Molecular Phylogenetics and Evolution
|
|
|
- Ashlie Kelly
- 6 years ago
- Views:
Transcription
1 Molecular Phylogenetics and Evolution xxx (2009) xxx xxx Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: Short Communication Phylogenetic relationships of Sardinian cave salamanders, genus Hydromantes, based on mitochondrial and nuclear DNA sequence data Arie van der Meijden a, *, Ylenia Chiari b,1, Mauro Mucedda c, Salvador Carranza d, Claudia Corti e, Michael Veith a a Department of Biogeography, University of Trier, Am Wissenschaftspark 25-27, Trier, Germany b Department of Ecology and Evolutionary Biology, YIBS-Molecular Systematics and Conservation Genetics Lab, ESC 158, Yale University, 21 Sachem St., New Haven, CT , USA c Via Leopardi 1, Sassari, Gruppo Speleologico Sassarese, Italy d Institute of Evolutionary Biology (UPF-CSIC), Passeig Maritim de la Barceloneta 37-49, E Barcelona, Spain e Museo di Storia Naturale dell Università di Firenze, Sezione di Zoologia La Specola, Via Romana, 17, Firenze, Italy article info Article history: Received 5 September 2008 Revised 12 November 2008 Accepted 10 December 2008 Available online xxxx Ó 2009 Elsevier Inc. All rights reserved. 1. Introduction The Tyrrhenian island of Sardinia is known for its high level of endemism (Oosterbroek and Arntzen, 1992; Grill et al., 2007). Beside the importance of this island as a refuge during the last glaciations, little is known about the origin and relationships of Sardinian species (reviewed in Grill et al., 2007). The origin of some Sardinian lineages is attributed to the isolation of the Sardo-Corsican plate (e.g. Lanza, 1983; Caccone et al., 1994; Oliverio et al., 2000; Omodeo and Rota, 2008), others to the Messinian salinity crisis (e.g. Marra, 2004) or to more recent events such as the severe climate changes from the early Pliocene and throughout the Pleistocene (Grill et al., 2007 and references therein). It still remains unclear which of these events are more relevant to explain the current distribution of the European members of the genus Hydromantes. Despite their common name, Hydromantes are not restricted to cave habitats, or even to limestone substrates, but are also found on exposed sites, even on granites and volcanic rocks (Lanza et al., 2005). Their range is therefore not directly restricted by these aspects of geology or even by plant cover. Humid subterranean hiding places, such as the space between the rocks of a rockslide, do seem to be correlated with their presence (personal observation). The salamanders of the genus Hydromantes have their highest diversity in Sardinia, being represented * Corresponding author. Fax: +49 (0) addresses: frog@arievandermeijden.nl (A. van der Meijden), yle@yleniachiari.it (Y. Chiari), mucedda@uniss.it (M. Mucedda), salvador.carranza@ibe.upfcsic.es (S. Carranza), claudia.corti@unifi.it (C. Corti), veith@uni-trier.de (M. Veith). 1 Present address: Institut des Sciences de l Evolution, Université Montpellier 2, Place Eugène Bataillon, Bat 22/RdC/CC064, Montpellier Cedex 5, France. by five species, versus only three species in mainland Europe, and three in the western USA. Recently, Vieites et al. (2007) have brought forward a compelling hypothesis for the highly disjunct distribution of the members of the genus Hydromantes between the western US and Europe by suggesting dispersal through the Bering land bridge. The eight European species are distributed in southeastern France, north and central Italy and throughout Sardinia. Previous phylogenetic works based on immunological data and allozymes (Wake et al., 1978), allozymes (Nascetti et al., 1996), morphology (Lanza and Caputo, 1995), morphology, caryology and cytochrome b sequences (Jackman et al., 1997) and cytochrome b, 12S rrna and 16S rrna sequences (Carranza et al., 2007) did not fully resolve the relationships among the European species of Hydromantes. Some of these studies (e.g., Nascetti et al., 1996; Nardi, 1991; but see also Carranza et al., 2007) support the hypothesis of H. (Atylodes) genei, which inhabits the southwest of Sardinia, as the most basal European species. The eastern Sardinian species (H. (Speleomantes) flavus, H. (S.) supramontis, H. (S.) imperialis and H. (S.) sarrabusensis) are thought to be more closely related to the mainland species (H. (S.) strinatii, H. (S.) ambrosii and H. (S.) italicus). Several biogeographic hypotheses have been put forward to explain this pattern (reviewed in Lanza et al., 2005). All these biogeographic hypotheses hinge on the basal position of H. (A.) genei, which is generally considered to have split off from the ancestral stock well before the other species. The recent publication of Carranza et al. (2007) addresses the phylogenetic relationships between the European species based on three mitochondrial genes, but could not resolve H. (A.) genei unambiguously as the most basal taxon, or the sister group relationship between the mainland species and the eastern Sardinian species. Also the placement of H. (S.) sarrabusensis could not be /$ - see front matter Ó 2009 Elsevier Inc. All rights reserved. doi: /j.ympev
2 2 A. van der Meijden et al. / Molecular Phylogenetics and Evolution xxx (2009) xxx xxx unambiguously resolved. Increased taxon sampling and enlargement of the dataset have been shown to increase phylogenetic resolution (Pollock et al., 2002; Brinkmann and Philippe, 2008). Therefore, in an effort to resolve these relationships, in the current study we extended the existing dataset considerably by adding two nuclear genes and additional mitochondrial sequence data, as well as greatly extend the number of Sardinian populations represented in the dataset from eight to 16 populations. Our data confirm the previous findings by Carranza et al. (2007), but despite the larger dataset, several relationships among the European members of the genus Hydromantes still remain poorly resolved. 2. Materials and methods 2.1. Specimen selection A total of 66 European Hydromantes specimens were used in this study. Of these 33 samples are the same as used in the study by Carranza et al. (2007), while the other 33 samples were collected in Sardinia in April 2007 (Table 1). We included a total of 27 populations. Sardinian species were represented by samples from across the range of each species from a total of 16 populations (Fig. 1). A maximum of three specimens from every sampling of each population were randomly selected to be included in this study (Table 1). Samples taken from sites at very close proximity were counted as a single population DNA extraction, PCR and sequencing DNA was extracted from tail tips preserved in 99% ethanol using a DNeasy Ò blood and tissue kit (Quiagen). Primers for one fragment of the 12S rrna gene and one fragment of the 16S rrna gene were 12SA-L and 12SB-H and 16SA-L and 16SB-H, used as in Palumbi et al. (1991). Primers for cytochrome b were designed based on an alignment of Hydromantes sequences available in GenBank. These were Cyt-BYf1 (5 0 -CAARTCCTYACCGGRYTATTT-3 0 ) as the forward primer and Cyt-BYr1 (5 0 -TTCGRTKTTTTGADGTRTTTA-3 0 ) as the reverse primer. This fragment was amplified using an initial denaturation step of 2 min at 94 C, followed by 35 cycles of denaturation at 94 C for 20 s, annealing at 52 C for 30 s, and extension at 72 C for 90 s. Two nested pairs of primers to amplify a single 1400 bp fragment of Rag-1 were designed based on an alignment of available salamander Rag-1 sequences from GenBank. The first PCR with forward primer Rag-1Yf1 (5 0 -CAGATTTTCCAGCCYTTACAYGC-3 0 ) and reverse Rag-1Yr1 (5 0 -CCATTTCATAAAACTTGGACTGC-3 0 ). The second PCR with forward primer Rag-1Yf2 (5 0 -CTWCCAGGMTAYCAYCYVTTYG- 3 0 ) and reverse primer Rag-1Yr2 (5 0 -CGGAAACGTCTGAACAGYTTC- 3 0 ). PCR conditions for these Rag-1 primers and the BDNF primers (Van der Meijden et al., 2007) were those as described for Rag-1 and Rag-2 in Van der Meijden et al. (2005). PCR was performed in 25 ll reactions using REDTaq Polymerase Readymix (Sigma, Taufkirchen, Germany). PCR products were purified via spin columns (Qiagen). Sequencing was performed directly using the corresponding PCR primers on an ABI3730XL sequencer. New sequences were combined with existing sequences taken from GenBank in the final dataset and additional five closely related plethodontid species and the salamandrid Salamandra were used as nested outgroups for the phylogenetic analyses (Table 1). New sequences were deposited in GenBank (accession numbers in Table 1) Phylogenetic analysis Chromatograms were checked by eye using FinchTV 1.4 (Geospiza.com) and the sequences were subsequently aligned using the Muscle alignment program version 3.6 (Edgar, 2004) using the default settings (best accuracy). The resulting alignments were checked by eye, but were not found to require additional editing. The final aligned dataset was deposited in the TreeBase database (Study accession number S2274). Missing genes were coded as missing (?) in the concatenated dataset (see Table 1). In Bayesian Inference (BI) and Maximum Parsimony (MP) analyses of large multi-gene data sets of hylid frogs, no relationship was found between completeness of the sequence data of a taxon, and the support values the taxon receives (Wiens et al., 2005), suggesting that the limited amount of missing data in our concatenated alignment is unlikely to distort the phylogenetic results. Furthermore, the placement of taxa with missing data in our analysis was congruent for well supported branches (bootstrap support >80%) between the single gene analyses in which the taxon was represented by a complete sequence, and the combined dataset, suggesting that omission of one or more genes from the combined set did not influence the placement of particular taxa (data not shown). A homogeneity partition test (Farris et al., 1994) as implemented in PAUP version 4.0b10 (Swofford, 2002) did not reject homogeneity of the five different markers (P = 0.17). Besides an analysis of the combined data set we also performed independent analyses for each gene and for the combined mitochondrial and the combined nuclear genes in the dataset. Phylogeny reconstruction based on the separate and combined datasets was performed using Maximum Likelihood (ML) and BI methods. The best fitting models of sequence evolution were determined by the AIC criterion in Modeltest 3.7 (Posada and Crandall, 1998). ML tree searches were performed using PhyML, version (Guindon and Gascuel, 2003). Bootstrap branch support values were calculated with 500 replicates for the combined datasets, and 200 replicates in the single gene analyses. The BI analyses of the combined and separate datasets were conducted with MrBayes (Huelsenbeck and Ronquist, 2001), using models estimated with Modeltest under the AIC criterion, with 2,500,000 generations, sampling trees every 100th generation (and calculating a consensus tree after omitting the first 6250 trees). Log likelihood scores for the remaining trees were graphed in Tracer 1.4 ( and checked for appropriateness of the burnin-period. The topologies resulting from the ML and BI analyses of the combined dataset were compared using a Shimodaira Hasegawa (SH) test as implemented in PAUP * (Swofford, 2002), with RELL optimization and bootstrap replicates. Between species genetic distances (uncorrected p-distances, see Table 2) were calculated based on the cytochrome b alignment using MEGA version 4 (Tamura et al., 2007). Tables of genetic distances based on the other genes are available from the authors on request. 3. Results and discussion The concatenated dataset contained a total of 3494 base pairs (375 bp of 12S rrna, 449 bp of 16S rrna, 755 bp of cytochrome b, 1230 bp of RAG-1 and 685 bp of BDNF). PCR amplification of some specimens failed for some genes despite several retries (see Table 1). The results of both the ML and BI analyses of the complete dataset as well as a separate mitochondrial and a nuclear dataset yielded similar groupings where branch support was high. Although not all support values were high, both the mitochondrial and nuclear dataset analyses showed all five Sardinian species as monophyletic (Fig. 2) Position of H. (A.) genei The basal position of H. (A.) genei is considered well supported based on, among other things, allozyme data (Wake et al., 1978;
3 A. van der Meijden et al. / Molecular Phylogenetics and Evolution xxx (2009) xxx xxx 3 Table 1 Specimens used in this study. Field numbers from Carranza et al. (2007) are listed for reference. Missing sequences are indicated by missing. H. = Hydromantes. A. = Atylodes. S. = Speleomantes. The H. (A.) genei A and B refer to the two subclades of that species, as shown in Fig. 2. Species name Specimen number Locality Carranza et al. (2007) 12S rrna 16S rrna Cytb Rag-1 BDNF H. (A.) genei B 1 Monte Tasua, Carbonia, Sardinia (Italy) E FJ missing FJ FJ FJ H. (A.) genei B 2 Monte Tasua, Carbonia, Sardinia (Italy) E FJ missing FJ missing missing H. (A.) genei B 3 Monte Tasua, Carbonia, Sardinia (Italy) E FJ missing FJ FJ FJ H. (A.) genei A 4 Grotta su Mannau, Fluminimaggiore, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (A.) genei A 5 Sanita, Fluminimaggiore, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (A.) genei A 6 Sanita, Fluminimaggiore, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (A.) genei A 7 Mine Terras Nieddas, Fluminimaggiore, Sardinia (Italy) FJ FJ FJ FJ FJ H. (A.) genei A 8 Mine Terras Nieddas, Fluminimaggiore, Sardinia (Italy) FJ FJ FJ FJ FJ H. (A.) genei A 9 Mine Terras Nieddas, Fluminimaggiore, Sardinia (Italy) FJ FJ missing FJ FJ H. (A.) genei A 10 Mine Su Corovau, Domusnovas, Sardinia (Italy) FJ FJ FJ FJ FJ H. (A.) genei A 11 Mine Su Corovau, Domusnovas, Sardinia (Italy) FJ FJ FJ FJ FJ H. (A.) genei A 12 Mine Su Corovau, Domusnovas, Sardinia (Italy) FJ FJ missing FJ FJ H. (A.) genei B 13 Grotta Cava Romana, Nuxis, Sardinia (Italy) FJ FJ FJ FJ FJ H. (A.) genei B 14 Grotta dei geotritoni, Nuxis, Sardinia (Italy) FJ FJ missing FJ missing H. (A.) genei B 15 Grotta dei geotritoni, Nuxis, Sardinia (Italy) FJ FJ FJ FJ FJ H. (A.) genei B 16 Grotta dei geotritoni, Nuxis, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) flavus 17 Grotta di Nurai, Lula, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) flavus 18 Grotta di Nurai, Lula, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) flavus 19 Grotta di Nurai, Lula, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) flavus 20 Conca e Crapa, Lula, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (S.) flavus 21 Conca e Crapa, Lula, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (S.) flavus 22 Conca e Crapa, Lula, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (S.) imperialis 23 Lago Omodeo, Ula Tirso, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 24 Lago Omodeo, Ula Tirso, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 25 Grotta Sa Turru, S. Nicolò Gerrei, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 26 Grotta Sa Turru, S. Nicolò Gerrei, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 27 Grotta Sa Turru, S. Nicolò Gerrei, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 28 Grotta Sa Rutta e Linus, Perdasdefogu, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 29 Grotta Sa Rutta e Linus, Perdasdefugu, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 30 Grotta Sa Rutta e Linus, Perdasdefugu, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 31 Grotta de Is Lianas, Ulassai, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 32 Grotta de Is Lianas, Ulassai, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 33 Grotta de Is Lianas, Ulassai, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 34 Grotta Orroli, Osini, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 35 Grotta Orroli, Osini, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) imperialis 36 Grotta Orroli, Osini, Sardinia (Italy) FJ FJ FJ missing FJ H. (S.) imperialis 37 Ulassai, Sardinia (Italy) E FJ FJ FJ missing missing H. (S.) imperialis 38 Ulassai, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (S.) imperialis 39 Ulassai, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (S.) sarrabusensis 40 Mount Sette Fratelli, Burcei, Cagliari, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) sarrabusensis 41 Mount Sette Fratelli, Burcei, Cagliari, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) sarrabusensis 42 Mount Sette Fratelli, Burcei, Cagliari, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) supramontis 43 Grotta Su Bentu, Oliena, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) supramontis 44 Grotta Su Bentu, Oliena, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) supramontis 45 Grotta Su Bentu, Oliena, Sardinia (Italy) FJ FJ FJ FJ FJ H. (S.) supramontis 46 Punta Cuccuttos, Urzulei, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) supramontis 47 Punta Cuccuttos, Urzulei, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) supramontis 48 Punta Cuccuttos, Urzulei, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) supramontis 49 Grotta su Bentu, Oliena, Sardinia (Italy) E FJ missing FJ FJ FJ H. (S.) supramontis 50 Grotta su Bentu, Oliena, Sardinia (Italy) E FJ FJ FJ FJ FJ H. (S.) supramontis 51 Grotta su Bentu, Oliena, Sardinia (Italy) E FJ FJ FJ missing FJ H. (S.) ambrosii 52 Grotta Fornace, Pignone, La Spezia, Liguria (Italy) E EU FJ FJ FJ FJ ambrosii H. (S.) ambrosii 53 Grotta Fornace, Pignone, La Spezia, Liguria (Italy) E EU FJ FJ FJ FJ ambrosii H. (S.) ambrosii 54 Apuan Alps, Massa-Carrara, Tuscany (Italy) E EU FJ FJ FJ FJ bianchii H. (S.) ambrosii 55 Apuan Alps, Massa-Carrara, Tuscany (Italy) E EU FJ FJ FJ FJ bianchii H. (S.) italicus 56 Sorgenti dell Esino, Esanatoglia, Ancona, Marche (Italy) E EU FJ FJ FJ FJ H. (S.) italicus 57 Abrutti, Farindola, Pescara (Italy) E EU FJ FJ FJ FJ H. (S.) italicus 58 Monte Tezio, Perugia, Umbria (Italy) E EU FJ FJ missing FJ H. (S.) italicus 59 Buca delle Fate (Coreglia Antelminelli), Lucca, Tuscany FJ FJ FJ FJ FJ (Italy) H. (S.) italicus 60 Buca delle Fate (Coreglia Antelminelli), Lucca, Tuscany FJ FJ FJ FJ FJ (Italy) H. (S.) italicus 61 Tana Buca di Maggiano (Maggiano), Lucca, Tuscany (Italy) FJ FJ FJ FJ FJ H. (S.) strinatii 62 W. Monte Groppi, Genova, Liguria (Italy) E EU EU FJ FJ FJ H. (S.) strinatii 63 Rio Tonno, Vallbrevenna, Genova, Liguria (Italy) E EU FJ FJ missing missing H. (S.) strinatii 64 S. Bartolomeo, Savignone, Genova, Liguria (Italy) E EU missing FJ missing missing H. (S.) strinatii 65 Maritime Alps (France) E EU missing FJ FJ FJ (continued on next page)
4 E8 15' E9 15' E9 45' ARTICLE IN PRESS 4 A. van der Meijden et al. / Molecular Phylogenetics and Evolution xxx (2009) xxx xxx Table 1 (continued) Species name Specimen number Locality Carranza et al. (2007) 12S rrna 16S rrna Cytb Rag-1 BDNF H. (S.) strinatii 66 Maritime Alps (France) E EU FJ FJ FJ FJ H. shastae missing missing missing EU EU H. brunus AY NC AY EU EU H. platycephalus DQ DQ missing EU EU Karsenia koreana missing missing missing AY EU Ensatina eschscholtzii AY EF AY EU EU Salamandra salamandra DQ AY AY AY EF km Sassari & Lago Omodeo Oristano Iglesias 1-3 Carbonia Cagliari Lanusei N40 45' N40 15' N39 45' N39 15' Fig. 1. Sampling localities for all Sardinian species. Numbers correspond to specimen numbers in Table , H. (A.) genei; 17 22, H. (S.) flavus; 23 39, H. (S.) imperialis; 40 42, H. (S.) sarrabusensis; 43 51, H. (S.) supramontis. Nascetti et al., 1996), the lack of differentiated sex chromosomes (Nardi, 1991) and parasites shared by eastern Sardinian species and mainland species, but not by H. (A.) genei (Lanza and Caputo, 1995). As in Jackman et al. (1997) and Carranza et al. (2007), the basal position of H. (A.) genei could not be unambiguously corroborated in our study despite a much larger dataset, and two additional populations. The BI tree showed H. (A.) genei as basal to all other European taxa, whereas the ML tree showed a sister group relationship between H. (A.) genei and the mainland species. Neither of these placements were highly supported. A SH test based on the combined dataset showed that neither of the two alternative placements of H. (A.) genei could be rejected (p = 0.43). In the single gene analyses, only Rag-1 highly supported the sister group relationship between the mainland species and the east Sardinian species to the exclusion of H. (A.) genei (93% bootstrap support, data not shown). As support for more derived as well as more basal nodes in the phylogeny based on the combined dataset are well resolved (see Fig. 2), it is possible that the lack of support basally in the clade of Hydromantes (Speleomantes) may have been the result of a relatively rapid succession of divergences during the cladogenesis of the European species of Hydromantes Divergence within H. (A.) genei Our data corroborate two well supported groups within H. (A.) genei, as first resolved based on allozymes by Nascetti et al. (1996). The genetic distance between these two groups is similar to the lowest distances between full species based on the cytochrome b sequences available. The p-distance for H. (A.) genei A H. (A.) genei B (sensu Lanza, 2005) is 0.061, a value identical to the distance between the full species H. (S.) italicus and H. (S.) ambrosii. These similar distances are consistent with the finding of Carranza et al. (2007), who estimated the divergence time between these two lineages of H. (A.) genei to be similar to the divergence times between the mainland species based on a shorter fragment of cytochrome b (398 bp) combined with 373 bp of 12S rrna. The next four lowest genetic distances between full species based on our cytochrome b dataset are H. (S.) sarrabusensis H. (S.) imperialis (0.078), H. (S.) sarrabusensis H. (S.) flavus (0.079), H. (S.) strinatii H. (S.) ambrosii (0.080) and H. (S.) supramontis H. (S.) flavus (0.081, see Table 2). We found the average genetic distance between full (European) species to be much higher (0.117). Specimens of H. (A.) genei B were found at Nuxis (Tattinu), a locality more than 17 kilometers ESE outside the range described for H. (A.) genei B by Lanza et al. (2005). Nascetti et al. (1996) placed specimens from close to this locality ( Near Nuxus, 5.7 km W, and Near Pula, 11.5 km SSE of the locality of specimens 13 16) in the H. (A.) genei A group based on allozyme data. Table 2 Genetic p-distance based on 755 basepairs of the cytochrome b gene. H. (S.) ambrosii H. (S.) flavus H. (A.) genei H. (S.) imperialis H. (S.) sarrabusensis H. (S.) italicus H. (S.) strinatii H. (S.) flavus H. (A.) genei H. (S.) imperialis H. (S.) sarrabusensis H. (S.) italicus H. (S.) strinatii H. (S.) supramontis
5 A. van der Meijden et al. / Molecular Phylogenetics and Evolution xxx (2009) xxx xxx / / / / /- 0.49/- 0.98/- 0.72/- 1.00/74 H. (S.) imperialis 28 H. (S.) imperialis 29 H. (S.) imperialis 30 H. (S.) imperialis 34 H. (S.) imperialis 35 H. (S.) imperialis 36 H. (S.) imperialis 25 H. (S.) imperialis 26 H. (S.) imperialis 27 H. (S.) imperialis 33 H. (S.) imperialis 39 H. (S.) imperialis 37 H. (S.) imperialis 38 H. (S.) imperialis 31 H. (S.) imperialis 32 H. (S.) imperialis 23 H. (S.) imperialis 24 H. (S.) sarrabusensis 40 H. (S.) sarrabusensis 41 H. (S.) sarrabusensis 42 H. (S.) supramontis 44 H. (S.) supramontis 49 H. (S.) supramontis 51 H. (S.) supramontis 43 H. (S.) supramontis 50 H. (S.) supramontis 45 H. (S.) supramontis 46 H. (S.) supramontis 48 H. (S.) supramontis 47 H. (S.) flavus 21 H. (S.) flavus 22 H. (S.) flavus 20 H. (S.) flavus 18 H. (S.) flavus 19 H. (S.) flavus 17 H. (S.) italicus 56 H. (S.) italicus 58 H. (S.) italicus 57 H. (S.) italicus 59 H. (S.) italicus 60 H. (S.) italicus 61 H. (S.) ambrosii ambrosii 52 H. (S.) ambrosii ambrosii 53 H. (S.) ambrosii bianchii 54 H. (S.) ambrosii bianchii 55 H. (S.) strinatii 63 H. (S.) strinatii 64 H. (S.) strinatii 62 H. (S.) strinatii 65 H. (S.) strinatii 66 H. (A.) genei 5 H. (A.) genei 4 H. (A.) genei 9 H. (A.) genei 8 H. (A.) genei 6 H. (A.) genei 10 H. (A.) genei 12 H. (A.) genei 11 H. (A.) genei 7 H. (A.) genei 13 H. (A.) genei 16 H. (A.) genei 15 H. (A.) genei 14 H. (A.) genei 2 H. (A.) genei 1 H. (A.) genei 3 H. shastae H. brunus H. platycephalus H. (A.) genei "A" H. (A.) genei "B" Fig. 2. Bayesian Inference tree based on all data combined. Numbers at branches represent Bayesian posterior probabilities/maximum Likelihood bootstrap percentages based on 500 bootstrap replicates. Branches not present in the ML tree are indicated with a dash ( ). Outgroup taxa not shown. It therefore seems that H. (A.) genei A and B live in close proximity in the southern part of their range. Further studies with a higher sampling density including the localities described in Nascetti et al. (1996) will be necessary to address this issue Position of H. (S.) sarrabusensis Although resolved to be monophyletic in all analyses, the position of the clade representing H. (S.) sarrabusensis is different
6 6 A. van der Meijden et al. / Molecular Phylogenetics and Evolution xxx (2009) xxx xxx between the ML and BI trees. It is placed with low support sister to the Eastern Sardinia group formed by H. (S.) flavus, H. (S.) supramontis and H. (S.) imperialis by the ML analysis, whereas the BI analysis places it as a sister group to the H. (S.) imperialis clade (Fig. 2). A SH test showed these two placements to be not significantly different (p = 0.23). More data will be required to resolve the position of this newly distinguished taxon within the highly supported eastern Sardinian group Hydromantes (Speleomantes) imperialis from the Lake Omodeo area The samples from the locality near Lake Omodeo (specimen numbers 23 24, Fig. 1 and Table 1) at the westernmost border of the range of H. (S.) imperialis are resolved as distinct from other populations with a genetic distance of (p-distance) in cytochrome b. This is considerably higher than the highest genetic distance found between two populations of H.(S.) imperialis excluding these samples (p-distance of 0.040, between specimens 28 and 31, Fig. 1). The sampling of H. (S.) imperialis in the current study is highly disjunct (see Fig. 1), including samples only from the eastern and western parts of the distribution range. Therefore, the high genetic distance between the specimens from Lake Omodeo and the remaining H. (S.) imperialis populations could be an artifact of the disjunct sampling in this study, and might be bridged by including samples from intermediate populations. However, a study based on allozyme data by Cimmaruta et al. (1998), which did included some samples from more intermediate localities, showed the populations from near Lake Omodeo to be highly distinct from, and basal to the remaining populations of H. (S.) imperialis. Further studies including a denser sampling across the whole distribution range will be necessary to resolve the genetic structure of this widespread species. Such a study would be particularly necessary to assess the status of the populations near Lake Omodeo which, based on the sampling in the current study, display a genetic distance approaching that between other species of the genus Hydromantes. Acknowledgments We are thankful to the Italian Ministero dell Ambiente e della Tutela del Territorio (Prot.DPN , DPN , DPN and /04/2007) for issuing the permits for tissue collection. We are grateful to the speleologists of the Gruppo Grotte Ogliastra, Renzo Tocco, Titino Salis, Luciano Cuccu and to Vasco Alderighi for helping with the sampling. Field work has been supported by a grant of the Societas Europea Herpetologica to A.vdM., Y.C., M.M., M.V., S.C. was supported by grant CGL /BOS from the Ministerio de Educación y Ciencia, Spain. This is the ISE-M publication * ISEM of Y.C. References Brinkmann, H., Philippe, H., Animal phylogeny and large-scale sequencing: progress and pitfalls. J. Syst. Evol. 46 (3), Caccone, A., Milinkovitch, M.C., Sbordoni, V., Powell, J.R., Molecular biogeography: using the Corsica-Sardinia microplate disjunction to calibrate mitochondrial rdna evolutionary rates in mountain newts (Euproctus). J. Evol. Biol. 7, Carranza, S., Romano, A., Arnold, E.N., Sotgiu, G., Biogeography and evolution of European cave salamanders, Hydromantes (Urodela: Plethodontidae), inferred from mtdna sequences. J. Biogeogr. 35 (4), Cimmaruta, R., Nascetti, G., Forti, G., Lanza, B., Bullini, L., Paleogeografia della Sardegna ed evoluzione degli Hydromantes (Amphibia, Plethodontidae). Biogeographia XIX (1997), Edgar, R.C., MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32 (5), Farris, J.S., Kallersjo, M., Kluge, A.G., Bult, C., Testing significance of incongruence. Cladistics 10, Grill, A., Casula, P., Lecis, R., Menken, S.B.J., Endemism in Sardinia. In: Weiss, S., Ferrand, N. (Eds.), Phylogeography of Southern European Refugia. Springer, Dordrecht, The Netherlands, pp Guindon, S., Gascuel, O., A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst. Biol. 52, Huelsenbeck, J.P., Ronquist, F., MRBAYES: Bayesian inference of phylogenetic trees. Bioinformatics 17, Jackman, T.R., Applebaum, G., Wake, D.B., Phylogenetic relationships of bolitoglossine salamanders: a demonstration of the effects of combining morphological and molecular data sets. Mol. Biol. Evol. 14, Lanza, B., Ipotesi sulle origini del popolamento erpetologico della Sardegna. Lav. Soc. Ital. Biogeog. (N.S.) 8, Lanza, B., Caputo, V. Nascetti, G., Bullini, L., Morphologic and genetic studies of the European plethodontid salamanders: taxonomic inferences (genus Hydromantes). Monografie XVI, Museo Regionale di Scienze Naturali, Torino. Lanza, B., Pastorelli, C., Laghi, P., Cimmaruta, R., A review of systematics, taxonomy, genetics, biogeography and natural history of the genus Speleomantes DUBOIS, 1984 (Amphibia Caudata Plethodontidae). Atti. Mus. Stor. Nat. Trieste (Suppl.) 52, Marra, A.C., Pleistocene mammals of Mediterranean islands. Quatern. Int. 129, Van der Meijden, A., Vences, M., Hoegg, S., Boistel, R., Channing, A., Meyer, A., Nuclear gene phylogeny of narrow-mouthed toads (Family Microhylidae) and a discussion of competing hypotheses concerning their biogeographical origins. Mol. Phylogenet. Evol. 44 (3), Van der Meijden, A., Vences, M., Hoegg, S., Meyer, A., A previously unrecognized radiation of ranid frogs in Southern Africa revealed by nuclear and mitochondrial DNA sequences. Mol. Phylogenet. Evol. 37 (3), Nardi, I., Cytogenetics of the European plethodontid salamanders, genus Hydromantes. In: Green, D.M., Sessions, S.K. (Eds.), Amphibian Cytogenetics and Evolution. Academic Press Inc., San Diego, CA, pp Nascetti, I., Cimmaruta, R., Lanza, B., Bullini, L., Molecular taxonomy of European plethodontid salamanders (genus Hydromantes). J. Herpetol. 30, Oliverio, M., Bologna, M.A., Mariottini, P., Molecular biogeography of the Mediterranean lizard Podarcis Wagler, 1830 and Teira Gray, 1838 (Reptilia, Lacertidae). J. Biogeogr. 27, Omodeo, P., Rota, E., Earthworm diversity and land evolution in three Mediterranean districts. Proc. Calif. Acad. Sci., Fourth Series, Suppl. I 59 (5), Oosterbroek, P., Arntzen, J.W., Area-cladograms of circum-mediterranean taxa in relation to Mediterranean paleogeography. J. Biogeogr. 19, Palumbi, S.R., Martin, A., Romano, S., McMillan, W.O., Stice, L., Grabowski, G., The Simple Fool s Guide to PCR, Version 2.0. University of Hawaii, Pollock, D.D., Zwickl, D.J., McGuire, J.A., Hillis, D.M., Increased taxon sampling is advantageous for phylogenetic inference. Syst. Biol. 51 (4), Posada, D., Crandall, K.A., Modeltest: testing the model of DNA substitution. Bioinformatics 14 (9), Swofford, D.L., PAUP *. Phylogenetic Analysis using Parsimony ( * and Other Methods). Sinauer Associates, Sunderland, MA. Tamura, K., Dudley, J., Nei, M., Kumar, S., MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 24, Vieites, D.R., Min, M.-S., Wake, D.B., Rapid diversification and dispersal during periods of global warming by plethodontid salamanders. Proc. Natl. Acad. Sci. USA 104, Wake, D.B., Maxson, L.R., Wurst, G.Z., Genetic differentiation, albumin evolution, and their biogeographic implications in plethodontid salamanders of California and southern Europe. Evolution 32, Wiens, J.J., Fetzner, J.W., Parkinson, C.L., Reeder, T.W., Hylid frog phylogeny and sampling strategies for speciose clades. Syst. Biol. 5,
Biogeography and evolution of European cave salamanders, Hydromantes (Urodela: Plethodontidae), inferred from mtdna sequences
 Journal of Biogeography (J. Biogeogr.) (2008) 35, 724 738 ORIGINAL ARTICLE Biogeography and evolution of European cave salamanders, Hydromantes (Urodela: Plethodontidae), inferred from mtdna sequences
Journal of Biogeography (J. Biogeogr.) (2008) 35, 724 738 ORIGINAL ARTICLE Biogeography and evolution of European cave salamanders, Hydromantes (Urodela: Plethodontidae), inferred from mtdna sequences
Phylogeography of Sardinian Cave Salamanders (Genus Hydromantes) Is Mainly Determined by Geomorphology
 (Genus Hydromantes) Is Mainly Determined by Geomorphology Ylenia Chiari 1,2 *, Arie van der Meijden 2, Mauro Mucedda 3, João M. Lourenço 1, Axel Hochkirch 4, Michael Veith 4 1 Institut des Sciences de
(Genus Hydromantes) Is Mainly Determined by Geomorphology Ylenia Chiari 1,2 *, Arie van der Meijden 2, Mauro Mucedda 3, João M. Lourenço 1, Axel Hochkirch 4, Michael Veith 4 1 Institut des Sciences de
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Consensus Methods. * You are only responsible for the first two
 Consensus Trees * consensus trees reconcile clades from different trees * consensus is a conservative estimate of phylogeny that emphasizes points of agreement * philosophy: agreement among data sets is
Consensus Trees * consensus trees reconcile clades from different trees * consensus is a conservative estimate of phylogeny that emphasizes points of agreement * philosophy: agreement among data sets is
Ontogenetic convergence and evolution of foot morphology in European cave salamanders (Family: Plethodontidae)
 RESEARCH ARTICLE Open Access Research article Ontogenetic convergence and evolution of foot morphology in European cave salamanders (Family: Plethodontidae) Dean C Adams* 1 and Annamaria Nistri 2 Abstract
RESEARCH ARTICLE Open Access Research article Ontogenetic convergence and evolution of foot morphology in European cave salamanders (Family: Plethodontidae) Dean C Adams* 1 and Annamaria Nistri 2 Abstract
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
Effects of Gap Open and Gap Extension Penalties
 Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
Brigham Young University BYU ScholarsArchive All Faculty Publications 200-10-01 Effects of Gap Open and Gap Extension Penalties Hyrum Carroll hyrumcarroll@gmail.com Mark J. Clement clement@cs.byu.edu See
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
Anatomy of a species tree
 Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Phylogenetic relationships of Pseudohynobius (Urodela, Hynobiidae) inferred from DNA barcoding analysis
 Phylogenetic relationships of Pseudohynobius (Urodela, Hynobiidae) inferred from DNA barcoding analysis Y.-Y. Zhao 1 *, L.-N. Su 3 *, Z.-M. Zhang 1 and X.-Y. Wang 2 1 Department of Genetics, Zunyi Medical
Phylogenetic relationships of Pseudohynobius (Urodela, Hynobiidae) inferred from DNA barcoding analysis Y.-Y. Zhao 1 *, L.-N. Su 3 *, Z.-M. Zhang 1 and X.-Y. Wang 2 1 Department of Genetics, Zunyi Medical
GIS Applications to Museum Specimens
 GIS Applications to Museum Specimens Joseph Grinnell (1877 1939) At this point I wish to emphasize what I believe will ultimately prove to be the greatest value of our museum. This value will not, however,
GIS Applications to Museum Specimens Joseph Grinnell (1877 1939) At this point I wish to emphasize what I believe will ultimately prove to be the greatest value of our museum. This value will not, however,
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
ESS 345 Ichthyology. Systematic Ichthyology Part II Not in Book
 ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
Identification of COI partial sequences in two closely related frog species, Rana dalmatina and Rana temporaria
 HERPETOLOGICA ROMANICA Vol. 4, 2010, pp.1-6 ISSN: 1842-9203 Article No. 041101 Identification of COI partial sequences in two closely related frog species, Rana dalmatina and Rana temporaria Béla MAROSI
HERPETOLOGICA ROMANICA Vol. 4, 2010, pp.1-6 ISSN: 1842-9203 Article No. 041101 Identification of COI partial sequences in two closely related frog species, Rana dalmatina and Rana temporaria Béla MAROSI
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Letter to the Editor. Department of Biology, Arizona State University
 Letter to the Editor Traditional Phylogenetic Reconstruction Methods Reconstruct Shallow and Deep Evolutionary Relationships Equally Well Michael S. Rosenberg and Sudhir Kumar Department of Biology, Arizona
Letter to the Editor Traditional Phylogenetic Reconstruction Methods Reconstruct Shallow and Deep Evolutionary Relationships Equally Well Michael S. Rosenberg and Sudhir Kumar Department of Biology, Arizona
Lecture 11 Friday, October 21, 2011
 Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Nesting of cave salamanders (Hydromantes flavus and H. italicus) in natural environments
 SALAMANDRA 50(2) 105 109 30 Nesting June 2014 of Hydromantes ISSN 0036 3375 flavus and H. italicus Nesting of cave salamanders (Hydromantes flavus and H. italicus) in natural environments Enrico Lunghi
SALAMANDRA 50(2) 105 109 30 Nesting June 2014 of Hydromantes ISSN 0036 3375 flavus and H. italicus Nesting of cave salamanders (Hydromantes flavus and H. italicus) in natural environments Enrico Lunghi
Historical Biogeography. Historical Biogeography. Systematics
 Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
Historical Biogeography I. Definitions II. Fossils: problems with fossil record why fossils are important III. Phylogeny IV. Phenetics VI. Phylogenetic Classification Disjunctions debunked: Examples VII.
How should we organize the diversity of animal life?
 How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
How should we organize the diversity of animal life? The difference between Taxonomy Linneaus, and Cladistics Darwin What are phylogenies? How do we read them? How do we estimate them? Classification (Taxonomy)
Introduction to characters and parsimony analysis
 Introduction to characters and parsimony analysis Genetic Relationships Genetic relationships exist between individuals within populations These include ancestordescendent relationships and more indirect
Introduction to characters and parsimony analysis Genetic Relationships Genetic relationships exist between individuals within populations These include ancestordescendent relationships and more indirect
Efficiencies of maximum likelihood methods of phylogenetic inferences when different substitution models are used
 Molecular Phylogenetics and Evolution 31 (2004) 865 873 MOLECULAR PHYLOGENETICS AND EVOLUTION www.elsevier.com/locate/ympev Efficiencies of maximum likelihood methods of phylogenetic inferences when different
Molecular Phylogenetics and Evolution 31 (2004) 865 873 MOLECULAR PHYLOGENETICS AND EVOLUTION www.elsevier.com/locate/ympev Efficiencies of maximum likelihood methods of phylogenetic inferences when different
Bootstrapping and Tree reliability. Biol4230 Tues, March 13, 2018 Bill Pearson Pinn 6-057
 Bootstrapping and Tree reliability Biol4230 Tues, March 13, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Rooting trees (outgroups) Bootstrapping given a set of sequences sample positions randomly,
Bootstrapping and Tree reliability Biol4230 Tues, March 13, 2018 Bill Pearson wrp@virginia.edu 4-2818 Pinn 6-057 Rooting trees (outgroups) Bootstrapping given a set of sequences sample positions randomly,
Today's project. Test input data Six alignments (from six independent markers) of Curcuma species
 DNA sequences II Analyses of multiple sequence data datasets, incongruence tests, gene trees vs. species tree reconstruction, networks, detection of hybrid species DNA sequences II Test of congruence of
DNA sequences II Analyses of multiple sequence data datasets, incongruence tests, gene trees vs. species tree reconstruction, networks, detection of hybrid species DNA sequences II Test of congruence of
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley B.D. Mishler Jan. 22, 2009. Trees I. Summary of previous lecture: Hennigian
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley B.D. Mishler Jan. 22, 2009. Trees I. Summary of previous lecture: Hennigian
Lecture V Phylogeny and Systematics Dr. Kopeny
 Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Applications of Genetics to Conservation Biology
 Applications of Genetics to Conservation Biology Molecular Taxonomy Populations, Gene Flow, Phylogeography Relatedness - Kinship, Paternity, Individual ID Conservation Biology Population biology Physiology
Applications of Genetics to Conservation Biology Molecular Taxonomy Populations, Gene Flow, Phylogeography Relatedness - Kinship, Paternity, Individual ID Conservation Biology Population biology Physiology
Phylogenies Scores for Exhaustive Maximum Likelihood and Parsimony Scores Searches
 Int. J. Bioinformatics Research and Applications, Vol. x, No. x, xxxx Phylogenies Scores for Exhaustive Maximum Likelihood and s Searches Hyrum D. Carroll, Perry G. Ridge, Mark J. Clement, Quinn O. Snell
Int. J. Bioinformatics Research and Applications, Vol. x, No. x, xxxx Phylogenies Scores for Exhaustive Maximum Likelihood and s Searches Hyrum D. Carroll, Perry G. Ridge, Mark J. Clement, Quinn O. Snell
SPECIATION. REPRODUCTIVE BARRIERS PREZYGOTIC: Barriers that prevent fertilization. Habitat isolation Populations can t get together
 SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
Bioinformatics tools for phylogeny and visualization. Yanbin Yin
 Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
Bioinformatics tools for phylogeny and visualization Yanbin Yin 1 Homework assignment 5 1. Take the MAFFT alignment http://cys.bios.niu.edu/yyin/teach/pbb/purdue.cellwall.list.lignin.f a.aln as input and
A data based parsimony method of cophylogenetic analysis
 Blackwell Science, Ltd A data based parsimony method of cophylogenetic analysis KEVIN P. JOHNSON, DEVIN M. DROWN & DALE H. CLAYTON Accepted: 20 October 2000 Johnson, K. P., Drown, D. M. & Clayton, D. H.
Blackwell Science, Ltd A data based parsimony method of cophylogenetic analysis KEVIN P. JOHNSON, DEVIN M. DROWN & DALE H. CLAYTON Accepted: 20 October 2000 Johnson, K. P., Drown, D. M. & Clayton, D. H.
Systematics - Bio 615
 Bayesian Phylogenetic Inference 1. Introduction, history 2. Advantages over ML 3. Bayes Rule 4. The Priors 5. Marginal vs Joint estimation 6. MCMC Derek S. Sikes University of Alaska 7. Posteriors vs Bootstrap
Bayesian Phylogenetic Inference 1. Introduction, history 2. Advantages over ML 3. Bayes Rule 4. The Priors 5. Marginal vs Joint estimation 6. MCMC Derek S. Sikes University of Alaska 7. Posteriors vs Bootstrap
Molecular Phylogenetics and Evolution
 Molecular Phylogenetics and Evolution 53 (2009) 340 344 Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: www.elsevier.com/locate/ympev Short Communication
Molecular Phylogenetics and Evolution 53 (2009) 340 344 Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: www.elsevier.com/locate/ympev Short Communication
Geography of Evolution
 Geography of Evolution Biogeography - the study of the geographic distribution of organisms. The current distribution of organisms can be explained by historical events and current climatic patterns. Darwin
Geography of Evolution Biogeography - the study of the geographic distribution of organisms. The current distribution of organisms can be explained by historical events and current climatic patterns. Darwin
Concepts and Methods in Molecular Divergence Time Estimation
 Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Phylogenetic inference
 Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Phylogenetic inference Bas E. Dutilh Systems Biology: Bioinformatic Data Analysis Utrecht University, March 7 th 016 After this lecture, you can discuss (dis-) advantages of different information types
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Macroevolution Part I: Phylogenies
 Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Macroevolution Part I: Phylogenies Taxonomy Classification originated with Carolus Linnaeus in the 18 th century. Based on structural (outward and inward) similarities Hierarchal scheme, the largest most
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft]
![Integrative Biology 200 PRINCIPLES OF PHYLOGENETICS Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft] Integrative Biology 200 PRINCIPLES OF PHYLOGENETICS Spring 2016 University of California, Berkeley. Parsimony & Likelihood [draft]](/thumbs/83/87443456.jpg) Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley K.W. Will Parsimony & Likelihood [draft] 1. Hennig and Parsimony: Hennig was not concerned with parsimony
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2016 University of California, Berkeley K.W. Will Parsimony & Likelihood [draft] 1. Hennig and Parsimony: Hennig was not concerned with parsimony
Phylogenomics. Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University. Tuesday, January 29, 13
 Phylogenomics Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University How may we improve our inferences? How may we improve our inferences? Inferences Data How may we improve
Phylogenomics Jeffrey P. Townsend Department of Ecology and Evolutionary Biology Yale University How may we improve our inferences? How may we improve our inferences? Inferences Data How may we improve
Reconstructing the history of lineages
 Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis
 Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Elements of Bioinformatics 14F01 TP5 -Phylogenetic analysis 10 December 2012 - Corrections - Exercise 1 Non-vertebrate chordates generally possess 2 homologs, vertebrates 3 or more gene copies; a Drosophila
Ph ylogeography. A guide to the study of the spatial distribution of Seahorses. By Leila Mougoui Bakhtiari
 Ph ylogeography A guide to the study of the spatial distribution of Seahorses By Leila Mougoui Bakhtiari Contents An Introduction to Phylogeography JT Bohem s Resarch Map of erectu s migration Conservation
Ph ylogeography A guide to the study of the spatial distribution of Seahorses By Leila Mougoui Bakhtiari Contents An Introduction to Phylogeography JT Bohem s Resarch Map of erectu s migration Conservation
INCREASED RATES OF MOLECULAR EVOLUTION IN AN EQUATORIAL PLANT CLADE: AN EFFECT OF ENVIRONMENT OR PHYLOGENETIC NONINDEPENDENCE?
 Evolution, 59(1), 2005, pp. 238 242 INCREASED RATES OF MOLECULAR EVOLUTION IN AN EQUATORIAL PLANT CLADE: AN EFFECT OF ENVIRONMENT OR PHYLOGENETIC NONINDEPENDENCE? JEREMY M. BROWN 1,2 AND GREGORY B. PAULY
Evolution, 59(1), 2005, pp. 238 242 INCREASED RATES OF MOLECULAR EVOLUTION IN AN EQUATORIAL PLANT CLADE: AN EFFECT OF ENVIRONMENT OR PHYLOGENETIC NONINDEPENDENCE? JEREMY M. BROWN 1,2 AND GREGORY B. PAULY
Bayesian Inference using Markov Chain Monte Carlo in Phylogenetic Studies
 Bayesian Inference using Markov Chain Monte Carlo in Phylogenetic Studies 1 What is phylogeny? Essay written for the course in Markov Chains 2004 Torbjörn Karfunkel Phylogeny is the evolutionary development
Bayesian Inference using Markov Chain Monte Carlo in Phylogenetic Studies 1 What is phylogeny? Essay written for the course in Markov Chains 2004 Torbjörn Karfunkel Phylogeny is the evolutionary development
Hexapods Resurrected
 Hexapods Resurrected (Technical comment on: "Hexapod Origins: Monophyletic or Paraphyletic?") Frédéric Delsuc, Matthew J. Phillips and David Penny The Allan Wilson Centre for Molecular Ecology and Evolution
Hexapods Resurrected (Technical comment on: "Hexapod Origins: Monophyletic or Paraphyletic?") Frédéric Delsuc, Matthew J. Phillips and David Penny The Allan Wilson Centre for Molecular Ecology and Evolution
Fields connected to Phylogeography Microevolutionary disciplines Ethology Demography Population genetics
 Stephen A. Roussos Fields connected to Phylogeography Microevolutionary disciplines Ethology Demography Population genetics Macrevolutionary disciplines Historical geography Paleontology Phylogenetic biology
Stephen A. Roussos Fields connected to Phylogeography Microevolutionary disciplines Ethology Demography Population genetics Macrevolutionary disciplines Historical geography Paleontology Phylogenetic biology
Phylogenetic position of Pelusios williamsi and a critique of current GenBank procedures (Reptilia: Testudines: Pelomedusidae)
 Amphibia-Reptilia 33 (2012): 150-154 Phylogenetic position of Pelusios williamsi and a critique of current GenBank procedures (Reptilia: Testudines: Pelomedusidae) Uwe Fritz 1,, Mario Vargas-Ramírez 1,
Amphibia-Reptilia 33 (2012): 150-154 Phylogenetic position of Pelusios williamsi and a critique of current GenBank procedures (Reptilia: Testudines: Pelomedusidae) Uwe Fritz 1,, Mario Vargas-Ramírez 1,
Assessing an Unknown Evolutionary Process: Effect of Increasing Site- Specific Knowledge Through Taxon Addition
 Assessing an Unknown Evolutionary Process: Effect of Increasing Site- Specific Knowledge Through Taxon Addition David D. Pollock* and William J. Bruno* *Theoretical Biology and Biophysics, Los Alamos National
Assessing an Unknown Evolutionary Process: Effect of Increasing Site- Specific Knowledge Through Taxon Addition David D. Pollock* and William J. Bruno* *Theoretical Biology and Biophysics, Los Alamos National
1/27/2010. Systematics and Phylogenetics of the. An Introduction. Taxonomy and Systematics
 Systematics and Phylogenetics of the Amphibia: An Introduction Taxonomy and Systematics Taxonomy, the science of describing biodiversity, mainly naming unnamed species, and arranging the diversity into
Systematics and Phylogenetics of the Amphibia: An Introduction Taxonomy and Systematics Taxonomy, the science of describing biodiversity, mainly naming unnamed species, and arranging the diversity into
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization)
 STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
Biology 211 (2) Week 1 KEY!
 Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Biology 211 (2) Week 1 KEY Chapter 1 KEY FIGURES: 1.2, 1.3, 1.4, 1.5, 1.6, 1.7 VOCABULARY: Adaptation: a trait that increases the fitness Cells: a developed, system bound with a thin outer layer made of
Phylogenetics. Applications of phylogenetics. Unrooted networks vs. rooted trees. Outline
 Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
Phylogenetics Todd Vision iology 522 March 26, 2007 pplications of phylogenetics Studying organismal or biogeographic history Systematics ating events in the fossil record onservation biology Studying
What is Phylogenetics
 What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
AP Biology. Cladistics
 Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
Cladistics Kingdom Summary Review slide Review slide Classification Old 5 Kingdom system Eukaryote Monera, Protists, Plants, Fungi, Animals New 3 Domain system reflects a greater understanding of evolution
Letter to the Editor. Temperature Hypotheses. David P. Mindell, Alec Knight,? Christine Baer,$ and Christopher J. Huddlestons
 Letter to the Editor Slow Rates of Molecular Evolution Temperature Hypotheses in Birds and the Metabolic Rate and Body David P. Mindell, Alec Knight,? Christine Baer,$ and Christopher J. Huddlestons *Department
Letter to the Editor Slow Rates of Molecular Evolution Temperature Hypotheses in Birds and the Metabolic Rate and Body David P. Mindell, Alec Knight,? Christine Baer,$ and Christopher J. Huddlestons *Department
Ratio of explanatory power (REP): A new measure of group support
 Molecular Phylogenetics and Evolution 44 (2007) 483 487 Short communication Ratio of explanatory power (REP): A new measure of group support Taran Grant a, *, Arnold G. Kluge b a Division of Vertebrate
Molecular Phylogenetics and Evolution 44 (2007) 483 487 Short communication Ratio of explanatory power (REP): A new measure of group support Taran Grant a, *, Arnold G. Kluge b a Division of Vertebrate
PHYLOGENY & THE TREE OF LIFE
 PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
PHYLOGENY & THE TREE OF LIFE PREFACE In this powerpoint we learn how biologists distinguish and categorize the millions of species on earth. Early we looked at the process of evolution here we look at
Biogeography expands:
 Biogeography expands: Phylogeography Ecobiogeography Due to advances in DNA sequencing and fingerprinting methods, historical biogeography has recently begun to integrate relationships of populations within
Biogeography expands: Phylogeography Ecobiogeography Due to advances in DNA sequencing and fingerprinting methods, historical biogeography has recently begun to integrate relationships of populations within
Using phylogenetics to estimate species divergence times... Basics and basic issues for Bayesian inference of divergence times (plus some digression)
 Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Combining Data Sets with Different Phylogenetic Histories
 Syst. Biol. 47(4):568 581, 1998 Combining Data Sets with Different Phylogenetic Histories JOHN J. WIENS Section of Amphibians and Reptiles, Carnegie Museum of Natural History, Pittsburgh, Pennsylvania
Syst. Biol. 47(4):568 581, 1998 Combining Data Sets with Different Phylogenetic Histories JOHN J. WIENS Section of Amphibians and Reptiles, Carnegie Museum of Natural History, Pittsburgh, Pennsylvania
Name. Ecology & Evolutionary Biology 2245/2245W Exam 2 1 March 2014
 Name 1 Ecology & Evolutionary Biology 2245/2245W Exam 2 1 March 2014 1. Use the following matrix of nucleotide sequence data and the corresponding tree to answer questions a. through h. below. (16 points)
Name 1 Ecology & Evolutionary Biology 2245/2245W Exam 2 1 March 2014 1. Use the following matrix of nucleotide sequence data and the corresponding tree to answer questions a. through h. below. (16 points)
The Phylogenetic Reconstruction of the Grass Family (Poaceae) Using matk Gene Sequences
 The Phylogenetic Reconstruction of the Grass Family (Poaceae) Using matk Gene Sequences by Hongping Liang Dissertation submitted to the Faculty of the Virginia Polytechnic Institute and State University
The Phylogenetic Reconstruction of the Grass Family (Poaceae) Using matk Gene Sequences by Hongping Liang Dissertation submitted to the Faculty of the Virginia Polytechnic Institute and State University
Letter to the Editor. The Effect of Taxonomic Sampling on Accuracy of Phylogeny Estimation: Test Case of a Known Phylogeny Steven Poe 1
 Letter to the Editor The Effect of Taxonomic Sampling on Accuracy of Phylogeny Estimation: Test Case of a Known Phylogeny Steven Poe 1 Department of Zoology and Texas Memorial Museum, University of Texas
Letter to the Editor The Effect of Taxonomic Sampling on Accuracy of Phylogeny Estimation: Test Case of a Known Phylogeny Steven Poe 1 Department of Zoology and Texas Memorial Museum, University of Texas
Diversity and Human Evolution. Homo neanderthalensis. Homo neanderthalensis. Homo neanderthalensis. Homo neanderthalensis. Part II
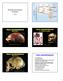 Diversity and Human Evolution Part II Neanderthal 1 La Chapelle-aux-Saints Photograph byrheinisches LandesmuseumBonn Photographs by John Reader Mount Circeo Photograph by Ministry of Culture, Italy An
Diversity and Human Evolution Part II Neanderthal 1 La Chapelle-aux-Saints Photograph byrheinisches LandesmuseumBonn Photographs by John Reader Mount Circeo Photograph by Ministry of Culture, Italy An
Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them?
 Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them? Carolus Linneaus:Systema Naturae (1735) Swedish botanist &
Phylogenies & Classifying species (AKA Cladistics & Taxonomy) What are phylogenies & cladograms? How do we read them? How do we estimate them? Carolus Linneaus:Systema Naturae (1735) Swedish botanist &
Molecular evidence for multiple origins of Insectivora and for a new order of endemic African insectivore mammals
 Project Report Molecular evidence for multiple origins of Insectivora and for a new order of endemic African insectivore mammals By Shubhra Gupta CBS 598 Phylogenetic Biology and Analysis Life Science
Project Report Molecular evidence for multiple origins of Insectivora and for a new order of endemic African insectivore mammals By Shubhra Gupta CBS 598 Phylogenetic Biology and Analysis Life Science
The California Hotspots Project: I.
 The California Hotspots Project: I. Identifying regions of rapid diversification of mammals Ed Davis, M. Koo, C. Conroy, J. Patton & C. Moritz Museum of Vertebrate Zoology, UC Berkeley *Funded by Resources
The California Hotspots Project: I. Identifying regions of rapid diversification of mammals Ed Davis, M. Koo, C. Conroy, J. Patton & C. Moritz Museum of Vertebrate Zoology, UC Berkeley *Funded by Resources
Chapter 26. Phylogeny and the Tree of Life. Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Pearson Education, Inc.
 Chapter 26 Phylogeny and the Tree of Life Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Investigating the Tree of Life Phylogeny is the evolutionary history of a species or group of
Chapter 26 Phylogeny and the Tree of Life Lecture Presentations by Nicole Tunbridge and Kathleen Fitzpatrick Investigating the Tree of Life Phylogeny is the evolutionary history of a species or group of
Application of 16S rrna, cytochrome b and control region sequences for understanding the phylogenetic relationships in Oryx species
 Short Communication Application of 16S rrna, cytochrome b and control region sequences for understanding the phylogenetic relationships in Oryx species H.A. Khan, I.A. Arif, A.A. Al Homaidan and A.H. Al
Short Communication Application of 16S rrna, cytochrome b and control region sequences for understanding the phylogenetic relationships in Oryx species H.A. Khan, I.A. Arif, A.A. Al Homaidan and A.H. Al
Inferring Molecular Phylogeny
 Dr. Walter Salzburger he tree of life, ustav Klimt (1907) Inferring Molecular Phylogeny Inferring Molecular Phylogeny 55 Maximum Parsimony (MP): objections long branches I!! B D long branch attraction
Dr. Walter Salzburger he tree of life, ustav Klimt (1907) Inferring Molecular Phylogeny Inferring Molecular Phylogeny 55 Maximum Parsimony (MP): objections long branches I!! B D long branch attraction
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Lecture 6 Phylogenetic Inference
 Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Chapter 19: Taxonomy, Systematics, and Phylogeny
 Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Algorithmic Methods Well-defined methodology Tree reconstruction those that are well-defined enough to be carried out by a computer. Felsenstein 2004,
 Tracing the Evolution of Numerical Phylogenetics: History, Philosophy, and Significance Adam W. Ferguson Phylogenetic Systematics 26 January 2009 Inferring Phylogenies Historical endeavor Darwin- 1837
Tracing the Evolution of Numerical Phylogenetics: History, Philosophy, and Significance Adam W. Ferguson Phylogenetic Systematics 26 January 2009 Inferring Phylogenies Historical endeavor Darwin- 1837
The Importance of Data Partitioning and the Utility of Bayes Factors in Bayesian Phylogenetics
 Syst. Biol. 56(4):643 655, 2007 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150701546249 The Importance of Data Partitioning and the Utility
Syst. Biol. 56(4):643 655, 2007 Copyright c Society of Systematic Biologists ISSN: 1063-5157 print / 1076-836X online DOI: 10.1080/10635150701546249 The Importance of Data Partitioning and the Utility
Classification and Phylogeny
 Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Molecular Markers, Natural History, and Evolution
 Molecular Markers, Natural History, and Evolution Second Edition JOHN C. AVISE University of Georgia Sinauer Associates, Inc. Publishers Sunderland, Massachusetts Contents PART I Background CHAPTER 1:
Molecular Markers, Natural History, and Evolution Second Edition JOHN C. AVISE University of Georgia Sinauer Associates, Inc. Publishers Sunderland, Massachusetts Contents PART I Background CHAPTER 1:
Smith et al. American Journal of Botany 98(3): Data Supplement S2 page 1
 Smith et al. American Journal of Botany 98(3):404-414. 2011. Data Supplement S1 page 1 Smith, Stephen A., Jeremy M. Beaulieu, Alexandros Stamatakis, and Michael J. Donoghue. 2011. Understanding angiosperm
Smith et al. American Journal of Botany 98(3):404-414. 2011. Data Supplement S1 page 1 Smith, Stephen A., Jeremy M. Beaulieu, Alexandros Stamatakis, and Michael J. Donoghue. 2011. Understanding angiosperm
Biology 1B Evolution Lecture 2 (February 26, 2010) Natural Selection, Phylogenies
 1 Natural Selection (Darwin-Wallace): There are three conditions for natural selection: 1. Variation: Individuals within a population have different characteristics/traits (or phenotypes). 2. Inheritance:
1 Natural Selection (Darwin-Wallace): There are three conditions for natural selection: 1. Variation: Individuals within a population have different characteristics/traits (or phenotypes). 2. Inheritance:
Anatomy of a tree. clade is group of organisms with a shared ancestor. a monophyletic group shares a single common ancestor = tapirs-rhinos-horses
 Anatomy of a tree outgroup: an early branching relative of the interest groups sister taxa: taxa derived from the same recent ancestor polytomy: >2 taxa emerge from a node Anatomy of a tree clade is group
Anatomy of a tree outgroup: an early branching relative of the interest groups sister taxa: taxa derived from the same recent ancestor polytomy: >2 taxa emerge from a node Anatomy of a tree clade is group
Classification and Phylogeny
 Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
Classification and Phylogeny The diversity it of life is great. To communicate about it, there must be a scheme for organization. There are many species that would be difficult to organize without a scheme
A new finding of Salamandra lanzai
 Acta Herpetologica 2(1): 53-58, 2007 A new finding of Salamandra lanzai in the Upper Sangone Valley (NW Italy) marks the species most disjunct population (Amphibia: Urodela: Salamandridae) Giulia Tessa
Acta Herpetologica 2(1): 53-58, 2007 A new finding of Salamandra lanzai in the Upper Sangone Valley (NW Italy) marks the species most disjunct population (Amphibia: Urodela: Salamandridae) Giulia Tessa
BIG4: Biosystematics, informatics and genomics of the big 4 insect groups- training tomorrow s researchers and entrepreneurs
 BIG4: Biosystematics, informatics and genomics of the big 4 insect groups- training tomorrow s researchers and entrepreneurs Kick-Off Meeting 14-18 September 2015 Copenhagen, Denmark This project has received
BIG4: Biosystematics, informatics and genomics of the big 4 insect groups- training tomorrow s researchers and entrepreneurs Kick-Off Meeting 14-18 September 2015 Copenhagen, Denmark This project has received
Homoplasy. Selection of models of molecular evolution. Evolutionary correction. Saturation
 Homoplasy Selection of models of molecular evolution David Posada Homoplasy indicates identity not produced by descent from a common ancestor. Graduate class in Phylogenetics, Campus Agrário de Vairão,
Homoplasy Selection of models of molecular evolution David Posada Homoplasy indicates identity not produced by descent from a common ancestor. Graduate class in Phylogenetics, Campus Agrário de Vairão,
Cladistics and Bioinformatics Questions 2013
 AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
AP Biology Name Cladistics and Bioinformatics Questions 2013 1. The following table shows the percentage similarity in sequences of nucleotides from a homologous gene derived from five different species
Evaluating phylogenetic hypotheses
 Evaluating phylogenetic hypotheses Methods for evaluating topologies Topological comparisons: e.g., parametric bootstrapping, constrained searches Methods for evaluating nodes Resampling techniques: bootstrapping,
Evaluating phylogenetic hypotheses Methods for evaluating topologies Topological comparisons: e.g., parametric bootstrapping, constrained searches Methods for evaluating nodes Resampling techniques: bootstrapping,
MOLECULAR SYSTEMATICS: A SYNTHESIS OF THE COMMON METHODS AND THE STATE OF KNOWLEDGE
 CELLULAR & MOLECULAR BIOLOGY LETTERS http://www.cmbl.org.pl Received: 16 August 2009 Volume 15 (2010) pp 311-341 Final form accepted: 01 March 2010 DOI: 10.2478/s11658-010-0010-8 Published online: 19 March
CELLULAR & MOLECULAR BIOLOGY LETTERS http://www.cmbl.org.pl Received: 16 August 2009 Volume 15 (2010) pp 311-341 Final form accepted: 01 March 2010 DOI: 10.2478/s11658-010-0010-8 Published online: 19 March
Phylogenetics in the Age of Genomics: Prospects and Challenges
 Phylogenetics in the Age of Genomics: Prospects and Challenges Antonis Rokas Department of Biological Sciences, Vanderbilt University http://as.vanderbilt.edu/rokaslab http://pubmed2wordle.appspot.com/
Phylogenetics in the Age of Genomics: Prospects and Challenges Antonis Rokas Department of Biological Sciences, Vanderbilt University http://as.vanderbilt.edu/rokaslab http://pubmed2wordle.appspot.com/
CHAPTER 10 Taxonomy and Phylogeny of Animals
 CHAPTER 10 Taxonomy and Phylogeny of Animals 10-1 10-2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Linnaeus and Taxonomy More than 1.5 million species of
CHAPTER 10 Taxonomy and Phylogeny of Animals 10-1 10-2 Copyright The McGraw-Hill Companies, Inc. Permission required for reproduction or display. Linnaeus and Taxonomy More than 1.5 million species of
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200 Spring 2018 University of California, Berkeley D.D. Ackerly Feb. 26, 2018 Maximum Likelihood Principles, and Applications to
OMICS Journals are welcoming Submissions
 OMICS Journals are welcoming Submissions OMICS International welcomes submissions that are original and technically so as to serve both the developing world and developed countries in the best possible
OMICS Journals are welcoming Submissions OMICS International welcomes submissions that are original and technically so as to serve both the developing world and developed countries in the best possible
Phylogenetic Tree Reconstruction
 I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
FIG S1: Plot of rarefaction curves for Borrelia garinii Clp A allelic richness for questing Ixodes
 FIG S1: Plot of rarefaction curves for Borrelia garinii Clp A allelic richness for questing Ixodes ricinus sampled in continental Europe, England, Scotland and grey squirrels (Sciurus carolinensis) from
FIG S1: Plot of rarefaction curves for Borrelia garinii Clp A allelic richness for questing Ixodes ricinus sampled in continental Europe, England, Scotland and grey squirrels (Sciurus carolinensis) from
The Effect of Ambiguous Data on Phylogenetic Estimates Obtained by Maximum Likelihood and Bayesian Inference
 Syst. Biol. 58(1):130 145, 2009 Copyright c Society of Systematic Biologists DOI:10.1093/sysbio/syp017 Advance Access publication on May 21, 2009 The Effect of Ambiguous Data on Phylogenetic Estimates
Syst. Biol. 58(1):130 145, 2009 Copyright c Society of Systematic Biologists DOI:10.1093/sysbio/syp017 Advance Access publication on May 21, 2009 The Effect of Ambiguous Data on Phylogenetic Estimates
