Protein Reorientation and Bound Water Molecules Measured by 1 H Magnetic Spin-Lattice Relaxation
|
|
|
- Everett Knight
- 6 years ago
- Views:
Transcription
1 558 Biophysical Journal Volume 84 January Protein Reorientation and Bound Water Molecules Measured by 1 H Magnetic Spin-Lattice Relaxation Alexandra Van-Quynh, Steven Willson, and Robert G. Bryant Chemistry Department, University of Virginia, Charlottesville, Virginia , USA ABSTRACT The water-proton spin-lattice relaxation rate constant, 1/T 1, was measured as a function of magnetic field strength for several dilute protein solutions. By separating the intermolecular contributions from the intramolecular contributions to the water-proton spin-lattice relaxation, the number of water molecules that bind to the protein for a time long compared with the rotational correlation time may be measured. We find a good correlation between the number of long-lived water molecules and the predictions based on available free volume in the proteins studied. The rotational correlation times of these proteins are larger than predicted by the Stokes-Einstein-Debye (SED) model for a sphere reorienting in a viscous liquid. The discrepancy between experiment and theory is usually attributed to hydration effects increasing the effective radius of the particle. However, the average lifetime of water molecules at the protein interface is far too short to justify such a picture. We suggest that surface roughness may be responsible for the retardation of rotational mobility and find that the SED model provides a reasonable representation of experiment if the radius assumed for the reorienting particle is the arithmetic mean of the crystallographic packing radius and the radius deduced from the effective surface area of the protein. INTRODUCTION Although water has been blamed for many mysterious aspects of macromolecular chemistry, several very different experiments have demonstrated that the vast majority of water at the protein surface has short residence times that are practically limited by the diffusion of water away from the surface sites; i.e., in the range of tens to hundreds of picoseconds. However, it is also now clear that there are a few specifically bound water molecules on most proteins that have lifetimes between 0.1 ms and 10 ns. These few water molecule sites affect the nuclear spin relaxation rates of the water protons through protein-water-proton dipolar couplings, which then affect other magnetic resonance observations, most notably, the contrast in nuclear magnetic images (Bryant, 1996a; Bryant, 1996b; Denisov and Halle, 1996; Halle et al., 1999). The same dipolar couplings may affect proton nuclear Overhauser effects in protein solutions and may attenuate signal intensities in high resolution experiments. The water-protein NMR literature includes complimentary contributions from proton, deuteron, and oxygen-17 spectroscopy. The interpretation of 1 H experiments has been somewhat troubled by the effects of labile proton chemical exchange and protein aggregation, but recent instrumental advances that provide both high sensitivity and high resolution (Wagner et al., 1999) permit experiments on dilute protein solutions and high isotope dilutions of protons so that it is possible to isolate the intramolecular contributions to the proton spin-lattice relaxation rate; i.e., the contribution from dipolar interaction between water protons. Submitted June 17, 2002, and accepted for publication September 19, Address reprint requests to Robert G. Bryant, Chemistry Dept., University of Virginia, McCormick Road, P. O. Box , Charlottesville, VA rgb4g@virginia.edu. Ó 2003 by the Biophysical Society /03/01/558/06 $2.00 (Kiihne and Bryant, 2000) Therefore, the proton experiments become practically equivalent in most ways to the more difficult oxygen or deuterium relaxation experiments except that the protons provide a considerable gain in sensitivity. Although the experiments are still technically demanding, the approach provides a means of exploring the number of water molecule binding sites for water molecule lifetimes in the range from 0.1 s to 10 ns. We report here the results of 1 H magnetic relaxation dispersion (MRD) experiments for ribonuclease A and cytochrome C, which yield the numbers of long-lived water molecules. We compare the number of bound water molecules with the number predicted based on free volume calculated using structural data obtained from x-ray diffraction, and then examine the dynamical consequences of these bound water molecules on the rotational and translational dynamics of the protein. Theoretical background Several studies have shown that the nuclear magnetic relaxation dispersion of 1 H, 2 H, and 17 O relaxation rate of bulk H 2 OorD 2 O is caused by a small number of water molecules bound to the protein for a time longer than the rotational correlation time of the protein (Koenig and Schillinger, 1969; Koenig et al., 1975; Halle et al., 1981; Koenig et al., 1993; Denisov and Halle, 1995; Denisov et al., 1995; Koenig, 1995; Bryant, 1996a; Denisov and Halle, 1996; Halle et al., 1999). Indeed, if these bound water molecules exchange with the bulk solvent with a rate that is fast compared to the spin-lattice relaxation rate of the bound water protons, the protein-bound-water-molecule sites act as relaxation sinks for the whole water population. In the case of dilute protein solutions where protein aggregation is minimized, the 1 H dispersion in the spin-lattice relaxation rate constant has a Lorentzian shape that permits accurate measurement of the rotational correlation time of the
2 Protein Reorientation and Bound Water 559 macromolecule, t rot (Kiihne and Bryant, 2000). The observed 1 H spin-lattice relaxation rate constant may be written (Denisov and Halle, 1998; Kiihne and Bryant, 2000): 1 ¼ P free þ P s P b þ þ H: (1) T 1obs T 1free T 1s T 1b þ t res 1/T 1free denotes the intramolecular H-H or H-D contributions of the free solvent H 2 O or HOD molecules and P free is essentially unity. The correlation times for reorientation of water in the bulk are short compared with the proton Larmor frequencies over the range studied, MHz; thus, 1/T 1free is independent of magnetic field strength over this range. The second term, 1/T 1s, comes from the hundreds of water molecules at the surface of the protein characterized by a probability P s. These molecules generally have a residence time much shorter than the rotational correlation time of the protein, and the relaxation dispersion for this contribution occurs above the largest magnetic field strength studied. Therefore, these short-lived surface interactions also add to the field independent relaxation rate. P b is the probability that water molecules are bound to the protein for times the order of or longer than the rotational correlation time, and may be expressed in terms of the number of bound water molecules per protein molecule, N b, and n prot =n H2 O, the ratio of the number of protein molecules to the total number of water molecules: n prot P b ¼ N b : (2) n H2 O We have assumed in Eq. 1 that a single spin-lattice relaxation time, T 1b, characterizes the bound environment. The mean residence time for water bound on the protein, t res, may be different for different sites on the protein, but its contribution is negligible if t rot < t res < T 1b, where t rot is the rotational correlation time of the protein. The last term in Eq. 1, H, represents the contribution from the exchange between protein ionizable groups and the protons of the solvent, which may be expressed as H ¼ + k P H k T 1k þ t ex;k ; (3) where the index k runs over all the protein-proton exchange sites occupied with a probability P H k and characterized by a relaxation time, T 1k, which is field dependent, and mean residence time, t ex;k. This contribution is generally a function of temperature and ph; it is often small because the mean residence times for many sites may be long relative to the relaxation times at the site, amide protons for example. The contribution of this term is independent of proton mole fraction and contributes to relaxation in the same way as the intermolecular contribution to 1/T 1b, which is discussed below. The bound water rate constant, 1/T 1b, may be decomposed as the sum of intermolecular and intramolecular dipolar contributions. In the H 2 O case, both the intramolecular (water proton-water proton) and intermolecular (water proton-protein proton) contributions are homonuclear and 1/T 1b may be written (Abragam, 1961): 1 ¼ intra½ T Jðv IÞþ4Jð2v I ÞŠþ inter½ Jðv IÞþ4Jð2v I ÞŠ; 1b H 2 O (4) where intra ¼ð2=5Þðg4 H h2 =rii 6 ÞIðI þ 1Þ characterizes the strength of the intramolecular dipole-dipole contribution of a proton pair separated by r II, which is 1.58 Å in the water molecule. g H is the magnetogyric ratio of the proton, h the Planck constant divided by 2p, and v I the proton Larmor frequency. inter ¼ Sð2=5Þðg4 H h2 =r 6 ij ÞIðI þ 1Þ is the intermolecular dipolar proton-proton contribution, which involves several proton-proton contacts characterized by different intermoment distances r ij. The minimum separation is determined by the van der Waals contact distance of 2.2 Å but a wide range of interproton distances may contribute. inter and intra may contain an order parameter, A 2 ð0 # A 2 # 1Þ, that may account for partial averaging of the dipolar interactions caused by high frequency motions of the bound water molecules. As previously noted (Denisov and Halle, 1998; Kiihne and Bryant, 2000), these relaxation dispersion experiments in the present field strength range do not provide a characterization of such high frequency motions, i.e., A 2. For simplicity we set the order parameter to 1 and make our calculations based on the assumption of rigidly bound water molecules. Jðv I Þ denotes the spectral density of magnetic fluctuations at v I. Assuming a single global correlation time for rotational diffusion, the spectral density function, Jðv I Þ has the Lorentzian form (Abragam, 1961) Jðv I Þ¼ t c 1 þðv I t c Þ 2 : (5) In the D 2 O solutions of proteins, the intermolecular contribution to the water 1 H spin-lattice relaxation rate comes from the residual HOD protons coupling to protein protons. The intramolecular contribution to proton relaxation caused by deuterons is heteronuclear and given by 1 ¼ B HD intra½ T Jðv I ÿ v S Þþ3Jðv I Þþ6Jðv I þ v S ÞŠ 1b HOD þ Jðv inter½ IÞþ4Jð2v I ÞŠ: (6) where B HD intra ¼ð2=15Þðg2 H g2 D h2 =ris 6 ÞSðS þ 1Þ, r IS is the proton-deuteron distance and S ¼ 1 for deuterium. In Eq. 4 6 we have neglected the intermolecular interaction between water protons and protein-deuterons because g D is small compared to g H and the protein-deuteron is relatively rare. Both in the H 2 O and D 2 O cases, the strength of the intermolecular contribution to the relaxation, inter, involves two terms that may be written inter ¼ + H i i þ + H k k ; (7)
3 560 Van-Quynh et al. where i represents the dipolar interaction between a nonexchangeable protein proton, H i, and a water proton; the sum runs over all the nonexchangeable protein protons. represents the strength of the dipolar interaction between a labile protein proton and a protein-bound water proton, which decreases linearly with the proton mole fraction, x H. To estimate the contribution of the second term of Eq. 7, we have used the coordinates based on the crystal structure of ribonuclease A (Wlodawer et al., 1988) to compute the ratio of the dipolar interaction between a fixed water proton and all the nonexchangeable protein protons to that of the same fixed water proton and all the protein protons, labile or not. This calculation demonstrates that ;10% of the coupling derives from the labile protein protons that are displaced in D 2 O solutions. In the following we neglect the intermolecular contribution coming from the labile protein-protons and discuss the error introduced by this term in the determination of the number of long-lived bound water molecules later. At low values of the Larmor frequency, Eqs. 4 6 reduce to k ðdr 1 Þ ¼ 1 H2 O T 1obs ÿ 1 H 2 O T 10 ðdr 1 Þ HOD ¼ 1 ÿ 1 ¼5P b t cd2 ð O T 1obs HOD T 10 HOD ¼ 5P b t ch2 ð OÞ þ intra BHH inter H 2 O (8) ; Þ 2B HD þ intra BHH inter where t cðh2 OÞ and t cðd2 OÞ are the rotational correlation times of the protein in H 2 O and D 2 O solutions respectively, which are different because the viscosities differ by ;20%. For both solvents, the intramolecular contribution of the bound water molecules involves a single interspin distance of 1.58 Å; then intra and BHD intra differ by the factor a a ¼ BHH intra ¼ 9 2 g H : (10) B HD intra 8 g D We then have the coupled equations ðdr 1 Þ ¼ 5P H2 O bt cðh2 OÞ (11) ðdr 1 Þ HOD ¼ 5P b t cðd2 OÞ 2 þ intra BHH inter (9) : (12) a BHH þ intra BHH inter Þ HOD, the ratio of the difference Let r ¼ ðdr 1 Þ H2 O = ð DR 1 between the high and low field relaxation rate constants in the two solvents, and c ¼ t cðh2 OÞ=t cðd2 OÞ the ratio of the rotational correlation time in the two solvents. Combining Eqs. 8 12, and solving for P b yields ð P b ¼ DR 1Þ H2 O 5t cðh2 OÞ intra aðc ÿ rþ rð2 ÿ aþ : (13) The numerical value of intra is calculated using 1.58 Å for the interproton distance in the H 2 O molecule and is equal to s ÿ2. Knowing all the other experimentally determined parameters, the number of long-lived bound water molecules is computed using Eq. 2. Aside from measurement noise, there are two sources of error in the calculation of the number of bound water molecules. The first comes from the intermolecular dipoledipole contribution of the labile protein protons, which may cause an overestimate of N b by 10% at maximum. The second is caused by the uncertainty in the order parameter, A 2, set to 1 in the above calculation. The quantity that we compute rigorously from our measurements is N b A 2. If A < 1, the number of bound water molecules, N b, increases as 1=A 2. The reduction of A is caused by restricted high frequency local motions that would, in principle, cause a second relaxation dispersion at very high field strengths. However, if the local correlation time is short, in the range of tens of picoseconds, for example, the contribution to the relaxation rate would be ;1000 times smaller than that from lower frequency motions. Thus, the primary effect of local motion of a bound water molecule in the binding site is reduction of the low field relaxation rate by the factor, A 2. EXPERIMENTAL Bovine pancreas ribonuclease A (R5555), bovine heart cytochrome C (C3131), bovine pancreas a-chymotrypsin (C4129), pepsin (6887), and thermolysin (protease type X P1512) were purchased from Sigma (St Louis, MO) as lyophilized powders. H 2 O and D 2 O solutions were made by dissolving the lyophilized proteins in dionized water and deuterium oxide (D, 99.9% Cambridge Isotope Laboratories, Andover, MA) respectively. For a-chymotrypsin and Thermolysin the ionic strength was maintained using 100 mm potassium chloride. Before NMR measurements, the H 2 O and D 2 O protein solutions were deoxygenated with a flowing nitrogen stream for one hour, to remove the paramagnetic contribution of the dissolved O 2 to the proton relaxation rate. Deoxygenated samples were sealed in a 5 mm o.d. glass sample tube utilizing a Delrin filler plug and a silicone rubber plug compressed between two threaded components similar to the design reported previously (Wagner et al., 1999). The glass tube is far superior to the Delrin tube because it does not leak oxygen as a function of time and is chemically much more inert. The MRD measurements were made in a dual magnet spectrometer described elsewhere (Wagner et al., 1999). The sample is allowed to achieve equilibrium in the high initial magnetic field, then pneumatically driven to a satellite magnet where it resides for variable relaxation period after which it is pneumatically driven back to the high field magnet where the magnetization is detected. The decay of the magnetization as a function of residence time in the satellite field is fit to an exponential time constant that is the relaxation time constant in the satellite field. The value of the satellite field strength is varied to map the relaxation
4 Protein Reorientation and Bound Water 561 dispersion over the range of proton Larmor frequencies from 0.01 to 70 MHz. The soak field with a proton Larmor frequency of 300 MHz provides the highest field relaxation rate constant. RESULTS Fig. 1 presents the relaxation dispersion curves obtained in H 2 O and D 2 O for ribonuclease A, 1.2 mm, and 0.60 mm respectively at ph ¼ 5.3. The two relaxation dispersions are Lorentzian. The fits according to Eq. 6 for the H 2 O case and Eq. 8 for the D 2 O case correspond to the solid lines and lead to a correlation time t cðh2 OÞ ¼ ns for the H 2 O solution and t cðd2 OÞ ¼ ns in the D 2 O case. We note that the viscosity in D 2 O is larger than that in H 2 O by 20%, which is only approximately consistent with the larger correlation time found in the D 2 O solution and is discussed below. The amplitudes of the dispersion in the relaxation rate constant used for the calculation of the number of bound water molecules, N b, are DðR 1 Þ H2 O ¼ 0:0216 sÿ1 and DðR 1 Þ D2 O ¼ 0:0219 sÿ1 when both data sets are normalized to 0.60 mm concentration of ribonuclease A. Knowing the value of t cðd2 OÞ and considering the ratio t cðh2 OÞ=t cðd2 OÞ, we find an intramolecular contribution to the relaxation of 30%. According to Eqs. 2 and 15 we find N b ¼ 3: The neglect of the dipolar contribution from labile protons causes this analysis to over estimate the number of bound water molecules by ;10%; thus, N b ¼ 3:5 6 1 which is not significantly different. Three internal water molecules have been reported based on x-ray diffraction data (Denisov and Halle, 1998). Rashin and coworkers report that there is space in the protein for internal water molecules based on calculations of free volume deduced from packing in the reported crystal structure. The value of the rotational correlation time for the water protons associated with ribonuclease A is short compared with expectations based on molecular volume and other measures of the protein reorientation time (Denisov and Halle, 1998). As pointed out by Denisov and Halle, the origin of this apparent discrepancy may derive from the interference between the rotational motion and the exchange of the water from the ribonuclease A binding environments. The water sites for long-lived water molecules on ribonuclease A are on surface pockets or crevasses not buried deeply inside the folded structure. The 17 O relaxation dispersion data agree with the proton relaxation dispersion data and imply that the rotational correlation time and the exchange times are of nearly the same size. Because the exchange event is uncorrelated with rotational diffusion, we may write the effective correlation time as 1=t c ¼ 1=t rot þ 1=t res : (14) If we assume that the rotational correlation is 6.6 ns as reported by Denisov and Halle based on deuterium relaxation data, which is also in agreement with the FIGURE 1 (Top) The 1 H spin-lattice relaxation rate constant for the residual protons, 1/T 1, are shown as a function of the magnetic field strength plotted as the proton Larmor frequency for a 0.60 mm solution of ribonuclease A in D 2 O at ambient laboratory temperature at a ph meter reading of 5.2. (Bottom) The 1 H spin-lattice relaxation rate constant, 1/T 1, shown as a function of the magnetic field strength plotted as the proton Larmor frequency for 1.2 mm solution of ribonuclease A in H 2 O at ph 5.2 and ambient laboratory temperature. calculation in Table 1 below based on molecular volume, then substitution of the measured rotational correlation time in Eq. 14 yields a value of 6.2 ns for the mean residence time of these bound water molecules. This value is in reasonable agreement with the value of 7.6 ns at 278C reported by Denisov and Halle (1998). One consequence of the short water-molecule residence times on ribonuclease A is that the deuterium and proton MRD inflections points are not simply related to the solution viscosity. In the deuterium case, the residual water-proton relaxation rate results from the sum of water-proton to protein-proton intermolecular contributions and from the exchange of labile protein protons with the water. The effective correlation time for the first contribution is reduced from
5 562 Van-Quynh et al. TABLE 1 Correlation time comparison Protein M w (kda) R V (nm) R S (nm) R av (nm) t th (ns) t c1 (ns) a-chymotrypsin Ribonuclease A Cytochrome C Thermolysin Pepsin BSA Comparison between the computed rotational correlation time, t th, obtained from the Stokes-Einstein-Debye relation and the experimental rotational correlation time, t c1, deduced from nuclear magnetic relaxation dispersion measurements. The radius used for the SED equation is the arithmetic mean of two measures of the effective protein radius. The first, R S, is the effective radius of the sphere that has the same surface area as the surface area of the protein deduced using the methods of Lee and Richards. The second, R V, is the radius deduced from the protein-protein contacts in the protein crystal. We list the molecular mass, M w, for reference. The experimental value t c1 for bovine serum albumin is extracted from previous work (Kiihne and Bryant, 2000). The comparisons are made for H 2 O solutions with h = 0.01 poise for T = 300 K. that for pure rotation because of the contribution of the short lifetime of the water on the protein according to Eq. 14. For the labile protein proton contribution, the effective correlation time is just the rotational correlation time of the protein. Because the weights of these contributions are different when the proton-proton intramolecular term is added for the bound H 2 O molecule, the observed MRD inflection frequencies are not simply proportional to the viscosities of the solutions. The same experiments and calculations were made for cytochrome C at 0.6 mm and ph ¼ 8. The intramolecular contribution to the relaxation is 64% and we find N b ¼ bound water molecules whereas the free volume analysis cited suggests that there should be two internal water molecules (Rashlin et al., 1986). Similar results were obtained for dilute solutions of thermolysin (0.040 mm, ph ¼ 6) and pepsin (0.28 mm, ph ¼ 7). The high field 1 H relaxation rate dispersions obtained in H 2 O solutions yield rotational correlation times t c1 ¼ 21:6 6 2:3 ns for thermolysin and t c1 ¼ 18:8 6 2:3 ns for pepsin. We discuss in the following section the magnitudes of these values in the context of prevalent ideas about protein volume and hydration. Fig. 2 shows the dispersion of the 1 H relaxation rate constant obtained for a H 2 O solution of a-chymotrypsin 0.3 mm, in 100 mm KCl and ph ¼ 6.9. Contrary to the ribonuclease A and cytochrome C results and those previously obtained in BSA (Kiihne and Bryant, 2000), the dispersion shape is not described by a single Lorentzian function. We have fitted this dispersion curve as the sum of two Lorentzian contributions to obtain the solid line; the correlation times are t c1 ¼ ns and t c2 ¼ ns. We attribute t c1 to the rotational correlation time of the monomeric protein. The value of t c2 is large and we attribute that to a low concentration of an impurity with large molecular mass. We note that the low field contributions to the relaxation rates from the particles of different size scale with the ratios of the rotational correlation times. Thus, the observed low field relaxation rate is consistent with only 2.6% of the bound water molecule sites derived from the larger molecule. Rotational correlation times and protein hydration As we show above, the MRD provides a direct report of the rotational motility of a protein as well as a quantitative measure of the number of long-lived water molecules that are associated with the protein. It is really an old (Koenig and Schillinger, 1969) but still remarkable result that the number of long-lived water molecules that hydrate proteins is a very small fraction of the total number of water molecules that are in contact with the protein. The vast majority of the surface contacts between the water and the protein are transient and characterized by short lifetimes, in the range of a few hundreds of picoseconds or shorter (Koenig, 1995; Bryant, 1996; Halle et al., 1999). Recognizing this fact, it is useful to compare the measured rotational correlation times with values predicted based on hydrodynamic theory. Proteins are large molecules that should be appropriate to Stokes- Einstein-Debye theory, which predicts that the rotational FIGURE 2 The 1 H spin-lattice relaxation rate constant, 1/T 1, as a function of the magnetic field strength plotted as the proton Larmor frequency for a 0.30 mm solution of a-chymotrypsin in H 2 O at ph 6.9 in 100 mm potassium chloride.
6 Protein Reorientation and Bound Water 563 correlation time, t th, is proportional to molecular volume and the viscosity, h, t th ¼ 4p hr 3 3 kt : (15) As commonly noted, the experimental values of t rot for proteins are larger than predicted by this relation and solvation has been blamed for the discrepancy (Yguerabide et al., 1970). A standard approach to understanding the failure of experiments to agree with theory is to ascribe a solvation layer of water molecules to the water-protein interface so that the effective size of the reorientational unit is larger than the volume of the protein presumed to be spherical. However, as the present and other measurements demonstrate, the mean residence time of water at the protein surface is compared with the rotational correlation time of the protein. This fact compromises the model that the hydration layer increases the effective radius of the protein and slows the rotational motion. An alterative hypothesis is that protein surface roughness retards the rotation of the protein (Garcia de la Torre and Bloomfield, 1981; Denisov and Halle, 1998). Indeed, the protein surface is not a smooth sphere when the different side chains are considered. One approach for measuring this roughness quantitatively is to compare radii computed from the effective surface area with that based on packing volume. The packing volume in a crystal provides a measure of the effective molecular volume from which a radius R V may be computed. Empirical relation between the molecular weight, M W, and the radius R V has been offered: R V ¼ 0:672M 1=3 W (Richards, 1977). This radius predicts a rotational correlation times that is smaller than that observed experimentally as shown in Table 1. An alternative way to consider the size of the protein is to examine the effective surface area S as considered by Lee and Richards using a probe molecule like water (Lee and Richards, 1971). This surface parea may be translated to an effective spherical radius, R S ¼ 4p=S ffiffiffiffiffiffiffiffiffiffiffi.for the proteins studied here, the value of R S is considerably larger than R V and if R S is used in Eq. 15, values of the rotational correlation time much larger than that observed experimentally are obtained. Although we have no fundamental or detailed theoretical justification for it, we find that a remarkably simple strategy provides an alternative approach to computing rotational correlation times for globular proteins. If we take the arithmetic mean between R S and R V to approximate the reorientational sphere in Eq. 15, reasonable agreement with the experiment is obtained as shown in Table 1. This procedure increases the effective reorientational radius by the factor 1.20 for the proteins listed. Although the concept of a protein as a smooth but enlarged sphere is difficult to defend, an alternative interpretation of this factor is that it represents the effective surface friction coefficient that is different from unity. The essence of the difference between this approach and assuming a bound hydration layer is that it springs from a reasonable physical picture of the macromolecule and avoids the unjustified assumption of ice-like water bound at the surface of the protein. REFERENCES Abragam, A The Principles of Nuclear Magnetism, Chapter VIII. The Clarendon Press, Oxford. Bryant, R. G. 1996a. The dynamics of water-protein interactions. Annu. Rev. Biophys. Biomol. Struct. 25: Bryant, R. G. 1996b. Magnetization transfer and cross relaxation in tissue. In Encyclopedia of Magnetic Resonance. E. D. M. Grant and R. K. Harris, Editors. John Wiley, New York Denisov, V. P., and B. Halle Protein hydration dynamics in aqueous solution: a comparison of bovine pancreatic trypsin inhibitor and ubiquitin by oxygen-17 spin relaxation dispersion. J. Mol. Biol. 245: Denisov, V. P., and B. Halle Protein hydration dynamics in aqueous solution. Faraday Discuss. 103: Denisov, V. P., and B. Halle Thermal denaturation of ribonuclease A characterized by water oxygen-17 and deuterium magnetic relaxation dispersion. Biochemistry. 37: Denisov, V. P., B. Halle, J. Peters, and H. D. Horlein Residence times of buried water molecules in bovine pancreatic trypsin inhibitor and its G36S mutant. Biochemistry. 34: Garcia de la Torre, J. G., and V. A. Bloomfield Hydrodynamic properties of complex, rigid, biological macromolecules: theory and applications. Q. Rev. Biophys. 14: Halle, B., T. Anderson, S. Forsen, and B. Lindman Protein hydration from water oxygen-17 magnetic relaxation. J. Am. Chem. Soc. 103: Halle, B., V. P. Denisov, and K. Venu Multinuclear relaxation dispersion studies of protein hydration. In Biological Magnetic Resonance, Vol. 17. L. J. Berliner and N. R. Krishna, editors. Klewer Academic/Plenum Press, New York Kiihne, S., and R. G. Bryant Protein-bound water molecule counting by resolution of (1)H spin- lattice relaxation mechanisms. Biophys. J. 78: Koenig, S. H Classes of hydration sites at protein-water interfaces: the source of contrast in magnetic resonance imaging. Biophys. J. 69: Koenig, S. H., R. D. I. Brown, and R. Ugolini A unified view of relaxation in protein solutions and tissue including hydration and magnetization transfer. Magn. Reson. Med. 29: Koenig, S. H., K. Hallenga, and M. Shporer Protein-water interactions studied by solvent proton, deuteron, and oxygen-17 magnetic relaxation. Proc. Natl. Acad. Sci. USA. 72: Koenig, S. H., and W. E. Schillinger Nuclear magnetic relaxation dispersion in protein solutions. I. Apotransferrin. J. Biol. Chem. 244: Lee, B., and F. M. Richards The interpretation of protein structures: estimation of static accessibility. J. Mol. Biol. 55: Rashlin, A. A., M. Iofin, and B. Honig Internal cavities and buried waters in globular proteins. Biochemistry. 25: Richards, F. M Areas, volumes, packing and protein Structure. Annu. Rev. Biophys. Bioeng. 6: Wagner, S., T. J. R. Dinesen, T. Rayner, and R. G. Bryant Magnetic relaxation dispersion measurements of solute spin probes using a dual magnet system. J. Magn. Reson. 140: Wlodawer, A., L. A. Svensson, L. Sjoln, and G. L. Gilleeland Structure of phosphate free ribonuclease A refined to 1.26 A. Biochemistry. 27: Yguerabide, J., H. Epstein, and L. Stryer Segmental flexibility in an antibody molecule. J. Mol. Biol. 51:
T 1, T 2, NOE (reminder)
 T 1, T 2, NOE (reminder) T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations of the system following perturbation
T 1, T 2, NOE (reminder) T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations of the system following perturbation
Asian Journal of Chemistry; Vol. 25, No. 4 (2013),
 Asian Journal of Chemistry; Vol. 25, No. 4 (213), 214-218 http://dx.doi.org/1.14233/ajchem.213.13346 Observation of Triplet Traces Obtained with Inversion Recovery Method in Both Residual Water- and H
Asian Journal of Chemistry; Vol. 25, No. 4 (213), 214-218 http://dx.doi.org/1.14233/ajchem.213.13346 Observation of Triplet Traces Obtained with Inversion Recovery Method in Both Residual Water- and H
Introduction to Relaxation Theory James Keeler
 EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
EUROMAR Zürich, 24 Introduction to Relaxation Theory James Keeler University of Cambridge Department of Chemistry What is relaxation? Why might it be interesting? relaxation is the process which drives
Magnetic Field Dependence of Proton Spin-Lattice Relaxation Times
 Magnetic Field Dependence of Proton Spin-Lattice Relaxation Times Jean-Pierre Korb and Robert G. Bryant 2 * Magnetic Resonance in Medicine 48:2 26 (2002) The magnetic field dependence of the water-proton
Magnetic Field Dependence of Proton Spin-Lattice Relaxation Times Jean-Pierre Korb and Robert G. Bryant 2 * Magnetic Resonance in Medicine 48:2 26 (2002) The magnetic field dependence of the water-proton
Spin Relaxation and NOEs BCMB/CHEM 8190
 Spin Relaxation and NOEs BCMB/CHEM 8190 T 1, T 2 (reminder), NOE T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations
Spin Relaxation and NOEs BCMB/CHEM 8190 T 1, T 2 (reminder), NOE T 1 is the time constant for longitudinal relaxation - the process of re-establishing the Boltzmann distribution of the energy level populations
Solid-state NMR and proteins : basic concepts (a pictorial introduction) Barth van Rossum,
 Solid-state NMR and proteins : basic concepts (a pictorial introduction) Barth van Rossum, 16.02.2009 Solid-state and solution NMR spectroscopy have many things in common Several concepts have been/will
Solid-state NMR and proteins : basic concepts (a pictorial introduction) Barth van Rossum, 16.02.2009 Solid-state and solution NMR spectroscopy have many things in common Several concepts have been/will
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Summary of 1D Experiment time domain data Fourier Transform (FT) frequency domain data or Transverse Relaxation Ensemble of Nuclear
K ex. Conformational equilibrium. equilibrium K B
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any yprocess in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Timescales of Protein Dynamics
 Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
Timescales of Protein Dynamics From Henzler-Wildman and Kern, Nature 2007 Dynamics from NMR Show spies Amide Nitrogen Spies Report On Conformational Dynamics Amide Hydrogen Transverse Relaxation Ensemble
File: {ELS_REV}Cavanagh X/Revises/Prelims.3d Creator: / Date/Time: /9:29pm Page: 1/26 PREFACE
 PREFACE The second edition of Protein NMR Spectroscopy: Principles and Practice reflects the continued rapid pace of development of biomolecular NMR spectroscopy since the original publication in 1996.
PREFACE The second edition of Protein NMR Spectroscopy: Principles and Practice reflects the continued rapid pace of development of biomolecular NMR spectroscopy since the original publication in 1996.
Simulation of the NMR Second Moment as a Function of Temperature in the Presence of Molecular Motion. Application to (CH 3
 Simulation of the NMR Second Moment as a Function of Temperature in the Presence of Molecular Motion. Application to (CH 3 ) 3 NBH 3 Roman Goc Institute of Physics, A. Mickiewicz University, Umultowska
Simulation of the NMR Second Moment as a Function of Temperature in the Presence of Molecular Motion. Application to (CH 3 ) 3 NBH 3 Roman Goc Institute of Physics, A. Mickiewicz University, Umultowska
Polarised Nucleon Targets for Europe, 2nd meeting, Bochum 2005
 Polarised Nucleon Targets for Europe, nd meeting, Bochum Temperature dependence of nuclear spin-lattice relaxations in liquid ethanol with dissolved TEMPO radicals H. Štěpánková, J. Englich, J. Kohout,
Polarised Nucleon Targets for Europe, nd meeting, Bochum Temperature dependence of nuclear spin-lattice relaxations in liquid ethanol with dissolved TEMPO radicals H. Štěpánková, J. Englich, J. Kohout,
Protein Dynamics, Allostery and Function
 Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
Protein Dynamics, Allostery and Function Lecture 3. Protein Dynamics Xiaolin Cheng UT/ORNL Center for Molecular Biophysics SJTU Summer School 2017 1 Obtaining Dynamic Information Experimental Approaches
Solid state 13 Cand 1 H MAS NMR investigations of C 60 (ferrocene-d 10 ) 2 complex
 Spectroscopy 17 (2003) 39 44 39 IOS Press Solid state 13 Cand 1 H MAS NMR investigations of C 60 (ferrocene-d 10 ) 2 complex E. Shabanova, K. Schaumburg and F.S. Kamounah CISMI, Department of Chemistry,
Spectroscopy 17 (2003) 39 44 39 IOS Press Solid state 13 Cand 1 H MAS NMR investigations of C 60 (ferrocene-d 10 ) 2 complex E. Shabanova, K. Schaumburg and F.S. Kamounah CISMI, Department of Chemistry,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
NMR SATELLITES AS A PROBE FOR CHEMICAL
 NMR SATELLITES AS A PROBE FOR CHEMICAL INVESTIGATIONS SHIzuo FUJIWARA, Yogi ARATA, HIR0sHI OZAWA and MASAYUKI KuruGI Department of Chemistry, The University of Tokyo, Japan ABSTRACT Satellite lines in
NMR SATELLITES AS A PROBE FOR CHEMICAL INVESTIGATIONS SHIzuo FUJIWARA, Yogi ARATA, HIR0sHI OZAWA and MASAYUKI KuruGI Department of Chemistry, The University of Tokyo, Japan ABSTRACT Satellite lines in
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Spin Dynamics Basics of Nuclear Magnetic Resonance. Malcolm H. Levitt
 Spin Dynamics Basics of Nuclear Magnetic Resonance Second edition Malcolm H. Levitt The University of Southampton, UK John Wiley &. Sons, Ltd Preface xxi Preface to the First Edition xxiii Introduction
Spin Dynamics Basics of Nuclear Magnetic Resonance Second edition Malcolm H. Levitt The University of Southampton, UK John Wiley &. Sons, Ltd Preface xxi Preface to the First Edition xxiii Introduction
Protein dynamics from NMR Relaxation data
 Protein dynamics from NMR Relaxation data Clubb 3/15/17 (S f2 ) ( e ) Nitrogen-15 relaxation ZZ-exchange R 1 = 1/T 1 Longitudinal relaxation (decay back to z-axis) R 2 = 1/T 2 Spin-spin relaxation (dephasing
Protein dynamics from NMR Relaxation data Clubb 3/15/17 (S f2 ) ( e ) Nitrogen-15 relaxation ZZ-exchange R 1 = 1/T 1 Longitudinal relaxation (decay back to z-axis) R 2 = 1/T 2 Spin-spin relaxation (dephasing
Protein folding. Today s Outline
 Protein folding Today s Outline Review of previous sessions Thermodynamics of folding and unfolding Determinants of folding Techniques for measuring folding The folding process The folding problem: Prediction
Protein folding Today s Outline Review of previous sessions Thermodynamics of folding and unfolding Determinants of folding Techniques for measuring folding The folding process The folding problem: Prediction
PRACTICAL ASPECTS OF NMR RELAXATION STUDIES OF BIOMOLECULAR DYNAMICS
 PRACTICAL ASPECTS OF MR RELAXATIO STUDIES OF BIOMOLECULAR DYAMICS Further reading: Can be downloaded from my web page Korzhnev D.E., Billeter M., Arseniev A.S., and Orekhov V. Y., MR Studies of Brownian
PRACTICAL ASPECTS OF MR RELAXATIO STUDIES OF BIOMOLECULAR DYAMICS Further reading: Can be downloaded from my web page Korzhnev D.E., Billeter M., Arseniev A.S., and Orekhov V. Y., MR Studies of Brownian
Principles of Physical Biochemistry
 Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
Principles of Physical Biochemistry Kensal E. van Hold e W. Curtis Johnso n P. Shing Ho Preface x i PART 1 MACROMOLECULAR STRUCTURE AND DYNAMICS 1 1 Biological Macromolecules 2 1.1 General Principles
7.88J Protein Folding Problem Fall 2007
 MIT OpenCourseWare http://ocw.mit.edu 7.88J Protein Folding Problem Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Lecture Notes - 3 7.24/7.88J/5.48J
MIT OpenCourseWare http://ocw.mit.edu 7.88J Protein Folding Problem Fall 2007 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Lecture Notes - 3 7.24/7.88J/5.48J
Lecture 34 Protein Unfolding Thermodynamics
 Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Supporting Information
 Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
Supporting Information Boehr et al. 10.1073/pnas.0914163107 SI Text Materials and Methods. R 2 relaxation dispersion experiments. 15 NR 2 relaxation dispersion data measured at 1 H Larmor frequencies of
EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December Suggested resolution
 page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
The change in free energy on transferring an ion from a medium of low dielectric constantε1 to one of high dielectric constant ε2:
 The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
The Born Energy of an Ion The free energy density of an electric field E arising from a charge is ½(ε 0 ε E 2 ) per unit volume Integrating the energy density of an ion over all of space = Born energy:
NMR Relaxation and Molecular Dynamics
 Ecole RMN Cargese Mars 2008 NMR Relaxation and Molecular Dynamics Martin Blackledge IBS Grenoble Carine van Heijenoort ICSN, CNRS Gif-sur-Yvette Solution NMR Timescales for Biomolecular Motion ps ns µs
Ecole RMN Cargese Mars 2008 NMR Relaxation and Molecular Dynamics Martin Blackledge IBS Grenoble Carine van Heijenoort ICSN, CNRS Gif-sur-Yvette Solution NMR Timescales for Biomolecular Motion ps ns µs
Chemistry 111 Syllabus
 Chemistry 111 Syllabus Chapter 1: Chemistry: The Science of Change The Study of Chemistry Chemistry You May Already Know The Scientific Method Classification of Matter Pure Substances States of Matter
Chemistry 111 Syllabus Chapter 1: Chemistry: The Science of Change The Study of Chemistry Chemistry You May Already Know The Scientific Method Classification of Matter Pure Substances States of Matter
The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals.
 Physical Metallurgy The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals. Crystal Binding In our discussions
Physical Metallurgy The broad topic of physical metallurgy provides a basis that links the structure of materials with their properties, focusing primarily on metals. Crystal Binding In our discussions
PROTEIN NMR SPECTROSCOPY
 List of Figures List of Tables xvii xxvi 1. NMR SPECTROSCOPY 1 1.1 Introduction to NMR Spectroscopy 2 1.2 One Dimensional NMR Spectroscopy 3 1.2.1 Classical Description of NMR Spectroscopy 3 1.2.2 Nuclear
List of Figures List of Tables xvii xxvi 1. NMR SPECTROSCOPY 1 1.1 Introduction to NMR Spectroscopy 2 1.2 One Dimensional NMR Spectroscopy 3 1.2.1 Classical Description of NMR Spectroscopy 3 1.2.2 Nuclear
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Lecture 2-3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions Van der Waals Interactions
Biophysical Chemistry: NMR Spectroscopy
 Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
Relaxation & Multidimensional Spectrocopy Vrije Universiteit Brussel 9th December 2011 Outline 1 Relaxation 2 Principles 3 Outline 1 Relaxation 2 Principles 3 Establishment of Thermal Equilibrium As previously
'H NMR Techniques in Studies of Transport of Paramagnetic Ions in Multicellular Systems
 Gen. Physiol. Biophys. (1987), 6, 609 615 609 'H NMR Techniques in Studies of Transport of Paramagnetic Ions in Multicellular Systems S. RATKOVIČ 1 AND G. BAČIČ 2 1 Department of Technology and Chemical
Gen. Physiol. Biophys. (1987), 6, 609 615 609 'H NMR Techniques in Studies of Transport of Paramagnetic Ions in Multicellular Systems S. RATKOVIČ 1 AND G. BAČIČ 2 1 Department of Technology and Chemical
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability
 Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
Proteins are not rigid structures: Protein dynamics, conformational variability, and thermodynamic stability Dr. Andrew Lee UNC School of Pharmacy (Div. Chemical Biology and Medicinal Chemistry) UNC Med
I690/B680 Structural Bioinformatics Spring Protein Structure Determination by NMR Spectroscopy
 I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
I690/B680 Structural Bioinformatics Spring 2006 Protein Structure Determination by NMR Spectroscopy Suggested Reading (1) Van Holde, Johnson, Ho. Principles of Physical Biochemistry, 2 nd Ed., Prentice
Measuring the size and shape of macromolecules. Hydrodynamics: study of the objects in water How do the move? Translation Rotation
 Measuring the size and shape of macromolecules Hydrodynamics: study of the objects in water How do the move? Translation Rotation 1) Movement with no external forcefree diffusion 2) Movement under the
Measuring the size and shape of macromolecules Hydrodynamics: study of the objects in water How do the move? Translation Rotation 1) Movement with no external forcefree diffusion 2) Movement under the
The Oxford Solid State Basics
 The Oxford Solid State Basics Steven H. Simon University of Oxford OXFORD UNIVERSITY PRESS Contents 1 About Condensed Matter Physics 1 1.1 What Is Condensed Matter Physics 1 1.2 Why Do We Study Condensed
The Oxford Solid State Basics Steven H. Simon University of Oxford OXFORD UNIVERSITY PRESS Contents 1 About Condensed Matter Physics 1 1.1 What Is Condensed Matter Physics 1 1.2 Why Do We Study Condensed
Experimental Techniques in Protein Structure Determination
 Experimental Techniques in Protein Structure Determination Homayoun Valafar Department of Computer Science and Engineering, USC Two Main Experimental Methods X-Ray crystallography Nuclear Magnetic Resonance
Experimental Techniques in Protein Structure Determination Homayoun Valafar Department of Computer Science and Engineering, USC Two Main Experimental Methods X-Ray crystallography Nuclear Magnetic Resonance
Biochemistry 530 NMR Theory and Practice
 Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington Lecturer: Gabriele Varani Biochemistry and Chemistry Room J479 and
Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington Lecturer: Gabriele Varani Biochemistry and Chemistry Room J479 and
Classes of Hydration Sites at Protein-Water Interfaces: The Source of Contrast in Magnetic Resonance Imaging
 Biophysical Journal Volume 69 August 1995 593-603 Classes of Hydration Sites at Protein-Water Interfaces: The Source of Contrast in Magnetic Resonance Imaging 593 Seymour H. Koenig Relaxometry Inc., Mahopac,
Biophysical Journal Volume 69 August 1995 593-603 Classes of Hydration Sites at Protein-Water Interfaces: The Source of Contrast in Magnetic Resonance Imaging 593 Seymour H. Koenig Relaxometry Inc., Mahopac,
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
BRIEF COMMUNICATION PROTONS IN METHEMOGLOBIN SOLUTIONS NMR RELAXATION OF PROTEIN AND WATER
 BRIEF COMMUNICATION NMR RELAXATION OF PROTEIN AND WATER PROTONS IN METHEMOGLOBIN SOLUTIONS MAURICE EISENSTADT, Department of Medicine, Albert Einstein College of Medicine, Bronx, New York 10461 U.S.A.
BRIEF COMMUNICATION NMR RELAXATION OF PROTEIN AND WATER PROTONS IN METHEMOGLOBIN SOLUTIONS MAURICE EISENSTADT, Department of Medicine, Albert Einstein College of Medicine, Bronx, New York 10461 U.S.A.
Suspended Long-Lived NMR Echo in Solids
 Suspended Long-Lived NMR Echo in Solids A. Turanov 1 and A.K. Khitrin 2 1 Zavoisky Physical-Technical Institute RAS, Kazan, 420029, Russia 2 Department of Chemistry, Kent State University, OH 44242, USA
Suspended Long-Lived NMR Echo in Solids A. Turanov 1 and A.K. Khitrin 2 1 Zavoisky Physical-Technical Institute RAS, Kazan, 420029, Russia 2 Department of Chemistry, Kent State University, OH 44242, USA
Hydrogen/Deuterium Exchange Mass Spectrometry: A Mini-Tutorial
 Florida State University National High Magnetic Field Laboratory Tallahassee-Florida Hydrogen/euterium Exchange Mass Spectrometry: A Mini-Tutorial George Bou-Assaf 56 th ASMS Conference June 2 nd, 2008
Florida State University National High Magnetic Field Laboratory Tallahassee-Florida Hydrogen/euterium Exchange Mass Spectrometry: A Mini-Tutorial George Bou-Assaf 56 th ASMS Conference June 2 nd, 2008
6 Hydrophobic interactions
 The Physics and Chemistry of Water 6 Hydrophobic interactions A non-polar molecule in water disrupts the H- bond structure by forcing some water molecules to give up their hydrogen bonds. As a result,
The Physics and Chemistry of Water 6 Hydrophobic interactions A non-polar molecule in water disrupts the H- bond structure by forcing some water molecules to give up their hydrogen bonds. As a result,
Lecture #7 In Vivo Water
 Lecture #7 In Vivo Water Topics Hydration layers Tissue relaxation times Magic angle effects Magnetization Transfer Contrast (MTC) CEST Handouts and Reading assignments Mathur-De Vre, R., The NMR studies
Lecture #7 In Vivo Water Topics Hydration layers Tissue relaxation times Magic angle effects Magnetization Transfer Contrast (MTC) CEST Handouts and Reading assignments Mathur-De Vre, R., The NMR studies
Magnetic Resonance Spectroscopy
 INTRODUCTION TO Magnetic Resonance Spectroscopy ESR, NMR, NQR D. N. SATHYANARAYANA Formerly, Chairman Department of Inorganic and Physical Chemistry Indian Institute of Science, Bangalore % I.K. International
INTRODUCTION TO Magnetic Resonance Spectroscopy ESR, NMR, NQR D. N. SATHYANARAYANA Formerly, Chairman Department of Inorganic and Physical Chemistry Indian Institute of Science, Bangalore % I.K. International
Chapter-2 (Page 22-37) Physical and Chemical Properties of Water
 Chapter-2 (Page 22-37) Physical and Chemical Properties of Water Introduction About 70% of the mass of the human body is water. Water is central to biochemistry for the following reasons: 1- Biological
Chapter-2 (Page 22-37) Physical and Chemical Properties of Water Introduction About 70% of the mass of the human body is water. Water is central to biochemistry for the following reasons: 1- Biological
Basic One- and Two-Dimensional NMR Spectroscopy
 Horst Friebolin Basic One- and Two-Dimensional NMR Spectroscopy Third Revised Edition Translated by Jack K. Becconsall WILEY-VCH Weinheim New York Chichester Brisbane Singapore Toronto Contents XV 1 The
Horst Friebolin Basic One- and Two-Dimensional NMR Spectroscopy Third Revised Edition Translated by Jack K. Becconsall WILEY-VCH Weinheim New York Chichester Brisbane Singapore Toronto Contents XV 1 The
SUPPLEMENTARY NOTE 1: ADDITIONAL CHARACTERIZATION OF NANODIAMOND SOLUTIONS AND THE OVERHAUSER EFFECT
 1 SUPPLEMENTARY NOTE 1: ADDITIONAL CHARACTERIZATION OF NANODIAMOND SOLUTIONS AND THE OVERHAUSER EFFECT Nanodiamond (ND) solutions were prepared using high power probe sonication and analyzed by dynamic
1 SUPPLEMENTARY NOTE 1: ADDITIONAL CHARACTERIZATION OF NANODIAMOND SOLUTIONS AND THE OVERHAUSER EFFECT Nanodiamond (ND) solutions were prepared using high power probe sonication and analyzed by dynamic
Macromolecular Crowding
 Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
Macromolecular Crowding Keng-Hwee Chiam Mathematical and Theoretical Biology Group Goodsell (1994) Macromolecular Crowding, Oct. 15, 2003 p.1/33 Outline What: introduction, definition Why: implications
NMR Spectroscopy. Guangjin Hou
 NMR Spectroscopy Guangjin Hou 22-04-2009 NMR History 1 H NMR spectra of water H NMR spectra of water (First NMR Spectra on Water, 1946) 1 H NMR spectra ethanol (First bservation of the Chemical Shift,
NMR Spectroscopy Guangjin Hou 22-04-2009 NMR History 1 H NMR spectra of water H NMR spectra of water (First NMR Spectra on Water, 1946) 1 H NMR spectra ethanol (First bservation of the Chemical Shift,
MULTINUCLEAR NMR. EDITED BY JOAN MASON The Open University Milton Keynes, Buckinghamshire, England PLENUM PRESS NEW YORK AND LONDON
 MULTINUCLEAR NMR EDITED BY JOAN MASON The Open University Milton Keynes, Buckinghamshire, England PLENUM PRESS NEW YORK AND LONDON CONTENTS Chapter 1 Introduction 1 Joan Mason Chapter 2 The Parameters
MULTINUCLEAR NMR EDITED BY JOAN MASON The Open University Milton Keynes, Buckinghamshire, England PLENUM PRESS NEW YORK AND LONDON CONTENTS Chapter 1 Introduction 1 Joan Mason Chapter 2 The Parameters
COMBINED EFFECT OF RESTRICTED ROTATIONAL
 COMBINED EFFECT OF RESTRICTED ROTATIONAL DIFFUSION PLUS JUMPS ON NUCLEAR MAGNETIC RESONANCE AND FLUORESCENCE PROBES OF AROMATIC RING MOTIONS IN PROTEINS RONALD M. LEVY AND ROBERT P. SHERIDAN Department
COMBINED EFFECT OF RESTRICTED ROTATIONAL DIFFUSION PLUS JUMPS ON NUCLEAR MAGNETIC RESONANCE AND FLUORESCENCE PROBES OF AROMATIC RING MOTIONS IN PROTEINS RONALD M. LEVY AND ROBERT P. SHERIDAN Department
Time-Dependent Statistical Mechanics 1. Introduction
 Time-Dependent Statistical Mechanics 1. Introduction c Hans C. Andersen Announcements September 24, 2009 Lecture 1 9/22/09 1 Topics of concern in the course We shall be concerned with the time dependent
Time-Dependent Statistical Mechanics 1. Introduction c Hans C. Andersen Announcements September 24, 2009 Lecture 1 9/22/09 1 Topics of concern in the course We shall be concerned with the time dependent
Effects of Chemical Exchange on NMR Spectra
 Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
Effects of Chemical Exchange on NMR Spectra Chemical exchange refers to any process in which a nucleus exchanges between two or more environments in which its NMR parameters (e.g. chemical shift, scalar
e 2m e c I, (7.1) = g e β B I(I +1), (7.2) = erg/gauss. (7.3)
 Chemistry 126 Molecular Spectra & Molecular Structure Week # 7 Electron Spin Resonance Spectroscopy, Supplement Like the hydrogen nucleus, an unpaired electron in a sample has a spin of I=1/2. The magnetic
Chemistry 126 Molecular Spectra & Molecular Structure Week # 7 Electron Spin Resonance Spectroscopy, Supplement Like the hydrogen nucleus, an unpaired electron in a sample has a spin of I=1/2. The magnetic
BMB/Bi/Ch 173 Winter 2018
 BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
BMB/Bi/Ch 173 Winter 2018 Homework Set 8.1 (100 Points) Assigned 2-27-18, due 3-6-18 by 10:30 a.m. TA: Rachael Kuintzle. Office hours: SFL 220, Friday 3/2 4:00-5:00pm and SFL 229, Monday 3/5 4:00-5:30pm.
NMR Resonance Assignment Assisted by Mass Spectrometry
 NMR Resonance Assignment Assisted by Mass Spectrometry This lecture talked about a NMR resonance assignment assisted by mass spectrometry [1, 2]. 1 Motivation Nuclear magnetic resonance (NMR) provides
NMR Resonance Assignment Assisted by Mass Spectrometry This lecture talked about a NMR resonance assignment assisted by mass spectrometry [1, 2]. 1 Motivation Nuclear magnetic resonance (NMR) provides
Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR
 Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR Residual Dipolar Couplings (RDC s) Relatively new technique ~ 1996 Nico Tjandra, Ad Bax- NIH, Jim Prestegard, UGA Combination of
Theory and Applications of Residual Dipolar Couplings in Biomolecular NMR Residual Dipolar Couplings (RDC s) Relatively new technique ~ 1996 Nico Tjandra, Ad Bax- NIH, Jim Prestegard, UGA Combination of
Protein Dynamics. The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron.
 Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Protein Dynamics The space-filling structures of myoglobin and hemoglobin show that there are no pathways for O 2 to reach the heme iron. Below is myoglobin hydrated with 350 water molecules. Only a small
Water Dynamics in Protein Hydration Shells: The Molecular Origins of the Dynamical Perturbation
 pubs.acs.org/jpcb Terms of Use Water Dynamics in Protein Hydration Shells: The Molecular Origins of the Dynamical Perturbation Aoife C. Fogarty and Damien Laage* Department of Chemistry, UMR ENS-CNRS-UPMC
pubs.acs.org/jpcb Terms of Use Water Dynamics in Protein Hydration Shells: The Molecular Origins of the Dynamical Perturbation Aoife C. Fogarty and Damien Laage* Department of Chemistry, UMR ENS-CNRS-UPMC
Chapter 10. Liquids and Solids
 Chapter 10 Liquids and Solids Chapter 10 Table of Contents 10.1 Intermolecular Forces 10.2 The Liquid State 10.3 An Introduction to Structures and Types of Solids 10.4 Structure and Bonding in Metals 10.5
Chapter 10 Liquids and Solids Chapter 10 Table of Contents 10.1 Intermolecular Forces 10.2 The Liquid State 10.3 An Introduction to Structures and Types of Solids 10.4 Structure and Bonding in Metals 10.5
Structure of the α-helix
 Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Structure of the α-helix Structure of the β Sheet Protein Dynamics Basics of Quenching HDX Hydrogen exchange of amide protons is catalyzed by H 2 O, OH -, and H 3 O +, but it s most dominated by base
Nature Structural & Molecular Biology: doi: /nsmb.3194
 Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Supplementary Figure 1 Mass spectrometry and solution NMR data for -syn samples used in this study. (a) Matrix-assisted laser-desorption and ionization time-of-flight (MALDI-TOF) mass spectrum of uniformly-
Lecture #6 (The NOE)
 Lecture #6 (The OE) 2/24/17 Clubb Determining Protein tructures by MR: Measure thousands of shorter inter-hydrogen atom distances. Use these to restrain the structure of protein computationally. Distances
Lecture #6 (The OE) 2/24/17 Clubb Determining Protein tructures by MR: Measure thousands of shorter inter-hydrogen atom distances. Use these to restrain the structure of protein computationally. Distances
Chapter 11. Intermolecular Forces and Liquids & Solids
 Chapter 11 Intermolecular Forces and Liquids & Solids The Kinetic Molecular Theory of Liquids & Solids Gases vs. Liquids & Solids difference is distance between molecules Liquids Molecules close together;
Chapter 11 Intermolecular Forces and Liquids & Solids The Kinetic Molecular Theory of Liquids & Solids Gases vs. Liquids & Solids difference is distance between molecules Liquids Molecules close together;
Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination
 Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Lecture 9 M230 Feigon Sequential resonance assignments in (small) proteins: homonuclear method 2º structure determination Reading resources v Roberts NMR of Macromolecules, Chap 4 by Christina Redfield
Lab Week 4 Module α 3. Polymer Conformation. Lab. Instructor : Francesco Stellacci
 3.014 Materials Laboratory Dec. 9 th Dec.14 th, 2004 Lab Week 4 Module α 3 Polymer Conformation Lab. Instructor : Francesco Stellacci OBJECTIVES 9 Review random walk model for polymer chains 9 Introduce
3.014 Materials Laboratory Dec. 9 th Dec.14 th, 2004 Lab Week 4 Module α 3 Polymer Conformation Lab. Instructor : Francesco Stellacci OBJECTIVES 9 Review random walk model for polymer chains 9 Introduce
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
CHEM Principles of Chemistry II Chapter 10 - Liquids and Solids
 CHEM 1212 - Principles of Chemistry II Chapter 10 - Liquids and Solids 10.1 Intermolecular Forces recall intramolecular (within the molecule) bonding whereby atoms can form stable units called molecules
CHEM 1212 - Principles of Chemistry II Chapter 10 - Liquids and Solids 10.1 Intermolecular Forces recall intramolecular (within the molecule) bonding whereby atoms can form stable units called molecules
CHAPTER ELEVEN KINETIC MOLECULAR THEORY OF LIQUIDS AND SOLIDS KINETIC MOLECULAR THEORY OF LIQUIDS AND SOLIDS
 CHAPTER ELEVEN AND LIQUIDS AND SOLIDS KINETIC MOLECULAR THEORY OF LIQUIDS AND SOLIDS Differences between condensed states and gases? KINETIC MOLECULAR THEORY OF LIQUIDS AND SOLIDS Phase Homogeneous part
CHAPTER ELEVEN AND LIQUIDS AND SOLIDS KINETIC MOLECULAR THEORY OF LIQUIDS AND SOLIDS Differences between condensed states and gases? KINETIC MOLECULAR THEORY OF LIQUIDS AND SOLIDS Phase Homogeneous part
How DLS Works: Interference of Light
 Static light scattering vs. Dynamic light scattering Static light scattering measures time-average intensities (mean square fluctuations) molecular weight radius of gyration second virial coefficient Dynamic
Static light scattering vs. Dynamic light scattering Static light scattering measures time-average intensities (mean square fluctuations) molecular weight radius of gyration second virial coefficient Dynamic
ν + = e2 qq φ = 36.9,andθ = Pankratov [4] obtained ν 0 = ν + ν = e2 qq φ = 34.1, and θ = Boogaarts et al. [5]
![ν + = e2 qq φ = 36.9,andθ = Pankratov [4] obtained ν 0 = ν + ν = e2 qq φ = 34.1, and θ = Boogaarts et al. [5] ν + = e2 qq φ = 36.9,andθ = Pankratov [4] obtained ν 0 = ν + ν = e2 qq φ = 34.1, and θ = Boogaarts et al. [5]](/thumbs/73/68415777.jpg) 14 N NQR Study of Diphenylamine Janez Seliger a,b and Veselko Žagar a a Jozef Stefan Institute, Jamova 39, 1000 Ljubljana, Slovenia b University of Ljubljana, Faculty of Mathematics and Physics, Department
14 N NQR Study of Diphenylamine Janez Seliger a,b and Veselko Žagar a a Jozef Stefan Institute, Jamova 39, 1000 Ljubljana, Slovenia b University of Ljubljana, Faculty of Mathematics and Physics, Department
Structure, dynamics and heterogeneity: solid-state NMR of polymers. Jeremy Titman, School of Chemistry, University of Nottingham
 Structure, dynamics and heterogeneity: solid-state NMR of polymers Jeremy Titman, School of Chemistry, University of Nottingham Structure, dynamics and heterogeneity Structure Dynamics conformation, tacticity,
Structure, dynamics and heterogeneity: solid-state NMR of polymers Jeremy Titman, School of Chemistry, University of Nottingham Structure, dynamics and heterogeneity Structure Dynamics conformation, tacticity,
Principles of Nuclear Magnetic Resonance Microscopy
 Principles of Nuclear Magnetic Resonance Microscopy Paul T. Callaghan Department of Physics and Biophysics Massey University New Zealand CLARENDON PRESS OXFORD CONTENTS 1 PRINCIPLES OF IMAGING 1 1.1 Introduction
Principles of Nuclear Magnetic Resonance Microscopy Paul T. Callaghan Department of Physics and Biophysics Massey University New Zealand CLARENDON PRESS OXFORD CONTENTS 1 PRINCIPLES OF IMAGING 1 1.1 Introduction
Polymer dynamics. Course M6 Lecture 5 26/1/2004 (JAE) 5.1 Introduction. Diffusion of polymers in melts and dilute solution.
 Course M6 Lecture 5 6//004 Polymer dynamics Diffusion of polymers in melts and dilute solution Dr James Elliott 5. Introduction So far, we have considered the static configurations and morphologies of
Course M6 Lecture 5 6//004 Polymer dynamics Diffusion of polymers in melts and dilute solution Dr James Elliott 5. Introduction So far, we have considered the static configurations and morphologies of
Christopher Pavlik Bioanalytical Chemistry March 2, 2011
 Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Nuclear Magnetic Resonance of Proteins Christopher Pavlik Bioanalytical Chemistry March 2, 2011 Nuclear Magnetic Resonance NMR Application of a magnetic field causes absorption of EM energy that induces
Long-Range Correlated Dynamics in Intrinsically Disordered Proteins
 SUPPORTING INFORMATION Long-Range Correlated Dynamics in Intrinsically Disordered Proteins Giacomo Parigi, Nasrollah Rezaei-Ghaleh, Andrea Giachetti, Stefan Becker, Claudio Fernandez, Martin Blackledge,
SUPPORTING INFORMATION Long-Range Correlated Dynamics in Intrinsically Disordered Proteins Giacomo Parigi, Nasrollah Rezaei-Ghaleh, Andrea Giachetti, Stefan Becker, Claudio Fernandez, Martin Blackledge,
A large number of enzymes display an absolute requirement
 Biochemistry 180, 1, 5481-5485 548 1 A Multinuclear Nuclear Magnetic Resonance Study of the Monovalent-Divalent Cation Sites of Pyruvate Kinase? Frank M. Raushelt and Joseph J. Villafranca* ABSTRACT: The
Biochemistry 180, 1, 5481-5485 548 1 A Multinuclear Nuclear Magnetic Resonance Study of the Monovalent-Divalent Cation Sites of Pyruvate Kinase? Frank M. Raushelt and Joseph J. Villafranca* ABSTRACT: The
NUCLEAR MAGNETIC RESONANCE. The phenomenon of nuclear magnetic resonance will be used to study magnetic moments of nuclei.
 14 Sep 11 NMR.1 NUCLEAR MAGNETIC RESONANCE The phenomenon of nuclear magnetic resonance will be used to study magnetic moments of nuclei. Theory: In addition to its well-known properties of mass, charge,
14 Sep 11 NMR.1 NUCLEAR MAGNETIC RESONANCE The phenomenon of nuclear magnetic resonance will be used to study magnetic moments of nuclei. Theory: In addition to its well-known properties of mass, charge,
Lecture #6 (The NOE)
 Lecture #6 (The OE) 2/18/15 Clubb Determining Protein tructures by MR: Measure thousands of shorter inter-hydrogen atom distances. Use these to restrain the structure of protein computationally. Distance
Lecture #6 (The OE) 2/18/15 Clubb Determining Protein tructures by MR: Measure thousands of shorter inter-hydrogen atom distances. Use these to restrain the structure of protein computationally. Distance
Hydrodynamics: Viscosity and Diffusion Hydrodynamics is the study of mechanics in a liquid, where the frictional drag of the liquid cannot be ignored
 Hydrodynamics: Viscosity and Diffusion Hydrodynamics is the study of mechanics in a liquid, where the frictional drag of the liquid cannot be ignored First let s just consider fluid flow, where the fluid
Hydrodynamics: Viscosity and Diffusion Hydrodynamics is the study of mechanics in a liquid, where the frictional drag of the liquid cannot be ignored First let s just consider fluid flow, where the fluid
Dynamics of Protein and Peptide Hydration
 Published on Web 12/12/2003 Dynamics of Protein and Peptide Hydration Kristofer Modig, Edvards Liepinsh, Gottfried Otting,, and Bertil Halle*, Contribution from the Department of Biophysical Chemistry,
Published on Web 12/12/2003 Dynamics of Protein and Peptide Hydration Kristofer Modig, Edvards Liepinsh, Gottfried Otting,, and Bertil Halle*, Contribution from the Department of Biophysical Chemistry,
Introduction solution NMR
 2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
2 NMR journey Introduction solution NMR Alexandre Bonvin Bijvoet Center for Biomolecular Research with thanks to Dr. Klaartje Houben EMBO Global Exchange course, IHEP, Beijing April 28 - May 5, 20 3 Topics
Muons in Chemistry Training School Dr N J Clayden School of Chemistry University of East Anglia Norwich
 Muons in Chemistry Training School 2014 Dr N J Clayden School of Chemistry University of East Anglia Norwich Why use muons? Extrinsic probe (Mu +, Mu, muoniated radical) Intrinsic interest Framing of the
Muons in Chemistry Training School 2014 Dr N J Clayden School of Chemistry University of East Anglia Norwich Why use muons? Extrinsic probe (Mu +, Mu, muoniated radical) Intrinsic interest Framing of the
WATER MOBILITY AND STRUCTURE IN NATURAL CLAY SYSTEMS
 The Clay Minerals Society Workshop Lectures Series, Vol. 21 (2016), Chapter 7, 79 86 WATER MOBILITY AND STRUCTURE IN NATURAL CLAY SYSTEMS MARC FLEURY,ERIC KOHLER, and LOIC BARRÉ IFP Energies nouvelles,
The Clay Minerals Society Workshop Lectures Series, Vol. 21 (2016), Chapter 7, 79 86 WATER MOBILITY AND STRUCTURE IN NATURAL CLAY SYSTEMS MARC FLEURY,ERIC KOHLER, and LOIC BARRÉ IFP Energies nouvelles,
Vol. 114 (2008) ACTA PHYSICA POLONICA A No. 6 A
 Vol. 114 (2008) ACTA PHYSICA POLONICA A No. 6 A Optical and Acoustical Methods in Science and Technology Effects of Solvation of 2-Methylpyridine and 2,6-Dimethylpyridine in Dilute Solutions in Water and
Vol. 114 (2008) ACTA PHYSICA POLONICA A No. 6 A Optical and Acoustical Methods in Science and Technology Effects of Solvation of 2-Methylpyridine and 2,6-Dimethylpyridine in Dilute Solutions in Water and
Supplementary Figure 1: Spin noise spectra of 55 Mn in bulk sample at BL =10.5 mt, before subtraction of the zero-frequency line. a, Contour plot of
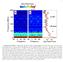 1 Supplementary Figure 1: Spin noise spectra of 55 Mn in bulk sample at BL =10.5 mt, before subtraction of the zero-frequency line. a, Contour plot of the spin noise spectra calculated with Eq. (2) for
1 Supplementary Figure 1: Spin noise spectra of 55 Mn in bulk sample at BL =10.5 mt, before subtraction of the zero-frequency line. a, Contour plot of the spin noise spectra calculated with Eq. (2) for
Nuclear Magnetic Resonance Studies of 35Cl, 37C1, 79Br, and 81Br in Aqueous Solution
 Nuclear Magnetic Resonance Studies of 35Cl, 37C, 79Br, and in Aqueous Solution J. BLASER, 0. L U T Z, a n d W. STEINKILBERG Physikalisches Institut der Universität T ü b i n g e n (Z. Naturforsch. 7 a,
Nuclear Magnetic Resonance Studies of 35Cl, 37C, 79Br, and in Aqueous Solution J. BLASER, 0. L U T Z, a n d W. STEINKILBERG Physikalisches Institut der Universität T ü b i n g e n (Z. Naturforsch. 7 a,
The Role of Metal Ions in Processes of Conformational Selection during Ligand- Macromolecule Interactions
 102 Bulletin of Magnetic Resonance The Role of Metal Ions in Processes of Conformational Selection during Ligand- Macromolecule Interactions E.Gaggelli, N.Gaggelli, G.Valensin Department of Chemistry,
102 Bulletin of Magnetic Resonance The Role of Metal Ions in Processes of Conformational Selection during Ligand- Macromolecule Interactions E.Gaggelli, N.Gaggelli, G.Valensin Department of Chemistry,
Orientational Disorder and Entropy of Water in Protein Cavities
 9380 J. Phys. Chem. B 1997, 101, 9380-9389 Orientational Disorder and Entropy of Water in Protein Cavities Vladimir P. Denisov, Kandadai Venu,, Jo1rg Peters, Hans Dietrich Ho1rlein, and Bertil Halle*,
9380 J. Phys. Chem. B 1997, 101, 9380-9389 Orientational Disorder and Entropy of Water in Protein Cavities Vladimir P. Denisov, Kandadai Venu,, Jo1rg Peters, Hans Dietrich Ho1rlein, and Bertil Halle*,
Coupling of Functional Hydrogen Bonds in Pyridoxal-5 -phosphate- Enzyme Model Systems Observed by Solid State NMR
 Supporting Information for Coupling of Functional ydrogen Bonds in Pyridoxal-5 -phosphate- Enzyme Model Systems bserved by Solid State NMR Shasad Sharif, David Schagen, Michael D. Toney, and ans-einrich
Supporting Information for Coupling of Functional ydrogen Bonds in Pyridoxal-5 -phosphate- Enzyme Model Systems bserved by Solid State NMR Shasad Sharif, David Schagen, Michael D. Toney, and ans-einrich
Protein Structure Determination using NMR Spectroscopy. Cesar Trinidad
 Protein Structure Determination using NMR Spectroscopy Cesar Trinidad Introduction Protein NMR Involves the analysis and calculation of data collected from multiple NMR techniques Utilizes Nuclear Magnetic
Protein Structure Determination using NMR Spectroscopy Cesar Trinidad Introduction Protein NMR Involves the analysis and calculation of data collected from multiple NMR techniques Utilizes Nuclear Magnetic
Proton MAS NMR of a protein in a frozen aqueous solution
 See discussions, stats, and author profiles for this publication at: https://www.researchgate.net/publication/224041726 Proton MAS NMR of a protein in a frozen aqueous solution Article in Chemical Physics
See discussions, stats, and author profiles for this publication at: https://www.researchgate.net/publication/224041726 Proton MAS NMR of a protein in a frozen aqueous solution Article in Chemical Physics
IMFA s. intermolecular forces of attraction Chez Chem, LLC All rights reserved.
 IMFA s intermolecular forces of attraction 2014 Chez Chem, LLC All rights reserved. **London Dispersion Forces Also know as Van der Waals forces A momentary non symmetrical electron distribution that can
IMFA s intermolecular forces of attraction 2014 Chez Chem, LLC All rights reserved. **London Dispersion Forces Also know as Van der Waals forces A momentary non symmetrical electron distribution that can
Medical Biophysics II. Final exam theoretical questions 2013.
 Medical Biophysics II. Final exam theoretical questions 2013. 1. Early atomic models. Rutherford-experiment. Franck-Hertz experiment. Bohr model of atom. 2. Quantum mechanical atomic model. Quantum numbers.
Medical Biophysics II. Final exam theoretical questions 2013. 1. Early atomic models. Rutherford-experiment. Franck-Hertz experiment. Bohr model of atom. 2. Quantum mechanical atomic model. Quantum numbers.
ALGORITHM OF ANALYSE OF SPIN-SPIN RELAXATION IN POLYBUTADIENE-C 6 H 12 AND POLYBUTADIENE-C 6 D 12 SOLUTIONS
 ALGORITHM OF ANALYSE OF SPIN-SPIN RELAXATION IN POLYBUTADIENE-C 6 H 12 AND POLYBUTADIENE-C 6 D 12 SOLUTIONS M. Todica Babes-Bolyai University, Faculty of Physics, 3400 Cluj-Napoca. Abstract The comparative
ALGORITHM OF ANALYSE OF SPIN-SPIN RELAXATION IN POLYBUTADIENE-C 6 H 12 AND POLYBUTADIENE-C 6 D 12 SOLUTIONS M. Todica Babes-Bolyai University, Faculty of Physics, 3400 Cluj-Napoca. Abstract The comparative
Biochemistry 530 NMR Theory and Practice
 Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington 1D spectra contain structural information.. but is hard to extract:
Biochemistry 530 NMR Theory and Practice Gabriele Varani Department of Biochemistry and Department of Chemistry University of Washington 1D spectra contain structural information.. but is hard to extract:
