High flexibility of DNA on short length scales probed by atomic force microscopy
|
|
|
- Clinton Joseph
- 5 years ago
- Views:
Transcription
1 High flexibility of DNA on short length scales probed by atomic force microscopy PAUL A. WIGGINS 1, THIJN VAN DER HEIJDEN, FERNANDO MORENO-HERRERO, ANDREW SPAKOWITZ 3, ROB PHILLIPS, JONATHAN WIDOM 5, CEES DEKKER AND PHILIP C. NELSON 6 * 1 Whitehead Institute, Cambridge, Massachusetts 1, USA Kavli Institute of NanoScience, Delft University of Technology, Lorentzweg 1, 68 CJ Delft, The Netherlands 3 Department of Chemical Engineering, Stanford University, Stanford, California 935, USA Division of Engineering and Applied Science, California Institute of Technology, Pasadena, California 9115, USA 5 Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, Illinois 68, USA 6 Department of Physics and Astronomy, University of Pennsylvania, Philadelphia, Pennsylvania 191, USA * nelson@physics.upenn.edu ARTICLES Published online: 3 November 6; doi:1.138/nnano.6.63 The mechanics of DNA bending on intermediate length scales (5 1 nm) plays a key role in many cellular processes, and is also important in the fabrication of artificial DNA structures, but previous experimental studies of DNA mechanics have focused on longer length scales than these. We use high-resolution atomic force microscopy on individual DNA molecules to obtain a direct measurement of the bending energy function appropriate for scales down to 5 nm. Our measurements imply that the elastic energy of highly bent DNA conformations is lower than predicted by classical elasticity models such as the worm-like chain () model. For example, we found that on short length scales, spontaneous large-angle are many times more prevalent than predicted by the model. We test our data and model with an interlocking set of consistency checks. Our analysis also shows how our model is compatible with previous experiments, which have sometimes been viewed as confirming the. The model of DNA conformation is an effective theory that idealizes the macromolecule as an inextensible elastic rod, and attributes to its bending deformations a classical (Hooke-lawtype) elastic energy cost. The has come to dominate physical discussions of double-stranded DNA mechanics, due in part to its simplicity and its successful description of experiments such as force spectroscopy on single DNA molecules 1 3. Because of these notable successes, classical elasticity models like the (and its generalizations to include twist stiffness) have been the framework for many studies of DNA mechanics. However, previous tests of the model using DNA stretching, atomic force microscopy (AFM) and other methods have been largely insensitive to the details of mechanics in the intermediate-scale regime crucial for cellular function, from chromosomal DNA packaging, to transcription, gene regulation and viral packaging 6. A quantitative understanding of such interactions requires a model of DNA bending that is valid on these biologically relevant length scales. In the following, we will argue that the successes of the model in fact arise from the long length scales probed by the classic experiments, rather than from any true underlying classical elasticity of DNA, and we will give short-scale measurements that do not agree with the predictions of the model. It may seem that a more general model, capable of embracing both our new data and the classic earlier experiments, would necessarily contain many more unknown parameters than the model. On the contrary, the model we propose is as simple as the. RESULTS THE The model can be formulated more precisely by representing each conformation as a chain of segments of length and assigning to it an elastic energy cost of the form E ¼ 1 k BT(j/ )(u i ), where u i is the angle between successive segment orientations t i and t iþ1 (in radians), k B T is the thermal energy, and j 5 nm is an effective elastic constant describing the chain s resistance to bending (see Supplementary Information). We call this bending-energy function classical (or harmonic ), because it is a quadratic function of the strain variable u i. The probability distribution of is then given by the Boltzmann distribution, g(t iþ1 jt i ) ¼ q 1 exp[e (u i )/k B T], where q is a normalization constant. The choice of a harmonic energy function E makes the angular distribution gaussian. However, even if E(u) is not harmonic, the angular distribution g(t iþn jt i ) will approach a gaussian form at large separations L ¼ n, because the iterated convolution of any distribution with itself converges to a gaussian form. That is, even if non-harmonic elastic behaviour is present, it will be hidden on long length scales by thermal fluctuations, nature nanotechnology VOL 1 NOVEMBER Nature Publishing Group
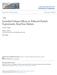
2 a b.9 1 nm a b cos θ s,s +L (R s,s +L ) (nm 1 ) Separation, L (nm) Height (nm) 5 nm 15 Position (nm) c In G (θ; 5 nm) Separation, L (nm), simulated c.5 1. Bend angle, θ (rad) nm L R Figure 1 High-resolution AFM images and tracing. Data shown were taken on V-grade mica with 1 mm Mg þ (see Supplementary Information for other surfaces and salt concentrations). a, A.5.5 mm AFM image (56 56 pixels) of,73-bp dsdna deposited on mica; z-range 1.5 nm. b, Crosssectional height profile of the DNA molecule, showing a half width at half maximum of only.5 nm. c, Detail of the square section indicated in a, showing also the chain of points determined by our automated image-analysis routine and illustrating the geometric quantities discussed in the text. Successive points are separated by ¼.5 nm. The end end distribution K(R; L ) is the probability distribution of real-space separation R among points separated by contour length L; in the example in the centre of the figure, we take L ¼.5 nm. The angle distribution G(u; L ) is the probability distribution of angles between tangents (short blue arrows) separated by L, which in the example on the right of the figure is 3.5 nm. giving rise to a theory that behaves like the. Thus, to investigate whether E(u) is harmonic, we must measure it directly on the length scale of interest, and not on some much longer scale. AFM MEASUREMENTS OF DNA CONTOURS We argued above that to look behind the model, we must examine length scales that are not much longer than the few nanometres scale of the molecule. It is difficult to obtain full 138 t t Figure Checks of equilibrium adsorption and failure of on short length scales. a,b, Plotsofk(R s,sþl ) l and kcos u s,sþl l from experimental data (dots), together with model-independent predictions assuming j ¼ 5 nm (curves). c, Dots: negative logarithm of the observed probability distribution function G(u; L ¼ 5 nm), a measure of the effective bending energy at this length scale in units of k B T. The graph is a histogram, computed using experimentally observed DNA contours with total length of about, nm, or a total of about 9, pairs of tangent vectors. The error bars represent expected p n error in bin populations due to finite sample size. Dashed curve: the same quantity for curves drawn from the distribution appropriate to the with persistence length equal to that used to draw the curves in (a and b). Although the two distributions agree qualitatively at low deflection u, they disagree at large angles: the model predicts far fewer such large deflections than were observed. Solid black curve: the same quantity when a sample of configurations was generated numerically and converted to simulated AFM data, then subjected to the same image analysis that yielded the experimental dots. three-dimensional views of equilibrium conformations of DNA in solution with the required resolution. However, under appropriate conditions, DNA adheres to a mica surface weakly enough that DNA mica interactions are believed not to affect the chain statistics 7. We will argue in the following that our molecules indeed adopt two-dimensional equilibrium conformations. We captured these by using AFM, following procedures outlined in ref. 8, but with significant improvements in image analysis (see Methods and Supplementary Information). Figure 1a is a representative image of DNA on mica, taken using tapping-mode AFM in air. The use of ultrasharp silicon tips allows a very high resolution to be obtained; for example, Fig. 1b displays a crosssectional DNA height profile with a half width at half maximum of only.5 nm. Despite our many precautions, we realized that potential gradients occur upon DNA adsorption that could potentially nature nanotechnology VOL 1 NOVEMBER Nature Publishing Group
3 generate spurious results. For this reason, we first made some detailed, model-independent theoretical predictions and checked that they were well obeyed by our data. As a first check, we measured the mean-square separation of pairs of points located at contour length s and s þ L from the end of the molecule, averaging over s and over all observed contours. We call this quantity k(r s,sþl ) l, and examine its behaviour as a function of the contour length separation L between the points (Fig. a). If the chain conformations are indeed equilibrated, and the effective elastic-energy function is local, then this function must take a particular form (see Supplementary Information). Indeed, as Fig. a shows, we found that the data follow this prediction very well. We can also define the tangent tangent correlation kcos u s,sþl l, where u s,sþl is the angle between tangent vectors at a pair of points separated by L. If the chains conformations are equilibrated, then this correlation must fall with contour separation L as e L/(j). Figure b shows that this prediction, too, is well satisfied, with j ¼ 5 nm. Together, these two tests confirm that, at least over length scales less than nm, our contours reflect equilibrium two-dimensional chain conformations, extending the observation of Rivetti et al. 7. The statistical measures in Fig. a, b are model-independent; they do not distinguish between different forms of the local elastic energy E(u). Therefore, we next measured the probability distribution function G(u; 5 nm) of various bend angles at points separated by contour length 5 nm. Figure c shows the negative logarithm of this histogram. The resulting curve measures the effective bending energy E(u)/k B T, coarse-grained to the length scale exp ¼ 5 nm probed by the experiment. As explained above, the model predicts that this function will be quadratic, regardless of the value of exp. Instead, Fig. c shows that the coarse-grained bending energy is far from being a quadratic function of the deflection angle u. At large angles we see instead that ln G(u) is nearly linear in u. These results can be stated differently by saying that large angular deflections between points separated by 5 nm were about 3 times more frequent in our data than the prediction of the model (Table 1). ANHARMONIC MODEL We constructed a local-elasticity model, with an effective bending-energy function chosen to mimic the behaviour seen in Fig. c. This model falls into a large class we have named sub-elastic chain (SEC), because it describes a polymer chain with a response to bending that, for large deflections, is softer than the usual harmonic model 9. One empirical energy function that summarizes the data is a linear SEC (LSEC): E LSEC ðuþ ¼ajujk B T; where a is a dimensionless constant that depends on the chosen segment length. We chose ¼.5 nm and adjusted a to fit the long-distance correlation G(u; 3 nm), yielding a ¼ 6.8. We then reasoned that, if indeed our contours represent equilibrium conformations of an elastic body, and in particular if successive chain elements are independently distributed according to equation (1), then the angle angle correlations at all separations L exp must be computable from that rule, with no further fitting. Figure 3 tests our predictions for G(u; L), and also for the end-to-end distribution K(R; L) defined in Fig. 1c. The remaining five curves in Fig. 3 are zero-fit-parameter predictions of our model and are observed to describe the data very well. The fact that our simple model passes the interlocking set of hurdles represented by the six curves in Fig. 3 is further strong evidence that we indeed measured spontaneous, equilibrium conformational fluctuations characterized by equation (1), and not an instrumental artifact or a surface-adsorption effect. Moreover, in keeping with the expectations raised earlier, these figures show that at large contour-length separations the distribution of bending angles does converge to that expected from the model. Our model correctly reproduces the existing successes of the model. For example, the force extension relation in our model is experimentally indistinguishable from that of the model, as is the rate of cyclization for linear constructs longer than a few persistence lengths (see Supplementary Information). ð1þ Table 1 Incidence of that are large (1.1 rad) or medium-large (.8 rad), at contour separations L 5 5 nm and 1 nm. The first three rows restate points made in the figures. Each of the columns labelled fraction show that our data disagree significantly with the predictions of the model (Fig. c), but agree with our model (Fig. 3a). The angles quoted refer to the angle between the vectors t and t in Fig. 1c. Row 1: experimental results from our main data. The large absolute numbers emphasize that our conclusions are not merely based on a handful of images. Row : expected results from a Monte Carlo evaluation of the model with persistence length j 5 5 nm. Row 3: expected results from Monte Carlo evaluation of our model with the same long-scale behaviour. The remaining rows show results of control experiments and a more detailed calculation (see Supplementary Information). Row : experimental data obtained using DNA incubated with ligase. Row 5: experimental data obtained when DNA was adsorbed to V1-grade mica. Row 6: numerical results when the configurations generated by the Monte Carlo code (row ) were converted to simulated AFM traces and then sent through our image analysis (solid curve in Fig. c and Supplementary Information, Fig. S8). Points separated by L ¼ 5 nm Points separated by L ¼ 1 nm pairs large 1 medium Experimental data 93, , ,1, ,88 3,15,187 17, LSEC equation (1),9,111, ,89 75,96,73 8, Experimental with 51, , ligase Experimental with 3, , V1-grade mica simulated, ,16 1, pairs large 1 nature nanotechnology VOL 1 NOVEMBER Nature Publishing Group
4 DISCUSSION a 1 In G (θ ; L) b In K (R ; L) 8 L = 5 nm L = 5 nm LSEC 1 θ (rad) LSEC L = 15 nm.7.8 L = 1 nm L = 3 nm L = 7.5 nm Figure 3 The non-harmonic elasticity model of equation (1) simultaneously fits many statistical properties of the experimental data. a, Negativelogarithm of the probability distribution function G(u; L ) for the angle u between tangents separated by contour length L, forl ¼ 5 nm (orange), 1 nm (purple) and 3 nm (blue). The dots are experimental data; the red dots, and the meaning of the error bars, are the same as in Fig. c. Solid curves: Monte Carlo evaluation of this correlation function in our model (equation (1)). Dashed curves: Monte Carlo evaluation of the same correlation in the model with persistence length j ¼ 5 nm. The dashed curves are all parabolas. The red dashed line is the same as in Fig. c. Although the model can reproduce the observed correlation at long separations, at short and medium separation it understates the prevalence of large-angle. b, Logarithm of the probability distribution function K(R ; L ), expressed in nm 1, for the real-space distance R between points separated by contour length L ¼ 7.5 nm (orange), 15 nm (purple) and 5 nm (blue). As in a, the model correctly captures the long-distance behaviour, but understates large bending (leading to large shortening) at separations less than about half a persistence length. Other authors have already reported that AFM images confirm that adsorbed DNA follows a two-dimensional distribution 7. Indeed we agree, when DNA is viewed on long length scales. However, we find significant deviations from the model on shorter, biologically relevant, length scales. It is possible in principle that surface adsorption and local defects on mica could conspire to modify the apparent elasticity of DNA. Our many checks and controls make this unlikely, however (for example, those shown in Fig. a, b). Other experiments not involving AFM, as well as molecular simulations, also point to a modification of the model qualitatively similar to the one we report here (see Supplementary Information). Our empirical effective bending-energy function, embodied in equation (1), is as simple as the model, and yet unlike the model it accurately describes the observed behaviour of DNA on multiple length scales, at both high and low curvatures. In particular, it is a useful starting point for the description of regulatory loops and other mesoscale DNA complexes. Our generic viewpoint, using models of the SEC type, may also be applicable to other stiff biopolymers. The form of the effective coarse-grained energy function that we find (equation (1)) may come as a surprise, but in fact there is a precedence for functions of this form. For example, 1 R /L.9 1. the overstretching transition of DNA reveals a plateau in the force extension relation, created by a transition between effective links of two different types, with different values of the rise per base pair 1. Another transition leads to a nearly constant axial torque as DNA under tension is twisted 11. Although these dramatic transitions occur in the laboratory at high external stresses, they will also occur spontaneously, albeit infrequently, and may in fact mediate important functions of DNA 1,13. By analogy, we can imagine a transition between two or more conformers with different bend angle (or stiffness), for example, the bent, but still base-paired, state constructed long ago by Crick and Klug 1. Increasing the bending stress on a tract of the molecule could then alter the coexistence between the conformers, leading to a plateau in the stress strain relation, or in other words an effectively linear energy function for bend. There may be a threshold for the onset of this approximately linear behaviour; the effective elastic energy function may have a harmonic (parabolic) region at small curvature. However, we found that the harmonic-elasticity regime, if any, is small (see Supplementary Information). In any case, the presence of such a regime is not needed to account for previously known facts about DNA mechanics (see Supplementary Information). An alternative possibility is that bare DNA elasticity may in fact be harmonic, but it is dressed into a non-harmonic form by electrostatic effects, which have already saturated at the lowest Mg þ concentrations we studied. For example, Rouzina and Bloomfield proposed that the presence of multivalent cations in solution could lead to transient kinking of DNA 15. Because their effect is non-local only over about six base pairs, it would appear local when coarse grained to length scales longer than about nm, and so can be described using the methods of this paper. Indeed, one benefit of our coarse-grained approach is that our empirical effective energy function is useful even before we resolve its mechanistic origin. One key application of this work is in biological systems, where DNA protein complexes are important for the molecularscale function of the cell. Many authors have assumed that for small loops, where thermal fluctuations may be neglected, classical harmonic rod elasticity is still applicable. On the contrary, we suggest that the model, when it is useful, actually owes its applicability to the existence of thermal fluctuations; when fluctuations can be neglected, then we must use a non-harmonic energy function like equation (1). In summary, we have argued that the short-length-scale bend distribution function must be measured directly, not extrapolated from long-length-scale measurements. We made such measurements, and showed that the probability of spontaneous sharp bending is orders of magnitude higher than predicted by the model. However, we found that a surprisingly simple coarse-grained elasticity theory is quantitatively accurate, both for describing spontaneous conformational fluctuations of DNA on length scales relevant for looping and nucleosome formation (5 nm and longer), and for the force extension of DNA and other long-length-scale phenomena. METHODS SAMPLE PREPARATION AND AFM DNA samples were prepared for AFM imaging by depositing 5 ng,73-bp linear double-stranded DNA (pgem-3z, Promega) diluted in 6 ml of 1 mm Tris HCl (ph 8.), supplemented with either 6, 1, 3 or 15 mm MgCl, onto freshly cleaved muscovite-form mica of grade V1 and V (SPI), following the methods of other authors 7,8,16. After approximately 3 s, the mica was washed with MilliQ-filtered water and blown dry in a gentle stream of nitrogen gas. Samples were imaged in air at room temperature and humidity with a NanoScope IIIa (Digital Instruments), operating in tapping mode with a nature nanotechnology VOL 1 NOVEMBER Nature Publishing Group
5 type-e scanner, with a pixel size (grid spacing) of 1.95 nm. Background correction consisted of fitting a second-order polynomial to every line in the AFM image. Tapping-mode SuperSharpSilicon tips, type SSS-NCH-8 (NanoSensors) were used. Some of the experiments were carried out using a commercial AFM from Nanotec Electronica operating in dynamic mode (see Supplementary Information). For both the Digital Instruments set-up and the configuration from Nanotec Electronica we obtained similar results, thus excluding artifacts caused by the imaging instrument. The surface and DNA were free of any salt deposits or protein impurities, and very clean images of DNA were obtained. Furthermore, we studied the effect of repairing possible nicks using ligase, examined various qualities of mica, and systematically varied the concentration of Mg þ in the solution. None of these variations altered our conclusions (see Supplementary Information and Table 1). We note that Mg þ concentrations at the lower end of the range we checked have been found to reduce the persistence length of DNA only slightly 17,18. IMAGE ANALYSIS Our image-analysis software was custom developed with the particular goal of analysing the local bend angles. The DNA molecules were traced automatically (but with human supervision) using a custom code in Matlab. Chain tracing was initiated at a user-determined initial point and trial tangent direction. (See Supplementary Information for a description of the algorithm.) Its output was a representation of the DNA contours as chains of xy pairs separated by contour length.5 nm (Fig. 1c). DATA ANALYSIS To reduce the effects of noise, we looked at the real-space separations of points separated by at least three steps (7.5 nm; see Fig. 3b), and the angular difference between tangent vectors defined by next-nearest neighbours and separated by at least two steps (5 nm; see Figs. 1c and 3a). We also applied the same procedures to virtual chains generated by our Monte Carlo code (see Supplementary Information), and made appropriate comparisons. We performed extensive tests to support our interpretation of the data. We checked by eye that our contour-tracking software faithfully followed the contours, and we sent simulated data, including noise and tip-induced broadening, through our image-processing and analysis software to confirm that the resulting -based contours did not generate distributions resembling our experimental data (see Fig. c and Supplementary Information). Moreover, we checked that the alternative hypothesis of non-equilibrium adsorption of a -distributed chain to the surface does not explain our experimental data (see Supplementary Information). Received 3 June 6; accepted 31 August 6; published 3 November 6. References 1. Bustamante, C., Marko, J. F., Siggia, E. D. & Smith, S. Entropic elasticity of lambda phage DNA. Science 65, (199).. Bustamante, C., Smith, S. B., Liphardt, J. & Smith, D. Single-molecule studies of DNA mechanics. Curr. Opin. Struct. Biol. 1, (). 3. Nelson, P. Biological Physics: Energy, Information, Life (W. H. Freeman, New York, ).. Widom, J. Role of DNA sequence in nucleosome stability and dynamics. Q. Rev. Biophys. 3, 69 3 (1). 5. Rippe, K., von Hippel, P. R. & Langowski, J. Action at a distance: DNA-looping and initiation of transcription. Trends Biochem. Sci., 5 56 (1995). 6. Shore, D., Langowski, J. & Baldwin, R. L. DNA flexibility studied by covalent closure of short fragments into circles. Proc. Natl Acad. Sci. USA 17, (1981). 7. Rivetti, C., Guthold, M. & Bustamante, C. Scanning force microscopy of DNA deposited onto mica: Equilibration versus kinetic trapping studied by statistical polymer chain analysis. J. Mol. Biol. 6, (1996). 8. van Noort, J. et al. The coiled-coil of the human rad5 DNA repair protein contains specific segments of increased flexibility. Proc. Natl Acad. Sci. USA 1, (3). 9. Wiggins, P. A. & Nelson, P. C. Generalized theory of semiflexible polymers. Phys. Rev. E 73, 3196 (6). 1. Storm, C. & Nelson, P. Theory of high-force DNA stretching and overstretching. Phys. Rev. E 67, 5196 (3). Erratum Phys. Rev E. 7, 139 (). 11. Bryant, Z. et al. Structural transitions and elasticity from torque measurements on DNA. Nature, (3). 1. Leger, J. F., Robert, J., Bourdieu, L., Chatenay, D. & Marko, J. F. RecA binding to a single double-stranded DNA molecule: A possible role of DNA conformational fluctuations. Proc. Natl Acad. Sci. USA 95, (1998). 13. Sobell, H. M., Tsai, C., Jain, S. C. & Gilbert, S. G. Visualization of drug nucleic acid interactions at atomic resolution. 3. Unifying structural concepts in understanding drug DNA interactions and their broader implications in understanding protein DNA interactions. J. Mol. Biol. 11, (1977). 1. Crick, F. H. C. & Klug, A. Kinky helix. Nature 55, (1975). 15. Rouzina, I. & Bloomfield, V. A. DNA bending by small, mobile multivalent cations. Biophys. J. 7, (1998). 16. Hansma, H., Revenko, I., Kim, K. & Laney, D. Atomic force microscopy of long and short double-stranded, single-stranded and triple-stranded nucleic acids. Nucleic Acids Res., (1996). 17. Wang, M. D., Yin, H., Landick, R., Gelles, J. & Block, S. M. Stretching DNA with optical tweezers. Biophys. J. 7, (1997). 18. van der Heijden, T. et al. Torque-limited RecA polymerization on dsdna. Nucleic Acids Res. 33, (5). Acknowledgements We thank N. R. Dan, N. H. Dekker, M. Inamdar, R. James, R. D. Kamien, I. Kulic, T. Laurence, R. Lavery, J. Maddocks J. Marko, A. Onufriev, P. Purohit, I. Rouzina, J. M. Schurr, R. Seidel, V. Soghomonian, Z.-G. Wang and Y. Zhang for helpful discussions and correspondence. P.A.W. acknowledges grant support from an NSF graduate fellowship. F.M.-H. acknowledges support from La Fundacien Ramon Areces as a postdoctoral fellow. A.J.S. acknowledges funding from the National Institutes of Health (NIH-GM7155). R.P. and P.A.W. acknowledge the Keck Foundation and NSF Grant CMS as well as the NSF-funded Center for Integrative Multiscale Modeling and Simulation. J.W. acknowledges support from NIH grants R1 GM569 and R1 GM C.D. acknowledges the Stichting voor Fundamenteel Onderzoek der Materie (FOM), which is financially supported by the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO). P.N. acknowledges NSF Grant DMR-67 and the NSF-funded NSEC on Molecular Function at the Nano/Bio Interface DMR-578. J.W., R.P. and P.C.N. acknowledge the hospitality of the Kavli Institute for Theoretical Physics, supported in part by the National Science Foundation under Grant PHY Correspondence and requests for materials should be addressed to P.C.N. Supplementary information accompanies this paper on Author contributions P.A.W., T.v.d.H., F.M.-H., C.D. and P.N. contributed to the experimental, theoretical and analysis strategy. T.v.d.H. and F.M.-H. carried out the AFM experiments, and with, C.D. and P.A.W. contributed to the custom image-processing software. P.A.W., T.v.d.H., F.M.-H., C.D. and P.C.N. contributed to the analysis. A.S. contributed the analysis and simulations of nonequilibrium adsorption. P.A.W., R.P., J.W. and P.C.N. contributed to the initial formulation of the hypothesis and its refinement. P.W., T.v.d.H., F.M.-H., A.S., C.D. and P.C.N. contributed to the written and graphical presentation; all authors discussed the results and commented on the manuscript. Competing financial interests The authors declare that they have no competing financial interests. Reprints and permission information is available online at nature nanotechnology VOL 1 NOVEMBER Nature Publishing Group
High flexibility of DNA on short length scales probed by atomic. force microscopy
 High flexibility of DNA on short length scales probed by atomic force microscopy Paul A. Wiggins, 1 Thijn van der Heijden, 2 Fernando Moreno Herrero, 2 Andrew Spakowitz, 3 Rob Phillips, 4 Jonathan Widom,
High flexibility of DNA on short length scales probed by atomic force microscopy Paul A. Wiggins, 1 Thijn van der Heijden, 2 Fernando Moreno Herrero, 2 Andrew Spakowitz, 3 Rob Phillips, 4 Jonathan Widom,
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Exact Theory of Kinkable Elastic Polymers
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 2-2005 Exact Theory of Kinkable Elastic Polymers Paul A. Wiggins Rob Phillips Philip C. Nelson University
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 2-2005 Exact Theory of Kinkable Elastic Polymers Paul A. Wiggins Rob Phillips Philip C. Nelson University
New Measurements of DNA Twist Elasticity
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA
 8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
Excluded-Volume Effects in Tethered-Particle Experiments: Bead Size Matters
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 3-2006 Excluded-Volume Effects in Tethered-Particle Experiments: Bead Size Matters Darren E. Segall Philip
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 3-2006 Excluded-Volume Effects in Tethered-Particle Experiments: Bead Size Matters Darren E. Segall Philip
Generalized theory of semiflexible polymers
 PHYSICAL REVIEW E 73, 031906 2006 Generalized theory of semiflexible polymers Paul A. Wiggins* Division of Physics, Mathematics, & Astronomy, California Institute of Technology, Pasadena, California 91125,
PHYSICAL REVIEW E 73, 031906 2006 Generalized theory of semiflexible polymers Paul A. Wiggins* Division of Physics, Mathematics, & Astronomy, California Institute of Technology, Pasadena, California 91125,
Curvature Distribution of Worm-like Chains in Two and Three Dimensions
 Curvature Distribution of Worm-like Chains in Two and Three Dimensions Shay M. Rappaport, Shlomi Medalion and Yitzhak Rabin Department of Physics, Bar-Ilan University, Ramat-Gan 59, Israel arxiv:8.383v
Curvature Distribution of Worm-like Chains in Two and Three Dimensions Shay M. Rappaport, Shlomi Medalion and Yitzhak Rabin Department of Physics, Bar-Ilan University, Ramat-Gan 59, Israel arxiv:8.383v
Magnetic tweezers and its application to DNA mechanics
 Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Exact theory of kinkable elastic polymers
 Exact theory of kinkable elastic polymers Paul A. Wiggins* Division of Physics, Mathematics, and Astronomy, California Institute of Technology, Pasadena, California 91125, USA Rob Phillips Division of
Exact theory of kinkable elastic polymers Paul A. Wiggins* Division of Physics, Mathematics, and Astronomy, California Institute of Technology, Pasadena, California 91125, USA Rob Phillips Division of
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
Atomic Force Microscopy Characterization of Room- Temperature Adlayers of Small Organic Molecules through Graphene Templating
 Atomic Force icroscopy Characterization of Room- Temperature Adlayers of Small Organic olecules through Graphene Templating Peigen Cao, Ke Xu,2, Joseph O. Varghese, and James R. Heath *. Kavli Nanoscience
Atomic Force icroscopy Characterization of Room- Temperature Adlayers of Small Organic olecules through Graphene Templating Peigen Cao, Ke Xu,2, Joseph O. Varghese, and James R. Heath *. Kavli Nanoscience
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
Chiral selection in wrapping, crossover, and braiding of DNA mediated by asymmetric bend-writhe elasticity
 http://www.aimspress.com/ AIMS Biophysics, 2(4): 666-694. DOI: 10.3934/biophy.2015.4.666 Received date 28 August 2015, Accepted date 29 October 2015, Published date 06 November 2015 Research article Chiral
http://www.aimspress.com/ AIMS Biophysics, 2(4): 666-694. DOI: 10.3934/biophy.2015.4.666 Received date 28 August 2015, Accepted date 29 October 2015, Published date 06 November 2015 Research article Chiral
DNA/RNA structure and packing
 DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
Supplementary Information. The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes
 Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Supplementary Information The Solution Structural Ensembles of RNA Kink-turn Motifs and Their Protein Complexes Xuesong Shi, a Lin Huang, b David M. J. Lilley, b Pehr B. Harbury a,c and Daniel Herschlag
Exact theory of kinkable elastic polymers
 Exact theory of kinkable elastic polymers Paul A. Wiggins, Rob Phillips, and Philip C. Nelson (Dated: August 31st, 2004) The importance of nonlinearities in material constitutive relations has long been
Exact theory of kinkable elastic polymers Paul A. Wiggins, Rob Phillips, and Philip C. Nelson (Dated: August 31st, 2004) The importance of nonlinearities in material constitutive relations has long been
Nature Methods: doi: /nmeth Supplementary Figure 1. Principle of force-distance (FD) curve based AFM.
 Supplementary Figure 1 Principle of force-distance (FD) curve based AFM. (a) FD-based AFM contours the sample surface while oscillating the AFM tip with a sine wave at a frequency of 0.25 khz. Pixel-by-pixel
Supplementary Figure 1 Principle of force-distance (FD) curve based AFM. (a) FD-based AFM contours the sample surface while oscillating the AFM tip with a sine wave at a frequency of 0.25 khz. Pixel-by-pixel
Weight and contact forces: Young's modulus, Hooke's law and material properties
 Weight and contact forces: Young's modulus, Hooke's law and material properties Many objects deform according to Hooke's law; many materials behave elastically and have a Young's modulus. In this section,
Weight and contact forces: Young's modulus, Hooke's law and material properties Many objects deform according to Hooke's law; many materials behave elastically and have a Young's modulus. In this section,
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi: 10.1038/nphys00 Peter Gross, Niels Laurens, Lene B. Oddershede, Ulrich Bockelmann 3, Erwin J. G. Peterman*, and Gijs J. L. Wuite* LaserLaB and Department of Physics and Astronomy,
SUPPLEMENTARY INFORMATION doi: 10.1038/nphys00 Peter Gross, Niels Laurens, Lene B. Oddershede, Ulrich Bockelmann 3, Erwin J. G. Peterman*, and Gijs J. L. Wuite* LaserLaB and Department of Physics and Astronomy,
S(l) bl + c log l + d, with c 1.8k B. (2.71)
 2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
SUPPLEMENTARY INFORMATION
 doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
doi:10.1038/nature09450 Supplementary Table 1 Summary of kinetic parameters. Kinetic parameters were V = V / 1 K / ATP and obtained using the relationships max ( + m [ ]) V d s /( 1/ k [ ATP] + 1 k ) =,
Supplementary Information
 Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Supplementary Information Switching of myosin-v motion between the lever-arm swing and Brownian search-and-catch Keisuke Fujita 1*, Mitsuhiro Iwaki 2,3*, Atsuko H. Iwane 1, Lorenzo Marcucci 1 & Toshio
Nucleosome Switching
 Nucleosome Switching David J. Schwab, Robijn F. Bruinsma, Joseph Rudnick Department of Physics and Astronomy, University of California, Los Angeles, Los Angeles, CA, 90024 & Jonathan Widom Department of
Nucleosome Switching David J. Schwab, Robijn F. Bruinsma, Joseph Rudnick Department of Physics and Astronomy, University of California, Los Angeles, Los Angeles, CA, 90024 & Jonathan Widom Department of
Electronic Supporting Information. Microcapsule Buckling Triggered by Compression- Induced Interfacial Chase Change
 Electronic Supporting Information Microcapsule Buckling Triggered by Compression- Induced Interfacial Chase Change Andrew R Salmon, 1 Richard M Parker, 1 Alexander S Groombridge, 1 Armando Maestro, 1 Roger
Electronic Supporting Information Microcapsule Buckling Triggered by Compression- Induced Interfacial Chase Change Andrew R Salmon, 1 Richard M Parker, 1 Alexander S Groombridge, 1 Armando Maestro, 1 Roger
2.4 DNA structure. S(l) bl + c log l + d, with c 1.8k B. (2.72)
 2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
2.4 DNA structure DNA molecules come in a wide range of length scales, from roughly 50,000 monomers in a λ-phage, 6 0 9 for human, to 9 0 0 nucleotides in the lily. The latter would be around thirty meters
A rule of seven in Watson-Crick base-pairing of mismatched sequences
 A rule of seven in Watson-Crick base-pairing of mismatched sequences Ibrahim I. Cisse 1,3, Hajin Kim 1,2, Taekjip Ha 1,2 1 Department of Physics and Center for the Physics of Living Cells, University of
A rule of seven in Watson-Crick base-pairing of mismatched sequences Ibrahim I. Cisse 1,3, Hajin Kim 1,2, Taekjip Ha 1,2 1 Department of Physics and Center for the Physics of Living Cells, University of
SUPPLEMENTARY INFORMATION
 1. Supplementary Methods Characterization of AFM resolution We employed amplitude-modulation AFM in non-contact mode to characterize the topography of the graphene samples. The measurements were performed
1. Supplementary Methods Characterization of AFM resolution We employed amplitude-modulation AFM in non-contact mode to characterize the topography of the graphene samples. The measurements were performed
Universal Scaling in Non-equilibrium Transport Through a Single- Channel Kondo Dot (EPAPS Supplementary Info)
 Universal Scaling in Non-equilibrium Transport Through a Single- Channel Kondo Dot (EPAPS Supplementary Info) M. Grobis, 1 I. G. Rau, 2 R. M. Potok, 1,3,* H. Shtrikman, 4 and D. Goldhaber-Gordon 1 1 Department
Universal Scaling in Non-equilibrium Transport Through a Single- Channel Kondo Dot (EPAPS Supplementary Info) M. Grobis, 1 I. G. Rau, 2 R. M. Potok, 1,3,* H. Shtrikman, 4 and D. Goldhaber-Gordon 1 1 Department
Basic Laboratory. Materials Science and Engineering. Atomic Force Microscopy (AFM)
 Basic Laboratory Materials Science and Engineering Atomic Force Microscopy (AFM) M108 Stand: 20.10.2015 Aim: Presentation of an application of the AFM for studying surface morphology. Inhalt 1.Introduction...
Basic Laboratory Materials Science and Engineering Atomic Force Microscopy (AFM) M108 Stand: 20.10.2015 Aim: Presentation of an application of the AFM for studying surface morphology. Inhalt 1.Introduction...
Measurements of interaction forces in (biological) model systems
 Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Measurements of interaction forces in (biological) model systems Marina Ruths Department of Chemistry, UMass Lowell What can force measurements tell us about a system? Depending on the technique, we might
Faculty of math and natural science. Universty of Split. Kratky-Porod model. Nataša Vučemilovid-Alagid. February, 2013.
 Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
SUPPLEMENTARY FIGURE 1. Force dependence of the unbinding rate: (a) Force-dependence
 (b) BSA-coated beads double exponential low force exponential high force exponential 1 unbinding time tb [sec] (a) unbinding time tb [sec] SUPPLEMENTARY FIGURES BSA-coated beads without BSA.2.1 5 1 load
(b) BSA-coated beads double exponential low force exponential high force exponential 1 unbinding time tb [sec] (a) unbinding time tb [sec] SUPPLEMENTARY FIGURES BSA-coated beads without BSA.2.1 5 1 load
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
Effect of Spontaneous Twist on DNA Minicircles
 Biophysical Journal Volume 99 November 2010 2987 2994 2987 Effect of Spontaneous Twist on DNA Minicircles Shlomi Medalion, * David A. Kessler, and Yitzhak Rabin { Department of Physics and Institute of
Biophysical Journal Volume 99 November 2010 2987 2994 2987 Effect of Spontaneous Twist on DNA Minicircles Shlomi Medalion, * David A. Kessler, and Yitzhak Rabin { Department of Physics and Institute of
Exchange of Counterions in DNA Condensation. Abstract
 Exchange of Counterions in DNA Condensation Yoshihiro Murayama and Masaki Sano Department of Physics, University of Tokyo, Tokyo 113-0033, Japan Abstract We measured the fluorescence intensity of DNA-bound
Exchange of Counterions in DNA Condensation Yoshihiro Murayama and Masaki Sano Department of Physics, University of Tokyo, Tokyo 113-0033, Japan Abstract We measured the fluorescence intensity of DNA-bound
Announcements. Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1.
 Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Elasticity of the human red blood cell skeleton
 Biorheology 40 (2003) 247 251 247 IOS Press Elasticity of the human red blood cell skeleton G. Lenormand, S. Hénon, A. Richert, J. Siméon and F. Gallet Laboratoire de Biorhéologie et d Hydrodynamique Physico-Chimique,
Biorheology 40 (2003) 247 251 247 IOS Press Elasticity of the human red blood cell skeleton G. Lenormand, S. Hénon, A. Richert, J. Siméon and F. Gallet Laboratoire de Biorhéologie et d Hydrodynamique Physico-Chimique,
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments
 Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
INTRODUCTION TO SINGLE-DNA MICROMECHANICS (AUGUST 9, 2004) John F. Marko. Department of Physics, University of Illinois at Chicago, Chicago, IL, USA
 INTRODUCTION TO SINGLE-DNA MICROMECHANICS AUGUST 9, 2004) John F. Marko Department of Physics, University of Illinois at Chicago, Chicago, IL, USA 1 Photo: width 7.5cm height 11cm Contents 1. Introduction
INTRODUCTION TO SINGLE-DNA MICROMECHANICS AUGUST 9, 2004) John F. Marko Department of Physics, University of Illinois at Chicago, Chicago, IL, USA 1 Photo: width 7.5cm height 11cm Contents 1. Introduction
Torsional directed walks, entropic elasticity, and DNA twist stiffness
 Proc. Natl. Acad. Sci. USA Vol. 94, pp. 14418 14422, December 1997 Biophysics Torsional directed walks, entropic elasticity, and DNA twist stiffness J. DAVID MOROZ AND PHILIP NELSON* Department of Physics
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 14418 14422, December 1997 Biophysics Torsional directed walks, entropic elasticity, and DNA twist stiffness J. DAVID MOROZ AND PHILIP NELSON* Department of Physics
Untangling the Mechanics of Entangled Biopolymers
 Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES
 Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
V = 2ze 2 n. . a. i=1
 IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
Explicit algebraic Reynolds stress models for internal flows
 5. Double Circular Arc (DCA) cascade blade flow, problem statement The second test case deals with a DCA compressor cascade, which is considered a severe challenge for the CFD codes, due to the presence
5. Double Circular Arc (DCA) cascade blade flow, problem statement The second test case deals with a DCA compressor cascade, which is considered a severe challenge for the CFD codes, due to the presence
Scanning Tunneling Microscopy and its Application
 Chunli Bai Scanning Tunneling Microscopy and its Application With 181 Figures SHANGHAI SCIENTIFIC & TECHNICAL PUBLISHERS Jpl Springer Contents 1. Introduction 1 1.1 Advantages of STM Compared with Other
Chunli Bai Scanning Tunneling Microscopy and its Application With 181 Figures SHANGHAI SCIENTIFIC & TECHNICAL PUBLISHERS Jpl Springer Contents 1. Introduction 1 1.1 Advantages of STM Compared with Other
SUPPLEMENTAL MATERIAL
 SUPPLEMENTAL MATERIAL Systematic Coarse-Grained Modeling of Complexation between Small Interfering RNA and Polycations Zonghui Wei 1 and Erik Luijten 1,2,3,4,a) 1 Graduate Program in Applied Physics, Northwestern
SUPPLEMENTAL MATERIAL Systematic Coarse-Grained Modeling of Complexation between Small Interfering RNA and Polycations Zonghui Wei 1 and Erik Luijten 1,2,3,4,a) 1 Graduate Program in Applied Physics, Northwestern
The bulk modulus of covalent random networks
 J. Phys.: Condens. Matter 9 (1997) 1983 1994. Printed in the UK PII: S0953-8984(97)77754-3 The bulk modulus of covalent random networks B R Djordjević and M F Thorpe Department of Physics and Astronomy
J. Phys.: Condens. Matter 9 (1997) 1983 1994. Printed in the UK PII: S0953-8984(97)77754-3 The bulk modulus of covalent random networks B R Djordjević and M F Thorpe Department of Physics and Astronomy
Measuring Colocalization within Fluorescence Microscopy Images
 from photonics.com: 03/01/2007 http://www.photonics.com/article.aspx?aid=39341 Measuring Colocalization within Fluorescence Microscopy Images Two-color fluorescence-based methods are uncovering molecular
from photonics.com: 03/01/2007 http://www.photonics.com/article.aspx?aid=39341 Measuring Colocalization within Fluorescence Microscopy Images Two-color fluorescence-based methods are uncovering molecular
Phys 450 Spring 2011 Solution set 6. A bimolecular reaction in which A and B combine to form the product P may be written as:
 Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Supporting Information. by Hexagonal Boron Nitride
 Supporting Information High Velocity Saturation in Graphene Encapsulated by Hexagonal Boron Nitride Megan A. Yamoah 1,2,, Wenmin Yang 1,3, Eric Pop 4,5,6, David Goldhaber-Gordon 1 * 1 Department of Physics,
Supporting Information High Velocity Saturation in Graphene Encapsulated by Hexagonal Boron Nitride Megan A. Yamoah 1,2,, Wenmin Yang 1,3, Eric Pop 4,5,6, David Goldhaber-Gordon 1 * 1 Department of Physics,
Origin of the Electrophoretic Force on DNA in a Nanopore
 Origin of the Electrophoretic Force on DNA in a Nanopore Stijn van Dorp 1 Ulrich F. Keyser 2, *Nynke H. Dekker 1, Cees Dekker 1, Serge G. Lemay 1 1 Kavli Institut of Nanoscience, Delft University of Technology,
Origin of the Electrophoretic Force on DNA in a Nanopore Stijn van Dorp 1 Ulrich F. Keyser 2, *Nynke H. Dekker 1, Cees Dekker 1, Serge G. Lemay 1 1 Kavli Institut of Nanoscience, Delft University of Technology,
Proteins polymer molecules, folded in complex structures. Konstantin Popov Department of Biochemistry and Biophysics
 Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Charge inversion accompanies DNA condensation. by multivalent ions. Construction, mechanics, and electronics. 11 May 2008.
 Charge inversion accompanies DNA condensation by multivalent ions DNA-based nanotechnology: Construction, mechanics, and electronics 11 May 2008 Serge Lemay Kavli Institute of Nanoscience Delft University
Charge inversion accompanies DNA condensation by multivalent ions DNA-based nanotechnology: Construction, mechanics, and electronics 11 May 2008 Serge Lemay Kavli Institute of Nanoscience Delft University
DNA supercoiling: plectonemes or curls?
 DNA supercoiling: plectonemes or curls? Sébastien Neukirch CNRS & UPMC Univ. Paris 6 (France) John F. Marko Northwestern University (IL. USA) Why study DNA mechanical properties? mechanical properties
DNA supercoiling: plectonemes or curls? Sébastien Neukirch CNRS & UPMC Univ. Paris 6 (France) John F. Marko Northwestern University (IL. USA) Why study DNA mechanical properties? mechanical properties
REMOVING A PIECE OF ADHESIVE TAPE FROM A HORIZONTAL SURFACE
 REMOVING A PIECE OF ADHESIVE TAPE FROM A HORIZONTAL SURFACE Hossein Azizinaghsh a, Hamid Ghaednia b a Sharif University of Technology, School of Computer Engineering, I. R. Iran Hossein.Azizi@gmail.com
REMOVING A PIECE OF ADHESIVE TAPE FROM A HORIZONTAL SURFACE Hossein Azizinaghsh a, Hamid Ghaednia b a Sharif University of Technology, School of Computer Engineering, I. R. Iran Hossein.Azizi@gmail.com
Supplementary Material
 1 2 3 Topological defects in confined populations of spindle-shaped cells by G. Duclos et al. Supplementary Material 4 5 6 7 8 9 10 11 12 13 Supplementary Note 1: Characteristic time associated with the
1 2 3 Topological defects in confined populations of spindle-shaped cells by G. Duclos et al. Supplementary Material 4 5 6 7 8 9 10 11 12 13 Supplementary Note 1: Characteristic time associated with the
TERRESTRIAL REDSHIFTS FROM A DIFFUSE LIGHT SOURCE. FOM Institute for Atomic and Molecular Physics, Kruislaan 407, 1098 SJ Amsterdam, The Netherlands
 March 25, 1999 TERRESTRIAL REDSHIFTS FROM A DIFFUSE LIGHT SOURCE Ad Lagendijk a FOM Institute for Atomic and Molecular Physics, Kruislaan 407, 1098 SJ Amsterdam, The Netherlands (Received ) Abstract Enhanced
March 25, 1999 TERRESTRIAL REDSHIFTS FROM A DIFFUSE LIGHT SOURCE Ad Lagendijk a FOM Institute for Atomic and Molecular Physics, Kruislaan 407, 1098 SJ Amsterdam, The Netherlands (Received ) Abstract Enhanced
Introduction The gramicidin A (ga) channel forms by head-to-head association of two monomers at their amino termini, one from each bilayer leaflet. Th
 Abstract When conductive, gramicidin monomers are linked by six hydrogen bonds. To understand the details of dissociation and how the channel transits from a state with 6H bonds to ones with 4H bonds or
Abstract When conductive, gramicidin monomers are linked by six hydrogen bonds. To understand the details of dissociation and how the channel transits from a state with 6H bonds to ones with 4H bonds or
Effect of protein shape on multibody interactions between membrane inclusions
 PHYSICAL REVIEW E VOLUME 61, NUMBER 4 APRIL 000 Effect of protein shape on multibody interactions between membrane inclusions K. S. Kim, 1, * John Neu, and George Oster 3, 1 Department of Physics, Graduate
PHYSICAL REVIEW E VOLUME 61, NUMBER 4 APRIL 000 Effect of protein shape on multibody interactions between membrane inclusions K. S. Kim, 1, * John Neu, and George Oster 3, 1 Department of Physics, Graduate
SUPPLEMENTARY INFORMATION
 med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
med!1,2 Wild-type (N2) end!3 elt!2 5 1 15 Time (minutes) 5 1 15 Time (minutes) med!1,2 end!3 5 1 15 Time (minutes) elt!2 5 1 15 Time (minutes) Supplementary Figure 1: Number of med-1,2, end-3, end-1 and
Supplemental Information - Glassy Dynamics in Composite Biopolymer Networks
 Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 Supplemental Information - Glassy Dynamics in Composite Biopolymer Networks Tom Golde, 1 Constantin
Electronic Supplementary Material (ESI) for Soft Matter. This journal is The Royal Society of Chemistry 2018 Supplemental Information - Glassy Dynamics in Composite Biopolymer Networks Tom Golde, 1 Constantin
Introduction to polymer physics Lecture 1
 Introduction to polymer physics Lecture 1 Boulder Summer School July August 3, 2012 A.Grosberg New York University Lecture 1: Ideal chains Course outline: Ideal chains (Grosberg) Real chains (Rubinstein)
Introduction to polymer physics Lecture 1 Boulder Summer School July August 3, 2012 A.Grosberg New York University Lecture 1: Ideal chains Course outline: Ideal chains (Grosberg) Real chains (Rubinstein)
Elasticity of biological gels
 Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Measurement of the Phase Diagram of DNA Unzipping in the Temperature- Force Plane.
 Measurement of the Phase Diagram - Danilowicz et al. 1 Measurement of the Phase Diagram of DNA Unzipping in the Temperature- Force Plane. C.Danilowicz, Y. Kafri, R.S. Conroy, V. W. Coljee, J. Weeks, M.
Measurement of the Phase Diagram - Danilowicz et al. 1 Measurement of the Phase Diagram of DNA Unzipping in the Temperature- Force Plane. C.Danilowicz, Y. Kafri, R.S. Conroy, V. W. Coljee, J. Weeks, M.
Swelling and Collapse of Single Polymer Molecules and Gels.
 Swelling and Collapse of Single Polymer Molecules and Gels. Coil-Globule Transition in Single Polymer Molecules. the coil-globule transition If polymer chains are not ideal, interactions of non-neighboring
Swelling and Collapse of Single Polymer Molecules and Gels. Coil-Globule Transition in Single Polymer Molecules. the coil-globule transition If polymer chains are not ideal, interactions of non-neighboring
STRAIN ASSESSMENT USFOS
 1 STRAIN ASSESSMENT IN USFOS 2 CONTENTS: 1 Introduction...3 2 Revised strain calculation model...3 3 Strain predictions for various characteristic cases...4 3.1 Beam with concentrated load at mid span...
1 STRAIN ASSESSMENT IN USFOS 2 CONTENTS: 1 Introduction...3 2 Revised strain calculation model...3 3 Strain predictions for various characteristic cases...4 3.1 Beam with concentrated load at mid span...
Lecture 34 Protein Unfolding Thermodynamics
 Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Physical Principles in Biology Biology 3550 Fall 2018 Lecture 34 Protein Unfolding Thermodynamics Wednesday, 21 November c David P. Goldenberg University of Utah goldenberg@biology.utah.edu Clicker Question
Lecture 1: Introduction. Keywords: Mathematics as a language, Need of learning mathematics, Applications of mathematics in BIology
 NPTEL Syllabus Biomathematics - Video course COURSE OUTLINE Graphs and functions, Derivative of a function, Techniques of differentiation Differentiation and its application in Biology, Finding maxima,
NPTEL Syllabus Biomathematics - Video course COURSE OUTLINE Graphs and functions, Derivative of a function, Techniques of differentiation Differentiation and its application in Biology, Finding maxima,
SUPPLEMENTARY INFORMATION
 In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2720 1 2 3 Tuning underwater adhesion with cation-π interactions Matthew A. Gebbie, Wei Wei, Alex M. Schrader,
In the format provided by the authors and unedited. SUPPLEMENTARY INFORMATION DOI: 10.1038/NCHEM.2720 1 2 3 Tuning underwater adhesion with cation-π interactions Matthew A. Gebbie, Wei Wei, Alex M. Schrader,
Effects of kinks on DNA elasticity
 PHYSICAL REVIEW E 71, 051905 2005 Effects of kinks on DNA elasticity Yuri O. Popov* and Alexei V. Tkachenko Department of Physics, University of Michigan, 450 Church Street, Ann Arbor, Michigan 48109,
PHYSICAL REVIEW E 71, 051905 2005 Effects of kinks on DNA elasticity Yuri O. Popov* and Alexei V. Tkachenko Department of Physics, University of Michigan, 450 Church Street, Ann Arbor, Michigan 48109,
Supplementary Figures
 Supplementary Figures Supplementary Figure 1: Stacking fault density is direction dependent: Illustration of the stacking fault multiplicity: lattice disorder is clearly direction specific, gradually zooming
Supplementary Figures Supplementary Figure 1: Stacking fault density is direction dependent: Illustration of the stacking fault multiplicity: lattice disorder is clearly direction specific, gradually zooming
Detailed AFM Force Spectroscopy of the Interaction between. CD44-IgG-Fusion Protein and Hyaluronan
 Electronic Supplementary Material to: Detailed AFM Force Spectroscopy of the Interaction between CD44-IgG-Fusion Protein and Hyaluronan Aernout A. Martens a, Marcel Bus a, Peter C. Thüne b,, Tjerk H. Oosterkamp,
Electronic Supplementary Material to: Detailed AFM Force Spectroscopy of the Interaction between CD44-IgG-Fusion Protein and Hyaluronan Aernout A. Martens a, Marcel Bus a, Peter C. Thüne b,, Tjerk H. Oosterkamp,
Simulation in Computer Graphics Elastic Solids. Matthias Teschner
 Simulation in Computer Graphics Elastic Solids Matthias Teschner Outline Introduction Elastic forces Miscellaneous Collision handling Visualization University of Freiburg Computer Science Department 2
Simulation in Computer Graphics Elastic Solids Matthias Teschner Outline Introduction Elastic forces Miscellaneous Collision handling Visualization University of Freiburg Computer Science Department 2
Intensity (a.u.) Intensity (a.u.) Raman Shift (cm -1 ) Oxygen plasma. 6 cm. 9 cm. 1mm. Single-layer graphene sheet. 10mm. 14 cm
 Intensity (a.u.) Intensity (a.u.) a Oxygen plasma b 6 cm 1mm 10mm Single-layer graphene sheet 14 cm 9 cm Flipped Si/SiO 2 Patterned chip Plasma-cleaned glass slides c d After 1 sec normal Oxygen plasma
Intensity (a.u.) Intensity (a.u.) a Oxygen plasma b 6 cm 1mm 10mm Single-layer graphene sheet 14 cm 9 cm Flipped Si/SiO 2 Patterned chip Plasma-cleaned glass slides c d After 1 sec normal Oxygen plasma
Structural investigation of single biomolecules
 Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like
Structural investigation of single biomolecules NMR spectroscopy and X-ray crystallography are currently the most common techniques capable of determining the structures of biological macromolecules like
DNA Structure. Voet & Voet: Chapter 29 Pages Slide 1
 DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
DNA Structure Voet & Voet: Chapter 29 Pages 1107-1122 Slide 1 Review The four DNA bases and their atom names The four common -D-ribose conformations All B-DNA ribose adopt the C2' endo conformation All
The Molecular Dynamics Method
 The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
The Molecular Dynamics Method Thermal motion of a lipid bilayer Water permeation through channels Selective sugar transport Potential Energy (hyper)surface What is Force? Energy U(x) F = d dx U(x) Conformation
Chapter 1 Introduction
 Chapter 1 Introduction This thesis is concerned with the behaviour of polymers in flow. Both polymers in solutions and polymer melts will be discussed. The field of research that studies the flow behaviour
Chapter 1 Introduction This thesis is concerned with the behaviour of polymers in flow. Both polymers in solutions and polymer melts will be discussed. The field of research that studies the flow behaviour
Direct Measurement of Electron Transfer through a Hydrogen Bond
 Supporting Information Direct Measurement of Electron Transfer through a Hydrogen Bond between Single Molecules Tomoaki Nishino,*, Nobuhiko Hayashi, and Phuc T. Bui Nanoscience and Nanotechnology Research
Supporting Information Direct Measurement of Electron Transfer through a Hydrogen Bond between Single Molecules Tomoaki Nishino,*, Nobuhiko Hayashi, and Phuc T. Bui Nanoscience and Nanotechnology Research
Single-molecule studies of DNA mechanics Carlos Bustamante*, Steven B Smith*, Jan Liphardt and Doug Smith
 279 Single-molecule studies of DNA mechanics Carlos Bustamante*, Steven B Smith*, Jan Liphardt and Doug Smith During the past decade, physical techniques such as optical tweezers and atomic force microscopy
279 Single-molecule studies of DNA mechanics Carlos Bustamante*, Steven B Smith*, Jan Liphardt and Doug Smith During the past decade, physical techniques such as optical tweezers and atomic force microscopy
Nanopores: Solid-state nanopores for these experiments were produced by using the
 Materials and Methods Nanopores: Solid-state nanopores for these experiments were produced by using the highly focused electron beam of a transmission electron microscope (TEM) to drill a single pore in
Materials and Methods Nanopores: Solid-state nanopores for these experiments were produced by using the highly focused electron beam of a transmission electron microscope (TEM) to drill a single pore in
arxiv:cond-mat/ v1 [cond-mat.stat-mech] 21 Dec 2004
![arxiv:cond-mat/ v1 [cond-mat.stat-mech] 21 Dec 2004 arxiv:cond-mat/ v1 [cond-mat.stat-mech] 21 Dec 2004](/thumbs/82/86926711.jpg) Theory of Bubble Nucleation and Cooperativity in DNA Melting arxiv:cond-mat/4259v [cond-mat.stat-mech] 2 Dec 24 Saúl Ares,, 2 N. K. Voulgarakis, 3 K. Ø. Rasmussen, and A. R. Bishop Theoretical Division
Theory of Bubble Nucleation and Cooperativity in DNA Melting arxiv:cond-mat/4259v [cond-mat.stat-mech] 2 Dec 24 Saúl Ares,, 2 N. K. Voulgarakis, 3 K. Ø. Rasmussen, and A. R. Bishop Theoretical Division
Fluctuating Elastic Filaments Under Distributed Loads
 Fluctuating Elastic Filaments Under Distributed Loads Tianxiang Su, Prashant K. Purohit Department of Mechanical Engineering and Applied Mechanics, University of Pennsylvania, Philadelphia, PA 94, USA.
Fluctuating Elastic Filaments Under Distributed Loads Tianxiang Su, Prashant K. Purohit Department of Mechanical Engineering and Applied Mechanics, University of Pennsylvania, Philadelphia, PA 94, USA.
Free energy recovery in single molecule experiments
 Supplementary Material Free energy recovery in single molecule experiments Single molecule force measurements (experimental setup shown in Fig. S1) can be used to determine free-energy differences between
Supplementary Material Free energy recovery in single molecule experiments Single molecule force measurements (experimental setup shown in Fig. S1) can be used to determine free-energy differences between
4. Objectives of Research work
 4. Objectives of Research work 4.1 Objectives of Study: The design of bellows is challenging looking to varieties of applications and evaluation of stresses is further difficult to approximate due to its
4. Objectives of Research work 4.1 Objectives of Study: The design of bellows is challenging looking to varieties of applications and evaluation of stresses is further difficult to approximate due to its
Supporting Information
 Supporting Information Beyond the Helix Pitch: Direct Visualization of Native DNA in Aqueous Solution Shinichiro Ido, Kenjiro Kimura, Noriaki Oyabu, Kei Kobayashi, Masaru Tsukada, Kazumi Matsushige and
Supporting Information Beyond the Helix Pitch: Direct Visualization of Native DNA in Aqueous Solution Shinichiro Ido, Kenjiro Kimura, Noriaki Oyabu, Kei Kobayashi, Masaru Tsukada, Kazumi Matsushige and
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION
 THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
THE TANGO ALGORITHM: SECONDARY STRUCTURE PROPENSITIES, STATISTICAL MECHANICS APPROXIMATION AND CALIBRATION Calculation of turn and beta intrinsic propensities. A statistical analysis of a protein structure
On the stall force for growing microtubules
 Eur Biophys J (2000) 29: 2±6 Ó Springer-Verlag 2000 ARTICLE G. Sander van Doorn á Catalin Tanase á Bela M. Mulder Marileen Dogterom On the stall force for growing microtubules Received: 27 September 1999
Eur Biophys J (2000) 29: 2±6 Ó Springer-Verlag 2000 ARTICLE G. Sander van Doorn á Catalin Tanase á Bela M. Mulder Marileen Dogterom On the stall force for growing microtubules Received: 27 September 1999
Extended description of the rigid base model (Supplement to Section 2.1: Model)
 Supplementary material for cgdna: a software package for the prediction of sequence-dependent coarse-grain free energies of B-form DNA by D. Petkevičiūtė, M. Pasi, O. Gonzalez and J.H. Maddocks S Extended
Supplementary material for cgdna: a software package for the prediction of sequence-dependent coarse-grain free energies of B-form DNA by D. Petkevičiūtė, M. Pasi, O. Gonzalez and J.H. Maddocks S Extended
A Computational Study of Nucleosomal DNA Flexibility
 Biophysical Journal Volume 91 December 2006 4121 4132 4121 A Computational Study of Nucleosomal DNA Flexibility Jory Z. Ruscio* and Alexey Onufriev y *Genetics, Bioinformatics & Computational Biology Program,
Biophysical Journal Volume 91 December 2006 4121 4132 4121 A Computational Study of Nucleosomal DNA Flexibility Jory Z. Ruscio* and Alexey Onufriev y *Genetics, Bioinformatics & Computational Biology Program,
The elastic theory of a single DNA molecule
 PRAMANA cfl Indian Academy of Sciences Vol. 61, No. 2 journal of August 2003 physics pp. 353 360 The elastic theory of a single DNA molecule HAIJUN HOU a;b, YANG HANG a and HANG-CAN OU-YANG a;c; * a Institute
PRAMANA cfl Indian Academy of Sciences Vol. 61, No. 2 journal of August 2003 physics pp. 353 360 The elastic theory of a single DNA molecule HAIJUN HOU a;b, YANG HANG a and HANG-CAN OU-YANG a;c; * a Institute
Polymer dynamics. Course M6 Lecture 5 26/1/2004 (JAE) 5.1 Introduction. Diffusion of polymers in melts and dilute solution.
 Course M6 Lecture 5 6//004 Polymer dynamics Diffusion of polymers in melts and dilute solution Dr James Elliott 5. Introduction So far, we have considered the static configurations and morphologies of
Course M6 Lecture 5 6//004 Polymer dynamics Diffusion of polymers in melts and dilute solution Dr James Elliott 5. Introduction So far, we have considered the static configurations and morphologies of
ATP binding controls distinct structural transitions. of Escherichia coli DNA gyrase in complex with DNA
 Supplementary Information ATP binding controls distinct structural transitions of Escherichia coli DNA gyrase in complex with DNA Aakash Basu, Allyn J. Schoeffler, James M. Berger, and Zev Bryant Table
Supplementary Information ATP binding controls distinct structural transitions of Escherichia coli DNA gyrase in complex with DNA Aakash Basu, Allyn J. Schoeffler, James M. Berger, and Zev Bryant Table
Lecture 4 Scanning Probe Microscopy (SPM)
 Lecture 4 Scanning Probe Microscopy (SPM) General components of SPM; Tip --- the probe; Cantilever --- the indicator of the tip; Tip-sample interaction --- the feedback system; Scanner --- piezoelectric
Lecture 4 Scanning Probe Microscopy (SPM) General components of SPM; Tip --- the probe; Cantilever --- the indicator of the tip; Tip-sample interaction --- the feedback system; Scanner --- piezoelectric
SUPPLEMENTARY INFORMATION
 Supplementary Information for Manuscript: Nanoscale wear as a stress-assisted chemical reaction Supplementary Methods For each wear increment, the diamond indenter was slid laterally relative to the silicon
Supplementary Information for Manuscript: Nanoscale wear as a stress-assisted chemical reaction Supplementary Methods For each wear increment, the diamond indenter was slid laterally relative to the silicon
Supporting information
 Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Self-assembled nanopatch with peptide-organic multilayers and mechanical
Electronic Supplementary Material (ESI) for Nanoscale. This journal is The Royal Society of Chemistry 2014 Supporting information Self-assembled nanopatch with peptide-organic multilayers and mechanical
Supporting information for. Direct imaging of kinetic pathways of atomic diffusion in. monolayer molybdenum disulfide
 Supporting information for Direct imaging of kinetic pathways of atomic diffusion in monolayer molybdenum disulfide Jinhua Hong,, Yuhao Pan,, Zhixin Hu, Danhui Lv, Chuanhong Jin, *, Wei Ji, *, Jun Yuan,,*,
Supporting information for Direct imaging of kinetic pathways of atomic diffusion in monolayer molybdenum disulfide Jinhua Hong,, Yuhao Pan,, Zhixin Hu, Danhui Lv, Chuanhong Jin, *, Wei Ji, *, Jun Yuan,,*,
Mechanical response of plectonemic DNA: an analytical solution
 Mechanical response of plectonemic DNA: an analytical solution N. Clauvelin, B. Audoly, S. Neukirch UPMC Univ. Paris 06 & CNRS, Institut Jean le Rond d Alembert, 4, place Jussieu, Paris, France February
Mechanical response of plectonemic DNA: an analytical solution N. Clauvelin, B. Audoly, S. Neukirch UPMC Univ. Paris 06 & CNRS, Institut Jean le Rond d Alembert, 4, place Jussieu, Paris, France February
