Temporal expression patterns of 39 Brachyurydownstream genes associated with notochord formation in the Ciona intestinalis embryo
|
|
|
- Baldric Cannon
- 6 years ago
- Views:
Transcription
1 Develop. Growth Differ. (1999) 41, Temporal expression patterns of 39 Brachyurydownstream genes associated with notochord formation in the Ciona intestinalis embryo Kohji Hotta, 1 * Hiroki Takahashi, 1 Albert Erives, 2 Michael Levine 2 and Nori Satoh 1 1 Department of Zoology, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto , Japan and 2 Division of Genetics, Department of Molecular and Cellular Biology, 21 Koshland Hall, University of California, Berkeley, CA 94720, USA Expression of the Brachyury (Ci-Bra) gene of the ascidian Ciona intestinalis is initiated at the 64-cell stage. Gene expression is restricted to notochord precursor cells, and Ci-Bra plays a key role in notochord differentiation. In a previous study, nearly 50 cdna clones for potential Ci-Bra-downstream genes that are expressed in notochord cells were isolated. The present determination, by whole-mount in situ hybridization, of the temporal expression patterns of 19 notochord-specific and 20 notochord-predominant genes demonstrated that the timings of initiation of the expression of various genes was not identical. The expression of several genes was initiated as early as the gastrula stage. However, the expression of most of the notochord-specific genes commenced at the neural plate stage. Partial nucleotide sequence data of these clones suggest that genes expressed earlier encode potential transcriptional factors and/or nuclear proteins, while those expressed later encode proteins implicated in cell adhesion, signal transduction, regulation of the cytoskeleton, and components of the extracellular matrix. These gene activities may be associated with changes in cell shape and adhesion during the intercalation and extension of the notochord cells. Key words: ascidian, Brachyury, downstream genes, notochord, temporal expression patterns. Introduction Brachyury encodes a sequence-specific activator that contains a T-box DNA-binding domain (reviewed by Herrmann & Kispert 1994; Smith 1997; Papaioannou & Silver 1998). The gene was originally cloned from mice, taking advantage of a Brachyury mutation (Herrmann et al. 1990). In vertebrates, Brachyury is initially expressed in the presumptive mesoderm of gastrulae, and during later stages the expression is gradually restricted to the developing notochord and tail-bud (Wilkinson et al. 1990; Smith et al. 1991; Schulte-Merker et al. 1992; Kispert et al. 1995). Brachyury expression is critical for notochord differentiation in all vertebrates that have been studied, including mice (Rashbass et al. 1991), frogs (Conlon et al. 1996) and zebrafish (Schulte-Merker et al. 1994). Ascidians (urochordates) constitute one of the three chordate groups. Because fertilized ascidian eggs *Author to whom all correspondence should be addressed. kozi@ascidian.zool.kyoto-u.ac.jp Received 9 April 1999; revised 21 May 1999; accepted 21 May develop rather quickly into tadpole-type larvae with several distinct types of tissues and organs, and because the lineage of embryonic cells is well documented, the ascidian embryo provides a good experimental system to explore the expression and function of developmental genes (reviewed by Satoh 1994, 1999). In addition, the ascidian tadpole is thought to represent the most simplified and primitive chordate body plan (reviewed by Satoh 1995; Satoh & Jeffery 1995; Di Gregorio & Levine 1998). It contains a notochord composed of just 40 cells, of which the lineage has been completely described (Conklin 1905; Nishida 1987). Previous studies in two divergent ascidians revealed that ascidian Brachyury genes, As-T of Halocynthia roretzi (Yasuo & Satoh 1993, 1994) and Ci-Bra of Ciona intestinalis (Corbo et al. 1997), are expressed exclusively in the notochord precursor cells and that the timing of gene expression coincides with clonal restriction of the notochord lineage at the 64-cell stage (cf, Fig. 1A). In H. roretzi, notochord formation is induced at the 32-cell stage by signals emanating from the adjacent endoderm cells as well as neighboring notochord cells (Nakatani & Nishida 1994). Overexpression of As-T via its messenger ribonucleic acid (mrna)
2 658 K. Hotta et al. injection results in notochord formation without a requirement for the inductive event at the 32-cell stage (Yasuo & Satoh 1998). In addition, misexpression of both As-T (Yasuo & Satoh 1998) and Ci-Bra (Takahashi et al. 1999) causes transformation of endodermal and neuronal lineages into notochord cells. These studies demonstrate that the ascidian Brachyury gene is critical in determination of the notochord. Figs 1 6. Temporal expression patterns of (1) Ci-Bra and (2 6) five Ci-Bra-downstream notochord-related genes during embryogenesis of Ciona intestinalis, as revealed by whole-mount in situ hybridization with a digoxigenin-labeled antisense probe. Genes encoded by cdna clones: (2) 403e, (3) 406g, (4) 210d, (5) 309h, and (6) 309g. Embryos at (A) 64-cell stage, vegetal pole view; (B) gastrula stage, vegetal pole view; (C) neural plate stage, dorsal (vegetal) view; (D) neurula stage, dorsal view; and (E) tail-bud stage, side view. ant, anterior; post, posterior side of the embryo. bp, blastopore; End, endoderm; N, notochord; np, neural plate. Bar, 50 µm 1(A); applicable for all panels.
3 Ciona Brachyury-downstream gene expression 659 In a previous study, we attempted the isolation of potential Ci-Bra target genes (Takahashi et al. 1999). A 2.6 kb genomic DNA fragment from the 5 -flanking region of the forkhead/hnf-3 gene of C. intestinalis (Ci-fkh) is sufficient to direct the expression of a lacz reporter gene in these tissues after electroporation into 1-cell embryos (Corbo et al. 1997). Attaching the Ci- Bra coding sequence to the Ci-fkh promoter region resulted in misexpression of the Ci-Bra gene in ectopic tissues. The resulting fusion gene causes extensive transformation of the endoderm into notochord, whereby mutant tail-bud embryos contain a large mass of notochord tissue in midtail regions. The efficiency of the electroporation method allowed us to obtain large quantities of mutant embryos, thereby facilitating subtractive hybridization reactions using mrna extracted from wild-type and mutant embryos at the neurula and early tail-bud stages. A subtractive cdna library was prepared that contained mrna that are overexpressed in the mutant embryos relative to the wild-type controls. The library contained 923 cdna clones, 501 of which were shown to be independent. Of the 501 independent cdna clones that were surveyed, nearly 50 genes were specifically or predominantly expressed in the notochord cells. The potential Ci-Bra target genes encode components of the extracellular matrix, as well as proteins implicated in cell adhesion, signal transduction, and regulation of the cytoskeleton. Therefore, we propose that those genes define one or more signaling pathways, which control changes in cell shape and adhesion during the intercalation and extension of the notochord (cf, Miyamoto & Crowther 1985). Even though the expression of all of these genes is induced by Ci-Bra misexpression, it remains to be determined whether these genes are direct or indirect targets of Ci-Bra. As a first step in studies to understand the Ci- Bra-downstream genetic cascade, the present study determined the temporal expression patterns of 19 notochord-specific and 20 notochord-predominant genes that are induced by Ci-Bra misexpression. Materials and Methods Ascidian eggs and embryos Eggs and sperm of C. intestinalis were obtained surgically from the gonoduct. After insemination, eggs were dechorionated by immersing them in seawater that contained 1% sodium thioglycolate (Wako Pure Chemical Industries Ltd, Osaka, Japan) and 0.05% actinase E (Kaken Pharmaceutical Co. Ltd, Tokyo, Japan). Naked eggs and embryos were then reared at about 18 C in agar-coated dishes with Millipore-filtered seawater (MFSW) containing 50 µg/ml streptomycin sulfate. Under our culture conditions, the first cleavage occurred about 1 h after insemination. Embryos developed to the 64-cell, gastrula, neural plate, neurula, and early tail-bud stages about 4, 5.5, 7, 8, and 9 h after fertilization, respectively. Tadpole larvae began to swim at about 18 h of development. Eggs and embryos at appropriate stages were fixed for in situ hybridization. cdna probes for Ciona notochord-related genes Details of isolation and characterization of cdna clones for Ci-Bra-downstream genes that are expressed in notochord cells of C. intestinalis tail-bud embryos were reported in Takahashi et al. (1999). In the present study, we examined the temporal expression patterns of 39 notochord-related genes (cf, Table 2). Nineteen genes represented by cdna clones 103f, 107g, 110g, 210d, 212g, 309h, 402c, 403e, 406g, 407e, 408h, 409d, 411d, 502a, 505a, 507d, 507h, 801h, and 807d are expressed specifically in notochord cells of the early tail-bud embryo. The other 20 genes, represented by 002d, 003b, 003d, 005d, 006c, 007d, 010b, 111a, 204d, 208e, 209c, 211g, 308d, 309g, 504g, 603d, 608g, 706e, 708g, and 903g, are predominantly expressed in notochord cells of the early tail-bud embryo (hereafter in this report we designate the clone number as the gene name). Determination of the sequences of about 500 nucleotides of both the 5 and 3 regions suggested that 21 of the 39 genes show sequence similarity to reported proteins, but the other 18 have no sequence similarity to known proteins (Takahashi et al. 1999; cf, Table 2). The cdna were inserted into the EcoRI/XhoI site of the plasmid vector pbluescript II SK( ) (Stratagene, La Jolla, CA, USA). Digoxigenin (DIG)-labeled sense and antisense probes were synthesized following the instructions of the suppliers of the kit (DIG RNA Labelling Kit; Boehringer Mannheim, Heidelberg, Germany). Their final sizes were reduced to approximately 150 nucleotides by alkaline hydrolysis. In situ hybridization Whole-mount specimens were hybridized in situ using DIG-labeled antisense and sense RNA probes essentially according to the method described by Corbo et al. (1997). Briefly, embryos were fixed in 4% paraformaldehyde in 0.1 M 3-(N-Morpholino) propanesulfonic acid (MOPS) buffer (ph 7.5), 0.5 M NaCl. After being thoroughly washed with phosphate-buffered saline containing 0.1% Tween 80 (PBT), the embryos were treated with 2 µg/ml proteinase K (Sigma Chemical Co., St Louis, MO, USA) in PBT for 30 min at 37 C, then postfixed with 2% paraformaldehyde in PBT for 30 min at room temperature. After 1.5 h of
4 660 K. Hotta et al. prehybridization at 55 C, the embryos were allowed to hybridize with the DIG-labeled probes at a concentration of 1 µg/ml for at least 18 h at 55 C. After hybridization, the embryos were washed 10 times for 15 min each with the hybridization solution at 60 C. Thereafter, the samples were incubated for 2 h with 500 µl anti-digalkaline phosphate conjugate, and color was developed according to the protocol of Boehringer Mannheim. After dehydration, some of the specimens were made transparent by placing them in 100% xylene. The samples were mounted on glass slides with entellan neu (Merck, Darmstadt, Germany) and photographed. Results Temporal expression patterns of 19 notochordspecific genes With the aid of in situ hybridization of whole-mount specimens, we first examined the appearance and distribution of transcripts of each of the 19 potential Ci- Bra downstream genes that are expressed specifically in notochord cells of C. intestinalis tail-bud embryos. As summarized in Table 1, the timings for the initiation of gene expression were not identical; two genes initiated their expression as early as the mid-gastrula stage, while the expression of 12 genes began at the neural plate stage and that of five genes at the neurula stage. No genes were expressed before the initiation of gastrulation. The 403e gene, for example, began to be expressed at the mid-gastrula stage (Fig. 2B). The timing of initiation of the 403e gene expression was about 1.5 h behind that of Ci-Bra expression at the 64-cell stage (Fig. 1A). At this stage, the 403e transcript was evident only in 10 notochord precursor cells (Fig. 2B). Although in later stages this gene seemed to be expressed in several neuronal cells as well as epidermal cells (Fig. 2C E), we categorized it as notochord specific, because its initial expression was restricted to the notochord precursor cells. Partial nucleotide sequence data suggest that the 403e gene encodes a nuclear protein with a LIM domain (Table 2). The transcript of another gene (507h) also became detectable in the notochord precursor cells at the gastrula stage. The 507h gene encodes a protein with no sequence similarity to any known protein. The present analysis demonstrated that the expression of 12 of the 19 Ci-Bra-downstream notochord-specific genes (63%) is initiated at the neural plate stage (Table 1; Figs 3,4); genes 107g, 110g, 406g, 408h, 502a, 807d, 103f, 407e, 210d, 212 g, 801h and 505a were categorized into this group. For example, 406g may encode a nuclear protein associated with cell division control (Table 2). The transcript of this gene became detectable in almost all of the primordial notochord cells at the neural plate stage (Fig. 3C), and later was restricted to notochord cells in the tail-bud embryo (Fig. 3E). On the other hand, 210d encoded a protein with no sequence similarity to any known protein. The 210d transcript was first evident in a fraction of the primordial notochord cells that were located in the posterior region of the notochord cell field (Fig. 4C,D), and later the transcript was found in all of the notochord cells (Fig. 4E). Partial nucleotide sequence data of the notochordspecific genes that initiate their expression at the neural plate stage suggested that these genes encode proteins that act as cell surface receptors, or play roles in signal transduction or regulation of the cytoskeleton. Expression of the remaining five notochord-specific genes was initiated at the neurula stage (Table 1). For example, 309h, of which the transcript became detectable at the neurula stage (Fig. 5D), may encode an integral membrane protein associated with a sulfate transporter protein (Table 2). The expression of four other genes, 402c, 409d, 507d and 411d, also began at this stage. In the present analysis, none of the potential Ci-Bra-downstream, notochord-specific genes started to be expressed at the early tail-bud or later stages. Temporal expression patterns of 20 notochordpredominant genes Twenty potential Ci-Bra-downstream genes were expressed predominantly in notochord cells of the tailbud embryos. In addition, transcripts were detectable in nerve cord cells, neuronal cells, or muscle cells, depending on the gene. Thus, these genes were expressed in the notochord and its neighboring cells. Table 1. Timing of initiation of expression of Ci-Bra-downstream notochord-related genes during embryogenesis of the ascidian Ciona intestinalis Stage of initiation of the gene expression Genes No. genes 64-cell Gastrula Neural plate Neurula Tail-bud Notochord-specific genes Notochord-predominant genes Total
5 Table 2. Timing of initiation of Ci-Bra-downstream notochord-related gene expression in Ciona embryos Developmental stage Clone Sequence similarity Possible features Figure Gastrula 403e LIM-only protein LIM domain 2 507h 006c Serpin proteins, transcriptional regulatory 308d 504g Myosin heavy chain Structure 706e 211g Inhibin B chain signature, C1q domain. speract receptor repeat 903g Neural plate 107g 110g ATP-citrate (pro-s-)-lyase C2HC-type zinc-finger signature, laminin-type EGF-like (LE) domain 406g Cell division control protein 45 Nuclear protein integral membrane signature 3 408h Prothrombinase precursor (fibrinogen-like protein) Fibrinogen and chains C-terminal domain 502a Cellular tumor antigen P53 EF-hand calcium-binding domain, recoverin family signature 807d 103f HSPG LDL-receptor class A (LDLRA) domain 407e LIM-only protein LIM domain 210d 4 212g 801h Extensin precursor (proline-rich glycoprotein) 505a Sulfate adenylyltransferase (SAT) Pheromone A receptor signature, tyrosine-specific protein phosphatases proteins 003b Agrin Cell adhesion, calcium-binding EGF-like domain pattern proteins 010b Collagen 1 (XI) chain 603d Max interacting protein 1 (MXI1 protein) myc-type, helix-loop-helix dimerization domain proteins Neurula 402c 409d 507d 309h Sulfate transporter Membrane, sulfate transporter proteins, integral membrane proteins 5 411d 002d Ezrin/radixin/moesin Cell adhesion, membrane 608g Protein-tyrosine phosphatase MEG1 Structure, extracellular 209c 007d 111a Reticulocalbin 1 EF-hand calcium-binding 204d P-selectin Cell adhesion, selectin superfamily complement-binding repeat, sushi domain 708g Initial tail-bud 003d 005d Tensin Cell adhesion 208e 309g N-acetyl-lactosamine synthase 6 Bold type, notochord-specific genes; roman type, notochord-predominant genes. ATP, adenosine triphosphate; HSPG, heparan sulfate proteoglycan; LDL, low density lipoprotein; MEG1, megakaryoblastic cell line cdna1. Ciona Brachyury-downstream gene expression 661
6 662 K. Hotta et al. In situ hybridization of whole-mount specimens demonstrated that these 20 potential Ci-Bra-downstream genes were expressed within a relatively broad time window, in contrast to the general pattern, in which notochord-specific downstream genes are mainly expressed at the neural plate stage, as mentioned above (Table 1). Namely, the expression of six genes was initiated at the mid-gastrula stage, while the expression of three, seven and four genes commenced at the neural plate, neurula, and tail-bud stages, respectively. Genes 006c, 211g, 308d, 504g, 706e and 903g began to be expressed at the mid-gastrula stage (Tables 1,2). Partial nucleotide sequence data suggest that the 006c gene may encode a protein associated with transcriptional regulation, while the 504g gene may encode a structural protein with sequence similarity to the myosin heavy chain. Transcripts of genes 003b, 010b and 603d became detectable in the primordial notochord cells at the neural plate stage. The 003b gene may encode a protein associated with cell adhesion, and 010b may encode the collagen 1 chain (Table 2). The expression of seven genes, 002d, 007d, 111a, 204d, 209c, 608 g and 708 g, was initiated at the neurula stage (Tables 1,2). For example, 002d may encode an ezrin/radixin/moesin (ERM)-like protein that is involved in cell adhesion (Table 2). The expression of four genes, 003d, 005d, 208e and 309 g, was initiated at the early tail-bud stage (Tables 1,2). For example, no transcripts of the 309g gene were detected at early developmental stages up to the neurula stage (Fig. 6A D). The transcript was first evident in notochord cells as well as a few neuronal cells in the early tail-bud embryo (Fig. 6E). The 309g gene may encode N-acetyl-lactosamine synthase (Table 2). The 005d gene may encode tensin, which is involved in cell adhesion (Table 2). Discussion Differentiation of notochord cells in the ascidian embryo has been studied intensively (reviewed by Satoh 1994, 1999). The studies include analyses of cell lineage (Conklin 1905; Nishida 1987), specification mechanisms at the cellular level (Nakatani & Nishida 1994; Nakatani et al. 1996) and at the molecular level (Yasuo & Satoh 1993, 1998; Corbo et al. 1997, 1998; Fujiwara et al. 1998), behavior of notochord cells in living C. intestinalis embryos investigated with time-lapse video photography (Miyamoto & Crowther 1985), and ultrastructural observations of changes in cell shape and notochord-sheath formation (reviewed by Cloney 1990). Altogether, these studies have revealed that the notochord of the ascidian embryo is formed as follows. The ascidian larval notochord consists of exactly 40 cells aligned at the midline, longitudinally in single file in the tail. Thirty-two of the 40 notochord cells in the anterior and middle portion of the tail are derived from the A4.1 pair of the 8-cell embryo, and the other eight cells, in the posterior region, from the B4.1 pair. At the 64-cell stage, the A7.3 and A7.7 pairs are restricted to give rise to notochord cells, and at the 110-cell stage, A8.5, A8.6, A8.13, A8.14 and B8.6 pairs are primordial notochord cells, giving rise to 40 cells after two divisions. Therefore, every notochord cell has a history of nine divisions from the zygotic stage until the ultimate differentiation. The positions of the notochord cells derived from each precursor cell are not predetermined within the tail. A cell lineage study revealed that the presumptive notochord cells from the left and right sides of the embryo can mix randomly at the midline (Nishida 1987). However, the A-line and B-line precursor cells do not interdigitate with one another. In H. roretzi, specification of notochord cells is induced at the 32-cell stage by signals emanating from the adjacent endoderm cells as well as neighboring notochord precursor cells (Nakatani & Nishida 1994). Then, at the 64-cell stage, the Brachyury gene (As-T) is expressed exclusively in the primordial notochord cells (Yasuo & Satoh 1993, 1994). Ci-Bra is also expressed exclusively in the notochord precursor cells in the 64-cell-stage Ciona embryo (Corbo et al. 1997). Molecules that may possibly be responsible for the ascidian Brachyury activation are bfgf for As-T (Nakatani et al. 1996) and Suppressor of Hairless protein for Ci-Bra (Corbo et al. 1998). During this stage, Ciona snail represses Ci-Bra expression in para-axial mesoderm cells of the mesenchyme and muscle (Fujiwara et al. 1998). The expression of As-T or Ci-Bra is sufficient for gene expression that is associated with the differentiation of notochord cells and the morphological formation of the notochord (Yasuo & Satoh 1998; Takahashi et al. 1999; G. Satoh et al. unpubl. data, 1999). Studies of the formation of the notochord in living C. intestinalis embryos with time-lapse video photography have demonstrated that cell proliferation, growth, migration, rearrangement, shape changes, and alteration of the extracellular environment occur as the notochord is transformed from a loose plate or mass of cells during neurulation and initial tail-bud formation into an elongated rod surrounded by a sheath of fibrous materials during larval formation (Miyamoto & Crowther 1985). The basal surfaces of the notochord cells rest on a basal lamina termed the notochordal sheath that completely surrounds the notochord. The
7 Ciona Brachyury-downstream gene expression 663 notochordal sheath contains many circumferentially orientated and longitudinally orientated filaments (Cloney 1969). Previous and present studies add further information on the process of notochord formation in the ascidian embryo. Namely, Ci-Bra expression in turn causes expression of a great number of genes that may be associated with formation of the notochord. As shown by the present study, the timings of initiation of expression of potential Ci-Bra-downstream notochord-related genes are not identical. The expression of several genes is initiated as early as the gastrula stage, about 1~1.5 h after Ci-Bra expression. Although the timing of expression of the Ci-Bra protein must be examined further, this time lag seems reasonable for inducing the activation of Ci-Bra direct target genes. However, the expression of most of the notochord-specific genes is initiated at the neural plate stage, suggesting that this is the stage critical for activation of structural genes associated with notochord formation. Partial nucleotide sequence data of these clones as well as the present determination of timing of gene expression demonstrated that genes expressed earlier encode potential transcriptional factors and/or nuclear proteins (such as 403e). Those expressed later (such as 406g, 003b, 010b, 002d and 005d) encode proteins implicated in cell adhesion, signal transduction, and regulation of the cytoskeleton, and components of the extracellular matrix, which may be associated with changes in cell shape and adhesion during the intercalation and extension of the notochord cells. These results suggest the possibility that Ci-Bra may directly activate a few target genes that encode other types of transcription factors. The transcription factors then activate the notochord-specific structural genes that are expressed around the time of neurulation. Therefore, most of the structural genes that are expressed around the time of neurulation are likely indirect targets of Ci-Bra. Keeping in mind such a possibility, details of the Ci-Bra-downstream genetic cascade for notochord formation in the ascidian embryo should be studied further in the future. Acknowledgements We thank all staff members of the Marine BioSource Education Center of Tohoku University and Kazuko Hirayama for their help in collection of C. intestinalis. This research was supported in part by a Grant-in-Aid for Specially Promoted Research (No ) from the Ministry of Education, Science, Sports and Culture of Japan to N. S. and the Human Frontier Science Program (RG212/1997) to M. L. and N. S. The nucleotide sequence data reported in this paper will appear in the homepage ( References Cloney, R. A Cytoplasmic filaments and morphogenesis: The role of the notochord in ascidian metamorphosis. Z. Zellforsch. 100, Cloney, R. A Urochordata Ascidiacea. In Reproductive Biology of Invertebrates (Eds K. G. Adiyodi & R. G. Adiyodi), pp Oxford & IBH Publishing, New Delhi. Conklin, E. G The organization and cell lineage of the ascidian egg. J. Acad. Natl Sci. 13, Conlon, F. L., Sedgwick, S. G., Weston, K. M. & Smith, J. C Inhibition of Xbra transcription activation causes defects in mesodermal patterning and reveals autoregulation of Xbra in dorsal mesoderm. Development 122, Corbo, J. C., Fujiwara, S., Levine, M. & Di Gregorio, A Suppressor of Hairless activates Brachyury expression in the Ciona embryo. Dev. Biol. 203, Corbo, J. C., Levine, M. & Zeller, R. W Characterization of a notochord-specific enhancer from the Brachyury promoter region of the ascidian, Ciona intestinalis. Development 124, Di Gregorio, A. & Levine, M Ascidian embryogenesis and the origins of the chordate body plan. Curr. Opin. Genet. Dev. 8, Fujiwara, S., Corbo, J. C. & Levine, M The Snail repressor establishes a muscle/notochord boundary in the Ciona embryo. Development 125, Herrmann, B. G. & Kispert, A The T genes in embryogenesis. Trends Genet. 10, Herrmann, B. G., Labeit, S., Poustka, A., King, T. R. & Lehrach, H Cloning of the T gene required in mesoderm formation in the mouse. Nature 343, Kispert, A., Ortner, H., Cooke, J. & Herrmann, B. G The chick Brachyury gene: Developmental expression pattern and response to axial induction by localized activin. Dev. Biol. 168, Miyamoto, D. M. & Crowther, R. J Formation of the notochord in living ascidian embryos. J. Embryol. Exp. Morphol. 86, Nakatani, Y. & Nishida, H Induction of notochord during ascidian embryogenesis. Dev. Biol. 166, Nakatani, Y., Yasuo, H., Satoh, N. & Nishida, H Basic fibroblast growth factor induces notochord formation and the expression of As-T, a Brachyury homolog, during ascidian embryogenesis. Development 122, Nishida, H Cell lineage analysis in ascidian embryos by intracellular injection of a tracer enzyme: III. Up to the tissue restricted stage. Dev. Biol. 121, Papaioannou, V. E. & Silver, L. M The T-box gene family. Bioessays 20, Rashbass, P., Cooke, L. A., Herrmann, B. G. & Beddington, R. S. P A cell autonomous function of Brachyury in T/T embryonic stem cell chimaeras. Nature 353, Satoh, N Developmental Biology of Ascidians. Cambridge University Press, New York. Satoh, N Towards a molecular understanding of developmental mechanisms underlying the origin and evolution of chordates. In Biodiversity and Evolution (Eds R. Arai, M. Kato & Y. Doi), pp National Science Museum Foundation, Tokyo.
8 664 K. Hotta et al. Satoh, N Cell fate determination in the ascidian embryo. In Cell Lineage and Fate Determination (Ed. S. A. Moody), pp Academic Press, San Diego. Satoh, N. & Jeffery, W. R Chasing tails in ascidians: Developmental insights into the origin and evolution of chordates. Trends Genet. 11, Schulte-Merker, S., Ho, R. K., Herrmann, B. G. & Nüsslein-Volhard, C The protein product of the zebrafish homologue of the mouse T gene is expressed in nuclei of the germ ring and the notochord of the early embryo. Development 116, Schulte-Merker, S., van Eeden, F. J. M., Halpern, M. E., Kimmel, C. B. & Nüsslein-Volhard, C no tail (ntl) is the zebrafish homologue of the mouse T (Brachyury) gene. Development 120, Smith, J Brachyury and the T-box genes. Curr. Opin. Genet. Dev. 7, Smith, J. C., Price, B. M. J., Green, J. B. A., Weigel, D. & Herrmann, B. G Expression of a Xenopus homolog of Brachyury (T) is an immediate-early response to mesoderm induction. Cell 67, Takahashi, H., Hotta, K., Erives, A. et al Brachyurydownstream notochord differentiation in the ascidian embryo. Genes Dev. 13, (in press). Wilkinson, D. G., Bhatt, S. & Herrmann, B. G Expression pattern of the mouse T gene and its role in mesoderm formation. Nature 343, Yasuo, H. & Satoh, N Function of vertebrate T gene. Nature 364, Yasuo, H. & Satoh, N An ascidian homolog of the mouse Brachyury (T) gene is expressed exclusively in notochord cells at the fate restricted stage. Develop. Growth Differ. 36, Yasuo, H. & Satoh, N Conservation of the developmental role of Brachyury in notochord formation in a urochordate, the ascidian Halocynthia roretzi. Dev. Biol. 200,
Brachyury downstream notochord differentiation in the ascidian embryo
 RESEARCH COMMUNICATION Brachyury downstream notochord differentiation in the ascidian embryo Hiroki Takahashi, 1,4 Kohji Hotta, 1,4 Albert Erives, 2,4 Anna Di Gregorio, 2 Robert W. Zeller, 3 Michael Levine,
RESEARCH COMMUNICATION Brachyury downstream notochord differentiation in the ascidian embryo Hiroki Takahashi, 1,4 Kohji Hotta, 1,4 Albert Erives, 2,4 Anna Di Gregorio, 2 Robert W. Zeller, 3 Michael Levine,
Spatio-Temporal Expression Patterns of Eight Epidermis-Specific Genes in the Ascidian Embryo
 Spatio-Temporal Expression Patterns of Eight Epidermis-Specific Genes in the Ascidian Embryo Author(s): Kouichi Ishida, Tatsuya Ueki, and Noriyuki Satoh Source: Zoological Science, 13(5):699-709. Published
Spatio-Temporal Expression Patterns of Eight Epidermis-Specific Genes in the Ascidian Embryo Author(s): Kouichi Ishida, Tatsuya Ueki, and Noriyuki Satoh Source: Zoological Science, 13(5):699-709. Published
Brachyury expression in ascidian embryos
 Development 126, 3725-3734 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV5331 3725 Evolutionary alterations of the minimal promoter for notochord-specific Brachyury expression
Development 126, 3725-3734 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV5331 3725 Evolutionary alterations of the minimal promoter for notochord-specific Brachyury expression
An essential role of a FoxD gene in notochord induction in Ciona embryos
 Development 129, 3441-3453 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV5028 3441 An essential role of a FoxD gene in notochord induction in Ciona embryos Kaoru S. Imai*, Nori
Development 129, 3441-3453 (2002) Printed in Great Britain The Company of Biologists Limited 2002 DEV5028 3441 An essential role of a FoxD gene in notochord induction in Ciona embryos Kaoru S. Imai*, Nori
posterior end mark, a novel maternal gene encoding a localized factor in the
 Development 122, 2005-2012 (1996) Printed in Great Britain The Company of Biologists Limited 1996 DEV5066 2005 posterior end mark, a novel maternal gene encoding a localized factor in the ascidian embryo
Development 122, 2005-2012 (1996) Printed in Great Britain The Company of Biologists Limited 1996 DEV5066 2005 posterior end mark, a novel maternal gene encoding a localized factor in the ascidian embryo
On the a clock' mechanism determining the time of tissue-specific enzyme development during ascidian embryogenesis
 /. Embryo!, exp. Morph. Vol. 54, pp. 131-139, 1979 Printed in Great Britain Company of Biologists Limited 1979 On the a clock' mechanism determining the time of tissue-specific enzyme development during
/. Embryo!, exp. Morph. Vol. 54, pp. 131-139, 1979 Printed in Great Britain Company of Biologists Limited 1979 On the a clock' mechanism determining the time of tissue-specific enzyme development during
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
doi:10.1038/nature11589 Supplementary Figure 1 Ciona intestinalis and Petromyzon marinus neural crest expression domain comparison. Cartoon shows dorsal views of Ciona mid gastrula (left) and Petromyzon
Developmental Biology Lecture Outlines
 Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Developmental Biology Lecture Outlines Lecture 01: Introduction Course content Developmental Biology Obsolete hypotheses Current theory Lecture 02: Gametogenesis Spermatozoa Spermatozoon function Spermatozoon
Research article. Summary. Introduction. Kaoru S. Imai*, Kyosuke Hino*, Kasumi Yagi, Nori Satoh and Yutaka Satou
 Research article 4047 Gene expression profiles of transcription factors and signaling molecules in the ascidian embryo: towards a comprehensive understanding of gene networks Kaoru S. Imai*, Kyosuke Hino*,
Research article 4047 Gene expression profiles of transcription factors and signaling molecules in the ascidian embryo: towards a comprehensive understanding of gene networks Kaoru S. Imai*, Kyosuke Hino*,
1. What are the three general areas of the developing vertebrate limb? 2. What embryonic regions contribute to the developing limb bud?
 Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Study Questions - Lecture 17 & 18 1. What are the three general areas of the developing vertebrate limb? The three general areas of the developing vertebrate limb are the proximal stylopod, zeugopod, and
Exam 1 ID#: October 4, 2007
 Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Biology 4361 Name: KEY Exam 1 ID#: October 4, 2007 Multiple choice (one point each) (1-25) 1. The process of cells forming tissues and organs is called a. morphogenesis. b. differentiation. c. allometry.
Characterization of a Maternal T-Box Gene in Ciona intestinalis
 Developmental Biology 225, 169 178 (2000) doi:10.1006/dbio.2000.9815, available online at http://www.idealibrary.com on Characterization of a Maternal T-Box Gene in Ciona intestinalis Albert Erives 1 and
Developmental Biology 225, 169 178 (2000) doi:10.1006/dbio.2000.9815, available online at http://www.idealibrary.com on Characterization of a Maternal T-Box Gene in Ciona intestinalis Albert Erives 1 and
Chapter 18 Lecture. Concepts of Genetics. Tenth Edition. Developmental Genetics
 Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Chapter 18 Lecture Concepts of Genetics Tenth Edition Developmental Genetics Chapter Contents 18.1 Differentiated States Develop from Coordinated Programs of Gene Expression 18.2 Evolutionary Conservation
Questions in developmental biology. Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration
 Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Questions in developmental biology Differentiation Morphogenesis Growth/apoptosis Reproduction Evolution Environmental integration Representative cell types of a vertebrate zygote => embryo => adult differentiation
Maternal Control of GermLayer Formation in Xenopus
 Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Maternal Control of GermLayer Formation in Xenopus The zygotic genome is activated at the mid-blastula transition mid-blastula fertilized egg Xenopus gastrulae early-gastrula 7 hrs 10 hrs control not VP
Question Set # 4 Answer Key 7.22 Nov. 2002
 Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Question Set # 4 Answer Key 7.22 Nov. 2002 1) A variety of reagents and approaches are frequently used by developmental biologists to understand the tissue interactions and molecular signaling pathways
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Sonic hedgehog (Shh) signalling in the rabbit embryo
 Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Sonic hedgehog (Shh) signalling in the rabbit embryo In the first part of this thesis work the physical properties of cilia-driven leftward flow were characterised in the rabbit embryo. Since its discovery
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its
 Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
Supplementary Figure 1: Mechanism of Lbx2 action on the Wnt/ -catenin signalling pathway. (a) The Wnt/ -catenin signalling pathway and its transcriptional activity in wild-type embryo. A gradient of canonical
9/4/2015 INDUCTION CHAPTER 1. Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology. Fig 1.
 INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
INDUCTION CHAPTER 1 Neurons are similar across phyla Thus, many different model systems are used in developmental neurobiology Fig 1.1 1 EVOLUTION OF METAZOAN BRAINS GASTRULATION MAKING THE 3 RD GERM LAYER
Temporal regulation of the muscle gene cascade by Macho1 and Tbx6 transcription factors in Ciona intestinalis
 Research Article 2453 Temporal regulation of the muscle gene cascade by Macho1 and Tbx6 transcription factors in Ciona intestinalis Jamie E. Kugler 1, Stefan Gazdoiu 1, Izumi Oda-Ishii 1, Yale J. Passamaneck
Research Article 2453 Temporal regulation of the muscle gene cascade by Macho1 and Tbx6 transcription factors in Ciona intestinalis Jamie E. Kugler 1, Stefan Gazdoiu 1, Izumi Oda-Ishii 1, Yale J. Passamaneck
Direct activation by Ets and Zic is required for initial expression of the Brachyury gene in the ascidian notochord
 Developmental Biology 306 (2007) 870 882 www.elsevier.com/locate/ydbio Genomes & Developmental Control Direct activation by Ets and Zic is required for initial expression of the Brachyury gene in the ascidian
Developmental Biology 306 (2007) 870 882 www.elsevier.com/locate/ydbio Genomes & Developmental Control Direct activation by Ets and Zic is required for initial expression of the Brachyury gene in the ascidian
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018
 Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Mesoderm Induction CBT, 2018 Hand-out CBT March 2018 Introduction 3. Books This module is based on the following books: - 'Principles of Developement', Lewis Wolpert, et al., fifth edition, 2015 - 'Developmental
Cell Cell Communication in Development
 Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
Biology 4361 Developmental Biology Cell Cell Communication in Development June 25, 2008 Cell Cell Communication Concepts Cells in developing organisms develop in the context of their environment, including
AP Biology Gene Regulation and Development Review
 AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
AP Biology Gene Regulation and Development Review 1. What does the regulatory gene code for? 2. Is the repressor by default active/inactive? 3. What changes the repressor activity? 4. What does repressor
Biology 218, practise Exam 2, 2011
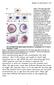 Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Figure 3 The long-range effect of Sqt does not depend on the induction of the endogenous cyc or sqt genes. a, Design and predictions for the experiments shown in b-e. b-e, Single-cell injection of 4 pg
Drosophila melanogaster- Morphogen Gradient
 NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
NPTEL Biotechnology - Systems Biology Drosophila melanogaster- Morphogen Gradient Dr. M. Vijayalakshmi School of Chemical and Biotechnology SASTRA University Joint Initiative of IITs and IISc Funded by
Chapter 11. Development: Differentiation and Determination
 KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
KAP Biology Dept Kenyon College Differential gene expression and development Mechanisms of cellular determination Induction Pattern formation Chapter 11. Development: Differentiation and Determination
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development June 23, 2009 Concepts Cell-Cell Communication Cells develop in the context of their environment, including: - their immediate
Biology 4361 - Developmental Biology Cell-Cell Communication in Development June 23, 2009 Concepts Cell-Cell Communication Cells develop in the context of their environment, including: - their immediate
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 2, 2006 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Life Sciences For NET & SLET Exams Of UGC-CSIR. Section B and C. Volume-08. Contents A. BASIC CONCEPT OF DEVELOPMENT 1
 Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Section B and C Volume-08 Contents 5. DEVELOPMENTAL BIOLOGY A. BASIC CONCEPT OF DEVELOPMENT 1 B. GAMETOGENESIS, FERTILIZATION AND EARLY DEVELOPMENT 23 C. MORPHOGENESIS AND ORGANOGENESIS IN ANIMALS 91 0
Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate
 Development 124, 2335-2344 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV9537 2335 Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate Joseph
Development 124, 2335-2344 (1997) Printed in Great Britain The Company of Biologists Limited 1997 DEV9537 2335 Dorsoventral patterning of the vertebrate neural tube is conserved in a protochordate Joseph
Ascidian Laboratory Woods Hole Embryology Course, 2010
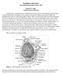 Ascidian Laboratory Woods Hole Embryology Course, 2010 Robert W. Zeller rzeller@sciences.sdsu.edu Ascidians have a long history as an experimental model system for development. Edwin Grant Conklin followed
Ascidian Laboratory Woods Hole Embryology Course, 2010 Robert W. Zeller rzeller@sciences.sdsu.edu Ascidians have a long history as an experimental model system for development. Edwin Grant Conklin followed
MBios 401/501: Lecture 14.2 Cell Differentiation I. Slide #1. Cell Differentiation
 MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
MBios 401/501: Lecture 14.2 Cell Differentiation I Slide #1 Cell Differentiation Cell Differentiation I -Basic principles of differentiation (p1305-1320) -C-elegans (p1321-1327) Cell Differentiation II
Mutations affecting tail and notochord development in the ascidian Ciona
 Development 126, 3293-3301 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV6405 3293 Mutations affecting tail and notochord development in the ascidian Ciona savignyi Yuki Nakatani,
Development 126, 3293-3301 (1999) Printed in Great Britain The Company of Biologists Limited 1999 DEV6405 3293 Mutations affecting tail and notochord development in the ascidian Ciona savignyi Yuki Nakatani,
Early Development in Invertebrates
 Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Developmental Biology Biology 4361 Early Development in Invertebrates October 25, 2006 Early Development Overview Cleavage rapid cell divisions divisions of fertilized egg into many cells Gastrulation
Unicellular: Cells change function in response to a temporal plan, such as the cell cycle.
 Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
Spatial organization is a key difference between unicellular organisms and metazoans Unicellular: Cells change function in response to a temporal plan, such as the cell cycle. Cells differentiate as a
T-box Genes in Early Embryogenesis
 DEVELOPMENTAL DYNAMICS 229:201 218, 2004 REVIEWS A PEER REVIEWED FORUM T-box Genes in Early Embryogenesis Chris Showell, Olav Binder, and Frank L. Conlon* The T-box gene family, encoding related DNA-binding
DEVELOPMENTAL DYNAMICS 229:201 218, 2004 REVIEWS A PEER REVIEWED FORUM T-box Genes in Early Embryogenesis Chris Showell, Olav Binder, and Frank L. Conlon* The T-box gene family, encoding related DNA-binding
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
Developmental Biology Biology 4361 Axis Specification in Drosophila November 6, 2007 Axis Specification in Drosophila Fertilization Superficial cleavage Gastrulation Drosophila body plan Oocyte formation
b. The maximum binding will decrease.
 Cell Signaling Receptors are a. proteins that change conformation upon interaction with a stimulus b. genes that change expression in response to a stimulus c. phosphorylation cascades that control cellular
Cell Signaling Receptors are a. proteins that change conformation upon interaction with a stimulus b. genes that change expression in response to a stimulus c. phosphorylation cascades that control cellular
Regulation of the Number of Cell Division Rounds by Tissue-Specific Transcription Factors and Cdk Inhibitor during Ascidian Embryogenesis
 Regulation of the Number of Cell Division Rounds by Tissue-Specific Transcription Factors and Cdk Inhibitor during Ascidian Embryogenesis Mami Kuwajima, Gaku Kumano, Hiroki Nishida* Department of Biological
Regulation of the Number of Cell Division Rounds by Tissue-Specific Transcription Factors and Cdk Inhibitor during Ascidian Embryogenesis Mami Kuwajima, Gaku Kumano, Hiroki Nishida* Department of Biological
Regionality of egg cytoplasm that promotes muscle differentiation in embryo of the ascidian, Halocynthia roretzi
 Development 116, 521-529 (1992) Printed in Great Britain The Company of Biologists Limited 1992 521 Regionality of egg cytoplasm that promotes muscle differentiation in embryo of the ascidian, Halocynthia
Development 116, 521-529 (1992) Printed in Great Britain The Company of Biologists Limited 1992 521 Regionality of egg cytoplasm that promotes muscle differentiation in embryo of the ascidian, Halocynthia
Supplementary Materials for
 www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
www.sciencesignaling.org/cgi/content/full/6/301/ra98/dc1 Supplementary Materials for Regulation of Epithelial Morphogenesis by the G Protein Coupled Receptor Mist and Its Ligand Fog Alyssa J. Manning,
Developmental Biology 3230 Midterm Exam 1 March 2006
 Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Name Developmental Biology 3230 Midterm Exam 1 March 2006 1. (20pts) Regeneration occurs to some degree to most metazoans. When you remove the head of a hydra a new one regenerates. Graph the inhibitor
Chapter 18 Regulation of Gene Expression
 Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Chapter 18 Regulation of Gene Expression Differential gene expression Every somatic cell in an individual organism contains the same genetic information and replicated from the same original fertilized
Conclusions. The experimental studies presented in this thesis provide the first molecular insights
 C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
C h a p t e r 5 Conclusions 5.1 Summary The experimental studies presented in this thesis provide the first molecular insights into the cellular processes of assembly, and aggregation of neural crest and
Novel Endostyle-Specific Genes in the Ascidian Ciona intestinalis
 ZOOLOGICAL SCIENCE 20: 1025 1030 (2003) 2003 Zoological Society of Japan Novel Endostyle-Specific Genes in the Ascidian Ciona intestinalis Akane Sasaki 1 *, Yuki Miyamoto 2, Yutaka Satou 1, Nori Satoh
ZOOLOGICAL SCIENCE 20: 1025 1030 (2003) 2003 Zoological Society of Japan Novel Endostyle-Specific Genes in the Ascidian Ciona intestinalis Akane Sasaki 1 *, Yuki Miyamoto 2, Yutaka Satou 1, Nori Satoh
PRACTICE EXAM. 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos.
 PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
PRACTICE EXAM 20 pts: 1. With the aid of a diagram, indicate how initial dorsal-ventral polarity is created in fruit fly and frog embryos. No Low [] Fly Embryo Embryo Non-neural Genes Neuroectoderm Genes
Nature Neuroscience: doi: /nn.2662
 Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
Supplementary Figure 1 Atlastin phylogeny and homology. (a) Maximum likelihood phylogenetic tree based on 18 Atlastin-1 sequences using the program Quicktree. Numbers at internal nodes correspond to bootstrap
8/23/2014. Introduction to Animal Diversity
 Introduction to Animal Diversity Chapter 32 Objectives List the characteristics that combine to define animals Summarize key events of the Paleozoic, Mesozoic, and Cenozoic eras Distinguish between the
Introduction to Animal Diversity Chapter 32 Objectives List the characteristics that combine to define animals Summarize key events of the Paleozoic, Mesozoic, and Cenozoic eras Distinguish between the
Maternal VegT and b-catenin: Patterning the Xenopus Blastula
 CHAPTER 1 Maternal VegT and b-catenin: Patterning the Xenopus Blastula Matthew Kofron, Jennifer Xanthos, and Janet Heasman 1 1.1 Introduction Loss of the maternal T-box transcription factor VegT has a
CHAPTER 1 Maternal VegT and b-catenin: Patterning the Xenopus Blastula Matthew Kofron, Jennifer Xanthos, and Janet Heasman 1 1.1 Introduction Loss of the maternal T-box transcription factor VegT has a
7.013 Problem Set
 7.013 Problem Set 5-2013 Question 1 During a summer hike you suddenly spot a huge grizzly bear. This emergency situation triggers a fight or flight response through a signaling pathway as shown below.
7.013 Problem Set 5-2013 Question 1 During a summer hike you suddenly spot a huge grizzly bear. This emergency situation triggers a fight or flight response through a signaling pathway as shown below.
Bio 127 Section I Introduction to Developmental Biology. Cell Cell Communication in Development. Developmental Activities Coordinated in this Way
 Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Bio 127 Section I Introduction to Developmental Biology Cell Cell Communication in Development Gilbert 9e Chapter 3 It has to be EXTREMELY well coordinated for the single celled fertilized ovum to develop
Expression of zebrafish goosecoid and no tail gene products in wild-type and mutant no tail embryos
 Development 120, 843-852 (1994) Printed in Great Britain The Company of Biologists Limited 1994 843 Expression of zebrafish goosecoid and no tail gene products in wild-type and mutant no tail embryos S.
Development 120, 843-852 (1994) Printed in Great Britain The Company of Biologists Limited 1994 843 Expression of zebrafish goosecoid and no tail gene products in wild-type and mutant no tail embryos S.
Exam 4 ID#: July 7, 2008
 Biology 4361 Name: KEY Exam 4 ID#: July 7, 2008 Multiple choice (one point each; indicate the best answer) 1. RNA polymerase II is not able to transcribe RNA unless a. it is first bound to TFIIB. b. its
Biology 4361 Name: KEY Exam 4 ID#: July 7, 2008 Multiple choice (one point each; indicate the best answer) 1. RNA polymerase II is not able to transcribe RNA unless a. it is first bound to TFIIB. b. its
!!!!!!!! DB3230 Midterm 2 12/13/2013 Name:
 1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
1. (10 pts) Draw or describe the fate map of a late blastula stage sea urchin embryo. Draw or describe the corresponding fate map of the pluteus stage larva. Describe the sequence of gastrulation events
Chapter 10 Development and Differentiation
 Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Part III Organization of Cell Populations Chapter Since ancient times, people have wondered how organisms are formed during the developmental process, and many researchers have worked tirelessly in search
Regulation and signaling. Overview. Control of gene expression. Cells need to regulate the amounts of different proteins they express, depending on
 Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
Regulation and signaling Overview Cells need to regulate the amounts of different proteins they express, depending on cell development (skin vs liver cell) cell stage environmental conditions (food, temperature,
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis
 18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
18.4 Embryonic development involves cell division, cell differentiation, and morphogenesis An organism arises from a fertilized egg cell as the result of three interrelated processes: cell division, cell
Exam 2 ID#: November 9, 2006
 Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Biology 4361 Name: KEY Exam 2 ID#: November 9, 2006 Multiple choice (one point each) Circle the best answer. 1. Inducers of Xenopus lens and optic vesicle include a. pharyngeal endoderm and anterior neural
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8
 Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Cellular Neurobiology BIPN 140 Fall 2016 Problem Set #8 1. Inductive signaling is a hallmark of vertebrate and mammalian development. In early neural development, there are multiple signaling pathways
Control of Gene Expression
 Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
Control of Gene Expression Mechanisms of Gene Control Gene Control in Eukaryotes Master Genes Gene Control In Prokaryotes Epigenetics Gene Expression The overall process by which information flows from
RNA Synthesis and Processing
 RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
RNA Synthesis and Processing Introduction Regulation of gene expression allows cells to adapt to environmental changes and is responsible for the distinct activities of the differentiated cell types that
2. Fertilization activates the egg and bring together the nuclei of sperm and egg
 2. Fertilization activates the egg and bring together the nuclei of sperm and egg Sea urchins (what phylum?) are models for the study of the early development of deuterostomes (like us, right?). Sea urchin
2. Fertilization activates the egg and bring together the nuclei of sperm and egg Sea urchins (what phylum?) are models for the study of the early development of deuterostomes (like us, right?). Sea urchin
3.a.2- Cell Cycle and Meiosis
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. 3.a.2- Cell Cycle and Meiosis EU 3.A: Heritable information provides for continuity of life.
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. 3.a.2- Cell Cycle and Meiosis EU 3.A: Heritable information provides for continuity of life.
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Biology 4361 Paraxial and Intermediate Mesoderm December 7, 2006 Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of BMPs more lateral mesoderm
Temporal regulation of the muscle gene cascade by Macho1 and Tbx6 transcription. factors in Ciona intestinalis.
 Temporal regulation of the muscle gene cascade by Macho1 and Tbx6 transcription factors in Ciona intestinalis. Jamie E. Kugler 1, Stefan Gazdoiu 1, Izumi Oda-Ishii 1, Yale J. Passamaneck 1, Albert J. Erives
Temporal regulation of the muscle gene cascade by Macho1 and Tbx6 transcription factors in Ciona intestinalis. Jamie E. Kugler 1, Stefan Gazdoiu 1, Izumi Oda-Ishii 1, Yale J. Passamaneck 1, Albert J. Erives
Role of Organizer Chages in Late Frog Embryos
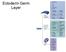 Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Ectoderm Germ Layer Frog Fate Map Frog Fate Map Role of Organizer Chages in Late Frog Embryos Organizer forms three distinct regions Notochord formation in chick Beta-catenin localization How does beta-catenin
Lecture 7. Development of the Fruit Fly Drosophila
 BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
BIOLOGY 205/SECTION 7 DEVELOPMENT- LILJEGREN Lecture 7 Development of the Fruit Fly Drosophila 1. The fruit fly- a highly successful, specialized organism a. Quick life cycle includes three larval stages
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Tuesday, December 27, 16
 Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Big Idea 3: Living systems store, retrieve, transmit and respond to information essential to life processes. Enduring understanding 3.B: Expression of genetic information involves cellular and molecular
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Cell Biology Review. The key components of cells that concern us are as follows: 1. Nucleus
 Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Cell Biology Review Development involves the collective behavior and activities of cells, working together in a coordinated manner to construct an organism. As such, the regulation of development is intimately
Neural development its all connected
 Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Neural development its all connected How do you build a complex nervous system? How do you build a complex nervous system? 1. Learn how tissue is instructed to become nervous system. Neural induction 2.
Paraxial and Intermediate Mesoderm
 Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Biology 4361 Paraxial and Intermediate Mesoderm December 6, 2007 Mesoderm Formation Chick Major Mesoderm Lineages Mesodermal subdivisions are specified along a mediolateral axis by increasing amounts of
Mechanisms of Human Health and Disease. Developmental Biology
 Mechanisms of Human Health and Developmental Biology Joe Schultz joe.schultz@nationwidechildrens.org D6 1 Dev Bio: Mysteries How do fertilized eggs become adults? How do adults make more adults? Why and
Mechanisms of Human Health and Developmental Biology Joe Schultz joe.schultz@nationwidechildrens.org D6 1 Dev Bio: Mysteries How do fertilized eggs become adults? How do adults make more adults? Why and
Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus:
 m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
m Eukaryotic mrna processing Newly made RNA is called primary transcript and is modified in three ways before leaving the nucleus: Cap structure a modified guanine base is added to the 5 end. Poly-A tail
Axis Specification in Drosophila
 Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Developmental Biology Biology 4361 Axis Specification in Drosophila July 9, 2008 Drosophila Development Overview Fertilization Cleavage Gastrulation Drosophila body plan Oocyte formation Genetic control
Development of Drosophila
 Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
Development of Drosophila Hand-out CBT Chapter 2 Wolpert, 5 th edition March 2018 Introduction 6. Introduction Drosophila melanogaster, the fruit fly, is found in all warm countries. In cooler regions,
the expression of alkaline PHosPHatase in Colonial ascidians
 the expression of alkaline PHosPHatase in Colonial ascidians Katherine Januszewicz, Heidi Swangler, Student Authors Dr. Elena Keeling, Research Advisor abstract The goal of this project was to better understand
the expression of alkaline PHosPHatase in Colonial ascidians Katherine Januszewicz, Heidi Swangler, Student Authors Dr. Elena Keeling, Research Advisor abstract The goal of this project was to better understand
Supplementary Figures
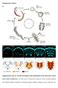 Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Supplementary Figures Supplementary Fig. S1: Normal development and organization of the embryonic ventral nerve cord in Platynereis. (A) Life cycle of Platynereis dumerilii. (B-F) Axonal scaffolds and
Early specification of ascidian larval motor neurons
 Developmental Biology 278 (2005) 310 322 www.elsevier.com/locate/ydbio Early specification of ascidian larval motor neurons Yu Katsuyama a,b, *, Toshiaki Okada a,c, Jun Matsumoto a,d, Yukio Ohtsuka a,
Developmental Biology 278 (2005) 310 322 www.elsevier.com/locate/ydbio Early specification of ascidian larval motor neurons Yu Katsuyama a,b, *, Toshiaki Okada a,c, Jun Matsumoto a,d, Yukio Ohtsuka a,
Is maternal mrna a determinant of tissue-specific proteins in ascidian embryos?
 J. Embryol. exp. Morph. 97 Supplement, 1-14 (1986) Printed in Great Britain The Company of Biologists Limited 1986 Is maternal mrna a determinant of tissue-specific proteins in ascidian embryos? W. R.
J. Embryol. exp. Morph. 97 Supplement, 1-14 (1986) Printed in Great Britain The Company of Biologists Limited 1986 Is maternal mrna a determinant of tissue-specific proteins in ascidian embryos? W. R.
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
SUPPLEMENTARY INFORMATION doi:1.138/nature1237 a b retinol retinal RA OH RDH (retinol dehydrogenase) O H Raldh2 O R/R.6.4.2 (retinaldehyde dehydrogenase 2) RA retinal retinol..1.1 1 Concentration (nm)
Axis determination in flies. Sem 9.3.B.5 Animal Science
 Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Axis determination in flies Sem 9.3.B.5 Animal Science All embryos are in lateral view (anterior to the left). Endoderm, midgut; mesoderm; central nervous system; foregut, hindgut and pole cells in yellow.
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A)
 Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Fig. S1. Proliferation and cell cycle exit are affected by the med mutation. (A,B) M-phase nuclei are visualized by a-ph3 labeling in wild-type (A) and mutant (B) 4 dpf retinae. The central retina of the
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday
 Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Complete all warm up questions Focus on operon functioning we will be creating operon models on Monday 1. What is the Central Dogma? 2. How does prokaryotic DNA compare to eukaryotic DNA? 3. How is DNA
Introduction. Gene expression is the combined process of :
 1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
1 To know and explain: Regulation of Bacterial Gene Expression Constitutive ( house keeping) vs. Controllable genes OPERON structure and its role in gene regulation Regulation of Eukaryotic Gene Expression
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION
 MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
MOLECULAR CONTROL OF EMBRYONIC PATTERN FORMATION Drosophila is the best understood of all developmental systems, especially at the genetic level, and although it is an invertebrate it has had an enormous
Advanced Higher Biology. Unit 1- Cells and Proteins 2c) Membrane Proteins
 Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
Advanced Higher Biology Unit 1- Cells and Proteins 2c) Membrane Proteins Membrane Structure Phospholipid bilayer Transmembrane protein Integral protein Movement of Molecules Across Membranes Phospholipid
Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio
 Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Urun 1 Comparison of the expression patterns of several sox genes between Oryzias latipes and Danio rerio Fatma Rabia URUN ilkent University, nkara - TURKEY High mobility group domain containing transcription
Biosc 41 9/10 Announcements
 Biosc 41 9/10 Announcements v Genetics review: group problem sets Groups of 3-4 Correct answer presented to class = 2 pts extra credit Incorrect attempt = 1 pt extra credit v Lecture: Animal Body Plans
Biosc 41 9/10 Announcements v Genetics review: group problem sets Groups of 3-4 Correct answer presented to class = 2 pts extra credit Incorrect attempt = 1 pt extra credit v Lecture: Animal Body Plans
FGF3 in the floor plate directs notochord convergent extension in the Ciona tadpole
 RESEARCH REPORT 23 Development 136, 23-28 (2009) doi:10.1242/dev.029157 FGF3 in the floor plate directs notochord convergent extension in the Ciona tadpole Weiyang Shi 1, *, Sara M. Peyrot 1, Edwin Munro
RESEARCH REPORT 23 Development 136, 23-28 (2009) doi:10.1242/dev.029157 FGF3 in the floor plate directs notochord convergent extension in the Ciona tadpole Weiyang Shi 1, *, Sara M. Peyrot 1, Edwin Munro
Lineage-Specific Regulation of the Ciona snail Gene in the Embryonic Mesoderm and Neuroectoderm
 DEVELOPMENTAL BIOLOGY 194, 213 225 (1998) ARTICLE NO. DB978810 Lineage-Specific Regulation of the Ciona snail Gene in the Embryonic Mesoderm and Neuroectoderm Albert Erives, 1 Joseph C. Corbo, 2 and Michael
DEVELOPMENTAL BIOLOGY 194, 213 225 (1998) ARTICLE NO. DB978810 Lineage-Specific Regulation of the Ciona snail Gene in the Embryonic Mesoderm and Neuroectoderm Albert Erives, 1 Joseph C. Corbo, 2 and Michael
Midterm 1. Average score: 74.4 Median score: 77
 Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
Midterm 1 Average score: 74.4 Median score: 77 NAME: TA (circle one) Jody Westbrook or Jessica Piel Section (circle one) Tue Wed Thur MCB 141 First Midterm Feb. 21, 2008 Only answer 4 of these 5 problems.
7.013 Spring 2005 Problem Set 4
 MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 4 FRIDAY April 8th,
MIT Department of Biology 7.013: Introductory Biology - Spring 2005 Instructors: Professor Hazel Sive, Professor Tyler Jacks, Dr. Claudette Gardel NAME TA 7.013 Spring 2005 Problem Set 4 FRIDAY April 8th,
Developmental genetics: finding the genes that regulate development
 Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Developmental Biology BY1101 P. Murphy Lecture 9 Developmental genetics: finding the genes that regulate development Introduction The application of genetic analysis and DNA technology to the study of
Outline. v Definition and major characteristics of animals v Dividing animals into groups based on: v Animal Phylogeny
 BIOSC 041 Overview of Animal Diversity: Animal Body Plans Reference: Chapter 32 Outline v Definition and major characteristics of animals v Dividing animals into groups based on: Body symmetry Tissues
BIOSC 041 Overview of Animal Diversity: Animal Body Plans Reference: Chapter 32 Outline v Definition and major characteristics of animals v Dividing animals into groups based on: Body symmetry Tissues
Nodal regulates neural tube formation in the Ciona intestinalis. embryo. Kaoru Mita* and Shigeki Fujiwara
 Nodal regulates neural tube formation in the Ciona intestinalis embryo Kaoru Mita* and Shigeki Fujiwara Department of Materials Science, Kochi University, 2-5-1 Akebono-cho, Kochi-shi, Kochi 780-8520,
Nodal regulates neural tube formation in the Ciona intestinalis embryo Kaoru Mita* and Shigeki Fujiwara Department of Materials Science, Kochi University, 2-5-1 Akebono-cho, Kochi-shi, Kochi 780-8520,
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning
 MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
MCDB 4777/5777 Molecular Neurobiology Lecture 29 Neural Development- In the beginning Learning Goals for Lecture 29 4.1 Describe the contributions of early developmental events in the embryo to the formation
v Scientists have identified 1.3 million living species of animals v The definition of an animal
 Biosc 41 9/10 Announcements BIOSC 041 v Genetics review: group problem sets Groups of 3-4 Correct answer presented to class = 2 pts extra credit Incorrect attempt = 1 pt extra credit v Lecture: Animal
Biosc 41 9/10 Announcements BIOSC 041 v Genetics review: group problem sets Groups of 3-4 Correct answer presented to class = 2 pts extra credit Incorrect attempt = 1 pt extra credit v Lecture: Animal
