False Discovery Rates of Protein Identifications: A Strike against the Two-Peptide Rule
|
|
|
- Molly Ellis
- 6 years ago
- Views:
Transcription
1 False Discovery Rates of Protein Identifications: A Strike against the Two-Peptide Rule Nitin Gupta*, and Pavel A. Pevzner Bioinformatics Program and Department of Computer Science and Engineering, University of California San Diego, La Jolla, California Received September 5, 2008 Most proteomics studies attempt to maximize the number of peptide identifications and subsequently infer proteins containing two or more peptides as reliable protein identifications. In this study, we evaluate the effect of this two-peptide rule on protein identifications, using multiple search tools and data sets. Contrary to the intuition, the two-peptide rule reduces the number of protein identifications in the target database more significantly than in the decoy database and results in increased false discovery rates, compared to the case when single-hit proteins are not discarded. We therefore recommend that the two-peptide rule should be abandoned, and instead, protein identifications should be subject to the estimation of error rates, as is the case with peptide identifications. We further extend the generating function approach (originally proposed for evaluating matches between a peptide and a single spectrum) to evaluating matches between a protein and an entire spectral data set. Keywords: two-peptide rule false discovery rate mass spectrometry peptide identification protein identification decoy database false positives 1. Introduction * To whom correspondence should be addressed. ngupta@ucsd.edu. Bioinformatics Program, University of California San Diego. Department of Computer Science and Engineering, University of California San Diego. Tandem mass spectrometry (MS/MS) database search tools are routinely used for peptide identifications. 1 The results often include many false positives, 2 and a common approach for estimating the False Discovery Rate (FDR) of peptide identifications is based on the use of randomized decoy databases. 3,4 The reported set of peptide identifications is determined by varying the score cutoff to achieve the desired FDR. Intuitively, a protein must be present in the sample if a peptide within it is identified (assuming the same peptide is not present elsewhere in the proteome). However, since peptide identification tools generate some false peptides, many researchers are cautious about the one-hit-wonders, 5-7 making it a common practice to report only proteins with at least two peptides as reliable identifications (proteins with single peptide identifications are often ignored, or delegated to the Supplemetary materials). The two-peptide rule has done the field a great service by providing a stringent criterion for reporting proteomic data. However, while the two peptide rule seems wellintentioned and reasonable, we are unaware of any theoretical studies supporting this rule. Indeed, the question whether two identified peptides with scores x and y (within the same protein) represent a better evidence for expression of this protein than a single peptide with score z (that is larger than x and y) depends on parameters x, y, z (and the protein length) and remains poorly addressed. Below, we show that the twopeptide rule is inferior to the single-peptide rule that takes into account one-hit-wonders (with appropriately chosen score threshold z). We therefore argue that the two-peptide bias should be removed and that protein identifications based on single peptides should be treated at par with identifications based on multiple peptides, instead of salvaging them through postprocessing. 8 Gupta et al., 9 estimated that 80% of the one-hit-wonders (proteins with a single identified peptide) in proteogenomics study of Shewanella oneidensis MR-1 are likely to be expressed. Comparative analysis of three Shewanella species revealed that many of these one-hit-wonders are actually observed as orthologs in multiple species, providing further support for their expression. 10 These observations indicate that the two-peptide rule for protein identification is an unsubstantiated heuristic that often results in loss of a large number of protein identifications (20%-25% of all expressed proteins). Another unsubstantiated assumption often made by proteomics researchers is that maximizing the number of peptide identifications automatically results in maximizing the number of protein identifications. While peptide and protein identification goals are closely related, there are cases when one is more important than the other. Optimizing protein-level FDR is critical in applications such as biomarker discovery or proteome profiling, while optimizing peptide-level FDR is important for label-free quantitations, 11 proteogenomics, 9,10 or peptidomics. 12,13 We demonstrate that peptide and protein identifications are two different computational problems that should be approached differently: maximizing the number of peptide identifications does not necessarily result in maximiz /pr CCC: $ American Chemical Society Journal of Proteome Research 2009, 8, Published on Web 07/23/2009
2 research articles ing the number of protein identifications (for example, in cases when most peptides hit a few proteins). While FDR among all protein identifications in a data set can be computed using the decoy database, computing the false positive rate (FPR) of individual protein identifications has been an open problem. The existing tools provide probabilistic scores, which may be correlated with the FPR, but do not provide its rigorous estimate taking into account the lengths of the proteins or the size of the spectral data sets. We extended the generating function framework 14,15 to suggest a different (and simple) approach to the problem of estimating the protein-level FPR. So far, we failed to find any evidence that the previously proposed techniques for evaluating protein identifications (often based on elaborate machine learning models and multiple peptides) improve over a simple singlepeptide rule combined with the generating function approach. 2. Methods MS/MS Data Sets. The MS/MS data sets used in this study were obtained from S. oneidensis MR-1 (generated in Dick Smith s laboratory at PNNL) and human (generated in Vivian Hook s laboratory at UCSD) samples. These data sets are described in the literature. 9,10,16 The data sets were generated on Thermo LCQ mass-spectrometer for Shewanella and Agilent XCT Ultra for human samples. The human data set includes nearly spectra, while the Shewanella data set has 14.5 million spectra. The Shewanella data set was searched against the Shewanella protein database (size 1.5 MB, containing 4928 sequences), and the human data set was searched against the IPI database version 3.41 (size 40 MB, containing sequences). Decoy databases were generated by randomly shuffling the sequence of each protein in the target database (preserving the background amino acid frequencies for each protein). Peptide Identification. Database searches were carried out using InsPecT 17 and X!Tandem. 18 InsPecT searches were run with the default parameter settings (fragment ion tolerance of 0.5 Da and parent mass tolerance of 2.5 Da). X!Tandem was run using default settings, without any protease specificity (allowing peptides of length up to 40). MS-GeneratingFunction (MS-GF) 14,15 was run on InsPecT results to evaluate the statistical significance of individual peptide identifications (spectral probabilities). We treat the spectral probability as a score, and since MS-GF was used to rescore all InsPecT identifications (including even the very low scoring ones that are typically never reported), InsPecTxMS-GF can be viewed as the third peptide identification tool, besides InsPecT and X!Tandem. MS-GF was recently shown to improve upon InsPecT, X!Tandem, and Sequest/PeptideProphet. 14,19 Protein Identification. Protein identifications are inferred by applying score-thresholds to the peptides identified in a protein. In case of the one-peptide rule (i.e., allowing onehit-wonders), any protein that has a peptide scoring above the chosen threshold is considered identified. This is equivalent to using a protein-level scoring scheme where the score of a protein is computed as the score of its highest scoring peptide. Similarly, in case of two-peptide rule, any protein that has two or more peptides scoring above the threshold is considered identified. Each point on the ROC curves is generated by changing this threshold, and computing the number of protein identifications by each rule, in the target and the decoy databases. Figure 1. Identification of peptides in the Shewanella data set using different approaches and scoring functions. Each point in the curves is generated by varying the scoring threshold and computing the number of hits in the target and the decoy database exceeding the threshold. ProteinProphet was run using the Trans Proteomics Pipeline, v4.0 JETSTREAM rev 2, to process X!Tandem search results on human data set. Both target and decoy protein sequences were included in the X!Tandem search results (without giving this information to ProteinProphet a priori). The final output thus included both target and decoy proteins with a probabilitybased score for each protein. The ROC curve was computed by varying the value of this score-threshold between 0 and 1. ProteinProphet performs better than the traditional twopeptide rule (as expected), but still worse than the one-peptide rule. 3. Results Gupta and Pevzner Comparison of Two-Peptide and Single-Peptide Rule. The intuition behind the two-peptide rule is that it may result in a more severe penalty to decoy hits as compared to the target hits. In other words, removing one-hit-wonders should improve the FDR of peptide and protein identifications. Figure 1 shows this trend for peptide identifications in the Shewanella data set for InsPecT and MS-GF scoring functions. Indeed, if we discard all peptides representing one-hit-wonders, we observe that the tradeoff between the number of peptides identified in the target and the decoy database for different score thresholds shifts in favor of the target hits, compared to the situation when one-hit-wonders are retained. Common sense suggests that the increased number of peptides, for a given FDR, should also increase the number of protein identifications. Therefore, it seems plausible that the two-peptide rule (discarding one-hit-wonders) should perform better than the single-peptide rule (retaining all proteins with one or more peptides). Note that, in common practice, the two-peptide rule is always used in the context of protein identifications, that is after the peptides have been identified for a chosen FDR at peptide level. Figure 2 demonstrates that this intuitive conclusion is not substantiated by the data as the simpler single-peptide rule has superior FDR as compared to the traditional two-peptide approach applied to protein identifications. For example, for the same number (21) of proteins identified in the decoy 4174 Journal of Proteome Research Vol. 8, No. 9, 2009
3 False Discovery Rates of Protein Identifications research articles Figure 2. (a) Identification of proteins in the human data set using different approaches and scoring functions. (b) Similar plot as in panel (a) for Shewanella data set. database, InsPecT 17 identifies 546 proteins in the target human database using the standard two-peptide approach and 742 proteins using the single-peptide approach, a 36% increase in the number of protein identifications. Similarly, the number of protein identifications with X!Tandem 18 increases from 350 to 414 (Figure 3), while the number of protein identifications with MS-GF 14 increases from 607 to 826. Similar trends are seen for the Shewanella data sets (Figure 2b,c). Besides the two-peptide rule, more complex approaches are sometimes used to combine evidence from multiple peptides into protein identifications We compare our results with ProteinProphet, 21 a popular tool that can be used to postprocess MS/MS database search results of programs like Sequest and X!Tandem. Benchmarking ProteinProphet (combined with X!Tandem) against the single-peptide approach (Figure 3) shows that the single-peptide rule has better sensitivity-specificity tradeoff than ProteinProphet, in both human and Shewanella data sets. This surprising result indicates that the simple single-peptide rule should not be discarded without benchmarking it against seemingly more reasonable ( two-peptide rule) and complex (ProteinProphet) approaches. We note that computing FDR using decoy databases (at the protein level) has some limitations because the number of Figure 3. (a) Protein identification in the human data set using X!Tandem search results with different scoring approaches at the protein level. (b) Similar plot as in panel (a) for an arbitrarily selected subset of Shewanella data set containing 1.25 million spectra. protein identifications in the decoy database may deviate from the number of incorrect protein identifications in the target database, particularly in data sets with high proteome coverage. However, the relative comparison of the single-peptide and the two-peptide approaches should not be significantly affected by this phenomenon. The next section describes an approach for computing the protein-level error-rates without using a decoy database. Inference of proteins from identified peptides is complicated by the presence of peptides that are present in multiple proteins. This problem is more pronounced in eukaryotes (since their proteomes have many repeated peptides) as compared to bacteria that have very few repeated peptides. The analysis of Shewanella in Gupta et al. 9 revealed that 98.5% of the identified peptides were unique, but the number can drop down to 40-50% in case of higher eukaryotes when using a sequence database like the human IPI database containing alternatively spliced variants of proteins. A number of approaches have been developed to address this problem, most of which are parsimony-based. 21,25-27 The computation of FDR in our method can depend on the exact method used for protein inference. For the sake of simplicity, we assign the ambiguous peptides (that map to multiple proteins) to one of Journal of Proteome Research Vol. 8, No. 9,
4 research articles Gupta and Pevzner Table 1. An Intuitive Description of Some Frequently-Used Terms a Term Description Figure 4. Identification of proteins, using the unique peptides only (peptides that are not shared between multiple proteins), in the human data set using InsPecT search results with different approaches. the matching proteins randomly. To check that our results are robust to this step, we tried limiting our attention to only the unique peptides that are identified by InsPecT in the human data set. We still find that the single-peptide approach works best for protein identifications (Figure 4). This shows that our conclusions are not significantly altered by peptides matching multiple proteins. Estimating Statistical Significance of Protein Identifications Using Spectral Dictionaries. We say that a protein matches a spectrum with score Score if one of its peptides matches the spectrum (with the same Score). Consider the following two problems: Peptide-Spectrum Matching Problem. Given a spectrum Spectrum and a score threshold Threshold for a spectrumpeptide scoring function, find the probability that a random peptide matches Spectrum with score equal to or larger than Threshold. Protein-Spectra Matching Problem. Given a spectral data set Spectra and a score threshold Threshold for a spectrumpeptide scoring function, find the probability that a random protein matches a spectrum in the set Spectra with score equal to or larger than Threshold. Recently, Kim et al. 14 suggested a generating function approach (MS-GF) for solving the Peptide-Spectrum Matching Problem and thereby evaluating the FPR of Peptide-Spectrum Matches (PSM). In contrast to FDRs empirically computed by the target-decoy approaches (that lack the ability to evaluate the statistical significance of individual PSMs), MS-GF rigorously evaluates individual PSMs using spectral probabilities. However, Kim et al. 14 defined the spectral probabilities for a single peptide and spectrum, while MS/MS searches compare multiple peptides (e.g., all peptides present in a protein database) against multiple spectra. Therefore, accurate estimation of protein-level FPRs requires a solution to the yet unsolved Protein-Spectra Matching Problem. In this section, we address this problem by extending the generating function framework from Peptide-Spectrum matches to Protein-Spectra matches. We start by reviewing the terminology related to spectral probability and spectral dictionaries 14,15 (see Table 1). Given a Dictionary(Spectrum, Threshold) SpectralProbability(Dictionary) Lexicon(Spectra, Threshold) Compression(Spectra, Threshold) a Please refer to the text for details. A set of peptides that match the Spectrum with scores greater than or equal to the Threshold The probability that a random peptide matches a sequence contained in the Dictionary Union of the dictionaries of all spectra in the set Spectra (for the given Threshold), after removing redundant peptides Ratio of the spectral probability of the Lexicon to the sum of the spectral probabilities of each spectral dictionary Dictionary(Spectrum, Threshold) for all spectra from the set Spectra scoring function for PSMs, Dictionary(Spectrum, Threshold) is defined as the set of all possible peptides with Score(Peptide, Spectrum) g Threshold. The spectral dictionary framework transforms the (difficult) problem of evaluating the statistical significance of a PSM with score Threshold into a (simple) problem of evaluating the statistical significance of matches between a set of strings (Dictionary) and a random peptide. Assuming probabilities of all amino acids in a random peptide equal to 1/20, the probability that a Dictionary contains a random peptide (more precisely, a prefix of a sufficiently long random string of amino acids) is given by SpectralProbability- (Dictionary) ) Peptide Dictionary 20 - Peptide, where Peptide stands for the length of Peptide. The dictionary corresponding to a PSM (Dictionary(Peptide, Spectrum)) is defined as Dictionary(Spectrum, Score(Peptide, Spectrum)). Similarly, the spectral probability of a PSM is defined as SpectralProbability(Peptide, Spectrum) ) SpectralProbability(Dictionary(Peptide, Spectrum)). This allows one to convert an arbitrary (additive) scoring function Score(Peptide, Spectrum) into a new scoring function represented by SpectralProbability(Peptide, Spectrum). Kim et al. 14 demonstrated that this new scoring function (varying between 0 and 1) results in a better sensitivity-specificity tradeoff than all other scoring functions they evaluated. Note that, in this new scoring function, the lower scores (spectral probabilities) represent better PSMs. As such, given a Spectrum, one can define Dictionary(Spectrum, SPThreshold), for a spectral probability threshold 0 e SPThreshold e 1, as the set of all peptides with SpectralProbability(Peptide, Spectrum) e SPThreshold when SpectralProbability serves as a scoring function. We alert the reader that the term dictionary refers to both dictionaries defined by the original scoring function Score and the new scoring function SpectralProbability. We further define SpectralxProbability(Spectrum, SPThreshold) as SpectralProbability(Dictionary(Spectrum, SPThreshold)). It is easy to see that the Spectral Probability score (in contrast to the original Score) satisfies the important property: SpectralProbability(Spectrum, SPThreshold) ) SPThreshold In the remainder of this paper, we assume that Spectral Probability is the scoring function being used Journal of Proteome Research Vol. 8, No. 9, 2009
5 False Discovery Rates of Protein Identifications The probability that a Dictionary contains a peptide from a random protein of length n can be computed as 1 - Pj(Dictionary, n), where Pj(Dictionary, n) is the probability that none of the peptides in the Dictionary are contained in a random string of length n. Computing Pj(Dictionary, n) is a nontrivial problem that was solved by Guibas and Odlyzko 28 using the generating function approach. While this approach allows one to compute Pj(Dictionary, n) precisely, the resulting expressions and recurrences are rather difficult to analyze due to correlations between different strings in the Dictionary (see ref 28 for details). We therefore prefer to use an approximation (that ignores correlations), a reasonable assumption for a rather large alphabet of 20 amino acids (the effect of correlations is reduced with the increase in the alphabet size 28 ). Under this assumption, Pj(Dictionary, n) is approximated as (1 - SpectralProbability(Dictionary)) n. Therefore, the probability that a Dictionary contains a peptide from a random protein Database can be approximated as 1 - (1 - SpectralProbability(Dictionary)) Database SpectralProbability(Dictionary) Database (under the condition that SpectralProbability(Dictionary) Database,1). 14 This condition is satisfied in practice since otherwise one ends up with spectral identifications that feature an unacceptably high FPR. Since we can represent a PSM by a Dictionary(Peptide, Spectrum), the probability of a Protein-Spectrum Match (with a random protein) can be similarly computed as SpectralProbability(Dictionary) Protein. Therefore, if one accepts peptides with spectral probability score SPThreshold and below (0 e SPThreshold e 1), then the probability of a match with a random Protein (spectral probability of Protein-Spectrum Matches) is approximated as SPThreshold Protein. Accounting for Protein Length. The score of a protein identification was previously computed as the score of its best scoring PSM, without taking into account the length of the protein. This is an oversimplification since longer proteins are more likely to have spurious matches than shorter ones. The spectral probability of Protein-Spectrum Matches (SpectralProbability(Dictionay) Protein ) suggests a natural normalization for computing protein-level FPR (note that while this normalization is applicable to the SpectralProbability score, it is not necessarily valid for other scoring functions). Figure 5 compares this Protein-Spectrum score (normalized by the protein length) with the original Peptide-Spectrum score (spectral probability before normalization) and shows that the proposed normalization makes sense (for both human and Shewanella data sets). While this normalization does not result in a large change in the sensitivity-specificity tradeoff (since mostproteinshavesimilarlengths),thenormalizedprotein-spectrum scoring function does show a modest increase in the number of protein identifications in the target database. For example, for 20 proteins identified in the decoy database (human data set), the Peptide-Spectrum scoring identifies 838 proteins in target database, while the Protein-Spectrum scoring (normalized by the protein length) identifies 865 proteins. Accounting for the Size of Spectral Data Set. We now extend the spectral dictionary approach from ref 15 to evaluate the statistical significance of matches between entire spectral data set Spectra and a protein. Like protein length, the size of the spectral data set also affects the statistical significance of a protein identification. While large spectral data sets are more likely to produce spurious identifications, some protein identification tools do not correct for the spectral data set size. When computing the probability of a match below the score Figure 5. (a) Identification of proteins in the human data set using MS-GF scores, without and with length correction. (b) Similar plot as in panel (a) for Shewanella data set. threshold 0 e SPThreshold e 1 for a single spectrum (we remind the reader that smaller SpectralProbability scores correspond to better Peptide-Spectrum matches), one could correct for the size of the Protein as SPThreshold Protein. However, when estimating the number of hits for multiple spectra, a similar correction SPThreshold Spectra does not apply because of the correlations between the spectra within a typical spectral data set (e.g., multiple spectra of the same peptide). To address this problem, we introduce the notion of a spectral lexicon below. We start by defining the dictionary of a spectral data set as Dictionary(Spectra, Threshold) ) Dictionary(Spectrum, Threshold) Spectrum Spectra research articles A string from Dictionary is called redundant if its proper substring also belongs to Dictionary (e.g., PEPTIDE is redundant if PEPTID, or PTIDE, or any of the shorter substrings belong to the dictionary). We define Lexicon(Dictionary(Spectra, Threshold)), represented more succinctly as Lexicon(Spectra, Threshold), as the set obtained by removal of all redundant strings from Dictionary(Spectra, Threshold). While Lexicon- (Spectra, Threshold) combines dictionaries of all spectra in the data set, for all practical purposes it can be treated as a Journal of Proteome Research Vol. 8, No. 9,
6 research articles dictionary of a single (virtual) spectrum. Moreover, the probability that a Lexicon contains a random peptide (i.e., that a spectrum from Spectra matches a random peptide with a score equal to or better than Threshold) is again given by SpectralProbability(Lexicon). Similarly, the probability that a Lexicon contains a peptide from a random Protein (i.e., the probability of a Protein-Spectra Match) can be approximated as 1 - (1 - SpectralProbability(Lexicon)) Protein 1 - e -SpectralProbability(Lexicon) Protein. Again, if SpectralProbability- (Lexicon) Protein,1, this probability can be approximated as SpectralProbability(Lexicon) Protein. We remark that while the expression 1 - (1 - SpectralProbability(Lexicon)) Protein is an approximation (the exact formula is given in 28 ), it nevertheless leads to a reasonable estimate of Protein-Spectra FPR in practice (see next section). While this simplified formula involves multiple levels of approximations, it remains reasonable for small values of spectral probability (such as or lower), for the typical sizes of spectral data sets (10 6 ) and protein lengths (10 3 ). For analyzing larger data sets, we recommend using more stringent (smaller) thresholds for spectral probability. While SpectralProbability(Lexicon) evaluates the statistical significance of Protein-Spectra matches (protein-level FPR), it remains unclear how to compute it. To address this problem, we define the notion of Compression of a spectral data set as a way to analyze dependencies between spectra. Loosely speaking, Compression is the ratio of the spectral probability of the lexicon (of a spectral data set) to the sum of spectral probabilities of individual dictionaries (of each spectrum). The ratio will often be less than 1 due to removal of redundant peptides from the lexicon and to the fact that many individual dictionaries share the same peptides. More precisely, Compression(Spectra, SPThreshold)) SpectralProbability(Lexicon(Spectra, SPThreshold)) SpectralProbability(Dictionary(Spectrum, SPThreshold)) Spectrum Spectra Since we use SpectralProbability as the scoring function, SpectralProbability(Dictionary(Spectrum, SPThreshold)) ) SPThreshold and therefore, Compression(Spectra, SPThreshold) ) SpectralProbability(Lexicon(Spectra, SPThreshold)) Spectra SPThreshold Two spectra are called independent if their dictionaries do not overlap and the union of their dictionaries does not contain redundant peptides. If all spectra in the spectral data set were independent, then Compression ) 1 and the quantity Spectra SPThreshold would provide a rigorous solution to the problem of the statistical significance of Peptide-Spectra Matches. In reality, however, spectra of the same and related peptides (e.g., peptides that represent subpeptides of other peptides) that are typically present in the spectral data set reduce Compression. While it can be explicitly computed for small spectral data sets, its efficient evaluation for large spectral data sets remains an open problem. Spectral Compression and Spectral Clustering. To estimate Compression of a spectral data set, we selected all high-accuracy Table 2. Compression for Different Values of Spectral Probability Threshold (SPThreshold) on a Small Spectral Data Set (Spectra) Consisting of Spectra, with Parent Mass between 1090 and 1100 Da a SPThreshold (FT) Shewanella spectra with parent mass varying between 1090 and 1100 Da resulting in the data set Spectra with spectra. For different values of SPThreshold, Dictionary(Spectrum, SPThreshold) were generated for each spectrum in Spectra and combined into Lexicon(Spectra, SPThreshold). Since all peptides in these dictionaries have similar parent mass (and thus do not have redundant peptides), Lexicon can be generated by simply taking a union of spectral dictionaries of individual spectra. We find that Compression values vary between 0.6 and 0.8 depending on the spectral probability threshold (Table 2). This experiment indicates that, while the expression Spectra SPThreshold overestimates FPR, it is still within 60-80% of the correct estimate, a reasonable approximation. Below we describe an alternative approach to estimating Compression via spectral clustering. We assume that a spectral sample Spectra represents the set of peptides Peptides. While the set Peptides is unknown, its size Peptides can be estimated by MS-Clustering tool 29 as the number of spectral clusters. Under the ideal scenario, the spectra of the same peptide (belonging to one cluster) have identical spectral dictionaries, spectra of different peptides do not overlap, and all spectra are independent. Under these assumptions, Therefore, S Spectra Dictionary (S, SPThreshold) Lexicon(Spectra, SPThreshold) Compression a The second column represents the sum of the sizes of the dictionaries of individual spectra and the third column represents the size of the combined Lexicon. SpectralProbability(Lexicon(Spectra, SPThreshold)) ) Peptides SPThreshold Compression ) Gupta and Pevzner Peptides SPThreshold Spectra SPThreshold ) Peptides Spectra In reality, dictionaries of spectra of the same peptides are not identical, dictionaries of spectra of different peptides may overlap, and spectra may be dependent. As a result, the parameter Peptides / Spectra typically underestimates Compression. For example, for the data set Spectra consisting of spectra with parent mass between 1090 and 1100 Da, MS- Clustering 29 found 3584 clusters from 8689 spectra, as shown in Table 3 (3928 out of spectra were discarded as lowquality spectra). The estimated Compression ) 3584/ is lower than the values observed in Table 2. Table 4 shows that large clusters have Compression much larger than 1/Cluster Size and illustrates that clustering underestimates Compression. While computing the FPR of Protein-Spectra matches enables one to evaluate the statistical significance of individual protein identifications, it also allows one to estimate the expected number of false identifications among all identifications, without requiring a decoy database. The following 4178 Journal of Proteome Research Vol. 8, No. 9, 2009
7 False Discovery Rates of Protein Identifications Table 3. Summary of Spectral Clusters Produced by MS-Clustering on the Set of Spectra with Parent Mass between 1090 and 1100 Da a average Compression cluster-size no. clusters no. spectra > Total a A total of 8689 spectra were clustered into 3584 clusters, while the remaining 3928 spectra were discarded by MS-Clustering due to low spectral quality. The last three columns indicate the average values of Compression for various cluster-sizes, for three different values of spectral probability threshold (SPThreshold). Table 4. Compression Computed for the 10 Largest Clusters in Table 3, Using Dictionaries Generated with the Spectral Probability Threshold of rank (by size) size Compression computational experiment shows that it is feasible to estimate the number of protein identifications in a decoy database without actually using a decoy database. The human spectra were searched with InsPecTxMS-GF against a database of randomly generated proteins of length 500 each, with equal probability of each amino acid at each position. A total of spectra (Spectra) got some InsPecT/MS-GF score against this database and were included in the analysis. Table 5 shows the observed number of proteins that have any peptide exceeding the score threshold (SPThreshold). The expected number of protein identifications, for different values of SPThreshold, is given by (1 - e -SpectralProbability(Lexicon) Protein ) where SpectralProbability(Lexicon) is estimated as Compression Spectra SPThreshold. This requires an estimate for the value of Compression for the spectral data set that we describe below. Suppose that the spectral data set Spectra is partitioned into nonoverlapping Clusters (i.e., Spectra ) Cluster Clusters Cluster), Table 5. Comparison between the Observed and the Expected Number of Protein Identifications for Different Values of Spectral Probability Threshold (SPThreshold), When Searching the Human Spectral Data Set against a Randomly Generated Decoy Database Consisting of Proteins of Size 500 Each a SPThreshold Compression expected no. IDs observed no. IDs a The values of Compression were estimated using the average values for different cluster-sizes from Table 3. where each Cluster represents the set of Cluster spectra originating from the same peptide. If we assume that Compression within each Sluster is 1/ Cluster, then SpectralProbability(Lexicon(Spectra, SPThreshold)) ) Clusters SPThreshold However, Table 4 reveals the limitations of this estimate. A more accurate estimate is given by Therefore, research articles SpectralProbability(Lexicon(Spectra, SPThreshold)) SpectralProbability(Lexicon(Cluster, SPThreshold)) Cluster Clusters Cluster Compression(Cluster, SPThreshold) Cluster Clusters SPThreshold Compression(Spectra, SPThreshold) Cluster Compression(Cluster, SPThreshold) Cluster Clusters Spectra While the parameter Compression(Cluster, SPThreshold) can be explicitly computed for each cluster, Table 3 shows that this parameter depends largely on the cluster size (and SPThreshold). Therefore, we can approximate Compression (Cluster, SPThreshold) by using the average value over the previously analyzed clusters of the same size. Using the cluster partitioning of the human data set determined by MS-Clustering, the expected number of proteins was estimated using the above formula by plugging in average Compression(Cluster, SPThreshold) values from Table 3. Table 5 compares this expected number of protein identifications with the number of proteins actually identified in the decoy database search and demonstrates that this approach allows one to get a reasonably close estimate of the number of false protein identifications without searching a decoy database. Using FPR of Protein Identifications. This paper recommends discarding the commonly used two-peptide rule and instead supports reporting protein identifications (including one-hit-wonders) according to their rigorous statistical significance (FPR). To illustrate this, Supplementary Table 1 provides a ranked list of protein identifications in the human data set, sorted by increasing FPRs. Denote the FPR of the i th ranked protein as FPR(i). Below we estimate the number of protein Journal of Proteome Research Vol. 8, No. 9,
8 research articles identifications in a decoy database (and thus FDR of protein identifications) if only proteins with FPRs equal to or exceeding FPR(i) are reported. If N is the total number of proteins in the sequence database (N ) for the human IPI database used here), one can estimate the expected number of false positives among top i protein identifications as False(i) ) N FPR(i), and therefore, the estimated FDR at that stringency level is FDR(i) ) (no. proteins in decoy database)/(no. proteins in target database) ) (N FPR(i))/(i). As i is increased from 1 to 600 in this data set, False(i) increases from 0 to 0.2 only, indicating that these top 600 protein identifications are almost error-free. It is worth noticing that 160 of these extremely reliable protein identifications are one-hit-wonders, a large fraction that may be missed by the traditional approaches favoring multiple peptides. When i is increased from 600 to 800, False(i) increases to 21, indicating that 1/10 ( 21/200) of the protein identifications in this range may be incorrect. Increasing i beyond 800 rapidly increases False(i), eventually at a rate higher than the rate of increase of i (see the last few rows in the table), thereby essentially reducing the number of true identifications. Therefore, one can choose to select the top 800 proteins in this list (at an FDR of 21/ %) as a reasonable set of protein identifications. This analysis shows how the knowledge of protein-level FPRs allows one to make informed judgment calls in selecting reliable protein identifications from many hits obtained in database searches. 4. Discussion We have demonstrated that the commonly used twopeptide rule jeopardizes the sensitivity-specificity tradeoff in protein identifications. This counterintuitive observation points out that we are more likely to get a protein with two mediocre peptide hits in the decoy database (by chance) than a single high-scoring peptide hit. Some software tools (e.g., Protein- Prophet, 21 Panoramics) 22 identify proteins if the combined score from all peptides exceeds the threshold, and thus allow scoring some single-hit proteins higher than proteins with multiple hits. However, these approaches are dependent on specific scoring models and require further theoretical justification to be suitable for use with all protein identification tools. In particular, our results indicate that the simpler singlepeptide rule results in better sensitivity/specificity tradeoff than ProteinProphet. One might expect that, since proteins are inferred from peptides, optimizing peptide identifications also optimizes protein identifications. We have provided evidence that this intuition does not hold ground. In reality, when one attempts to maximize the number of peptides in the target database, the additional peptides often come from the already covered proteins (i.e., from proteins with more than 1 match) and thus do not increase the overall number of protein identifications. However, the corresponding increase in the number of peptides in the decoy database significantly raises the number of decoy protein identifications (and thus increases FDR). Therefore, we emphasize that optimizing FDRs for peptides and proteins must be considered as different problems that are best addressed by different approaches. This study does not recommend accepting all proteins with single peptide hits but instead argues that the single-peptide approach must be used in conjunction with control of the FDR. While it may be surprising that the single-peptide approach generates a larger set of identified proteins than the seemingly more reliable two-peptide approach (without sacrificing FDR), our results indicate that it is the case. We demonstrated that for any set of proteins identified by the two-peptide approach (with peptide score threshold x), there is a larger set of protein identified by the single-peptide approach with the same FDR (with a more stringent peptide score threshold x + ɛ). Therefore, one has to choose the peptide-level score thresholds carefully to ensure that the single-peptide approach and two-peptide approach are being used for the same level of FDR. We also discussed how to estimate the FPR of individual Protein-Spectra matches using the generating function framework. Spectral clustering appears as a promising, albeit an indirect, approach for approximating spectral compression and should be explored further. While the proteomics community often takes great care in evaluating peptide-level error rates, the protein-level FDRs and FPRs are rarely computed. We suggest that publications reporting protein identifications should also report protein-level FPRs and/or FDRs. Lack of this checkpoint raises concerns about the validity of studies, such as biomarker discovery, where the number of identified proteins as well as the reliability of each individual identification is of critical importance. Acknowledgment. This work was supported by NIH 5R01RR and 1-P41-RR grants. We are indebted to Yong Li, Nuno Bandeira, and Sangtae Kim for many helpful discussions. We are grateful to Sangtae Kim for helping with MS-Dictionary, Ari Frank for helping with MS-Clustering, Seungjin Na and Jongwha Joo for help in running ProteinProphet, and Richard D. Smith, Vivian Hook, and Steven Bark for making MS/MS data sets generated in their laboratories available. Supporting Information Available: Table showing ranked list of protein identifications in the human database sorted by their increasing FPR. This material is available free of charge via the Internet at References Gupta and Pevzner (1) Aebersold, R.; Mann, M. Mass spectrometry-based proteomics. Nature 2003, 422, (2) Cargile, B. J.; Bundy, J. L.; Stephenson, J. L., Jr. Potential for false positive identifications from large databases through tandem mass spectrometry. J. Proteome Res. 2004, 3, (3) Elias, J. E.; Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 2007, 4, (4) Kall, L.; Storey, J. D.; MacCoss, M. J.; Noble, S. W. Assigning significance to peptides identified by tandem mass spectrometry using decoy databases. J. Proteome Res. 2008, 7, (5) Omenn, G. S.; States, D. J.; Adamski, M.; Blackwell, T. W.; Menon, R.; Hermjakob, H.; Apweiler, R.; Haab, B. B.; Simpson, R. J.; Eddes, J. S. Overview of the HUPO Plasma Proteome Project: results from the pilot phase with 35 collaborating laboratories and multiple analytical groups, generating a core dataset of 3020 proteins and a publicly-available database. Proteomics 2005, 5, (6) Carr, S.; Aebersold, R.; Baldwin, M.; Burlingame, A.; Clauser, K.; Nesvizhskii, A. The Need for Guidelines in Publication of Peptide and Protein Identification Data: Working Group on Publication Guidelines for Peptide and Protein Identification Data. Mol. Cell. Proteomics 2004, 3, (7) Bradshaw, R. A.; Burlingame, A. L.; Carr, S.; Aebersold, R. Reporting protein identification data: the next generation of guidelines. Mol. Cell. Proteomics 2006, 5, (8) Higdon, R.; Kolker, E. A predictive model for identifying proteins by a single peptide match. Bioinformatics 2007, 23, (9) Gupta, N.; Tanner, S.; Jaitly, N.; Adkins, J.; Lipton, M.; Edwards, R.; Romine, M.; Osterman, A.; Bafna, V.; Smith, R. D.; Pevzner, P. A. Whole proteome analysis of post-translational modifications: 4180 Journal of Proteome Research Vol. 8, No. 9, 2009
9 False Discovery Rates of Protein Identifications applications of mass-spectrometry for proteogenomic annotation. Genome Res. 2007, 17, (10) Gupta, N.; Benhamida, J.; Bhargava, V.; Goodman, D.; Kain, E.; Nguyen, N.; Ollikainen, N.; Rodriguez, J.; Wang, J.; Lipton, M. S.; Romine, M.; Bafna, V.; Smith, R. D.; Pevzner., P. A. Comparative proteogenomics: combining mass spectrometry and comparative genomics to analyze multiple genomes. Genome Res. 2008, 18, (11) Lu, P.; Vogel, C.; Wang, R.; Yao, X.; Marcotte, E. M. Absolute protein expression profiling estimates the relative contributions of transcriptional and translational regulation. Nat. Biotechnol. 2007, 25, (12) Boonen, K.; Landuyt, B.; Baggerman, G.; Husson, S. J.; Huybrechts, J.; Schoofs, L. Peptidomics: the integrated approach of MS, hyphenated techniques and bioinformatics for neuropeptide analysis. J. Sep. Sci. 2008, 31, (13) Falth, M.; Skold, K.; Norrman, M.; Svensson, M.; Fenyo, D.; Andren., P. E. SwePep, a database designed for endogenous peptides and mass spectrometry. Mol. Cell. Proteomics 2006, 5, (14) Kim, S.; Gupta, N.; Pevzner, P. A. Spectral probabilities and generating functions of tandem mass spectra: a strike against decoy databases. J. Proteome Res. 2008, 7, (15) Kim, S.; Gupta, N.; Banderia, N.; Pevzner, P. A. Spectral dictionaries: Integrating de novo peptide sequencing with database search of tandem mass spectra. Mol. Cell. Proteomics 2009, 8, (16) Gupta, N. Bark, S. J. Lu, W. D. Taupenot, L. O Connor, D. T. Pevzner, P. A. Hook, V. Evaluation of alternative neuropeptide processing in human and bovine dense-core secretory granules by mass spectrometry-based neuropeptidomics. Manuscript in preparation, (17) Tanner, S.; Shu, H.; Frank, A.; Wang, L. C.; Zandi, E.; Mumby, M.; Pevzner, P. A.; Bafna, V. InsPecT: identification of posttranslationally modified peptides from tandem mass spectra. Anal. Chem. 2005, 77, (18) Craig, R.; Beavis., R. C. TANDEM: matching proteins with tandem mass-spectra. Bioinformatics 2004, 20, (19) Keller, A.; Nesvizhskii, A. I.; Kolker, E.; Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 2002, 74, (20) Kall, L.; Canterbury, J. D.; Weston, J.; Noble, W. S.; MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 2007, 4, (21) Nesvizhskii, A. I.; Keller, A.; Kolker, E.; Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 2003, 75, (22) Feng, J.; Naiman, D. Q.; Cooper, B. Probability model for assessing proteins assembled from peptide sequences inferred from tandem mass spectrometry data. Anal. Chem. 2007, 79, (23) Moore, R. E.; Young, M. K.; Lee, T. D. Qscore: an algorithm for evaluating SEQUEST database search results. J. Am. Soc. Mass Spectrom. 2002, 13, (24) Weatherly, D. B.; Atwood, J. A.; Minning, T. A.; Cavola, C.; Tarleton, R. L.; Orlando, R. A Heuristic Method for Assigning a Falsediscovery Rate for Protein Identifications from Mascot Database Search Results. Mol. Cell. Proteomics 2005, 4, (25) Nesvizhskii, A. I.; Aebersold., R. Interpretation of shotgun proteomic data: the protein inference problem. Mol. Cell. Proteomics 2005, 4, (26) Alves, P.; Arnold, R. J.; Novotny, M. V.; Radivojac, P.; Reilly, J. P.; Tang, H. Advancement in protein inference from shotgun proteomics using peptide detectability. Pac. Symp. Biocomput. 2007, (27) Li, Y. F.; Arnold, R. J.; Li, Y.; Radivojac, P.; Sheng, Q.; Tang, H. A Bayesian approach to protein inference problem in shotgun proteomics. In Research in Computational Molecular Biology; Springer: Berlin, 2008; pp (28) Guibas, L. J.; Odlyzko., A. M. String overlaps, pattern matching, and nontransitive games. J. Comb. Theory, Ser. A 1981, 30, (29) Frank, A. M.; Bandeira, N.; Shen, Z.; Tanner, S.; Briggs, S. P.; Smith, R. D.; Pevzner, P. A. Clustering millions of tandem mass spectra. J. Proteome Res. 2008, 7, PR research articles Journal of Proteome Research Vol. 8, No. 9,
Supplementary Material for: Clustering Millions of Tandem Mass Spectra
 Supplementary Material for: Clustering Millions of Tandem Mass Spectra Ari M. Frank 1 Nuno Bandeira 1 Zhouxin Shen 2 Stephen Tanner 3 Steven P. Briggs 2 Richard D. Smith 4 Pavel A. Pevzner 1 October 4,
Supplementary Material for: Clustering Millions of Tandem Mass Spectra Ari M. Frank 1 Nuno Bandeira 1 Zhouxin Shen 2 Stephen Tanner 3 Steven P. Briggs 2 Richard D. Smith 4 Pavel A. Pevzner 1 October 4,
Overview - MS Proteomics in One Slide. MS masses of peptides. MS/MS fragments of a peptide. Results! Match to sequence database
 Overview - MS Proteomics in One Slide Obtain protein Digest into peptides Acquire spectra in mass spectrometer MS masses of peptides MS/MS fragments of a peptide Results! Match to sequence database 2 But
Overview - MS Proteomics in One Slide Obtain protein Digest into peptides Acquire spectra in mass spectrometer MS masses of peptides MS/MS fragments of a peptide Results! Match to sequence database 2 But
Tandem Mass Spectrometry: Generating function, alignment and assembly
 Tandem Mass Spectrometry: Generating function, alignment and assembly With slides from Sangtae Kim and from Jones & Pevzner 2004 Determining reliability of identifications Can we use Target/Decoy to estimate
Tandem Mass Spectrometry: Generating function, alignment and assembly With slides from Sangtae Kim and from Jones & Pevzner 2004 Determining reliability of identifications Can we use Target/Decoy to estimate
Modeling Mass Spectrometry-Based Protein Analysis
 Chapter 8 Jan Eriksson and David Fenyö Abstract The success of mass spectrometry based proteomics depends on efficient methods for data analysis. These methods require a detailed understanding of the information
Chapter 8 Jan Eriksson and David Fenyö Abstract The success of mass spectrometry based proteomics depends on efficient methods for data analysis. These methods require a detailed understanding of the information
In shotgun proteomics, a complex protein mixture derived from a biological sample is directly analyzed. Research Article
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 16, Number 8, 2009 # Mary Ann Liebert, Inc. Pp. 1 11 DOI: 10.1089/cmb.2009.0018 Research Article A Bayesian Approach to Protein Inference Problem in Shotgun Proteomics
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 16, Number 8, 2009 # Mary Ann Liebert, Inc. Pp. 1 11 DOI: 10.1089/cmb.2009.0018 Research Article A Bayesian Approach to Protein Inference Problem in Shotgun Proteomics
Key questions of proteomics. Bioinformatics 2. Proteomics. Foundation of proteomics. What proteins are there? Protein digestion
 s s Key questions of proteomics What proteins are there? Bioinformatics 2 Lecture 2 roteomics How much is there of each of the proteins? - Absolute quantitation - Stoichiometry What (modification/splice)
s s Key questions of proteomics What proteins are there? Bioinformatics 2 Lecture 2 roteomics How much is there of each of the proteins? - Absolute quantitation - Stoichiometry What (modification/splice)
Mass spectrometry in proteomics
 I519 Introduction to Bioinformatics, Fall, 2013 Mass spectrometry in proteomics Haixu Tang School of Informatics and Computing Indiana University, Bloomington Modified from: www.bioalgorithms.info Outline
I519 Introduction to Bioinformatics, Fall, 2013 Mass spectrometry in proteomics Haixu Tang School of Informatics and Computing Indiana University, Bloomington Modified from: www.bioalgorithms.info Outline
ADVANCEMENT IN PROTEIN INFERENCE FROM SHOTGUN PROTEOMICS USING PEPTIDE DETECTABILITY
 ADVANCEMENT IN PROTEIN INFERENCE FROM SHOTGUN PROTEOMICS USING PEPTIDE DETECTABILITY PEDRO ALVES, 1 RANDY J. ARNOLD, 2 MILOS V. NOVOTNY, 2 PREDRAG RADIVOJAC, 1 JAMES P. REILLY, 2 HAIXU TANG 1, 3* 1) School
ADVANCEMENT IN PROTEIN INFERENCE FROM SHOTGUN PROTEOMICS USING PEPTIDE DETECTABILITY PEDRO ALVES, 1 RANDY J. ARNOLD, 2 MILOS V. NOVOTNY, 2 PREDRAG RADIVOJAC, 1 JAMES P. REILLY, 2 HAIXU TANG 1, 3* 1) School
A NESTED MIXTURE MODEL FOR PROTEIN IDENTIFICATION USING MASS SPECTROMETRY
 Submitted to the Annals of Applied Statistics A NESTED MIXTURE MODEL FOR PROTEIN IDENTIFICATION USING MASS SPECTROMETRY By Qunhua Li,, Michael MacCoss and Matthew Stephens University of Washington and
Submitted to the Annals of Applied Statistics A NESTED MIXTURE MODEL FOR PROTEIN IDENTIFICATION USING MASS SPECTROMETRY By Qunhua Li,, Michael MacCoss and Matthew Stephens University of Washington and
Learning Score Function Parameters for Improved Spectrum Identification in Tandem Mass Spectrometry Experiments
 pubs.acs.org/jpr Learning Score Function Parameters for Improved Spectrum Identification in Tandem Mass Spectrometry Experiments Marina Spivak, Michael S. Bereman, Michael J. MacCoss, and William Stafford
pubs.acs.org/jpr Learning Score Function Parameters for Improved Spectrum Identification in Tandem Mass Spectrometry Experiments Marina Spivak, Michael S. Bereman, Michael J. MacCoss, and William Stafford
A statistical approach to peptide identification from clustered tandem mass spectrometry data
 A statistical approach to peptide identification from clustered tandem mass spectrometry data Soyoung Ryu, David R. Goodlett, William S. Noble and Vladimir N. Minin Department of Statistics, University
A statistical approach to peptide identification from clustered tandem mass spectrometry data Soyoung Ryu, David R. Goodlett, William S. Noble and Vladimir N. Minin Department of Statistics, University
PeptideProphet: Validation of Peptide Assignments to MS/MS Spectra
 PeptideProphet: Validation of Peptide Assignments to MS/MS Spectra Andrew Keller Day 2 October 17, 2006 Andrew Keller Rosetta Bioinformatics, Seattle Outline Need to validate peptide assignments to MS/MS
PeptideProphet: Validation of Peptide Assignments to MS/MS Spectra Andrew Keller Day 2 October 17, 2006 Andrew Keller Rosetta Bioinformatics, Seattle Outline Need to validate peptide assignments to MS/MS
Was T. rex Just a Big Chicken? Computational Proteomics
 Was T. rex Just a Big Chicken? Computational Proteomics Phillip Compeau and Pavel Pevzner adjusted by Jovana Kovačević Bioinformatics Algorithms: an Active Learning Approach 215 by Compeau and Pevzner.
Was T. rex Just a Big Chicken? Computational Proteomics Phillip Compeau and Pavel Pevzner adjusted by Jovana Kovačević Bioinformatics Algorithms: an Active Learning Approach 215 by Compeau and Pevzner.
Protein inference based on peptides identified from. tandem mass spectra
 Protein inference based on peptides identified from tandem mass spectra A Thesis Submitted to the College of Graduate Studies and Research in Partial Fulfillment of the Requirements for the degree of Doctor
Protein inference based on peptides identified from tandem mass spectra A Thesis Submitted to the College of Graduate Studies and Research in Partial Fulfillment of the Requirements for the degree of Doctor
MSblender: a probabilistic approach for integrating peptide identifications from multiple database search engines
 Article Subscriber access provided by University of Texas Libraries MSblender: a probabilistic approach for integrating peptide identifications from multiple database search engines Taejoon Kwon, Hyungwon
Article Subscriber access provided by University of Texas Libraries MSblender: a probabilistic approach for integrating peptide identifications from multiple database search engines Taejoon Kwon, Hyungwon
DE NOVO PEPTIDE SEQUENCING FOR MASS SPECTRA BASED ON MULTI-CHARGE STRONG TAGS
 DE NOVO PEPTIDE SEQUENCING FO MASS SPECTA BASED ON MULTI-CHAGE STONG TAGS KANG NING, KET FAH CHONG, HON WAI LEONG Department of Computer Science, National University of Singapore, 3 Science Drive 2, Singapore
DE NOVO PEPTIDE SEQUENCING FO MASS SPECTA BASED ON MULTI-CHAGE STONG TAGS KANG NING, KET FAH CHONG, HON WAI LEONG Department of Computer Science, National University of Singapore, 3 Science Drive 2, Singapore
Mass Spectrometry and Proteomics - Lecture 5 - Matthias Trost Newcastle University
 Mass Spectrometry and Proteomics - Lecture 5 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Proteomics Sample prep 144 Lecture 5 Quantitation techniques Search Algorithms Proteomics
Mass Spectrometry and Proteomics - Lecture 5 - Matthias Trost Newcastle University matthias.trost@ncl.ac.uk Previously Proteomics Sample prep 144 Lecture 5 Quantitation techniques Search Algorithms Proteomics
Identification of proteins by enzyme digestion, mass
 Method for Screening Peptide Fragment Ion Mass Spectra Prior to Database Searching Roger E. Moore, Mary K. Young, and Terry D. Lee Beckman Research Institute of the City of Hope, Duarte, California, USA
Method for Screening Peptide Fragment Ion Mass Spectra Prior to Database Searching Roger E. Moore, Mary K. Young, and Terry D. Lee Beckman Research Institute of the City of Hope, Duarte, California, USA
Transferred Subgroup False Discovery Rate for Rare Post-translational Modifications Detected by Mass Spectrometry* S
 Transferred Subgroup False Discovery Rate for Rare Post-translational Modifications Detected by Mass Spectrometry* S Yan Fu and Xiaohong Qian Technological Innovation and Resources 2014 by The American
Transferred Subgroup False Discovery Rate for Rare Post-translational Modifications Detected by Mass Spectrometry* S Yan Fu and Xiaohong Qian Technological Innovation and Resources 2014 by The American
PepHMM: A Hidden Markov Model Based Scoring Function for Mass Spectrometry Database Search
 PepHMM: A Hidden Markov Model Based Scoring Function for Mass Spectrometry Database Search Yunhu Wan, Austin Yang, and Ting Chen*, Department of Mathematics, Department of Pharmaceutical Sciences, and
PepHMM: A Hidden Markov Model Based Scoring Function for Mass Spectrometry Database Search Yunhu Wan, Austin Yang, and Ting Chen*, Department of Mathematics, Department of Pharmaceutical Sciences, and
De Novo Peptide Identification Via Mixed-Integer Linear Optimization And Tandem Mass Spectrometry
 17 th European Symposium on Computer Aided Process Engineering ESCAPE17 V. Plesu and P.S. Agachi (Editors) 2007 Elsevier B.V. All rights reserved. 1 De Novo Peptide Identification Via Mixed-Integer Linear
17 th European Symposium on Computer Aided Process Engineering ESCAPE17 V. Plesu and P.S. Agachi (Editors) 2007 Elsevier B.V. All rights reserved. 1 De Novo Peptide Identification Via Mixed-Integer Linear
MS-MS Analysis Programs
 MS-MS Analysis Programs Basic Process Genome - Gives AA sequences of proteins Use this to predict spectra Compare data to prediction Determine degree of correctness Make assignment Did we see the protein?
MS-MS Analysis Programs Basic Process Genome - Gives AA sequences of proteins Use this to predict spectra Compare data to prediction Determine degree of correctness Make assignment Did we see the protein?
HOWTO, example workflow and data files. (Version )
 HOWTO, example workflow and data files. (Version 20 09 2017) 1 Introduction: SugarQb is a collection of software tools (Nodes) which enable the automated identification of intact glycopeptides from HCD
HOWTO, example workflow and data files. (Version 20 09 2017) 1 Introduction: SugarQb is a collection of software tools (Nodes) which enable the automated identification of intact glycopeptides from HCD
Computational Methods for Mass Spectrometry Proteomics
 Computational Methods for Mass Spectrometry Proteomics Eidhammer, Ingvar ISBN-13: 9780470512975 Table of Contents Preface. Acknowledgements. 1 Protein, Proteome, and Proteomics. 1.1 Primary goals for studying
Computational Methods for Mass Spectrometry Proteomics Eidhammer, Ingvar ISBN-13: 9780470512975 Table of Contents Preface. Acknowledgements. 1 Protein, Proteome, and Proteomics. 1.1 Primary goals for studying
Introduction to pepxmltab
 Introduction to pepxmltab Xiaojing Wang October 30, 2018 Contents 1 Introduction 1 2 Convert pepxml to a tabular format 1 3 PSMs Filtering 4 4 Session Information 5 1 Introduction Mass spectrometry (MS)-based
Introduction to pepxmltab Xiaojing Wang October 30, 2018 Contents 1 Introduction 1 2 Convert pepxml to a tabular format 1 3 PSMs Filtering 4 4 Session Information 5 1 Introduction Mass spectrometry (MS)-based
Peptide Sequence Tags for Fast Database Search in Mass-Spectrometry
 Peptide Sequence Tags for Fast Database Search in Mass-Spectrometry Ari Frank,*, Stephen Tanner, Vineet Bafna, and Pavel Pevzner Department of Computer Science & Engineering, University of California,
Peptide Sequence Tags for Fast Database Search in Mass-Spectrometry Ari Frank,*, Stephen Tanner, Vineet Bafna, and Pavel Pevzner Department of Computer Science & Engineering, University of California,
Efficient Marginalization to Compute Protein Posterior Probabilities from Shotgun Mass Spectrometry Data
 Efficient Marginalization to Compute Protein Posterior Probabilities from Shotgun Mass Spectrometry Data Oliver Serang Department of Genome Sciences, University of Washington, Seattle, Washington Michael
Efficient Marginalization to Compute Protein Posterior Probabilities from Shotgun Mass Spectrometry Data Oliver Serang Department of Genome Sciences, University of Washington, Seattle, Washington Michael
Properties of Average Score Distributions of SEQUEST
 Research Properties of Average Score Distributions of SEQUEST THE PROBABILITY RATIO METHOD* S Salvador Martínez-Bartolomé, Pedro Navarro, Fernando Martín-Maroto, Daniel López-Ferrer **, Antonio Ramos-Fernández,
Research Properties of Average Score Distributions of SEQUEST THE PROBABILITY RATIO METHOD* S Salvador Martínez-Bartolomé, Pedro Navarro, Fernando Martín-Maroto, Daniel López-Ferrer **, Antonio Ramos-Fernández,
Efficiency of Database Search for Identification of Mutated and Modified Proteins via Mass Spectrometry
 Methods Efficiency of Database Search for Identification of Mutated and Modified Proteins via Mass Spectrometry Pavel A. Pevzner, 1,3 Zufar Mulyukov, 1 Vlado Dancik, 2 and Chris L Tang 2 Department of
Methods Efficiency of Database Search for Identification of Mutated and Modified Proteins via Mass Spectrometry Pavel A. Pevzner, 1,3 Zufar Mulyukov, 1 Vlado Dancik, 2 and Chris L Tang 2 Department of
iprophet: Multi-level integrative analysis of shotgun proteomic data improves peptide and protein identification rates and error estimates
 MCP Papers in Press. Published on August 29, 2011 as Manuscript M111.007690 This is the Pre-Published Version iprophet: Multi-level integrative analysis of shotgun proteomic data improves peptide and protein
MCP Papers in Press. Published on August 29, 2011 as Manuscript M111.007690 This is the Pre-Published Version iprophet: Multi-level integrative analysis of shotgun proteomic data improves peptide and protein
Workflow concept. Data goes through the workflow. A Node contains an operation An edge represents data flow The results are brought together in tables
 PROTEOME DISCOVERER Workflow concept Data goes through the workflow Spectra Peptides Quantitation A Node contains an operation An edge represents data flow The results are brought together in tables Protein
PROTEOME DISCOVERER Workflow concept Data goes through the workflow Spectra Peptides Quantitation A Node contains an operation An edge represents data flow The results are brought together in tables Protein
De novo Protein Sequencing by Combining Top-Down and Bottom-Up Tandem Mass Spectra. Xiaowen Liu
 De novo Protein Sequencing by Combining Top-Down and Bottom-Up Tandem Mass Spectra Xiaowen Liu Department of BioHealth Informatics, Department of Computer and Information Sciences, Indiana University-Purdue
De novo Protein Sequencing by Combining Top-Down and Bottom-Up Tandem Mass Spectra Xiaowen Liu Department of BioHealth Informatics, Department of Computer and Information Sciences, Indiana University-Purdue
HMMatch: Peptide Identification by Spectral Matching. of Tandem Mass Spectra using Hidden Markov Models
 HMMatch: Peptide Identification by Spectral Matching of Tandem Mass Spectra using Hidden Markov Models Xue Wu 1 Chau-Wen Tseng 1 Nathan Edwards 2 1 Department of Computer Science University of Maryland,
HMMatch: Peptide Identification by Spectral Matching of Tandem Mass Spectra using Hidden Markov Models Xue Wu 1 Chau-Wen Tseng 1 Nathan Edwards 2 1 Department of Computer Science University of Maryland,
Quality Assessment of Tandem Mass Spectra Based on Cumulative Intensity Normalization
 Quality Assessment of Tandem Mass Spectra Based on Cumulative Intensity Normalization Seungjin Na and Eunok Paek* Department of Mechanical and Information Engineering, University of Seoul, Seoul, Korea
Quality Assessment of Tandem Mass Spectra Based on Cumulative Intensity Normalization Seungjin Na and Eunok Paek* Department of Mechanical and Information Engineering, University of Seoul, Seoul, Korea
Identification of Post-translational Modifications via Blind Search of Mass-Spectra
 Identification of Post-translational Modifications via Blind Search of Mass-Spectra Dekel Tsur Computer Science and Engineering UC San Diego dtsur@cs.ucsd.edu Vineet Bafna Computer Science and Engineering
Identification of Post-translational Modifications via Blind Search of Mass-Spectra Dekel Tsur Computer Science and Engineering UC San Diego dtsur@cs.ucsd.edu Vineet Bafna Computer Science and Engineering
An SVM Scorer for More Sensitive and Reliable Peptide Identification via Tandem Mass Spectrometry
 An SVM Scorer for More Sensitive and Reliable Peptide Identification via Tandem Mass Spectrometry Haipeng Wang, Yan Fu, Ruixiang Sun, Simin He, Rong Zeng, and Wen Gao Pacific Symposium on Biocomputing
An SVM Scorer for More Sensitive and Reliable Peptide Identification via Tandem Mass Spectrometry Haipeng Wang, Yan Fu, Ruixiang Sun, Simin He, Rong Zeng, and Wen Gao Pacific Symposium on Biocomputing
MaSS-Simulator: A highly configurable MS/MS simulator for generating test datasets for big data algorithms.
 MaSS-Simulator: A highly configurable MS/MS simulator for generating test datasets for big data algorithms. Muaaz Gul Awan 1 and Fahad Saeed 1 1 Department of Computer Science, Western Michigan University,
MaSS-Simulator: A highly configurable MS/MS simulator for generating test datasets for big data algorithms. Muaaz Gul Awan 1 and Fahad Saeed 1 1 Department of Computer Science, Western Michigan University,
Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries
 Anal. Chem. 2006, 78, 5678-5684 Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries Barbara E. Frewen, Gennifer E. Merrihew, Christine C. Wu, William Stafford
Anal. Chem. 2006, 78, 5678-5684 Analysis of Peptide MS/MS Spectra from Large-Scale Proteomics Experiments Using Spectrum Libraries Barbara E. Frewen, Gennifer E. Merrihew, Christine C. Wu, William Stafford
PeptideProphet: Validation of Peptide Assignments to MS/MS Spectra. Andrew Keller
 PeptideProphet: Validation of Peptide Assignments to MS/MS Spectra Andrew Keller Outline Need to validate peptide assignments to MS/MS spectra Statistical approach to validation Running PeptideProphet
PeptideProphet: Validation of Peptide Assignments to MS/MS Spectra Andrew Keller Outline Need to validate peptide assignments to MS/MS spectra Statistical approach to validation Running PeptideProphet
UC San Diego UC San Diego Electronic Theses and Dissertations
 UC San Diego UC San Diego Electronic Theses and Dissertations Title Algorithms for tandem mass spectrometry-based proteomics Permalink https://escholarship.org/uc/item/89f7x81r Author Frank, Ari Michael
UC San Diego UC San Diego Electronic Theses and Dissertations Title Algorithms for tandem mass spectrometry-based proteomics Permalink https://escholarship.org/uc/item/89f7x81r Author Frank, Ari Michael
Spectrum-to-Spectrum Searching Using a. Proteome-wide Spectral Library
 MCP Papers in Press. Published on April 30, 2011 as Manuscript M111.007666 Spectrum-to-Spectrum Searching Using a Proteome-wide Spectral Library Chia-Yu Yen, Stephane Houel, Natalie G. Ahn, and William
MCP Papers in Press. Published on April 30, 2011 as Manuscript M111.007666 Spectrum-to-Spectrum Searching Using a Proteome-wide Spectral Library Chia-Yu Yen, Stephane Houel, Natalie G. Ahn, and William
Speeding up Scoring Module of Mass Spectrometry Based Protein Identification by GPUs
 Speeding up Scoring Module of Mass Spectrometry Based Protein Identification by GPUs Li You Abstract Database searching is a main method for protein identification in shotgun proteomics, and till now most
Speeding up Scoring Module of Mass Spectrometry Based Protein Identification by GPUs Li You Abstract Database searching is a main method for protein identification in shotgun proteomics, and till now most
A Description of the CPTAC Common Data Analysis Pipeline (CDAP)
 A Description of the CPTAC Common Data Analysis Pipeline (CDAP) v. 01/14/2014 Summary The purpose of this document is to describe the software programs and output files of the Common Data Analysis Pipeline
A Description of the CPTAC Common Data Analysis Pipeline (CDAP) v. 01/14/2014 Summary The purpose of this document is to describe the software programs and output files of the Common Data Analysis Pipeline
Effective Strategies for Improving Peptide Identification with Tandem Mass Spectrometry
 Effective Strategies for Improving Peptide Identification with Tandem Mass Spectrometry by Xi Han A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree
Effective Strategies for Improving Peptide Identification with Tandem Mass Spectrometry by Xi Han A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree
PRIDE Cluster: building the consensus of proteomics data
 Supplementary Materials PRIDE Cluster: building the consensus of proteomics data Johannes Griss, Joseph Michael Foster, Henning Hermjakob and Juan Antonio Vizcaíno EMBL-European Bioinformatics Institute,
Supplementary Materials PRIDE Cluster: building the consensus of proteomics data Johannes Griss, Joseph Michael Foster, Henning Hermjakob and Juan Antonio Vizcaíno EMBL-European Bioinformatics Institute,
An Unsupervised, Model-Free, Machine-Learning Combiner for Peptide Identifications from Tandem Mass Spectra
 Clin Proteom (2009) 5:23 36 DOI 0.007/s204-009-9024-5 An Unsupervised, Model-Free, Machine-Learning Combiner for Peptide Identifications from Tandem Mass Spectra Nathan Edwards Xue Wu Chau-Wen Tseng Published
Clin Proteom (2009) 5:23 36 DOI 0.007/s204-009-9024-5 An Unsupervised, Model-Free, Machine-Learning Combiner for Peptide Identifications from Tandem Mass Spectra Nathan Edwards Xue Wu Chau-Wen Tseng Published
Introduction to spectral alignment
 SI Appendix C. Introduction to spectral alignment Due to the complexity of the anti-symmetric spectral alignment algorithm described in Appendix A, this appendix provides an extended introduction to the
SI Appendix C. Introduction to spectral alignment Due to the complexity of the anti-symmetric spectral alignment algorithm described in Appendix A, this appendix provides an extended introduction to the
Mass spectrometry has been used a lot in biology since the late 1950 s. However it really came into play in the late 1980 s once methods were
 Mass spectrometry has been used a lot in biology since the late 1950 s. However it really came into play in the late 1980 s once methods were developed to allow the analysis of large intact (bigger than
Mass spectrometry has been used a lot in biology since the late 1950 s. However it really came into play in the late 1980 s once methods were developed to allow the analysis of large intact (bigger than
Towards the Prediction of Protein Abundance from Tandem Mass Spectrometry Data
 Towards the Prediction of Protein Abundance from Tandem Mass Spectrometry Data Anthony J Bonner Han Liu Abstract This paper addresses a central problem of Proteomics: estimating the amounts of each of
Towards the Prediction of Protein Abundance from Tandem Mass Spectrometry Data Anthony J Bonner Han Liu Abstract This paper addresses a central problem of Proteomics: estimating the amounts of each of
NIH Public Access Author Manuscript IEEE/ACM Trans Comput Biol Bioinform. Author manuscript; available in PMC 2013 May 01.
 NIH Public Access Author Manuscript Published in final edited form as: IEEE/ACM Trans Comput Biol Bioinform. 2012 ; 9(3): 809 817. doi:10.1109/tcbb.2012.26. Faster Mass Spectrometry-based Protein Inference:
NIH Public Access Author Manuscript Published in final edited form as: IEEE/ACM Trans Comput Biol Bioinform. 2012 ; 9(3): 809 817. doi:10.1109/tcbb.2012.26. Faster Mass Spectrometry-based Protein Inference:
MSnID Package for Handling MS/MS Identifications
 Vladislav A. Petyuk December 1, 2018 Contents 1 Introduction.............................. 1 2 Starting the project.......................... 3 3 Reading MS/MS data........................ 3 4 Updating
Vladislav A. Petyuk December 1, 2018 Contents 1 Introduction.............................. 1 2 Starting the project.......................... 3 3 Reading MS/MS data........................ 3 4 Updating
NovoHMM: A Hidden Markov Model for de Novo Peptide Sequencing
 Anal. Chem. 2005, 77, 7265-7273 NovoHMM: A Hidden Markov Model for de Novo Peptide Sequencing Bernd Fischer, Volker Roth, Franz Roos, Jonas Grossmann, Sacha Baginsky, Peter Widmayer, Wilhelm Gruissem,
Anal. Chem. 2005, 77, 7265-7273 NovoHMM: A Hidden Markov Model for de Novo Peptide Sequencing Bernd Fischer, Volker Roth, Franz Roos, Jonas Grossmann, Sacha Baginsky, Peter Widmayer, Wilhelm Gruissem,
Nature Methods: doi: /nmeth Supplementary Figure 1. Fragment indexing allows efficient spectra similarity comparisons.
 Supplementary Figure 1 Fragment indexing allows efficient spectra similarity comparisons. The cost and efficiency of spectra similarity calculations can be approximated by the number of fragment comparisons
Supplementary Figure 1 Fragment indexing allows efficient spectra similarity comparisons. The cost and efficiency of spectra similarity calculations can be approximated by the number of fragment comparisons
JUMP: a tag-based database search tool for peptide identification with high sensitivity
 MCP Papers in Press. Published on September 8, 2014 as Manuscript O114.039586 JUMP: a tag-based database search tool for peptide identification with high sensitivity and accuracy Xusheng Wang 1, Yuxin
MCP Papers in Press. Published on September 8, 2014 as Manuscript O114.039586 JUMP: a tag-based database search tool for peptide identification with high sensitivity and accuracy Xusheng Wang 1, Yuxin
DIA-Umpire: comprehensive computational framework for data independent acquisition proteomics
 DIA-Umpire: comprehensive computational framework for data independent acquisition proteomics Chih-Chiang Tsou 1,2, Dmitry Avtonomov 2, Brett Larsen 3, Monika Tucholska 3, Hyungwon Choi 4 Anne-Claude Gingras
DIA-Umpire: comprehensive computational framework for data independent acquisition proteomics Chih-Chiang Tsou 1,2, Dmitry Avtonomov 2, Brett Larsen 3, Monika Tucholska 3, Hyungwon Choi 4 Anne-Claude Gingras
HMMatch: Peptide Identification by Spectral Matching of Tandem Mass Spectra Using Hidden Markov Models ABSTRACT
 JOURNAL OF COMPUTATIONAL BIOLOGY Volume 14, Number 8, 2007 Mary Ann Liebert, Inc. Pp. 1025 1043 DOI: 10.1089/cmb.2007.0071 HMMatch: Peptide Identification by Spectral Matching of Tandem Mass Spectra Using
JOURNAL OF COMPUTATIONAL BIOLOGY Volume 14, Number 8, 2007 Mary Ann Liebert, Inc. Pp. 1025 1043 DOI: 10.1089/cmb.2007.0071 HMMatch: Peptide Identification by Spectral Matching of Tandem Mass Spectra Using
Protein Sequencing and Identification by Mass Spectrometry
 Protein Sequencing and Identification by Mass Spectrometry Tandem Mass Spectrometry De Novo Peptide Sequencing Spectrum Graph Protein Identification via Database Search Identifying Post Translationally
Protein Sequencing and Identification by Mass Spectrometry Tandem Mass Spectrometry De Novo Peptide Sequencing Spectrum Graph Protein Identification via Database Search Identifying Post Translationally
The Pitfalls of Peaklist Generation Software Performance on Database Searches
 Proceedings of the 56th ASMS Conference on Mass Spectrometry and Allied Topics, Denver, CO, June 1-5, 2008 The Pitfalls of Peaklist Generation Software Performance on Database Searches Aenoch J. Lynn,
Proceedings of the 56th ASMS Conference on Mass Spectrometry and Allied Topics, Denver, CO, June 1-5, 2008 The Pitfalls of Peaklist Generation Software Performance on Database Searches Aenoch J. Lynn,
Improved Validation of Peptide MS/MS Assignments. Using Spectral Intensity Prediction
 MCP Papers in Press. Published on October 2, 2006 as Manuscript M600320-MCP200 Improved Validation of Peptide MS/MS Assignments Using Spectral Intensity Prediction Shaojun Sun 1, Karen Meyer-Arendt 2,
MCP Papers in Press. Published on October 2, 2006 as Manuscript M600320-MCP200 Improved Validation of Peptide MS/MS Assignments Using Spectral Intensity Prediction Shaojun Sun 1, Karen Meyer-Arendt 2,
Model Accuracy Measures
 Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
Model Accuracy Measures Master in Bioinformatics UPF 2017-2018 Eduardo Eyras Computational Genomics Pompeu Fabra University - ICREA Barcelona, Spain Variables What we can measure (attributes) Hypotheses
STATISTICAL METHODS FOR THE ANALYSIS OF MASS SPECTROMETRY- BASED PROTEOMICS DATA. A Dissertation XUAN WANG
 STATISTICAL METHODS FOR THE ANALYSIS OF MASS SPECTROMETRY- BASED PROTEOMICS DATA A Dissertation by XUAN WANG Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of
STATISTICAL METHODS FOR THE ANALYSIS OF MASS SPECTROMETRY- BASED PROTEOMICS DATA A Dissertation by XUAN WANG Submitted to the Office of Graduate Studies of Texas A&M University in partial fulfillment of
Mass Spectrometry Based De Novo Peptide Sequencing Error Correction
 Mass Spectrometry Based De Novo Peptide Sequencing Error Correction by Chenyu Yao A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree of Master of Mathematics
Mass Spectrometry Based De Novo Peptide Sequencing Error Correction by Chenyu Yao A thesis presented to the University of Waterloo in fulfillment of the thesis requirement for the degree of Master of Mathematics
BIOINFORMATICS ORIGINAL PAPER
 BIOINFORMATICS ORIGINAL PAPER Vol. 27 no. 6 2011, pages 797 806 doi:10.1093/bioinformatics/btr017 Gene expression Advance Access publication January 22, 2011 Computational refinement of post-translational
BIOINFORMATICS ORIGINAL PAPER Vol. 27 no. 6 2011, pages 797 806 doi:10.1093/bioinformatics/btr017 Gene expression Advance Access publication January 22, 2011 Computational refinement of post-translational
In order to compare the proteins of the phylogenomic matrix, we needed a similarity
 Similarity Matrix Generation In order to compare the proteins of the phylogenomic matrix, we needed a similarity measure. Hamming distances between phylogenetic profiles require the use of thresholds for
Similarity Matrix Generation In order to compare the proteins of the phylogenomic matrix, we needed a similarity measure. Hamming distances between phylogenetic profiles require the use of thresholds for
InsPecT: Identification of Posttranslationally Modified Peptides from Tandem Mass Spectra
 Anal. Chem. 2005, 77, 4626-4639 InsPecT: Identification of Posttranslationally Modified Peptides from Tandem Mass Spectra Stephen Tanner,*, Hongjun Shu, Ari Frank, Ling-Chi Wang, Ebrahim Zandi, Marc Mumby,
Anal. Chem. 2005, 77, 4626-4639 InsPecT: Identification of Posttranslationally Modified Peptides from Tandem Mass Spectra Stephen Tanner,*, Hongjun Shu, Ari Frank, Ling-Chi Wang, Ebrahim Zandi, Marc Mumby,
Intensity-based protein identification by machine learning from a library of tandem mass spectra
 Intensity-based protein identification by machine learning from a library of tandem mass spectra Joshua E Elias 1,Francis D Gibbons 2,Oliver D King 2,Frederick P Roth 2,4 & Steven P Gygi 1,3,4 Tandem mass
Intensity-based protein identification by machine learning from a library of tandem mass spectra Joshua E Elias 1,Francis D Gibbons 2,Oliver D King 2,Frederick P Roth 2,4 & Steven P Gygi 1,3,4 Tandem mass
De Novo Peptide Sequencing: Informatics and Pattern Recognition applied to Proteomics
 De Novo Peptide Sequencing: Informatics and Pattern Recognition applied to Proteomics John R. Rose Computer Science and Engineering University of South Carolina 1 Overview Background Information Theoretic
De Novo Peptide Sequencing: Informatics and Pattern Recognition applied to Proteomics John R. Rose Computer Science and Engineering University of South Carolina 1 Overview Background Information Theoretic
PepHMM: A Hidden Markov Model Based Scoring Function for Mass Spectrometry Database Search
 PepHMM: A Hidden Markov Model Based Scoring Function for Mass Spectrometry Database Search Yunhu Wan, Austin Yang, and Ting Chen Department of Mathematics, Department of Pharmaceutical Sciences, Department
PepHMM: A Hidden Markov Model Based Scoring Function for Mass Spectrometry Database Search Yunhu Wan, Austin Yang, and Ting Chen Department of Mathematics, Department of Pharmaceutical Sciences, Department
Biological Mass Spectrometry
 Biochemistry 412 Biological Mass Spectrometry February 13 th, 2007 Proteomics The study of the complete complement of proteins found in an organism Degrees of Freedom for Protein Variability Covalent Modifications
Biochemistry 412 Biological Mass Spectrometry February 13 th, 2007 Proteomics The study of the complete complement of proteins found in an organism Degrees of Freedom for Protein Variability Covalent Modifications
Learning Peptide-Spectrum Alignment Models for Tandem Mass Spectrometry
 Learning Peptide-Spectrum Alignment Models for Tandem Mass Spectrometry John T. Halloran Dept. of Electrical Engineering University of Washington Seattle, WA 99, USA Jeff A. Bilmes Dept. of Electrical
Learning Peptide-Spectrum Alignment Models for Tandem Mass Spectrometry John T. Halloran Dept. of Electrical Engineering University of Washington Seattle, WA 99, USA Jeff A. Bilmes Dept. of Electrical
Last updated: Copyright
 Last updated: 2012-08-20 Copyright 2004-2012 plabel (v2.4) User s Manual by Bioinformatics Group, Institute of Computing Technology, Chinese Academy of Sciences Tel: 86-10-62601016 Email: zhangkun01@ict.ac.cn,
Last updated: 2012-08-20 Copyright 2004-2012 plabel (v2.4) User s Manual by Bioinformatics Group, Institute of Computing Technology, Chinese Academy of Sciences Tel: 86-10-62601016 Email: zhangkun01@ict.ac.cn,
Computational Tasks and Models
 1 Computational Tasks and Models Overview: We assume that the reader is familiar with computing devices but may associate the notion of computation with specific incarnations of it. Our first goal is to
1 Computational Tasks and Models Overview: We assume that the reader is familiar with computing devices but may associate the notion of computation with specific incarnations of it. Our first goal is to
MixGF: Spectral Probabilities for Mixture Spectra from more than One Peptide* S
 MixGF: Spectral Probabilities for Mixture Spectra from more than One Peptide* S Jian Wang, Philip E. Bourne, and Nuno Bandeira ** Technological Innovation and Resources 2014 by The American Society for
MixGF: Spectral Probabilities for Mixture Spectra from more than One Peptide* S Jian Wang, Philip E. Bourne, and Nuno Bandeira ** Technological Innovation and Resources 2014 by The American Society for
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment
 Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Bioinformatics (GLOBEX, Summer 2015) Pairwise sequence alignment Substitution score matrices, PAM, BLOSUM Needleman-Wunsch algorithm (Global) Smith-Waterman algorithm (Local) BLAST (local, heuristic) E-value
Protein Identification Using Tandem Mass Spectrometry. Nathan Edwards Informatics Research Applied Biosystems
 Protein Identification Using Tandem Mass Spectrometry Nathan Edwards Informatics Research Applied Biosystems Outline Proteomics context Tandem mass spectrometry Peptide fragmentation Peptide identification
Protein Identification Using Tandem Mass Spectrometry Nathan Edwards Informatics Research Applied Biosystems Outline Proteomics context Tandem mass spectrometry Peptide fragmentation Peptide identification
Supplementary text for the section Interactions conserved across species: can one select the conserved interactions?
 1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
1 Supporting Information: What Evidence is There for the Homology of Protein-Protein Interactions? Anna C. F. Lewis, Nick S. Jones, Mason A. Porter, Charlotte M. Deane Supplementary text for the section
Proteomics: the first decade and beyond. (2003) Patterson and Aebersold Nat Genet 33 Suppl: from
 Advances in mass spectrometry and the generation of large quantities of nucleotide sequence information, combined with computational algorithms that could correlate the two, led to the emergence of proteomics
Advances in mass spectrometry and the generation of large quantities of nucleotide sequence information, combined with computational algorithms that could correlate the two, led to the emergence of proteomics
On Optimizing the Non-metric Similarity Search in Tandem Mass Spectra by Clustering
 On Optimizing the Non-metric Similarity Search in Tandem Mass Spectra by Clustering Jiří Novák, David Hoksza, Jakub Lokoč, and Tomáš Skopal Siret Research Group, Faculty of Mathematics and Physics, Charles
On Optimizing the Non-metric Similarity Search in Tandem Mass Spectra by Clustering Jiří Novák, David Hoksza, Jakub Lokoč, and Tomáš Skopal Siret Research Group, Faculty of Mathematics and Physics, Charles
BLAST. Varieties of BLAST
 BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
BLAST Basic Local Alignment Search Tool (1990) Altschul, Gish, Miller, Myers, & Lipman Uses short-cuts or heuristics to improve search speed Like speed-reading, does not examine every nucleotide of database
Protein identification problem from a Bayesian pointofview
 Statistics and Its Interface Volume 5 (2012 21 37 Protein identification problem from a Bayesian pointofview YongFugaLi,RandyJ.Arnold,PredragRadivojac and Haixu Tang We present a generic Bayesian framework
Statistics and Its Interface Volume 5 (2012 21 37 Protein identification problem from a Bayesian pointofview YongFugaLi,RandyJ.Arnold,PredragRadivojac and Haixu Tang We present a generic Bayesian framework
profileanalysis Innovation with Integrity Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research
 profileanalysis Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research Innovation with Integrity Omics Research Biomarker Discovery Made Easy by ProfileAnalysis
profileanalysis Quickly pinpointing and identifying potential biomarkers in Proteomics and Metabolomics research Innovation with Integrity Omics Research Biomarker Discovery Made Easy by ProfileAnalysis
Algorithms in Bioinformatics FOUR Pairwise Sequence Alignment. Pairwise Sequence Alignment. Convention: DNA Sequences 5. Sequence Alignment
 Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Algorithms in Bioinformatics FOUR Sami Khuri Department of Computer Science San José State University Pairwise Sequence Alignment Homology Similarity Global string alignment Local string alignment Dot
Fast and Accurate Causal Inference from Time Series Data
 Fast and Accurate Causal Inference from Time Series Data Yuxiao Huang and Samantha Kleinberg Stevens Institute of Technology Hoboken, NJ {yuxiao.huang, samantha.kleinberg}@stevens.edu Abstract Causal inference
Fast and Accurate Causal Inference from Time Series Data Yuxiao Huang and Samantha Kleinberg Stevens Institute of Technology Hoboken, NJ {yuxiao.huang, samantha.kleinberg}@stevens.edu Abstract Causal inference
Proteomics. November 13, 2007
 Proteomics November 13, 2007 Acknowledgement Slides presented here have been borrowed from presentations by : Dr. Mark A. Knepper (LKEM, NHLBI, NIH) Dr. Nathan Edwards (Center for Bioinformatics and Computational
Proteomics November 13, 2007 Acknowledgement Slides presented here have been borrowed from presentations by : Dr. Mark A. Knepper (LKEM, NHLBI, NIH) Dr. Nathan Edwards (Center for Bioinformatics and Computational
BLAST Database Searching. BME 110: CompBio Tools Todd Lowe April 8, 2010
 BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
BLAST Database Searching BME 110: CompBio Tools Todd Lowe April 8, 2010 Admin Reading: Read chapter 7, and the NCBI Blast Guide and tutorial http://www.ncbi.nlm.nih.gov/blast/why.shtml Read Chapter 8 for
SQID: An Intensity-Incorporated Protein Identification Algorithm for Tandem Mass Spectrometry
 pubs.acs.org/jpr SQID: An Intensity-Incorporated Protein Identification Algorithm for Tandem Mass Spectrometry Wenzhou Li, Li Ji, Jonathan Goya, Guanhong Tan, and Vicki H. Wysocki* Department of Chemistry
pubs.acs.org/jpr SQID: An Intensity-Incorporated Protein Identification Algorithm for Tandem Mass Spectrometry Wenzhou Li, Li Ji, Jonathan Goya, Guanhong Tan, and Vicki H. Wysocki* Department of Chemistry
Empirical Statistical Model To Estimate the Accuracy of Peptide Identifications Made by MS/MS and Database Search
 Anal. Chem. 2002, 74, 5383-5392 Empirical Statistical Model To Estimate the Accuracy of Peptide Identifications Made by MS/MS and Database Search Andrew Keller,*, Alexey I. Nesvizhskii,*, Eugene Kolker,
Anal. Chem. 2002, 74, 5383-5392 Empirical Statistical Model To Estimate the Accuracy of Peptide Identifications Made by MS/MS and Database Search Andrew Keller,*, Alexey I. Nesvizhskii,*, Eugene Kolker,
Compound Identification Using Penalized Linear Regression on Metabolomics
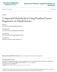 Journal of Modern Applied Statistical Methods Volume 15 Issue 1 Article 20 5-2016 Compound Identification Using Penalized Linear Regression on Metabolomics Ruiqi Liu University of Louisville, r0liu009@louisville.edu
Journal of Modern Applied Statistical Methods Volume 15 Issue 1 Article 20 5-2016 Compound Identification Using Penalized Linear Regression on Metabolomics Ruiqi Liu University of Louisville, r0liu009@louisville.edu
Faster SEQUEST Searching for Peptide Identification from Tandem Mass Spectra
 pubs.acs.org/jpr Faster SEQUEST Searching for Peptide Identification from Tandem Mass Spectra Benjamin J. Diament Department of Computer Science and Engineering, University of Washington, Seattle, Washington,
pubs.acs.org/jpr Faster SEQUEST Searching for Peptide Identification from Tandem Mass Spectra Benjamin J. Diament Department of Computer Science and Engineering, University of Washington, Seattle, Washington,
Bioinformatics and BLAST
 Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Bioinformatics and BLAST Overview Recap of last time Similarity discussion Algorithms: Needleman-Wunsch Smith-Waterman BLAST Implementation issues and current research Recap from Last Time Genome consists
Supplementary materials Quantitative assessment of ribosome drop-off in E. coli
 Supplementary materials Quantitative assessment of ribosome drop-off in E. coli Celine Sin, Davide Chiarugi, Angelo Valleriani 1 Downstream Analysis Supplementary Figure 1: Illustration of the core steps
Supplementary materials Quantitative assessment of ribosome drop-off in E. coli Celine Sin, Davide Chiarugi, Angelo Valleriani 1 Downstream Analysis Supplementary Figure 1: Illustration of the core steps
via Tandem Mass Spectrometry and Propositional Satisfiability De Novo Peptide Sequencing Renato Bruni University of Perugia
 De Novo Peptide Sequencing via Tandem Mass Spectrometry and Propositional Satisfiability Renato Bruni bruni@diei.unipg.it or bruni@dis.uniroma1.it University of Perugia I FIMA International Conference
De Novo Peptide Sequencing via Tandem Mass Spectrometry and Propositional Satisfiability Renato Bruni bruni@diei.unipg.it or bruni@dis.uniroma1.it University of Perugia I FIMA International Conference
Database search of tandem mass spectra is a central component
 B American Society for Mass Spectrometry, 2015 J. Am. Soc. Mass Spectrom. (2015) 26:1077Y1084 DOI: 10.1007/s13361-015-1120-3 RESEARCH ARTICLE Crescendo: A Protein Sequence Database Search Engine for Tandem
B American Society for Mass Spectrometry, 2015 J. Am. Soc. Mass Spectrom. (2015) 26:1077Y1084 DOI: 10.1007/s13361-015-1120-3 RESEARCH ARTICLE Crescendo: A Protein Sequence Database Search Engine for Tandem
Supplementary Materials for mplr-loc Web-server
 Supplementary Materials for mplr-loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to mplr-loc Server Contents 1 Introduction to mplr-loc
Supplementary Materials for mplr-loc Web-server Shibiao Wan and Man-Wai Mak email: shibiao.wan@connect.polyu.hk, enmwmak@polyu.edu.hk June 2014 Back to mplr-loc Server Contents 1 Introduction to mplr-loc
Discovery, Identification and Localization of Post-Translational Modifications. Nuno Bandeira, Ph.D.
 Discovery, Identification and Localization of Post-Translational Modifications Nuno Bandeira, Ph.D. Assistant Professor Dept. Computer Science and Engineering Skaggs School of Pharmacy and Pharmaceutical
Discovery, Identification and Localization of Post-Translational Modifications Nuno Bandeira, Ph.D. Assistant Professor Dept. Computer Science and Engineering Skaggs School of Pharmacy and Pharmaceutical
Electrospray ionization mass spectrometry (ESI-
 Automated Charge State Determination of Complex Isotope-Resolved Mass Spectra by Peak-Target Fourier Transform Li Chen a and Yee Leng Yap b a Bioinformatics Institute, 30 Biopolis Street, Singapore b Davos
Automated Charge State Determination of Complex Isotope-Resolved Mass Spectra by Peak-Target Fourier Transform Li Chen a and Yee Leng Yap b a Bioinformatics Institute, 30 Biopolis Street, Singapore b Davos
On the Monotonicity of the String Correction Factor for Words with Mismatches
 On the Monotonicity of the String Correction Factor for Words with Mismatches (extended abstract) Alberto Apostolico Georgia Tech & Univ. of Padova Cinzia Pizzi Univ. of Padova & Univ. of Helsinki Abstract.
On the Monotonicity of the String Correction Factor for Words with Mismatches (extended abstract) Alberto Apostolico Georgia Tech & Univ. of Padova Cinzia Pizzi Univ. of Padova & Univ. of Helsinki Abstract.
Proteomics. Yeast two hybrid. Proteomics - PAGE techniques. Data obtained. What is it?
 Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
Proteomics What is it? Reveal protein interactions Protein profiling in a sample Yeast two hybrid screening High throughput 2D PAGE Automatic analysis of 2D Page Yeast two hybrid Use two mating strains
hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference
 CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
CS 229 Project Report (TR# MSB2010) Submitted 12/10/2010 hsnim: Hyper Scalable Network Inference Machine for Scale-Free Protein-Protein Interaction Networks Inference Muhammad Shoaib Sehgal Computer Science
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of omputer Science San José State University San José, alifornia, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Pairwise Sequence Alignment Homology
Algorithms in Bioinformatics Sami Khuri Department of omputer Science San José State University San José, alifornia, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Pairwise Sequence Alignment Homology
