DNA barcode-based delineation of putative species: efficient start for taxonomic workflows
|
|
|
- Ashlie Miles
- 6 years ago
- Views:
Transcription
1 Molecular Ecology Resources (2014) 14, doi: / DNA barcode-based delineation of putative species: efficient start for taxonomic workflows MARI KEKKONEN* and PAUL D. N. HEBERT *Zoology Unit, Finnish Museum of Natural History, University of Helsinki, P.O. Box 17, FI-00014, Helsinki, Finland, Biodiversity Institute of Ontario, University of Guelph, Guelph, ON N1G 2W1, Canada Abstract The analysis of DNA barcode sequences with varying techniques for cluster recognition provides an efficient approach for recognizing putative species (operational taxonomic units, OTUs). This approach accelerates and improves taxonomic workflows by exposing cryptic species and decreasing the risk of synonymy. This study tested the congruence of OTUs resulting from the application of three analytical methods (ABGD, BIN, GMYC) to sequence data for Australian hypertrophine moths. OTUs supported by all three approaches were viewed as robust, but 20% of the OTUs were only recognized by one or two of the methods. These OTUs were examined for three criteria to clarify their status. Monophyly and diagnostic nucleotides were both uninformative, but information on ranges was useful as sympatric sister OTUs were viewed as distinct, while allopatric OTUs were merged. This approach revealed 124 OTUs of Hypertrophinae, a more than twofold increase from the currently recognized 51 species. Because this analytical protocol is both fast and repeatable, it provides a valuable tool for establishing a basic understanding of species boundaries that can be validated with subsequent studies. Keywords: Australia, Automatic Barcode Gap Discovery, Barcode Index Number, DNA barcoding, General Mixed Yulecoalescent, Hypertrophinae Received 27 September 2013; revision received 2 January 2014; accepted 18 January 2014 Introduction Species delimitation studies have traditionally focused on fine-tuning problematic es by compiling varied types of data (e.g. DNA sequences, morphological characters, karyotypes) and examining multiple individuals of each species. Although analyses of this type are appropriate for well-studied groups such as European butterflies (e.g. Dinca et al. 2011), baseline knowledge is much less for many taxonomic assemblages (Common 1990; Raven & Yeates 2007). As a result, there is a critical need for an approach which enables the simultaneous analysis of large numbers of putative species, even if it delivers a less precise outcome. Prior work has shown that preliminary species delineation can often be achieved by analysing single-locus data from suitable genomic regions, such as the 648bp region of the mitochondrial cytochrome c oxidase subunit I selected as DNA barcodes (Hebert et al. 2003a,b; Hausmann et al. 2011; Collins et al. 2012a; Magnacca & Brown 2012). Because the use of any mtdna marker Correspondence: Mari Kekkonen, Fax: ; mari.kekkonen@helsinki.fi risks exposure to complications such as introgression and incomplete lineage sorting, particularly for closely related species (Funk & Omland 2003; Dupuis et al. 2012; Talavera et al. 2013), sequence clusters revealed by the analysis of single-locus data should be considered as operational taxonomic units (OTUs). DNA barcodebased delimitation of species is best viewed as a quick start for the taxonomic process. Several analytical methods support species delineation with single-locus data, partitioning sequences into genetic clusters without adopting a rigid sequence threshold. One popular approach, the General Mixed Yule-coalescent (GMYC; Pons et al. 2006; Fujisawa & Barraclough 2013), takes advantage of both Yule s (1924) and Kingman s (1982) models for calculating the maximum-likelihood solution for the transition point between the speciation and coalescence processes on an ultrametric gene tree. Under GMYC, the number of OTUs (putative species) equals the number of lineages crossing the threshold line. Although Monaghan et al. (2009) modified the original single-threshold model to incorporate variable threshold values throughout a tree, the single-threshold approach is generally preferred (e.g. Brewer et al. 2012; Paz & Crawford 2012). This is an open access article under the terms of the Creative Commons Attribution License, which permits use, distribution and reproduction in any medium, provided the original work is properly cited.
2 DELINEATING SPECIES WITH DNA BARCODES 707 Two methods designed for the analysis of DNA barcode data, Automatic Barcode Gap Discovery (ABGD; Puillandre et al. 2012a) and Barcode Index Number System (BIN; Ratnasingham & Hebert 2013) employ a different approach. Both ABGD and BIN apply clustering algorithms to distinguish partitions in the genetic distances among a group of individuals, using a twophased procedure to create a final array of OTUs. ABGD first divides the data into groups based on a statistically inferred barcode gap and then recursively applies the same procedure to the groups obtained in the first step. By comparison, the BIN approach initially employs single linkage clustering coupled with a 2.2% threshold to establish preliminary OTU boundaries followed by secondary analysis using Markov clustering. The biphasic process has the same goal for both methods: improving and, if needed, redefining groups recovered in the first phase. The congruence among the three methods can be viewed as supporting the robustness of any particular OTU due to their differing analytical approaches and theoretical basis (Carstens et al. 2013). Furthermore, comparison of these methods aids understanding of their tendency to either split or merge clusters. Their performance was contrasted in an earlier study that examined eight data sets covering several taxonomic groups including three well-studied lepidopteran assemblages (Ratnasingham & Hebert 2013). This analysis indicated that the three approaches had similar success in recognizing OTUs that matched known species, but that none delivered perfect correspondence. The results from GMYC and ABGD have been compared in several other studies with general congruence although GMYC tends to deliver a higher OTU count than ABGD, especially as the number of species rises (J orger et al. 2012; Pantaleoni & Badano 2012; Paz & Crawford 2012; Puillandre et al. 2012b; Tang et al. 2012; Hendrixson et al. 2013; Weigand et al. 2013). When these methods have been examined for their capacity to re-cover previously recognized species, the results have been divergent with preference towards GMYC in some cases (Tang et al. 2012) and ABGD in others (Paz & Crawford 2012). If congruence is viewed as a measure of the robustness of any OTU, how should cases of discordance be interpreted? Conservative (Weigand et al. 2013) and minimum consensus (J orger et al. 2012) approaches have been adopted in the past, but both discard much information. Because the proportion of abandoned data will likely increase as the number of species rises (because there will be more chances for mismatches), such approaches are not ideal for large data sets. In this study, we employ three criteria derived from different species concepts to aid a final decision on the status of any controversial OTU: monophyly, diagnostic characters (nucleotide substitutions) and the sympatry of sister OTUs. The inclusion of these parameters reflects their importance as a criterion for one or more species concepts. For example, the phylogenetic species concept (Rosen 1979; Mishler & Donoghue 1982; Donoghue 1985; Mishler 1985) requires that members of a species form a monophyletic unit, motivating our inclusion of this criterion. Another variant of the phylogenetic species concept demands that each species possess diagnostic characters lacking from its sister taxa (Nelson & Platnick 1981; Cracraft 1983; Nixon & Wheeler 1990), justifying our test for such characters. Finally, the biological species concept (Dobzhansky 1937; Mayr 1940; Wright 1940) requires that members of a species comprise a reproductively isolated group, a criterion that can only be tested in nature when species are sympatric (Coyne & Orr 2004). Despite their varied perspectives, different species concepts usually deliver congruent decisions when the taxa being considered have evolved independently for a substantial interval (de Queiroz 2005). This study represents one of the first efforts to use DNA barcode data as a taxonomic exploration tool, grouping specimens into OTUs that can be viewed as the first step towards a framework for subsequent phylogenetic and taxonomic work. The study focuses on the Hypertrophinae, a group of poorly known moths endemic to Australia. We employ a novel combination of methods to reach this goal, examining the congruence of OTUs resulting from three delimitation methods (GMYC, ABGD, BIN). We subsequently evaluate cases of discordance in OTU boundaries employing monophyly, diagnostic characters and sympatry as criteria for clarifying their status (Fig. 1). Materials and methods Sampling The Hypertrophinae was chosen for study due to its high endemism and many undescribed species (Common 1996). With the exception of two species from New Guinea, the group is only known from 51 described species in 12 genera endemic to Australia (Common 1980). The biology and distributions of its component species are very poorly known, and compilation of this information will be constrained until the taxonomy of the group is improved. A total of 864 specimens of Hypertrophinae were analysed, covering all described Australian species, selecting representatives from across the known distribution of each taxon including all biogeographical regions of Australia (Ebach et al. 2013). A large proportion of these specimens were sampled in the Australian National Insect Collection (ANIC) during 2010 and 2011.
3 708 M.KEKKONENandP.D.N.HEBERT Delineation methods Congruence between methods Evaluation of conflicting results BIN ABGD GMYC FULL MATCH PARTIAL MATCH Diagnostics* Monophyly* Sympatry Final OTUs for hypertrophines DISCORDANT Fig. 1 A flowchart describing the protocol, starting with the use of three delineation methods and followed by the division of resultant OTUs into three categories. OTUs assigned to FULL MATCH are included within the final OTU counts, while OTUs in the PARTIAL MATCH and DISCORDANT categories are evaluated against three criteria: sympatry, diagnostic characters, and monophyly (the latter two were tested, but uninformative). Description of FULL MATCH, PARTIAL MATCH, and DISCORDANT categories are provided in the Material and Methods. Additional specimens were analysed from the Agricultural Scientific Collections Unit (ASCU); the Australian Museum, Sydney (AMS); the Biodiversity Institute of Ontario, University of Guelph (BIOUG); the Finnish Museum of Natural History, University of Helsinki (MZH), and the private collections of Graeme Cocks and Doug Hilton. Identifications follow original species descriptions (listed in Appendix S1, Supporting information) and taxonomic assignments for specimens in ANIC, mainly reflecting curatorial activity by Ian Common. No type specimens were examined. DNA extraction, PCR amplification and sequencing DNA extraction, PCR and sequencing were performed at the Canadian Centre for DNA Barcoding following standard high-throughput protocols (dewaard et al. 2008). The first round of PCR employed the primers LepF1 and LepR1 (Hebert et al. 2003a) which generate a 658bp amplicon that spans the barcode region of CO1. In cases of failure, two additional PCR reactions were carried out to re-cover 306bp amplicon and 407bp amplicon using a standard primer set (Hajibabaei et al. 2006). If one of these reactions was successful, an effort was made to obtain a barcode compliant record (>497bp) by amplifying shorter regions of CO1 using the primer sets described in Hebert et al. (2013). All sequences were aligned using the BOLD Aligner in the Barcode of Life Data Systems (BOLD; Ratnasingham & Hebert 2007) and then inspected visually for stop codons and frameshift mutations in MEGA 5 (Tamura et al. 2011). Data analyses Sequences were automatically assigned to a BIN on the BOLD Workbench v3.6 ( analyses performed on 9 May 2013 and repeated on 8 December 2013) where assignments are easily visualized using the Taxon ID Tree. ABGD analyses were performed at the web interface ( jussieu.fr/public/abgd/, web version April , performed on 31 August 2012, repeated on 6 December 2013; source code for ABGD is provided in Appendix S2, Supporting information) using a default value of relative gap width (X = 1.5) and both available distance metrics [JC69 (Jukes & Cantor 1969), K2P (Kimura 1980)] together with p-distance. All assignments for intraspecific divergence (P) values between and were recorded, while other parameter values employed defaults. The General Mixed Yule-coalescent (GMYC) method requires a fully resolved ultrametric gene tree as input for the analysis. We constructed a Bayesian inference tree in BEAST (Drummond et al. 2006; Drummond & Rambaut 2007) employing a Yule pure birth model (Gernhard 2008) tree prior. XML file (Appendix S3, Supporting information) was made with BEAUti v1.7.1 interface with the following settings: GTR+G+I substitution model, empirical base frequencies, four gamma categories, all codon positions partitioned with unlinked base frequencies and substitution rates. An uncorrelated relaxed lognormal clock model was used with rate estimated from the data and ucldmean parameter with uniform prior to value 0 as a lower and 10 as an upper boundary. All other settings were left as defaults. The length of MCMC chain was sampling every All BEAST runs were executed in Bioportal (Kumar et al. 2009), and the ESS values and trace files of runs were evaluated in Tracer v Two independent runs were merged using Log- Combiner v1.7.1 with 20% burn-in. Maximum clade credibility trees with a 0.5 posterior probability limit, and node heights of target tree were constructed in TreeAnnotator v Both single- and multiple-threshold GMYC analyses were conducted in R (R Core Team. 2012) using the APE (Paradis et al. 2004) and SPLITS (Ezard et al. 2009) packages (for R code used for GMYC analyses, see Appendix S4, Supporting information). GMYC analyses were performed with haplotype data collapsed in ALTER (Glez-Pe~na et al. 2010, performed on 11 December 2012). Maximum-likelihood analysis was also performed with haplotype data to compare the results of Bayesian inference and maximum likelihood using RAxML BlackBox (Stamatakis et al. 2008,
4 DELINEATING SPECIES WITH DNA BARCODES 709 performed on 12 May 2013) with GTR+G model and default bootstrap settings. Comparison of resulting OTUs The congruence of the three species delimitation methods was evaluated by comparing the composition of the clusters recognized by each method. To aid comparison, the OTUs were divided into three categories: FULL MATCH where all methods generated the same partition, PARTIAL MATCH where two of three methods generated similar results and DISCORDANT where all three led to a different result. OTUs in the PARTIAL MATCH and DISCORDANT categories were analysed for diagnostic characters between sister OTUs based on application of the phylogenetic species concept using function nucdiag in the R package SPIDER (Brown et al. 2012). This function only considers pure diagnostic characters sensu Sarkar et al. (2008). Although a search for diagnostic characters was conducted for all clusters, its validity is questionable for clusters with few representatives. In addition, monophyly over a NJ tree was studied with the function monophyly in SPIDER. To evaluate putative species status from the context of the biological species concept, we compared the range for members of each distinct OTU based on the coordinates for these specimens in BOLD. Sister OTUs were considered as sympatric when they occupied the same biogeographical region (terrestrial zoogeographical subregions in Ebach et al. 2013). Both range comparison and the search for diagnostic characters were conducted for pairs of sister taxa based on the topology of the Bayesian inference tree. For range estimation, all barcode compliant sequences of hypertrophines in BOLD were included for OTUs in the PARTIAL MATCH and DISCORDANT categories. Results Sequence data were recovered from 702 of the 864 specimens, but some records from older specimens were incomplete. The collection year of successfully sampled specimens varied from 1958 to 2012, but most specimens were collected in the last decade (Fig. S5, Supporting information). Subsequent analysis of OTU diversity focused on 502 full-length (654bp as the BOLD aligner reduces the original length of 658bp by omitting the first and three last bases) barcode sequences which included 294 haplotypes. These records provided coverage for 47 of the 51 known hypertrophine species from Australia (Oxytropha ametalla, Thudaca cymatistis, T. monolechria and T. ophiosema lacked coverage). The sequences used here are publicly available on BOLD and GenBank (see Table S6, Supporting information for Accession nos; DOI: dx.doi.org/ /DS-HOTUS). We only used full-length sequences to remove complications introduced by missing data. Overall pairwise distances (K2P) indicated a clear barcode gap between 0.01 and 0.05 (n = , mean = 0.104) (Fig. S7, Supporting information). A comparison between Bayesian inference (Fig. 2) and maximum-likelihood (Fig. S8, Supporting information) gene trees did not reveal obvious differences in the OTUs. The count of OTUs varied from 73 to 222 with both the lowest and highest result produced by ABGD (Fig. 3). ABGD analysis with JC69 produced two initial partitions with OTU counts of 73 (P = ) and 83 (P = ), whereas use of K2P returned only one initial value of 127 OTUs (P = ) (Table 1). Because the use of p-distance produced strongly discordant outcomes with the initial partition including 140 OTUs (P = ) and 177 (P = ) OTUs (Table 1), it was omitted. BIN (120 OTUs) and GMYC (123 OTUs) with a single-threshold model produced very similar results, and values close to the 127 OTUs obtained with ABGD and the initial partition of K2P. Similar to many earlier studies, the implementation of GMYC with a multiple-threshold model produced a higher OTU count (139) than the single-threshold model, but it failed to improve the fit of the GMYC model to the data (v 2 = 12.73, d.f. = 15, P < 0.62) (Table 2). Also, the likelihood-ratio test rejected the null model denoting the presence of more than one species in the data (Table 2). To examine the congruence of putative species, we assigned each cluster into one of three categories (FULL MATCH, PARTIAL MATCH, DISCORDANT). In making these assignments, we only considered results from the initial partition with K2P from ABGD as the OTU count was closest to those from the other methods. The results obtained with JC69 were excluded due to the extensive merging of clusters which was in strong conflict with the results from the other two methods. From the two GMYC analyses, we only included the single threshold for the above-mentioned reason. Comparison of the assignments showed that 96 OTUs (80%) were recognized by all three methods (i.e. FULL MATCH). Another 22 OTUs (18.3%) were PARTIAL MATCHES, while only two OTUs (1.7%) were DISCORDANT (splits within Hypertropha tortriciformis and Callizyga dispar) (Fig. 2). Diagnostic characters were discovered for all different OTU boundaries within the PARTIAL MATCH and DISCORDANT categories, although this outcome was undoubtedly due, at least in part, to the fact that most conflicting OTUs were represented by few individuals. The test for monophyly revealed that the OTUs delimited by ABGD and BIN each included one paraphyletic group (Eupselia sp. ANICMK238 of beatella ), whereas two groups were paraphyletic with GMYC
5 710 M.KEKKONENandP.D.N.HEBERT GMYC ABGD BIN FINAL Callizyga dispar D Eupselia sp. ANICMK106 Acraephnes litodes P Acraephnes sp. ANICMK30 Acraephnes nivea Acraephnes cryeropis Eupselia metabola Eupselia sp. ANICMK41 Eupselia sp. ANICMK222 Eupselia sp. ANICMK42 Thudaca campylota/ T. heterastis Eupselia sp. ANICMK9 Eupselia sp. ANICMK216 Eupselia sp. ANICMK211 Eupselia sp. ANICMK212 Eupselia sp. ANICMK213 Eupselia sp. ANICMK209 Thudaca obliquella Eupselia sp. ANICMK206 Eupselia sp. ANICMK207 Eupselia sp. ANICMK210 Eupselia sp. ANICMK215 carpocapsella Eupselia sp. ANICMK214 Thudaca haplonota Eupselia sp. ANICMK208 P Thudaca sp. ANICMK24 Thudaca sp. ANICMK22 Thudaca sp. ANICMK231 Acraephnes innubila Thudaca calliphrontis Eupselia sp. ANICMK238 P Eupselia sp. ANICMK237 P Eupselia sp. ANICIC5 beatella Acraephnes sp. ANICMK2 Thudaca circumdatella Thudaca trabeata Thudaca sp. ANICMK234 Thudaca sp. ANICMK235 Thudaca sp. ANICMK232 Thudaca sp. ANICMK236 Acraephnes sp. ANICMK200 Thudaca stadiaula Thudaca sp. ANICMK233 Thudaca orthodroma Acraephnes inscripta Eupselia hypsichora Eupselia sp. ANICMK59 Eupselia sp. ANICMK220 P Epithetica typhoscia Eupselia axiepaena P Eupselia sp. ANICMK202 Eupselia sp. ANICIC7 Eupselia sp. ANICMK221 P Eupselia sp. ANICMK134 Eupselia sp. ANICIC53 Eupselia sp. ANICMK45 Eupselia leucaspis belterairidizonasatrapella belterairidizonasatrapella Acraephnes sp. ANICMK1 Acraephnes nitida Acraephnes sulfurata Eupselia philomorpha P Eupselia sp. ANICMK204 Eupselia sp. ANICMK224 P Thudaca monolinea Eupselia sp. ANICMK62 P Eupselia sp. ANICMK18 Eupselia sp. ANICMK19 Eupselia callidyas belterairidizonasatrapella Eupselia sp. ANICMK205 Eupselia sp. ANICMK105 P Eupselia sp. ANICMK223 P Eupselia sp. ANICMK48 Eupselia sp. ANICMK203 P Eupselia sp. ANICMK219 belterairidizonasatrapella Eupselia sp. ANICMK201 Eupselia sp. ANICIC32 Eupselia aristonica Eupselia sp. ANICMK43 Eupselia sp. ANICMK228 Eupselia sp. ANICMK7 P Eupselia tristephana Eupselia sp. ANICMK44 P Progonica rhothias Eupselia sp. ANICMK12 Eupselia sp. ANICMK49 Eupselia sp. ANICMK46 melanostreptatheorella Hypertropha chlaenota P Eupselia sp. ANICIC70 Eupselia holoxantha Eupselia sp. ANICMK218 Hypertropha sp. ANICMK20 P Eupselia sp. ANICMK217 Eupselia sp. ANICMK40 holoxanthasyncapna Eupselia sp. ANICMK51 Eupselia sp. ANICMK52 Eupselia sp. ANICMK55 Eupselia sp. ANICMK225 Eupselia sp. ANICIC59 Eupselia anommata Eupselia sp. ANICMK11 Eupselia sp. ANICMK58 Eupselia sp. ANICMK14 Eupselia sp. ANICMK15 melanostreptatheorella Eupselia sp. ANICMK61 Eupselia sp. ANICMK60 Eupselia sp. ANICMK13 Eupselia sp. ANICIC24 Hypertropha tortriciformis D Eupselia sp. ANICMK226 Eupselia sp. ANICIC61 Eupselia sp. ANICMK227 holoxanthasyncapna Hypertropha sp. ANICMK229 Hypertropha desumptana Hypertropha thesaurella P Hypertropha sp.anicmk38p Peritropha sp. ANICMK230 Eomystis rhodopis P Eomystis sp. ANICMK35 P Peritropha oligodrachma Fig. 2 Bayesian inference gene tree with delineated OTUs. OTUs in the PARTIAL MATCH and DISCORDANT categories are marked with red letters P and D, respectively. (Eupselia sp. ANICMK105 of beltera-satrapella-iridizona and a split from Hypertropha tortriciformis). The two OTUs (Hypertropha tortriciformis and Callizyga dispar) in the DISCORDANT category may well include more than one species, but each was treated as a single OTU due to the conflicting results. The status of the 22 OTUs in the PARTIAL MATCH category was evaluated by examining the sympatry criterion for sister groups (Fig. 4) similar to the integrative taxonomic approach (ITAX) of Miralles & Vences (2013). Eight of the OTUs partitioned by one of the methods failed to meet the sympatry criterion. Three of these eight OTUs included allopatric subgroups (i.e. restricted to different biogeographical regions), while five other OTUs were represented by a single specimen allopatric from a sister OTU composed of multiple specimens. All eight of these PARTIAL MATCHES were treated as a single OTU on the conservative presumption that the sequence divergence apparent between their allopatric lineages reflected phylogeographic variation in a single taxon. The remaining 14 PARTIAL MATCHES involved cases of sister OTUs which occurred in sympatry, so they were recognized as distinct (Eomystis rhodopis and Eomystis sp. ANICMK35; Eupselia sp. ANICMK237 and E. sp.
6 DELINEATING SPECIES WITH DNA BARCODES 711 OTU count JC Init.JC Init. JC JC JC JC K2P Init Single Mult K2P K2P JC JC Simp. Init Simp. K2P K2P Simp. Init. ABGD BIN GMYC ABGD GMYC ABGD Delimitation method JC Simp. Simp. K2P Simp. Fig. 3 OTU counts resulting from three delineation methods. Figures below the results for ABGD indicate prior intraspecific divergence (P) values. The two OTU counts for GMYC result from single- and multiple-threshold models. Table 1 Results of the Automatic Barcode Gap Discovery (ABGD) analyses Prior intraspecific divergence (P) Subst. model X Partition Simple 1.5 Initial Recursive JC 1.5 Initial Recursive K2P 1.5 Initial Recursive X, relative gap width; Simple, p-distance; JC69, Jukes-Cantor substitution model; K2P, Kimura 2-parameter substitution model. Table 2 Results of the General Mixed Yule-coalescent (GMYC) analyses Analysis Clusters (CI) Entities (CI) Likelihood null Likelihood GMYC Likelihood ratio Threshold Single 76 (75 77) 123 ( ) *** Multiple 70 (70 72) 139 ( ) *** Clusters, OTUs delineated by GMYC with more than one specimen; Entities, singleton OTUs delineated by GMYC; CI, confidence interval; Likelihood null, likelihood of the null model; Likelihood GMYC, likelihood of the GMYC model; Threshold, the threshold between speciation and coalescence processes; Single, single-threshold model; Multiple, multiple-threshold model; ***P < ANICMK238 of beatella ; E. sp. ANICMK208 and E. sp. ANICMK104 of carpocapsella ; E. sp. ANICMK105 and E. sp. ANICMK223 of E. satrapella- E. beltera-e. iridizona; E. sp. ANICMK44 and E. sp. ANICMK7; Hypertropha chlaenota and H. sp. ANICMK20; H. thesaurella and H. sp. ANICMK38). A final count of 120 putative species was obtained by recognizing FULL MATCH clusters (96) as distinct OTUs, and augmenting this total with those from the PARTIAL MATCH (3 + 5 allopatric, 14 sympatric) and DISCORDANT (2) categories. Discussion Estimating the number of Australian species of Hypertrophinae The 502 specimens of Hypertrophinae examined in this study include 120 OTUs that are likely to represent distinct species. Because four known, morphologically distinctive species were not included in our study, the probable species count is at least 124, a more than two-fold increase from the currently recognized fauna.
7 712 M.KEKKONENandP.D.N.HEBERT Allopatric Sympatric Region A Region B Region A Region B 1 final OTU 2 final OTUs Because nearly 50% of the OTUs in this study were represented by just one or two individuals, it is likely that many additional taxa await discovery. An accumulation curve for OTUs (Fig. S9, Supporting information) shows reduced steepness, but indicates the likely presence of additional species. However, based on current results, it is already clear that two closely related genera, Allotropha and Eupselia, will rise in diversity (19 current species vs. 78 OTUs). While Eupselia carpocapsella provides a particularly striking example of cryptic species with 12 OTUs, four other lineages (Allotropha percussana-eupselia aristonica, E. holoxantha-e. syncapna, E. satrapella-e. beltera-e. iridizona and E. melanostrepta-e. theorella) also likely form multispecies es. Interestingly, many of these es show polyphyly in the Bayesian gene tree (Fig. 2), a result which might be an artefact of the gene tree, but the situation certainly calls for further investigation. Two other genera also appear to include unrecognized species with Acraephnes rising from 7 to 11 OTUs and Thudaca from 15 to 20 OTUs (Thudaca crypsidesma and T. mimodora were probably analysed, but none of the OTUs was assigned to these species because they lack clear morphological diagnostics). Thudaca heterastis showed no sequence difference from T. campylota, so these taxa may be synonyms and were treated as one OTU. OTUs were also added to Hypertropha (from 4 to 7), Peritropha (from 1 to 2) and Eomystis (from 1 to 2). No evidence of unrecognized species was obtained in the other four genera (Callizyga, Epithetica, Oxytropha and Progonica). Comparing the performance of OTU delineation methods OTU1 OTU2 Fig. 4 Sympatry criterion for sister OTUs in PARTIAL MATCH category. OTUs found in separate biogeographical regions are merged to form one final OTU while sister OTUs sympatric in one or several regions are recognized as two final OTUs. ABGD, GMYC and BIN showed good concordance with the same assignment for 80% of OTUs, supporting earlier studies (Puillandre et al. 2012b; Ratnasingham & Hebert 2013). However, congruence would have been much lower if other outcomes of ABGD were included. For example, the initial partition with JC69 merged many clusters, while recursive partitions with JC69 and K2P created many splits. This difference between distance metrics is strongly discordant from the results obtained by Collins et al. (2012b), a situation requiring further investigation. We adopted the initial partition of ABGD with K2P due to its closer correspondence with the results from BIN and GMYC, simplifying the comparison. Although recursive partitions of ABGD were excluded from the correspondence check, they revealed subgroups which may be useful in certain taxonomic contexts. Automatic Barcode Gap Discovery generates diverse outcomes, and it is difficult to select the most appropriate one. Puillandre et al. (2012a) proposed adoption of a single value of P = 0.01 as it produced the strongest congruence with previous studies examining the same data with different approaches. In our analysis, this value was only produced by JC69, but the OTUs with this distance metric showed strong discordance to those obtained with the other methods. Because we selected the outcome from ABGD which delivered the closest OTU count to the other two methods, our test for the robustness of OTU boundaries (i.e. all methods assigning particular specimens to the same OTU) is partially compromised. However, it needs emphasis that the overall OTU count and the specimens composing each OTU are not strictly associated. As results from the three approaches diverged in 20% of all OTUs, they certainly provide some insights into the stability of the OTUs. However, more investigations are needed to strengthen the use of ABGD so that the adoption of a particular value of P is made without the a posteriori approach employed in this study. General Mixed Yule-coalescent has a strong theoretical basis, but it typically generates more OTUs than other methods (Esselstyn et al. 2012; Paz & Crawford 2012; Sauer & Hausdorf 2012; Miralles & Vences 2013; Talavera et al. 2013) and errors in the ultrametric gene tree that underpins the analysis will influence final results. In addition, GMYC calculations are very timeconsuming for large data sets due to their requirement for an input tree (the multiple-threshold model is a particular challenge). BIN is the fastest and most userfriendly of the methods as it delivers only one result, making clear the OTU boundaries which need evaluation. All three methods have the tendency to split outliers, but, as indicated above, we treated these as a probable artefact of geographical distance and not as reflective of a species boundary. Employing three analytical approaches improves confidence in the validity of OTUs delineated by all
8 DELINEATING SPECIES WITH DNA BARCODES 713 approaches, although Carstens et al. (2013) encourage using even more methods. Conflicting results can be viewed as indicators for OTU boundaries which deserve detailed inspection. The use of several methods does have one disadvantage; it increases the ity and time required for OTU evaluation. Adequate sample sizes are critical for any effort to delineate species (e.g. Lohse 2009). If the current species count (51) for Australian Hypertrophinae was complete, our analysis of 502 specimens would have provided nearly 109 coverage if each taxon had equal representation. However, 46% of the OTUs (55 of 120) in our analyses were represented by just one or two specimens, reflecting the commonness of rarity in nature (Lim et al. 2012). This fact emphasizes the need for analytical methods that deal effectively with low taxon coverage. Because simulation studies indicate that ABGD performs poorly unless there are 3 5 samples per species (Puillandre et al. 2012a), its use for explorations of species diversity in poorly known groups is problematic, because the number of samples per species is impossible to know a priori. Interestingly, despite this limitation, one analytical option of ABGD generated results that were relatively congruent with other methods despite the low numbers of specimens. Criteria for discordant OTU boundaries To be useful, each test criterion requires differences between OTUs assigned to the PARTIAL MATCH and DISCORDANT categories. For example, cases of monophyly or the presence of diagnostic characters would support the validity on a controversial OTU, while the detection of paraphyly or the lack of diagnostic characters would not. Because all discordant OTUs in our study possessed diagnostic nucleotide substitutions, this criterion did not help to clarify their status. This criterion may be useful in other situations, but its utility will often be compromised by the rarity of many taxa. The test for monophyly revealed few cases of paraphyly, so it was also of little value in clarifying OTU boundaries. Apparently, the three delimitation methods typically recognize breaks in sequence space associated with monophyly, so secondary inspection reveals few exceptions. By contrast, the sympatry criterion provided a useful tool for the evaluation of conflicts in OTU boundaries. When two allopatric populations are only assigned to distinct OTUs by certain methods, their status as distinct species becomes questionable (e.g. Mutanen et al. 2012). By comparison, when sister OTUs show range overlap, this provides presumptive evidence for their reproductive isolation although it should be confirmed by nuclear markers. We add two provisos. Because sympatry was imprecisely evaluated in this study, sister OTUs viewed as sympatric may actually be microallopatric. Conversely, because sampling efforts were not comprehensive, future sampling may reveal that sister OTUs currently viewed as allopatric are actually sympatric. Delineating species with DNA barcodes This study describes an efficient protocol for obtaining an initial taxonomic framework. Puillandre et al. (2012b) adopted a more strategy for delimiting species of marine molluscs which coupled testing initial OTUs (primary species hypotheses) for differences in morphology, sequence divergence at additional loci and the dispersal capacity of larvae before creating secondary species hypotheses. Riedel et al. (2013) presented an even more approach covering the whole taxonomic procedure. Our scheme has the advantage of keeping the initial step of OTU designation separate from the detailed analysis required for full-blown taxonomic characterization. Because the varying steps in species delineation require different sampling strategies and types of data, the primary delineation of OTUs with single-locus data has the advantage of employing one extensive data set with clearly defined criteria to produce a stable outcome. We emphasize that the delimitation of putative species based on DNA barcode data not only increases objectivity, but accelerates work on poorly studied groups and enables inexperienced taxonomists to make a valuable contribution. As many groups of arthropods lack expert taxonomists, the need to recruit new experts is obvious and barcode-based approaches provide an easy path for initial engagement. Even without detailed study, an accurate estimate of the species count is obtained through the simple algorithmic processing of barcode data. While decisions based on analysis of single-locus mtdna data and on small sample sizes do pose interpretational risks, they are inconsequential if the outcome is viewed as a scaffold for taxonomy rather than as the sole criterion for species description. Acknowledgements We are extremely grateful to staff and researchers at the Australian National Insect Collection (ANIC), especially Marianne Horak, Ted Edwards, You Ning Su, Beth Mantle, and John LaSalle, for their assistance and advice. This work would have been difficult without the past curatorial efforts of Ian Common on the Hypertrophinae at ANIC. We also thank Graeme Cocks and Ian McMillan for their efforts in collecting some of the specimens analysed. Andrew Mitchell kindly permitted the use of sequences from the Australian Museum and the Agricultural Scientific Collections Unit. Rupert Collins, Lauri Kaila, Marko
9 714 M.KEKKONENandP.D.N.HEBERT Mutanen, Marko Nieminen, and an anonymous referee provided insightful comments on the manuscript. This work was supported by Research Foundation of the University of Helsinki, Finnish Concordia Fund, and the Ella and Georg Ehrnrooth Foundation to MK. Sequence analysis was enabled by funds from the government of Canada through Genome Canada and the Ontario Genomics Institute in support of the International Barcode of Life project, while a NSERC grant to PDNH supported collecting activity. References Brewer MS, Spruill CL, Rao NS, Bond JE (2012) Phylogenetics of the millipede genus Brachycybe Wood 1864 (Diplopoda: Platydesmida: Andrognathidae): patterns of deep evolutionary history and recent speciation. Molecular Phylogenetics and Evolution, 64, Brown SDJ, Collins RA, Boyer S et al. (2012) Spider: an R package for the analysis of species identity and evolution with particular reference to DNA barcoding. Molecular Ecology Resources, 12, Carstens BC, Pelletier TA, Reid NM, Satler JD (2013) How to fail at species delimitation. Molecular Ecology, 22, Collins RA, Armstrong KF, Meier R et al. (2012a) Barcoding and border biosecurity: identifying cyprinid fishes in the aquarium trade. PLoS ONE, 7, e Collins RA, Boykin LM, Cruickshank RH, Armstrong KF (2012b) Barcoding s next top model: an evaluation of nucleotide substitution models for specimen identification. Methods in Ecology and Evolution, 3, Common IFB (1980) The systematic position of Hypertropha (Lepidoptera) and related Australian genera. Entomologica Scandinavica, 11, Common IFB(1990) Moths of Australia. Melbourne University Press, Victoria. Common IFB (1996) Hypertrophidae. In: Checklist of the Lepidoptera of Australia(eds Nielsen ES, Edwards ED & Rangsi TV), pp CSIRO, Melbourne. Coyne JA, Orr HA (2004) Speciation. Sinauer Associates, Sunderland,MA USA. Cracraft J (1983) Species concepts and speciation analysis. Current Ornithology, 1, Dinca V, Lukhtanov VA, Talavera G, Vila R (2011) Unexpected layers of cryptic diversity in wood white Leptidea butterflies. Nature Communications, 2, 324. Dobzhansky T (1937) Genetics and the Origin of Species. Columbia University Press, New York. Donoghue M (1985) A critique of the biological species concept and recommendations for a phylogenetic alternative. Bryologist, 88, Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evolutionary Biology, 7, 214. Drummond AJ, Ho SYW, Phillips MJ, Rambaut A (2006) Relaxed phylogenetics and dating with confidence. PLoS Biology, 4, Dupuis JR, Roe AD, Sperling FAH (2012) Multi-locus species delimitation in closely related animals and fungi: one marker is not enough. Molecular Ecology, 21, Ebach MC, Gill AC, Kwan A, Ahyong ST, Murphy DJ, Cassis G (2013) Towards an Australian bioregionalisation atlas: a provisional area taxonomy of Australia s biogeographical regions. Zootaxa, 3619, Esselstyn JA, Evans BJ, Sedlock JL, Khan FAA, Heaney LR (2012) Singlelocus species delimitation: a test of the mixed Yule-coalescent model with an empirical application to Philippine round-leaf bats. Proceedings of the Royal Society of London series B - Biological Sciences, 279, Ezard T, Fujisawa T, Barraclough TG(2009) splits: SPecies LImits by Threshold Statistics. R package version /r31. Fujisawa T, Barraclough TG (2013) Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: a revised method and evolution on simulated data sets. Systematic Biology, 62, Funk DJ, Omland KE (2003) Species-level paraphyly and polyphyly: frequency, causes, and consequences, with insights from animal mitochondrial DNA. Annual Review of Ecology, Evolution, and Systematics, 34, Gernhard T (2008) The conditioned reconstructed process. Journal of Theoretical Biology, 253, Glez-Pe~na D, Gomez-Blanco D, Reboiro-Jato M, Fdez-Riverola F, Posada D (2010) ALTER: program-oriented conversion of DNA and protein alignments. Nucleic Acids Research, 38, W14 W18. Hajibabaei M, Janzen D, Burns J, Hallwachs W, Hebert PDN (2006) DNA barcodes distinguish species of tropical Lepidoptera. Proceedings of the National Academy of Sciences of the United States of America, 103, Hausmann A, Haszprunar G, Hebert PDN (2011) DNA barcoding the geometrid fauna of Bavaria (Lepidoptera): successes, surprises, and questions. PLoS ONE, 6, e Hebert PDN, Cywinska A, Ball S, dewaard J (2003a) Biological identifications through DNA barcodes. Proceedings of the Royal Society of London series B - Biological Sciences, 270, Hebert PDN, Ratnasingham S, dewaard J (2003b) Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proceedings of the Royal Society of London series B - Biological Sciences, 270, Hebert PDN, dewaard JR, Zakharov EV et al. (2013) A DNA Barcode Blitz : rapid digitization and sequencing of a natural history collection. PLoS ONE, 8, e Hendrixson BE, DeRussy BM, Hamilton CA, Bond JE (2013) An exploration of species boundaries in turret-building tarantulas of the Mojave Desert (Araneae, Mygalomorphae, Theraphosidae, Aphonopelma). Molecular Phylogenetics and Evolution, 66, J orger KM, Norenburg JL, Wilson NG, Schr odl M (2012) Barcoding against a paradox? Combined molecular species delineations reveal multiple cryptic lineages in elusive meiofaunal sea slugs. BMC Evolutionary Biology, 12, 245. Jukes TH, Cantor CR (1969) Evolution of protein molecules. In: Mammalian Protein Metabolism(ed Munro NH), pp Academic Press, New York. Kimura M (1980) A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution, 16, Kingman JFC (1982) The coalescent. Stochastic Processes and their Applications, 13, Kumar S, Skjaeveland A, Orr RJS et al. (2009) AIR: a batch-oriented web program package for construction of supermatrices ready for phylogenomic analyses. BMC Bioinformatics, 10, 357. Lim GS, Balke M, Meier R (2012) Determining species boundaries in a world full of rarity: singletons, species delimitation methods. Systematic Biology, 61, Lohse K (2009) Can mtdna barcodes be used to delimit species? A response to Pons et al (2006) Systematic Biology, 58, Magnacca KN, Brown MJF (2012) DNA barcoding a regional fauna: Irish solitary bees. Molecular Ecology Resources, 12, Mayr E (1940) Speciation phenomena in birds. The American Naturalist, 74, Miralles A, Vences M (2013) New metrics for comparison of taxonomies reveal striking discrepancies among species delimitation methods in Madascincus lizards. PLoS ONE, 8, e Mishler B (1985) The morphological developmental and phylogenetic basis of species concepts in Bryophytes. Bryologist, 88, Mishler B, Donoghue M (1982) Species concepts - a case for pluralism. Systematic Zoology, 31, Monaghan MT, Wild R, Elliot M et al. (2009) Accelerated species inventory on Madagascar using coalescent-based models of species delineation. Systematic Biology, 58, Mutanen M, Hausmann A, Hebert PDN, Landry J-F, dewaard JF, Huemer P (2012) Allopatry as a Gordian Knot for taxonomists:
10 DELINEATING SPECIES WITH DNA BARCODES 715 patterns of DNA barcode divergence in Arctic-Alpine Lepidoptera. PLoS ONE, 7, e Nelson GJ, Platnick NI (1981) Systematics and Biogeography: Cladistics and Vicariance. Columbia University Press, New York. Nixon K, Wheeler Q (1990) An amplification of the phylogenetic species concept. Cladistics - the International Journal of the Willi Hennig Society, 6, Pantaleoni RA, Badano D (2012) Myrmeleon punicanus n. sp., a new pitbuilding antlion (Neuroptera Myrmeleontidae) from Sicily and Pantelleria. Bulletin of Insectology, 65, Paradis E, Claude J, Strimmer K (2004) APE: analyses of phylogenetics and evolution in R language. Bioinformatics, 20, Paz A, Crawford AJ (2012) Molecular-based rapid inventories of sympatric diversity: a comparison of DNA barcode clustering methods applied to geography-based vs clade-based sampling of amphibians. Journal of Biosciences, 37, Pons J, Barraclough TG, Gomez-Zurita J et al. (2006) Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Systematic Biology, 55, Puillandre N, Lambert A, Brouillet S, Achaz G (2012a) ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Molecular Ecology, 21, Puillandre N, Modica MV, Zhang Y et al. (2012b) Large-scale species delimitation method for hyperdiverse groups. Molecular Ecology, 21, de Queiroz K (2005) Different species problems and their resolution. BioEssays, 27, R Core Team. (2012) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. Ratnasingham S, Hebert PDN (2007) BOLD: the Barcode of Life Data System ( Molecular Ecology Notes, 7, Ratnasingham S, Hebert PDN (2013) A DNA-based registry for all animal species: the Barcode Index Number (BIN) system. PLoS ONE, 8, e Raven PH, Yeates DK (2007) Australian biodiversity: threats for the present, opportunities for the future. Australian Journal of Entomology, 46, Riedel A, Sagata K, Suhardjono YR, T anzler R, Balke M (2013) Integrative taxonomy on the fast track towards more sustainability in biodiversity research. Frontiers in Zoology, 10, 15. Rosen DE (1979) Fishes from the uplands and intermontane basins of Guatemala: revisionary studies and comparative geography. Bulletin of the American Museum of Natural History, 162, 5. Sarkar IN, Planet PJ, DeSalle R (2008) CAOS software for use in character-based DNA barcoding. Molecular Ecology Resources, 8, Sauer J, Hausdorf B (2012) A comparison of DNA-based methods for delimiting species in a Cretan land snail radiation reveals shortcomings of exclusively molecular taxonomy. Cladistics - the International Journal of the Willi Hennig Society, 28, Stamatakis A, Hoover P, Rougemont J (2008) A rapid bootstrap algorithm for the RAxML web-servers. Systematic Biology, 75, Talavera G, Dinca V, Vila R (2013) Factors affecting species delimitations with the GMYC model: insights from a butterfly survey. Methods in Ecology and Evolution, 4, Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Molecular Biology and Evolution, 28, Tang CQ, Leasi F, Obertegger U, Kieneke A, Barraclough TG, Fontaneto D (2012) The widely used small subunit 18S rdna molecule greatly underestimates true diversity in biodiversity surveys of the meiofauna. Proceedings of the National Academy of Sciences of the United States of America, 109, dewaard JR, Ivanova NV, Hajibabaei M, Hebert PDN (2008) Assembling DNA Barcodes: analytical Protocols. In: Methods in Molecular Biology: Environmental Genetics(ed Cristofre Martin C), pp Humana Press Inc., Totowa USA. Weigand AM, Jochum A, Slapnik R, Schnitzler J, Zarza E, Klussmann- Kolb A (2013) Evolution of microgastropods (Ellobioidea, Carychiidae): integrating taxonomic, phylogenetic and evolutionary hypotheses. BMC Evolutionary Biology, 13, 18. Wright S (1940) The statistical consequences of Mendelian heredity in relation to speciation. In: The New Systematics(ed Huxley J), pp Oxford University Press, London. Yule GU (1924) A mathematical theory of evolution based on the conclusions of Dr JC Willis FRS. Philosophical Transactions of the Royal Society of London series B-Biological Sciences, 213, M.K. designed the study, sampled most of the specimens at ANIC, analysed the data, and wrote the manuscript. P.D.N.H. was responsible for the sequence analysis and wrote the manuscript. Data Accessibility DNA sequences: GenBank accessions see Table S6, Supporting information. BOLD data set Hypertrophinae OTUs (DS-HOTUS) DOI: Supporting Information Additional Supporting Information may be found in the online version of this article: Appendix S1 The reference list of original species descriptions. Appendix S2 The source code for ABGD. Downloaded from on 6 December Appendix S3 XML file for BEAST. Appendix S4 R code used for GMYC analyses. Fig. S5 The collection years of specimens with barcode sequences (any length). Table S6 Samples used for OTU delineation. OTU names, BOLD Sample and Process IDs, GenBank accessions and storing institutions are provided. Fig. S7 Pairwise distances (K2P) for 502 full-length (654bp) DNA barcode sequences from Australian Hypertrophinae. Fig. S8 Maximum-likelihood tree with bootstrap values. Fig. S9 An accumulation curve of putative species (OTUs).
Factors affecting species delimitations with the GMYC model: insights from a butterfly survey
 Methods in Ecology and Evolution 2013, 4, 1101 1110 doi: 10.1111/2041-210X.12107 Factors affecting species delimitations with the GMYC model: insights from a butterfly survey Gerard Talavera 1 *, Vlad
Methods in Ecology and Evolution 2013, 4, 1101 1110 doi: 10.1111/2041-210X.12107 Factors affecting species delimitations with the GMYC model: insights from a butterfly survey Gerard Talavera 1 *, Vlad
Bayesian species identification under the multispecies coalescent provides significant improvements to DNA barcoding analyses
 Molecular Ecology (2017 26, 3028 3036 doi: 10.1111/mec.14093 Bayesian species identification under the multispecies coalescent provides significant improvements to DNA barcoding analyses ZIHENG YANG* and
Molecular Ecology (2017 26, 3028 3036 doi: 10.1111/mec.14093 Bayesian species identification under the multispecies coalescent provides significant improvements to DNA barcoding analyses ZIHENG YANG* and
A minimalist barcode can identify a specimen whose DNA is degraded
 Molecular Ecology Notes (2006) 6, 959 964 doi: 10.1111/j.1471-8286.2006.01470.x Blackwell Publishing Ltd BARCODING A minimalist barcode can identify a specimen whose DNA is degraded MEHRDAD HAJIBABAEI,*
Molecular Ecology Notes (2006) 6, 959 964 doi: 10.1111/j.1471-8286.2006.01470.x Blackwell Publishing Ltd BARCODING A minimalist barcode can identify a specimen whose DNA is degraded MEHRDAD HAJIBABAEI,*
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley
 Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Integrative Biology 200 "PRINCIPLES OF PHYLOGENETICS" Spring 2018 University of California, Berkeley B.D. Mishler Feb. 14, 2018. Phylogenetic trees VI: Dating in the 21st century: clocks, & calibrations;
Using phylogenetics to estimate species divergence times... Basics and basic issues for Bayesian inference of divergence times (plus some digression)
 Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Using phylogenetics to estimate species divergence times... More accurately... Basics and basic issues for Bayesian inference of divergence times (plus some digression) "A comparison of the structures
Dr. Amira A. AL-Hosary
 Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
Phylogenetic analysis Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic Basics: Biological
8/23/2014. Phylogeny and the Tree of Life
 Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Phylogeny and the Tree of Life Chapter 26 Objectives Explain the following characteristics of the Linnaean system of classification: a. binomial nomenclature b. hierarchical classification List the major
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut
 Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amira A. AL-Hosary PhD of infectious diseases Department of Animal Medicine (Infectious Diseases) Faculty of Veterinary Medicine Assiut University-Egypt Phylogenetic analysis Phylogenetic Basics: Biological
Amy Driskell. Laboratories of Analytical Biology National Museum of Natural History Smithsonian Institution, Wash. DC
 DNA Barcoding Amy Driskell Laboratories of Analytical Biology National Museum of Natural History Smithsonian Institution, Wash. DC 1 Outline 1. Barcoding in general 2. Uses & Examples 3. Barcoding Bocas
DNA Barcoding Amy Driskell Laboratories of Analytical Biology National Museum of Natural History Smithsonian Institution, Wash. DC 1 Outline 1. Barcoding in general 2. Uses & Examples 3. Barcoding Bocas
Species Identification and Barcoding. Brendan Reid Wildlife Conservation Genetics February 9th, 2010
 Species Identification and Barcoding Brendan Reid Wildlife Conservation Genetics February 9th, 2010 Why do we need a genetic method of species identification? Black-knobbed Map Turtle (Graptemys nigrinoda)
Species Identification and Barcoding Brendan Reid Wildlife Conservation Genetics February 9th, 2010 Why do we need a genetic method of species identification? Black-knobbed Map Turtle (Graptemys nigrinoda)
Other matters: * Term paper: 11 (Monday) Nov 15 (Friday), 16, 17? Final * Wednesday, 11 December (8 11 AM)
 Nov 4-Nov 8 Nov 11-15 DNA barcoding Molecular systematics in plants (Les) Lab 8: likelihood (Garli and/or RAxML) and Bayesian analyses (Wagner in Austin) Molecular systematics in animals (Simon) (Wagner
Nov 4-Nov 8 Nov 11-15 DNA barcoding Molecular systematics in plants (Les) Lab 8: likelihood (Garli and/or RAxML) and Bayesian analyses (Wagner in Austin) Molecular systematics in animals (Simon) (Wagner
DNA Barcoding and taxonomy of Glossina
 DNA Barcoding and taxonomy of Glossina Dan Masiga Molecular Biology and Biotechnology Department, icipe & Johnson Ouma Trypanosomiasis Research Centre, KARI The taxonomic problem Following ~250 years of
DNA Barcoding and taxonomy of Glossina Dan Masiga Molecular Biology and Biotechnology Department, icipe & Johnson Ouma Trypanosomiasis Research Centre, KARI The taxonomic problem Following ~250 years of
Reconstructing the history of lineages
 Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
Reconstructing the history of lineages Class outline Systematics Phylogenetic systematics Phylogenetic trees and maps Class outline Definitions Systematics Phylogenetic systematics/cladistics Systematics
How to use CAOS software for taxonomy? quick guide to extract diagnostic nucleotides or amino acids for species descriptions.
 SPIXIANA 37 1 21-26 München, August 2014 ISSN 0341-8391 How to use CAOS software for taxonomy? A quick guide to extract diagnostic nucleotides or amino acids for species descriptions Katharina M. Jörger
SPIXIANA 37 1 21-26 München, August 2014 ISSN 0341-8391 How to use CAOS software for taxonomy? A quick guide to extract diagnostic nucleotides or amino acids for species descriptions Katharina M. Jörger
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley B.D. Mishler Jan. 22, 2009. Trees I. Summary of previous lecture: Hennigian
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2009 University of California, Berkeley B.D. Mishler Jan. 22, 2009. Trees I. Summary of previous lecture: Hennigian
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics
 POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
POPULATION GENETICS Winter 2005 Lecture 17 Molecular phylogenetics - in deriving a phylogeny our goal is simply to reconstruct the historical relationships between a group of taxa. - before we review the
A (short) introduction to phylogenetics
 A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
A (short) introduction to phylogenetics Thibaut Jombart, Marie-Pauline Beugin MRC Centre for Outbreak Analysis and Modelling Imperial College London Genetic data analysis with PR Statistics, Millport Field
How to read and make phylogenetic trees Zuzana Starostová
 How to read and make phylogenetic trees Zuzana Starostová How to make phylogenetic trees? Workflow: obtain DNA sequence quality check sequence alignment calculating genetic distances phylogeny estimation
How to read and make phylogenetic trees Zuzana Starostová How to make phylogenetic trees? Workflow: obtain DNA sequence quality check sequence alignment calculating genetic distances phylogeny estimation
Concepts and Methods in Molecular Divergence Time Estimation
 Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Concepts and Methods in Molecular Divergence Time Estimation 26 November 2012 Prashant P. Sharma American Museum of Natural History Overview 1. Why do we date trees? 2. The molecular clock 3. Local clocks
Molecular Markers, Natural History, and Evolution
 Molecular Markers, Natural History, and Evolution Second Edition JOHN C. AVISE University of Georgia Sinauer Associates, Inc. Publishers Sunderland, Massachusetts Contents PART I Background CHAPTER 1:
Molecular Markers, Natural History, and Evolution Second Edition JOHN C. AVISE University of Georgia Sinauer Associates, Inc. Publishers Sunderland, Massachusetts Contents PART I Background CHAPTER 1:
SPECIATION. REPRODUCTIVE BARRIERS PREZYGOTIC: Barriers that prevent fertilization. Habitat isolation Populations can t get together
 SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
SPECIATION Origin of new species=speciation -Process by which one species splits into two or more species, accounts for both the unity and diversity of life SPECIES BIOLOGICAL CONCEPT Population or groups
Chapter 19: Taxonomy, Systematics, and Phylogeny
 Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
Chapter 19: Taxonomy, Systematics, and Phylogeny AP Curriculum Alignment Chapter 19 expands on the topics of phylogenies and cladograms, which are important to Big Idea 1. In order for students to understand
PHYLOGENY AND SYSTEMATICS
 AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
AP BIOLOGY EVOLUTION/HEREDITY UNIT Unit 1 Part 11 Chapter 26 Activity #15 NAME DATE PERIOD PHYLOGENY AND SYSTEMATICS PHYLOGENY Evolutionary history of species or group of related species SYSTEMATICS Study
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks!
 Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
Some of these slides have been borrowed from Dr. Paul Lewis, Dr. Joe Felsenstein. Thanks! Paul has many great tools for teaching phylogenetics at his web site: http://hydrodictyon.eeb.uconn.edu/people/plewis
DNA-based species delimitation
 DNA-based species delimitation Phylogenetic species concept based on tree topologies Ø How to set species boundaries? Ø Automatic species delimitation? druhů? DNA barcoding Species boundaries recognized
DNA-based species delimitation Phylogenetic species concept based on tree topologies Ø How to set species boundaries? Ø Automatic species delimitation? druhů? DNA barcoding Species boundaries recognized
CHAPTERS 24-25: Evidence for Evolution and Phylogeny
 CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
CHAPTERS 24-25: Evidence for Evolution and Phylogeny 1. For each of the following, indicate how it is used as evidence of evolution by natural selection or shown as an evolutionary trend: a. Paleontology
Taming the Beast Workshop
 Workshop and Chi Zhang June 28, 2016 1 / 19 Species tree Species tree the phylogeny representing the relationships among a group of species Figure adapted from [Rogers and Gibbs, 2014] Gene tree the phylogeny
Workshop and Chi Zhang June 28, 2016 1 / 19 Species tree Species tree the phylogeny representing the relationships among a group of species Figure adapted from [Rogers and Gibbs, 2014] Gene tree the phylogeny
Intraspecific gene genealogies: trees grafting into networks
 Intraspecific gene genealogies: trees grafting into networks by David Posada & Keith A. Crandall Kessy Abarenkov Tartu, 2004 Article describes: Population genetics principles Intraspecific genetic variation
Intraspecific gene genealogies: trees grafting into networks by David Posada & Keith A. Crandall Kessy Abarenkov Tartu, 2004 Article describes: Population genetics principles Intraspecific genetic variation
A preliminary framework for DNA barcoding, incorporating the multispecies coalescent
 University of Wollongong Research Online Faculty of Science, Medicine and Health - Papers Faculty of Science, Medicine and Health 2014 A preliminary framework for DNA barcoding, incorporating the multispecies
University of Wollongong Research Online Faculty of Science, Medicine and Health - Papers Faculty of Science, Medicine and Health 2014 A preliminary framework for DNA barcoding, incorporating the multispecies
Cover Page. The handle holds various files of this Leiden University dissertation.
 Cover Page The handle http://hdl.handle.net/1887/65602 holds various files of this Leiden University dissertation. Author: Ruchisansakun, S. Title: Balsaminaceae in Southeast Asia: systematics, evolution,
Cover Page The handle http://hdl.handle.net/1887/65602 holds various files of this Leiden University dissertation. Author: Ruchisansakun, S. Title: Balsaminaceae in Southeast Asia: systematics, evolution,
Constructing Evolutionary/Phylogenetic Trees
 Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Constructing Evolutionary/Phylogenetic Trees 2 broad categories: istance-based methods Ultrametric Additive: UPGMA Transformed istance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Speciation. Today s OUTLINE: Mechanisms of Speciation. Mechanisms of Speciation. Geographic Models of speciation. (1) Mechanisms of Speciation
 Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Phylogenetic diversity and conservation
 Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Phylogenetic diversity and conservation Dan Faith The Australian Museum Applied ecology and human dimensions in biological conservation Biota Program/ FAPESP Nov. 9-10, 2009 BioGENESIS Providing an evolutionary
Speciation. Today s OUTLINE: Mechanisms of Speciation. Mechanisms of Speciation. Geographic Models of speciation. (1) Mechanisms of Speciation
 Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Introduction to Phylogenetic Analysis
 Introduction to Phylogenetic Analysis Tuesday 24 Wednesday 25 July, 2012 School of Biological Sciences Overview This free workshop provides an introduction to phylogenetic analysis, with a focus on the
Introduction to Phylogenetic Analysis Tuesday 24 Wednesday 25 July, 2012 School of Biological Sciences Overview This free workshop provides an introduction to phylogenetic analysis, with a focus on the
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization)
 STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
STEM-hy: Species Tree Estimation using Maximum likelihood (with hybridization) Laura Salter Kubatko Departments of Statistics and Evolution, Ecology, and Organismal Biology The Ohio State University kubatko.2@osu.edu
Speciation. Today s OUTLINE: Mechanisms of Speciation. Mechanisms of Speciation. Geographic Models of speciation. (1) Mechanisms of Speciation
 Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
Speciation Today s OUTLINE: (1) Geographic Mechanisms of Speciation (What circumstances lead to the formation of new species?) (2) Species Concepts (How are Species Defined?) Mechanisms of Speciation Last
UoN, CAS, DBSC BIOL102 lecture notes by: Dr. Mustafa A. Mansi. The Phylogenetic Systematics (Phylogeny and Systematics)
 - Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
- Phylogeny? - Systematics? The Phylogenetic Systematics (Phylogeny and Systematics) - Phylogenetic systematics? Connection between phylogeny and classification. - Phylogenetic systematics informs the
ESS 345 Ichthyology. Systematic Ichthyology Part II Not in Book
 ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
ESS 345 Ichthyology Systematic Ichthyology Part II Not in Book Thought for today: Now, here, you see, it takes all the running you can do, to keep in the same place. If you want to get somewhere else,
Anatomy of a species tree
 Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Anatomy of a species tree T 1 Size of current and ancestral Populations (N) N Confidence in branches of species tree t/2n = 1 coalescent unit T 2 Branch lengths and divergence times of species & populations
Lecture 11 Friday, October 21, 2011
 Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
Lecture 11 Friday, October 21, 2011 Phylogenetic tree (phylogeny) Darwin and classification: In the Origin, Darwin said that descent from a common ancestral species could explain why the Linnaean system
C3020 Molecular Evolution. Exercises #3: Phylogenetics
 C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
C3020 Molecular Evolution Exercises #3: Phylogenetics Consider the following sequences for five taxa 1-5 and the known outgroup O, which has the ancestral states (note that sequence 3 has changed from
Algorithms in Bioinformatics
 Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Algorithms in Bioinformatics Sami Khuri Department of Computer Science San José State University San José, California, USA khuri@cs.sjsu.edu www.cs.sjsu.edu/faculty/khuri Distance Methods Character Methods
Letter to the Editor. Department of Biology, Arizona State University
 Letter to the Editor Traditional Phylogenetic Reconstruction Methods Reconstruct Shallow and Deep Evolutionary Relationships Equally Well Michael S. Rosenberg and Sudhir Kumar Department of Biology, Arizona
Letter to the Editor Traditional Phylogenetic Reconstruction Methods Reconstruct Shallow and Deep Evolutionary Relationships Equally Well Michael S. Rosenberg and Sudhir Kumar Department of Biology, Arizona
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley
 Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley B.D. Mishler Feb. 7, 2012. Morphological data IV -- ontogeny & structure of plants The last frontier
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley B.D. Mishler Feb. 7, 2012. Morphological data IV -- ontogeny & structure of plants The last frontier
Phylogeny and systematics. Why are these disciplines important in evolutionary biology and how are they related to each other?
 Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
Phylogeny and systematics Why are these disciplines important in evolutionary biology and how are they related to each other? Phylogeny and systematics Phylogeny: the evolutionary history of a species
What do we mean by a species? Morphological species concept. Morphological species concept BIOL2007 SPECIES AND BIODIVERSITY. Kanchon Dasmahapatra
 BIOL2007 SPECIES AND BIODIVERSITY Kanchon Dasmahapatra What are species? How do species differ from each other? Biodiversity: How many species are there? What do we mean by a species? Darwin proved species
BIOL2007 SPECIES AND BIODIVERSITY Kanchon Dasmahapatra What are species? How do species differ from each other? Biodiversity: How many species are there? What do we mean by a species? Darwin proved species
Phyletic groups on networks. Ward C. Wheeler*
 Cladistics Cladistics 30 (2014) 447 451 10.1111/cla.12062 Phyletic groups on networks Ward C. Wheeler* Division of Invertebrate Zoology, American Museum of Natural History, Central Park West @ 79th Street,
Cladistics Cladistics 30 (2014) 447 451 10.1111/cla.12062 Phyletic groups on networks Ward C. Wheeler* Division of Invertebrate Zoology, American Museum of Natural History, Central Park West @ 79th Street,
BINF6201/8201. Molecular phylogenetic methods
 BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
BINF60/80 Molecular phylogenetic methods 0-7-06 Phylogenetics Ø According to the evolutionary theory, all life forms on this planet are related to one another by descent. Ø Traditionally, phylogenetics
SpeciesNetwork Tutorial
 SpeciesNetwork Tutorial Inferring Species Networks from Multilocus Data Chi Zhang and Huw A. Ogilvie E-mail: zhangchi@ivpp.ac.cn January 21, 2018 Introduction This tutorial covers SpeciesNetwork, a fully
SpeciesNetwork Tutorial Inferring Species Networks from Multilocus Data Chi Zhang and Huw A. Ogilvie E-mail: zhangchi@ivpp.ac.cn January 21, 2018 Introduction This tutorial covers SpeciesNetwork, a fully
Chapter 26: Phylogeny and the Tree of Life Phylogenies Show Evolutionary Relationships
 Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Chapter 26: Phylogeny and the Tree of Life You Must Know The taxonomic categories and how they indicate relatedness. How systematics is used to develop phylogenetic trees. How to construct a phylogenetic
Algorithmic Methods Well-defined methodology Tree reconstruction those that are well-defined enough to be carried out by a computer. Felsenstein 2004,
 Tracing the Evolution of Numerical Phylogenetics: History, Philosophy, and Significance Adam W. Ferguson Phylogenetic Systematics 26 January 2009 Inferring Phylogenies Historical endeavor Darwin- 1837
Tracing the Evolution of Numerical Phylogenetics: History, Philosophy, and Significance Adam W. Ferguson Phylogenetic Systematics 26 January 2009 Inferring Phylogenies Historical endeavor Darwin- 1837
5/31/17. Week 10; Monday MEMORIAL DAY NO CLASS. Page 88
 Week 10; Monday MEMORIAL DAY NO CLASS Page 88 Week 10; Wednesday Announcements: Family ID final in lab Today Final exam next Tuesday at 8:30 am here Lecture: Species concepts & Speciation. What are species?
Week 10; Monday MEMORIAL DAY NO CLASS Page 88 Week 10; Wednesday Announcements: Family ID final in lab Today Final exam next Tuesday at 8:30 am here Lecture: Species concepts & Speciation. What are species?
Evolutionary Significant Units (ESUs) & Management Units (MUs)
 Evolutionary Significant Units (ESUs) & Management Units (MUs) Diversity is Diverse and Complex Defining Management Units Within Species Genetic Distinctiveness & ESU s definition Measuring & Managing
Evolutionary Significant Units (ESUs) & Management Units (MUs) Diversity is Diverse and Complex Defining Management Units Within Species Genetic Distinctiveness & ESU s definition Measuring & Managing
Constructing Evolutionary/Phylogenetic Trees
 Constructing Evolutionary/Phylogenetic Trees 2 broad categories: Distance-based methods Ultrametric Additive: UPGMA Transformed Distance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Constructing Evolutionary/Phylogenetic Trees 2 broad categories: Distance-based methods Ultrametric Additive: UPGMA Transformed Distance Neighbor-Joining Character-based Maximum Parsimony Maximum Likelihood
Chapter 16: Reconstructing and Using Phylogenies
 Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Chapter Review 1. Use the phylogenetic tree shown at the right to complete the following. a. Explain how many clades are indicated: Three: (1) chimpanzee/human, (2) chimpanzee/ human/gorilla, and (3)chimpanzee/human/
Phylogenies Scores for Exhaustive Maximum Likelihood and Parsimony Scores Searches
 Int. J. Bioinformatics Research and Applications, Vol. x, No. x, xxxx Phylogenies Scores for Exhaustive Maximum Likelihood and s Searches Hyrum D. Carroll, Perry G. Ridge, Mark J. Clement, Quinn O. Snell
Int. J. Bioinformatics Research and Applications, Vol. x, No. x, xxxx Phylogenies Scores for Exhaustive Maximum Likelihood and s Searches Hyrum D. Carroll, Perry G. Ridge, Mark J. Clement, Quinn O. Snell
The Phylogenetic Reconstruction of the Grass Family (Poaceae) Using matk Gene Sequences
 The Phylogenetic Reconstruction of the Grass Family (Poaceae) Using matk Gene Sequences by Hongping Liang Dissertation submitted to the Faculty of the Virginia Polytechnic Institute and State University
The Phylogenetic Reconstruction of the Grass Family (Poaceae) Using matk Gene Sequences by Hongping Liang Dissertation submitted to the Faculty of the Virginia Polytechnic Institute and State University
Lecture 6 Phylogenetic Inference
 Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Lecture 6 Phylogenetic Inference From Darwin s notebook in 1837 Charles Darwin Willi Hennig From The Origin in 1859 Cladistics Phylogenetic inference Willi Hennig, Cladistics 1. Clade, Monophyletic group,
Special features / Articles spéciaux
 DNA barcoding makes an impact on the other side of the globe Muhammad Ashfaq Special features / Articles spéciaux those found in individuals from other species (Pereira et al. 2008). A wide range of molecular
DNA barcoding makes an impact on the other side of the globe Muhammad Ashfaq Special features / Articles spéciaux those found in individuals from other species (Pereira et al. 2008). A wide range of molecular
Introduction to Phylogenetic Analysis
 Introduction to Phylogenetic Analysis Tuesday 23 Wednesday 24 July, 2013 School of Biological Sciences Overview This workshop will provide an introduction to phylogenetic analysis, leading to a focus on
Introduction to Phylogenetic Analysis Tuesday 23 Wednesday 24 July, 2013 School of Biological Sciences Overview This workshop will provide an introduction to phylogenetic analysis, leading to a focus on
Creating an e-flora for South Africa
 SANBI POLICY DOCUMENT DIVISION: Biosystematics Research and Biodiversity Collections EFFECTIVE DATE: 1 April 2014 Compiler: Marianne le Roux & Janine Victor POLICY NUMBER: LAST AMENDED: Creating an e-flora
SANBI POLICY DOCUMENT DIVISION: Biosystematics Research and Biodiversity Collections EFFECTIVE DATE: 1 April 2014 Compiler: Marianne le Roux & Janine Victor POLICY NUMBER: LAST AMENDED: Creating an e-flora
BMC Evolutionary Biology 2013, 13:6
 BMC Evolutionary Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Correction: Male-killing
BMC Evolutionary Biology This Provisional PDF corresponds to the article as it appeared upon acceptance. Fully formatted PDF and full text (HTML) versions will be made available soon. Correction: Male-killing
Supplementary Materials for
 advances.sciencemag.org/cgi/content/full/1/8/e1500527/dc1 Supplementary Materials for A phylogenomic data-driven exploration of viral origins and evolution The PDF file includes: Arshan Nasir and Gustavo
advances.sciencemag.org/cgi/content/full/1/8/e1500527/dc1 Supplementary Materials for A phylogenomic data-driven exploration of viral origins and evolution The PDF file includes: Arshan Nasir and Gustavo
Microbial Taxonomy. Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Systematics - Bio 615
 Bayesian Phylogenetic Inference 1. Introduction, history 2. Advantages over ML 3. Bayes Rule 4. The Priors 5. Marginal vs Joint estimation 6. MCMC Derek S. Sikes University of Alaska 7. Posteriors vs Bootstrap
Bayesian Phylogenetic Inference 1. Introduction, history 2. Advantages over ML 3. Bayes Rule 4. The Priors 5. Marginal vs Joint estimation 6. MCMC Derek S. Sikes University of Alaska 7. Posteriors vs Bootstrap
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley
 "PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler March 31, 2011. Reticulation,"Phylogeography," and Population Biology:
"PRINCIPLES OF PHYLOGENETICS: ECOLOGY AND EVOLUTION" Integrative Biology 200B Spring 2011 University of California, Berkeley B.D. Mishler March 31, 2011. Reticulation,"Phylogeography," and Population Biology:
Computational approaches for functional genomics
 Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Computational approaches for functional genomics Kalin Vetsigian October 31, 2001 The rapidly increasing number of completely sequenced genomes have stimulated the development of new methods for finding
Likelihood Ratio Tests for Detecting Positive Selection and Application to Primate Lysozyme Evolution
 Likelihood Ratio Tests for Detecting Positive Selection and Application to Primate Lysozyme Evolution Ziheng Yang Department of Biology, University College, London An excess of nonsynonymous substitutions
Likelihood Ratio Tests for Detecting Positive Selection and Application to Primate Lysozyme Evolution Ziheng Yang Department of Biology, University College, London An excess of nonsynonymous substitutions
Taxon: generally refers to any named group of organisms, such as species, genus, family, order, etc.. Node: represents the hypothetical ancestor
 A quick review Taxon: generally refers to any named group of organisms, such as species, genus, family, order, etc.. Node: represents the hypothetical ancestor Branches: lines diverging from a node Root:
A quick review Taxon: generally refers to any named group of organisms, such as species, genus, family, order, etc.. Node: represents the hypothetical ancestor Branches: lines diverging from a node Root:
Phylogenetic Tree Reconstruction
 I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
I519 Introduction to Bioinformatics, 2011 Phylogenetic Tree Reconstruction Yuzhen Ye (yye@indiana.edu) School of Informatics & Computing, IUB Evolution theory Speciation Evolution of new organisms is driven
Lecture V Phylogeny and Systematics Dr. Kopeny
 Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
Delivered 1/30 and 2/1 Lecture V Phylogeny and Systematics Dr. Kopeny Lecture V How to Determine Evolutionary Relationships: Concepts in Phylogeny and Systematics Textbook Reading: pp 425-433, 435-437
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time
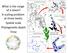 What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
What is the range of a taxon? A scaling problem at three levels: Spa9al scale Phylogene9c depth Time 1 5 0.25 0.15 5 0.05 0.05 0.10 2 0.10 0.10 0.20 4 Reminder of what a range-weighted tree is Actual Tree
1/27/2010. Systematics and Phylogenetics of the. An Introduction. Taxonomy and Systematics
 Systematics and Phylogenetics of the Amphibia: An Introduction Taxonomy and Systematics Taxonomy, the science of describing biodiversity, mainly naming unnamed species, and arranging the diversity into
Systematics and Phylogenetics of the Amphibia: An Introduction Taxonomy and Systematics Taxonomy, the science of describing biodiversity, mainly naming unnamed species, and arranging the diversity into
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS
 ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
ESTIMATION OF CONSERVATISM OF CHARACTERS BY CONSTANCY WITHIN BIOLOGICAL POPULATIONS JAMES S. FARRIS Museum of Zoology, The University of Michigan, Ann Arbor Accepted March 30, 1966 The concept of conservatism
DNA Sequencing as a Method for Larval Identification in Odonates
 DNA Sequencing as a Method for Larval Identification in Odonates Adeline Harris 121 North St Apt 3 Farmington, ME 04938 Adeline.harris@maine.edu Christopher Stevens 147 Main St Apt 1 Farmington, ME 04938
DNA Sequencing as a Method for Larval Identification in Odonates Adeline Harris 121 North St Apt 3 Farmington, ME 04938 Adeline.harris@maine.edu Christopher Stevens 147 Main St Apt 1 Farmington, ME 04938
Chapter 26 Phylogeny and the Tree of Life
 Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
Chapter 26 Phylogeny and the Tree of Life Biologists estimate that there are about 5 to 100 million species of organisms living on Earth today. Evidence from morphological, biochemical, and gene sequence
 that of Phylotree.org, mtdna tree Build 1756 (Supplementary TableS2). is resulted in 78 individuals allocated to the hg B4a1a1 and three individuals to hg Q. e control region (nps 57372 and nps 1602416526)
that of Phylotree.org, mtdna tree Build 1756 (Supplementary TableS2). is resulted in 78 individuals allocated to the hg B4a1a1 and three individuals to hg Q. e control region (nps 57372 and nps 1602416526)
Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
 1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
1 2 3 4 5 6 7 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 25 26 27 28 29 30 31 32 33 34 35 36 37 38 39 40 41 42 43 44 45 Title ghost-tree: creating hybrid-gene phylogenetic trees for diversity analyses
Effects of phylogenetic reconstruction method on the robustness of species delimitation using single-locus data
 Methods in Ecology and Evolution 2014, 5, 1086 1094 doi: 10.1111/2041-210X.12246 Effects of phylogenetic reconstruction method on the robustness of species delimitation using single-locus data Cuong Q.
Methods in Ecology and Evolution 2014, 5, 1086 1094 doi: 10.1111/2041-210X.12246 Effects of phylogenetic reconstruction method on the robustness of species delimitation using single-locus data Cuong Q.
Homework Assignment, Evolutionary Systems Biology, Spring Homework Part I: Phylogenetics:
 Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Homework Assignment, Evolutionary Systems Biology, Spring 2009. Homework Part I: Phylogenetics: Introduction. The objective of this assignment is to understand the basics of phylogenetic relationships
Plant Names and Classification
 Plant Names and Classification Science of Taxonomy Identification (necessary!!) Classification (order out of chaos!) Nomenclature (why not use common names?) Reasons NOT to use common names Theophrastus
Plant Names and Classification Science of Taxonomy Identification (necessary!!) Classification (order out of chaos!) Nomenclature (why not use common names?) Reasons NOT to use common names Theophrastus
7. Tests for selection
 Sequence analysis and genomics 7. Tests for selection Dr. Katja Nowick Group leader TFome and Transcriptome Evolution Bioinformatics group Paul-Flechsig-Institute for Brain Research www. nowicklab.info
Sequence analysis and genomics 7. Tests for selection Dr. Katja Nowick Group leader TFome and Transcriptome Evolution Bioinformatics group Paul-Flechsig-Institute for Brain Research www. nowicklab.info
Post-doc fellowships to non-eu researchers FINAL REPORT. Home Institute: Centro de Investigaciones Marinas, Universidad de La Habana, CUBA
 Recipient: Maickel Armenteros Almanza. Post-doc fellowships to non-eu researchers FINAL REPORT Home Institute: Centro de Investigaciones Marinas, Universidad de La Habana, CUBA Promoter: Prof. Dr. Wilfrida
Recipient: Maickel Armenteros Almanza. Post-doc fellowships to non-eu researchers FINAL REPORT Home Institute: Centro de Investigaciones Marinas, Universidad de La Habana, CUBA Promoter: Prof. Dr. Wilfrida
Today's project. Test input data Six alignments (from six independent markers) of Curcuma species
 DNA sequences II Analyses of multiple sequence data datasets, incongruence tests, gene trees vs. species tree reconstruction, networks, detection of hybrid species DNA sequences II Test of congruence of
DNA sequences II Analyses of multiple sequence data datasets, incongruence tests, gene trees vs. species tree reconstruction, networks, detection of hybrid species DNA sequences II Test of congruence of
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Jan 27 & 29):
 Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Jan 27 & 29): Statistical estimation of models of sequence evolution Phylogenetic inference using maximum likelihood:
Biology 559R: Introduction to Phylogenetic Comparative Methods Topics for this week (Jan 27 & 29): Statistical estimation of models of sequence evolution Phylogenetic inference using maximum likelihood:
Integrating Fossils into Phylogenies. Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
 IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
IB 200B Principals of Phylogenetic Systematics Spring 2011 Integrating Fossils into Phylogenies Throughout the 20th century, the relationship between paleontology and evolutionary biology has been strained.
Phylogenetics. BIOL 7711 Computational Bioscience
 Consortium for Comparative Genomics! University of Colorado School of Medicine Phylogenetics BIOL 7711 Computational Bioscience Biochemistry and Molecular Genetics Computational Bioscience Program Consortium
Consortium for Comparative Genomics! University of Colorado School of Medicine Phylogenetics BIOL 7711 Computational Bioscience Biochemistry and Molecular Genetics Computational Bioscience Program Consortium
Phylogenetics: Bayesian Phylogenetic Analysis. COMP Spring 2015 Luay Nakhleh, Rice University
 Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Phylogenetics: Bayesian Phylogenetic Analysis COMP 571 - Spring 2015 Luay Nakhleh, Rice University Bayes Rule P(X = x Y = y) = P(X = x, Y = y) P(Y = y) = P(X = x)p(y = y X = x) P x P(X = x 0 )P(Y = y X
Phylogeny 9/8/2014. Evolutionary Relationships. Data Supporting Phylogeny. Chapter 26
 Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Phylogeny Chapter 26 Taxonomy Taxonomy: ordered division of organisms into categories based on a set of characteristics used to assess similarities and differences Carolus Linnaeus developed binomial nomenclature,
Estimating Evolutionary Trees. Phylogenetic Methods
 Estimating Evolutionary Trees v if the data are consistent with infinite sites then all methods should yield the same tree v it gets more complicated when there is homoplasy, i.e., parallel or convergent
Estimating Evolutionary Trees v if the data are consistent with infinite sites then all methods should yield the same tree v it gets more complicated when there is homoplasy, i.e., parallel or convergent
The practice of naming and classifying organisms is called taxonomy.
 Chapter 18 Key Idea: Biologists use taxonomic systems to organize their knowledge of organisms. These systems attempt to provide consistent ways to name and categorize organisms. The practice of naming
Chapter 18 Key Idea: Biologists use taxonomic systems to organize their knowledge of organisms. These systems attempt to provide consistent ways to name and categorize organisms. The practice of naming
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley
 Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley B.D. Mishler April 12, 2012. Phylogenetic trees IX: Below the "species level;" phylogeography; dealing
Integrative Biology 200A "PRINCIPLES OF PHYLOGENETICS" Spring 2012 University of California, Berkeley B.D. Mishler April 12, 2012. Phylogenetic trees IX: Below the "species level;" phylogeography; dealing
AToL: Collaborative research on ant phylogeny: a comprehensive evolutionary tree for the world s premier social organisms
 AToL: Collaborative research on ant phylogeny: a comprehensive evolutionary tree for the world s premier social organisms NSF EF-0431330; 10/01/2004 09/30/2009 P.S.Ward1, Seán Brady2, Brian Fisher3 & Ted
AToL: Collaborative research on ant phylogeny: a comprehensive evolutionary tree for the world s premier social organisms NSF EF-0431330; 10/01/2004 09/30/2009 P.S.Ward1, Seán Brady2, Brian Fisher3 & Ted
Microbes usually have few distinguishing properties that relate them, so a hierarchical taxonomy mainly has not been possible.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy. Slowly evolving molecules (e.g., rrna) used for large-scale structure; "fast- clock" molecules for fine-structure.
 Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
Microbial Taxonomy Traditional taxonomy or the classification through identification and nomenclature of microbes, both "prokaryote" and eukaryote, has been in a mess we were stuck with it for traditional
What is Phylogenetics
 What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
What is Phylogenetics Phylogenetics is the area of research concerned with finding the genetic connections and relationships between species. The basic idea is to compare specific characters (features)
Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Information #
 Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Details of PRF Methodology In the Poisson Random Field PRF) model, it is assumed that non-synonymous mutations at a given gene are either
Bustamante et al., Supplementary Nature Manuscript # 1 out of 9 Details of PRF Methodology In the Poisson Random Field PRF) model, it is assumed that non-synonymous mutations at a given gene are either
 Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
Tree of Life iological Sequence nalysis Chapter http://tolweb.org/tree/ Phylogenetic Prediction ll organisms on Earth have a common ancestor. ll species are related. The relationship is called a phylogeny
JML: testing hybridization from species trees
 Molecular Ecology Resources (2012) 12, 179 184 doi: 10.1111/j.1755-0998.2011.03065.x JML: testing hybridization from species trees SIMON JOLY Institut de recherche en biologie végétale, Université de Montréal
Molecular Ecology Resources (2012) 12, 179 184 doi: 10.1111/j.1755-0998.2011.03065.x JML: testing hybridization from species trees SIMON JOLY Institut de recherche en biologie végétale, Université de Montréal
