Self-Assembling Cyclic Peptides: Molecular Dynamics Studies of Dimers in Polar and Nonpolar Solvents
|
|
|
- Madeline Edwards
- 6 years ago
- Views:
Transcription
1 J. Phys. Chem. B 2006, 110, Self-Assembling Cyclic Peptides: Molecular Dynamics Studies of Dimers in Polar and Nonpolar Solvents Ekta Khurana,* Steven O. Nielsen, Bernd Ensing, and Michael L. Klein Center for Molecular Modeling and Department of Chemistry, UniVersity of PennsylVania, Philadelphia, PennsylVania ReceiVed: December 22, 2005; In Final Form: March 17, 2006 The self-assembly of cyclic D,L-R-peptides into hollow nanotubes is a crucial mechanistic step in their application as antibacterial and drug-delivery agents. To understand this process, molecular dynamics (MD) simulations were performed on dimers of cyclic peptides formed from cyclo [(-L-Trp-D-N-MeLeu-) 4 -] 2 and cyclo [(-L-Trp-D-Leu-) 4 -] 2 subunits in nonpolar (nonane) and polar (water) solvent. The dimers were observed to be stable only in nonpolar solvent over the full 10 ns length of the MD trajectory. The behavior of the dimers in different solvents is rationalized in terms of the intersubunit hydrogen bonding, hydrogen bonding with the solvent, and planarity of the rings. It is shown that the φ and ψ dihedral angles of a single uncapped ring in nonane lie in the β-sheet region of the Ramachandran plot, and the ring stays in a flat conformation. Steered MD (SMD) simulations based on Jarzynski s equality were performed to obtain the potential of mean force as a function of the distance between the two rings of the capped dimer in nonane. It is also shown that a single peptide subunit prefers to reside close to the nonane/water interface rather than in bulk solvent because of the amphiphilic character of the peptide ring. The present MD results build the foundation for using MD simulations to study the mechanism of the formation of cyclic peptide nanotubes in lipid bilayers. Introduction Ghadiri and co-workers have designed remarkable synthetic nanotubes, which are formed by the self-assembly of cyclic peptides with an even number of alternating D- and L-amino acid residues. The cyclic peptides in question adopt a planar ring conformation and stack, under favorable conditions, to form hydrogen-bonded nanotubes, which can act as selective antibacterial agents. Moreover, the cyclic D,L-R-peptide building blocks are proteolytically stable, easy to synthesize, and can be derived from a potentially vast membrane-active sequence space. 1 Such self-assembling peptide nanotubes have been studied experimentally by high-resolution X-ray crystallography, cryoelectron microscopy, electron diffraction, and Fourier transform infrared (FT-IR) spectroscopy, 2-7 as well as theoretically by ab initio and semiempirical approaches Molecular dynamics (MD) simulations focusing on the structure and dynamics of pre-assembled nanotubes have also been performed. 11,12 The computational studies have thrown light on the structure of these nanotubes and their action as ion channels. In an ideal supramolecular nanotube assembly, rings stack through backbone-backbone β-sheet-like hydrogen bonding. The individual peptide rings adopt a planar conformation in which all amide backbone functionalities reside approximately perpendicular to the plane of the ring structure. Steric considerations and the alternating amino acid configuration force all the peptide side chains to lie on the outside of the ensemble, Part of the special issue Robert J. Silbey Festschrift. * Corresponding author. Present address: Department of Chemistry, University of Texas at Dallas, P.O. Box , Richardson TX Present address: Department of Chemistry and Applied Biosciences, ETH Zurich USI-Campus, Via Giuseppe Buffi 13, Lugano, CH-6900 Switzerland. which results in a hollow tubular core. 6 The side chains influence the self-assembly process and functionality of the nanotubes. If the peptide subunits have appropriate hydrophobic surface characteristics, they undergo rapid partitioning into lipid bilayers and spontaneously self-assemble into ion-transport-competent membrane channel structures. These nanotubes then display good channel-mediated ion-transport activity with rates exceeding 10 7 ions per second. 4 Details of the assembly process of these tubes and their mode of action as antibacterial agents are lacking. Although it is thought that peptide nanotubes operating through the carpetlike mechanism would have the greatest potential for membrane discrimination because of electrostatic interactions with various membrane constituents, 1 an understanding of the self-assembly process will help determine what factors favor this mode of action. The goal of the present study is therefore to understand elements of the self-assembly process generating the cyclic peptide nanotubes. To this end, classical MD simulations have been performed on cyclic peptide dimers composed of two cyclo [(-L-Trp-D-N-RLeu-) 4 -] 2 subunits (R ) H, CH 3 ). In the case of R ) CH 3, via selective methylation of backbone amide nitrogen functionalities, the cyclic peptide subunit is devoid of hydrogenbond donation from one face of the ring, so that it is predisposed toward an antiparallel stacked, cylindrical dimer. The capped dimer, which contains the key fundamental repeating structural motif of longer peptide nanotubes, has been studied experimentally. 5 The (R ) H) model dimer system, with uncapped nitrogen ends, is also studied. This subunit, which is known to self-assemble in a lipid bilayer environment and acts as an antibacterial agent, will be important in future studies. 3,4,13,14 Besides their behavior in a lipid bilayer environment, cyclic rings made of alternating L-Trp and D-Leu residues have also been shown to self-assemble in organosulfur self-assembled monolayers on gold film /jp057471y CCC: $ American Chemical Society Published on Web 04/19/2006
2 18966 J. Phys. Chem. B, Vol. 110, No. 38, 2006 Khurana et al. The nanotubes formed by the self-assembly of cyclic peptide subunits in a lipid bilayer are known to possess a β-sheet structure from various experimental studies. Here we show that the φ and ψ dihedral angles of a single uncapped peptide ring in a nonpolar solvent also lie in the β-sheet region of the Ramachandran plot. The thermodynamic basis for the self-assembly process comes from an experimental study performed on a cyclic peptide with a selectively N-methylated backbone that was shown to form discrete soluble cylindrical dimers in CCl 4 and CDCl 3. 5 It was found that the peptide subunit cyclo [(-L-Phe-D-N-MeAla-) 4 -] adopts a flat-ring-shaped conformation, which energetically favors ring stacking and intermolecular hydrogen-bonding interactions over the nonassembled form by kcal mol -1, depending on the solvent. The free energy of the dimerization process is accessible via steered MD (SMD) simulations using Crooks theorem 16,17 in the form of the Jarzynski equality Understanding the behavior of a single peptide subunit at the hydrophobic/hydrophilic interface is crucial to the understanding of the mechanism of formation of a nanotube in a hydrated lipid bilayer. Along these lines, we conducted MD simulations of a single uncapped peptide subunit in a nonane/water interfacial system. Irrespective of the initial starting configuration, the amphiphilic peptide subunit prefers to reside close to the interface in the nonane phase. However, when extrapolating this observation to the hydrated lipid bilayer, it should be kept in mind that the hydrophobic/hydrophilic interface in a bilayer is much more complex and broader than a simple nonane/water interface. The results of the present study on dimerization indicate that the nanotube assembly probably takes place in the hydrophobic region of the hydrated lipid bilayer. This study in a hydrophobic and a hydrophilic solvent builds the foundation for an understanding of the tube elongation process in a heterogeneous membrane environment. Simulation Details A hydrated lipid bilayer consists of two sheets of amphiphilic phospholipids oriented tail-to-tail so that the hydrocarbon region is sequestered from water exposure by the hydrophilic heads. Before attempting to study peptide nanotube self-assembly in a lipid bilayer, it was thought prudent to begin with studies of the cyclic peptides in pure solvents that mimic the two environments of the bilayer. Nonane and water were used as the hydrophobic and hydrophilic solvents, respectively. The starting peptide coordinates for the MD simulations were taken from the X-ray crystallographic coordinates provided by M. Reza Ghadiri (private communication). The X-ray coordinates were for the cyclo [(-L-Gln-D-Leu-) 4 -] 2 dimer in an antiparallel β-sheet structure. The L-Gln side chains were changed to L-Trp side chains using the ACCELRYS visualization package Insight II. Alternating D- and L-conformations of the backbone constrain the β-sheet structure so that only homochiral residues can form the cross-strand β-sheet nearest pairs. 21 MD simulations were carried out using the program NAMD version 2.5b1 22 and the CHARMM force field. 23 Periodic boundary conditions were used in three dimensions. The Nosé-Hoover Langevin piston pressure control method was used to maintain the pressure of the cell at 1 atm during isobaric simulation runs, and, in all the simulations, Langevin dynamics was used to control the temperature at 300 K. A time step of 2 fs was used. Nonbonded interactions were calculated every time step, and full electrostatic interactions were calculated every two time steps. Bonds between hydrogens and heavy atoms were constrained to their equilibrium value by means of the SHAKE/ RATTLE algorithm. 24,25 Long-range electrostatic forces were taken into account using the particle mesh Ewald approach with a real space cutoff of 12 Å. 26,27 The water molecules were described using the TIP3P model. 28 The following four MD simulations (designated by Roman numerals I-IV) were performed on the cyclic peptide dimers: (I) the capped dimer cyclo [(-L-Trp-D-N-MeLeu-) 4 -] 2 in 3024 water molecules; (II) the uncapped dimer cyclo [(-L-Trp- D-Leu-) 4 -] 2 in 3031 water molecules; (III) the capped dimer cyclo [(-L-Trp-D-N-MeLeu-) 4 -] 2 in 300 nonane molecules; and (IV) the uncapped dimer cyclo [(-L-Trp-D-Leu-) 4 -] 2 in 293 nonane molecules. A box consisting of 316 molecules of nonane was created and equilibrated for 1 ns using the isothermalisobaric (NPT) ensemble and allowing isotropic cell fluctuations (maintaining a constant ratio of x:y:z). The box dimensions converged to Å after about 100 ps. The respective dimers were then added to this box, resulting in the removal of a few nonane molecules to avoid overlap. All the systems for cases I-IV were equilibrated for 1 ns by constraining the CR atoms of the backbone to their initial positions, using the NPT ensemble, and allowing isotropic cell fluctuations. This allowed equilibration of the side chains and the solvent molecules. The box sizes converged after a few picoseconds. After 1 ns of equilibration, the constraints on the CR atoms were removed, and the NVT ensemble was used for production runs. The box dimensions for cases I and II were Å, while, for case III the box dimensions were Å, and for case IV they were Å. In addition, we conducted an MD simulation of one uncapped ring in 303 nonane molecules for 3 ns using the NPT ensemble with isotropic cell fluctuations. The free energy of dimerization was studied for the capped dimer in nonane (case III) in the following manner. The (x,y,z) coordinates of the center of mass of one ring were kept fixed. The center of mass of the second ring was pulled away from that of the first ring in the direction parallel to the intersubunit hydrogen bonding (z) direction. Force was applied to all the atoms of the rings to constrain the respective centers of mass. The (x,y) coordinates of the pulled ring were not constrained. Initial conditions were drawn from constrained MD runs of 1 ns at separation distances of 6.5, 7.5, 8.5,..., and 12.5 Å between the z coordinates of the two rings. Every 50 ps, a frame was saved to initiate a total of 20 SMD pulling runs for each separation distance. In this fashion, a total of 140 pulling runs corresponding to pulling for 1 Å each were launched, resulting in a total pulling distance of 7 Å. To join the different freeenergy profiles at the overlapping values of z, the curve that had better sampling (more data points) for that region was chosen. At the maximum separation distance of 13.5 Å, the backbones of the two rings are not interacting with each other. In every case, the reaction coordinate was linked to the pulling coordinate with a harmonic spring of stiffness 250 kcal/mol/å 2 ( pn Å). The force constant was chosen such that it was large enough to ensure small deviation of the reaction coordinate from the constraint position. The pulling velocity in each case was 5 Å/ns. The free energy calculation was based on Jarzynski s equality and the stiff-spring approximation described by Park and Schulten. 29 The free energy was calculated based on the first order cumulant expansion formula, the second order cumulant expansion formula, and the full exponential formula. The convergence of the free energy curves derived from the second order cumulant and the full exponential formula was used as an indication that the work distribution is
3 Self-Assembling Cyclic Peptides J. Phys. Chem. B, Vol. 110, No. 38, Gaussian. 29,30 For the regions corresponding to poor convergence of the two curves, further equilibration was performed for 1 ns under constrained MD conditions, and 20 snapshots were taken every 50 ps as the initial conditions for SMD runs with the same velocity as that used for the previous runs (5 Å/ns). The free energy analysis was then performed on the data collected from 40 trajectories for the same pulling length. This did not improve the convergence of the free energy curves derived from the second order cumulant and the full exponential formula. However, when the velocity of pulling was lowered to onequarter of the original velocity (1.25 Å/ns), the convergence improved significantly. Hence the error, which was a systematic one arising from the non-gaussian behavior of the probability densities of work, was decreased significantly by pulling with a lower velocity. The calculation in the regions where the convergence was good at the original pulling velocity was not repeated. To study the behavior of the uncapped ring at the nonane/ water interface, a box containing 126 nonane molecules and 1598 water molecules was created. The box was equilibrated for 1 ns using the NPT ensemble, allowing isotropic cell fluctuations. After a few picoseconds, the cell dimensions converged to about Å. The equilibration was followed by two 4 ns simulations: one starting with the entire uncapped peptide ring immersed in water and the other inside nonane. During these simulations, the dimensions of the unit cell were kept constant in the x-y plane while allowing fluctuations along the z-axis (the interfacial direction) using Langevin piston pressure control with a pressure of 1 atm. Results and Discussion The present MD simulations indicate that the uncapped and the capped dimers stay together by means of intersubunit hydrogen bonds in nonpolar solvent, whereas they fall apart in polar solvent. Figure 1a shows an uncapped ring, while Figure 1b shows a capped ring. Figure 1c,d shows the uncapped dimer in nonane after 1 ns and 10 ns of MD simulations, respectively. Figure 1e,f shows the uncapped dimer in water at the end of 1 ns and 10 ns of MD, respectively. In both the cases, the CR atoms of the backbones were fixed to their initial positions during the first nanosecond of MD. Cases I and III involve capped dimers in polar and nonpolar solvents, respectively, and correspond to systems accessible experimentally. Here, the present results are consistent with the experimental data. Simulation cases II and IV represent pure dimeric systems that are inaccessible to experiment: there are only two peptide rings in the simulation unit cell. Experimentally, when self-assembly occurs, not only would the dimer form, but the trimer and longer assemblies would also be present. The 1 H NMR and IR studies by Ghadiri et al. 5 showed that cyclo [(-L-Phe-D-N-MeAla-) 4 -] displays several conformational isomers in polar solvents, such as deuterated methanol or dimethyl sulfoxide (DMSO), which slowly interconvert as a result of the cis-trans isomerization of secondary amides. This inhibits the dimerization process in polar solvent for the capped dimer. The factors that prevent the dimerization of the capped subunit in polar solvent should also inhibit the dimerization of the uncapped subunit in polar solvent. Experimental studies have confirmed the self-assembly of the peptide subunit studied here in a lipid bilayer environment. This leads us to predict that the uncapped dimer should be stable in a nonpolar solvent while being unstable in a polar solvent. Our results agree with this prediction. Experimental studies have indicated that the peptide nanotubes and the dimer formed by the self-assembly of cyclic peptide rings possess a β-sheet structure. Figure 2a shows the Ramachandran plot obtained from the MD simulation of a single uncapped peptide ring in nonane. This plot clearly shows that the φ and ψ angles of all the residues lie in the β-sheet region. The signs of the φ s and ψ s for the D-amino acids are simply the inverse of what they should be for the L-amino acids. This tells us that the geometry of the single peptide ring dictates the preference for a β-sheet structure, even when it is not hydrogen bonded to another subunit. Figure 2b shows the radius of gyration of the backbone atoms of the ring versus time. The ring has a flat conformation in the beginning of the simulation. The radius of gyration stays close to the initial value throughout the simulation, indicating the ring stays flat in nonane. Figure 3 shows the time evolution of the number of hydrogen bonds between the subunits constituting the dimer under different conditions, following 1 ns of constrained MD. Concerning the definition of a hydrogen bond, we say a hydrogen bond exists if the distance between the donor and acceptor atoms is less than 3.4 Å and the angle D-H-A is greater than 130, where D represents the donor atom, H represents the hydrogen atom and A represents the acceptor atom. The number of hydrogen bonds shown is the average number of hydrogen bonds calculated every 100 ps. The maximum possible number of hydrogen bonds between the two subunits is eight. As shown in the figure, the number of intersubunit hydrogen bonds remains close to eight for the uncapped dimer in nonane. Although the number of hydrogen bonds for the capped dimer in nonane fluctuates more than for the uncapped dimer in nonane, at least six intersubunit hydrogen bonds always exist between the two rings during the entire 9 ns length of the trajectory. The number of hydrogen bonds during the initial 1 ns of constrained MD is not plotted in the figure, as it is expected to be eight for all the systems. The quantum mechanical studies by Chen et al. 10 showed that the conformation for the optimized cyclo [(-L-Phe- D-N-MeAla-) 4 -] is distorted to a certain extent compared to that for cyclo [(-L-Phe-D-Ala-) 4 -]. It was observed that the angles of NH and CO with respect to the ring plane in the capped ring depart farther than that found for the uncapped ring. A departure of ( 10 from the normal value of 180 for ω was also observed. Thus, distortion of the capped ring compared to that of the uncapped ring could be the possible reason for the slight reduction in the number of intersubunit hydrogen bonds for the capped dimer versus the uncapped dimer. Figure 3 also shows that the number of hydrogen bonds between the two rings falls to zero after about 1 ns of unconstrained MD for the capped dimer in water. Although it takes much longer for the hydrogen bonds to break between the two monomers of the uncapped dimer in water, the monomeric state is more stable as the dimer breaks at the end of 10 ns. We stop the simulation at the end of 10 ns, even though the two rings have not completely fallen apart, as the breaking of the remaining hydrogen bonds is just a matter of running for more time. One possible reason for the faster breaking of the intersubunit hydrogen bonds in the capped dimer, compared to those in the uncapped dimer, could be the conformational distortion of the capped ring compared to the uncapped ring as explained previously. The exposure of the hydrophobic methyl groups of the capped dimer to the hydrophilic water molecules could be another reason for the higher instability of the capped dimer. Figure 4 shows the number of hydrogen bonds between the backbone atoms of the peptide rings and the water molecules using the same definition of the hydrogen bond as previously defined for intersubunit hydrogen bonds. Here again, the number of hydrogen bonds shown is the average number of hydrogen bonds calculated every 100
4 18968 J. Phys. Chem. B, Vol. 110, No. 38, 2006 Khurana et al. Figure 1. Representative configurations taken from MD simulations of cyclic peptide dimers: (a) one uncapped ring; (b) one capped ring. Panel a shows the hydrogen atoms of the N-H groups of the backbone, while panel b shows the hydrogen atoms of the Leu N-H groups substituted by methyl groups. (c) The uncapped dimer in nonane at t ) 1 ns. (d) The uncapped dimer in nonane at t ) 10 ns. Panels c and d show that the planar rings stay close to each other in the nonpolar solvent. Intersubunit hydrogen bonds contribute to the stability of the dimer. (e) The uncapped dimer in water at t ) 1 ns. (f) The uncapped dimer in water at t ) 10 ns. Panel e shows that the rings exist in the form of a dimer after 1 ns of constrained MD during which the backbone CR atoms are fixed, while panel f shows that the intersubunit hydrogen bonds break at the end of 10 ns. Carbon atoms are green, oxygen atoms are red, and nitrogen atoms are blue. The cyclic peptide backbones are rendered as tubes, and the side chains are shown as stick and ball. The solvent molecules are shown in purple. The hydrogen bonds between the two rings are shown as dashed black lines. In panels a and b, only the hydrogen atoms of the N-H groups of the backbones are shown; in all other cases, all hydrogen atoms have been omitted for clarity. The images were made using VMD. 33 ps. In the case of capped dimer, only the uncapped side of the ring can participate in hydrogen bonding with water, whereas, for the uncapped ring, both faces of the ring can take part in hydrogen bonding with water. However, we measure the number of hydrogen bonds formed only by that side of the ring that takes part in intersubunit hydrogen bonding. This gives eight hydrogen-bonding sites on each ring: four N-H sites and four C-O sites. Each backbone oxygen atom can participate in two hydrogen bonds with water molecules. Hence, each ring can form a maximum of 12 hydrogen bonds with water if all the sites participate in hydrogen bonding with water. From Figures 3 and 4 we can clearly see that, as the number of intersubunit hydrogen bonds goes down, the number of hydrogen bonds with water increases for both the capped and the uncapped dimers in water. The competition between intersubunit hydrogen bonding and hydrogen bonding with the solvent has also been discussed in another study. 31 In that study, it was observed that, under rigorously anhydrous conditions, the association constant for cyclo [(-L-Phe-D-N-MeAla-) 4 -] 2 in deuteriochloroform approxi-
5 Self-Assembling Cyclic Peptides J. Phys. Chem. B, Vol. 110, No. 38, Figure 3. Time evolution of the average number of intersubunit hydrogen bonds in the four MD simulations. A hydrogen bond exists between two atoms if the distance between the donor and acceptor atoms is less than 3.4 Å and the angle D-H-A is greater than 130, where D represents the donor atom, H represents the hydrogen atom, and A represents the acceptor atom. The figure shows that the number of hydrogen bonds remains almost constant for both the uncapped dimer and the capped dimer in nonane. All the hydrogen bonds break after about 2 ns for the capped dimer in water, while it takes about 4.5 ns for most of the intersubunit hydrogen bonds to break for the uncapped dimer in water. The number of hydrogen bonds during the first nanosecond of constrained MD is not shown. Figure 2. (a) Ramachandran plot for a single uncapped peptide ring in nonane. The dihedrals for L-Trp residues are shown with diamondshaped symbols, while the dihedrals for D-Leu residues are shown with crosses. The plot clearly shows that the ring has a β-sheet structure. (b) Radius of gyration of the backbone atoms of one uncapped ring in nonane. The radius of gyration stays close to the initial value throughout the simulation, indicating that the ring stays in a flat conformation in nonane. The range of the y axis is chosen for easy comparison with Figure 5. mately doubled, suggesting competition by water for intermolecular hydrogen-bonding sites. Weak low-energy bands were observed in FT-IR spectra of peptides that do not form dimers, presumably arising from intramolecular hydrogen bonding. We measured the distance between the centers of mass of the backbone atoms of the two rings. The average intersubunit distance over the last 5 ns was found to be 4.78 ( 0.08 Å for the uncapped dimer in nonane and 4.96 ( 0.10 Å for the capped dimer in nonane. These values agree well with the values of Å obtained by electron diffraction patterns, X-ray analyses, and IR measurements. 2,32 On the other hand, the average intersubunit distance calculated during the last 5 ns of the MD simulations was measured to be about 6.4 Å for the uncapped dimer in water and Å for the capped dimer in water. Thus, the simulations demonstrate that both the capped and the uncapped dimers remain stable in a hydrophobic solvent. This result agrees with the 1 H NMR and IR results obtained by Ghadiri et al. 5 for the capped dimer. They observed that cyclo [(-L-Phe-D-N-MeAla-) 4 -] displays several conformational isomers in polar solvents such as deuterated methanol or DMSO, which slowly interconvert as a result of cis-trans isomerization. However, in nonpolar solvents such as carbon tetrachloride Figure 4. Hydrogen bonds between water molecules and peptide rings. Upon comparing Figures 3 and 4, we can clearly see that, as the number of intersubunit hydrogen bonds decreases, the number of hydrogen bonds with water molecules increases. The definition of hydrogen bonds used is the same as that for intersubunit hydrogen bonds. (CCl 4 ), the peptide exists in an all-trans conformation with a flat-ring-shaped backbone, which is in dynamic equilibrium with the expected cylindrical dimer. Regarding the uncapped dimer, as discussed previously, we expected the dimer to be stable in nonpolar solvent because a nanotube made of the peptide subunit is stable in a lipid bilayer. Our observation here agrees with this prediction. Figure 5 shows the radius of gyration of the backbone atoms of the ring versus time. The radius of gyration should be the maximum for the ring in a flat conformation. As seen clearly in the figure, the radius of gyration stays close to the initial value, corresponding to a flat conformation for both the capped and the uncapped dimers in nonane. In Figure 2b, it was seen that the backbone of a single uncapped ring also stays in a flat conformation in nonane. Figure 5 shows that the ring does not stay flat in water after the intersubunit hydrogen bonds break. Hence our results agree with the experimental results.
6 18970 J. Phys. Chem. B, Vol. 110, No. 38, 2006 Khurana et al. Figure 5. Radius of gyration of the backbone atoms of one ring of the dimer studied under four different conditions. The radius of gyration should be the maximum, corresponding to a flat conformation of the ring. It is clear from the figure that, as the intersubunit hydrogen bonds break and the rings separate from each other in water, the radius of gyration decreases and the ring does not stay in a flat conformation. Figure 7. Change in free energy associated with pulling one capped ring away from the other capped ring of the stable dimer in nonane. The free energy is calculated by using the second-order cumulant approximation of Jarzynski s equality and the full exponential formula. Figure 6. The z coordinates of the centers of mass (c.o.m) of the two rings vs time. The blue line shows the z coordinate of the fixed ring 1. The red line shows the position of the dummy atom attached to the z coordinate of the pulled ring 2 (reaction coordinate). The z trajectory of the c.o.m of ring 2 (pulling coordinate) is shown in black for regions where the velocity of pulling was 1.25 Å/ns and green for regions where the velocity of pulling was 5 Å/ns. The complete pulling trajectory was divided into seven sections. Each section consisted of 20 pulling runs. Data from only one trajectory for each section are shown as an example. As seen from the figure, the pulling coordinate closely follows the reaction coordinate for ring 2. Figure 6 shows the z coordinate of the center of mass of the fixed ring 1 versus time and the z coordinate of the center of mass of the pulled ring 2 versus time. As seen in the figure, the z coordinate of ring 2 closely follows the reaction coordinate, that is, the z coordinate of the dummy atom attached to it. As explained in the simulation details, the whole pulling trajectory was divided into seven sections, each consisting of 20 pulling runs. The plots shown in Figure 6 are only for one pulling run chosen independently for each section. The free energy profile of the distance between the two capped rings in nonane as a function of the z coordinate of the pulled ring is shown in Figure 7. As explained in the simulation details, for certain sections, 40 pulling runs were performed and/or lower pulling velocities were used. Plots showing the effect of the number of pulling runs and the velocities on the free energy profile are available in the Supporting Information for the interested reader. The Figure 8. (a) Starting configuration for the uncapped peptide ring in the water phase and (b) starting configuration for the uncapped peptide ring in the nonane phase. Carbon atoms are green, oxygen atoms are red, and nitrogen atoms are blue. The cyclic peptide backbones are rendered as tubes, and the side chains are shown as stick and ball. Nonane molecules are shown as thin purple lines, and water molecules are shown as purple spheres. Hydrogen atoms have been omitted for clarity. The images were made using VMD. 33 availability of experimental data for a capped dimer was the reason for our choice of the capped dimer for the free energy calculation, although both the capped dimer and the uncapped dimer are stable in nonane. The initial position of the pulled ring corresponds to a distance of 6.5 Å between the z coordinates of the centers of mass of the two rings. After pulling for about 2 Å, the backbones of the two rings stop interacting with each other. All the intersubunit hydrogen bonds are broken at this point. Further changes in the free energy occur as a result of the formation and disruption of hydrogen bonds between the Trp side chains of one ring (either ring 1 or ring 2) and the
7 Self-Assembling Cyclic Peptides J. Phys. Chem. B, Vol. 110, No. 38, energy difference between the initial state of the two rings when they are hydrogen bonded to each other and when they are separated is 7 ( 1 kcal/mol, derived from the second order cumulant approximation and the full exponential formula. This result is reasonable, given the value of kcal/mol obtained by Ghadiri et al., 5 with a dimer made up of different amino acid residues and a different solvent. As mentioned by Kobayashi et al., 21 it should be expected that the changes in the polarity of the medium may produce significant and additive energetic contributions, emanating from cross-strand interactions between side chains, toward stabilizing a particular β-sheet arrangement. To further understand the process of self-assembly of peptide rings in a lipid bilayer environment, we studied the behavior of a single ring in a box with a nonane/water interface. Figure 9a shows the number density profiles obtained from a 1 ns simulation of a pure nonane/water interface along the direction normal to the interface (z direction). A single uncapped peptide ring was immersed completely in the water phase with the center of the peptide backbone at a distance of about 6 Å from the nonane phase, as shown in Figure 8a. Figure 9b shows the density profiles obtained from the last 2 ns of the 4 ns MD simulation of this system along the z direction. In another simulation, a single uncapped peptide ring was immersed completely in the nonane phase with the center of the peptide backbone at a distance of about 9 Å from the water phase as shown in Figure 8b. Figure 9c shows the density profiles obtained from the last 2 ns of the 4 ns MD simulation of this system. From Figures 9b and 9c it is clear that the peptide ring prefers to sit close to the interface in the nonane phase, irrespective of the initial configuration. This observation can be explained as a result of the amphiphilic nature of the peptide ring. This tells us that a single peptide ring in a lipid bilayer environment should prefer to sit close to the hydrophobic/ hydrophilic interface. However, the hydrophobic/hydrophilic interface in a hydrated lipid bilayer is not as well-defined as a pure nonane/water interface. It can be said from our simulation results that the peptide ring behaves like a surfactant at a nonane/ water interface. The lipid molecules also possess surfactant properties, with a hydrophilic headgroup and a hydrophobic tail. The behavior of the peptide ring in the presence of another surfactant molecule in a lipid bilayer might be different than its behavior at a pure hydrophobic/hydrophilic interface. Upon comparing Figure 9a-c, we can clearly see that the water phase extends deeper into the nonane phase in panels b and c compared to that in panel a. These water molecules entering the nonane phase are associated with the hydrophilic sites on the peptide ring by hydrogen bonding. Figure 9. Number density profiles for the different components of the system along the normal to the nonane/water interface: (a) the pure nonane/water interface; (b) the density profiles obtained from the last 2 ns of the 4 ns simulation corresponding to the initial condition of a single uncapped peptide ring completely immersed in the water phase; and (c) the density profiles obtained from the last 2 ns of the 4 ns simulation corresponding to the initial condition of a single uncapped peptide ring completely immersed in the nonane phase. As seen clearly from panels b and c, the ring prefers to reside close to the interface in the nonane phase, irrespective of the initial configuration. Also, more water molecules penetrate the nonane phase in panels b and c compared to those in panel a (see the insets). These water molecules are associated with the hydrophilic sites on the peptide ring via hydrogen bonds. backbone of the other ring. A figure showing the intersubunit hydrogen bonds during the pulling simulation is available in the Supporting Information for the interested reader. The free Conclusions This study characterizes the behavior of the dimers formed from cyclo [(-L-Trp-D-Leu) 4 ] 2 and cyclo [(-L-Trp-D-N- MeLeu-) 4 -] 2 in polar and nonpolar environments under the CHARMM force field. The observed stability of the dimers and the flat conformation of the peptide ring in a hydrophobic solvent matches well with the experimental results. The dimers are observed to be unstable in a hydrophilic solvent. We also observe that a single peptide ring prefers to sit close to the nonane/water interface. This suggests that the self-assembly of the peptide rings to form a nanotube starts after the rings enter the hydrophobic tail region of a lipid bilayer. We also show that the φ and ψ angles of a single uncapped peptide ring in nonane lie in the β-sheet region of the Ramachandran plot. The potential of mean force was computed as a function of the
8 18972 J. Phys. Chem. B, Vol. 110, No. 38, 2006 Khurana et al. distance between the two rings using SMD and Jarzynski s equality. This simulation study is a necessary prelude to the assembly of the complete nanotube. The understanding of the mechanism of the formation of these nanotubes will likely cast light on their mode of action as antibacterial agents and on other applications such as molecular inclusion devices and drugdelivery agents. Acknowledgment. The authors thank M. Reza Ghadiri for providing the coordinates for the dimer and Ivaylo Ivanov for useful discussions. We acknowledge the Pittsburgh Supercomputing Center through the Pennsylvania research initiative for computer time used for these simulations. This work was funded by the National Institute of Health, Grant GM Supporting Information Available: Plots showing the effect of the number of pulling runs and the velocities on free energy profiles and a figure showing the intersubunit hydrogen bonds during the pulling run. This material is available free of charge via the Internet at References and Notes (1) Lopez, S. F.; Kim, H. S.; Choi, E. C.; Delgado, M.; Granja, J. R.; Khasanov, A.; Kraehenbuehl, K.; Long, G.; Weinberger, D. A.; Wilcoxen, K. M.; Ghadiri, M. R. Nature 2001, 412, 452. (2) Ghadiri, M. R.; Granja, J. R.; Milligan, R. A.; McRee, D. E.; Khazanovich, N. Nature 1993, 366, 324. (3) Kim, H. S.; Hartgerink, J. D.; Ghadiri, M. R. J. Am. Chem. Soc. 1998, 120, (4) Ghadiri, M. R.; Granja, J. R.; Buehler, L. K. Nature 1994, 369, 301. (5) Ghadiri, M. R.; Kobayashi, K.; Granja, J. R.; Chadha, R. K.; McRee, D. E. Angew. Chem., Int. Ed. Engl. 1995, 34, 93. (6) Ghadiri, M. R. AdV. Mater. 1995, 7, 675. (7) Hartgerink, J. D.; Clark, T. D.; Ghadiri, M. R. Chem.sEur. J. 1998, 4, (8) Carloni, P.; Andreoni, W.; Parinello, M. Phys. ReV. Lett. 1997, 106, (9) Lewis, J. P.; Pawley, N. H.; Sankey, O. F. J. Phys. Chem. B. 1997, 101, (10) Chen, G.; Su, S.; Liu, R. J. Phys. Chem. B. 2002, 106, (11) Engels, M.; Bashford, D.; Ghadiri, M. R. J. Am. Chem. Soc. 1995, 117, (12) Tarek, M.; Maigret, B.; Chipot, C. Biophys. J. 2003, 85, (13) Quesada, J. S.; Isler, M. P.; Ghadiri, M. R. J. Am. Chem. Soc. 2002, 124, (14) Quesada, J. S.; Kim, H. S.; Ghadiri, M. R. Angew. Chem., Int. Ed. Engl. 2001, 40, (15) Motesharei, K.; Ghadiri, M. R. J. Am. Chem. Soc. 1997, 119, (16) Crooks, G. E. Phys. ReV. E1999, 60, (17) Collin, D.; Ritort, F.; Jarzynski, C.; Smith, S. B.; Tinoco, I., Jr.; Bustamante, C. Nature 2005, 437, 231. (18) Jarzynski, C. Phys. ReV. Lett. 1997, 78, (19) Jensen, M. O.; Park, S.; Tajkhorshid, E.; Schulten, K. Proc. Natl. Acad. Sci. U.S.A. 2002, 99, (20) Amaro, R.; Tajkhorshid, E.; Schulten, Z. L. Proc. Natl. Acad. Sci. U.S.A. 2003, 100, (21) Kobayashi, K.; Granja, J. R.; Ghadiri, M. R. Angew. Chem., Int. Ed. Engl. 1995, 34, 95. (22) Kale, L.; Skeel, R.; Bhandarkar, M.; Brunner, R.; Gursoy, A.; Krawetz, N.; Phillips, J.; Shinozaki, A.; Varadarajan, K.; Schulten, K. J. Comput. Phys. 1999, 151, 283. (23) MacKerell, A. D.; Bashford, D., Jr.; Bellott, M.; Dunbrack, R. L.; Evanseck, J. D., Jr.; Field, M. J.; Fischer, S.; Gao, J.; Guo, H.; Ha, S.; Joseph-McCarthy, D.; Kuchnir, L.; Kuczera, K.; Lau, F. T. K.; Mattos, C.; Michnick, S.; Ngo, T.; Nguyen, D. T.; Prodhom, B.; Reiher, W. E.; Roux, B., III; Schlenkrich, M.; Smith, J. C.; Stote, R.; Straub, J.; Watanabe, M.; Wiórkiewicz-Kuczera, J.; Yin, D.; Karplus, M. J. Phys. Chem. B. 1998, 102, (24) Ryckaert, J. P.; Ciccotti, G.; Berendson, H. J. C. J. Comput. Phys. 1977, 23, 327. (25) Andersen, H. C. J. Comput. Phys. 1983, 52, 24. (26) Darden, T.; York, D.; Pederson, L. J. Chem. Phys. 1993, 98, (27) Essmann, U.; Perera, L.; Berkowitz, M. L.; Darden, T.; Lee, H.; Pederson, L. G. J. Chem. Phys. 1995, 103, (28) Jorgensen, W. L.; Chandrasekhar, J.; Madura, J. D.; Impey, R. W.; Klein, M. L. J. Chem. Phys. 1983, 79, 926. (29) Park, S.; Khalilli-Araghi, F.; Tajkhorshid, E.; Schulten, K. J. Chem. Phys. 2003, 119, (30) Hummer, G. J. Chem. Phys. 2001, 114, (31) Clark, T. D.; Buriak, J. M.; Kobayashi, K.; Isler, M. P.; McRee, D. E.; Ghadiri, M. R. J. Am. Chem. Soc. 1998, 120, (32) Hartgerink, J. D.; Granja, J. R.; Milligan, R. A.; Ghadiri, M. R. J. Am. Chem. Soc. 1996, 118, 43. (33) Humphrey, W.; Dalke, A.; Schulten, K. J. Mol. Graphics 1996, 14, 33.
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein
 Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Improved Resolution of Tertiary Structure Elasticity in Muscle Protein Jen Hsin and Klaus Schulten* Department of Physics and Beckman Institute, University of Illinois at Urbana-Champaign, Urbana, Illinois
Molecular Dynamics Investigation of the ω-current in the Kv1.2 Voltage Sensor Domains
 Molecular Dynamics Investigation of the ω-current in the Kv1.2 Voltage Sensor Domains Fatemeh Khalili-Araghi, Emad Tajkhorshid, Benoît Roux, and Klaus Schulten Department of Physics, Department of Biochemistry,
Molecular Dynamics Investigation of the ω-current in the Kv1.2 Voltage Sensor Domains Fatemeh Khalili-Araghi, Emad Tajkhorshid, Benoît Roux, and Klaus Schulten Department of Physics, Department of Biochemistry,
Gemini Surfactants at the Air/Water Interface: A Fully Atomistic Molecular Dynamics Study
 22136 J. Phys. Chem. B 2006, 110, 22136-22142 Gemini Surfactants at the Air/Water Interface: A Fully Atomistic Molecular Dynamics Study Ekta Khurana,* Steven O. Nielsen, and Michael L. Klein Center for
22136 J. Phys. Chem. B 2006, 110, 22136-22142 Gemini Surfactants at the Air/Water Interface: A Fully Atomistic Molecular Dynamics Study Ekta Khurana,* Steven O. Nielsen, and Michael L. Klein Center for
W tot (z) = W (z) k B T ln e i q iφ mp(z i )/k B T, [1]
![W tot (z) = W (z) k B T ln e i q iφ mp(z i )/k B T, [1] W tot (z) = W (z) k B T ln e i q iφ mp(z i )/k B T, [1]](/thumbs/87/95587399.jpg) Supporting Text Simulation Details. Simulations were carried out with the program charmm (1) using the PARAM27 force field with standard lipid (2), protein (3), and TIP3P water model (4) parameters. The
Supporting Text Simulation Details. Simulations were carried out with the program charmm (1) using the PARAM27 force field with standard lipid (2), protein (3), and TIP3P water model (4) parameters. The
Supporting Information. for An atomistic modeling of F-actin mechanical responses and determination of. mechanical properties
 Supporting Information for An atomistic modeling of F-actin mechanical responses and determination of mechanical properties Authors: Si Li 1, Jin Zhang 2, Chengyuan Wang 1, Perumal Nithiarasu 1 1. Zienkiewicz
Supporting Information for An atomistic modeling of F-actin mechanical responses and determination of mechanical properties Authors: Si Li 1, Jin Zhang 2, Chengyuan Wang 1, Perumal Nithiarasu 1 1. Zienkiewicz
Pressure-Induced Water Transport in Membrane Channels Studied by Molecular Dynamics
 154 Biophysical Journal Volume 83 July 2002 154 160 Pressure-Induced Water Transport in Membrane Channels Studied by Molecular Dynamics Fangqiang Zhu, Emad Tajkhorshid, and Klaus Schulten Beckman Institute,
154 Biophysical Journal Volume 83 July 2002 154 160 Pressure-Induced Water Transport in Membrane Channels Studied by Molecular Dynamics Fangqiang Zhu, Emad Tajkhorshid, and Klaus Schulten Beckman Institute,
Universal Repulsive Contribution to the. Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information
 Universal Repulsive Contribution to the Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information B. Shadrack Jabes, Dusan Bratko, and Alenka Luzar Department of Chemistry,
Universal Repulsive Contribution to the Solvent-Induced Interaction Between Sizable, Curved Hydrophobes: Supporting Information B. Shadrack Jabes, Dusan Bratko, and Alenka Luzar Department of Chemistry,
Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality
 JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 6 8 AUGUST 2003 Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality Sanghyun Park and Fatemeh Khalili-Araghi Beckman
JOURNAL OF CHEMICAL PHYSICS VOLUME 119, NUMBER 6 8 AUGUST 2003 Free energy calculation from steered molecular dynamics simulations using Jarzynski s equality Sanghyun Park and Fatemeh Khalili-Araghi Beckman
Supporting Information
 Supporting Information Comparative Study of Materials-Binding Peptide Interactions with Gold and Silver Surfaces and Nanostructures: A Thermodynamic Basis for Biological Selectivity of Inorganic Materials
Supporting Information Comparative Study of Materials-Binding Peptide Interactions with Gold and Silver Surfaces and Nanostructures: A Thermodynamic Basis for Biological Selectivity of Inorganic Materials
The Dominant Interaction Between Peptide and Urea is Electrostatic in Nature: A Molecular Dynamics Simulation Study
 Dror Tobi 1 Ron Elber 1,2 Devarajan Thirumalai 3 1 Department of Biological Chemistry, The Hebrew University, Jerusalem 91904, Israel 2 Department of Computer Science, Cornell University, Ithaca, NY 14853
Dror Tobi 1 Ron Elber 1,2 Devarajan Thirumalai 3 1 Department of Biological Chemistry, The Hebrew University, Jerusalem 91904, Israel 2 Department of Computer Science, Cornell University, Ithaca, NY 14853
implications for different reactivity of identical subunits
 Supplementary Material Finding molecular dioxygen tunnels in homoprotocatechuate 2,3-dioxygenase: implications for different reactivity of identical subunits Liang Xu 1, 2, Weijie Zhao 3, Xicheng Wang
Supplementary Material Finding molecular dioxygen tunnels in homoprotocatechuate 2,3-dioxygenase: implications for different reactivity of identical subunits Liang Xu 1, 2, Weijie Zhao 3, Xicheng Wang
Approach to Thermal Equilibrium in Biomolecular
 Approach to Thermal Equilibrium in Biomolecular Simulation Eric Barth 1, Ben Leimkuhler 2, and Chris Sweet 2 1 Department of Mathematics Kalamazoo College Kalamazoo, Michigan, USA 49006 2 Centre for Mathematical
Approach to Thermal Equilibrium in Biomolecular Simulation Eric Barth 1, Ben Leimkuhler 2, and Chris Sweet 2 1 Department of Mathematics Kalamazoo College Kalamazoo, Michigan, USA 49006 2 Centre for Mathematical
Supporting Material for. Microscopic origin of gating current fluctuations in a potassium channel voltage sensor
 Supporting Material for Microscopic origin of gating current fluctuations in a potassium channel voltage sensor J. Alfredo Freites, * Eric V. Schow, * Stephen H. White, and Douglas J. Tobias * * Department
Supporting Material for Microscopic origin of gating current fluctuations in a potassium channel voltage sensor J. Alfredo Freites, * Eric V. Schow, * Stephen H. White, and Douglas J. Tobias * * Department
Molecular Dynamics Simulation of a Nanoconfined Water Film
 Molecular Dynamics Simulation of a Nanoconfined Water Film Kyle Lindquist, Shu-Han Chao May 7, 2013 1 Introduction The behavior of water confined in nano-scale environment is of interest in many applications.
Molecular Dynamics Simulation of a Nanoconfined Water Film Kyle Lindquist, Shu-Han Chao May 7, 2013 1 Introduction The behavior of water confined in nano-scale environment is of interest in many applications.
Computational Predictions of 1-Octanol/Water Partition Coefficient for Imidazolium based Ionic Liquids.
 Computational Predictions of 1-Octanol/Water Partition Coefficient for Imidazolium based Ionic Liquids. Ganesh Kamath,* a Navendu Bhatnagar b, Gary A. Baker a, Sheila N. Baker c and Jeffrey J. Potoff b
Computational Predictions of 1-Octanol/Water Partition Coefficient for Imidazolium based Ionic Liquids. Ganesh Kamath,* a Navendu Bhatnagar b, Gary A. Baker a, Sheila N. Baker c and Jeffrey J. Potoff b
The dependence of hydrophobic interactions on the solute size
 The dependence of hydrophobic interactions on the solute size Q. Sun * Key Laboratory of Orogenic Belts and Crustal Evolution, Ministry of Education, The School of Earth and Space Sciences, Peking University
The dependence of hydrophobic interactions on the solute size Q. Sun * Key Laboratory of Orogenic Belts and Crustal Evolution, Ministry of Education, The School of Earth and Space Sciences, Peking University
Biophysics II. Hydrophobic Bio-molecules. Key points to be covered. Molecular Interactions in Bio-molecular Structures - van der Waals Interaction
 Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
Biophysics II Key points to be covered By A/Prof. Xiang Yang Liu Biophysics & Micro/nanostructures Lab Department of Physics, NUS 1. van der Waals Interaction 2. Hydrogen bond 3. Hydrophilic vs hydrophobic
arxiv: v1 [physics.chem-ph] 8 Mar 2010
![arxiv: v1 [physics.chem-ph] 8 Mar 2010 arxiv: v1 [physics.chem-ph] 8 Mar 2010](/thumbs/88/116239288.jpg) arxiv:1003.1678v1 [physics.chem-ph] 8 Mar 2010 A new battery-charging method suggested by molecular dynamics simulations Ibrahim Abou Hamad 1,, M. A. Novotny 1,2, D. Wipf 3, and P. A. Rikvold 4 1 HPC 2,
arxiv:1003.1678v1 [physics.chem-ph] 8 Mar 2010 A new battery-charging method suggested by molecular dynamics simulations Ibrahim Abou Hamad 1,, M. A. Novotny 1,2, D. Wipf 3, and P. A. Rikvold 4 1 HPC 2,
Introduction to Comparative Protein Modeling. Chapter 4 Part I
 Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Introduction to Comparative Protein Modeling Chapter 4 Part I 1 Information on Proteins Each modeling study depends on the quality of the known experimental data. Basis of the model Search in the literature
Molecular Mechanics. I. Quantum mechanical treatment of molecular systems
 Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Molecular Mechanics I. Quantum mechanical treatment of molecular systems The first principle approach for describing the properties of molecules, including proteins, involves quantum mechanics. For example,
Molecular dynamics simulations of a single stranded (ss) DNA
 Molecular dynamics simulations of a single stranded (ss) DNA Subhasish Chatterjee 1, Bonnie Gersten 1, Siddarth Thakur 2, Alexander Burin 2 1 Department of Chemistry, Queens College and the Graduate Center
Molecular dynamics simulations of a single stranded (ss) DNA Subhasish Chatterjee 1, Bonnie Gersten 1, Siddarth Thakur 2, Alexander Burin 2 1 Department of Chemistry, Queens College and the Graduate Center
Molecular dynamics simulations of gramicidin A in a lipid bilayer: From structure function relations to force fields
 Chemistry and Physics of Lipids 141 (2006) 197 204 Molecular dynamics simulations of gramicidin A in a lipid bilayer: From structure function relations to force fields Turgut Baştuğ, Swarna M. Patra, Serdar
Chemistry and Physics of Lipids 141 (2006) 197 204 Molecular dynamics simulations of gramicidin A in a lipid bilayer: From structure function relations to force fields Turgut Baştuğ, Swarna M. Patra, Serdar
SUPPLEMENTARY INFORMATION An Empirical IR Frequency Map for Ester C=O Stretching Vibrations
 SUPPLEMENTARY INFORMATION An Empirical IR Frequency Map for Ester C=O Stretching Vibrations Sean C. Edington, Jennifer C. Flanagan, Carlos R. Baiz* Department of Chemistry, University of Texas at Austin
SUPPLEMENTARY INFORMATION An Empirical IR Frequency Map for Ester C=O Stretching Vibrations Sean C. Edington, Jennifer C. Flanagan, Carlos R. Baiz* Department of Chemistry, University of Texas at Austin
Supporting Information for. Hydrogen Bonding Structure at Zwitterionic. Lipid/Water Interface
 Supporting Information for Hydrogen Bonding Structure at Zwitterionic Lipid/Water Interface Tatsuya Ishiyama,, Daichi Terada, and Akihiro Morita,, Department of Applied Chemistry, Graduate School of Science
Supporting Information for Hydrogen Bonding Structure at Zwitterionic Lipid/Water Interface Tatsuya Ishiyama,, Daichi Terada, and Akihiro Morita,, Department of Applied Chemistry, Graduate School of Science
Molecular dynamics simulations of anti-aggregation effect of ibuprofen. Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov
 Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Biophysical Journal, Volume 98 Supporting Material Molecular dynamics simulations of anti-aggregation effect of ibuprofen Wenling E. Chang, Takako Takeda, E. Prabhu Raman, and Dmitri Klimov Supplemental
Benchmarking the Thermodynamic Analysis of Water Molecules Around a Model Beta Sheet. David J. Huggins. Cambridge, CB2 0XZ, United Kingdom
 Benchmarking the Thermodynamic Analysis of Water Molecules Around a Model Beta Sheet a, b, c David J. Huggins Affiliations: a University of Cambridge, Cambridge Molecular Therapeutics Programme, Hutchison/MRC
Benchmarking the Thermodynamic Analysis of Water Molecules Around a Model Beta Sheet a, b, c David J. Huggins Affiliations: a University of Cambridge, Cambridge Molecular Therapeutics Programme, Hutchison/MRC
Current address: Department of Chemistry, Hong Kong Baptist University, Kowloon Tong, Hong Kong,
 Hydrolysis of Cisplatin - A Metadynamics Study Supporting Information Justin Kai-Chi Lau a and Bernd Ensing* b Department of Chemistry and Applied Bioscience, ETH Zurich, USI Campus, Computational Science,
Hydrolysis of Cisplatin - A Metadynamics Study Supporting Information Justin Kai-Chi Lau a and Bernd Ensing* b Department of Chemistry and Applied Bioscience, ETH Zurich, USI Campus, Computational Science,
Equilibrated atomic models of outward-facing P-glycoprotein and effect of ATP binding on structural dynamics (Supplementary Information)
 Equilibrated atomic models of outward-facing P-glycoprotein and effect of ATP binding on structural dynamics (Supplementary Information) Lurong Pan 1 and Stephen G. Aller 2 * 1,2 Department of Pharmacology
Equilibrated atomic models of outward-facing P-glycoprotein and effect of ATP binding on structural dynamics (Supplementary Information) Lurong Pan 1 and Stephen G. Aller 2 * 1,2 Department of Pharmacology
PROOF. Forced Dissociation of the Strand Dimer Interface between C-Cadherin Ectodomains. M.V. Bayas 1, K.Schulten 2 and D.
 Tech Science Press Paper Galley Proof Only Please Return in 48 Hours. Forced Dissociation of the Strand Dimer Interface between C-Cadherin Ectodomains Abstract: The force-induced dissociation of the strand
Tech Science Press Paper Galley Proof Only Please Return in 48 Hours. Forced Dissociation of the Strand Dimer Interface between C-Cadherin Ectodomains Abstract: The force-induced dissociation of the strand
Structure and Stability of Cyclic Peptide Based Nanotubes: An Molecular Dynamics Study of the Influence of Amino Acid Composition
 Structure and Stability of Cyclic Peptide Based Nanotubes: An Molecular Dynamics Study of the Influence of Amino Acid Composition Ramadoss Vijayaraj, Sofie Van Damme, Patrick Bultinck, and Venkatesan Subramanian
Structure and Stability of Cyclic Peptide Based Nanotubes: An Molecular Dynamics Study of the Influence of Amino Acid Composition Ramadoss Vijayaraj, Sofie Van Damme, Patrick Bultinck, and Venkatesan Subramanian
A Study of the Molecular Conformations and the Electronic Structures of Peptide Nanorings and Nanotubes Hajime Okamoto
 A Study of the Molecular Conformations and the Electronic Structures of Peptide Nanorings and Nanotubes by Hajime Okamoto March, 2004 Contents 1 Introduction 1 1.1 Introduction.................................
A Study of the Molecular Conformations and the Electronic Structures of Peptide Nanorings and Nanotubes by Hajime Okamoto March, 2004 Contents 1 Introduction 1 1.1 Introduction.................................
T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y. jgp
 S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
S u p p l e m e n ta l m at e r i a l jgp Lee et al., http://www.jgp.org/cgi/content/full/jgp.201411219/dc1 T H E J O U R N A L O F G E N E R A L P H Y S I O L O G Y S u p p l e m e n ta l D I S C U S
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015,
 Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Biochemistry,530:,, Introduc5on,to,Structural,Biology, Autumn,Quarter,2015, Course,Informa5on, BIOC%530% GraduateAlevel,discussion,of,the,structure,,func5on,,and,chemistry,of,proteins,and, nucleic,acids,,control,of,enzyma5c,reac5ons.,please,see,the,course,syllabus,and,
Ion Selectivity in the KcsA Potassium Channel from the Perspective of the Ion Binding Site
 2138 Biophysical Journal Volume 96 March 2009 2138 2145 Ion Selectivity in the KcsA Potassium Channel from the Perspective of the Ion Binding Site Purushottam D. Dixit, Safir Merchant, and D. Asthagiri
2138 Biophysical Journal Volume 96 March 2009 2138 2145 Ion Selectivity in the KcsA Potassium Channel from the Perspective of the Ion Binding Site Purushottam D. Dixit, Safir Merchant, and D. Asthagiri
AN AB INITIO STUDY OF INTERMOLECULAR INTERACTIONS OF GLYCINE, ALANINE AND VALINE DIPEPTIDE-FORMALDEHYDE DIMERS
 Journal of Undergraduate Chemistry Research, 2004, 1, 15 AN AB INITIO STUDY OF INTERMOLECULAR INTERACTIONS OF GLYCINE, ALANINE AND VALINE DIPEPTIDE-FORMALDEHYDE DIMERS J.R. Foley* and R.D. Parra Chemistry
Journal of Undergraduate Chemistry Research, 2004, 1, 15 AN AB INITIO STUDY OF INTERMOLECULAR INTERACTIONS OF GLYCINE, ALANINE AND VALINE DIPEPTIDE-FORMALDEHYDE DIMERS J.R. Foley* and R.D. Parra Chemistry
Forced Dissociation of the Strand Dimer Interface between C-Cadherin Ectodomains
 Copyright c 2004 Tech Science Press MCB, vol.1, no.2, pp.101-111, 2004 Forced Dissociation of the Strand Dimer Interface between C-Cadherin Ectodomains M.V. Bayas 1, K.Schulten 2 and D. Leckband 3 Abstract:
Copyright c 2004 Tech Science Press MCB, vol.1, no.2, pp.101-111, 2004 Forced Dissociation of the Strand Dimer Interface between C-Cadherin Ectodomains M.V. Bayas 1, K.Schulten 2 and D. Leckband 3 Abstract:
Alkanes. Introduction
 Introduction Alkanes Recall that alkanes are aliphatic hydrocarbons having C C and C H bonds. They can be categorized as acyclic or cyclic. Acyclic alkanes have the molecular formula C n H 2n+2 (where
Introduction Alkanes Recall that alkanes are aliphatic hydrocarbons having C C and C H bonds. They can be categorized as acyclic or cyclic. Acyclic alkanes have the molecular formula C n H 2n+2 (where
REPORT TITLE: Final Report: Facially Amphiphilic Polymers with Cationic Groups that Mimic Beta-Sheet Structure
 AR-STIR-Final Report MEMRANDUM F TRANSMITTAL U.S. Army Research ffice ATTN: AMSRL-R-BI (TR) P.. Box 12211 Research Triangle Park, NC 27709-2211 Reprint (rig + 2 copies) Manuscript (1 copy) Technical Report
AR-STIR-Final Report MEMRANDUM F TRANSMITTAL U.S. Army Research ffice ATTN: AMSRL-R-BI (TR) P.. Box 12211 Research Triangle Park, NC 27709-2211 Reprint (rig + 2 copies) Manuscript (1 copy) Technical Report
Solutions and Non-Covalent Binding Forces
 Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
Chapter 3 Solutions and Non-Covalent Binding Forces 3.1 Solvent and solution properties Molecules stick together using the following forces: dipole-dipole, dipole-induced dipole, hydrogen bond, van der
FACTORS IN MECHANICAL STABILITY OF PROTEIN L : A STEERED MOLECULAR DYNAMICS STUDY R. ADITAMA, R. HERTADI *
 FACTORS IN MECHANICAL STABILITY OF PROTEIN L : A STEERED MOLECULAR DYNAMICS STUDY R. ADITAMA, R. HERTADI * Biochemistry Research Group, Faculty of Mathematics and Natural Sciences Institut Teknologi Bandung,
FACTORS IN MECHANICAL STABILITY OF PROTEIN L : A STEERED MOLECULAR DYNAMICS STUDY R. ADITAMA, R. HERTADI * Biochemistry Research Group, Faculty of Mathematics and Natural Sciences Institut Teknologi Bandung,
Chapter 6 Cyclic urea - a new central unit in bent-core compounds
 82 Chapter 6 Cyclic urea - a new central unit in bent-core compounds A new class of five-ring bent-core molecules with a cyclic urea group as a central unit was synthesized [94]. A significant difference
82 Chapter 6 Cyclic urea - a new central unit in bent-core compounds A new class of five-ring bent-core molecules with a cyclic urea group as a central unit was synthesized [94]. A significant difference
MODIFIED PROXIMITY CRITERIA FOR THE ANALYSIS OF THE SOLVATION OF A POLYFUNCTIONAL SOLUTE.
 Molecular Simulation 1988, Vol. 1 pp.327-332 c 1988 Gordon and Breach Science Publishers S.A. MODIFIED PROXIMITY CRITERIA FOR THE ANALYSIS OF THE SOLVATION OF A POLYFUNCTIONAL SOLUTE. MIHALY MEZEI Department
Molecular Simulation 1988, Vol. 1 pp.327-332 c 1988 Gordon and Breach Science Publishers S.A. MODIFIED PROXIMITY CRITERIA FOR THE ANALYSIS OF THE SOLVATION OF A POLYFUNCTIONAL SOLUTE. MIHALY MEZEI Department
Supplementary Information. Surface Microstructure Engenders Unusual Hydrophobicity in. Phyllosilicates
 Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2018 Supplementary Information Surface Microstructure Engenders Unusual Hydrophobicity in Phyllosilicates
Electronic Supplementary Material (ESI) for ChemComm. This journal is The Royal Society of Chemistry 2018 Supplementary Information Surface Microstructure Engenders Unusual Hydrophobicity in Phyllosilicates
Useful background reading
 Overview of lecture * General comment on peptide bond * Discussion of backbone dihedral angles * Discussion of Ramachandran plots * Description of helix types. * Description of structures * NMR patterns
Overview of lecture * General comment on peptide bond * Discussion of backbone dihedral angles * Discussion of Ramachandran plots * Description of helix types. * Description of structures * NMR patterns
Amino Acids and Proteins at ZnO-water Interfaces in Molecular Dynamics Simulations: Electronic Supplementary Information
 Amino Acids and Proteins at ZnO-water Interfaces in Molecular Dynamics Simulations: Electronic Supplementary Information Grzegorz Nawrocki and Marek Cieplak Institute of Physics, Polish Academy of Sciences,
Amino Acids and Proteins at ZnO-water Interfaces in Molecular Dynamics Simulations: Electronic Supplementary Information Grzegorz Nawrocki and Marek Cieplak Institute of Physics, Polish Academy of Sciences,
Dynamic hydration valve controlled ion permeability and
 1 Dynamic hydration valve controlled ion permeability and stability of NaK channel Rong Shen 1, *, Wanlin Guo 1, * & Wenyu Zhong 1 1 Institute of Nano Science, Nanjing University of Aeronautics and Astronautics,
1 Dynamic hydration valve controlled ion permeability and stability of NaK channel Rong Shen 1, *, Wanlin Guo 1, * & Wenyu Zhong 1 1 Institute of Nano Science, Nanjing University of Aeronautics and Astronautics,
 There are two types of polysaccharides in cell: glycogen and starch Starch and glycogen are polysaccharides that function to store energy Glycogen Glucose obtained from primary sources either remains soluble
There are two types of polysaccharides in cell: glycogen and starch Starch and glycogen are polysaccharides that function to store energy Glycogen Glucose obtained from primary sources either remains soluble
Molecular Insights on the Cyclic Peptide Nanotube-Mediated Transportation of Antitumor Drug 5-Fluorouracil
 articles Molecular Insights on the Cyclic Peptide Nanotube-Mediated Transportation of Antitumor Drug 5-Fluorouracil Huifang Liu, Jian Chen, Qing Shen, Wei Fu,* and Wei Wu* School of Pharmacy, Fudan UniVersity,
articles Molecular Insights on the Cyclic Peptide Nanotube-Mediated Transportation of Antitumor Drug 5-Fluorouracil Huifang Liu, Jian Chen, Qing Shen, Wei Fu,* and Wei Wu* School of Pharmacy, Fudan UniVersity,
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Biochimica et Biophysica Acta
 Biochimica et Biophysica Acta 1798 (2010) 1474 1479 Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamem Hydration valve controlled non-selective
Biochimica et Biophysica Acta 1798 (2010) 1474 1479 Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamem Hydration valve controlled non-selective
Membrane insertion profiles of peptides probed by molecular dynamics simulations
 Membrane insertion profiles of peptides probed by molecular dynamics simulations In-Chul Yeh, * Mark A. Olson, # Michael S. Lee, *# and Anders Wallqvist * * Biotechnology High Performance Computing Software
Membrane insertion profiles of peptides probed by molecular dynamics simulations In-Chul Yeh, * Mark A. Olson, # Michael S. Lee, *# and Anders Wallqvist * * Biotechnology High Performance Computing Software
Structure and Bonding of Organic Molecules
 Chem 220 Notes Page 1 Structure and Bonding of Organic Molecules I. Types of Chemical Bonds A. Why do atoms forms bonds? Atoms want to have the same number of electrons as the nearest noble gas atom (noble
Chem 220 Notes Page 1 Structure and Bonding of Organic Molecules I. Types of Chemical Bonds A. Why do atoms forms bonds? Atoms want to have the same number of electrons as the nearest noble gas atom (noble
Bulk behaviour. Alanine. FIG. 1. Chemical structure of the RKLPDA peptide. Numbers on the left mark alpha carbons.
 Bulk behaviour To characterise the conformational behaviour of the peptide, first we looked at the statistics of alpha carbons and the torsion angles. Then they were correlated with positions of positively
Bulk behaviour To characterise the conformational behaviour of the peptide, first we looked at the statistics of alpha carbons and the torsion angles. Then they were correlated with positions of positively
Supporting Information
 Supporting Information ph-responsive self-assembly of polysaccharide through a rugged energy landscape Brian H. Morrow, Gregory F. Payne, and Jana Shen Department of Pharmaceutical Sciences, School of
Supporting Information ph-responsive self-assembly of polysaccharide through a rugged energy landscape Brian H. Morrow, Gregory F. Payne, and Jana Shen Department of Pharmaceutical Sciences, School of
Vibrational Energy Relaxation of Tailored Hemes in Myoglobin Following Ligand Photolysis Supports Energy Funneling Mechanism of Heme Cooling
 10634 J. Phys. Chem. B 2003, 107, 10634-10639 Vibrational Energy Relaxation of Tailored Hemes in Myoglobin Following Ligand Photolysis Supports Energy Funneling Mechanism of Heme Cooling Lintao Bu and
10634 J. Phys. Chem. B 2003, 107, 10634-10639 Vibrational Energy Relaxation of Tailored Hemes in Myoglobin Following Ligand Photolysis Supports Energy Funneling Mechanism of Heme Cooling Lintao Bu and
CHAPTER 2. Structure and Reactivity: Acids and Bases, Polar and Nonpolar Molecules
 CHAPTER 2 Structure and Reactivity: Acids and Bases, Polar and Nonpolar Molecules 2-1 Kinetics and Thermodynamics of Simple Chemical Processes Chemical thermodynamics: Is concerned with the extent that
CHAPTER 2 Structure and Reactivity: Acids and Bases, Polar and Nonpolar Molecules 2-1 Kinetics and Thermodynamics of Simple Chemical Processes Chemical thermodynamics: Is concerned with the extent that
New Perspective on structure and bonding in water using XAS and XRS
 New Perspective on structure and bonding in water using XAS and XRS Anders Nilsson Stanford Synchrotron Radiation Laboratory (SSRL) and Stockholm University, Sweden R. Ludwig Angew. Chem. 40, 1808 (2001)
New Perspective on structure and bonding in water using XAS and XRS Anders Nilsson Stanford Synchrotron Radiation Laboratory (SSRL) and Stockholm University, Sweden R. Ludwig Angew. Chem. 40, 1808 (2001)
Organic Chemistry, Second Edition. Janice Gorzynski Smith University of Hawai i. Chapter 4 Alkanes
 Organic Chemistry, Second Edition Janice Gorzynski Smith University of Hawai i Chapter 4 Alkanes Prepared by Rabi Ann Musah State University of New York at Albany Copyright The McGraw-Hill Companies, Inc.
Organic Chemistry, Second Edition Janice Gorzynski Smith University of Hawai i Chapter 4 Alkanes Prepared by Rabi Ann Musah State University of New York at Albany Copyright The McGraw-Hill Companies, Inc.
A Molecular Dynamics Simulation of the Human Lysozyme Camelid VHH HL6 Antibody System
 Int. J. Mol. Sci. 2009, 10, 1719-1727; doi:10.3390/ijms10041719 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article A Molecular Dynamics Simulation
Int. J. Mol. Sci. 2009, 10, 1719-1727; doi:10.3390/ijms10041719 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article A Molecular Dynamics Simulation
Diffusion of Water and Diatomic Oxygen in Poly(3-hexylthiophene) Melt: A Molecular Dynamics Simulation Study
 Diffusion of Water and Diatomic Oxygen in Poly(3-hexylthiophene) Melt: A Molecular Dynamics Simulation Study Julia Deitz, Yeneneh Yimer, and Mesfin Tsige Department of Polymer Science University of Akron
Diffusion of Water and Diatomic Oxygen in Poly(3-hexylthiophene) Melt: A Molecular Dynamics Simulation Study Julia Deitz, Yeneneh Yimer, and Mesfin Tsige Department of Polymer Science University of Akron
Structural and mechanistic insight into the substrate. binding from the conformational dynamics in apo. and substrate-bound DapE enzyme
 Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
Electronic Supplementary Material (ESI) for Physical Chemistry Chemical Physics. This journal is the Owner Societies 215 Structural and mechanistic insight into the substrate binding from the conformational
The Ionization State and the Conformation of Glu-71 in the KcsA K Channel
 772 Biophysical Journal Volume 82 February 2002 772 780 The Ionization State and the Conformation of Glu-71 in the KcsA K Channel Simon Bernèche* and Benoît Roux* *Department of Biochemistry, Weill Medical
772 Biophysical Journal Volume 82 February 2002 772 780 The Ionization State and the Conformation of Glu-71 in the KcsA K Channel Simon Bernèche* and Benoît Roux* *Department of Biochemistry, Weill Medical
Secondary and sidechain structures
 Lecture 2 Secondary and sidechain structures James Chou BCMP201 Spring 2008 Images from Petsko & Ringe, Protein Structure and Function. Branden & Tooze, Introduction to Protein Structure. Richardson, J.
Lecture 2 Secondary and sidechain structures James Chou BCMP201 Spring 2008 Images from Petsko & Ringe, Protein Structure and Function. Branden & Tooze, Introduction to Protein Structure. Richardson, J.
Secondary Structure. Bioch/BIMS 503 Lecture 2. Structure and Function of Proteins. Further Reading. Φ, Ψ angles alone determine protein structure
 Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
Bioch/BIMS 503 Lecture 2 Structure and Function of Proteins August 28, 2008 Robert Nakamoto rkn3c@virginia.edu 2-0279 Secondary Structure Φ Ψ angles determine protein structure Φ Ψ angles are restricted
The biomolecules of terrestrial life
 Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Functional groups in biomolecules Groups of atoms that are responsible for the chemical properties of biomolecules The biomolecules of terrestrial life Planets and Astrobiology (2017-2018) G. Vladilo 1
Mechanical Proteins. Stretching imunoglobulin and fibronectin. domains of the muscle protein titin. Adhesion Proteins of the Immune System
 Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Mechanical Proteins F C D B A domains of the muscle protein titin E Stretching imunoglobulin and fibronectin G NIH Resource for Macromolecular Modeling and Bioinformatics Theoretical Biophysics Group,
Supporting Information
 Supporting Information Interface-Induced Affinity Sieving in Nanoporous Graphenes for Liquid-Phase Mixtures Yanan Hou, Zhijun Xu, Xiaoning Yang * State Key Laboratory of Material-Orientated Chemical Engineering,
Supporting Information Interface-Induced Affinity Sieving in Nanoporous Graphenes for Liquid-Phase Mixtures Yanan Hou, Zhijun Xu, Xiaoning Yang * State Key Laboratory of Material-Orientated Chemical Engineering,
What makes a good graphene-binding peptide? Adsorption of amino acids and peptides at aqueous graphene interfaces: Electronic Supplementary
 Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Electronic Supplementary Material (ESI) for Journal of Materials Chemistry B. This journal is The Royal Society of Chemistry 21 What makes a good graphene-binding peptide? Adsorption of amino acids and
Molecular dynamics simulation of Aquaporin-1. 4 nm
 Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Molecular dynamics simulation of Aquaporin-1 4 nm Molecular Dynamics Simulations Schrödinger equation i~@ t (r, R) =H (r, R) Born-Oppenheimer approximation H e e(r; R) =E e (R) e(r; R) Nucleic motion described
Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2015 Destruction of Amyloid Fibrils by Graphene through Penetration and Extraction of Peptides Zaixing
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability
 Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Lecture 2 and 3: Review of forces (ctd.) and elementary statistical mechanics. Contributions to protein stability Part I. Review of forces Covalent bonds Non-covalent Interactions: Van der Waals Interactions
Protein Structure Basics
 Protein Structure Basics Presented by Alison Fraser, Christine Lee, Pradhuman Jhala, Corban Rivera Importance of Proteins Muscle structure depends on protein-protein interactions Transport across membranes
Protein Structure Basics Presented by Alison Fraser, Christine Lee, Pradhuman Jhala, Corban Rivera Importance of Proteins Muscle structure depends on protein-protein interactions Transport across membranes
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL Systematic Coarse-Grained Modeling of Complexation between Small Interfering RNA and Polycations Zonghui Wei 1 and Erik Luijten 1,2,3,4,a) 1 Graduate Program in Applied Physics, Northwestern
SUPPLEMENTAL MATERIAL Systematic Coarse-Grained Modeling of Complexation between Small Interfering RNA and Polycations Zonghui Wei 1 and Erik Luijten 1,2,3,4,a) 1 Graduate Program in Applied Physics, Northwestern
Don t forget to bring your MD tutorial. Potential Energy (hyper)surface
 Don t forget to bring your MD tutorial Lab session starts at 1pm You will have to finish an MD/SMD exercise on α-conotoxin in oxidized and reduced forms Potential Energy (hyper)surface What is Force? Energy
Don t forget to bring your MD tutorial Lab session starts at 1pm You will have to finish an MD/SMD exercise on α-conotoxin in oxidized and reduced forms Potential Energy (hyper)surface What is Force? Energy
Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached
 Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached to HpUreI. Urea hydrolysis products 2NH 3 and 1CO 2
Supplementary Figure S1. Urea-mediated buffering mechanism of H. pylori. Gastric urea is funneled to a cytoplasmic urease that is presumably attached to HpUreI. Urea hydrolysis products 2NH 3 and 1CO 2
Problem Set 1
 2006 7.012 Problem Set 1 Due before 5 PM on FRIDAY, September 15, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. For each of the following parts, pick
2006 7.012 Problem Set 1 Due before 5 PM on FRIDAY, September 15, 2006. Turn answers in to the box outside of 68-120. PLEASE WRITE YOUR ANSWERS ON THIS PRINTOUT. 1. For each of the following parts, pick
Introduction to Alkenes. Structure and Reactivity
 4 4 Introduction to Alkenes. Structure and Reactivity Alkenes are hydrocarbons that contain one or more carbon carbon double bonds. Alkenes are sometimes called olefins, particularly in the chemical industry.
4 4 Introduction to Alkenes. Structure and Reactivity Alkenes are hydrocarbons that contain one or more carbon carbon double bonds. Alkenes are sometimes called olefins, particularly in the chemical industry.
Dihedral Angles. Homayoun Valafar. Department of Computer Science and Engineering, USC 02/03/10 CSCE 769
 Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Dihedral Angles Homayoun Valafar Department of Computer Science and Engineering, USC The precise definition of a dihedral or torsion angle can be found in spatial geometry Angle between to planes Dihedral
Supporting information for. DNA Origami-Graphene Hybrid Nanopore for DNA Detection
 Supporting information for DNA Origami-Graphene Hybrid Nanopore for DNA Detection Amir Barati Farimani, Payam Dibaeinia, Narayana R. Aluru Department of Mechanical Science and Engineering Beckman Institute
Supporting information for DNA Origami-Graphene Hybrid Nanopore for DNA Detection Amir Barati Farimani, Payam Dibaeinia, Narayana R. Aluru Department of Mechanical Science and Engineering Beckman Institute
Supplementary Information
 1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
1 Supplementary Information Figure S1 The V=0.5 Harker section of an anomalous difference Patterson map calculated using diffraction data from the NNQQNY crystal at 1.3 Å resolution. The position of the
Supporting Information for. The Origin of Coupled Chloride and Proton Transport in a Cl /H + Antiporter
 Supporting Information for The Origin of Coupled Chloride and Proton Transport in a Cl /H + Antiporter Sangyun Lee, Heather B. Mayes, Jessica M. J. Swanson, and Gregory A. Voth * Department of Chemistry,
Supporting Information for The Origin of Coupled Chloride and Proton Transport in a Cl /H + Antiporter Sangyun Lee, Heather B. Mayes, Jessica M. J. Swanson, and Gregory A. Voth * Department of Chemistry,
Molecular Dynamics Simulations. Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia
 Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Molecular Dynamics Simulations Dr. Noelia Faginas Lago Dipartimento di Chimica,Biologia e Biotecnologie Università di Perugia 1 An Introduction to Molecular Dynamics Simulations Macroscopic properties
Exploring the free-energy landscapes of biological systems with steered molecular dynamics
 Exploring the free-energy landscapes of biological systems with steered molecular dynamics Guodong Hu and L. Y. Chen* Department of Physics, University of Texas at San Antonio, One UTSA Circle, San Antonio,
Exploring the free-energy landscapes of biological systems with steered molecular dynamics Guodong Hu and L. Y. Chen* Department of Physics, University of Texas at San Antonio, One UTSA Circle, San Antonio,
Govardhan Reddy. John E. Straub. D. Thirumalai*
 1162 J. Phys. Chem. B 2009, 113, 1162 1172 Influence of Preformed Asp23-Lys28 Salt Bridge on the Conformational Fluctuations of Monomers and Dimers of Aβ Peptides with Implications for ates of Fibril Formation
1162 J. Phys. Chem. B 2009, 113, 1162 1172 Influence of Preformed Asp23-Lys28 Salt Bridge on the Conformational Fluctuations of Monomers and Dimers of Aβ Peptides with Implications for ates of Fibril Formation
Molecular Dynamics Simulation and Coarse-Grained Analysis of the Arp2/3 Complex
 5324 Biophysical Journal Volume 95 December 2008 5324 5333 Molecular Dynamics Simulation and Coarse-Grained Analysis of the Arp2/3 Complex Jim Pfaendtner* y and Gregory A. Voth* y *Center for Biophysical
5324 Biophysical Journal Volume 95 December 2008 5324 5333 Molecular Dynamics Simulation and Coarse-Grained Analysis of the Arp2/3 Complex Jim Pfaendtner* y and Gregory A. Voth* y *Center for Biophysical
Supplementary Figure 1 Irregular arrangement of E,E-8-mer on TMA. STM height images formed when
 Supplementary Figure 1 Irregular arrangement of E,E-8-mer on TMA. STM height images formed when a 5 µl heptanoic acid solution of E,E-8-mer is applied on: (a) a TMA templated HOPG substrate, U s = +1.00
Supplementary Figure 1 Irregular arrangement of E,E-8-mer on TMA. STM height images formed when a 5 µl heptanoic acid solution of E,E-8-mer is applied on: (a) a TMA templated HOPG substrate, U s = +1.00
This Answer/Solution Example is taken from a student s actual homework report. I thank him for permission to use it here. JG
 This Answer/Solution Example is taken from a student s actual homework report. I thank him for permission to use it here. JG Chem 8021 Spring 2005 Project II Calculation of Self-Diffusion Coeffecient of
This Answer/Solution Example is taken from a student s actual homework report. I thank him for permission to use it here. JG Chem 8021 Spring 2005 Project II Calculation of Self-Diffusion Coeffecient of
Supporting Information
 Supporting Information Constant ph molecular dynamics reveals ph-modulated binding of two small-molecule BACE1 inhibitors Christopher R. Ellis 1,, Cheng-Chieh Tsai 1,, Xinjun Hou 2, and Jana Shen 1, 1
Supporting Information Constant ph molecular dynamics reveals ph-modulated binding of two small-molecule BACE1 inhibitors Christopher R. Ellis 1,, Cheng-Chieh Tsai 1,, Xinjun Hou 2, and Jana Shen 1, 1
Controllable Water Channel Gating of Nanometer Dimensions
 Published on Web 04/21/2005 Controllable Water Channel Gating of Nanometer Dimensions Rongzheng Wan, Jingyuan Li, Hangjun Lu, and Haiping Fang*, Contribution from Shanghai Institute of Applied Physics,
Published on Web 04/21/2005 Controllable Water Channel Gating of Nanometer Dimensions Rongzheng Wan, Jingyuan Li, Hangjun Lu, and Haiping Fang*, Contribution from Shanghai Institute of Applied Physics,
MARTINI simulation details
 S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
S1 Appendix MARTINI simulation details MARTINI simulation initialization and equilibration In this section, we describe the initialization of simulations from Main Text section Residue-based coarsegrained
LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor
 LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
LS1a Fall 2014 Problem Set #2 Due Monday 10/6 at 6 pm in the drop boxes on the Science Center 2 nd Floor Note: Adequate space is given for each answer. Questions that require a brief explanation should
Free energy calculations and the potential of mean force
 Free energy calculations and the potential of mean force IMA Workshop on Classical and Quantum Approaches in Molecular Modeling Mark Tuckerman Dept. of Chemistry and Courant Institute of Mathematical Science
Free energy calculations and the potential of mean force IMA Workshop on Classical and Quantum Approaches in Molecular Modeling Mark Tuckerman Dept. of Chemistry and Courant Institute of Mathematical Science
Lecture 34 Protein Unfolding Thermodynamics
 Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Routine access to millisecond timescale events with accelerated molecular dynamics
 Routine access to millisecond timescale events with accelerated molecular dynamics Levi C.T. Pierce, Romelia Salomon-Ferrer, Cesar Augusto F. de Oliveira #, J. Andrew McCammon #, Ross C. Walker * SUPPORTING
Routine access to millisecond timescale events with accelerated molecular dynamics Levi C.T. Pierce, Romelia Salomon-Ferrer, Cesar Augusto F. de Oliveira #, J. Andrew McCammon #, Ross C. Walker * SUPPORTING
Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia
 University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
University of Groningen Peptide folding in non-aqueous environments investigated with molecular dynamics simulations Soto Becerra, Patricia IMPORTANT NOTE: You are advised to consult the publisher's version
The Structure and Functions of Proteins
 Wright State University CORE Scholar Computer Science and Engineering Faculty Publications Computer Science and Engineering 2003 The Structure and Functions of Proteins Dan E. Krane Wright State University
Wright State University CORE Scholar Computer Science and Engineering Faculty Publications Computer Science and Engineering 2003 The Structure and Functions of Proteins Dan E. Krane Wright State University
Chapter 13 Conjugated Unsaturated Systems
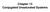 Chapter 13 Conjugated Unsaturated Systems Introduction Conjugated unsaturated systems have a p orbital on a carbon adjacent to a double bond The p orbital can come from another double or triple bond The
Chapter 13 Conjugated Unsaturated Systems Introduction Conjugated unsaturated systems have a p orbital on a carbon adjacent to a double bond The p orbital can come from another double or triple bond The
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY-
 BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
BIOCHEMISTRY GUIDED NOTES - AP BIOLOGY- ELEMENTS AND COMPOUNDS - anything that has mass and takes up space. - cannot be broken down to other substances. - substance containing two or more different elements
Coherent Two Dimensional Infrared Spectroscopy of a Cyclic Decapeptide Antamanide. A Simulation Study of the Amide-I and A Bands
 J. Phys. Chem. B 2008, 112, 12479 12490 12479 Coherent Two Dimensional Infrared Spectroscopy of a Cyclic Decapeptide Antamanide. A Simulation Study of the Amide-I and A Bands Cyril Falvo, Tomoyuki Hayashi,
J. Phys. Chem. B 2008, 112, 12479 12490 12479 Coherent Two Dimensional Infrared Spectroscopy of a Cyclic Decapeptide Antamanide. A Simulation Study of the Amide-I and A Bands Cyril Falvo, Tomoyuki Hayashi,
Conjugated Systems, Orbital Symmetry and UV Spectroscopy
 Conjugated Systems, Orbital Symmetry and UV Spectroscopy Introduction There are several possible arrangements for a molecule which contains two double bonds (diene): Isolated: (two or more single bonds
Conjugated Systems, Orbital Symmetry and UV Spectroscopy Introduction There are several possible arrangements for a molecule which contains two double bonds (diene): Isolated: (two or more single bonds
