominence IC System Craig Young, PhD, Sr. Tech Support Specialist Introduction Ophthalmic
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi
 Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Small-Angle X-ray Scattering (SAXS) SPring-8/JASRI Naoto Yagi 1 Wikipedia Small-angle X-ray scattering (SAXS) is a small-angle scattering (SAS) technique where the elastic scattering of X-rays (wavelength
Experiment title: SAXS measurement of waterborne polymer/clay nanocomposites.
 Beamline: BM16 Shifts: 6 Experiment title: SAXS measurement of waterborne polymer/clay nanocomposites. Date of experiment: from: 02/05/07 to: 31/07/08 Local contact(s): Dr. Francois FAUTH Experiment number:
Beamline: BM16 Shifts: 6 Experiment title: SAXS measurement of waterborne polymer/clay nanocomposites. Date of experiment: from: 02/05/07 to: 31/07/08 Local contact(s): Dr. Francois FAUTH Experiment number:
Method Transfer between HPLC and UHPLC Instruments Equipment-related challenges and solutions
 Method Transfer between HPLC and UHPLC Instruments Equipment-related challenges and solutions Today, ultra-high-performance liquid chromatography (UHPLC) has taken a firm foothold in the analytical laboratory.
Method Transfer between HPLC and UHPLC Instruments Equipment-related challenges and solutions Today, ultra-high-performance liquid chromatography (UHPLC) has taken a firm foothold in the analytical laboratory.
PHYCOLINK CONJUGATE EVALUATIONS
 TechNote #TNPJ200 PHYCOLINK CONJUGATE EVALUATIONS INTRODUCTION PhycoLink Conjugation Kits provide a quick and simple method to attach fluorescent pigments to antibodies or other proteins of interest. When
TechNote #TNPJ200 PHYCOLINK CONJUGATE EVALUATIONS INTRODUCTION PhycoLink Conjugation Kits provide a quick and simple method to attach fluorescent pigments to antibodies or other proteins of interest. When
type GroEL-GroES complex. Crystals were grown in buffer D (100 mm HEPES, ph 7.5,
 Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Supplementary Material Supplementary Materials and Methods Structure Determination of SR1-GroES-ADP AlF x SR1-GroES-ADP AlF x was purified as described in Materials and Methods for the wild type GroEL-GroES
Chromatographic Analysis
 Chromatographic Analysis Distribution of Analytes between Phases An analyte is in equilibrium between the two phases [S 1 ] [S 2 ] (in phase 1) (in phase 2) AS [S2 ] K 2 A S [S1 ] 1 AS, A 1 S Activity
Chromatographic Analysis Distribution of Analytes between Phases An analyte is in equilibrium between the two phases [S 1 ] [S 2 ] (in phase 1) (in phase 2) AS [S2 ] K 2 A S [S1 ] 1 AS, A 1 S Activity
Performance evaluation of the Agilent 1290 Infinity 2D-LC Solution for comprehensive two-dimensional liquid chromatography
 Performance evaluation of the Agilent 1290 Infinity 2D-LC Solution for comprehensive two-dimensional liquid chromatography Technical Overview 2D-LC Conventional 1D-LC Abstract This Technical Overview presents
Performance evaluation of the Agilent 1290 Infinity 2D-LC Solution for comprehensive two-dimensional liquid chromatography Technical Overview 2D-LC Conventional 1D-LC Abstract This Technical Overview presents
Ion Chromatography. Anion Exchange. Chromatography Ion Exchange Theory. Dr. Shulamit Levin
 Ion Exchange Chromatography Chromatographic Process BA Mobile phase Stationary Phase A Shula Levin Bioforum B Distribution: K = C s/c m B shulal@zahav.net.il http://shulalc.co.il/ A Elution through the
Ion Exchange Chromatography Chromatographic Process BA Mobile phase Stationary Phase A Shula Levin Bioforum B Distribution: K = C s/c m B shulal@zahav.net.il http://shulalc.co.il/ A Elution through the
Application Note Pharmaceutical QA/QC. Agilent Application Solution. Authors. Abstract. Syed Salman Lateef Agilent Technologies, Inc.
 Agilent Application Solution Transfer of a USP method for tolazamide from normal phase HPLC to SFC using the Agilent 126 Infinity Hybrid SFC/UHPLC System Improving peak shape and providing wider UV selectivity
Agilent Application Solution Transfer of a USP method for tolazamide from normal phase HPLC to SFC using the Agilent 126 Infinity Hybrid SFC/UHPLC System Improving peak shape and providing wider UV selectivity
Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking
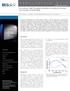 A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
A p p l i c a t i o n N o t e Low-volume, High Throughput Workflow for Analysis of Nucleic Acid Samples for Biobanking Peter J. Brescia, Jr., Chris Wilson, and Peter Banks, BioTek Instruments, Inc., Winooski,
Data sheet. UV-Tracer TM Biotin-Maleimide. For Labeling of Thiol-groups with UV-detectable Biotin CLK-B Description
 Cat. No. CLK-B105-10 CLK-B105-100 Amount 10 mg 100 mg For in vitro use only! Quality guaranteed for 12 months Store at -20 C 1. Description UV-Tracer TM Biotin Maleimide for biotinylation reactions of
Cat. No. CLK-B105-10 CLK-B105-100 Amount 10 mg 100 mg For in vitro use only! Quality guaranteed for 12 months Store at -20 C 1. Description UV-Tracer TM Biotin Maleimide for biotinylation reactions of
Luminescence transitions. Fluorescence spectroscopy
 Luminescence transitions Fluorescence spectroscopy Advantages: High sensitivity (single molecule detection!) Measuring increment in signal against a dark (zero) background Emission is proportional to excitation
Luminescence transitions Fluorescence spectroscopy Advantages: High sensitivity (single molecule detection!) Measuring increment in signal against a dark (zero) background Emission is proportional to excitation
High Brilliance SAXS on Synchrotrons
 High Brilliance SAXS on Synchrotrons Manfred Roessle Luebeck University of Applied Science Lab of X-ray Engineering X-ray generation Synchrotron Sources Shanghai Synchrotron Radiation Facility China Deutsches
High Brilliance SAXS on Synchrotrons Manfred Roessle Luebeck University of Applied Science Lab of X-ray Engineering X-ray generation Synchrotron Sources Shanghai Synchrotron Radiation Facility China Deutsches
Gel Permeation Chromatography (GPC) or Size Exclusion Chromatography (SEC)
 Gel Permeation Chromatography (GPC) or Size Exclusion Chromatography (SEC) Size Exclusion Chromatography (SEC) is a non-interaction based separation mechanism in which compounds are retained for different
Gel Permeation Chromatography (GPC) or Size Exclusion Chromatography (SEC) Size Exclusion Chromatography (SEC) is a non-interaction based separation mechanism in which compounds are retained for different
P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France
 Scattering of X-rays P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France SAXS measurement Sample SAXS measuring cell SAXS measurement Scattering experiment X-ray beam?? Detector SAXS measurement
Scattering of X-rays P. Vachette IBBMC (CNRS-Université Paris-Sud), Orsay, France SAXS measurement Sample SAXS measuring cell SAXS measurement Scattering experiment X-ray beam?? Detector SAXS measurement
HPLC Background Chem 250 F 2008 Page 1 of 24
 HPLC Background Chem 250 F 2008 Page 1 of 24 Outline: General and descriptive aspects of chromatographic retention and separation: phenomenological k, efficiency, selectivity. Quantitative description
HPLC Background Chem 250 F 2008 Page 1 of 24 Outline: General and descriptive aspects of chromatographic retention and separation: phenomenological k, efficiency, selectivity. Quantitative description
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition SUPPLEMENTARY INFORMATION
 SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
SOCS3 binds specific receptor JAK complexes to control cytokine signaling by direct kinase inhibition Nadia J. Kershaw 1,2, James M. Murphy 1,2, Nicholas P.D. Liau 1,2, Leila N. Varghese 1,2, Artem Laktyushin
Harris: Quantitative Chemical Analysis, Eight Edition CHAPTER 25: CHROMATOGRAPHIC METHODS AND CAPILLARY ELECTROPHORESIS
 Harris: Quantitative Chemical Analysis, Eight Edition CHAPTER 25: CHROMATOGRAPHIC METHODS AND CAPILLARY ELECTROPHORESIS CHAPTER 25: Opener Aa CHAPTER 25: Opener Ab CHAPTER 25: Opener B 25-1 Ion-Exchange
Harris: Quantitative Chemical Analysis, Eight Edition CHAPTER 25: CHROMATOGRAPHIC METHODS AND CAPILLARY ELECTROPHORESIS CHAPTER 25: Opener Aa CHAPTER 25: Opener Ab CHAPTER 25: Opener B 25-1 Ion-Exchange
Beers Law Instructor s Guide David T. Harvey
 Beers Law Instructor s Guide David T. Harvey Introduction This learning module provides an introduction to Beer s law that is designed for an introductory course in analytical chemistry. The module consists
Beers Law Instructor s Guide David T. Harvey Introduction This learning module provides an introduction to Beer s law that is designed for an introductory course in analytical chemistry. The module consists
Abstract: An minimalist overview of chromatography for the person who would conduct chromatographic experiments, but not design experiments.
 Chromatography Primer Abstract: An minimalist overview of chromatography for the person who would conduct chromatographic experiments, but not design experiments. At its heart, chromatography is a technique
Chromatography Primer Abstract: An minimalist overview of chromatography for the person who would conduct chromatographic experiments, but not design experiments. At its heart, chromatography is a technique
Amphiphilic diselenide-containing supramolecular polymers
 Electronic Supplementary Material (ESI) for Polymer Chemistry. This journal is The Royal Society of Chemistry 2014 Amphiphilic diselenide-containing supramolecular polymers Xinxin Tan, Liulin Yang, Zehuan
Electronic Supplementary Material (ESI) for Polymer Chemistry. This journal is The Royal Society of Chemistry 2014 Amphiphilic diselenide-containing supramolecular polymers Xinxin Tan, Liulin Yang, Zehuan
Supplemental Data SUPPLEMENTAL FIGURES
 Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
Supplemental Data CRYSTAL STRUCTURE OF THE MG.ADP-INHIBITED STATE OF THE YEAST F 1 C 10 ATP SYNTHASE Alain Dautant*, Jean Velours and Marie-France Giraud* From Université Bordeaux 2, CNRS; Institut de
