Conversion of the LIN-1 ETS protein of C. elegans from a sumoylated transcriptional repressor to a phosphorylated transcriptional activator.
|
|
|
- Fay Smith
- 5 years ago
- Views:
Transcription
1 Genetics: Early Online, published on January 7, 2015 as /genetics Conversion of the LIN-1 ETS protein of C. elegans from a sumoylated transcriptional repressor to a phosphorylated transcriptional activator. Elizabeth R. Leight *, John T. Murphy *, Douglas A. Fantz *,1, Danielle Pepin *, Daniel L. Schneider *, Thomas M. Ratliff &, Duaa H. Mohammad &, Michael A. Herman &, and Kerry Kornfeld * * Department of Developmental Biology, Washington University School of Medicine, St. Louis, MO & Division of Biology, Kansas State University, Manhattan, KS Current address: Department of Chemistry, Agnes Scott College, Decatur, GA Copyright 2015.
2 Running Title: ERK changes LIN-1 repressor to activator Key words: LIN-1, SUMO, ETS, chromatin, vulval development, transcription Corresponding Author: Kerry Kornfeld Campus Box 8103 Washington University School of Medicine 660 S. Euclid Ave. St. Louis, MO Phone: FAX:
3 ARTICLE SUMMARY The LIN-1 ETS transcription factor plays a pivotal role in controlling cell fate decisions during development of the Caenorhabditis elegans vulva. Prior to activation of the RTK/Ras/ERK signaling pathway, LIN-1 functions as a SUMOylated transcriptional repressor that inhibits the primary vulval cell fate. Here we demonstrate that LIN-1 binds two proteins predicted to repress transcription: RAD-26 binds the SUMOylated amino-terminus and EGL-27 binds the unsumoylated carboxy terminus. We propose a model in which LIN-1 recruits multiple transcriptional repressors to inhibit the primary vulval cell fate, and phosphorylation by ERK converts LIN-1 to a transcriptional activator that promotes the primary vulval cell fate. 3
4 ABSTRACT The LIN-1 ETS transcription factor plays a pivotal role in controlling cell fate decisions during development of the Caenorhabditis elegans vulva. Prior to activation of the RTK/Ras/ERK signaling pathway, LIN-1 functions as a SUMOylated transcriptional repressor that inhibits vulval cell fates. Here we demonstrate using the yeast two-hybrid system that SUMOylation of LIN-1 mediates interactions with a protein predicted to be involved in transcriptional repression: the RAD-26 Mi-2β/CHD4 component of the nucleosome remodeling and histone deacetylation (NuRD) transcriptional repression complex. Genetic studies indicated that rad-26 functions to inhibit vulval cell fates in worms. Using the yeast two-hybrid system, we showed that the EGL-27/MTA1 component of the NuRD complex binds the carboxyterminus of LIN-1 independent of LIN-1 SUMOylation. EGL-27 also binds UBC-9, an enzyme involved in SUMOylation, and MEP-1, a zinc finger protein previously shown to bind LIN-1. Genetic studies indicated that egl-27 inhibits vulval cell fates in worms. These results suggest that LIN-1 recruits multiple proteins that repress transcription via both the SUMOylated aminoterminus and the unsumoylated carboxy-terminus. Assays in cultured cells showed that the carboxy-terminus of LIN-1 was converted to a potent transcriptional activator in response to active ERK. We propose a model in which LIN-1 recruits multiple transcriptional repressors to inhibit the 1 vulval cell fate, and phosphorylation by ERK converts LIN-1 to a transcriptional activator that promotes the 1 vulval cell fate. 4
5 INTRODUCTION Cell fates are established during development by spatially controlled activation of signal transduction pathways that regulate transcription factors. The Elk subfamily of ETS transcription factors are critical targets of the receptor tyrosine kinase (RTK)/Ras/extracellularregulated kinase (ERK) signaling pathway (Treisman, 1994). This subfamily includes extensively characterized proteins from three divergent animals: vertebrate Elk-1, Drosophila Aop/Yan and C. elegans LIN-1. The mechanisms that enable ETS proteins to switch cell fates in response to signaling pathway activation are not fully defined, and to address this important issue it is critical to elucidate the function of these proteins both prior to ERK activation and following phosphorylation by ERK. These issues have significant implications for human health, since ETS proteins contribute to human diseases including cancer (Dittmer, 2003). lin-1 has been extensively characterized during development of the hermaphrodite vulva, an epidermal structure used for egg laying and sperm entry (reviewed by Horvitz and Sternberg, 1991; Kornfeld, 1997; Greenwald, 1997; Sternberg, 2005; Sundaram, 2013). In third larval stage hermaphrodites, six ventral epidermal blast cells called P3.p - P8.p (Pn.p cells) lie along the anterior-posterior axis. These Pn.p cells are an equivalence group, since each cell can adopt the 1 vulval cell fate (eight descendents), the 2 vulval cell fate (seven descendents), or the nonvulval 3 cell fate (two descendents). During vulval induction, the anchor cell of the somatic gonad secretes LIN-3 ligand, an epidermal growth factor (EGF) homolog, thereby activating the LET-23/RTK on the adjacent P6.p cell (Hill and Sternberg, 1992; Aroian et al., 1990). Activated LET-23 recruits the SEM-5 adaptor (GRB2) and the LET-341/SOS-1 guanine nucleotide exchange factor (SOS), followed by sequential activation of the GTPase LET-60 (RAS) and the protein kinases LIN-45 (RAF), MEK-2 (MAPK kinase, or MEK), and MPK-1 5
6 (ERK) (Clark et al., 1992; Chang et al., 2000; Han and Sternberg, 1990; Beitel et al., 1990; Han et al., 1993; Hsu et al., 2002; Kornfeld et al., 1995; Wu et al., 1995; Lackner et al., 1994; Wu and Han, 1994). Activated MPK-1 phosphorylates substrates including the LIN-1 ETS transcription factor, thereby promoting the 1 fate in P6.p. P6.p signals P5.p and P7.p to adopt 2 fates by activating a lateral signal involving LIN-12/Notch. The 22 descendents of P5.p, P6.p and P7.p invaginate and differentiate to form the vulval structure. P3.p, P4.p and P8.p receive neither signal and therefore adopt 3 fates. Mutations that reduce activation of RTK/Ras/MAPK signaling cause a vulvaless (Vul) phenotype, whereas mutations that constitutively activate this pathway cause more than three Pn.p cells to adopt the 1 or 2 vulval cell fate, resulting in a multivulval (Muv) phenotype characterized by ectopic patches of vulval tissue. LIN-1 is a transcription factor that plays a critical role in establishing vulval cell fates. Lossof-function mutations that abrogate sequence-specific DNA binding activity cause a strong Muv phenotype, demonstrating that DNA binding is necessary for lin-1 function and lin-1 function is necessary to inhibit the 1º fate (Sulston and Horvitz, 1981; Beitel et al., 1995; Miley et al., 2004). Genetic epistasis studies established that lin-1 functions downstream of mpk-1 ERK (Ferguson et al., 1987; Lackner et al., 1994; Wu and Han, 1994). LIN-1 contains two docking sites for ERK, the D-domain and FQFP motif, and 17 S/TP motifs that are potential ERK phosphorylation sites (Tan et al., 1998; Jacobs et al., 1998; Jacobs et al., 1999). Mutations of LIN-1 that decrease phosphorylation by ERK cause a gain-of-function Vul phenotype, indicating that phosphorylation is necessary to stop LIN-1 from inhibiting the 1º fate (Jacobs et al., 1998). These studies are consistent with two possible models. Phosphorylation by ERK may abrogate LIN-1 activity in P6.p. Alternatively, phosphorylation may convert LIN-1 from an inhibitor to an activator of the 1 cell fate. Here we address these models by analyzing how phosphorylation 6
7 by ERK affects the transcriptional activity of LIN-1. Our results indicate that the carboxyterminus of LIN-1 is converted from a transcriptional repression domain to a potent transcriptional activation domain by ERK phosphorylation. LIN-1 contains two consensus SUMOylation motifs. The E2 small ubiquitin-related modifier (SUMO) conjugating enzyme, UBC-9, binds these motifs and mediates the covalent attachment of SUMO to LIN-1 (Leight et al., 2005). C. elegans smo-1, the gene encoding SUMO, and ubc-9 inhibit vulval cell fates (Poulin et al., 2005) and genetically interact with lin-1 (Leight et al., 2005), indicating that SUMOylation of LIN-1 mediates inhibition of vulval cell fates. SUMOylation of LIN-1 promotes transcriptional repression (Leight et al., 2005), but the mechanism has not been well defined. We previously showed that SUMOylated LIN-1 interacts with MEP-1, a component of the NuRD transcriptional repression complex, suggesting this interaction is one mechanism of LIN-1 transcriptional repression (Leight et al., 2005). The HOX gene lin-39 is a positive regulator of vulval cell fates and a direct transcriptional target of LIN-1 (Wagmaister et al., 2006). Guerry et al., (2007) showed that LIN-1 interacts with the Mi-2 nucleosome remodeling protein LET-418 on the lin-39 promoter. Here we identify proteins that bind LIN-1 and may contribute to transcriptional repression by LIN-1. SUMOylation of LIN-1 mediated recruitment of two chromatin regulatory enzymes in yeast cells: SET-6 and RAD-26. SET-6 is homologous to SUV39H1, a histone methyltransferase that mediates transcriptional repression (Lachner et al., 2001; Bannister et al., 2001; Nielsen et al., 2001). RAD-26 shares homology with the Mi-2β/CHD4 component of the NuRD transcriptional repression complex. In addition, the EGL-27/MTA1 component of the NuRD complex binds the carboxy-terminus of LIN-1 in yeast cells independent of SUMOylation. These findings suggest that LIN-1 recruits cofactors via both its SUMOylated 7
8 amino-terminus and unsumoylated carboxy-terminus to inhibit vulval cell fates. rad-26 and egl-27 function to inhibit vulval cell fates in worms. These data support a model in which SUMOylated LIN-1 recruits multiple chromatin regulatory proteins to repress transcription of vulval cell fate genes, and phosphorylation by ERK MAP kinase in the P6.p cell converts LIN-1 to a transcriptional activator that promotes vulval cell fates. MATERIALS AND METHODS Yeast two-hybrid screens and reporter gene assays: The YEL3 strain encoding a LexADBD:LIN-1(1-252) fusion protein was described by Leight et al. (2005). The YEL2 strain was generated by transforming the L40 strain (Vojtek et al., 1993) with a bait plasmid encoding LIN-1( ) cloned into pbtm116 (Bartel et al., 1993). A random-primed cdna library from mixed-stage hermaphrodites (kindly provided by R. Barstead), containing cdnas fused to the GAL4AD, was transformed into YEL2 and YEL3 (Schiestl and Gietz, 1989). Prey plasmids were isolated from positive colonies. cdnas that failed to PCR amplify with MEP-1 or MPK-1 gene-specific primers were sequenced using standard techniques. Bait plasmids encoding EGL-27(1-1129), which corresponds to the largest predicted egl- 27 transcript, EGL-27(1-637) and EGL-27( ) (Herman et al., 1999; Solari et al., 1999) were cloned into pas2-1 (CLONTECH, Palo Alto, CA) and transfected into S. cerevisiae strain Hf7c (Feilotter et al., 1994). Positive clones growing on His dropout media were assayed for β- galactosidase activity. To monitor activation of the LexA-dependent lacz reporter, we prepared lysates from at least six independent yeast transformants of equivalent size and measured β-galactosidase activity using the Galacto-Light Plus System (Applied Biosystems). For Figure 2, yeast 8
9 transformants were grown in selective media at 30 C to an O.D. of approximately 1.0 before analysis. Analysis of C. elegans strains: C. elegans strains were cultured at 20 C as described by Brenner (1974) unless otherwise noted. The following mutations were analyzed: let-60(n1046) (Beitel et al., 1990), lin-1(e1275 R175Opal) (Beitel et al., 1995), lin-1(n1790 R352Opal) (Jacobs et al., 1998), egl-27(n170) (Herman et al., 1999), egl-27(we3) (Solari et al., 1999), lin-15a(n767) and lin-15b(n744) (Ferguson and Horvitz, 1989), mpk-1(oz140) (Lee et al., 2007), and set- 6(tm1611) (Andersen and Horvitz 2007). rad-26(tm1991) was generated by the National Bioresource Project by screening animals mutagenized by TMP/UV for a deletion of the C27B7.4 gene. We outcrossed tm1991 to the N2 wild-type strain twice and constructed double mutants using standard techniques. set-6(ok2195) was obtained from the CGC and outcrossed multiple times to the N2 wild-type strain. cdnas encoding RAD-26(17-655), SET-6(1-708) and EGL-27( ) were cloned into the L4440 plasmid (Timmons and Fire, 1998) for expression of double-stranded RNA (dsrna). Plasmids were transformed into HT115(DE3) E. coli. L4 hermaphrodites were placed on Petri dishes containing bacteria that expressed dsrna or control bacteria with the L4440 plasmid and transferred to new Petri dishes daily; progeny were scored for the Muv phenotype. Protein purification and GST pull-down assays: 6xHis- and FLAG-tagged RAD- 26(17-655) (HF:RAD-26) and SET-6(22-538) (HF:SET-6) were expressed in Sf9 cells using a baculovirus expression system (Invitrogen, Bac-to-Bac Baculovirus Expression Systems manual). DNA encoding EGL-27( ) was cloned in pet-28a(+) plasmid (EMD 9
10 Biosciences) with the 6xHis- and T7 tag, and the protein was expressed in E. coli. HF:RAD-26, HF:SET-6 and T7:EGL-27 were purified using native conditions as described in Protocol 16 of the QIAexpressionist handbook (Qiagen, August 2002), with minor modifications. DNAs encoding LIN-1(1-64) and the SMO-1(1-88) were cloned in-frame with GST into the plasmid pgex-4t-1 (Amersham Pharmacia Biotech). DNA encoding LIN-1( ) was cloned inframe with GST in the plasmid pgex-2t (Amersham Pharmacia Biotech). GST and GST fusion proteins were expressed in E. coli and partially purified under native conditions using glutathione sepharose as described (Jacobs et al., 1998). For pull-down assays in Figure 3, equimolar amounts of HF:RAD-26 and HF:SET-6 were incubated with approximately 5 µg of glutathione sepharose-bound GST, GST:LIN-1(1-64), or GST:SMO-1 for 1 hour at 4 C in binding buffer (PBS + 0.1% NP-40). The glutathione sepharose was washed twice; bound proteins were eluted by boiling in sample buffer and separated by SDS-PAGE (Sambrook et al., 1989). HF:RAD-26 and HF:SET-6 were detected by immunoblotting using the α-flag M2 antibody (Sigma). Pull-down assays in Figure 5C were performed similarly, except T7:EGL-27 was incubated with approximately 1 µg of glutathione sepharose-bound GST, GST:LIN-1(1-64) or GST:LIN-1( ). Bound T7:EGL-27 was detected by immunoblotting using the α-t7 Tag monoclonal antibody (EMD Biosciences). For pull-down assays in Figure 5D, the egl-27 cdna was cloned into pgex-4t-1 to encode GST:EGL-27(1-1129), and a full-length mep-1 cdna was cloned into ptrchis (Invitrogen) to encode 6xHis:MEP-1. Fusion proteins were expressed in E. coli strain BL21(DE3) and purified by the method of Frangioni and Neel (1993) with minor modifications. An equimolar amount of 6xHis:MEP-1 was added to GST and GST:EGL-27(1-1129) bound to GSH beads. 6xHis:MEP-1 10
11 was detected with HRP conjugated anti-his antibody (Santa Cruz) using West Pico Super Signal Chemiluminescent detection system (Pierce). Reporter gene assays in mammalian cultured cells: The MC3T3 mouse fibroblast cell line was grown in α-mem with 10% fetal calf serum and transfected as described by Fantz et al. (2001) with (1) the pfr-luc reporter plasmid (Stratagene), (2) an expression plasmid that encodes GAL4 DNA binding domain (G4) alone or fused to fragments of LIN-1 or Elk-1 (pfa- Elk, Stratagene) and (3) the CMV-β-galactosidase control plasmid. Cells were treated with 5% glycerol one day after transfection, incubated an additional two days, and harvested for cell lysates; luciferase and β-galactosidase activity were quantified. Cells were exposed to 50 µg/ml recombinant basic fibroblast growth factor (bfgf) (Upstate Biotechnology) for 20 hours beginning two days post-transfection. Site-directed mutagenesis was used to generate mut. D (K280A, K283A, L287A, L289A), mut. F (F381A, Q382A, F383A) and the double mutant (mut. D/F). RESULTS LIN-1 interacted with RAD-26, SET-6 and EGL-27 in yeast cells: To identify proteins that associate with LIN-1 (Figure 1A), we conducted yeast two-hybrid screens. In the first screen, the bait was the LexA DNA binding domain (LA) fused to the amino-terminal 252 residues of LIN-1, which includes the ETS DNA binding domain and two consensus SUMOylation motifs. 233 cdnas that encode proteins that bind LIN-1(1-252) specifically were identified by screening 4x10 6 cdnas from a mixed-stage C. elegans cdna library. 123 cdnas encoded the zinc finger protein MEP-1 that we described previously (Leight et al., 2005). Four 11
12 cdnas derived from the C27B7.4/rad-26 gene. The predicted RAD-26 protein contains a DNAdependent ATPase domain of the SNF2 family N-terminal class and a helicase C-terminal domain (Figure 1B). RAD-26 shares 23% identity over 883 amino acids with LET-418/CHD-4, a component of the NuRD transcriptional repression complex. Interestingly LET-418/CHD-4 binds MEP-1 (Unhavaithaya et al., 2002). Two cdnas derived from the C49F5.2/set-6 gene. The predicted SET-6 protein contains a SET lysine methyltransferase domain and is predicted to function as a histone H3 lysine 9 methyltransferase based upon its homology to the transcriptional repressor SUV39H1 (Lachner et al., 2001; Bannister et al., 2001; Nielsen et al., 2001) (Figure 1B). In the second screen, the bait was the LexA DNA binding domain fused to the LIN-1 carboxy-terminus (residues ), which includes the D domain and FQFP docking sites for ERK MAP kinase. 143 cdnas that encode proteins that bind LIN-1( ) specifically were identified by screening 1.3x10 6 cdnas. 37 cdnas encoded MPK-1 ERK MAP kinase. Seven cdnas encoded EGL-27. EGL-27 shares 24% identity and 41% similarity over 467 amino acids with the MTA-1 component of the NuRD transcriptional repression complex (Herman et al., 1999). EGL-27 contains an ELM1 domain that is predicted to mediate protein-protein interactions, an ELM2 domain that is predicted to mediate DNA binding or protein-protein interactions, a SANT DNA binding domain and a GATA zinc finger domain. Two SUMOylation motifs of LIN-1 are necessary to promote binding of RAD-26 and SET-6 in yeast cells: To define regions of LIN-1 that are sufficient to bind SET-6 and RAD-26, we analyzed fragments of LIN-1. LA:LIN-1(1-64) and LA:LIN-1( ) mediated robust binding of SET-6 and RAD-26, indicating that LIN-1 contains two separable binding sites 12
13 for SET-6 and RAD-26 (Figure 1C). To identify amino acids necessary for binding, we analyzed mutants of LIN-1(1-64) with eight consecutive amino acids changed to alanine. Substitution of LIN-1 residues 9-16 dramatically reduced binding of SET-6 and RAD-26 by 106-fold and 44- fold, respectively (Figure 1D, line 3, and 1F). LIN-1 contains a consensus SUMOylation motif in this region, VK 10 KE (Sampson et al., 2001). Substitution of the SUMO acceptor lysine (K10A) dramatically reduced binding of SET-6 and RAD-26 by 62-fold and 37-fold, respectively (Figure 1D, line 11). Substitution of the highly conserved glutamic acid (E12A) reduced binding of SET-6 and RAD-26 by 40-fold and 30-fold, respectively (Figure 1D, line 13). Substitution of the moderately conserved valine (V9A) decreased binding of SET-6 and RAD-26 by 6-fold and 5-fold respectively, whereas mutation of the non-conserved lysine (K11A) had minimal effects (Figure 1D, lines 10 and 12). These results demonstrate a correlation between the function of each residue in the ψkxe motif in promoting SUMOylation and in promoting binding of SET-6 and RAD-26. To investigate SET-6 and RAD-26 binding to LIN-1 residues , we analyzed the LIN-1( ) fragment that contains the VK 169 DE SUMOylation motif. SET-6 and RAD-26 robustly bound LIN-1( ) (Figure 1E). Mutations of the entire motif ( A) or the predicted SUMO acceptor lysine (K169A) markedly reduced binding of LIN-1 to SET-6 and RAD-26 (Figure 1E). Thus, 64 residues of LIN-1 containing the SUMOylation motif VK 10 KE and 25 residues of LIN-1 containing the SUMOylation motif VK 169 DE were sufficient to bind SET-6 and RAD-26, and for both LIN-1 fragments the SUMO acceptor lysine was necessary for binding. 13
14 SUMOylation of LIN-1 promotes binding of RAD-26 and SET-6 in yeast cells: The ψkxe consensus SUMOylation motifs of LIN-1 are post-translationally modified by SUMO-1 in yeast cells (Leight et al., 2005). If SUMOylation of the ψkxe motifs promotes binding of SET- 6 and RAD-26, then the addition of SUMO-1 to a SUMOylation-defective LIN-1 mutant might restore binding. To mimic the SUMOylated isoform of LIN-1, we generated a translational fusion of C. elegans SMO-1 and the LIN-1(1-64; 9-16A) SUMOylation-defective mutant. This fusion protein functioned as efficiently as SUMOylated LIN-1(1-64) to repress transcription in cultured cells, indicating that this translational fusion of SMO-1 can be functional (Leight et al., 2005). The interaction of SET-6 and RAD-26 with LA:SMO-1:LIN-1(1-64; 9-16A) was increased by 24-fold and 3-fold, respectively (Figure 2, lines 2 and 3; data not shown). These results indicate that SET-6 and RAD-26 bind SUMOylated LIN-1. RAD-26 directly binds SUMO and LIN-1: To determine if LIN-1 directly binds SET-6 and RAD-26, we expressed histidine- and FLAG-tagged SET-6(22-538) (HF:SET-6) and RAD- 26(17-655) (HF:RAD-26) proteins in insect cells and purified these proteins by metal affinity chromatography. Glutathione-S-transferase (GST)-tagged C. elegans SMO-1 and LIN-1(1-64) were expressed in E. coli and purified by glutathione affinity chromatography. Binding was monitored by the GST pull-down method. HF:SET-6 did not bind GST:SMO-1 nor GST:LIN- 1(1-64), indicating that additional proteins may facilitate the interaction between SET-6 and LIN-1 in yeast cells or the appropriate pull-down conditions were not identified. HF:RAD-26 did bind GST:SMO-1 and GST:LIN-1(1-64) specifically (Figure 3). These findings demonstrate that RAD-26 can directly bind both SMO-1 and LIN-1. 14
15 rad-26 functions in worms to inhibit vulval cell fates: To investigate the possibility that these protein interactions are functionally significant in worms, we reduced the activity of rad-26 and set-6 and monitored vulval development in wild-type and sensitized genetic backgrounds. rad-26 and set-6 have been analyzed in genome-wide screens using RNAi, but phenotypes have not been reported (Piano et al., 2002; Kamath et al., 2003; Sonnichsen et al., 2005). Wild-type hermaphrodites exposed to rad-26 dsrna did not display a Muv phenotype (Table 1, line 15). The rad-26(tm1991) allele is deleted for exon 6 and a portion of exon 7 and is predicted to encode a truncated protein of 412 amino acids that lacks half of the ATPase domain and the entire helicase domain, suggesting the mutant protein has reduced activity. rad- 26(tm1991) mutants grown at 15ºC and 20ºC did not display a Muv phenotype (Table 1, line 2 and 6). To analyze interactions of rad-26 with the EGFR/Ras/MAPK pathway, we utilized sensitized genetic backgrounds. let-60(n1046gf) is a gain-of-function mutation that causes a temperature-sensitive Muv phenotype (Beitel et al., 1990). When let-60(n1046gf) hermaphrodites were fed rad-26 dsrna, the penetrance of the Muv phenotype increased significantly from 16% to 85% (Table 1, line 17), indicating that rad-26 inhibits vulval cell fates. mpk-1(oz140) is a partial loss-of-function mutation that affects ERK MAP kinase and inhibits vulval induction: rad-26(tm1991) did not cause a Muv phenotype in this genetic background (Table 1, line 11). lin-1(e1275 R175Opal) is a partial loss-of-function mutation that causes a temperature-sensitive Muv phenotype. A double mutant with rad-26(tm1991) increased the penetrance of the lin-1(e1275) Muv phenotype significantly from 41% to 89% (Table 1, line 4). The lin-1 Muv phenotype is caused by a transformation of the cell fates of P3.p, P4.p and P8.p from nonvulval to vulval (Ferguson et al., 1987). Nomarski optics were used to confirm that the 15
16 lin-1(e1275) rad-26(tm1991) double mutant animals displayed ectopic vulval cell fates (Figure 4A,B). These results demonstrate that rad-26 functions to inhibit vulval cell fates, and are consistent with the model that rad-26 is positioned in the EGFR/Ras/MAPK pathway or a parallel signaling pathway. To investigate the position of the rad-26 gene in the Ras pathway, we analyzed a lin-1 gain-of-function mutation. lin-1(n1790gf R352Opal) causes a weak vulvaless phenotype and partially suppresses the Muv phenotype caused by activated let-60 ras; the LIN-1(1-351) protein lacks the FQFP ERK docking site and is partially resistant to negative regulation by MPK-1 ERK (Jacobs et al., 1999). The lin-1(n1790gf) allele also causes a low penetrance Muv phenotype because the lin-1 mrna contains a premature stop codon and is subject to nonsensemediated decay. If rad-26 functions upstream of lin-1 in a linear signaling pathway, then we predict that the rad-26 mutation will not significantly affect the lin-1(n1790) phenotype. By contrast, if rad-26 functions downstream or at the level of lin-1, then we predict that the rad-26 mutation will promote a Muv phenotype in the lin-1(n1790) background. Specifically, if RAD- 26 is recruited to SUMOylated LIN-1(1-351) and contributes to LIN-1 function, then removing RAD-26 protein is predicted to impair the function of LIN-1(1-351) as a constitutive inhibitor of vulval cell fates, resulting in a Muv phenotype. The lin-1(n1790gf) rad-26(tm1991) double mutant hermaphrodites displayed a Muv phenotype that was 29% penetrant, significantly greater than the Muv phenotype of n1790 and tm1991 single mutant animals (Table 1, line 8). Consistent with this observation, lin-1(n1790gf) hermaphrodites fed rad-26 dsrna displayed a weak but significant enhancement of the Muv phenotype (Table 1, line 19). These findings indicate that the LIN-1(1-351) protein may require rad-26 function to inhibit vulval cell fates efficiently. 16
17 To analyze the function of set-6, we reduced the activity of set-6 by feeding dsrna. Wild-type hermaphrodites exposed to set-6 dsrna did not display obvious vulval defects (data not shown). The set-6(tm1611) and set-6(ok2195) deletion mutations did not cause an independent Muv phenotype (data not shown). By contrast to rad-26, set-6(lf) mutations did not appear to interact with the EGFR/Ras/MAPK pathway. The Muv phenotype of lin-1(e1275); set-6(ok2195) double mutant animals was similar to single mutant lin-1(e1275) animals (data not shown). The Muv phenotype of let-60(n1046); set-6(tm1611) and let-60(n1046); set-6(ok2195) double mutant animals was similar to let-60(n1046) single mutant animals (data not shown). These finding indicate that set-6(lf) mutations do not interact strongly with mutations that affect the EGFR/Ras/MAPK pathway. set-6(ok2195) did not cause a Muv phenotype in combination with rad-26(tm1991) (Table 1, line 9) or egl-27(we3) (Table 2, line 8). These results do not provide evidence that set-6 plays a significant functional role in lin-1 mediated inhibition of vulval cell fates and raise the possibility that that the interaction between SET-6 and LIN-1 proteins may reflect a role in a different tissue The synthetic multivulva (synmuv) pathway is composed of two major groups of genes called class A and class B (Ferguson and Horvitz, 1989). Loss-of-function mutations in a single class A gene or a single class B gene do not cause an independent Muv phenotype, but a double mutant animal containing any class A mutation and any class B mutation displays a strong Muv phenotype. Thus, the class A and class B synmuv genes appear to function redundantly to inhibit vulval cell fates. The synmuv genes appear to function non cell autonmously in the hyp7 hypodermal cell to inhibit the expression and secretion of the lin-3 ligand; in class AB double mutant animals, ectopic secretion of lin-3 causes the Muv phenotype (Herman and Hedgecock 1990; Cui et al., 2006). Several genes associated with the NURD complex have been proposed to 17
18 function in the synmuv pathway, including let-418 and egr-1/lin-40 (von Zelewsky et al., 2000; Solari and Ahringer 2000). Andersen and Horvitz (2007) analyzed set-6 for a role in the synmuv pathway; the set-6(tm1611) deletion allele did not cause a Muv phenotype in combination the class A synmuv gene lin-15a or the class B synmuv gene lin-15b, indicating set-6 is not a synmuv gene. To investigate the possibility that rad-26 is a synmuv gene, we generated double mutant animals with lin-15a or lin-15b mutations. rad-26(tm1991) did not cause a Muv phenotype in combination with either class A or class B synmuv mutations, indicating rad-26 is not a synmuv gene (Table 1, lines 12-13). EGL-27 binds two domains of LIN-1 that are separable from the SUMOylation motifs and ERK docking sites: EGL-27 was identified based on an interaction with LIN- 1( ) that contains the carboxy-terminus. To characterize the interaction with EGL-27, we used the yeast two-hybrid system to define domains of LIN-1 that are necessary and sufficient to mediate binding. We mutagenized one or both of the ERK docking sites and monitored the association of these LIN-1 derivatives with EGL-27 and MPK-1. MPK-1 displayed reduced binding to LA:LIN-1( ; mut. D) and LA:LIN-1( ; mut. F), and failed to bind LA:LIN-1( ; mut. D/F) (Figure 5B, lines 2-4). By contrast, EGL-27 efficiently bound each of these LIN-1 derivatives, indicating that the ERK docking sites in LIN-1 are not necessary for EGL-27 binding. To define regions of LIN-1 that are sufficient to bind EGL-27, we monitored the association of EGL-27 to fragments of LIN-1. These studies demonstrated that EGL-27 binds two domains in LIN-1 contained in residues and residues (Figure 5B, lines 6 and 8). 18
19 To determine if EGL-27 can directly bind LIN-1, histidine- and T7-tagged EGL-27( ) was expressed in E. coli, purified using metal affinity chromatography, and monitored for its binding to purified GST:LIN-1( ) using a GST pulldown assay. EGL-27 reproducibly bound GST:LIN-1( ) more efficiently than GST:LIN-1(1-64) or GST alone (Figure 5C), demonstrating that EGL-27 can directly bind the carboxy-terminus of LIN-1. To further characterize the function of EGL-27, we used the yeast two-hybrid system to identify C. elegans proteins that interact with EGL-27. EGL-27(1-1129) was used to screen 3.4x10 6 cdnas, and eight positive cdnas were identified; three of these cdnas corresponded to MEP-1. EGL-27( ) was used to screen 6.9x10 4 cdnas, and 49 positive cdnas were identified; two of these cdnas corresponded to UBC-9. MEP-1 and UBC-9 were previously demonstrated to interact with LIN-1 and function to inhibit vulval cell fates during development (Leight et al., 2005). A fragment of MEP-1 containing residues , which includes all seven zinc-finger domains, was sufficient to interact with full-length EGL-27 in yeast. In addition, MEP-1 residues were sufficient to interact with the amino-terminal 637 residues of EGL-27 in yeast (data not shown). The interaction of MEP-1 with full-length EGL- 27 was demonstrated to be a direct interaction by a GST-pulldown assay (Figure 5D). egl-27 inhibits vulval cell fates: egl-27 loss-of-function mutations were first identified based on egg-laying defects (Trent et al., 1983). egl-27 mutations also cause defects in TL and TR cell polarity, migration of HSN cells and QL neuroblast descendants (Herman et al., 1999), and embryonic patterning (Solari et al., 1999). egl-27 has been demonstrated to inhibit vulval cell fates, since egl-27 RNAi was reported to enhance the Muv phenotype of lin-37(n758) egr- 1(RNAi) animals from 21% to 50% (Solari and Ahringer, 2000). egl-27(n170) was reported to enhance the let-60(n1046) Muv phenotype, increasing the percent of induced P3.p cells from 19
20 47% to 79%, induced P4.p cells from 46% to 74%, and induced P8.p cells from 28% to 62% (Chen and Han, 2001a). We hypothesized that LIN-1 recruits EGL-27 to repress transcription of vulval cell fate genes. To test this prediction, we reduced the function of egl-27 using loss-offunction mutations or RNAi and monitored vulval development in different genetic backgrounds. egl-27(we3) is a nonsense mutation that results in a truncated protein of 811 amino acids (Solari et al., 1999). egl-27(n170) is a deletion of at least the first eight exons of egl-27 and affects the full-length transcript, but not the smaller transcripts (Herman et al., 1999). egl-27(n170) and egl- 27(we3) hermaphrodites both displayed a very low penetrance Muv phenotype of 1% (Table 2, lines 2, 3). Likewise, 1% of wild-type hermaphrodites exposed to egl-27 RNAi displayed a Muv phenotype (Table 2, line 10). These results indicate that egl-27 functions to inhibit vulval cell fates, but the effect is difficult to measure in an otherwise wild-type background. To investigate the possibility that egl-27 functions in the synmuv pathway, we generated double mutant animals containing egl-27(we3) and a mutation of the class A synmuv gene lin- 15A or the class B synmuv gene lin-15b. These double mutant animals did not display a Muv phenotype, indicating that egl-27 is not a synmuv gene (Table 2, lines 6-7). This is similar to a previous report showing egl-27 RNAi did not cause a Muv phenotype in combination with lin- 15A and caused only a low penetrance Muv phenotype (5%) in combination with the synmuv B gene lin-37 (Solari and Ahringer, 2000) To investigate the possibility that egl-27 functions in the EGFR/Ras/MAPK pathway, we examined egl-27 function in sensitized genetic backgrounds. The penetrance of the let- 60(n1046gf) Muv phenotype was significantly increased from 27% to 72% in hermaphrodites fed egl-27 dsrna (Table 2, line 12), consistent with previous studies demonstrating that egl- 27(n170) enhanced the frequency of vulval cell fates of P3.p, P4.p and P8.p (Chen and Han, 20
21 2001a). The egl-27(we3); lin-1(n1790gf) double mutant animals displayed a Muv phenotype that was 21% penetrant, significantly greater than the Muv phenotype of n1790 and we3 single mutants (Table 2, line 5). An analysis using DIC optics confirmed that these double mutant animals displayed ectopic protrusions of vulval tissue (Figure 4C,D). Likewise, 28% of lin- 1(n1790gf) hermaphrodites fed egl-27 dsrna displayed a Muv phenotype, which was significantly higher that the 3% of lin-1(n1790gf) hermaphrodites fed control bacteria that displayed a Muv phenotype (Table 2, line 14). Thus, two independent methods of reducing egl- 27 function promoted vulval cell fates in the presence of lin-1(n1790), indicating this is a robust result although the effect is not fully penetrant. These results are consistent with the model that EGL-27 may promote the ability of gain-of-function LIN-1(1-351) protein to inhibit vulval cell fates. egl-27(we3) did not cause a Muv phenotype in combination with rad-26(tm1991) (Table 1, line 10) or set-6(ok2195) (Table 2, line 8). The carboxy-terminus of LIN-1 functions as an ERK-dependent transcriptional activation domain and an ERK-independent transcriptional repression domain: To analyze how phosphorylation by ERK regulates LIN-1, we analyzed transcriptional activity of the carboxy-terminus of LIN-1. Fragments of LIN-1 fused to the GAL4 DNA binding domain were expressed in the MC3T3 murine fibroblast cell line, and transcription of a GAL4-dependent reporter gene was monitored. To increase ERK activity, we treated cells with fibroblast growth factor (bfgf). To decrease the ability of ERK to phosphorylate LIN-1, we mutated the ERK docking sites (Jacobs et al., 1998; Jacobs et al., 1999). LIN-1( ) functioned as a potent activator of transcription, activating transcription by 81-fold relative to the G4 only control (Figure 6A, line 2). Treatment of cells with bfgf resulted in a 2-fold increase in the 21
22 transcriptional activation potential of G4:LIN-1( ), suggesting that phosphorylation of LIN-1 by ERK promotes transcriptional activation. The activity of the LIN-1 carboxy-terminus in this assay is similar to the activity of the Elk-1 carboxy-terminus (Figure 6A, line 6). One possible explanation for the relatively small effect of adding bfgf is that ERK is already highly active in the standard culture conditions. This is likely, since the cells were cultured with 10% fetal calf serum that contains a variety of growth factors that activate ERK. To address the possibility that LIN-1 is highly phosphorylated by ERK in standard culture conditions, we investigated the effect of reducing phosphorylation of LIN-1 by mutating the docking sites for ERK. LIN-1 contains two docking sites for ERK, and we mutated only the D domain, G4:LIN-1( , mut. D), only the FQFP motif, G4:LIN-1( , mut. F), and both the D domain and the FQFP motif, G4:LIN-1( , mut. D/F). Compared to wild-type LIN-1, these mutations reduced transcriptional activity by 251 fold, 75-fold and 380 fold, respectively. These results indicate that under standard culture conditions, binding of ERK to the D-domain and FQFP motif of LIN-1 and phosphorylation of LIN-1 dramatically increases the transcriptional activation potential of the carboxy-terminus of LIN-1. These experiments allowed us to measure the transcriptional activity of the LIN-1 carboxy-terminus in the absence of ERK phosphorylation. The LIN-1( ) fragment contains one consensus SUMOylation motif and two binding sites for EGL-27. In the absence of ERK docking sites, this fragment of LIN-1 repressed transcription by 4.7-fold relative to G4 alone (Figure 6B, line 2). These results are consistent with the model that SUMOylation of LIN- 1 and/or EGL-27 binding to the carboxy-terminus of LIN-1 mediate transcriptional repression, and phosphorylation by ERK converts LIN-1 to a transcriptional activator (Figure 7). 22
23 DISCUSSION SUMOylated LIN-1 recruits a nucleosome-remodeling enzyme to inhibit vulval cell fates: Genetic analyses demonstrate that lin-1 inhibits Pn.p cells from adopting vulval cell fates (Sulston and Horvitz, 1981; Beitel et al., 1995; Jacobs et al., 1998). DNA binding and SUMOylation are necessary for this LIN-1 activity, suggesting that SUMOylated LIN-1 represses transcription of vulval cell fate genes (Miley et al., 2004; Leight et al., 2005). Here we define a novel mechanism for transcriptional repression by SUMOylated LIN-1, recruitment of RAD-26. We discovered RAD-26 as a LIN-1-interacting protein in yeast cells. SUMOylation of LIN-1 was necessary and sufficient to promote binding of RAD-26 in yeast cells. Furthermore, RAD-26 directly binds both LIN-1 and SUMO in purified extracts. These observations suggest RAD-26 binds SUMOylated LIN-1 in worms; however, this biochemical interaction in worms has not been tested directly. Our genetic analyses demonstrated that rad-26 inhibits vulval cell fates. rad-26 interacted genetically with multiple genes in the Ras signaling pathway that function in P6.p, and rad-26 did not cause a Muv phenotype in combination with class A or class B synmuv genes that function in hyp7. These genetic results are consistent with the model that rad-26 functions in the EGFR/Ras/MAPK signaling pathway or that rad-26 functions in parallel to this signaling pathway, and they do not establish the cellular site of action of rad-26. Together, these biochemical and genetic results are consistent with the model that RAD-26 binding to LIN-1 promotes the ability of LIN-1 to repress primary vulval cell fates. Whereas reducing the activity of rad-26 or lin-1 causes ectopic vulval cell fates, the frequency of ectopic 23
24 vulval cell fates is higher when lin-1 activity is reduced. This observation suggests that lin-1 retains some ability to inhibit vulval cell fates in the absence of rad-26. RAD-26 is a homolog of vertebrate Mi-2β/CHD4, a core component of the NuRD transcriptional repression complex. This complex possesses ATP-dependent nucleosome remodeling activity that is dependent upon Mi-2β/CHD4 and histone deacetylase activity provided by HDAC-1 and HDAC-2; both of these activities promote transcriptional silencing (Tong et al., 1998; Wade et al., 1998; Xue et al., 1998; Zhang et al., 1998; Wang and Zhang, 2001). Multiple components of the C. elegans NuRD complex have been shown to inhibit vulval cell fates, including CHD-3 and LET-418 that are homologs of vertebrate Mi-2β/CHD4 (Solari and Ahringer, 2000; von Zelewsky et al., 2000; Chen and Han, 2001a; Chen and Han, 2001b; Lu and Horvitz, 1998). Guerry et al. (2007) showed that LET-418 interacts with LIN-1 to repress transcription of the lin-39 target gene during vulval induction, although the role of SUMOylation was not described. Thus, LIN-1 interacts with two different homologs of vertebrate Mi-2: RAD- 26 and LET-418. One possible explanation for the relatively mild vulval defects caused by rad- 26(lf) mutations is that one or more genes function redundantly with rad-26. In particular, chd-3 and let-418 are highly similar to rad-26, and von Zelewsky et al. (2000) showed that let-418 and chd-3 function redundantly during vulval development. An important goal of future studies is to determine the unique and overlapping activities of rad-26, let-418 and chd-3. Together, these studies indicate that recruitment of the NuRD complex by LIN-1, and perhaps by additional transcription factors, is critical for inhibition of vulval cell fates. The identification of RAD-26 provides important new insights into the mechanisms by which SUMOylated LIN-1 mediates transcriptional repression, since this protein has not been previously reported to interact with LIN-1 or contribute to vulval cell fate specification. 24
25 Furthermore, these findings have implications for other transcription factors. Over 30 transcription factors, including seven ETS proteins, have been demonstrated to be posttranslationally modified by SUMO-1 (Gill, 2003; Hay, 2005). In the majority of these cases, SUMOylation promotes repression of transcription. A critical question in this field is: how does SUMOylation mediate transcriptional repression? SUMOylation of p300 and Elk-1 promotes an association with histone deacetylases (HDACs) that repress transcription (Girdwood et al., 2003; Yang and Sharrocks, 2004). However, these HDACs were not shown to directly bind the SUMO moieties. Furthermore, the HDAC inhibitor trichostatin A (TSA) blocks repression at some, but not all, promoters (Yang et al., 2002). Thus, SUMO is likely to mediate transcriptional repression by additional, undefined mechanisms. In Drosophila cells, Mi-2, MEP-1 and a polycomb protein are important for SUMO-dependent transcriptional repression (Stielow et al., 2007). Our studies define a new mechanisms by which SUMO mediates transcriptional repression: recruitment of a nucleosome-remodeling enzyme. Given that the NuRD complex possesses both nucleosome remodeling and histone deacetylase activities, the direct binding of SUMO-1 to RAD-26/Mi-2β may account for SUMO-1-mediated recruitment of HDAC activity observed in vertebrates (Girdwood et al., 2003; Yang and Sharrocks, 2004). The EGL-27/MTA1 component of the NuRD complex binds LIN-1 independent of LIN-1 SUMOylation and inhibits vulval cell fates: We discovered EGL-27/MTA1 as a LIN- 1-interacting protein in yeast cells. EGL-27 binds two domains in the carboxy-terminus of LIN- 1 in yeast cells, and binding is independent of SUMOylation. Furthermore, EGL-27 directly binds LIN-1 in purified extracts. These results suggest EGL-27 binds LIN-1 in worms; however, this biochemical interaction in worms has not been analyzed directly. The EGL-27 expression 25
26 pattern was determined using an EGL-27::GFP reporter gene by Solari et al. (1999) and Herman et al. (1999). EGL-27 is expressed in somatic nuclei of most or all cells from the 50-cell stage of embryogenesis through adulthood. The results indicate that EGL-27 is expressed in the vulval precursor cells, which is consistent with the model that it interacts with LIN-1. EGL-27 is a homolog of the MTA1 component of the NuRD transcriptional repression complex, and we hypothesize that recruitment of EGL-27 to LIN-1 mediates transcriptional repression of vulval cell fate genes. Consistent with this hypothesis, previous genetic studies showed that egl-27 inhibits vulval cell fates (Solari and Ahringer, 2000; Chen and Han, 2001a). Our genetic analyses extend these findings by providing evidence that that egl-27 interacts with multiple genes in the EGFR/Ras/MAPK signaling pathway that function in Pn.p cells, including lin-1. egl-27 did not cause a Muv phenotype in combination with a class A or class B synmuv genes that function in hyp7. These genetic results are consistent with the model that egl-27 functions in the Ras signaling pathway or that egl-27 functions in parallel to the Ras signaling pathway, and they do not establish the cellular site of action of egl-27. The Muv phenotype caused by reducing the activity of egl-27 is not as strong as the Muv phenotype caused by reducing the activity of lin-1, suggesting that lin-1 retains some function to inhibit vulval cell fates in the absence of egl- 27. One possible explanation for the relatively mild vulval defects caused by egl-27(lf) mutations is that one or more genes function redundantly with egl-27. In particular, lin-40/egr-1 is highly similar to egl-27. Notably, Chen and Han (2001) showed that lin-40 negatively regulates vulval fate specification, and Solari et al (1999) showed that egl-27 and lin-40/egr-1 function redundantly during embryonic patterning. An important goal of future studies is to determine the unique and overlapping activities of egl-27 and lin
27 By searching for proteins that interact with EGL-27, we demonstrated that EGL-27 could interact with UBC-9 and MEP-1. UBC-9 is an E2 SUMO conjugating enzyme that binds consensus SUMOylation motifs and mediates the covalent attachment of SUMO-1. EGL-27 contains five consensus SUMOylation motifs, including four in the EGL-27( ) fragment that binds UBC-9, suggesting that EGL-27 may be SUMOylated. MEP-1 is a component of the NURD transcriptional repression complex (Unhavaithaya et al., 2002). We previously demonstrated that MEP-1 interacts with the SUMOylated N-terminus of LIN-1 and functions to inhibit vulval cell fates (Leight et al., 2005). This interaction of MEP-1 with SUMO may be conserved, since Drosophila MEP-1 mediates SUMO-dependent transcriptional repression (Stielow et al., 2007). These observations suggest that MEP-1 may be recruited by two mechanisms: directly by binding to the SUMOylated N-terminus of LIN-1 and indirectly by binding to EGL-27. Our studies demonstrate that LIN-1 binds multiple proteins that are predicted to function to repress transcription: RAD-26 and MEP-1 bind two SUMOylation motifs in the aminoterminus of LIN-1, and EGL-27 binds two domains in the carboxy-terminus of LIN-1. LIN-1 also binds the LIN-31 transcription factor (Tan et al., 1998) and LET-418 (Guerry et al., 2007). If these proteins function independently to modify chromatin and repress transcription, then elimination of one component might impair, but not abrogate, the function of the complex, resulting in a weak Muv phenotype. Mutations of rad-26, egl-27 and mep-1 all cause weak Muv phenotypes, consistent with the model that LIN-1 employs multiple independent mechanisms to repress transcription. 27
28 Phosphorylation of LIN-1 by ERK converts LIN-1 from a transcriptional repressor to a transcriptional activator: We examined how ERK affects the transcriptional activity of LIN-1 to determine if phosphorylation by ERK abrogates LIN-1 activity or converts LIN-1 to a transcriptional activator. If LIN-1 cannot be phosphorylated by ERK because the docking sites for ERK are mutated, then the carboxy-terminus of LIN-1 repressed transcription by about fivefold. By contrast, the carboxy-terminus of LIN-1 activated transcription by about 80-fold when ERK docking sites were intact. Stimulating ERK activity with bfgf growth factor caused a further two-fold increase in transcriptional activation. Overall the difference between transcriptional activation by LIN-1 that cannot be phosphorylated and LIN-1 that is maximally phosphorylated is about 900-fold. The dramatic ERK-responsive transcriptional activation potential of the carboxy-terminus of LIN-1 strongly supports the model that ERK switches LIN- 1 from a transcriptional repressor to a phosphorylated transcriptional activator. These results extend the similarities between LIN-1 and vertebrate Elk-1, documenting the relevance of LIN-1 as a model for understanding vertebrate biology. Elk-1 was initially characterized as a ternary complex factor that binds the serum response element present in the promoters of immediate early genes, such as c-fos (Treisman, 1994). Upon activation of the RTK/Ras/ERK pathway, ERK phosphorylates multiple sites in the carboxy-terminus of Elk-1, resulting in potent transcriptional activation of immediate early genes (Janknecht et al., 1993; Marais et al., 1993; Cruzalegui et al., 1999). By contrast, studies of the Drosophila ETS protein Aop/Yan indicated that in the absence of activated ERK, Aop/Yan functions as an inhibitor of neuronal cell fates by mediating transcriptional repression (O Neill et al., 1994). ERK-mediated phosphorylation of Aop/Yan results in its nuclear export and degradation, thus abrogating the 28
29 repressor activity of Aop/Yan and allowing cells to adopt neuronal cell fates (Rebay and Rubin, 1995). Our results indicate that LIN-1 regulation is different from that of Aop/Yan. These molecular studies of LIN-1 provide a unifying hypothesis that explains diverse genetic observations of lin-1. In two instances where the EGFR/Ras/MAPK pathway functions during early development, lin-1 appears to be a positive mediator of this pathway. Howard et al. (2002) showed that lin-1(lf) mutations could cause a larval lethal phenotype similar to loss-offunction mutations in positive regulators of EGFR/Ras/MAPK signaling, suggesting that lin-1 is a positive mediator of this signaling pathway during larval development. Jiang and Wu (2014) recently reported that the lin-3, let-23, let-60, mpk-1, lin-1 pathway promotes specific programmed cell death events; phosphorylated LIN-1 directly binds the egl-1 promoter and activates transcription of egl-1, consistent with a transcriptional activation function of phosphorylated LIN-1. The function of lin-1 during vulval development is more complex. lin-1 null mutations cause P3.p, P4.p and P8.p to adopt vulval cell fates, suggesting that lin-1 is necessary to inhibit vulval cell fates and may not be necessary to promote vulval cell fates (Sulston and Horvitz, 1981; Ferguson et al., 1987). However, Tiensuu et al. (2005) showed that egl-17 expression in P6.p is abnormal in lin-1(lf) mutants, indicating that lin-1 function is required in P6.p for normal gene expression. Wagmaister et al. (2006) identified lin-39 as a directly regulated LIN-1 target gene. In lin-1(lf) mutants, lin-39 gene expression is upregulated in P3.p, P4.p and P8.p, consistent with LIN-1 functioning as a transcriptional repressor in these cells. In addition, lin-39 gene expression is down regulated in P6.p, consistent with LIN-1 functioning as a transcriptional activator in P6.p. Zhang and Greenwald (2011) demonstrated that lag-2, which encodes the ligand for Notch, is transcriptionaly repressed in all six Pn.p cells prior to ERK activation, and this repression requires lin-1. Farooqui et al. (2012) proposed that 29
30 non-phosphorylated LIN-1 directly binds and activates the let-502 promoter in the secondary cells P5.p and P7.p, suggesting that LIN-1 may have different regulatory mechanisms in the primary cell P6.p, the secondary cells P5.p and P7.p, and tertiary cells P3.p, P4.p and P8.p. We propose that SUMOylated LIN-1 represses transcription of target genes in P3.p, P4.p and P8.p, and phosphorylated LIN-1 activates transcription of target genes in P6.p (Figure 7). In lin-1(lf) mutants, the absence of SUMOylated LIN-1 causes derepression of target genes in P3.p, P4.p and P8.p, and these derepressed target genes are sufficient to promote a vulval cell fate. Although the absence of phosphorylated LIN-1 in P6.p results in diminished activation of target genes, P6.p can still adopt the proliferative vulval cell fate. An important question that is raised by these findings is: how does phosphorylation by ERK convert LIN-1 from a transcriptional repressor to a transcriptional activator? Activation of ERK might result in decreased SUMOylation of LIN-1. For example, the steady-state level of SUMOylated Elk-1 decreases when ERK is activated in vertebrate cells (Yang et al., 2003). Phosphorylation of LIN-1 by ERK might disrupt the binding interaction of LIN-1 with transcriptional repressors. In particular, the EGL-27 transcriptional repressor binds two domains in the carboxy-terminus of LIN-1 that contain multiple consensus phosphorylation sites for ERK. Phosphorylated LIN-1 might recruit transcriptional activators. Phosphorylation of Ets-1 and Ets- 2 by ERK mediates recruitment of the transcriptional coactivators CREB binding protein (CBP) and p300 (Foulds et al., 2004). Phosphorylated Elk-1 interacts with the mediator subunit Sur2 to promote transcriptional activation (Stevens et al., 2002; Cantin et al., 2003), and the C. elegans sur-2 gene promotes vulval cell fates (Singh and Han, 1995), suggesting that phosphorylated LIN-1 might recruit SUR-2 to promote transcriptional activation. Proteins recruited by phosphorylated LIN-1 might reverse the chromatin modifications caused by transcriptional 30
31 repressors, such as the histone deacetylation and nucleosome remodeling caused by the NuRD complex. These mechanisms are not mutually exclusive, and important goals of future studies are to determine the relative importance of these mechanisms and identify transcriptional activators recruited to LIN-1. ACKNOWLEDGEMENTS rad-26(tm1991) was provided by Shohei Mitani of the National Bioresource Project. We thank Tim Schedl, Kurt Warnhoff and members of the Kornfeld laboratory for insightful comments on the manuscript. This research was supported by NIH grants GM068598, CA84271 and F32 GM66605 (K.K.). K.K. was a Scholar of the Leukemia and Lymphoma Society and a Senior Scholar of the Ellison Medical Foundation. E.R.L. was supported by Keck and NRSA postdoctoral fellowships. 31
32 REFERENCES Andersen, E.C., and H.R. Horvitz, 2007 Two C. elegans histone methyltransferases repress lin-3 EGF transcription to inhibit vulval development. Development 134: Aroian, R.V., Koga, M., Mendel, J.E., Ohshima, Y., and P.W. Sternberg, 1990 The let-23 gene necessary for Caenorhabditis elegans vulval induction encodes a tyrosine kinase of the EGF receptor subfamily. Nature 348: Bannister, A. J., Zegerman, P., Partridge, J. F., Miska, E. A., Thomas, J. O., Allshire, R. C., and T. Kouzarides, 2001 Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410: Bartel, P. L., Chien, C.-T., Sternglanz, R., and S. Fields, 1993 Cellular Interactions in Development: A Practical Approach, (ed. D. A. Hartley), p Oxford: Oxford University Press. Beitel, G. J., Clark, S. G., and H. R. Horvitz, 1990 Caenorhabditis elegans ras gene let-60 acts as a switch in the pathway of vulval induction. Nature 348: Beitel, G. J., Tuck, S., Greenwald, I., and H. R. Horvitz, 1995 The Caenorhabditis elegans gene lin-1 encodes an ETS-domain protein and defines a branch of the vulval induction pathway. Genes Dev 9: Brenner, S., 1974 The genetics of Caenorhabditis elegans. Genetics 77: Cantin, G.T., Stevens, J.L., and A.J. Berk, 2003 Activation domain-mediator interactions promote transcription preinitiation complex assembly on promoter DNA. Proc. Natl. Acad. Sci. USA 100: Chang, C., Hopper, N.A., and P. W. Sternberg, 2000 Caenorhabditis elegans SOS-1 is necessary for multiple RAS-mediated developmental signals. EMBO J. 19:
33 Chen, Z., and M. Han, 2001a Role of C. elegans lin-40 MTA in vulval fate specification and morphogenesis. Development 128: Chen, Z., and M. Han, 2001b C. elegans Rb, NuRD, and Ras regulate lin-39-mediated cell fusion during vulval fate specification. Curr Biol. 11: Clark, S.G., Stern, M.J., and H.R. Horvitz, 1992 C. elegans cell-signaling gene sem-5 encodes a protein with SH2 and SH3 domains. Nature 356: Cruzalegui, F. H., Cano, E., and R. Treisman, 1999 ERK activation induces phosphorylation of Elk-1 at multiple S/T-P motifs to high stoichiometry. Oncogene 18: Cui, M., Chen, J., Myers, T.R., Hwang, B.J., Sternberg, P.W., Greenwald, I., and M. Han, 2006 SynMuv genes redundantly inhibit lin-3/egf expression to prevent inappropriate vulval induction in C. elegans. Dev. Cell 10: Dittmer, J., 2003 The biology of the Ets1 proto-oncogene. Molecular Cancer 2: 29. Fantz, D. A., Jacobs, D., Glossip, D., and K. Kornfeld, 2001 Docking sites on substrate proteins direct extracellular signal-regulated kinase to phosphorylate specific residues. J Biol Chem 276: Farooqui, S., Pellegrino, M.W., Rimann, I., Morf, M.K., Muller, L., Frohli, E., and A. Hajnal, 2012 Coordinated lumen contraction and expansion during vulval tube morphogenesis in Caenorhabditis elegans. Dev. Cell 23: Feilotter, H. E., Hannon, G. J., Ruddell, C. J., and D. Beach, 1994 Construction of an improved host strain for two hybrid screening. Nucleic Acids Res. 22: Ferguson, E.L., and H. R. Horvitz, 1989 The multivulva phenotype of certain Caenorhabditis elegans mutants results from defects in two functionally redundant pathways. Genetics 123:
34 Ferguson, E. L., Sternberg, P. W., and H. R. Horvitz, 1987 A genetic pathway for the specification of the vulval cell lineages of Caenorhabditis elegans. Nature 326: Foulds, C. E., Nelson, M. L., Blaszczak, A. G., and B. J. Graves, 2004 Ras/mitogen-activated protein kinase signaling activates Ets-1 and Ets-2 by CBP/p300 recruitment. Mol Cell Biol. 24: Frangioni, J. V., and B. G. Neel, 1993 Solubilization and purification of enzymatically active glutathione S-transferase (pgex) fusion proteins. Anal Biochem. 210: Gill, G., 2003 Post-translational modification by the small ubiquitin-related modifier SUMO has big effects on transcription factor activity. Curr Opin Genet Dev 13: Girdwood, D., Bumpass, D., Vaughan, O. A., Thain, A., Anderson, L. A., Snowden, A. W., Garcia-Wilson, E., Perkins, N. D., and R. T. Hay, 2003 P300 transcriptional repression is mediated by SUMO modification. Mol Cell 11: Greenwald, I. S., 1997 Development of the vulva, pp in C. elegans II, edited by D. L. Riddle, T. Blumenthal, B. J. Meyer and J. R. Preiss. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY. Guerry, F., Marti, C., Zhang, Y., Moroni, P. S., Jaquiery, E., and F. Muller, 2007 The Mi-2 nucleosome-remodeling protein LET-418 is targeted via LIN-1/ETS to the promoter of lin- 39/Hox during vulval development in C. elegans. Dev Biol. 306: Han, M., and P.W. Sternberg, 1990 let-60, a gene that specifies cell fates during C. elegans vulval induction, encodes a ras protein. Cell 63: Han, M., Golden, A., Han, Y., and P.W. Sternberg, 1993 C. elegans lin-45 raf gene participates in let-60 ras-stimulated vulval differentiation. Nature 363: Hay, R. T., 2005 SUMO: a history of modification. Mol Cell 18:
35 Herman, M. A., Ch'ng, Q., Hettenbach, S. M., Ratliff, T. M., Kenyon, C., and R. K Herman, 1999 EGL-27 is similar to a metastasis-associated factor and controls cell polarity and cell migration in C. elegans. Development 126: Herman, R.K., and E.M. Hedgecock, 1990 Limitation of the size of the vulval primordium of Caenorhabditis elegans by lin-15 expression in surrounding hypodermis. Nature 348: Hill, R.J., and P.W. Sternberg, 1992 The gene lin-3 encodes an inductive signal for vulval development in C. elegans. Nature 358: Horvitz, H.R., and P.W. Sternberg, 1991 Multiple intercellular signalling systems control the development of the Caenorhabditis elegans vulva. Nature 351: Howard, R. M., and M. V. Sundaram, 2002 C. elegans EOR-1/PLZF and EOR-2 positively regulate Ras and Wnt signaling and function redundantly with LIN-25 and the SUR-2 Mediator component. Genes Dev. 16: Hsu, V., Zobel, C.L., Lambie, E.J., Schedl, T., and K. Kornfeld, 2002 Caenorhabditis elegans lin-45 raf is essential for larval viability, fertility and the induction of vulval cell fates. Genetics 160: Jacobs, D., Beitel, G. J., Clark, S. G., Horvitz, H. R., and K. Kornfeld, 1998 Gain-of-function mutations in the Caenorhabditis elegans lin-1 ETS gene identify a C-terminal regulatory domain phosphorylated by ERK MAP kinase. Genetics 149: Jacobs, D., Glossip, D., Xing, H., Muslin, A. J., and K. Kornfeld, 1999 Multiple docking sites on substrate proteins form a modular system that mediates recognition by ERK MAP kinase. Genes Dev 13: Janknecht, R., Ernst, W. H., Pingoud, V., and A. Nordheim, 1993 Activation of ternary complex factor Elk-1 by MAP kinases. EMBO J 12:
36 Jiang, H-S., and Y-C. Wu, 2014 LIN-3/EGF promotes the programmed cell death of specific cells in Caenorhabditis elegans by transcriptional activation of the pro-apoptotic gene egl-1. PLOS Genetics 10: e Kamath, R. S., Fraser, A. G., Dong, Y., Poulin, G., Durbin, R., Gotta, M., Kanapin, A., Le Bot, N., Moreno, S., Sohrmann, M., Welchman, D. P., Ziperlen, P., and J. Ahringer, 2003 Systemic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421: Kornfeld, K., 1997 Vulval development in Caenorhabditis elegans. Trends Genet 13: Kornfeld, K., Guan, K.L., and H.R. Horvitz, 1995 The Caenorhabditis elegans gene mek-2 is required for vulval induction and encodes a protein similar to the protein kinase MEK. Genes Dev. 9: Lackner, M. R., Kornfeld, K., Miller, L. M., Horvitz, H. R., and S. K. Kim, 1994 A MAP kinase homolog, mpk-1, is involved in ras-mediated induction of vulval cell fates in Caenorhabditis elegans. Genes Dev 8: Lachner, M., O'Carroll, D., Rea, S., Mechtler, K., and T. Jenuwein, 2001 Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410: Lee, M. H, Ohmachi, M., Arur, S., Nayak, S., Francis, R., Church, D., Lambie, E., and T. Schedl, 2007 Multiple functions and dynamic activation of MPK-1 extracellular signal-regulated kinase signaling in Caenorhabditis elegans germline development. Genetics 177: Leight, E. R., Glossip, D., and K. Kornfeld, 2005 Sumoylation of LIN-1 promotes transcriptional repression and inhibition of vulval cell fates. Development 132: Lu, X., and H.R. Horvitz, 1998 lin-35 and lin-53, two genes that antagonize a C. elegans Ras pathway, encode proteins similar to Rb and its binding protein RbAp48. Cell 95:
37 Marais, R., Wynne, J., and R. Treisman, 1993 The SRF accessory protein Elk-1 contains a growth factor-regulated transcriptional activation domain. Cell 73: Miley, G. R., Fantz, D., Glossip, D., Lu, X., Saito, R. M., Palmer, R. E., Inoue, T., van den Heuvel, S., Sternberg, P. W., and K. Kornfeld, 2004 Identification of residues of the Caenorhabditis elegans LIN-1 ETS domain that are necessary for DNA binding and regulation of vulval cell fates. Genetics 167: Nielsen, S. J., Schneider, R., Bauer, U. M., Bannister, A. J., Morrison, A., O'Carroll, D., Firestein, R., Cleary, M., Jenuwein, T., Herrera, R. E., and T. Kouzarides, 2001 Rb targets histone H3 methylation and HP1 to promoters. Nature 412: O'Neill, E. M., Rebay, I., Tjian, R., and G. M. Rubin, 1994 The activities of two Ets-related transcription factors required for Drosophila eye development are modulated by the Ras/MAPK pathway. Cell 78: Piano, F., Schetter, A. J., Morton, D. G., Gunsalus, K. C., Reinke, V., Kim, S. K., and K. J. Kemphues, 2002 Gene clustering based on RNAi phenotypes of ovary-enriched genes in C. elegans. Curr Biol. 12: Poulin, G., Dong, Y., Fraser, A. G., Hopper, N. A., and J. Ahringer, 2005 Chromatin regulation and sumoylation in the inhibition of Ras-induced vulval development in Caenorhabditis elegans. EMBO J. 24: Rebay, I., and G. M. Rubin, 1995 Yan functions as a general inhibitor of differentiation and is negatively regulated by activation of the Ras1/MAPK pathway. Cell 81: Sambrook, J., Fritsch, E. F., and T. Maniatis, 1989 Molecular Cloning: A Laboratory Manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY. 37
38 Sampson, D. A., Wang, M., and M. J. Matunis, 2001 The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J Biol Chem 276: Schiestl, R. H., and R. D. Gietz, 1989 High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Current Genetics 16: Singh, N., and M. Han, 1995 sur-2, a novel gene, functions late in the let-60 ras-mediated signaling pathway during Caenorhabditis elegans vulval induction. Genes Dev 9: Solari, F., Bateman, A., and J. Ahringer, 1999 The Caenorhabditis elegans genes egl-27 and egr-1 are similar to MTA1, a member of a chromatin regulatory complex, and are redundantly required for embryonic patterning. Development 126: Solari, F., and J. Ahringer, 2000 NURD-complex genes antagonise Ras-induced vulval development in Caenorhabditis elegans. Curr Biol 10: Sonnichsen, B., Koski, L. B., Walsh, A., Marschall, P., Neumann, B., Brehm, M., Alleaume, A. M., Artelt, J., Bettencourt, P., Cassin, E., Hewitson, M., Holz, C., Khan, M., Lazik, S., Martin, C., Nitzsche, B., Ruer, M., Stamford, J., Winzi, M., Heinkel, R., Roder, M., Finell, J., Hantsch, H., Jones, S. J., Jones, M., Piano, F., Gunsalus, K. C., Oegema, K., Gonczy, P., Coulson, A., Hyman, A. A., and C. J. Echeverri, 2005 Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans. Nature 434: Sternberg, P.W., 2005 Vulval development. WormBook 25:1-28. Stevens, J.L., Cantin, G.T., Wang, G., Shevchenko, A., Shevchenko, A., and A.J. Berk, 2002 Transcription control by E1A and MAP kinase pathway via Sur2 mediator subunit. Science 296:
39 Stielow, B., Sapetschnig, A., Kruger, I., Kunert, N., Brehm, A., Boutros, M., and G. Suske, 2007 Identification of SUMO-dependent chromatin-associated transcriptional repression components by a genome-wide RNAi screen. Mol. Cell 29: Sulston, J. E., and H. R. Horvitz, 1981 Abnormal cell lineages in mutants of the nematode Caenorhabditis elegans. Dev Biol 82: Sundaram, M.V., 2013 Canonical RTK-Ras-ERK signaling and related alternative pathways.wormbook. 11:1-38. Tan, P. B., Lackner, M. R., and S. K. Kim, 1998 MAP kinase signaling specificity mediated by the LIN -1 Ets/LIN-31 WH transcription factor complex during C. elegans vulval induction. Cell 93: Tiensuu, T., Larsen, M. K., Vernersson, E., and S. Tuck, 2005 lin-1 has both positive and negative functions in specifying multiple cell fates induced by Ras/MAP kinase signaling in C. elegans. Dev Biol. 286: Timmons, L., and A. Fire, 1998 Specific interference by ingested dsrna. Nature 395: 854. Tong, J. K., Hassig, C. A., Schnitzler, G. R., Kingston, R. E., and S. L. Schreiber, 1998 Chromatin deacetylation by an ATP-dependent nucleosome remodelling complex. Nature 395: Treisman, R Ternary complex factors: growth factor regulated transcriptional activators. Curr Opin Genet Dev 4: Trent, C., Tsung, N., and H.R. Horvitz, 1983 Egg-laying defective mutants of the nematode Caenorhabditis elegans. Genetics 104:
40 Unhavaithaya, Y., Shin, T. H., Miliaras, N., Lee, J., Oyama, T., and C. C. Mello, 2002 MEP-1 and a homolog of the NURD complex component Mi-2 act together to maintain germline-soma distinctions in C. elegans. Cell 111: Vojtek, A. B., Hollenberg, S. M., and J. A. Cooper, 1993 Mammalian Ras interacts directly with the serine/threonine kinase Raf. Cell 74: von Zelewsky, T., Palladino, F., Brunschwig, K., Tobler, H., Hajnal, A., and F. Muller, 2000 The C. elegans Mi-2 chromatin-remodelling proteins function in vulval cell fate determination. Development 127: Wade, P. A., Jones, P. L., Vermaak, D., and A. P. Wolffe, 1998 A multiple subunit Mi-2 histone deacetylase from Xenopus laevis cofractionates with an associated Snf2 superfamily ATPase. Curr Biol 8: Wagmaister, J. A., Miley, G. R., Morris, C. A., Gleason, J. E., Miller, L. M., Kornfeld, K., and D. M. Eisenmann, 2006 Identification of cis-regulatory elements from the C. elegans Hox gene lin-39 required for embryonic expression and for regulation by the transcription factors LIN-1, LIN-31 and LIN-39. Dev Biol. 297: Wang, H. B., and Y. Zhang 2001 Mi2, an auto-antigen for dermatomyositis, is an ATPdependent nucleosome remodeling factor. Nucleic Acids Res. 29: Wu, Y., and M. Han, 1994 Suppression of activated Let-60 ras protein defines a role of Caenorhabditis elegans Sur-1 MAP kinase in vulval differentiation. Genes Dev. 8: Wu, Y., Han, M., and K.L. Guan, 1995 MEK-2, a Caenorhabditis elegans MAP kinase kinase, functions in Ras-mediated vulval induction and other developmental events. Genes Dev. 9: Xue, Y., Wong, J., Moreno, G. T., Young, M. K., Cote, J., and W. Wang, 1998 NURD, a novel 40
41 complex with both ATP-dependent chromatin-remodeling and histone deacetylase activities. Mol Cell 2: Yang, S. H., Bumpass, D. C., Perkins, N. D., and A. D. Sharrocks, 2002 The ETS domain transcription factor Elk-1 contains a novel class of repression domain. Mol Cell Biol. 22: Yang, S. H., Jaffray, E., Hay, R. T., and A. D. Sharrocks, 2003 Dynamic interplay of the SUMO and ERK pathways in regulating Elk-1 transcriptional activity. Mol Cell 12: Yang, S. H., and A. D. Sharrocks, 2004 SUMO promotes HDAC-mediated transcriptional repression. Mol Cell 13: Zhang, X., and I. Greenwald, 2011 Spatial regulation of lag-2 transcription during vulval precursor cell fate patterning in Caenorhabditis elegans. Genetics 188: Zhang, Y., LeRoy, G., Seelig, H. P., Lane, W. S., and D. Reinberg, 1998 The dermatomyositisspecific autoantigen Mi2 is a component of a complex containing histone deacetylase and nucleosome remodeling activities. Cell 95:
42 FIGURE LEGENDS Figure 1: LIN-1 binds RAD-26 and SET-6 in yeast cells. (A) Schematic of LIN-1 showing the ETS DNA binding domain (dotted), consensus SUMOylation motifs (ψkxe), and docking sites for ERK MAP kinase (D-domain [white] and FQFP motif [black]). Amino acid numbers are shown below. (B) SET-6 contains a SET lysine methyltransferase domain (gray). RAD-26 contains a DNA-dependent ATPase domain of the SNF2 family N-terminal class (white) and a helicase C-terminal domain (gray). (C) Interactions between LexA DNA binding domain (LA):LIN-1 fusion proteins and SET-6(22-538) or RAD- 26(17-655) fused to the GAL4 activation domain (GAL4AD) were measured qualitatively. Symbols indicate robust activation (+++) and no detectable activation (-) of a LexA-dependent lacz reporter gene. (D, E) Interactions were measured quantitatively by assaying LexAdependent β-galactosidase activity. Bars represent the average and standard deviation of 3-6 independent yeast transformants. Values were normalized by setting the signal with wild-type LA:LIN-1 to 100. (F) Expression of LA:LIN-1 proteins was analyzed by Western blotting using an anti-la antibody. Lanes 1-14 corresponds to proteins in columns 1-14 of panel D. Figure 2: SUMO was sufficient to increase binding of SET-6 to LIN-1 in yeast cells. Associations of SET-6 with LA fusion proteins were monitored using the yeast two-hybrid system. Bars represent the average and standard deviation of LexA-dependent β-galactosidase activity from three independent yeast transformants. Values were normalized by setting the interaction of SET-6 with LA:LIN-1(1-64) to 100. The LA fusion proteins were expressed at similar levels as determined by Western blotting (data not shown). 42
43 Figure 3: RAD-26 binds SMO-1 and LIN-1 in purified extracts. 6xHis- and FLAG-tagged RAD-26(17-655) (HF:RAD-26) was incubated with equivalent amounts of glutathione agarose-bound GST, GST:LIN-1(1-64) or GST:SMO-1. Bound HF:RAD-26 was visualized by immunoblotting using α-flag antibody (upper gel). Lane 4 shows 20% of the HF:RAD-26 input. HF:RAD-26 did not bind GST alone as assessed by prolonged exposure of the immunoblot. Input GST fusion proteins were visualized by SDS- PAGE and Simply Blue staining (lower gel). Molecular weight markers (in kda) are indicated. Figure 4: Ectopic vulval protrusions in rad-26 and egl-27 mutant animals. Panels show DIC images of the mid body region. The genotypes are lin-1(e1275) rad- 26(tm1991) (A, B) and egl-27(we3); lin-1(n1790) (C, D). Arrows mark protruding ectopic vulval tissue. Magnification is 400 X. Figure 5: LIN-1 binds EGL-27 in yeast cells and purified extracts. (A) EGL-27 protein (Solari et al., 1999) contains ELM1 (gray), ELM2 (black), SANT (white) and GATA-like zinc finger (gray) domains. (B) The interactions between LA:LIN-1 proteins and MPK-1(1-444) and EGL-27( ) fused to the GAL4 activation domain were measured. Symbols indicate robust (+++), moderate (++), weak (+), and no detectable (-) activation of a LexA-dependent lacz reporter gene. (C) 6xHis- and T7-tagged EGL-27( ) (T7:EGL- 27) was incubated with equivalent amounts of glutathione agarose-bound GST, GST:LIN-1(1-64) or GST:LIN-1( ). Bound T7:EGL-27 was visualized by immunoblotting using α-t7 antibody (upper gel). Lane 4 shows 5% of the T7:EGL-27 input. Input GST fusion proteins 43
44 were visualized by SDS-PAGE and Simply Blue staining (lower gel). Lower molecular weight proteins in lanes 2 and 3 represent degradation products. (D) 6xHis tagged MEP-1 (MEP-1) was incubated with equivalent amounts of glutathione agarose-bound GST or GST:EGL-27(1-1129). Bound MEP-1 was visualized by immunoblotting using anti-his antibody. Lane 1 shows 10% of the MEP-1 input. Figure 6: The carboxy-terminus of LIN-1 contains an ERK-dependent transcriptional activation domain and an ERK-independent repression domain. MC3T3 murine fibroblast cells were transfected with (1) the pfr-luc reporter plasmid that contains five tandem GAL4 binding sites upstream of a basal promoter that regulates expression of luciferase, (2) an expression plasmid that encodes GAL4 DNA binding domain (G4) alone or fused to the indicated fragment of LIN-1 or Elk-1 and (3) a reporter plasmid that encodes β- galactosidase to measure transfection efficiency. Cells were untreated (-) or treated with basic fibroblast growth factor (FGF) to activate ERK (+). (A) The schematics illustrate the ERK MAP kinase docking sites, the D-domain (D) and the FQFP motif (F). Asterisks indicate 15 S/TP motifs in LIN-1 that are potential ERK phosphorylation sites and six S/TP motifs in Elk-1 that are phosphorylated by ERK (Cruzalegui et al., 1999). Bars indicate luciferase activity divided by β-galactosidase activity. Values were normalized by setting the value for G4 alone in the absence of FGF to 1.0. Values represent the average and standard deviation of 2-5 independent transfections conducted in parallel. (B) The schematic of LIN-1 illustrates the VK 169 DE consensus SUMOylation motif (above) and two domains that are sufficient to bind EGL-27 (hatched). 44
45 Figure 7: ERK converts LIN-1 from a SUMOylated transcriptional repressor to a phosphorylated transcriptional activator. A model for the regulation of vulval cell fates by LIN-1. LIN-1 represses 1 vulval cell fate genes by recruiting RAD-26 and MEP-1 to its SUMOylated amino-terminus and EGL-27 to its unsumoylated carboxy-terminus. RAD-26 remodels nucleosomes, and EGL-27 and MEP-1 facilitate histone deacetylation through their association with HDAC-1 and HDAC-2. Following activation of the EGFR/Ras/MAPK pathway in P6.p, ERK converts LIN-1 to a phosphorylated transcriptional activator of 1 vulval cell fate genes. 45
46 Table 1 Genetic analysis of rad-26 Genotype a RNAi b %Muv c n d 1. Wild type e NA rad-26(tm1991) e NA lin-1(e1275) e NA lin-1(e1275) rad-26(tm1991) e NA Wild type NA rad-26(tm1991) NA lin-1(n1790) NA lin-1(n1790) rad-26(tm1991) NA rad-26(tm1991); set-6(ok2195) NA egl-27(we3); rad-26(tm1991) NA mpk-1(oz140); rad-26(tm1991) NA rad-26(tm1911); lin15a(n767) NA rad-26(tm1911); lin-15b(n744) NA Wild type Control Wild type rad let-60(n1046) e Control let-60(n1046) e rad lin-1(n1790) Control lin-1(n1790) rad a To facilitate comparisons, the following experiments were conducted in parallel: 1-4, 5-8, 14-15, 16-17, and The difference between the percentage of the lin-1(n1790) Muv phenotype in line 7 and line 18 might reflect day-to-day variability or the difference between control RNAi bacteria and standard OP50 bacteria as a food source. b L4 hermaphrodites were fed E. coli containing a control plasmid or a plasmid that expresses double-stranded RNA from the rad-26 gene, and the vulval phenotype of the progeny that grew on these dishes was scored. NA, not applicable. c Adult hermaphrodites were scored as multivulval (Muv) if they displayed one or more ventral protrusions displaced from the site of the vulva. Hermaphrodites were propagated at 20ºC unless otherwise indicated. The chi-square test was used to compare two values. The following comparisons were significantly different (p<0.05): 1 and 3, 2 and 4, 3 and 4, 5 and 7, 6 and 8, 7 and 8, 16 and 17, 18 and 19. d n, number of hermaphrodites examined. e Hermaphrodites were propagated at 15ºC because lin-1(e1275) and let-60(n1046) cause heat-sensitive Muv phenotypes.
47 Table 2 Genetic analysis of egl-27 Genotype a RNAi %Muv c n 1. Wild type NA egl-27(n170) NA egl-27(we3) NA lin-1(n1790) NA egl-27(we3); lin-1(n1790) NA egl-27(we3); lin-15a(n767) NA egl-27(we3); lin-15b(n744) NA egl-27(we3); set-6(ok2195) NA Wild type Control Wild type egl let-60(n1046) b Control let-60(n1046) b egl lin-1(n1790) Control lin-1(n1790) egl Columns are as described in Table 1. a To facilitate comparisons, the following experiments were conducted in parallel; 4-5, 9-10,11-12, and The difference between the percentage of the let-60(n1046) Muv phenotype shown in line 11 compared to line 16 of Table 1 may reflect day-to-day variability in temperature or humidity. b Hermaphrodites were propagated and scored at 15º c The chi-square test was used to compare two values. The following comparisons were significantly different (p<0.05): 1 and 2, 1 and 3, 1 and 4, 3 and 5, 4 and 5, 9 and 10, 11 and 12, 13 and 14.
48 Figure 1 A LIN-1 B SET-6 D SET RAD-26 ATPase H LA:LIN-1(1-64) 2. LA:LIN-1(1-64; 1-8A) 3. LA:LIN-1(1-64; 9-16A) 4. LA:LIN-1(1-64; 17-24A) 5. LA:LIN-1(1-64; 25-32A) 6. LA:LIN-1(1-64; 33-40A) 7. LA:LIN-1(1-64; 41-48A) 8. LA:LIN-1(1-64; 49-56A) 9. LA:LIN-1(1-64; 57-64A) 10. LA:LIN-1(1-64: V9A) 11. LA:LIN-1(1-64: K10A) 12. LA:LIN-1(1-64: K11A) 13. LA:LIN-1(1-64: E12A) 14. LA E VK 10 KE ETS D F LA:LIN-1 ( ) VK 169 DE C Interaction with SET-6 LA:LIN-1(1-252) Relative Light Units VK 169 DE AA LA:LIN-1 ( ; A) 169 AA VK 10 KE VK 169 DE ETS SET-6 RAD LA:LIN-1(1-64) LA:LIN-1(65-145) ETS LA:LIN-1( ) LA Interaction with RAD Relative Light Units F LA:LIN-1 ( ; K169A) VA 169 DE LA Interaction with SET-6 Interaction with RAD Relative Light Units
49 Figure 2 1. LA:LIN-1(1-64) 2. LA:LIN-1(1-64; 9-16A) 3. LA:SMO-1:LIN-1(1-64; 9-16A) 4. LA:SMO-1 5. LA VK 10 KE 1 8A 8A 64 Interaction with SET Relative Light Units
50 Figure 3 GST GST:LIN-1(1-64) GST:SUMO-1 HF:RAD-26 input (20%) HF:RAD Input GST fusion
51 Figure 4 A B lin-1; rad-26 lin-1; rad-26 C D egl-27; lin-1 egl-27; lin-1
52 5. LA:LIN-1( ) D LA:LIN-1( ) F LA:LIN-1( ) 10. LA LA:LIN-1( ) 9. LA:LIN-1(1-252) T:L ) IN -1 ( ) 5% T7 : inp EGL ut D 441 ETS GS N- Input GST fusion ) 4. LA:LIN-1( ; mut. D/F) D (1 3. LA:LIN-1( ; mut. F) F +++ L- 2. LA:LIN-1( ; mut. D) +++ ST :E G 441 G F D t 1. LA:LIN-1( ) T7:EGL-27 pu MPK-1 EGL-27 in B C G ST 1129 M EP -1 SANT Zn ELM G ST GS T:L I 87 % ELM A EGL-27 1( Figure 5 MEP
INTERCELLULAR signaling is one of the primary descendants) (reviewed by Horvitz and Sternberg
 Copyright 1998 by the Genetics Society of America Gain-of-Function Mutations in the Caenorhabditis elegans lin-1 ETS Gene Identify a C-Terminal Regulatory Domain Phosphorylated by ERK MAP Kinase Dave Jacobs,*
Copyright 1998 by the Genetics Society of America Gain-of-Function Mutations in the Caenorhabditis elegans lin-1 ETS Gene Identify a C-Terminal Regulatory Domain Phosphorylated by ERK MAP Kinase Dave Jacobs,*
Bypass and interaction suppressors; pathway analysis
 Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Bypass and interaction suppressors; pathway analysis The isolation of extragenic suppressors is a powerful tool for identifying genes that encode proteins that function in the same process as a gene of
Cell fate specification by Ras-mediated cell signalling in C. elegans. Teresa Tiensuu
 Cell fate specification by Ras-mediated cell signalling in C. elegans Teresa Tiensuu Umeå Centre for Molecular Pathogenesis Umeå University Umeå Sweden 2003 Copyright Teresa Tiensuu 2003 ISBN 91-7305-525-5
Cell fate specification by Ras-mediated cell signalling in C. elegans Teresa Tiensuu Umeå Centre for Molecular Pathogenesis Umeå University Umeå Sweden 2003 Copyright Teresa Tiensuu 2003 ISBN 91-7305-525-5
Wan-Ju Liu 1,2, John S Reece-Hoyes 3, Albertha JM Walhout 3 and David M Eisenmann 1*
 Liu et al. BMC Developmental Biology 2014, 14:17 RESEARCH ARTICLE Open Access Multiple transcription factors directly regulate Hox gene lin-39 expression in ventral hypodermal cells of the C. elegans embryo
Liu et al. BMC Developmental Biology 2014, 14:17 RESEARCH ARTICLE Open Access Multiple transcription factors directly regulate Hox gene lin-39 expression in ventral hypodermal cells of the C. elegans embryo
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
University of Massachusetts Medical School Wan-Ju Liu University of Maryland
 University of Massachusetts Medical School escholarship@umms Program in Systems Biology Publications and Presentations Program in Systems Biology 5-13-2014 Multiple transcription factors directly regulate
University of Massachusetts Medical School escholarship@umms Program in Systems Biology Publications and Presentations Program in Systems Biology 5-13-2014 Multiple transcription factors directly regulate
CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON
 PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
PROKARYOTE GENES: E. COLI LAC OPERON CHAPTER 13 CHAPTER 13 PROKARYOTE GENES: E. COLI LAC OPERON Figure 1. Electron micrograph of growing E. coli. Some show the constriction at the location where daughter
Eukaryotic Gene Expression
 Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
Eukaryotic Gene Expression Lectures 22-23 Several Features Distinguish Eukaryotic Processes From Mechanisms in Bacteria 123 Eukaryotic Gene Expression Several Features Distinguish Eukaryotic Processes
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles
 Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
Illegitimate translation causes unexpected gene expression from on-target out-of-frame alleles created by CRISPR-Cas9 Shigeru Makino, Ryutaro Fukumura, Yoichi Gondo* Mutagenesis and Genomics Team, RIKEN
A complementation test would be done by crossing the haploid strains and scoring the phenotype in the diploids.
 Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Problem set H answers 1. To study DNA repair mechanisms, geneticists isolated yeast mutants that were sensitive to various types of radiation; for example, mutants that were more sensitive to UV light.
Genetics: Early Online, published on December 29, 2015 as /genetics
 Genetics: Early Online, published on December, 01 as.1/genetics..10 1 The Mediator kinase module restrains epidermal growth factor receptor signaling and represses vulval cell fate specification in Caenorhabditis
Genetics: Early Online, published on December, 01 as.1/genetics..10 1 The Mediator kinase module restrains epidermal growth factor receptor signaling and represses vulval cell fate specification in Caenorhabditis
Regulation of gene Expression in Prokaryotes & Eukaryotes
 Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Regulation of gene Expression in Prokaryotes & Eukaryotes 1 The trp Operon Contains 5 genes coding for proteins (enzymes) required for the synthesis of the amino acid tryptophan. Also contains a promoter
Lecture 18 June 2 nd, Gene Expression Regulation Mutations
 Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
Lecture 18 June 2 nd, 2016 Gene Expression Regulation Mutations From Gene to Protein Central Dogma Replication DNA RNA PROTEIN Transcription Translation RNA Viruses: genome is RNA Reverse Transcriptase
GENETIC CONTROL OF CELL INTERACTIONS IN NEMATODE DEVELOPMENT
 Annu. Rev. Genet. 1991. 25. 411-36 Copyright 199l by Annual Reviews Inc. All rights reserved GENETIC CONTROL OF CELL INTERACTIONS IN NEMATODE DEVELOPMENT E. J. Lambie and Judith Kimble Laboratory of Molecular
Annu. Rev. Genet. 1991. 25. 411-36 Copyright 199l by Annual Reviews Inc. All rights reserved GENETIC CONTROL OF CELL INTERACTIONS IN NEMATODE DEVELOPMENT E. J. Lambie and Judith Kimble Laboratory of Molecular
Cytokines regulate interactions between cells of the hemapoietic system
 Cytokines regulate interactions between cells of the hemapoietic system Some well-known cytokines: Erythropoietin (Epo) G-CSF Thrombopoietin IL-2 INF thrombopoietin Abbas et al. Cellular & Molecular Immunology
Cytokines regulate interactions between cells of the hemapoietic system Some well-known cytokines: Erythropoietin (Epo) G-CSF Thrombopoietin IL-2 INF thrombopoietin Abbas et al. Cellular & Molecular Immunology
REVIEW SESSION. Wednesday, September 15 5:30 PM SHANTZ 242 E
 REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
REVIEW SESSION Wednesday, September 15 5:30 PM SHANTZ 242 E Gene Regulation Gene Regulation Gene expression can be turned on, turned off, turned up or turned down! For example, as test time approaches,
with%dr.%van%buskirk%%%
 with%dr.%van%buskirk%%% How$to$do$well?$ Before$class:$read$the$corresponding$chapter$ Come$to$class$ready$to$par9cipate$in$Top$Hat$ Don t$miss$an$exam!!!!!!!!!!!!!!!!!!!!!!!!!!$ But$I m$not$good$with$science
with%dr.%van%buskirk%%% How$to$do$well?$ Before$class:$read$the$corresponding$chapter$ Come$to$class$ready$to$par9cipate$in$Top$Hat$ Don t$miss$an$exam!!!!!!!!!!!!!!!!!!!!!!!!!!$ But$I m$not$good$with$science
The β-catenin homolog BAR-1 and LET-60 Ras coordinately regulate the Hox gene lin-39 during Caenorhabditis elegans vulval development
 Development 125, 3667-3680 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV5223 3667 The β-catenin homolog BAR-1 and LET-60 Ras coordinately regulate the Hox gene lin-39 during
Development 125, 3667-3680 (1998) Printed in Great Britain The Company of Biologists Limited 1998 DEV5223 3667 The β-catenin homolog BAR-1 and LET-60 Ras coordinately regulate the Hox gene lin-39 during
Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
Chapter 18 Regulation of Gene Expression Edited by Shawn Lester PowerPoint Lecture Presentations for Biology Eighth Edition Neil Campbell and Jane Reece Lectures by Chris Romero, updated by Erin Barley
2. Yeast two-hybrid system
 2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
2. Yeast two-hybrid system I. Process workflow a. Mating of haploid two-hybrid strains on YPD plates b. Replica-plating of diploids on selective plates c. Two-hydrid experiment plating on selective plates
CHAPTER 3. EGF and Wnt signaling during patterning of the C. elegans
 III-1 CHAPTER 3 EGF and Wnt signaling during patterning of the C. elegans Bγ/δ Equivalence Group Adeline Seah and Paul W. Sternberg Abstract III-2 During development, different signaling pathways interact
III-1 CHAPTER 3 EGF and Wnt signaling during patterning of the C. elegans Bγ/δ Equivalence Group Adeline Seah and Paul W. Sternberg Abstract III-2 During development, different signaling pathways interact
Predictive Modeling of Signaling Crosstalk... Model execution and model checking can be used to test a biological hypothesis
 Model execution and model checking can be used to test a biological hypothesis The model is an explanation of a biological mechanism and its agreement with experimental results is used to either validate
Model execution and model checking can be used to test a biological hypothesis The model is an explanation of a biological mechanism and its agreement with experimental results is used to either validate
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
7.06 Problem Set #4, Spring 2005
 7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
7.06 Problem Set #4, Spring 2005 1. You re doing a mutant hunt in S. cerevisiae (budding yeast), looking for temperaturesensitive mutants that are defective in the cell cycle. You discover a mutant strain
DPL-1 DP, LIN-35 Rb, and EFL-1 E2F act with the MCD-1 Zinc-finger protein to promote programmed cell death in C. elegans
 Genetics: Published Articles Ahead of Print, published on January 21, 2007 as 10.1534/genetics.106.068148 DPL-1 DP, LIN-35 Rb, and EFL-1 E2F act with the MCD-1 Zinc-finger protein to promote programmed
Genetics: Published Articles Ahead of Print, published on January 21, 2007 as 10.1534/genetics.106.068148 DPL-1 DP, LIN-35 Rb, and EFL-1 E2F act with the MCD-1 Zinc-finger protein to promote programmed
EGF Signal Propagation during C. elegans Vulval Development Mediated by ROM-1 Rhomboid
 Open access, freely available online EGF Signal Propagation during C. elegans Vulval Development Mediated by ROM-1 Rhomboid PLoS BIOLOGY Amit Dutt [, Stefano Canevascini [, Erika Froehli-Hoier, Alex Hajnal
Open access, freely available online EGF Signal Propagation during C. elegans Vulval Development Mediated by ROM-1 Rhomboid PLoS BIOLOGY Amit Dutt [, Stefano Canevascini [, Erika Froehli-Hoier, Alex Hajnal
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Caenorhabditis elegans
 Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Caenorhabditis elegans Why C. elegans? Sea urchins have told us much about embryogenesis. They are suited well for study in the lab; however, they do not tell us much about the genetics involved in embryogenesis.
Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc
 OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
OPTIMIZATION OF IMMUNOBLOT PROTOCOL 121 Optimization of Immunoblot Protocol for Use with a Yeast Strain Containing the CDC7 Gene Tagged with myc Jacqueline Bjornton and John Wheeler Faculty Sponsor: Anne
Caenorhabditis elegans Development
 MOLECULAR REPRODUCTION AND DEVELOPMENT 42523528 (1995) LET-23-Mediated Signal Transduction During Caenorhabditis elegans Development PAUL W. STERNBERG, GIOVANNI LESA, JUNHO LEE, WENDY S. KATZ, CHARLES
MOLECULAR REPRODUCTION AND DEVELOPMENT 42523528 (1995) LET-23-Mediated Signal Transduction During Caenorhabditis elegans Development PAUL W. STERNBERG, GIOVANNI LESA, JUNHO LEE, WENDY S. KATZ, CHARLES
CHAPTER 2. Wnt and EGF pathways act together to induce C. elegans. male hook development
 II-1 CHAPTER 2 Wnt and EGF pathways act together to induce C. elegans male hook development Adeline Seah *, Hui Yu *, Michael A. Herman, Edwin L. Ferguson, H. Robert Horvitz and Paul W. Sternberg *Equal
II-1 CHAPTER 2 Wnt and EGF pathways act together to induce C. elegans male hook development Adeline Seah *, Hui Yu *, Michael A. Herman, Edwin L. Ferguson, H. Robert Horvitz and Paul W. Sternberg *Equal
Conservation and diversification of Wnt signaling function during the evolution of nematode vulva development
 Conservation and diversification of Wnt signaling function during the evolution of nematode vulva development Min Zheng 1, Daniel Messerschmidt 1, Benno Jungblut 1,2 & Ralf J Sommer 1 Cell-fate specification
Conservation and diversification of Wnt signaling function during the evolution of nematode vulva development Min Zheng 1, Daniel Messerschmidt 1, Benno Jungblut 1,2 & Ralf J Sommer 1 Cell-fate specification
Prokaryotic Regulation
 Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
Prokaryotic Regulation Control of transcription initiation can be: Positive control increases transcription when activators bind DNA Negative control reduces transcription when repressors bind to DNA regulatory
The two steps of vulval induction in Oscheius tipulae CEW1 recruit common regulators including a MEK kinase
 Developmental Biology 265 (2004) 113 126 www.elsevier.com/locate/ydbio The two steps of vulval induction in Oscheius tipulae CEW1 recruit common regulators including a MEK kinase Marie-Laure Dichtel-Danjoy
Developmental Biology 265 (2004) 113 126 www.elsevier.com/locate/ydbio The two steps of vulval induction in Oscheius tipulae CEW1 recruit common regulators including a MEK kinase Marie-Laure Dichtel-Danjoy
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
Javier A. Wagmaister a,1, Ginger R. Miley b,2, Corey A. Morris c,3, Julie E. Gleason a, Leilani M. Miller c, Kerry Kornfeld b, David M.
 Developmental Biology 297 (2006) 550 565 www.elsevier.com/locate/ydbio Genomes & Developmental Control Identification of cis-regulatory elements from the C. elegans Hox gene lin-39 required for embryonic
Developmental Biology 297 (2006) 550 565 www.elsevier.com/locate/ydbio Genomes & Developmental Control Identification of cis-regulatory elements from the C. elegans Hox gene lin-39 required for embryonic
CHARACTERIZATION OF let-765/nsh-1 AND ITS ROLE IN RAS SIGNALLING IN Caenorhabditis elegans
 CHARACTERIZATION OF let-765/nsh-1 AND ITS ROLE IN RAS SIGNALLING IN Caenorhabditis elegans by Carrie Leanne Simms MSc, Simon Fraser University 2004 BSc, Simon Fraser University 1999 THESIS SUBMITTED IN
CHARACTERIZATION OF let-765/nsh-1 AND ITS ROLE IN RAS SIGNALLING IN Caenorhabditis elegans by Carrie Leanne Simms MSc, Simon Fraser University 2004 BSc, Simon Fraser University 1999 THESIS SUBMITTED IN
Regulation of Transcription in Eukaryotes
 Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
Regulation of Transcription in Eukaryotes Leucine zipper and helix-loop-helix proteins contain DNA-binding domains formed by dimerization of two polypeptide chains. Different members of each family can
PROGRAMMED cell death is important for many
 Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.106.068148 DPL-1 DP, LIN-35 Rb and EFL-1 E2F Act With the MCD-1 Zinc-Finger Protein to Promote Programmed Cell Death in Caenorhabditis
Copyright Ó 2007 by the Genetics Society of America DOI: 10.1534/genetics.106.068148 DPL-1 DP, LIN-35 Rb and EFL-1 E2F Act With the MCD-1 Zinc-Finger Protein to Promote Programmed Cell Death in Caenorhabditis
Mutations in cye-1, a Caenorhabditis elegans cyclin E homolog, reveal coordination between cell-cycle control and vulval development
 Development 127, 4049-4060 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV8754 4049 Mutations in cye-1, a Caenorhabditis elegans cyclin E homolog, reveal coordination between
Development 127, 4049-4060 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV8754 4049 Mutations in cye-1, a Caenorhabditis elegans cyclin E homolog, reveal coordination between
16 CONTROL OF GENE EXPRESSION
 16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
16 CONTROL OF GENE EXPRESSION Chapter Outline 16.1 REGULATION OF GENE EXPRESSION IN PROKARYOTES The operon is the unit of transcription in prokaryotes The lac operon for lactose metabolism is transcribed
SUPPLEMENTARY MATERIAL
 1 SUPPLEMENTARY MATERIAL TABLE OF CONTENTS: Overview of VPC fate specification Biological sources of information for the Core Model Model structure: Objects GUI objects Internal objects Allele representation
1 SUPPLEMENTARY MATERIAL TABLE OF CONTENTS: Overview of VPC fate specification Biological sources of information for the Core Model Model structure: Objects GUI objects Internal objects Allele representation
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Afundamental issue in developmental biology is how
 Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.056465 Identification and Classification of Genes That Act Antagonistically to let-60 Ras Signaling in Caenorhabditis elegans
Copyright Ó 2006 by the Genetics Society of America DOI: 10.1534/genetics.106.056465 Identification and Classification of Genes That Act Antagonistically to let-60 Ras Signaling in Caenorhabditis elegans
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
 Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Supplementary Information to Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition Nadine Czudnochowski 1,2, *, Christian A. Bösken 1, * & Matthias Geyer 1 1 Max-Planck-Institut
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
BIS &003 Answers to Assigned Problems May 23, Week /18.6 How would you distinguish between an enhancer and a promoter?
 Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
Week 9 Study Questions from the textbook: 6 th Edition: Chapter 19-19.6, 19.7, 19.15, 19.17 OR 7 th Edition: Chapter 18-18.6 18.7, 18.15, 18.17 19.6/18.6 How would you distinguish between an enhancer and
The C. elegans Mi-2 chromatin-remodelling proteins function in vulval cell fate determination
 Development 127, 5277-5284 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV2639 5277 The C. elegans Mi-2 chromatin-remodelling proteins function in vulval cell fate determination
Development 127, 5277-5284 (2000) Printed in Great Britain The Company of Biologists Limited 2000 DEV2639 5277 The C. elegans Mi-2 chromatin-remodelling proteins function in vulval cell fate determination
Ribosome readthrough
 Ribosome readthrough Starting from the base PROTEIN SYNTHESIS Eukaryotic translation can be divided into four stages: Initiation, Elongation, Termination and Recycling During translation, the ribosome
Ribosome readthrough Starting from the base PROTEIN SYNTHESIS Eukaryotic translation can be divided into four stages: Initiation, Elongation, Termination and Recycling During translation, the ribosome
Why Flies? stages of embryogenesis. The Fly in History
 The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
The Fly in History 1859 Darwin 1866 Mendel c. 1890 Driesch, Roux (experimental embryology) 1900 rediscovery of Mendel (birth of genetics) 1910 first mutant (white) (Morgan) 1913 first genetic map (Sturtevant
Activation of a receptor. Assembly of the complex
 Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
Activation of a receptor ligand inactive, monomeric active, dimeric When activated by growth factor binding, the growth factor receptor tyrosine kinase phosphorylates the neighboring receptor. Assembly
CHAPTER : Prokaryotic Genetics
 CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
CHAPTER 13.3 13.5: Prokaryotic Genetics 1. Most bacteria are not pathogenic. Identify several important roles they play in the ecosystem and human culture. 2. How do variations arise in bacteria considering
Follow this and additional works at:
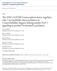 Washington University School of Medicine Digital Commons@Becker Open Access Publications 2014 The DAF-16 FOXO transcription factor regulates natc-1 to modulate stress resistance in Caenorhabditis elegans,
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2014 The DAF-16 FOXO transcription factor regulates natc-1 to modulate stress resistance in Caenorhabditis elegans,
Controlling Gene Expression
 Controlling Gene Expression Control Mechanisms Gene regulation involves turning on or off specific genes as required by the cell Determine when to make more proteins and when to stop making more Housekeeping
Controlling Gene Expression Control Mechanisms Gene regulation involves turning on or off specific genes as required by the cell Determine when to make more proteins and when to stop making more Housekeeping
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
32 Gene regulation, continued Lecture Outline 11/21/05
 32 Gene regulation, continued Lecture Outline 11/21/05 Review the operon concept Repressible operons (e.g. trp) Inducible operons (e.g. lac) Positive regulation of lac () Practice applying the operon concept
32 Gene regulation, continued Lecture Outline 11/21/05 Review the operon concept Repressible operons (e.g. trp) Inducible operons (e.g. lac) Positive regulation of lac () Practice applying the operon concept
Zinc Ions and Cation Diffusion Facilitator Proteins Regulate Ras-Mediated Signaling
 Developmental Cell, Vol. 2, 567 578, May, 2002, Copyright 2002 by Cell Press Zinc Ions and Cation Diffusion Facilitator Proteins Regulate Ras-Mediated Signaling Janelle J. Bruinsma, 1 Tanawat Jirakulaporn,
Developmental Cell, Vol. 2, 567 578, May, 2002, Copyright 2002 by Cell Press Zinc Ions and Cation Diffusion Facilitator Proteins Regulate Ras-Mediated Signaling Janelle J. Bruinsma, 1 Tanawat Jirakulaporn,
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Principles of Genetics
 Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Principles of Genetics Snustad, D ISBN-13: 9780470903599 Table of Contents C H A P T E R 1 The Science of Genetics 1 An Invitation 2 Three Great Milestones in Genetics 2 DNA as the Genetic Material 6 Genetics
Upstream Elements Regulating mir-241 and mir-48 Abstract Introduction
 Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
Upstream Elements Regulating mir-241 and mir-48 Hanna Vollbrecht, Tamar Resnick, and Ann Rougvie University of Minnesota: Twin Cities Undergraduate Research Scholarship 2012-2013 Abstract Caenorhabditis
Hypodermis to Oppose Ras-Mediated Vulval Induction in C. elegans
 Developmental Cell, Vol. 8, 117 123, January, 2005, Copyright 2005 by Elsevier Inc. DOI 10.1016/j.devcel.2004.11.015 lin-35 Rb Acts in the Major Hypodermis to Oppose Ras-Mediated Vulval Induction in C.
Developmental Cell, Vol. 8, 117 123, January, 2005, Copyright 2005 by Elsevier Inc. DOI 10.1016/j.devcel.2004.11.015 lin-35 Rb Acts in the Major Hypodermis to Oppose Ras-Mediated Vulval Induction in C.
Chapter 16 Lecture. Concepts Of Genetics. Tenth Edition. Regulation of Gene Expression in Prokaryotes
 Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
Chapter 16 Lecture Concepts Of Genetics Tenth Edition Regulation of Gene Expression in Prokaryotes Chapter Contents 16.1 Prokaryotes Regulate Gene Expression in Response to Environmental Conditions 16.2
Welcome to Class 21!
 Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Welcome to Class 21! Introductory Biochemistry! Lecture 21: Outline and Objectives l Regulation of Gene Expression in Prokaryotes! l transcriptional regulation! l principles! l lac operon! l trp attenuation!
Regulation of gene expression. Premedical - Biology
 Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Regulation of gene expression Premedical - Biology Regulation of gene expression in prokaryotic cell Operon units system of negative feedback positive and negative regulation in eukaryotic cell - at any
Gene Control Mechanisms at Transcription and Translation Levels
 Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Gene Control Mechanisms at Transcription and Translation Levels Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by MHRD Page 1 of 9
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam
 Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
Chapter 10, 11, 14: Gene Expression, Regulation, and Development Exam Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme
OSM-11 Facilitates LIN-12 Notch Signaling during Caenorhabditis elegans Vulval Development
 OSM-11 Facilitates LIN-12 Notch Signaling during Caenorhabditis elegans Vulval Development Hidetoshi Komatsu 1,2[ a, Michael Y. Chao 3[, Jonah Larkins-Ford 1 b, Mark E. Corkins 1, Gerard A. Somers 1, Tim
OSM-11 Facilitates LIN-12 Notch Signaling during Caenorhabditis elegans Vulval Development Hidetoshi Komatsu 1,2[ a, Michael Y. Chao 3[, Jonah Larkins-Ford 1 b, Mark E. Corkins 1, Gerard A. Somers 1, Tim
purpose of this Chapter is to highlight some problems that will likely provide new
 119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
119 Chapter 6 Future Directions Besides our contributions discussed in previous chapters to the problem of developmental pattern formation, this work has also brought new questions that remain unanswered.
Lesson Overview. Gene Regulation and Expression. Lesson Overview Gene Regulation and Expression
 13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
13.4 Gene Regulation and Expression THINK ABOUT IT Think of a library filled with how-to books. Would you ever need to use all of those books at the same time? Of course not. Now picture a tiny bacterium
Abl Kinase Inhibits the Engulfment of Apoptotic [corrected] Cells in Caenorhabditis elegans
![Abl Kinase Inhibits the Engulfment of Apoptotic [corrected] Cells in Caenorhabditis elegans Abl Kinase Inhibits the Engulfment of Apoptotic [corrected] Cells in Caenorhabditis elegans](/thumbs/74/71020578.jpg) Abl Kinase Inhibits the Engulfment of Apoptotic [corrected] Cells in Caenorhabditis elegans The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story
Abl Kinase Inhibits the Engulfment of Apoptotic [corrected] Cells in Caenorhabditis elegans The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story
Ch 10, 11 &14 Preview
 Ch 10, 11 &14 Preview Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme hypothesis have to be modified? a. Some
Ch 10, 11 &14 Preview Multiple Choice Identify the choice that best completes the statement or answers the question. 1. Why did the original one-gene, one-enzyme hypothesis have to be modified? a. Some
COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION
 COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION I. Aksan 1, M. Sen 2, M. K. Araz 3, and M. L. Kurnaz 3 1 School of Biological Sciences, University of Manchester,
COMPUTER SIMULATION OF DIFFERENTIAL KINETICS OF MAPK ACTIVATION UPON EGF RECEPTOR OVEREXPRESSION I. Aksan 1, M. Sen 2, M. K. Araz 3, and M. L. Kurnaz 3 1 School of Biological Sciences, University of Manchester,
Cell Death & Trophic Factors II. Steven McLoon Department of Neuroscience University of Minnesota
 Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Cell Death & Trophic Factors II Steven McLoon Department of Neuroscience University of Minnesota 1 Remember? Neurotrophins are cell survival factors that neurons get from their target cells! There is a
Developmental Biology
 Developmental Biology 327 (2009) 419 432 Contents lists available at ScienceDirect Developmental Biology jour nal homepage: www. elsevier. com/ developmentalbiology Wnt and EGF pathways act together to
Developmental Biology 327 (2009) 419 432 Contents lists available at ScienceDirect Developmental Biology jour nal homepage: www. elsevier. com/ developmentalbiology Wnt and EGF pathways act together to
The geneticist s questions
 The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
The geneticist s questions a) What is consequence of reduced gene function? 1) gene knockout (deletion, RNAi) b) What is the consequence of increased gene function? 2) gene overexpression c) What does
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
The Eukaryotic Genome and Its Expression. The Eukaryotic Genome and Its Expression. A. The Eukaryotic Genome. Lecture Series 11
 The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
The Eukaryotic Genome and Its Expression Lecture Series 11 The Eukaryotic Genome and Its Expression A. The Eukaryotic Genome B. Repetitive Sequences (rem: teleomeres) C. The Structures of Protein-Coding
Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence
 Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence www.plantcell.org/cgi/doi/10.1105/tpc.110.tt0110 Epigenetics Usually
Epigenetics and Flowering Any potentially stable and heritable change in gene expression that occurs without a change in DNA sequence www.plantcell.org/cgi/doi/10.1105/tpc.110.tt0110 Epigenetics Usually
MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition in C. Elegans
 Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition
Marquette University e-publications@marquette Master's Theses (2009 -) Dissertations, Theses, and Professional Projects MIR-237 is Likely a Developmental Timing Gene that Regulates the L2-to-L3 Transition
Name: SBI 4U. Gene Expression Quiz. Overall Expectation:
 Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
Gene Expression Quiz Overall Expectation: - Demonstrate an understanding of concepts related to molecular genetics, and how genetic modification is applied in industry and agriculture Specific Expectation(s):
1. In most cases, genes code for and it is that
 Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Name Chapter 10 Reading Guide From DNA to Protein: Gene Expression Concept 10.1 Genetics Shows That Genes Code for Proteins 1. In most cases, genes code for and it is that determine. 2. Describe what Garrod
Gene regulation I Biochemistry 302. Bob Kelm February 25, 2005
 Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Gene regulation I Biochemistry 302 Bob Kelm February 25, 2005 Principles of gene regulation (cellular versus molecular level) Extracellular signals Chemical (e.g. hormones, growth factors) Environmental
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology. Biology. Slide 1 of 26. End Show. Copyright Pearson Prentice Hall
 Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Biology Biology 1 of 26 Fruit fly chromosome 12-5 Gene Regulation Mouse chromosomes Fruit fly embryo Mouse embryo Adult fruit fly Adult mouse 2 of 26 Gene Regulation: An Example Gene Regulation: An Example
Chapter 12. Genes: Expression and Regulation
 Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Chapter 12 Genes: Expression and Regulation 1 DNA Transcription or RNA Synthesis produces three types of RNA trna carries amino acids during protein synthesis rrna component of ribosomes mrna directs protein
Molecular Biology (9)
 Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
Molecular Biology (9) Translation Mamoun Ahram, PhD Second semester, 2017-2018 1 Resources This lecture Cooper, Ch. 8 (297-319) 2 General information Protein synthesis involves interactions between three
The Worm, Ceanorhabditis elegans
 1 1 Institute of Biology University of Iceland October, 2005 Lecture outline The problem of phenotype Dear Max Sidney Brenner A Nobel Prize in Medicine Genome sequence Some tools Gene structure Genomic
1 1 Institute of Biology University of Iceland October, 2005 Lecture outline The problem of phenotype Dear Max Sidney Brenner A Nobel Prize in Medicine Genome sequence Some tools Gene structure Genomic
Lecture 10: Cyclins, cyclin kinases and cell division
 Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem*3560 Lecture 10: Cyclins, cyclin kinases and cell division The eukaryotic cell cycle Actively growing mammalian cells divide roughly every 24 hours, and follow a precise sequence of events know as
Chem Lecture 10 Signal Transduction
 Chem 452 - Lecture 10 Signal Transduction 111202 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
Chem 452 - Lecture 10 Signal Transduction 111202 Here we look at the movement of a signal from the outside of a cell to its inside, where it elicits changes within the cell. These changes are usually mediated
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The geneticist s questions. Deleting yeast genes. Functional genomics. From Wikipedia, the free encyclopedia
 From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
From Wikipedia, the free encyclopedia Functional genomics..is a field of molecular biology that attempts to make use of the vast wealth of data produced by genomic projects (such as genome sequencing projects)
GENE REGULATION AND PROBLEMS OF DEVELOPMENT
 GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
GENE REGULATION AND PROBLEMS OF DEVELOPMENT By Surinder Kaur DIET Ropar Surinder_1998@ yahoo.in Mob No 9988530775 GENE REGULATION Gene is a segment of DNA that codes for a unit of function (polypeptide,
Chapter 4 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays.
 Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Evaluating a potential interaction between deltex and git in Drosophila: genetic interaction, gene overexpression and cell biology assays. The data described in chapter 3 presented evidence that endogenous
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
RECEPTOR tyrosine kinase (RTK)-Ras-extracellu- ricks 2000), which then cooperate with or antagonize
 Copyright 2002 by the Genetics Society of America A lin-45 raf Enhancer Screen Identifies eor-1, eor-2 and Unusual Alleles of Ras Pathway Genes in Caenorhabditis elegans Christian E. Rocheleau, 1 Robyn
Copyright 2002 by the Genetics Society of America A lin-45 raf Enhancer Screen Identifies eor-1, eor-2 and Unusual Alleles of Ras Pathway Genes in Caenorhabditis elegans Christian E. Rocheleau, 1 Robyn
Stochastic simulations
 Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
Stochastic simulations Application to molecular networks Literature overview Noise in genetic networks Origins How to measure and distinguish between the two types of noise (intrinsic vs extrinsic)? What
