Mechanical response of plectonemic DNA: an analytical solution
|
|
|
- Timothy Stevens
- 5 years ago
- Views:
Transcription
1 Mechanical response of plectonemic DNA: an analytical solution N. Clauvelin, B. Audoly, S. Neukirch UPMC Univ. Paris 06 & CNRS, Institut Jean le Rond d Alembert, 4, place Jussieu, Paris, France February 21, 2008 Abstract We consider an elastic rod model for twisted DNA in the plectonemic regime. The molecule is treated as an impenetrable tube with an effective, adjustable radius. The model is solved analytically and we derive formulas for the contact pressure, twisting moment and geometrical parameters of the supercoiled region. We apply our model to magnetic tweezer experiments of a DNA molecule subjected to a tensile force and a torque, and extract mechanical and geometrical quantities from the linear part of the experimental response curve. These reconstructed values are derived in a self-contained manner, and are found to be consistent with those available in the literature. 1 Introduction Mechanical properties of the DNA molecule play an important role in the biological processes involved in the cell, yet we only have an imprecise view of these properties. Advances in nanotechnologies make it possible to exert forces onto isolated DNA filaments: mechanical response of the molecule is now widely studied. Single molecule experiments provide a powerful way to investigate the behavior of DNA subjected to mechanical stress. In such experiments, the molecule is held by optical or magnetic tweezers and forces and torques are applied to it [28, 8]. The interaction between DNA and proteins is actively investigated; for instance, the chemical and mechanical action of an enzyme on a molecule can be inferred from the global deformation of the molecule [24]. In this paper we focus on a specific type of experiments: a double stranded DNA molecule is fixed by one end to a glass surface while the other end is attached to a magnetic bead; using a magnet, a pulling force and a torque are applied on the DNA filament [30]. Large ranges of pulling 1
2 forces, from one tenth to one hundred piconewton, and number of turns can be explored in the experiments, and the molecule displays a variety of behaviors and conformations [1, 27, 7]. We study the response of the molecule to moderate forces, below 10 pn, and moderate to large number of turns, equivalent to a positive supercoiling ratio of the order of 0.1. In experiments, the pulling force is kept constant while the bead is rotated gradually. The vertical extension of the molecule is recorded and plotted as a function of the number of turns. Experimental rotation-extension curves have a characteristic shape and are called hat curves [5, 29]. At zero number of turns these curves exhibit a maximum, the value of which is explained by the worm-like chain (WLC) model [18] and its variants. For small number of turns the vertical extension decreases and the curve takes a rounded shape. Above a threshold value of the number of turns, the extension of the molecule decreases linearly. This linear part of the curve is obtained when the molecule wraps around itself in a helical way, giving rise to a structure comprising plectonemes. The plectonemic structure is made of two interwound helical filaments whose geometry is characterized by the so-called superhelical angle and radius; note that each of these filaments is itself made of a double-stranded DNA molecule. The superhelical angle and the twisting moment in the filaments are key parameters that control the action of topoisomerases [12], RNA polymerase [23], or other enzymes [32] on DNA. The distance of self-approach of DNA in supercoiled regime has been the subject of a number of studies [3, 25, 26, 9]. In previous analytical and numerical work, the double stranded DNA molecule has been modeled as a twist-storing elastic filament. These approaches have been successful at reproducing the response of DNA to moderate torque [4, 19], given by the central region of the experimental curves. The analysis of the linear regions of these curves, based on a detailed model of plectonemes, was lacking until recently: in Ref. [17], a composite model based on an empirical free energy of supercoiled DNA is proposed. Here we present an elastic rod model for helical supercoiling of the DNA molecule, which is relevant to large number of turns. Our model is selfcontained and provides a mechanically accurate description of elastic filaments in contact. The molecule is divided in two domains: one where the configuration is a worm-like-chain, dominated by thermal fluctuations, and the other one, a superhelical region dominated by elasticity, where the molecule contacts itself. Several plectonemic regions may lie at various places of the molecule, but as this does not change the mechanical response of the system, we refer to these regions as if they were in one piece. We deal with self-contact by introducing an effective superhelical radius (distinct from the crystallographic radius of 1 nm, from the size of the Manning condensate and from the Debye length, although in the same range of values), which varies with external loads and salinity of the solution. The effective radius is defined as the radius of a chargeless, impenetrable and elastic 2
3 tube having the same mechanical response as the molecule. This radius is not given as a parameter of the model and is extracted from experimental data. Using an energy approach, we relate geometrical variables (superhelical radius and angle) to applied force and torque. We also characterize the response of the molecule in the plectonemic regime, extend former numerical results [20], and show how geometrical and mechanical parameters can be extracted from experimental data. 2 Model The present model investigates the equilibrium behavior of an elastic rod with bending rigidity K 0 (the bending persistence length is A = K 0 /(kt), where k is the Boltzmann constant and T the absolute temperature) and twisting rigidity K 3 under traction and torsion as shown in Fig. 1. This is a coarse-grained model for DNA where base-pairs details are neglected. For instance, the anisotropic flexibility of the molecule, originating from base pairing and major-minor grove geometry, is smoothed out at a scale of several base pairs: a highly twisted anisotropic rod can be replaced by an equivalent isotropic rod with effective bending rigidity [11]. Geometry We start with a geometric description of the rod configurations relevant to the plectonemic regime. This defines a reduced set of configurations (Ansatz), over which we shall minimize the elastic strain energy associated with deformations. The rod, of length l, is considered inextensible and has circular cross-section; let s denote the arclength along the rod. The strain energy involves, at lowest order, the geometric curvature κ(s) of the centerline of the rod as well as the twist τ(s). We emphasize that the twist τ(s) is different from the geometrical (Frénet-Serret) torsion of the centerline as it takes into account the rotation of material cross-sections around the centerline. It allows one to distinguish between twisted and untwisted configurations of the rod having the same centerline. The rod centerline is parameterized by r(s) and its unit tangent t def = dr/ds can be described with spherical angles, as shown in Fig. 1: α(s) is the zenith angle and ψ(s) the azimuth angle with respect to the direction e x along the common axis of the two superhelices in the plectonemic region. We consider the following configurations, relevant to a large applied number of turns, n. The tails are assumed to be straight but twisted (thermal fluctuations will be accounted for by using the rescaled tail length predicted by WLC theory). The plectonemes are described by two identical and uniform helices where, again, each of these helices is itself a double-stranded DNA molecule. Both the end loop of the plectonemes and the matching 3
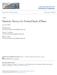
4 Figure 1: Sketch of the magnetic tweezers experiment. A B-DNA molecule of total contour length l is fixed in s = 0 to a glass surface while the other end in s = l is attached to a magnetic bead. A pulling force F ext and a torque M ext are applied at the upper end by using a magnet. The superhelical angle and radius are denoted α and R respectively. The zenith angle α and the azimuth angle ψ of the tangent vector with regard to the superhelical axis e x are also shown. 4
5 region between the tails and the plectonemic part are neglected. Consequently the rod comprises two phases: one made up of straight and twisted tails and the other one of plectonemic structures. The plectonemic phase is not necessarily made of a single component but, for the sake of simplicity, we discuss the case of a single plectonemic structure (our results are still valid if the plectonemes are split into several components). In the tails the rod is straight and aligned with the e z axis: t = e z. The geometric curvature κ def = dt/ds is zero, κ(s) = 0. In each filament of the plectonemes, the position vector r(s) and the tangent vector t(s) describe a superhelix of axis e x : r x (s) = s cos α r y (s) = χr sin ψ(s) r z (s) = χr cos ψ(s) and t x (s) = cos α t y (s) = sin α cos ψ(s) t z (s) = sin α sin ψ(s) The other filament of the plectonemes is obtained by a rotation of 180 around the axis e x. Here χ = ±1 stands for the chirality of the two helices and R and α denote the superhelical radius and angle, respectively. In equation (1), the condition dr/ds = t yields dψ/ds = χ sin α/r. The curvature in the plectonemes is κ(s) def = dt/ds = sin2 α R. Noting l p the contour length spent in the plectonemes, we obtain the following expression for the integral of the squared curvature over the whole length of the rod: l 0 (1) κ 2 ds = sin4 α R 2 l p. (2) The end torque twists the filament. For a rod with circular cross-section, the twist τ(s) at equilibrium is uniform [2], dτ/ds = 0 for all s. As a result, the equilibrium configuration of the rod is fully specified by the centerline, through the variables α, R and l p, and an additional scalar τ describing twist. The twist τ is geometrically related to the number of turns imposed on the magnetic bead, n, which is equal to the link of the DNA molecule, n = Lk. In the present case the link reads [20]: Lk = Tw + Wr = 1 2π l 0 τ ds χ as we neglect the writhe of the tails. Energy formulation sin 2α 4π R l p = 1 2π ( ) sin 2α τ l χ 2R l p, (3) Using the above notations the rod is described by four variables: α the superhelical angle, R the superhelical radius, τ the twist and l p the contour length spent in the plectonemes. We proceed to derive the total energy 5
6 of the system as a function of these four variables. It is the sum of three terms, V = V el +V ext +V int, where the first term is the strain elastic energy, the second is the potential energy associated with the external loads F ext and M ext, and the third accounts for interaction of the filaments in the plectonemes. The strain elastic energy for the rod of total contour length l is : V el = K 0 2 l 0 κ 2 ds + K 3 2 l 0 τ 2 ds. (4) We do not take into account the reduction of the effective torsional rigidity in the tails due to fluctuations [19]. The potential energy is given by: V ext = F ext (z(l) z(0)) 2π M ext n, (5) where z(l) z(0) = l l p for straight tails and n = Lk. If the DNA-DNA interaction was clearly established, we would include the corresponding interaction energy V int in the total energy V [10]. This is not the case and we model the filaments in electrostatic interaction as effective chargeless hard-core tubes. The effective radius a of these tubes accounts for a variety of physical mechanisms, including for example the presence of counter-ions or thermal fluctuations in the plectonemes, which we do not attempt to model. As in Refs. [33, 25], we do not try to predict the actual radius a but simply follow its variation under changing experimental conditions (applied load, salinity, etc.). Doing so, we replace the actual (unknown) interaction potential V int (R,α) by a hard-core interaction with adjustable radius a, and optimize a to best fit a given experiment. In section 3 we show how a can be extracted from experimental measurements. The parameter a must certainly be larger than the crystallographic DNA radius 1 nm. It is different from the radius of the Manning condensate [14, 15, 16] since approximately a quarter of the charge remains outside of the Manning condensate. The equilibrium is the solution of a constrained minimization problem for the elastic energy, subjected to the impenetrability condition R a. (6) We anticipate on the fact that there is contact, R = a, for typical experimental conditions. Consequently we replace the actual interaction energy with a constraint term: V int = λ(r a), (7) where λ is a Lagrange multiplier. Note that this term is not a regular energy but comes from the constraint: the multiplier λ has to be set at the end of the procedure and chosen in such a way that the constraint R = a is satisfied. Combining Eqs. (2 7), we write the total potential energy of the system 6
7 as: V (α,r,τ,l p ) = K 0 2 sin 4 α R 2 l p + K 3 2 τ2 l F ext (l l p ) ) sin 2α 2R l p λ(r a). (8) M ext ( τ l χ In Ref. [13] a similar energy function has been introduced but the rest of analysis differs from ours. Indeed, their approach focuses on statistical mechanics and the analysis of the state of lowest energy is overlooked. Moreover, the parameter a is fixed a priori to the crystallographic radius of DNA, a = 1 nm, which is a strong restriction and an underestimation of the actual distance of self-approach of DNA in saline solution. In contrast, we undertake a detailed analysis of the equilibrium solutions, with thermal fluctuations considered in the tails; this allows us to derive simple formulas for the force and the moment as a function of the superhelical variables, applicable to magnetic tweezers experiments. 3 Results Mechanical equilibrium is given by the Euler-Lagrange condition for the stationarity of the potential V (α,r,τ,l p ) in Eq. (8) with respect to its variables, ( V τ, V α, V, V ) = 0. l p R The first condition V/ τ allows one to recover the constitutive relation for twist deformations, M ext = K 3 τ, given that the twisting moment is uniform in the filament and equal to the applied torque M ext. Variation of the total energy with respect to α gives the expression of the applied torque M ext in terms of the superhelical variables α and R: M ext = 2χK 0 R cos α sin 3 α cos 2α, (9) which is what was found for purely plectonemic solution (no tails) [31]. The condition V/ l p = 0, combined with Eq. (9), allows one to relate the pulling force F ext to the superhelical geometry: F ext = K 0 R 2 sin4 α ( cos 2α ). (10) This formula justifies and extends the numerical fit F ext K 0 α 4 /R 2 found in Ref. [20] for small values of α. The Euler-Lagrange condition with respect to R yields an equation involving the Lagrange multiplier λ. The quantity λ/l p can be interpreted as 7
8 the contact force per unit length, p, of one filament onto the other. Eqs. (8 9), together with the condition V/ R = 0, yields: p = λ l p = K 0 R 3 sin 4 α cos 2α. (11) Note that this pressure (more accurately, force per unit length) is positive for α π/4; if our assumption of contact R = a was incorrect, this would be indicated by a negative pressure value here. In magnetic tweezers experiments, the pulling force F ext is imposed although the applied torque M ext is unknown. The two unknowns R and α are then related by Eq. (10); in the next Section, a second equation relating those unknowns and the extension z is given, which makes it possible to solve for R and α. The twisting moment can then be found from Eq. (9). Vertical extension of the filament In magnetic tweezers experiments, the measurable quantities are the vertical extension z and the number of turns n imposed on the bead. Using Eq. (3) for n = Lk, the equation z = l l p and the constitutive relation τ = M ext /K 3 where M ext is found from Eq. (9), we obtain the vertical extension of the filament as a linear function of the number of turns n: z = ( 1 + 2K 0 K 3 sin 2 α cos 2α ) l + χn 4π R sin 2α. (12) Thermal fluctuations dominantly affect the tails and make the end-to-end distance z of the molecule smaller than the contour length l l p of the tail parts, by a factor ρ wlc [0,1]: z = ρ wlc (l l p ). This factor depends on both the pulling force F ext and the bending persistence length A = K 0 /(kt) and can either be read off an experimental hat curve from the value z(n = 0) = ρ wlc l, or computed from theoretical formulas [18, 6]. The dependence of ρ wlc on the pulling force makes the tails effectively extensible (this is the classical entropic stiffness of a chain). To account for these thermal effects, we replace Eq. (12) with: ( z = ρ wlc 1 + 2K 0 sin 2 ) α 4π R l + χρ wlc K 3 cos 2α sin2α n. (13) One of the main features of the experimental hat curves is the linear decrease of the vertical extension with the number of turns. We define the slope q in the linear part of the hat curve as: q def = dz dn = ρ 4π R wlc sin 2α. (14) Given experimental values of F ext and q, Eqs. (10) and (14) can be solved for R and α. Since q (and F ext ) are constant along the linear part of a hat 8
9 curve, the values of R and α thus determined will be constant as well. As a result, the twisting moment in the molecule, given by Eq. (9), is constant, for a given experiment, along the linear region of the hat curve, a property that has been previously reported in the literature [5, 17] and which is a clear outcome of the present model. An interpretation of the fact that R and α are constant in the linear region of the hat curve is that each additional turn of the bead is used to convert a small piece of tail into plectonemes. Twisting moment The twisting moment in the molecule, which is uniform and equal to M ext at equilibrium, cannot be measured in magnetic tweezers experiments. However, it has been shown that enzyme activity such as RNA polymerase depends on the value of the twisting moment in DNA [23]. The value of M ext can be determined from Eq. (9) once R and α are known, as explained above. Here, we give a formula for M ext directly as function of the experimental slope q and the external force F ext. Indeed, using Eq. (14) to eliminate R in Eqs. (9) and (10), one obtains M ext (q,α) and F ext (q,α) as functions of q and α. It is then possible to eliminate α, which yields: M ext = m+ ( m 2 + 2K 0 F ext ) 1/2, q F ext where m = 3π ρ wlc K 0 4π ρ wlc 2q (15) In the limit of small α, one can expand the functions M ext (q,α) and F ext (q,α) prior to elimination of α, and this leads to a simplified formula: M ext 2q 3π ρ wlc F ext, (16) where, as explained above, ρ wlc = z(n = 0)/l. As shown in Fig. 4, this approximation is accurate when used with typical experimental values. Eq. (16) provides a simple and direct mean of evaluating the twisting moment in magnetic tweezers experiments, based on the slope of the linear part of the hat curve only. Note that it should not be inferred from Eq. (16) that M ext depends linearly on F ext, as the slope q is itself a function of F ext. Superhelical angle limit It is known that the topology of contact between two impenetrable helical tubes winding along a common axis changes when α becomes larger than π/4 [21]. The possibility of such a change of topology is not considered in our model (being specific to hard-core repulsion between tubes, it is not relevant to DNA molecules undergoing long-range electrostatic repulsion anyway). Nevertheless, the equilibrium solutions found here are all such that α < π/4. This upper bound has a mechanical origin, and not a geometrical one: the expressions for F ext in Eq. (10) and for M ext in Eq. (9) both diverge 9
10 ÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌÌ ÌÌÌ Ì ÌÌÌ ÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁÁ ÁÁÁÁ ÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙ ÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙÙ Ì ÙÙ Á ÌÌÌ Ù ÁÁÁ Ù Ù Ù Ì Á Ù ÌÌ Ù Ù Ù Ù Ù Ù Ù Ù Ù Á Ù Ù Ì ÌÌ Ì ÌÌ Ì ÌÌ Figure 2: Experimental hat curves showing the vertical extension of a lambda phage DNA 48kbp molecule as a function of the number of turns imposed on the magnetic bead (salt concentration 10 mm, temperature 298 K). Experimentally measured persistence length of the molecule is A = 51.35nm. Each curve corresponds to a fixed pulling force F ext : 0.25, 0.33, 0.44, 0.57, 0.74, 1.10, 1.31, 2.20, 2.95 pn. Triangles represent the fit for the slope q of the linear region. Data kindly provided by V. Croquette (CNRS, France). at α = π/4 and plectonemic solutions with a superhelical angle larger than π/4 are unstable. Application to experiments The model is used to extract mechanical and geometrical parameters from experimental data. To allow comparison with previous work, we use the same data as in [20]. These data are shown in Fig. 2; they were obtained on a 48kbp lambda phage DNA molecule in a 10mM phosphate buffer. For each curve in Fig. 2, corresponding to a given value of the external force F ext, we extract the slope q by fitting the linear region. The superhelical variables R and α are found by solving Eqs. (10) and (14), and are plotted in Fig. 3 as a function of F ext. The reconstructed values of R are in the nanometric range; they decrease with the pulling force, from approximately 6 to 2 times the DNA crystallographic radius in this particular experiment. At large values of the force, R is close to (and actually smaller than) the Debye length, 3.07 nm in 10 mm salt, and the Manning condensation radius, 3.18 nm in 10 mm salt [22]. We note that the values of R found here in the presence of a pulling force are smaller than (and in the same range as) in 10
11 Figure 3: Reconstructed values of the plectonemic radius R as a function of the pulling force, from the data in Fig. 2 by solving Eqs. (10) and (14). The angle α is shown in the inset. Ref. [26] where no force is applied, which is consistent. The reconstructed values of the twisting moment M ext and of the contact pressure p are given in Fig. 4, based on the same experimental data. The values of M ext are determined both by Eq. (9) using the previously computed values of R and α, and by the approximate formula (16) directly. A good agreement is obtained, which validates the proposed approximation. The values of M ext are also compared to those predicted by a composite analytical model, see Eq. (17) in Ref. [17] (this model uses effective parameters determined from Monte-Carlo simulations [34]). 4 Conclusion We have shown that, under the approximation that thermal fluctuations are neglected in the plectonemes, one can calculate analytically the response of twisted DNA: supercoils are described by a mechanically exact and selfcontained model. Self-contact in the plectonemic region is treated with a hard-core potential; an expression for the contact pressure between the two dsdna is derived. The hard-core radius is an effective parameter determined, for a given value of the applied force, from the slope of the linear region of the experimental curve. A formula for the twisting moment is proposed, as a function of the slope of the linear region of the experimental hat curve only. We apply this analysis to experimental data from which we extract the mechanical quantities: superhelical radius and angle, contact 11
12 Á Á Á Á Á Á Á Á Á Figure 4: Reconstructed values for the twisting moment in the molecule based on the data shown in Fig. 2, using the exact formula in Eq. (9) (solid squares), and the small angles approximation in Eq. (16) (open circles). Comparison with the prediction of the composite model in Ref. [17] (curve). Contact pressure is shown in inset. pressure and twisting moment. We compared these values with predictions from previous analyses, when available, and found that they are consistent. In future work, we shall extend the present model to deal with long-range interaction potentials, predict the superhelical radius, and utilize magnetic tweezers experiments to probe DNA-DNA electrostatic interaction. The present paper is a first step towards a mechanically accurate description of bare dsdna subjected to tensile and torsional loads, a problem relevant to the architecture of DNA in the cell nucleus where proteins come into play. Acknowledgment: We thank V. Croquette for providing us with the experimental data used here. References [1] J. F. Allemand, D. Bensimon, R. Lavery, and V. Croquette. Stretched and overwound DNA forms a pauling-like structure with exposed bases. Proc. Natl. Acad. Sci. USA, 95: , [2] S. S. Antman. Nonlinear Problems of Elasticity. Springer Verlag, New York,
13 [3] J. Bednar, P. Furrer, A. Stasiak, J. Dubochet, E. H. Egelman, and A. D. Bates. The twist, writhe and overall shape of supercoiled DNA change during counterion-induced transition from a loosely to a tightly interwound superhelix. possible implications for DNA structure in vivo. J. Mol. Biol., 235(3): , [4] C. Bouchiat and M. Mézard. Elasticity model of a supercoiled DNA molecule. Physical Review Letters, 80(7): , [5] C. Bouchiat and M. Mézard. Elastic rod model of a supercoiled DNA molecule. European Physical Journal E, 2: , [6] C. Bouchiat, M. D. Wang, J.-F. Allemand, T. Strick, S. M. Block, and V. Croquette. Estimating the persistence length of a worm-like chain molecule from force-extension measurements. Biophysical Journal, 76: , [7] Zev Bryant, Michael D. Stone, Jeff Gore, Steven B. Smith, Nicholas R. Cozzarelli, and Carlos Bustamante. Structural transitions and elasticity from torque measurements on DNA. Nature, 424: , [8] C. Bustamente, J. F. Marko, E. D. Siggia, and S. Smith. Entropic elasticity of lambda-phage DNA. Science, 265: , [9] G. Charvin, A. Vologodskii, D. Bensimon, and V. Croquette. Braiding DNA: Experiments, simulations, and models. Biophysical Journal, 88: , [10] N. Clauvelin, B. Audoly, and S. Neukirch. Plectonemic DNA experiment to test longe-range interaction potential. submitted. [11] S. Kehrbaum and J. H. Maddocks. Effective properties of elastic rods with high intrinsic twist. Proceedings of the 16th IMACS World Congress, [12] Daniel A. Koster, Vincent Croquette, Cees Dekker, Stewart Shuman, and Nynke H. Dekker. Friction and torque govern the relaxation of DNA supercoils by eukaryotic topoisomerase IB. Nature, 434: , [13] S. Kutter and E. M. Terentjev. Helix coil mixing in twist-storing polymers. European Physical Journal B, 21: , [14] G. S. Manning. Limiting laws and counterion condensation in polyelectrolyte solutions - I. colligative properties. Journal of Chemical Physics, 51(3): ,
14 [15] G. S. Manning. Limiting laws and counterion condensation in polyelectrolyte solutions - II. self-diffusion of the small ions. Journal of Chemical Physics, 51(3): , [16] G. S. Manning. Limiting laws and counterion condensation in polyelectrolyte solutions - III. an analysis based on the mayer ionic solution theory. Journal of Chemical Physics, 51(8): , [17] J. F. Marko. Torque and dynamics of linking number relaxation in stretched supercoiled DNA. Physical Review E, 76:021926, [18] J. F. Marko and E. D. Siggia. Streching DNA. Macromolecules, 28: , [19] J. David Moroz and Philip Nelson. Torsional directed walks, entropic elasticity, and DNA twist stiffness. Proceedings of the National Academy of Sciences, USA, 94:14418, [20] S. Neukirch. Extracting DNA twist rigidity from experimental supercoiling data. Physical Review Letters, 93(19):198107, [21] Sébastien Neukirch and Gert van der Heijden. Geometry and mechanics of uniform n-plies: from engineering ropes to biological filaments. Journal of Elasticity, 69:41 72, [22] B. O Shaughnessy and Q. Yang. Manning-Osawa counterion condensation. Physical Review Letters, 94(048302):1 4, [23] Andrey Revyakin, Richard H. Ebright, and Terence R. Strick. Promoter unwinding and promoter clearance by RNA polymerase: Detection by single-molecule DNA nanomanipulation. Proc. Natl. Acad. Sci. USA, 101(14): , [24] Andrey Revyakin, Chenyu Liu, Richard H. Ebright, and Terence R. Strick. Abortive Initiation and Productive Initiation by RNA Polymerase Involve DNA Scrunching. Science, 314(5802): , [25] V. V. Rybenkov, A. V. Vologodskii, and N. R. Cozzarelli. The effect of ionic conditions on DNA helical repeat, effective diameter and free energy of supercoiling. Nucleic Acids Research, 25(7), [26] Valentin V. Rybenkov, Nicholas R. Cozzarelli, and Alexander V. Vologodskii. Probability of DNA knotting and the effective diameter of the DNA double-helix. Proc. Natl. Acad. Sci. USA, 90: , [27] A. Sarkar, J. F. Leger, D. Chatenay, and J. F. Marko. Structural transitions in DNA driven by external force and torque. Physical Review E, 63:051903,
15 [28] S. B. Smith, L. Finzi, and C. Bustamante. Direct mechanical measurements of the elasticity of single DNA molecules by using magnetic beads. Science, 258: , [29] T. Strick, J.-F. Allemand, D. Bensimon, and V. Croquette. Behavior of supercoiled DNA. Biophysical Journal, 74( ), [30] T. R. Strick, J.-F. Allemand, D. Bensimon, A. Bensimon, and V. Croquette. The elasticity of a single supercoiled DNA molecule. Science, 271: , [31] J. M. T. Thompson, G. H. M. van der Heijden, and S. Neukirch. Supercoiling of DNA plasmids: mechanics of the generalized ply. Proc. R. Soc. Lond. A, 458: , [32] Thijn van der Heijden, John van Noort, Hendrikje van Leest, Roland Kanaar, Claire Wyman, Nynke Dekker, and Cees Dekker. Torquelimited RecA polymerization on dsdna. Nucleic Acids Research, 33(7): , [33] Alexander Vologodskii and Nicholas Cozzarelli. Modeling of long-range electrostatic interactions in DNA. Biopolymers, 35(3): , [34] Alexander V. Vologodskii and John F. Marko. Extension of torsionally stressed DNA by external force. Biophysical Journal, 73: ,
16 For Table of Contents use only N. Clauvelin, B. Audoly, S. Neukirch Mechanical response of plectonemic DNA: an analytical solution 16
DNA supercoiling: plectonemes or curls?
 DNA supercoiling: plectonemes or curls? Sébastien Neukirch CNRS & UPMC Univ. Paris 6 (France) John F. Marko Northwestern University (IL. USA) Why study DNA mechanical properties? mechanical properties
DNA supercoiling: plectonemes or curls? Sébastien Neukirch CNRS & UPMC Univ. Paris 6 (France) John F. Marko Northwestern University (IL. USA) Why study DNA mechanical properties? mechanical properties
La molécule d'adn vue comme une poutre élastique : application aux expériences de pinces magnétiques
 La molécule d'adn vue comme une poutre élastique : application aux expériences de pinces magnétiques joint work with: Sébastien Neukirch CNRS & UPMC Univ Paris 6 Institut Jean le Rond d Alembert Michael
La molécule d'adn vue comme une poutre élastique : application aux expériences de pinces magnétiques joint work with: Sébastien Neukirch CNRS & UPMC Univ Paris 6 Institut Jean le Rond d Alembert Michael
New Measurements of DNA Twist Elasticity
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 5-1998 New Measurements of DNA Twist Elasticity Philip C. Nelson University of Pennsylvania, nelson@physics.upenn.edu
Magnetic tweezers and its application to DNA mechanics
 Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Biotechnological Center Research group DNA motors (Seidel group) Handout for Practical Course Magnetic tweezers and its application to DNA mechanics When: 9.00 am Where: Biotec, 3 rd Level, Room 317 Tutors:
Curvature Distribution of Worm-like Chains in Two and Three Dimensions
 Curvature Distribution of Worm-like Chains in Two and Three Dimensions Shay M. Rappaport, Shlomi Medalion and Yitzhak Rabin Department of Physics, Bar-Ilan University, Ramat-Gan 59, Israel arxiv:8.383v
Curvature Distribution of Worm-like Chains in Two and Three Dimensions Shay M. Rappaport, Shlomi Medalion and Yitzhak Rabin Department of Physics, Bar-Ilan University, Ramat-Gan 59, Israel arxiv:8.383v
Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA
 8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
8 th International LS-DYNA Users Conference Simulation echnology (3) Investigation of dsdna Stretching Meso-Mechanics Using LS-DYNA C. A. Yuan and K. N. Chiang Department of Power Mechanical Engineering,
Torsional directed walks, entropic elasticity, and DNA twist stiffness
 Proc. Natl. Acad. Sci. USA Vol. 94, pp. 14418 14422, December 1997 Biophysics Torsional directed walks, entropic elasticity, and DNA twist stiffness J. DAVID MOROZ AND PHILIP NELSON* Department of Physics
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 14418 14422, December 1997 Biophysics Torsional directed walks, entropic elasticity, and DNA twist stiffness J. DAVID MOROZ AND PHILIP NELSON* Department of Physics
Chirality of coiled-coils: elasticity matters
 Chirality of coiled-coils: elasticity matters Sébastien Neukirch Institut Jean le Rond d Alembert, CNRS & Université Pierre et Marie Curie, Paris, France Alain Goriely Department of Mathematics and Program
Chirality of coiled-coils: elasticity matters Sébastien Neukirch Institut Jean le Rond d Alembert, CNRS & Université Pierre et Marie Curie, Paris, France Alain Goriely Department of Mathematics and Program
Chiral selection in wrapping, crossover, and braiding of DNA mediated by asymmetric bend-writhe elasticity
 http://www.aimspress.com/ AIMS Biophysics, 2(4): 666-694. DOI: 10.3934/biophy.2015.4.666 Received date 28 August 2015, Accepted date 29 October 2015, Published date 06 November 2015 Research article Chiral
http://www.aimspress.com/ AIMS Biophysics, 2(4): 666-694. DOI: 10.3934/biophy.2015.4.666 Received date 28 August 2015, Accepted date 29 October 2015, Published date 06 November 2015 Research article Chiral
arxiv:cond-mat/ v1 [cond-mat.soft] 29 May 2002
![arxiv:cond-mat/ v1 [cond-mat.soft] 29 May 2002 arxiv:cond-mat/ v1 [cond-mat.soft] 29 May 2002](/thumbs/90/102416155.jpg) Stretching Instability of Helical Springs David A. Kessler and Yitzhak Rabin Dept. of Physics, Bar-Ilan University, Ramat-Gan, Israel (Dated: October 31, 18) arxiv:cond-mat/05612v1 [cond-mat.soft] 29 May
Stretching Instability of Helical Springs David A. Kessler and Yitzhak Rabin Dept. of Physics, Bar-Ilan University, Ramat-Gan, Israel (Dated: October 31, 18) arxiv:cond-mat/05612v1 [cond-mat.soft] 29 May
Faculty of math and natural science. Universty of Split. Kratky-Porod model. Nataša Vučemilovid-Alagid. February, 2013.
 Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
Faculty of math and natural science Universty of Split Kratky-Porod model Nataša Vučemilovid-Alagid February, 2013. Table of contents 1.Introduction:... 3 1. DNA:... 4 2. Models of polymer elasticity:...
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments
 Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Magnetic Torque Tweezers: measuring torsional stiffness in DNA and RecA DNA filaments Lipfert, J., Kerssemakers, J. W., Jager, T. & Dekker, N. H. Magnetic torque tweezers: measuring torsional stiffness
Effect of Spontaneous Twist on DNA Minicircles
 Biophysical Journal Volume 99 November 2010 2987 2994 2987 Effect of Spontaneous Twist on DNA Minicircles Shlomi Medalion, * David A. Kessler, and Yitzhak Rabin { Department of Physics and Institute of
Biophysical Journal Volume 99 November 2010 2987 2994 2987 Effect of Spontaneous Twist on DNA Minicircles Shlomi Medalion, * David A. Kessler, and Yitzhak Rabin { Department of Physics and Institute of
, follows from these assumptions.
 Untangling the mechanics and topology in the frictional response of long overhand elastic knots Supplemental Information M.K. Jawed, P. Dieleman, B. Audoly, P. M. Reis S1 Experimental details In this section,
Untangling the mechanics and topology in the frictional response of long overhand elastic knots Supplemental Information M.K. Jawed, P. Dieleman, B. Audoly, P. M. Reis S1 Experimental details In this section,
Elastic property of single double-stranded DNA molecules: Theoretical study and comparison with experiments
 PHYSICA REVIEW E VOUME 62, NUMBER 1 JUY 2 Elastic property of single double-stranded DNA molecules: Theoretical study and comparison with experiments Haijun Zhou, 1,2, * Yang Zhang, 1 and Zhong-can Ou-Yang
PHYSICA REVIEW E VOUME 62, NUMBER 1 JUY 2 Elastic property of single double-stranded DNA molecules: Theoretical study and comparison with experiments Haijun Zhou, 1,2, * Yang Zhang, 1 and Zhong-can Ou-Yang
The elastic theory of a single DNA molecule
 PRAMANA cfl Indian Academy of Sciences Vol. 61, No. 2 journal of August 2003 physics pp. 353 360 The elastic theory of a single DNA molecule HAIJUN HOU a;b, YANG HANG a and HANG-CAN OU-YANG a;c; * a Institute
PRAMANA cfl Indian Academy of Sciences Vol. 61, No. 2 journal of August 2003 physics pp. 353 360 The elastic theory of a single DNA molecule HAIJUN HOU a;b, YANG HANG a and HANG-CAN OU-YANG a;c; * a Institute
Supplementary Information
 Supplementary Information Gi-Moon Nam and Gaurav Arya Department of NanoEngineering, University of California at San Diego, 9500 Gilman Drive, La Jolla, CA 9093-0448 1 Coarse-grained (CG) model of nucleosome
Supplementary Information Gi-Moon Nam and Gaurav Arya Department of NanoEngineering, University of California at San Diego, 9500 Gilman Drive, La Jolla, CA 9093-0448 1 Coarse-grained (CG) model of nucleosome
On the Topology of Chromatin Fibers
 On the Topology of Chromatin Fibers Maria Barbi Multiscale modeling of living matter (M3V) group" Laboratoire de Physique Théorique de la Matière Condensée (LPTMC), " CNRS-Université Paris VI, France"
On the Topology of Chromatin Fibers Maria Barbi Multiscale modeling of living matter (M3V) group" Laboratoire de Physique Théorique de la Matière Condensée (LPTMC), " CNRS-Université Paris VI, France"
3 Biopolymers Uncorrelated chains - Freely jointed chain model
 3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
3.4 Entropy When we talk about biopolymers, it is important to realize that the free energy of a biopolymer in thermal equilibrium is not constant. Unlike solids, biopolymers are characterized through
Multimedia : Fibronectin and Titin unfolding simulation movies.
 I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
I LECTURE 21: SINGLE CHAIN ELASTICITY OF BIOMACROMOLECULES: THE GIANT PROTEIN TITIN AND DNA Outline : REVIEW LECTURE #2 : EXTENSIBLE FJC AND WLC... 2 STRUCTURE OF MUSCLE AND TITIN... 3 SINGLE MOLECULE
Friction of F-actin knots
 Friction of F-actin knots Helmut O. Kirchner a, Sebastien Neukirch b a INM Leibniz Institute for New Materials Campus D, D-6613 Saarbrücken, Germany b CNS & UPMC Univ Paris 06 UM 7190, Institut Jean Le
Friction of F-actin knots Helmut O. Kirchner a, Sebastien Neukirch b a INM Leibniz Institute for New Materials Campus D, D-6613 Saarbrücken, Germany b CNS & UPMC Univ Paris 06 UM 7190, Institut Jean Le
Elasticity of the human red blood cell skeleton
 Biorheology 40 (2003) 247 251 247 IOS Press Elasticity of the human red blood cell skeleton G. Lenormand, S. Hénon, A. Richert, J. Siméon and F. Gallet Laboratoire de Biorhéologie et d Hydrodynamique Physico-Chimique,
Biorheology 40 (2003) 247 251 247 IOS Press Elasticity of the human red blood cell skeleton G. Lenormand, S. Hénon, A. Richert, J. Siméon and F. Gallet Laboratoire de Biorhéologie et d Hydrodynamique Physico-Chimique,
Untangling the Mechanics of Entangled Biopolymers
 Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
Untangling the Mechanics of Entangled Biopolymers Rae M. Robertson-Anderson Physics Department University of San Diego students/postdocs: Cole Chapman, PhD Tobias Falzone, PhD Stephanie Gorczyca, USD 16
8.592J HST.452J: Statistical Physics in Biology
 Assignment # 4 8.592J HST.452J: Statistical Physics in Biology Coulomb Interactions 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions scales as Q 2 E c.
Assignment # 4 8.592J HST.452J: Statistical Physics in Biology Coulomb Interactions 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions scales as Q 2 E c.
V = 2ze 2 n. . a. i=1
 IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
IITS: Statistical Physics in Biology Assignment # 3 KU Leuven 5/29/2013 Coulomb Interactions & Polymers 1. Flory Theory: The Coulomb energy of a ball of charge Q and dimension R in d spacial dimensions
Single molecule force spectroscopy reveals a highly compliant helical
 Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Supplementary Information Single molecule force spectroscopy reveals a highly compliant helical folding for the 30 nm chromatin fiber Maarten Kruithof, Fan-Tso Chien, Andrew Routh, Colin Logie, Daniela
Announcements. Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1.
 Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Announcements Homework 3 (Klaus Schulten s Lecture): Due Wednesday at noon. Next homework assigned. Due Wednesday March 1. No lecture next Monday, Feb. 27 th! (Homework is a bit longer.) Marco will have
Elasticity Theory of a Twisted Stack of Plates
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 7-1997 Elasticity Theory of a Twisted Stack of Plates Corey S. O'Hern Randall Kamien University of Pennsylvania,
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 7-1997 Elasticity Theory of a Twisted Stack of Plates Corey S. O'Hern Randall Kamien University of Pennsylvania,
On end rotation for open rods undergoing. large deformations. G.H.M. van der Heijden, M.A. Peletier and. R. Planqué
 ARMA manuscript No. (will be inserted by the editor) On end rotation for open rods undergoing large deformations G.H.M. van der Heijden, M.A. Peletier and R. Planqué Abstract We give a careful discussion
ARMA manuscript No. (will be inserted by the editor) On end rotation for open rods undergoing large deformations G.H.M. van der Heijden, M.A. Peletier and R. Planqué Abstract We give a careful discussion
Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102:
 Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102: 6783-6788. Background: The lac operon is a cluster of genes in the E. coli genome
Villa et al. (2005) Structural dynamics of the lac repressor-dna complex revealed by a multiscale simulation. PNAS 102: 6783-6788. Background: The lac operon is a cluster of genes in the E. coli genome
Single-molecule studies of DNA mechanics Carlos Bustamante*, Steven B Smith*, Jan Liphardt and Doug Smith
 279 Single-molecule studies of DNA mechanics Carlos Bustamante*, Steven B Smith*, Jan Liphardt and Doug Smith During the past decade, physical techniques such as optical tweezers and atomic force microscopy
279 Single-molecule studies of DNA mechanics Carlos Bustamante*, Steven B Smith*, Jan Liphardt and Doug Smith During the past decade, physical techniques such as optical tweezers and atomic force microscopy
Introduction to Polymer Physics
 Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
Introduction to Polymer Physics Enrico Carlon, KU Leuven, Belgium February-May, 2016 Enrico Carlon, KU Leuven, Belgium Introduction to Polymer Physics February-May, 2016 1 / 28 Polymers in Chemistry and
DNA Conformational Changes and Phase Transitions Induced by Tension and Twist
 University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 1-1-2014 DNA Conformational Changes and Phase Transitions Induced by Tension and Twist David E. Argudo University of Pennsylvania,
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 1-1-2014 DNA Conformational Changes and Phase Transitions Induced by Tension and Twist David E. Argudo University of Pennsylvania,
3.22 Mechanical Properties of Materials Spring 2008
 MIT OpenCourseWare http://ocw.mit.edu 3.22 Mechanical Properties of Materials Spring 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Problem Set
MIT OpenCourseWare http://ocw.mit.edu 3.22 Mechanical Properties of Materials Spring 2008 For information about citing these materials or our Terms of Use, visit: http://ocw.mit.edu/terms. Problem Set
Elastic theories of single DNA molecules
 Physica A 36 (22) 359 367 www.elsevier.com/locate/physa Elastic theories of single DNA molecules HaijunZhou a;b; 1, Yang Zhang a; 2, Zhong-can Ou-Yang a;c; a Institute of Theoretical Physics, The Chinese
Physica A 36 (22) 359 367 www.elsevier.com/locate/physa Elastic theories of single DNA molecules HaijunZhou a;b; 1, Yang Zhang a; 2, Zhong-can Ou-Yang a;c; a Institute of Theoretical Physics, The Chinese
Dynamical simulations of DNA supercoiling and compression
 554 Biochemical Society Transactions (2013) Volume 41, part 2 Dynamical simulations of DNA supercoiling and compression David Swigon* 1, Sookkyung Lim and Yongsam Kim *Department of Mathematics, University
554 Biochemical Society Transactions (2013) Volume 41, part 2 Dynamical simulations of DNA supercoiling and compression David Swigon* 1, Sookkyung Lim and Yongsam Kim *Department of Mathematics, University
Measurement of the Phase Diagram of DNA Unzipping in the Temperature- Force Plane.
 Measurement of the Phase Diagram - Danilowicz et al. 1 Measurement of the Phase Diagram of DNA Unzipping in the Temperature- Force Plane. C.Danilowicz, Y. Kafri, R.S. Conroy, V. W. Coljee, J. Weeks, M.
Measurement of the Phase Diagram - Danilowicz et al. 1 Measurement of the Phase Diagram of DNA Unzipping in the Temperature- Force Plane. C.Danilowicz, Y. Kafri, R.S. Conroy, V. W. Coljee, J. Weeks, M.
Mechanical Design in Optical Engineering
 Torsion Torsion: Torsion refers to the twisting of a structural member that is loaded by couples (torque) that produce rotation about the member s longitudinal axis. In other words, the member is loaded
Torsion Torsion: Torsion refers to the twisting of a structural member that is loaded by couples (torque) that produce rotation about the member s longitudinal axis. In other words, the member is loaded
Modeling DNA loops using the theory of elasticity
 Modeling DNA loops using the theory of elasticity Alexander Balaeff* Beckman Institute and Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, Urbana, Illinois
Modeling DNA loops using the theory of elasticity Alexander Balaeff* Beckman Institute and Center for Biophysics and Computational Biology, University of Illinois at Urbana-Champaign, Urbana, Illinois
Magnetic tweezers for DNA manipulation
 University of Ljubljana Faculty of Mathematics and Physics Seminar Magnetic tweezers for DNA manipulation Ivna Kavre Adviser: prof. dr. Rudolf Podgornik Ljubljana, March 2011 Abstract There are several
University of Ljubljana Faculty of Mathematics and Physics Seminar Magnetic tweezers for DNA manipulation Ivna Kavre Adviser: prof. dr. Rudolf Podgornik Ljubljana, March 2011 Abstract There are several
Super-Coiling of DNA Plasmids: Mechanics of the Generalised Ply
 Super-Coiling of DNA Plasmids: Mechanics of the Generalised Ply J. M. T. Thompson, G. H. M. van der Heijden & S. Neukirch Centre for Nonlinear Dynamics, Civil Engineering Building, University College London,
Super-Coiling of DNA Plasmids: Mechanics of the Generalised Ply J. M. T. Thompson, G. H. M. van der Heijden & S. Neukirch Centre for Nonlinear Dynamics, Civil Engineering Building, University College London,
Brownian Dynamics Simulation of DNA Condensation
 1858 Biophysical Journal Volume 77 October 1999 1858 1870 Brownian Dynamics Simulation of DNA Condensation Pierre-Edouard Sottas, Eric Larquet, Andrzej Stasiak, and Jacques Dubochet Laboratoire d Analyse
1858 Biophysical Journal Volume 77 October 1999 1858 1870 Brownian Dynamics Simulation of DNA Condensation Pierre-Edouard Sottas, Eric Larquet, Andrzej Stasiak, and Jacques Dubochet Laboratoire d Analyse
ON END ROTATION FOR OPEN RODS UNDERGOING LARGE DEFORMATIONS
 QUARTERLY OF APPLIED MATHEMATICS VOLUME LXV, NUMBER 2 JUNE 27, PAGES 385 42 S 33-569X(7)149-X Article electronically published on April 1, 27 ON END ROTATION FOR OPEN RODS UNDERGOING LARGE DEFORMATIONS
QUARTERLY OF APPLIED MATHEMATICS VOLUME LXV, NUMBER 2 JUNE 27, PAGES 385 42 S 33-569X(7)149-X Article electronically published on April 1, 27 ON END ROTATION FOR OPEN RODS UNDERGOING LARGE DEFORMATIONS
SECOND PUBLIC EXAMINATION. Honour School of Physics Part C: 4 Year Course. Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS
 2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
2757 SECOND PUBLIC EXAMINATION Honour School of Physics Part C: 4 Year Course Honour School of Physics and Philosophy Part C C7: BIOLOGICAL PHYSICS TRINITY TERM 2013 Monday, 17 June, 2.30 pm 5.45 pm 15
INTRODUCTION TO SINGLE-DNA MICROMECHANICS (AUGUST 9, 2004) John F. Marko. Department of Physics, University of Illinois at Chicago, Chicago, IL, USA
 INTRODUCTION TO SINGLE-DNA MICROMECHANICS AUGUST 9, 2004) John F. Marko Department of Physics, University of Illinois at Chicago, Chicago, IL, USA 1 Photo: width 7.5cm height 11cm Contents 1. Introduction
INTRODUCTION TO SINGLE-DNA MICROMECHANICS AUGUST 9, 2004) John F. Marko Department of Physics, University of Illinois at Chicago, Chicago, IL, USA 1 Photo: width 7.5cm height 11cm Contents 1. Introduction
Theory of Self-contact in DNA Molecules Modeled as Elastic Rods
 1 Theory of Self-contact in DNA Molecules Modeled as Elastic Rods Bernard D. Coleman and David Swigon To be published in 001 by the Accademia Nazionale dei Lincei in the volume "Nuovi progressi nella fisica
1 Theory of Self-contact in DNA Molecules Modeled as Elastic Rods Bernard D. Coleman and David Swigon To be published in 001 by the Accademia Nazionale dei Lincei in the volume "Nuovi progressi nella fisica
Electromagnetic Torque Tweezers: A Versatile Approach for Measurement of Single-Molecule Twist and Torque
 Supplementary information for: Electromagnetic Torque Tweezers: A Versatile Approach for Measurement of Single-Molecule Twist and Torque Xander J.A. Janssen 1, Jan Lipfert 1, Tessa Jager 1, Renier Daudey
Supplementary information for: Electromagnetic Torque Tweezers: A Versatile Approach for Measurement of Single-Molecule Twist and Torque Xander J.A. Janssen 1, Jan Lipfert 1, Tessa Jager 1, Renier Daudey
Macroscopic modeling and simulations of supercoiled DNA with bound proteins
 JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Macroscopic modeling and simulations of supercoiled DNA with bound proteins Jing Huang a) and Tamar Schlick b) Department of Chemistry
JOURNAL OF CHEMICAL PHYSICS VOLUME 117, NUMBER 18 8 NOVEMBER 2002 Macroscopic modeling and simulations of supercoiled DNA with bound proteins Jing Huang a) and Tamar Schlick b) Department of Chemistry
Stretching of macromolecules and proteins
 Home Search Collections Journals About Contact us My IOPscience Stretching of macromolecules and proteins This content has been downloaded from IOPscience. Please scroll down to see the full text. 2003
Home Search Collections Journals About Contact us My IOPscience Stretching of macromolecules and proteins This content has been downloaded from IOPscience. Please scroll down to see the full text. 2003
Pulling hairpinned polynucleotide chains: Does base-pair stacking interaction matter?
 JOURNAL OF CHEMICAL PHYSICS VOLUME 114, NUMBER 19 15 MAY 2001 Pulling hairpinned polynucleotide chains: Does base-pair stacking interaction matter? Haijun Zhou a) Max-Planck-Institute of Colloids and Interfaces,
JOURNAL OF CHEMICAL PHYSICS VOLUME 114, NUMBER 19 15 MAY 2001 Pulling hairpinned polynucleotide chains: Does base-pair stacking interaction matter? Haijun Zhou a) Max-Planck-Institute of Colloids and Interfaces,
No. of Pages 12, Model 5G. Contents lists available at SciVerse ScienceDirect. Acta Biomaterialia
 ACTBIO 2074 10 February 2012 No. of Pages 12, Model 5G Acta Biomaterialia xxx (2012) xxx xxx 1 Contents lists available at SciVerse ScienceDirect Acta Biomaterialia journal homepage: www.elsevier.com/locate/actabiomat
ACTBIO 2074 10 February 2012 No. of Pages 12, Model 5G Acta Biomaterialia xxx (2012) xxx xxx 1 Contents lists available at SciVerse ScienceDirect Acta Biomaterialia journal homepage: www.elsevier.com/locate/actabiomat
arxiv: v1 [cond-mat.soft] 26 Sep 2016
![arxiv: v1 [cond-mat.soft] 26 Sep 2016 arxiv: v1 [cond-mat.soft] 26 Sep 2016](/thumbs/88/117259531.jpg) APS/123-QED Formation of Non-uniform Double Helices for Elastic Rods under Torsion Hongyuan Li, 1 Shumin Zhao, 1, Minggang Xia, 1, 2, 3 Siyu He, 1 Qifan Yang, 4 Yuming Yan, 5 and Hanqiao Zhao 4 1 Department
APS/123-QED Formation of Non-uniform Double Helices for Elastic Rods under Torsion Hongyuan Li, 1 Shumin Zhao, 1, Minggang Xia, 1, 2, 3 Siyu He, 1 Qifan Yang, 4 Yuming Yan, 5 and Hanqiao Zhao 4 1 Department
Soft Matter and Biological Physics
 Dr. Ulrich F. Keyser - ufk20 (at) cam.ac.uk Soft Matter and Biological Physics Question Sheet Michaelmas 2011 Version: November 2, 2011 Question 0: Sedimentation Initially consider identical small particles
Dr. Ulrich F. Keyser - ufk20 (at) cam.ac.uk Soft Matter and Biological Physics Question Sheet Michaelmas 2011 Version: November 2, 2011 Question 0: Sedimentation Initially consider identical small particles
Supporting Online Material for
 www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
www.sciencemag.org/cgi/content/full/314/5801/1001/dc1 Supporting Online Material for Direct Measurement of the Full, Sequence-Dependent Folding Landscape of a Nucleic Acid Michael T. Woodside, Peter C.
Writhe formulas and antipodal points in plectonemic DNA configurations
 PHYSIL RVIW 78, 041912 2008 Writhe formulas and antipodal points in plectonemic N configurations Sébastien Neukirch Institut Jean le Rond d lembert, NRS and UPM Univ. Paris 6, 4 Place Jussieu (case 162),
PHYSIL RVIW 78, 041912 2008 Writhe formulas and antipodal points in plectonemic N configurations Sébastien Neukirch Institut Jean le Rond d lembert, NRS and UPM Univ. Paris 6, 4 Place Jussieu (case 162),
Multiple folding and packing in DNA modeling
 omputers and Mathematics with Applications 55 (2008) 1044 1053 www.elsevier.com/locate/camwa Multiple folding and packing in DNA modeling Renzo L. Ricca a,b,, Francesca Maggioni a a Department of Mathematics
omputers and Mathematics with Applications 55 (2008) 1044 1053 www.elsevier.com/locate/camwa Multiple folding and packing in DNA modeling Renzo L. Ricca a,b,, Francesca Maggioni a a Department of Mathematics
Physics of RecA-mediated homologous recognition
 Submitted to: Biophysical Journal, Jan. 05, 004 Physics of RecA-mediated homologous recognition Kevin Klapstein, 1 Tom Chou, 1 and Robijn Bruinsma 1 Dept. of Biomathematics, UCLA, Los Angeles, CA 90095-1766
Submitted to: Biophysical Journal, Jan. 05, 004 Physics of RecA-mediated homologous recognition Kevin Klapstein, 1 Tom Chou, 1 and Robijn Bruinsma 1 Dept. of Biomathematics, UCLA, Los Angeles, CA 90095-1766
Introduction to polymer physics Lecture 1
 Introduction to polymer physics Lecture 1 Boulder Summer School July August 3, 2012 A.Grosberg New York University Lecture 1: Ideal chains Course outline: Ideal chains (Grosberg) Real chains (Rubinstein)
Introduction to polymer physics Lecture 1 Boulder Summer School July August 3, 2012 A.Grosberg New York University Lecture 1: Ideal chains Course outline: Ideal chains (Grosberg) Real chains (Rubinstein)
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES
 Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
Sec. 2.1 Filaments in the cell 21 PART I - RODS AND ROPES Sec. 2.1 Filaments in the cell 22 CHAPTER 2 - POLYMERS The structural elements of the cell can be broadly classified as filaments or sheets, where
An experimental study of the temperature-dependent DNA elasticity using optical tweezers.
 An experimental study of the temperature-dependent DNA elasticity using optical tweezers. Author: Álvaro Martínez Monge Facultat de Física, Universitat de Barcelona, Diagonal 645, 08028 Barcelona, Spain.
An experimental study of the temperature-dependent DNA elasticity using optical tweezers. Author: Álvaro Martínez Monge Facultat de Física, Universitat de Barcelona, Diagonal 645, 08028 Barcelona, Spain.
Link, Twist, Energy, and the Stability of DNA Minicircles.
 Link, Twist, Energy, and the Stability of DNA Minicircles Kathleen A. Hoffman 1, Robert S. Manning 2, & John H. Maddocks 3 khoffman@math.umbc.edu, rmanning@haverford.edu, john.maddocks@epfl.ch November
Link, Twist, Energy, and the Stability of DNA Minicircles Kathleen A. Hoffman 1, Robert S. Manning 2, & John H. Maddocks 3 khoffman@math.umbc.edu, rmanning@haverford.edu, john.maddocks@epfl.ch November
EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December Suggested resolution
 page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
page 1 of 7 EXAM I COURSE TFY4310 MOLECULAR BIOPHYSICS December 2013 Suggested resolution Exercise 1. [total: 25 p] a) [t: 5 p] Describe the bonding [1.5 p] and the molecular orbitals [1.5 p] of the ethylene
Phys 450 Spring 2011 Solution set 6. A bimolecular reaction in which A and B combine to form the product P may be written as:
 Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Problem Phys 45 Spring Solution set 6 A bimolecular reaction in which A and combine to form the product P may be written as: k d A + A P k d k a where k d is a diffusion-limited, bimolecular rate constant
Magnetic Tweezers for Single-Molecule Experiments
 Magnetic Tweezers for Single-Molecule Experiments I. D. Vilfan, J. Lipfert, D. A. Koster, S. G. Lemay, and N. H. Dekker 13 Abstract Over the last decade, single-molecule techniques have proven their wide
Magnetic Tweezers for Single-Molecule Experiments I. D. Vilfan, J. Lipfert, D. A. Koster, S. G. Lemay, and N. H. Dekker 13 Abstract Over the last decade, single-molecule techniques have proven their wide
SUPPLEMENTARY INFORMATION
 SUPPLEMENTARY INFORMATION doi: 10.1038/nphys00 Peter Gross, Niels Laurens, Lene B. Oddershede, Ulrich Bockelmann 3, Erwin J. G. Peterman*, and Gijs J. L. Wuite* LaserLaB and Department of Physics and Astronomy,
SUPPLEMENTARY INFORMATION doi: 10.1038/nphys00 Peter Gross, Niels Laurens, Lene B. Oddershede, Ulrich Bockelmann 3, Erwin J. G. Peterman*, and Gijs J. L. Wuite* LaserLaB and Department of Physics and Astronomy,
Origin of the Electrophoretic Force on DNA in Nanopores. Biological and Soft Systems - Cavendish Laboratory
 Origin of the Electrophoretic Force on DNA in Nanopores Ulrich F. Keyser Biological and Soft Systems - Cavendish Laboratory Acknowledgements Delft Cees Dekker, Nynke H. Dekker, Serge G. Lemay R. Smeets,
Origin of the Electrophoretic Force on DNA in Nanopores Ulrich F. Keyser Biological and Soft Systems - Cavendish Laboratory Acknowledgements Delft Cees Dekker, Nynke H. Dekker, Serge G. Lemay R. Smeets,
Proteins polymer molecules, folded in complex structures. Konstantin Popov Department of Biochemistry and Biophysics
 Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Proteins polymer molecules, folded in complex structures Konstantin Popov Department of Biochemistry and Biophysics Outline General aspects of polymer theory Size and persistent length of ideal linear
Effects of kinks on DNA elasticity
 PHYSICAL REVIEW E 71, 051905 2005 Effects of kinks on DNA elasticity Yuri O. Popov* and Alexei V. Tkachenko Department of Physics, University of Michigan, 450 Church Street, Ann Arbor, Michigan 48109,
PHYSICAL REVIEW E 71, 051905 2005 Effects of kinks on DNA elasticity Yuri O. Popov* and Alexei V. Tkachenko Department of Physics, University of Michigan, 450 Church Street, Ann Arbor, Michigan 48109,
The Effect of Ionic Conditions on the Conformations of Supercoiled DNA. I. Sedimentation Analysis
 J. Mol. Biol. (1997) 267, 299±311 The Effect of Ionic Conditions on the Conformations of Supercoiled DNA. I. Sedimentation Analysis Valentin V. Rybenkov 1, Alexander V. Vologodskii 2 and Nicholas R. Cozzarelli
J. Mol. Biol. (1997) 267, 299±311 The Effect of Ionic Conditions on the Conformations of Supercoiled DNA. I. Sedimentation Analysis Valentin V. Rybenkov 1, Alexander V. Vologodskii 2 and Nicholas R. Cozzarelli
Mechanical Engineering Ph.D. Preliminary Qualifying Examination Solid Mechanics February 25, 2002
 student personal identification (ID) number on each sheet. Do not write your name on any sheet. #1. A homogeneous, isotropic, linear elastic bar has rectangular cross sectional area A, modulus of elasticity
student personal identification (ID) number on each sheet. Do not write your name on any sheet. #1. A homogeneous, isotropic, linear elastic bar has rectangular cross sectional area A, modulus of elasticity
DNA/RNA structure and packing
 DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
DNA/RNA structure and packing Reminder: Nucleic acids one oxygen atom distinguishes RNA from DNA, increases reactivity (so DNA is more stable) base attaches at 1, phosphate at 5 purines pyrimidines Replace
Mechanics of elastic rods
 Mechanics of elastic rods Khalid Jawed www.khalidjawed.com Postdoctoral Fellow Integrated Soft Materials Laboratory Carnegie Mellon University PhD (Mechanics) 2016 Massachusetts Institute of Technology
Mechanics of elastic rods Khalid Jawed www.khalidjawed.com Postdoctoral Fellow Integrated Soft Materials Laboratory Carnegie Mellon University PhD (Mechanics) 2016 Massachusetts Institute of Technology
A Continuum Model for Condensation of DNA in Solution
 A Continuum Model for Condensation of DNA in Solution YONGZHAO WANG School of Mechanical Engineering Tianjin Key Laboratory of Nonlinear Dynamics and Chaos Control Tianjin University Weijin Road, Nankai
A Continuum Model for Condensation of DNA in Solution YONGZHAO WANG School of Mechanical Engineering Tianjin Key Laboratory of Nonlinear Dynamics and Chaos Control Tianjin University Weijin Road, Nankai
Virtual Work & Energy Methods. External Energy-Work Transformation
 External Energy-Work Transformation Virtual Work Many structural problems are statically determinate (support reactions & internal forces can be found by simple statics) Other methods are required when
External Energy-Work Transformation Virtual Work Many structural problems are statically determinate (support reactions & internal forces can be found by simple statics) Other methods are required when
Modeling DNA structure, elasticity, and deformations at the base-pair level
 Modeling DNA structure, elasticity, and deformations at the base-pair level Boris Mergell,* Mohammad R. Ejtehadi, and Ralf Everaers Max-Planck-Institut für Polymerforschung, Postfach 3148, D-55021 Mainz,
Modeling DNA structure, elasticity, and deformations at the base-pair level Boris Mergell,* Mohammad R. Ejtehadi, and Ralf Everaers Max-Planck-Institut für Polymerforschung, Postfach 3148, D-55021 Mainz,
Models for Single Molecule Experiments
 Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
Models for Single Molecule Experiments Binding Unbinding Folding - Unfolding -> Kramers Bell theory 1 Polymer elasticity: - Hooke s law (?) - FJC (Freely joined chain) - WLC (Worm-like chain) 3 Molecular
DNA topology in chromosomes: a quantitative survey and its physiological implications
 J. Math. Biol. DOI 10.1007/s00285-012-0621-y Mathematical Biology DNA topology in chromosomes: a quantitative survey and its physiological implications Maria Barbi Julien Mozziconacci Hua Wong Jean-Marc
J. Math. Biol. DOI 10.1007/s00285-012-0621-y Mathematical Biology DNA topology in chromosomes: a quantitative survey and its physiological implications Maria Barbi Julien Mozziconacci Hua Wong Jean-Marc
Writhe distribution of stiff biopolymers
 Writhe distribution of stiff biopolymers Abhijit Ghosh Raman Research Institute, Bangalore, India 560080 Received 5 November 2007; published 30 April 2008 Motivated by the interest in the role of stiff
Writhe distribution of stiff biopolymers Abhijit Ghosh Raman Research Institute, Bangalore, India 560080 Received 5 November 2007; published 30 April 2008 Motivated by the interest in the role of stiff
Nonlinear Analytical Model for Wire Strands
 Advances in Engineering Research, volume 4th International Conference on Renewable Energy and Environmental Technology (ICREET 06) Nonlinear Analytical Model for Wire Strands Chun-Lei Yu, a, Wen-Guang
Advances in Engineering Research, volume 4th International Conference on Renewable Energy and Environmental Technology (ICREET 06) Nonlinear Analytical Model for Wire Strands Chun-Lei Yu, a, Wen-Guang
Contents. xiii. Preface v
 Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Contents Preface Chapter 1 Biological Macromolecules 1.1 General PrincipIes 1.1.1 Macrornolecules 1.2 1.1.2 Configuration and Conformation Molecular lnteractions in Macromolecular Structures 1.2.1 Weak
Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation
 Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation Ion:ion interactions What is the free energy of ion:ion interactions ΔG i-i? Consider an ion in a solution
Bchem 675 Lecture 9 Electrostatics-Lecture 2 Debye-Hückel: Continued Counter ion condensation Ion:ion interactions What is the free energy of ion:ion interactions ΔG i-i? Consider an ion in a solution
Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of µm
 University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 12-2007 Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of 0.6 7 µm Yeonee Seol
University of Pennsylvania ScholarlyCommons Department of Physics Papers Department of Physics 12-2007 Elasticity of Short DNA Molecules: Theory and Experiment for Contour Lengths of 0.6 7 µm Yeonee Seol
For an imposed stress history consisting of a rapidly applied step-function jump in
 Problem 2 (20 points) MASSACHUSETTS INSTITUTE OF TECHNOLOGY DEPARTMENT OF MECHANICAL ENGINEERING CAMBRIDGE, MASSACHUSETTS 0239 2.002 MECHANICS AND MATERIALS II SOLUTION for QUIZ NO. October 5, 2003 For
Problem 2 (20 points) MASSACHUSETTS INSTITUTE OF TECHNOLOGY DEPARTMENT OF MECHANICAL ENGINEERING CAMBRIDGE, MASSACHUSETTS 0239 2.002 MECHANICS AND MATERIALS II SOLUTION for QUIZ NO. October 5, 2003 For
Week 3: Differential Geometry of Curves
 Week 3: Differential Geometry of Curves Introduction We now know how to differentiate and integrate along curves. This week we explore some of the geometrical properties of curves that can be addressed
Week 3: Differential Geometry of Curves Introduction We now know how to differentiate and integrate along curves. This week we explore some of the geometrical properties of curves that can be addressed
The Effects of Mechanical Properties of DNA on the Thermodynamic Stability of the DNA-Dendronized Polymer Nanocluster
 Journal of the Iranian Chemical Society Vol 3, No, pp 185-190, June 006 DOI: 101007/BF0345948 The Effects of Mechanical Properties of DNA on the Thermodynamic Stability of the DNA-Dendronized Polymer Nanocluster
Journal of the Iranian Chemical Society Vol 3, No, pp 185-190, June 006 DOI: 101007/BF0345948 The Effects of Mechanical Properties of DNA on the Thermodynamic Stability of the DNA-Dendronized Polymer Nanocluster
Elasticity of biological gels
 Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Seminar II Elasticity of biological gels Author: Gašper Gregorič Mentor: assoc. prof. Primož Ziherl Ljubljana, February 2014 Abstract In the seminar we discuss the elastic behavior of biological gels,
Experimental Soft Matter (M. Durand, G. Foffi)
 Master 2 PCS/PTSC 2016-2017 10/01/2017 Experimental Soft Matter (M. Durand, G. Foffi) Nota Bene Exam duration : 3H ecture notes are not allowed. Electronic devices (including cell phones) are prohibited,
Master 2 PCS/PTSC 2016-2017 10/01/2017 Experimental Soft Matter (M. Durand, G. Foffi) Nota Bene Exam duration : 3H ecture notes are not allowed. Electronic devices (including cell phones) are prohibited,
High flexibility of DNA on short length scales probed by atomic force microscopy
 High flexibility of DNA on short length scales probed by atomic force microscopy PAUL A. WIGGINS 1, THIJN VAN DER HEIJDEN, FERNANDO MORENO-HERRERO, ANDREW SPAKOWITZ 3, ROB PHILLIPS, JONATHAN WIDOM 5, CEES
High flexibility of DNA on short length scales probed by atomic force microscopy PAUL A. WIGGINS 1, THIJN VAN DER HEIJDEN, FERNANDO MORENO-HERRERO, ANDREW SPAKOWITZ 3, ROB PHILLIPS, JONATHAN WIDOM 5, CEES
Bending and Twisting Elasticity of DNA
 Macromolecules 1994,27, 981-988 981 Bending and Twisting Elasticity of DNA J. F. Marko and E. D. Siggia Laboratory of Atomic and Solid State Physics, Clark Hall, Cornell University, Zthaca, New York 14853-2501
Macromolecules 1994,27, 981-988 981 Bending and Twisting Elasticity of DNA J. F. Marko and E. D. Siggia Laboratory of Atomic and Solid State Physics, Clark Hall, Cornell University, Zthaca, New York 14853-2501
Force-Induced Melting of the DNA Double Helix. 2. Effect of Solution Conditions*
 894 Biophysical Journal Volume 80 February 2001 894 900 Force-Induced Melting of the DNA Double Helix. 2. Effect of Solution Conditions* Ioulia Rouzina and Victor A. Bloomfield Department of Biochemistry,
894 Biophysical Journal Volume 80 February 2001 894 900 Force-Induced Melting of the DNA Double Helix. 2. Effect of Solution Conditions* Ioulia Rouzina and Victor A. Bloomfield Department of Biochemistry,
Consider an elastic spring as shown in the Fig.2.4. When the spring is slowly
 .3 Strain Energy Consider an elastic spring as shown in the Fig..4. When the spring is slowly pulled, it deflects by a small amount u 1. When the load is removed from the spring, it goes back to the original
.3 Strain Energy Consider an elastic spring as shown in the Fig..4. When the spring is slowly pulled, it deflects by a small amount u 1. When the load is removed from the spring, it goes back to the original
Relating Single-Molecule Measurements to Thermodynamics
 Biophysical Journal Volume 84 February 23 733 738 733 Relating Single-Molecule Measurements to Thermodynamics David Keller,* David Swigon, y and Carlos Bustamante z { *Department of Chemistry, University
Biophysical Journal Volume 84 February 23 733 738 733 Relating Single-Molecule Measurements to Thermodynamics David Keller,* David Swigon, y and Carlos Bustamante z { *Department of Chemistry, University
GEM4 Summer School OpenCourseWare
 GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Polymer Chains by Ju Li. Given August 16, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use the
GEM4 Summer School OpenCourseWare http://gem4.educommons.net/ http://www.gem4.org/ Lecture: Polymer Chains by Ju Li. Given August 16, 2006 during the GEM4 session at MIT in Cambridge, MA. Please use the
Magnetic torque tweezers: measuring torsional stiffness in DNA and
 1 Nature America, Inc. All rights reserved. Magnetic torque tweezers: measuring torsional stiffness in DNA and RecA-DNA filaments Jan Lipfert, Jacob W J Kerssemakers, Tessa Jager & Nynke H Dekker We introduce
1 Nature America, Inc. All rights reserved. Magnetic torque tweezers: measuring torsional stiffness in DNA and RecA-DNA filaments Jan Lipfert, Jacob W J Kerssemakers, Tessa Jager & Nynke H Dekker We introduce
Lecture 4: viscoelasticity and cell mechanics
 Teaser movie: flexible robots! R. Shepherd, Whitesides group, Harvard 1 Lecture 4: viscoelasticity and cell mechanics S-RSI Physics Lectures: Soft Condensed Matter Physics Jacinta C. Conrad University
Teaser movie: flexible robots! R. Shepherd, Whitesides group, Harvard 1 Lecture 4: viscoelasticity and cell mechanics S-RSI Physics Lectures: Soft Condensed Matter Physics Jacinta C. Conrad University
Supplementary Information for: The Temperature Dependence of the Helical Twist of DNA
 Supplementary Information for: The Temperature Dependence of the Helical Twist of DNA Franziska Kriegel 1, Christian Matek 2, Tomáš Dršata 3, Klara Kulenkampff 1, Sophie Tschirpke 1, Martin Zacharias 4,
Supplementary Information for: The Temperature Dependence of the Helical Twist of DNA Franziska Kriegel 1, Christian Matek 2, Tomáš Dršata 3, Klara Kulenkampff 1, Sophie Tschirpke 1, Martin Zacharias 4,
Liquid Crystal Formation in Supercoiled DNA Solutions
 Biophysical Journal Volume 83 August 2002 1119 1129 1119 Liquid Crystal Formation in Supercoiled DNA Solutions Svetlana S. Zakharova, Wim Jesse, Claude Backendorf, and Johan R. C. van der Maarel Leiden
Biophysical Journal Volume 83 August 2002 1119 1129 1119 Liquid Crystal Formation in Supercoiled DNA Solutions Svetlana S. Zakharova, Wim Jesse, Claude Backendorf, and Johan R. C. van der Maarel Leiden
MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins
 MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins Ideal freely jointed chain N identical unstretchable links (Kuhn segments) of length a with freely rotating joints ~r 2 a ~r ~R ~r N In first
MAE 545: Lecture 4 (9/29) Statistical mechanics of proteins Ideal freely jointed chain N identical unstretchable links (Kuhn segments) of length a with freely rotating joints ~r 2 a ~r ~R ~r N In first
Adaptive elastic properties of chromatinber
 Physica A 314 (2002) 592 599 www.elsevier.com/locate/physa Adaptive elastic properties of chromatinber Eli Ben-Ham, Annick Lesne, Jean-Marc Victor Laboratoire de Physique Theorique des Liquides, Universite
Physica A 314 (2002) 592 599 www.elsevier.com/locate/physa Adaptive elastic properties of chromatinber Eli Ben-Ham, Annick Lesne, Jean-Marc Victor Laboratoire de Physique Theorique des Liquides, Universite
