Applications of Latin Hypercube Sampling Scheme and Partial Rank Correlation Coefficient Analysis to Mathematical Models on Wound Healing
|
|
|
- Leon Hampton
- 5 years ago
- Views:
Transcription
1 Western Kentucky University TopSCHOLAR Honors College Capstone Experience/Thesis Projects Honors College at WKU Applications of Latin Hypercube Sampling Scheme and Partial Rank Correlation Coefficient Analysis to Mathematical Models on Wound Healing Hannah M. Pennington Western Kentucky University, Follow this and additional works at: Part of the Applied Mathematics Commons, and the Biology Commons Recommended Citation Pennington, Hannah M., "Applications of Latin Hypercube Sampling Scheme and Partial Rank Correlation Coefficient Analysis to Mathematical Models on Wound Healing" (215). Honors College Capstone Experience/Thesis Projects. Paper This Thesis is brought to you for free and open access by TopSCHOLAR. It has been accepted for inclusion in Honors College Capstone Experience/ Thesis Projects by an authorized administrator of TopSCHOLAR. For more information, please contact
2 APPLICATIONS OF LATIN HYPERCUBE SAMPLING SCHEME AND PARTIAL RANK CORRELATION COEFFICIENT ANALYSIS TO MATHEMATICAL MODELS IN WOUND HEALING. A Capstone Experience/Thesis Project Presented in Partial Fulfillment of the Requirements for the Degree Bachelor of Science with Honors College Graduate Distinction at Western Kentucky University By Hannah M. Pennington * * * * * Western Kentucky University 215 CE/T Committee: Approved by Professor Richard Schugart, Advisor Professor Marisa Eisenberg Professor Michael Smith Advisor Department of Mathematics
3 Copyright by Hannah M. Pennington 215
4 ABSTRACT Latin hypercube sampling and Partial Rank Correlation Coefficient procedure (LHS/ PRCC) can be used in combination to perform a sensitivity analysis that assesses a model over a global parameter space. Through this analysis, the uncertainty of the parameters and therefore the variability of the model output in response to this uncertainty can be observed. Latin hypercube sampling divides the parameter space into equiprobable regions and samples without replacement, producing a global, unbiased selection of parameter values. For monotonic, non-linear relationships, the correlation between the outputs and parameters can be understood by performing a Partial Rank Correlation Coefficient procedure. This sensitivity analysis is applied to ordinary differential equation models describing the interactions of various biological components in the healing of a diabetic foot ulcer. The results of the LHS/PRCC sensitivity analysis are used to assess the biological significance of the parameters in relation to each compartment of the model to further understand its biological implications. Keywords: Wound Healing, Latin hypercube sampling, Partial Rank Correlation Coefficient procedure, Uncertainty, Sensitivity Analysis ii
5 I dedicate this to my family and friends. aut viam inveniam aut faciam iii
6 ACKNOWLEDGMENTS This research would not have been impossible without the support of many. My mentor, Dr. Richard Schugart, shaped my future by showing me the true potential of mathematics and research. Through his guidance and encouragment, I grew as a speaker and academic. His invaluable direction and discussion pushed me to think beyond the textbook and taught me to be a true scientist. I would also like to thank the rest of my committee, Dr. Marisa Eisenberg and Dr. Michael Smith, for their help and guidance in completing my thesis. I was fortunate to have the constant camaraderie of my colleague and research partner, Nitin Krishna. Together, we forged an understanding of research and supported each other through times when all logical reasoning seemed out of reach. I thank my family and friends for their continual support and words of encouragement when I reached the peak of my frustration. Financial support was provided by a Western Kentucky University (WKU) Faculty- Undergraduate Student Engagement (FUSE) Award #14-SP141, a WKU Honors Development grant, and a Gatton Alumni Scholarship Award. Additional financial support was also provided by The Gatton Academy of Mathematics and Science in Kentucky. Permission was granted by Wiley to use the data given in Muller et al. iv
7 v VITA June 21, Born - Cincinnati, Ohio Conner High School, Hebron, KY Graduated Honors with Distinction Gatton Academy, Bowling Green, KY UREP Wood Hudson Cancer Research Laboratory, Newport, KY Western Kentucky University Summa Cum Laude PRESENTATIONS Pennington, Hannah M. (214 June). A mathematical model for the interaction of the proteins MMP-1, TIMP-1, and ECM in a wound. Poster presented at European Conference of Mathematical and Theoretical Biology, Gothenburg, Sweden. Pennington, Hannah M. (214 October). A mathematical model for the interaction of the proteins MMP-1, TIMP-1, and ECM in a wound. Paper presented at Western Kentucky University Mathematics Symposium, Bowling Green, KY. Pennington, Hannah M. (214 November). A mathematical model for the interaction of the proteins MMP-1, TIMP-1, and ECM in a wound. Paper presented at National Institute for Mathematical and Biological Synthesis Undergraduate Research Conference at the Interface of Biology and Mathematics, Knoxville, TN. Pennington, Hannah M. (215 Jan). Latin hypercube sampling and Partial Rank Correlation Coefficient procedure as applied to a mathematical model for wound healing. Paper presented at Joint Mathematics Meetings, San Antonio, TX. Pennington, Hannah M. (215 March). Applications Of Latin Hypercube Sampling Scheme And Partial Rank Correlation Coefficient Analysis To Mathematical Models In Wound Healing. Paper presented at WKU Student Research Conference, Bowling Green, KY.
8 vi PUBLICATIONS Krishna, NK., Pennington, HP. Coppola, C., Eisenberg, MC., Schugart, RC. (215). Connecting Local and Global Sensitivities for a Mathematical Model in Wound Healing. Manuscript submitted for publication to Bulletin of Mathematical Biology. Major Fields: Biology, Chemistry Minor Field: Mathematics FIELDS OF STUDY
9 TABLE OF CONTENTS Abstract Dedication Acknowledgments Vita Presentations Publications Fields of Study ii iii iv v v v vi 1 Introduction 1 2 MMP 1/TIMP 1 Model 4 2.1Introduction Materials and methods Clinical data Mathematical Framework Model Equations Nondimensionalization Parameter Estimation Steady-State Analysis Global sensitivity analysis Local sensitivity analyses Classical sensitivity analysis SVD-QR subset selection Local-to-global analysis Results Steady-state Analysis Sensitivity analysis Global sensitivity analysis Local sensitivity analysis Local-to-global analysis vii
10 TABLE OF CONTENTS viii 2.4Discussion Parameter Estimate Interpretation Steady-state analysis Comparison of sensitivity analyses Limitations Future work Extended TGF-β Model Model Equations Nondimensionalization Parameter Estimation Global Sensitivity Analysis Discussion Bacterial Infection Model Model Equations Global Sensitivity Analysis Discussion Conclusions and Further Work 72
11 LIST OF FIGURES 2.1 Simulations of the best fit parameter sets for the good healers, MMP 1/TIMP 1 model Simulations of the best fit parameter sets for the poor healers, MMP 1/TIMP 1 model Exemplary monotonicity plot PRCC Plots for M, MMP 1/TIMP 1 model PRCC Plots for T, MMP 1/TIMP 1 model PRCC Plots for E, MMP 1/TIMP 1 model PRCC Plots for f, MMP 1/TIMP 1 model Parameter sensitivities in the good healer model using the boundary method, MMP 1/TIMP 1 model Parameter sensitivities in the poor healer model using the boundary method, MMP 1/TIMP 1 model Parameter sensitivities in the good healer model using the percentage method, MMP 1/TIMP 1 model Parameter sensitivities in the poor healer model using the percentage method, MMP 1/TIMP 1 model Simulations of the best fit parameter sets for the good healers, extended TGF β model Simulations of the best fit parameter sets for the poor healers, extended TGF β model PRCC plots for M, extended TGF-β model PRCC plots for T, extended TGF-β model PRCC plots for E, extended TGF-β model PRCC plots for f, extended TGF-β model PRCC plots for β, extended TGF-β model PRCC plots for bacteria, bacterial infection model PRCC plots for neutrophils, bacterial infection model PRCC plots for oxygen, bacterial infection model ix
12 LIST OF TABLES 2.1 Patient data collected by Muller et al. (28) Nondimensionalized data for modeling Nondimensionalized parameter estimates from curve fitting Global sensitivity analysis results, MMP 1/TIMP 1 model Ranked total sensitivity values computed over observed states, MMP 1/TIMP 1 model Classical sensitivity analysis results, MMP 1/TIMP 1 model Normalized changes in sensitivities for both patient categories and parameter space constructions Parameter estimates for the extended TGF-β model Total sensitivity values, extended TGF-β model Ranked total sensitivity values, extended TGF-β model Parameter values and ranges for bacterial infection model Total sensitivity values, bacterial infection model Ranked total sensitivity values, bacterial infection model x
13 CHAPTER 1 Introduction Mathematical models have recently become increasingly popular tools to describe real world phenomena. Through these models, we can efficiently and inexpensively simulate, understand, and establish baseline characteristics of a system. In biology, these models are often used to describe population ecology (such as predator-prey models) or disease transmission (such as SIR models). At the molecular level, we can use mathematical models to understand processes such as tumorogenesis or wound healing. Applications of mathematics to these and other biological processes provide an entire understanding of the complex systems. Models in molecular biology often describe the concentration of proteins or cell count as they change over time and space. Parameters in these types of models represent empirically observable quantities such as production and decay rates. A qualitative model uses estimates for parameter values that can capture the general behavior of how the cells and proteins change over time. In contrast, a quantitative model is constructed using data; comparisons allow the accuracy of the model to be objectively evaluated. While information can be obtained from both types of model, we are more confident in our model predictions for experimental procedures when using quantitative models. In this work we use a mathematical model to describe the systems that comprise the healing wound. Specifically, we focus on chronic wounds, which are wounds that do not heal in a timely manner. While there are several options for treatment, 1
14 CHAPTER 1. INTRODUCTION 2 including structure dressing and oxygen therapy, treatment is often defined by the type of chronic wound, such as a diabetic foot or leg ulcer. However, the response to treatment varies significantly from patient to patient. Developing strategies to understand the healing response due to a particular treatment for each individual patient should be of primary importance. Identifying strategies to individualize treatment for each patient can significantly reduce the cost of treatment, estimated at reducting $5 1 billion per year in the United States mathematical modeling provides a means. However, current mathematical models for wound healing are limited by the qualitative approach for model developemnt, preventing such usage in the clinical setting; the healing of chronic wounds has been seldom modeled using quantitative methods that rely on patient data. By incorporating individual patient data through model parameters, we can formulate a model to be used in a clinical setting to analyze possibilties for personalized treatment. In this work, three different models are presented. The first, part of a paper submitted for publication, describes the interactions of matrix metalloproteinase and their inhibitors with extracellular matrix during dermal repair of a diabetic foot ulcer. We solve and analyze an inverse problem for the constructed model. Through numerical methods which attempt to minimize the difference between data and model curve, estimates can be determined for each parameter. The results can provide insight into the underlying mechanisms and interactions. In the second chapter, an extended model is suggested which incorporates a universally functional growth factor, transforming growth factor beta (TGF β). TGF β is responsible for the regulation of production for many proteins and cells, further coupling the previously presented equations. Parameters were re estimated for the extended model. Using a different approach to the wound healing process, a model formulated by my mentor, Dr. Richard Schugart, models the process by describing the immune
15 CHAPTER 1. INTRODUCTION 3 system response to a bacterial infection with respect to oxygen concentration. Many parameters for this model were previously estimated in the literature. The formulation of any mathematical model is incomplete without analysis. The noisy nature of biological data demands the use of uncertainty analyses to determine the propensity of this noise to propogate to the model output the ability of uncertainty of an input vector to cause uncertainty in the output. By replicating the noise in the original data, we can observe the changes in the output and determine which inputs have the strongest effect and on which parts of the model. Interpretation of the results of this analysis can reveal limitations of the model and allow for modification to minimize uncertainty. We assess the three models discussed using a Latin Hypercube Sampling/Partial Rank Correlation Coefficient procedure to determine parameter uncertainty and sensitivity.
16 CHAPTER 2 MMP 1/TIMP 1 Model The work shown below details the construction, parameter estimation, and analysis of a model describing the interactions for matrix metalloproteinase and its inhibitors. My work on this model includes parameter estimation, a global sensitivity analysis, and the local to global analysis, as well as the expanded introduction. Work by Nitin Krishna, an alumnus of the Gatton Academy who also worked under Dr. Schugart, is included. His contributions include the local sensitivity analyses, a steady state analysis, and work on the parameter estimation. This work, titled Connecting Local and Global Sensitivities for a Mathematical Model in Wound Healing, was submitted to the Bulletin of Mathematical Biology (Manuscript Number BMAB D ) and is in review. 2.1 Introduction Injury to the dermal layer of the skin requires the body to respond quickly in order to repair and regenerate broken skin. In an acute wound, healing typically progresses through four stages: hemostasis, inflammation, proliferation, and remodeling [22]. Each stage depends on the proper balance of several proteins and cells in the wound. The careful maintenance of the levels of these components contributes to the complex interactions that define the healing response of a wound. A perturbation to this biological system can have a serious effect on the healing of the wound. 4
17 CHAPTER 2. MMP 1/TIMP 1 MODEL 5 The first stage, hemostasis, is characterized by the clotting of the wound, prevention of blood loss, and blockage of damaged vessels. The resulting vasoconstriction at the wound site is immediately followed by histamine-regulated vasodilation, which increases the permeability of the blood vessels in the area [17]. Simultaneously, platelet aggregation leads to the release of many cytokines, especially growth factors such as transforming growth factor beta (TGF-β), acting as chemoattractants for the recruitment of neutrophils and monocytes [39]. The arriving neutrophils may then permeate the vascular membrane to enter the damaged tissue, functioning as an immunological reaction to exposure by phagocytizing waste and dead tissue. This migration marks the beginning of the inflammatory phase. The later arrival of monocytes is concurrent with their transformation into macrophages. These leukocytes phagocytize present neutrophils, bacteria, and dead tissue following activation through T lymphocyte signaling [17]. The growth factors secreted by these activated macrophages recruit mesenchymal and endothelial cells, marking the beginning of the proliferative phase. Fibroblasts, a type of mesenchymal cell, must migrate from nearby tissue to the wound site, where they take on the primary role of producing collagen and other fibrous proteins during dermal repair [17]. This type of repair denotes the reconstruction of the deeper reticular dermis, the layer of skin comprised of extracellular matrix (ECM) that serves as as connective tissue [22]. Collagen, the most abundant protein of ECM, is produced by fibroblasts and acts as the major support structure for the surrounding layers of skin. The formation of the new matrix requires the degradation of damaged matrix components or obstructive bodies in the ECM. Mediators, proteins that activate these cells by acting as a molecular signal to control their release, are responsible for the timing of such events during each stage. Many mediators, such as TGF-β, stimulate the secretion of proteolytic enzymes by fibroblasts, which can clear a path for migration through the ECM [17].
18 CHAPTER 2. MMP 1/TIMP 1 MODEL 6 Among these enzymes are matrix metalloproteinases (MMPs), a family of endopeptidases present at low levels except during wound healing [44]. Once they are activated, MMPs are able to degrade the ECM obstructing the fibroblasts and thus become critical to the proliferation and remodeling of new skin. MMPs are characterized by a catalytic zinc ion; they also contain multiple structural calcium and zinc ions integrated into their catalytic site [21]. The dissociation of a cysteine-zinc ion bond frees the active site, activating MMPs and allowing for degradation of proteins [16]. There are 25 isoforms of MMPs, each specializing in the breakdown of specific proteins comprising the matrix [27]. In the wound-healing process, the subgroups containing collagenases (MMP-1 and MMP-8) and gelatinases (MMP-2 and MMP-9) play a crucial role [31]. Particularly, MMP-1, also known as fibroblast collagenase, acts as the major collagenase in wound healing. MMP-1 lyses type-i, -II, and -III collagen in order to allow fibroblast migration during remodeling and proliferation of the tissue [25]. The heavy deposition of type-i collagen, constituting roughly 9% of collagen in load-bearing structures such as the skin, contributes to the importance of collagenases in wound healing [17]. Successful progression of healing heavily relies on the proper balance between the various proteins interacting in a wound. As a major factor in the reconstruction of the skin at the wound site, MMPs regulate formation of new ECM. Even so, excessive levels of the enzyme may be deleterious. Conversely, lowered concentrations of MMPs allow surplus, uncontrolled production of new ECM, which can lead to hypertrophic scarring or fibrotic conditions such as dermal fibrosis or atherosclerosis [18]. The implied limitation of MMP activity requires precise regulation of MMP expression, crucial to the proper remodeling of skin at the site of the wound during dermal repair. Most regulation of MMP occurs at the transcriptional level through signaling [27]. Expression of select isoforms of MMPs (1, 3, 8, 24) by the fibroblasts can be increased
19 CHAPTER 2. MMP 1/TIMP 1 MODEL 7 by factors such as stratifin, shown to activate the pathway for production of MMP-1 and -3 [42]. These proteins are released by keratinocytes, a prominent epidermal skin cell, and promote the production of various enzymes and matrix deposition by fibroblasts [18]. Atypical communications between keratinocytes and fibroblasts due to low levels or distance can affect the interactions between the two, contributing to a possible significant imbalance in the levels of MMPs [31]. Regulation of MMPs is not only controlled at the level of production, but also on a direct inhibitory level. Tissue inhibitors of metalloproteinases (TIMPs) function by binding the catalytic zinc atom of MMPs in a tight one-to-one complex [4]. TIMPs are comprised of a family of four proteins (TIMP-1, -2, -3, and -4), all of which can inhibit a majority of the MMP family. This inhibition controls the rate of breakdown of ECM during tissue remodeling and angiogenesis. Some TIMPs have strong affinities for specific MMPs, e.g. the relationship between concentrations of TIMP-1 and MMP- 1 [16]. The interactions of fibroblasts, MMPs, and TIMPs are critical to the successful reconstruction of damaged skin. During the proliferative stage, fibroblasts must produce collagen at an increased rate. The heightened activity of these cells begins about four days following injury and continues for four weeks [17]. TGF-β plays a central role by activating fibroblasts for the aforementioned collagen production and regulating protease production to allow the proliferation and gradual growth of new skin [22]. The remodeling stage follows this period of intense epithelialization as MMPs and other enzymes interact with the fibroblasts to reorganize and reconstruct the ECM in the new skin. The collagen composition is structurally modified through an increase in diameter of produced collagen and an intensified presence of cross-linking, strengthening the wound tissue. As the healing progresses in this stage, the rate of collagen synthesis eventually balances with that of collagen destruction, allowing the skin to reach equilibrium once again. The skin begins to become less cellular as the
20 CHAPTER 2. MMP 1/TIMP 1 MODEL 8 excess cells recruited for healing undergo apoptosis. Even so, remodeling of the scar tissue may continue for many years to promote the structural integrity of the skin [43]. In a chronic wound, such as a diabetic foot ulcer, healing may not occur in the systematic manner seen in acute wounds; instead, these wounds often linger in the inflammatory stage of the healing process. The prolonged presence of neutrophils and macrophages in a chronic wound may contribute to the increased length of the inflammatory stage [25]. In this case, neutrophils produce abnormally high levels of MMP, leading to the break down of ECM at a faster rate than the fibroblasts are able to construct it. The resulting imbalance between proliferation and degradation plays a major role in the length of the inflammatory phase and late epithelialization in chronic wounds. The careful regulation of MMPs, especially in chronic wounds, is critical to the nature of wound healing [19]. TIMPs must precisely inhibit MMPs to maintain the production of the ECM at a functional rate, or else risk excessive proteolysis. This role of the TIMPs is therefore extremely important in imbalanced chronic wounds, marking the ratio of MMPs-to-TIMPs as vital to the control of the breakdown of ECM during effective wound healing [25]. Muller et al. (28) showed that the initial ratio of MMP-1-to-TIMP-1 is considered a useful predictive measure of the healing response. Higher levels of MMP-1 early in the wound-healing process favor good healing and may play a part in the successful progression of the proliferation stage. Accordingly, a high level of TIMP-1 later during the healing process indicates the regulation of the MMPs, representing the prevention of excessive breakdown of the ECM. In Muller et al. (28), the authors measured the levels of select isoforms of MMPs and TIMPs (specifically, MMP-1, -2, -8, and -9 and TIMP-1) from wound fluid. In order to identify which of these key proteins contributes significantly to a positive
21 CHAPTER 2. MMP 1/TIMP 1 MODEL 9 healing outcome, the authors divided the patients into groups, identified as good healers and bad healers, and compared the average healing responses of the two groups. A large prospective trial study (Sheehan et al. 23) showed that the percent change in wound surface area over a 4-week period is indicative of how long-term healing will occur. Specifically, the study found the mean percentage reduction in wound area at four weeks for those who healed within 12 weeks was 82%. Therefore, Muller classified good healers by a reduction of at least 82% in initial wound surface area at 4 weeks, and poor healers by a reduction in wound surface area of less than 82% at 4 weeks. The authors concluded that the ratio of MMP-1-to-TIMP-1, as opposed to just taking either measurement separately, measured at Weeks and 2, best correlate and predict a positive healing response at Week 4 and complete healing response at Week 12. Muller hypothesizes an up-regulation of MMP-1 completes the proliferative phase of healing as it has also been shown to be a key mediator for keratinocyte migration re-epithelialization of the epidermal layer [28, 36]. Furthermore, Muller suggests that an increase in TIMP-1 between Weeks and 2 is needed to regulate MMP-1 levels for successful healing. In this work, a mathematical model is formulated, which describes the interactions of MMP-1 and their inhibitors, specifically TIMP-1, and their effect on the healing response, measured by ECM levels. Mathematical models have been formulated to describe and analyze specific aspects of the wound-healing process (for review, see [35] and [1]). While many of these models use differential equations, which often include protein-cell, protein-protein, or cell-cell interactions, they are often qualitative in nature due to the sparseness of temporal data [1]. Although these models can still be used to analyze a physiological process or a treatment response, a model that uses data to measures parameters is, in general, preferable. Ideally, data from a single patient can be used to quantify the model and then the model can be used to predict a future response and individualize treatment. Yet, even averaged patient data can
22 CHAPTER 2. MMP 1/TIMP 1 MODEL 1 still be useful in trying to make model predictions. Averaged data from Muller et al. (28) is used to parameterize our model. After the model has been curve fit to data, the model is then analyzed through steady-state and multiple sensitivity analyses. Sensitivity analysis can help to elucidate which parameters may play a key role in determining or altering the model behavior. Given the range of realistic potential parameter values depending on whether an individual mounts a good or poor healing response, it is important to consider both local and global sensitivity approaches [2]. 2.2 Materials and methods Clinical data Experimental data for modeling were taken from Muller et al. (28), a 12-week prospective trial study that monitored and treated sixteen patients with Type-2 diabetes and neuropathy, ranging in age from 47 to 84, with a chronic diabetic foot ulcer rated 1 to 3, stage A according to the University of Texas Wound Classification. At each visit (weeks, 1, 2, 4, 8, and 12), the percent of wound closure and concentrations of specific isoforms of MMPs (MMP-1, -2, -8, and -9) and TIMP-1 were measured. Proteins were collected in wound fluid using sterile absorbent paper strips placed on the end of the wound for five minutes. In order to avoid variations in the amount of collected fluid, all proteins were measured as ratios of the concentration of the particular isoform of the MMP or TIMP (in picograms) to the total number of proteins (in micrograms). Wound area was measured by examining numeric photography and analytical software. Median data estimated from line graphs in Muller et al. (28) is given in Table 1. In their study, patients were carefully selected and treated in order to minimize excess protein levels in the data collection. For example, patients with soft tissue infections in the wound were excluded from the study since bacteria may secrete
23 CHAPTER 2. MMP 1/TIMP 1 MODEL 11 Table 2.1: Patient data collected by Muller et al. (28). By week 8 7/7 good healers completely healed, whereas by week 12 3/9 poor healers had not healed [25]. Week Good Healers Poor Healers MMP-1 TIMP-1 Wound MMP-1 TIMP-1 Wound (pg/µg) (pg/µg) Closure (pg/µg) (pg/µg) Closure % % % % % % % % % % % excess MMPs. Yet, two patients with osteomyelitis were included since osteomyelitis is not necessarily associated with soft-tissue infection. Wound dressings (as well as oral antibiotics for patients with osteomyelitis) were also chosen to not interfere with MMP levels. To relate our model with healing prediction described in Muller et al. (28), we adopt the same good healers and poor healers categorization. MMP-1 and TIMP-1 data were nondimensionalized with respect to the TIMP-1 concentration at inclusion (Week ). This scaling gives a direct way of identifying the MMP-1 to TIMP-1 ratio that is critical in wound-healing prediction. ECM levels were taken to be proportional to wound closure. Nondimensionalized data is given in Table 2. Table 2.2: Nondimensionalized data for modeling. Good Healers Poor Healers Week MMP-1 TIMP-1 ECM MMP-1 TIMP-1 ECM
24 CHAPTER 2. MMP 1/TIMP 1 MODEL Mathematical Framework We introduce here a data-driven mechanistic model to follow the evolution of matrix metalloproteinases (M ), their inhibitors (T ), extracellular matrix (E), and fibroblasts (f ) with respect to time to describe their interactions in a healing wound. A general mathematical model describing the biological processes is the initial value problem d x dt = f(t, x, θ), x() = x, (2.1) where f : R 1+n+q R n, t R denotes time, x R n denotes the state vector, θ R q denotes the model parameters, and x is the initial state vector at time t =. Model outputs corresponding to available data are computed as y = g(t, x, θ), (2.2) where g : R 1+n+q R m and m is the number of output variables. Data for modeling sampled at times t i are given in a set of data Y, with associated model outputs from Eq. (2.2) in vector y. These vectors are represented by: Y = ( Y 1 (t 11 ), Y 1 (t 12 ),..., Y 1 (t 1k1 ),..., Y m (t m1 ), Y m (t m2 ),..., Y m (t mkm )) T, (2.3) y = ( y 1 (t 11 ), y 1 (t 12 ),..., y 1 (t 1k1 ),..., y m (t m1 ), y m (t m2 ),..., y m (t mkm )) T, (2.4) where Y l and y l are the l th components of the vectors with length K, or the number of observations.
25 CHAPTER 2. MMP 1/TIMP 1 MODEL 13 Model Equations Equation for matrix metalloproteinases (M). We model fibroblast production of MMP-1 by a Hill function of order α [1]. During the wound-healing process, TIMPs bind to MMPs in a one-to-one inhibitor-to-enzyme ratio [4]. This regulatory effect is modeled by an inhibition term, characterized by a rate constant k 4, and is proportional to MMP and TIMP concentrations. Lastly, we assume that MMPs linearly decay with rate constant k 3. With these assumptions, we have: dm dt = k 1M α f k2 α + M }{{ α } Production k 3 M }{{} Decay k 4 MT. (2.5) }{{} Inhibition Equation for tissue inhibitors of matrix metalloproteinases (T). We assume that fibroblasts produce TIMP-1 in the presence of MMPs [27]. We also represent TIMP production by a Hill function of order β [1]. Since the binding of TIMPs to MMPs also reduces TIMP activity, the inhibition term in equation (2.5) is also used to model the inhibition of TIMPs, with the same rate constant k 4. The decay of this protein is assumed to be linear with rate constant k 7. We thus have: dt dt = k 5T β fm k β 6 + T }{{ β } Production k 7 T }{{} Decay k 4 MT. (2.6) }{{} Inhibition Equation for extracellular matrix (E). Collagen, the main component of ECM, is synthesized by fibroblasts [17]. We model production of ECM with a rate constant k 8 and a carrying capacity of E [17, 22]. Since MMP activity degrades ECM [19, 36], we assume this process occurs with rate constant k 9. Taking natural decay of ECM
26 CHAPTER 2. MMP 1/TIMP 1 MODEL 14 to occur linearly with rate constant k 1, we obtain: ) de dt = k 8f (1 EE k 9 ME }{{}}{{} Degradation Production k 1 E. (2.7) }{{} Decay Equation for fibroblasts (f). Although there is no data for fibroblasts, it is included in the model because we need a cell type to produce the proteins MMP-1, TIMP-1, and ECM. In order to do so, we model the proliferation of fibroblasts with a standard logistic growth term with carrying capacity f. We also assume cells die at constant rate k 12 proportional to cell count. We thus have: ) df dt = k 11f (1 ff k 12 f. (2.8) }{{}}{{} Death Production Although the proposed model for fibroblasts is simple due to the absence of cell data, we chose this form of the equation because it still allows us to reliably model the data. While a closed-form solution can be found for this equation, we leave the fibroblast equation in differential-equation form for subsequent work. Nondimensionalization To nondimensionalize equations (2.5) (2.8), we scale the MMP-1 and TIMP-1 levels by the initial concentration of TIMP-1 at Week (T init ), the ECM and fibroblasts by their carrying capacities E and f, respectively, and time by t = 1 day. Under these conditions, nondimensionalized variables and parameters (labeled with a *) are: {M, T, E, f, t } = { M T,, E, f, t }, (2.9) T init T init E f t
27 CHAPTER 2. MMP 1/TIMP 1 MODEL 15 { {k1, k2, k3, k4, k5, k6} k1 f t =, T init {k 7, k 8, k 9, k 1, k 11, k 12} = k 2 T init, k 3 t, k 4 t T init, k 5 f t, k 6 T init { k 7 t, k 8f t E, k 9 t T init, k 1 t, k 11 t, k 12 t }, }. (2.1) Rewriting our model in terms of the dimensionless variables and parameters (and removing * for simplicity) gives the following system of differential equations: dm dt dt dt de dt df dt = k 1M α f k α 2 + M α k 3M k 4 MT, (2.11) = k 5T β fm k β 6 + T β k 7T k 4 MT, (2.12) = k 8 f(1 E) k 9 ME k 1 E, (2.13) = k 11 f(1 f) k 12 f, (2.14) together with the initial conditions M() = M init T init, T () = 1, E() =, f() = f i. (2.15) In terms of equations (2.1) and (2.2), the state vector is x = (M, T, E, f ) with length n = 4, and the model output function gives (M, T, E ) with length m = 3. The initial state vector is x = (M init /T init, 1,, f i ). Our model has q = 12 parameters, given by θ = (k 1, k 2, k 3, k 4, k 5, k 6, k 7, k 8, k 9, k 1, k 11, k 12) T. (2.16) Nondimensionalized equations (2.11) (2.14) and initial conditions (2.15) were chosen as the simplest system of equations that captures the key fundamental mechanisms
28 CHAPTER 2. MMP 1/TIMP 1 MODEL 16 of healing in diabetic foot ulcers Parameter Estimation Curve fitting each state variable to the data was done by minimizing the leastsquares residuals between collected data and model output (Eq. (2.3) and (2.4)). Matlab was used to solve the nonlinear optimization problem of minimizing the functional (J -value) K J = y l Y l 2. (2.17) l=1 Values for y l were approximated by the stiff differential equation solver ode15s with an absolute and relative error tolerance of O(1 6 ). Matlab s GlobalSearch and fmincon were used to find the global minimum of (2.17). GlobalSearch uses a scatter search algorithm to generate a programmed number of trial parameters subject to given optimizing constraints, and fmincon rejects those that were unlikely to improve the best local minimum found so far [23]. To keep parameters within a numerically feasible range, GlobalSearch searched for parameter values between and 2. The sets of parameters corresponding to the smallest J -values for good healers and poor healers were recorded, and graphs of the curve-fits were used to evaluate sources of error and compare good and poor healers Steady-State Analysis Analyzing the mathematical model independently of parameter values or patient category is important in interpreting the biology of the system of equations, as the results can then be used to draw conclusions for patients not included in the parameter estimation. A steady-state analysis was conducted to find the levels of the proteins and cells as t. We solved for the model equilibria, and stability was assessed by computing the Jacobian matrix. When the steady-state equations could not be solved algebraically, steady-states and their stabilities were identified by intersections
29 CHAPTER 2. MMP 1/TIMP 1 MODEL 17 of the nullclines of the differential equations Global sensitivity analysis The existence of multiple steady-states implies that the convergence to an end state is dependent upon the values of specific parameters. A small change in a parameter value could be the difference between convergence to a healed state or a non-healed state. Sensitivity analyses were conducted to determine how sensitive the system of equations is to changes in parameter value(s). A parameter whose change has a large impact on the model output is said to be sensitive and a parameter whose change has a negligible impact on the model output is said to be insensitive. A global sensitivity analysis is independent of parameter estimates and covers a wide range of the parameter space, while a local analysis examines a small neighborhood of a particular point in the parameter space. Both global and local analyses were conducted in this work. A global analysis was conducted using a Latin Hypercube Sampling (LHS) scheme combined with Partial Rank Correlation Coefficient (PRCC) analysis. LHS, a type of stratified Monte Carlo sampling scheme, assigns a probability distribution to each parameter, partitions the intervals in the distribution into a given number of equiprobable regions, and samples each of the intervals without replacement. The sampling provides a global analysis of the parameter space by treating each model parameter as a random variable [13, 24, 37]. Partial rank correlation measures the strength of the relationship between the output of each state function and the model parameters [15, 2]. Standard correlation measures the strength of a linear association between variables (e.g. a parameter and output). The PRCC value measures the correlation of the ranked ordering of the variable values rather than the variable values themselves [5]. The advantage of the PRCC over traditional correlation measures is that the rank transformation allows
30 CHAPTER 2. MMP 1/TIMP 1 MODEL 18 us to consider non-linear, monotonic relationships [12, 14]. Partial rank correlation coefficients provide a measure of the strength of the statistical relationship between each parameter and state function, while discounting the effects of the other parameters on the outputs. PRCC values range from 1 to 1, with the magnitude indicating the sensitivity of the state function to the parameter uncertainty and the sign indicating whether the correlation is positive or negative. The p-value of the PRCC indicates the significance of the value. A state variable is sensitive to a parameter if the absolute value of the PRCC is greater than.5 and the corresponding p-value is less than.5. If each observation in Eq. (2.4) (i.e. each time point for each observed state variable) is monotonic (described below) with respect to each input, the algorithm for the LHS/PRCC method is described as follows: 1. Assign a probability density function to each model input of interest. In this work, we are concerned with parameter sensitivity, so for the i th parameter, assign a probability density function on the interval [l i, u i ], where l i and u i are the lower and upper bounds in the parameter estimation (1 i q). 2. Partition each [l i, u i ] into N equally probable subintervals. As in [3, 24], we require N > 4q For the i th parameter, randomly sample the subintervals without replacement. Then sample a random number from each subinterval and enter randomized list into the ith column of the N q matrix L. Each row of matrix L can be considered to be a set of parameter values. 4. Compute the model outputs for all N sets of parameter values at a given time point. 5. Rank the N entries in each column of L from smallest to largest. Separately, for each state variable, rank the N model outputs from smallest to largest.
31 CHAPTER 2. MMP 1/TIMP 1 MODEL Perform a multiple linear regression for each set of ranked parameters and each ranked output in relation to all other parameter rankings. 7. To calculate the PRCC for a parameter and output combination, plot the residuals from the corresponding regressions and compute the Pearson linear correlation coefficient, our PRCC value, and associated p-value for the correlation of the residuals [2]. 8. Repeat steps times. Let PRCC i,j,k and p i,j,k be the PRCC value and p-value of the i th parameter with respect to the j th state variable in the k th PRCC calculation (1 i q, 1 j n, 1 k 1). 9. Compute the sensitivity S i,j of the i th parameter with respect to the j th state variable using a weighted norm: S i,j = 1 (PRCC i,j,k (1 p i,j,k )) 2. (2.18) k=1 1. Compute the total sensitivity S i of the i th parameter. Let I be the set of indices of the states we are interested in using to determine sensitivity. Then S i = Si,j 2. (2.19) j I To verify monotonicity of the state variables with respect to the m th parameter, we constructed an N q matrix M = (M ij ), where N is the number of simulations (Step 2) and M ij is the baseline value for the j th parameter. We replaced the m th column of this matrix with the m th column of a Latin Hypercube Matrix. Next, we generated model outputs at a fixed time point using each of the N sets of parameters. Plotting the output against the corresponding values for the m th parameter, we visually observed if the relationship was monotonic. If a state function was not monotonic
32 CHAPTER 2. MMP 1/TIMP 1 MODEL 2 with respect to a parameter over the entire parameter space, we conducted the PRCC analysis over the intervals for which the relationship was monotonic Local sensitivity analyses Classical sensitivity analysis A classical sensitivity analysis described in [8] and [9] quantifies and ranks the sensitivities of individual model parameters with respect to other parameters. The sensitivity of the model output y with respect to a single parameter θ i can be represented by the columns of a sensitivity matrix, S = y θ = y 1 / θ 1 (t 11 ) y 1 / θ q (t 11 )... y 1 / θ 1 (t 1k1 ) y 1 / θ q (t 1k1 ).... (2.2) y m / θ 1 (t m1 ) y m / θ q (t m1 )... y m / θ 1 (t mkm ) y m / θ q (t mkm ) Methods used in the sensitivity analysis are more efficient if parameter values are the same order of magnitude [29], so we scaled the parameters by the natural logarithm, i.e. θ = ln θ. Relative sensitivities were computed by multiplying each element by a nondimensionalizing factor S = y 1 θ y. (2.21) Partial derivatives were computed using forward finite-difference approximations y k y k(t, x, θ + h e j ) y k (t, x, θ), θ j h
33 CHAPTER 2. MMP 1/TIMP 1 MODEL 21 where e j is the j th standard basis vector in R q. A scaled 2-norm is used to evaluate the total sensitivity, S j, of the j th parameter ( 1 S j = K 1/2 K Si,j) 2. i=1 To separate sensitive and insensitive parameters we consider the computational accuracy of finite-difference approximation and hence the sensitivity matrix. Following Pope et al. (29) and Olufsen et al. (213), the error in computing the sensitivity matrix is one order of magnitude greater than the square root of the absolute error tolerance of the ode solver, or O(1 2 ). Parameters with sensitivity values greater than 1 2 were classified as sensitive. Although sensitivity analyses employing analytical or automatic differentiation (see [7] and [38]) are more accurate, forward finite difference approximation using scaled parameter values is computationally efficient for a complex system of equations. SVD-QR subset selection While the classical sensitivity analysis provides a way of identifying the sensitivity of parameters individually, it is also advantageous to find a subset of parameters sensitive as a group. The subset selection uses the singular value decomposition to determine the numerical rank ρ of the sensitivity matrix. The rank is used to determine the number of parameters in the subset. The QR factorization of the right matrix of eigenvectors after column pivoting determines which ρ parameters are sensitive as a group. As stated in the classical sensitivity analysis section, the errors of the sensitivities are O(1 2 ), so the smallest allowable singular value in determining the rank was set as 1 2. The SVD-QR algorithm for subset selection described in Olufsen et al. (213) is as follows:
34 CHAPTER 2. MMP 1/TIMP 1 MODEL Using parameter estimates in θ and the sensitivity matrix (2.21), calculate the singular value decomposition, S = UΣV T. 2. Calculate ρ, the rank of the scaled sensitivity matrix. Here, ρ is the number of singular values (extracted from Σ) greater than Partition V into V = [V ρ V n ρ ]. 4. Determine a permutation matrix P such that V T ρ P = QR, where R is an upper triangular matrix with diagonal elements in decreasing order of magnitude. This is done with column pivoting after the maximal element for V T ρ [11]. 5. Use P to re-order the vector of parameters θ as θ = P T θ. 6. The first ρ elements of the re-ordered vector of parameters are sensitive as a group Local-to-global analysis To compare the results of the local and global sensitivity analyses, we developed a local-to-global analysis to study the sensitivities of the parameters as the parameter space in the LHS/PRCC analysis shifts from local to global. Given the parameter estimates in vector θ = (θ 1,..., θ q ), we construct 1 spaces in two ways: Boundary method: the nth parameter space is given by the set of all points P (n) = (θ (n) 1,..., θ q (n) ) whose coordinates satisfy the inequalities θ i n (θ i l i )/1 θ (n) i θ i + n (u i θ i )/1, where l i and u i are the lower and upper bounds for θ i in the parameter estimation. Percentage method: the nth parameter space is given by the set of all points P (n) = (θ (n) 1,..., θ q (n) ) whose coordinates satisfy the inequalities θ i (1 n/1) θ (n) i θ i (1 + n/1).
35 CHAPTER 2. MMP 1/TIMP 1 MODEL 23 The first method increases the bounds of the parameter spaces linearly until the global bounds are reached, while the second method constructs intervals around the parameter estimates that grow from 1% to 1% of the estimate in each direction. We conduct an LHS/PRCC analysis for each parameter space and plot the total sensitivities and sensitivity rankings of each parameter over the 1 iterations. We quantify the magnitude of the change in sensitivity of the ith parameter by S i = max S i,n min S i,n, 1 n 1 1 n 1 where S i,n is the total sensitivity of the ith parameter in the nth parameter space computed using Eqs. (2.18) and (2.19). The changes in sensitivity are normalized by the smallest change for all parameters. That is, S i = S i min 1 i q S i. The relative changes, together with the sensitivity plots, were used to directly compare the results of the two types of sensitivity analyses done in this work and to determine parameters whose sensitivity varied the most as the parameter space expanded from local to global. 2.3 Results Using the numerical methods and curve fitting procedures described in the methods section, we found optimal parameter values for the data from each patient group. Because there is no data for fibroblasts, an initial condition for the ODE solver was found through simulations. Different initial values for f() = f i were tested, and.1 most consistently gave the best J -value. Similarly, we determined Hill function orders in the MMP and TIMP equations to be α = β = 3. The parameter sets corresponding to the minimized least-squares residual for both patient groups are given
36 CHAPTER 2. MMP 1/TIMP 1 MODEL 24 Table 2.3: Nondimensionalized parameter estimates from curve fitting. The J -values indicate there is very good agreement between the model and data for the good healers and excellent agreement for the poor healers. Graphs in Figures 2.1 and 2.2 illustrate the agreement as well. Parameter Good Healers Poor Healers k k k k k k k k k k k k J in Table 2.3, with corresponding plots of the model fitted to the data in Figures 2.1 and 2.2. The best sets of parameters for both good healers and poor healers were found when GlobalSearch routine selected 6 sets of values for θ. Simulations run for more than 6 trial points generated parameters that corresponded to the same model outputs. The parameter sets generated by 6 trial points were selected because they contained the least number of parameters close to the boundary of the parameter space and were more likely to be biologically feasible.
An Investigation of the Accuracy of Parallel Analysis for Determining the Number of Factors in a Factor Analysis
 Western Kentucky University TopSCHOLAR Honors College Capstone Experience/Thesis Projects Honors College at WKU 6-28-2017 An Investigation of the Accuracy of Parallel Analysis for Determining the Number
Western Kentucky University TopSCHOLAR Honors College Capstone Experience/Thesis Projects Honors College at WKU 6-28-2017 An Investigation of the Accuracy of Parallel Analysis for Determining the Number
Experimental designs for multiple responses with different models
 Graduate Theses and Dissertations Graduate College 2015 Experimental designs for multiple responses with different models Wilmina Mary Marget Iowa State University Follow this and additional works at:
Graduate Theses and Dissertations Graduate College 2015 Experimental designs for multiple responses with different models Wilmina Mary Marget Iowa State University Follow this and additional works at:
Supplementary Figures
 Supplementary Figures a x 1 2 1.5 1 a 1 = 1.5 a 1 = 1.0 0.5 a 1 = 1.0 b 0 0 20 40 60 80 100 120 140 160 2 t 1.5 x 2 1 0.5 a 1 = 1.0 a 1 = 1.5 a 1 = 1.0 0 0 20 40 60 80 100 120 140 160 t Supplementary Figure
Supplementary Figures a x 1 2 1.5 1 a 1 = 1.5 a 1 = 1.0 0.5 a 1 = 1.0 b 0 0 20 40 60 80 100 120 140 160 2 t 1.5 x 2 1 0.5 a 1 = 1.0 a 1 = 1.5 a 1 = 1.0 0 0 20 40 60 80 100 120 140 160 t Supplementary Figure
M U LT I C E L L U L A R O R G A N I Z AT I O N
 LESSON 6 M U LT I C E L L U L A R O R G A N I Z AT I O N LEVELS OF ORGANIZATION Multicellular organisms have five levels of cellular organization Cells-} Tissues-} Organs-} Organ System-} Organism LEVELS
LESSON 6 M U LT I C E L L U L A R O R G A N I Z AT I O N LEVELS OF ORGANIZATION Multicellular organisms have five levels of cellular organization Cells-} Tissues-} Organs-} Organ System-} Organism LEVELS
A mathematical and computational model of necrotizing enterocolitis
 A mathematical and computational model of necrotizing enterocolitis Ivan Yotov Department of Mathematics, University of Pittsburgh McGowan Institute Scientific Retreat March 10-12, 2008 Acknowledgment:
A mathematical and computational model of necrotizing enterocolitis Ivan Yotov Department of Mathematics, University of Pittsburgh McGowan Institute Scientific Retreat March 10-12, 2008 Acknowledgment:
CONTROLLING WOUND HEALING THROUGH DEBRIDEMENT
 CONTROLLING WOUND HEALING THROUGH DEBRIDEMENT MICHAEL A JONES AND DIANA M THOMAS Abstract The formation of slough (dead tissue) on a wound is widely accepted as an inhibitor to natural wound healing In
CONTROLLING WOUND HEALING THROUGH DEBRIDEMENT MICHAEL A JONES AND DIANA M THOMAS Abstract The formation of slough (dead tissue) on a wound is widely accepted as an inhibitor to natural wound healing In
Controlling Systemic Inflammation Using NMPC. Using Nonlinear Model Predictive Control with State Estimation
 Controlling Systemic Inflammation Using Nonlinear Model Predictive Control with State Estimation Gregory Zitelli, Judy Day July 2013 With generous support from the NSF, Award 1122462 Motivation We re going
Controlling Systemic Inflammation Using Nonlinear Model Predictive Control with State Estimation Gregory Zitelli, Judy Day July 2013 With generous support from the NSF, Award 1122462 Motivation We re going
Electrostatic Breakdown Analysis
 Utah State University DigitalCommons@USU Senior Theses and Projects Materials Physics 11-18-2014 Electrostatic Breakdown Analysis Sam Hansen Utah State University Follow this and additional works at: https://digitalcommons.usu.edu/mp_seniorthesesprojects
Utah State University DigitalCommons@USU Senior Theses and Projects Materials Physics 11-18-2014 Electrostatic Breakdown Analysis Sam Hansen Utah State University Follow this and additional works at: https://digitalcommons.usu.edu/mp_seniorthesesprojects
Comparing Group Means When Nonresponse Rates Differ
 UNF Digital Commons UNF Theses and Dissertations Student Scholarship 2015 Comparing Group Means When Nonresponse Rates Differ Gabriela M. Stegmann University of North Florida Suggested Citation Stegmann,
UNF Digital Commons UNF Theses and Dissertations Student Scholarship 2015 Comparing Group Means When Nonresponse Rates Differ Gabriela M. Stegmann University of North Florida Suggested Citation Stegmann,
Inferences about Parameters of Trivariate Normal Distribution with Missing Data
 Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 7-5-3 Inferences about Parameters of Trivariate Normal Distribution with Missing
Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 7-5-3 Inferences about Parameters of Trivariate Normal Distribution with Missing
Problem Set # 3
 20.320 Problem Set # 3 October 1 st, 2010 Due on October 8 th, 2010 at 11:59am. No extensions, no electronic submissions. General Instructions: 1. You are expected to state all your assumptions and provide
20.320 Problem Set # 3 October 1 st, 2010 Due on October 8 th, 2010 at 11:59am. No extensions, no electronic submissions. General Instructions: 1. You are expected to state all your assumptions and provide
ADVANCED PLACEMENT BIOLOGY
 ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
ADVANCED PLACEMENT BIOLOGY Description Advanced Placement Biology is designed to be the equivalent of a two-semester college introductory course for Biology majors. The course meets seven periods per week
Networks in systems biology
 Networks in systems biology Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2017 M. Macauley (Clemson) Networks in systems
Networks in systems biology Matthew Macauley Department of Mathematical Sciences Clemson University http://www.math.clemson.edu/~macaule/ Math 4500, Spring 2017 M. Macauley (Clemson) Networks in systems
Modeling the Immune System W9. Ordinary Differential Equations as Macroscopic Modeling Tool
 Modeling the Immune System W9 Ordinary Differential Equations as Macroscopic Modeling Tool 1 Lecture Notes for ODE Models We use the lecture notes Theoretical Fysiology 2006 by Rob de Boer, U. Utrecht
Modeling the Immune System W9 Ordinary Differential Equations as Macroscopic Modeling Tool 1 Lecture Notes for ODE Models We use the lecture notes Theoretical Fysiology 2006 by Rob de Boer, U. Utrecht
COURSE NUMBER: EH 590R SECTION: 1 SEMESTER: Fall COURSE TITLE: Computational Systems Biology: Modeling Biological Responses
 DEPARTMENT: Environmental Health COURSE NUMBER: EH 590R SECTION: 1 SEMESTER: Fall 2017 CREDIT HOURS: 2 COURSE TITLE: Computational Systems Biology: Modeling Biological Responses COURSE LOCATION: TBD PREREQUISITE:
DEPARTMENT: Environmental Health COURSE NUMBER: EH 590R SECTION: 1 SEMESTER: Fall 2017 CREDIT HOURS: 2 COURSE TITLE: Computational Systems Biology: Modeling Biological Responses COURSE LOCATION: TBD PREREQUISITE:
Taguchi Method and Robust Design: Tutorial and Guideline
 Taguchi Method and Robust Design: Tutorial and Guideline CONTENT 1. Introduction 2. Microsoft Excel: graphing 3. Microsoft Excel: Regression 4. Microsoft Excel: Variance analysis 5. Robust Design: An Example
Taguchi Method and Robust Design: Tutorial and Guideline CONTENT 1. Introduction 2. Microsoft Excel: graphing 3. Microsoft Excel: Regression 4. Microsoft Excel: Variance analysis 5. Robust Design: An Example
Enduring understanding 1.A: Change in the genetic makeup of a population over time is evolution.
 The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
The AP Biology course is designed to enable you to develop advanced inquiry and reasoning skills, such as designing a plan for collecting data, analyzing data, applying mathematical routines, and connecting
Supplemental table S7.
 Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
Supplemental table S7. GO terms significantly enriched in significantly up-regulated genes of the microarray. K: number of genes from the input cluster in the given category. F: number of total genes in
CHAPTER 2 Estimating Probabilities
 CHAPTER 2 Estimating Probabilities Machine Learning Copyright c 2017. Tom M. Mitchell. All rights reserved. *DRAFT OF September 16, 2017* *PLEASE DO NOT DISTRIBUTE WITHOUT AUTHOR S PERMISSION* This is
CHAPTER 2 Estimating Probabilities Machine Learning Copyright c 2017. Tom M. Mitchell. All rights reserved. *DRAFT OF September 16, 2017* *PLEASE DO NOT DISTRIBUTE WITHOUT AUTHOR S PERMISSION* This is
Computational Biology Course Descriptions 12-14
 Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
Computational Biology Course Descriptions 12-14 Course Number and Title INTRODUCTORY COURSES BIO 311C: Introductory Biology I BIO 311D: Introductory Biology II BIO 325: Genetics CH 301: Principles of Chemistry
A. Incorrect! The Cell Cycle contains 4 distinct phases: (1) G 1, (2) S Phase, (3) G 2 and (4) M Phase.
 Molecular Cell Biology - Problem Drill 21: Cell Cycle and Cell Death Question No. 1 of 10 1. Which of the following statements about the cell cycle is correct? Question #1 (A) The Cell Cycle contains 3
Molecular Cell Biology - Problem Drill 21: Cell Cycle and Cell Death Question No. 1 of 10 1. Which of the following statements about the cell cycle is correct? Question #1 (A) The Cell Cycle contains 3
Analysing data: regression and correlation S6 and S7
 Basic medical statistics for clinical and experimental research Analysing data: regression and correlation S6 and S7 K. Jozwiak k.jozwiak@nki.nl 2 / 49 Correlation So far we have looked at the association
Basic medical statistics for clinical and experimental research Analysing data: regression and correlation S6 and S7 K. Jozwiak k.jozwiak@nki.nl 2 / 49 Correlation So far we have looked at the association
Programmed Cell Death
 Programmed Cell Death Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id What is apoptosis? a normal component of the development
Programmed Cell Death Dewajani Purnomosari Department of Histology and Cell Biology Faculty of Medicine Universitas Gadjah Mada d.purnomosari@ugm.ac.id What is apoptosis? a normal component of the development
Why This Class? James K. Peterson. August 22, Department of Biological Sciences and Department of Mathematical Sciences Clemson University
 Why This Class? James K. Peterson Department of Biological Sciences and Department of Mathematical Sciences Clemson University August 22, 2013 Outline 1 Our Point of View Mathematics, Science and Computer
Why This Class? James K. Peterson Department of Biological Sciences and Department of Mathematical Sciences Clemson University August 22, 2013 Outline 1 Our Point of View Mathematics, Science and Computer
Self-healing tile sets: rates of regrowth in damaged assemblies
 University of Arkansas, Fayetteville ScholarWorks@UARK Mathematical Sciences Undergraduate Honors Theses Mathematical Sciences 5-2015 Self-healing tile sets: rates of regrowth in damaged assemblies Kevin
University of Arkansas, Fayetteville ScholarWorks@UARK Mathematical Sciences Undergraduate Honors Theses Mathematical Sciences 5-2015 Self-healing tile sets: rates of regrowth in damaged assemblies Kevin
Principles of Synthetic Biology: Midterm Exam
 Principles of Synthetic Biology: Midterm Exam October 28, 2010 1 Conceptual Simple Circuits 1.1 Consider the plots in figure 1. Identify all critical points with an x. Put a circle around the x for each
Principles of Synthetic Biology: Midterm Exam October 28, 2010 1 Conceptual Simple Circuits 1.1 Consider the plots in figure 1. Identify all critical points with an x. Put a circle around the x for each
Toxicological Targets. Russell L. Carr Department of Basic Sciences College of Veterinary Medicine
 Toxicological Targets Russell L. Carr Department of Basic Sciences College of Veterinary Medicine Toxicology Definitions = study of poisons Poison = any agent capable of producing a deleterious response
Toxicological Targets Russell L. Carr Department of Basic Sciences College of Veterinary Medicine Toxicology Definitions = study of poisons Poison = any agent capable of producing a deleterious response
Integrated reliable and robust design
 Scholars' Mine Masters Theses Student Research & Creative Works Spring 011 Integrated reliable and robust design Gowrishankar Ravichandran Follow this and additional works at: http://scholarsmine.mst.edu/masters_theses
Scholars' Mine Masters Theses Student Research & Creative Works Spring 011 Integrated reliable and robust design Gowrishankar Ravichandran Follow this and additional works at: http://scholarsmine.mst.edu/masters_theses
Biology Assessment. Eligible Texas Essential Knowledge and Skills
 Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Biology Assessment Eligible Texas Essential Knowledge and Skills STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules
Mathematical Models of Biological Systems
 Mathematical Models of Biological Systems CH924 November 25, 2012 Course outline Syllabus: Week 6: Stability of first-order autonomous differential equation, genetic switch/clock, introduction to enzyme
Mathematical Models of Biological Systems CH924 November 25, 2012 Course outline Syllabus: Week 6: Stability of first-order autonomous differential equation, genetic switch/clock, introduction to enzyme
RISK- AND RELIABILITY ANALYSIS WITH APPLICATIONS
 RISK- AND RELIABILITY ANALYSIS WITH APPLICATIONS by ARNE BANG HUSEBY and KRISTINA ROGNLIEN DAHL Department of Mathematics Faculty of Mathematics and Natural Sciences University of Oslo ii iii Abstract
RISK- AND RELIABILITY ANALYSIS WITH APPLICATIONS by ARNE BANG HUSEBY and KRISTINA ROGNLIEN DAHL Department of Mathematics Faculty of Mathematics and Natural Sciences University of Oslo ii iii Abstract
Organizing Diversity Taxonomy is the discipline of biology that identifies, names, and classifies organisms according to certain rules.
 1 2 3 4 5 6 7 8 9 10 Outline 1.1 Introduction to AP Biology 1.2 Big Idea 1: Evolution 1.3 Big Idea 2: Energy and Molecular Building Blocks 1.4 Big Idea 3: Information Storage, Transmission, and Response
1 2 3 4 5 6 7 8 9 10 Outline 1.1 Introduction to AP Biology 1.2 Big Idea 1: Evolution 1.3 Big Idea 2: Energy and Molecular Building Blocks 1.4 Big Idea 3: Information Storage, Transmission, and Response
M469, Fall 2010, Practice Problems for the Final
 M469 Fall 00 Practice Problems for the Final The final exam for M469 will be Friday December 0 3:00-5:00 pm in the usual classroom Blocker 60 The final will cover the following topics from nonlinear systems
M469 Fall 00 Practice Problems for the Final The final exam for M469 will be Friday December 0 3:00-5:00 pm in the usual classroom Blocker 60 The final will cover the following topics from nonlinear systems
Savannah State University New Programs and Curriculum Committee Summary Page Form I
 Summary Page Form I 1. Submitting College: COST 2. Department(s) Generating The Proposal: Natural Sciences Choose an item. (if needed) 3. Proposal Title: Revised Chemistry Program Curriculum 4. Course
Summary Page Form I 1. Submitting College: COST 2. Department(s) Generating The Proposal: Natural Sciences Choose an item. (if needed) 3. Proposal Title: Revised Chemistry Program Curriculum 4. Course
Confidence Intervals for the Ratio of Two Exponential Means with Applications to Quality Control
 Western Kentucky University TopSCHOLAR Student Research Conference Select Presentations Student Research Conference 6-009 Confidence Intervals for the Ratio of Two Exponential Means with Applications to
Western Kentucky University TopSCHOLAR Student Research Conference Select Presentations Student Research Conference 6-009 Confidence Intervals for the Ratio of Two Exponential Means with Applications to
Big Idea 1: The process of evolution drives the diversity and unity of life.
 Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
Big Idea 1: The process of evolution drives the diversity and unity of life. understanding 1.A: Change in the genetic makeup of a population over time is evolution. 1.A.1: Natural selection is a major
LESSON 2.2 WORKBOOK. How is a cell born? Workbook Lesson 2.2
 For a complete list of defined terms, see the Glossary. Cell cycle the progression of events that prepares a cell to replicate, and then leads to division into two daughter cells. Mitosis the phase of
For a complete list of defined terms, see the Glossary. Cell cycle the progression of events that prepares a cell to replicate, and then leads to division into two daughter cells. Mitosis the phase of
Campbell Biology AP Edition 11 th Edition, 2018
 A Correlation and Narrative Summary of Campbell Biology AP Edition 11 th Edition, 2018 To the AP Biology Curriculum Framework AP is a trademark registered and/or owned by the College Board, which was not
A Correlation and Narrative Summary of Campbell Biology AP Edition 11 th Edition, 2018 To the AP Biology Curriculum Framework AP is a trademark registered and/or owned by the College Board, which was not
An Introduction to Metabolism
 An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
An Introduction to Metabolism I. All of an organism=s chemical reactions taken together is called metabolism. A. Metabolic pathways begin with a specific molecule, which is then altered in a series of
Linear Regression and Its Applications
 Linear Regression and Its Applications Predrag Radivojac October 13, 2014 Given a data set D = {(x i, y i )} n the objective is to learn the relationship between features and the target. We usually start
Linear Regression and Its Applications Predrag Radivojac October 13, 2014 Given a data set D = {(x i, y i )} n the objective is to learn the relationship between features and the target. We usually start
General Model of the Innate Immune Response
 General Model of the Innate Immune Response Katherine Reed, Kathryn Schalla, Souad Sosa, Jackie Tran, Thuy-My Truong, Alicia Prieto Langarica, Betty Scarbrough, Hristo Kojouharov, James Grover Technical
General Model of the Innate Immune Response Katherine Reed, Kathryn Schalla, Souad Sosa, Jackie Tran, Thuy-My Truong, Alicia Prieto Langarica, Betty Scarbrough, Hristo Kojouharov, James Grover Technical
Delivery. Delivery Processes. Delivery Processes: Distribution. Ultimate Toxicant
 Delivery Ultimate Toxicant The chemical species that reacts with the endogenous target. Toxicity depends on the concentration (dose) of the ultimate toxicant at the target site Delivery Processes Absorption
Delivery Ultimate Toxicant The chemical species that reacts with the endogenous target. Toxicity depends on the concentration (dose) of the ultimate toxicant at the target site Delivery Processes Absorption
16 The Cell Cycle. Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization
 The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
The Cell Cycle 16 The Cell Cycle Chapter Outline The Eukaryotic Cell Cycle Regulators of Cell Cycle Progression The Events of M Phase Meiosis and Fertilization Introduction Self-reproduction is perhaps
STAAR Biology Assessment
 STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
STAAR Biology Assessment Reporting Category 1: Cell Structure and Function The student will demonstrate an understanding of biomolecules as building blocks of cells, and that cells are the basic unit of
MITOCW enzyme_kinetics
 MITOCW enzyme_kinetics In beer and wine production, enzymes in yeast aid the conversion of sugar into ethanol. Enzymes are used in cheese-making to degrade proteins in milk, changing their solubility,
MITOCW enzyme_kinetics In beer and wine production, enzymes in yeast aid the conversion of sugar into ethanol. Enzymes are used in cheese-making to degrade proteins in milk, changing their solubility,
Explain your answer:
 Biology Midterm Exam Review Introduction to Biology and the Scientific Method Name: Date: Hour: 1. Biology is the study of: 2. A living thing is called a(n): 3. All organisms are composed of: 4. The smallest
Biology Midterm Exam Review Introduction to Biology and the Scientific Method Name: Date: Hour: 1. Biology is the study of: 2. A living thing is called a(n): 3. All organisms are composed of: 4. The smallest
Comparison of Some Improved Estimators for Linear Regression Model under Different Conditions
 Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 3-24-2015 Comparison of Some Improved Estimators for Linear Regression Model under
Florida International University FIU Digital Commons FIU Electronic Theses and Dissertations University Graduate School 3-24-2015 Comparison of Some Improved Estimators for Linear Regression Model under
1 Mathematics and Statistics in Science
 1 Mathematics and Statistics in Science Overview Science students encounter mathematics and statistics in three main areas: Understanding and using theory. Carrying out experiments and analysing results.
1 Mathematics and Statistics in Science Overview Science students encounter mathematics and statistics in three main areas: Understanding and using theory. Carrying out experiments and analysing results.
Signal Transduction. Dr. Chaidir, Apt
 Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Signal Transduction Dr. Chaidir, Apt Background Complex unicellular organisms existed on Earth for approximately 2.5 billion years before the first multicellular organisms appeared.this long period for
Atherosclerosis Initiation Modeled as an Inflammatory Process
 Math. Model. Nat. Phenom. Vol. 2, No. 2, 2007, pp. 126-141 Atherosclerosis Initiation Modeled as an Inflammatory Process N. El Khatib 1, S. Génieys and V. Volpert Université Lyon 1, Institut Camille Jordan,
Math. Model. Nat. Phenom. Vol. 2, No. 2, 2007, pp. 126-141 Atherosclerosis Initiation Modeled as an Inflammatory Process N. El Khatib 1, S. Génieys and V. Volpert Université Lyon 1, Institut Camille Jordan,
arxiv: v1 [q-bio.to] 15 Apr 2016
![arxiv: v1 [q-bio.to] 15 Apr 2016 arxiv: v1 [q-bio.to] 15 Apr 2016](/thumbs/78/77676506.jpg) Spatio-temporal Models of Lymphangiogenesis in Wound Healing Arianna Bianchi, Kevin J. Painter, Jonathan A. Sherratt 2016 arxiv:1604.04654v1 [q-bio.to] 15 Apr 2016 ABSTRACT: Several studies suggest that
Spatio-temporal Models of Lymphangiogenesis in Wound Healing Arianna Bianchi, Kevin J. Painter, Jonathan A. Sherratt 2016 arxiv:1604.04654v1 [q-bio.to] 15 Apr 2016 ABSTRACT: Several studies suggest that
A Simple Model s Best Hope: A Brief Introduction to Universality
 A Simple Model s Best Hope: A Brief Introduction to Universality Benjamin Good Swarthmore College (Dated: May 5, 2008) For certain classes of systems operating at a critical point, the concept of universality
A Simple Model s Best Hope: A Brief Introduction to Universality Benjamin Good Swarthmore College (Dated: May 5, 2008) For certain classes of systems operating at a critical point, the concept of universality
LABETTE COMMUNITY COLLEGE BRIEF SYLLABUS. ANATOMY AND PHYSIOLOGY, lecture and lab
 LABETTE COMMUNITY COLLEGE BRIEF SYLLABUS SPECIAL NOTE: This brief syllabus is not intended to be a legal contract. A full syllabus will be distributed to students at the first class session. TEXT AND SUPPLEMENTARY
LABETTE COMMUNITY COLLEGE BRIEF SYLLABUS SPECIAL NOTE: This brief syllabus is not intended to be a legal contract. A full syllabus will be distributed to students at the first class session. TEXT AND SUPPLEMENTARY
The effects of histone and glycosaminoglycans on human factor Xa and antithrombin III interactions
 Bowling Green State University ScholarWorks@BGSU Honors Projects Honors College Fall 12-12-2013 The effects of histone and glycosaminoglycans on human factor and antithrombin III interactions Amy Thomas
Bowling Green State University ScholarWorks@BGSU Honors Projects Honors College Fall 12-12-2013 The effects of histone and glycosaminoglycans on human factor and antithrombin III interactions Amy Thomas
BIOL 51A - Biostatistics 1 1. Lecture 1: Intro to Biostatistics. Smoking: hazardous? FEV (l) Smoke
 BIOL 51A - Biostatistics 1 1 Lecture 1: Intro to Biostatistics Smoking: hazardous? FEV (l) 1 2 3 4 5 No Yes Smoke BIOL 51A - Biostatistics 1 2 Box Plot a.k.a box-and-whisker diagram or candlestick chart
BIOL 51A - Biostatistics 1 1 Lecture 1: Intro to Biostatistics Smoking: hazardous? FEV (l) 1 2 3 4 5 No Yes Smoke BIOL 51A - Biostatistics 1 2 Box Plot a.k.a box-and-whisker diagram or candlestick chart
Tutorial 2: Power and Sample Size for the Paired Sample t-test
 Tutorial 2: Power and Sample Size for the Paired Sample t-test Preface Power is the probability that a study will reject the null hypothesis. The estimated probability is a function of sample size, variability,
Tutorial 2: Power and Sample Size for the Paired Sample t-test Preface Power is the probability that a study will reject the null hypothesis. The estimated probability is a function of sample size, variability,
AP Curriculum Framework with Learning Objectives
 Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Big Ideas Big Idea 1: The process of evolution drives the diversity and unity of life. AP Curriculum Framework with Learning Objectives Understanding 1.A: Change in the genetic makeup of a population over
Written Exam 15 December Course name: Introduction to Systems Biology Course no
 Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
Technical University of Denmark Written Exam 15 December 2008 Course name: Introduction to Systems Biology Course no. 27041 Aids allowed: Open book exam Provide your answers and calculations on separate
AP Biology Curriculum Framework
 AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
AP Biology Curriculum Framework This chart correlates the College Board s Advanced Placement Biology Curriculum Framework to the corresponding chapters and Key Concept numbers in Campbell BIOLOGY IN FOCUS,
Confidence Estimation Methods for Neural Networks: A Practical Comparison
 , 6-8 000, Confidence Estimation Methods for : A Practical Comparison G. Papadopoulos, P.J. Edwards, A.F. Murray Department of Electronics and Electrical Engineering, University of Edinburgh Abstract.
, 6-8 000, Confidence Estimation Methods for : A Practical Comparison G. Papadopoulos, P.J. Edwards, A.F. Murray Department of Electronics and Electrical Engineering, University of Edinburgh Abstract.
Tutorial 5: Power and Sample Size for One-way Analysis of Variance (ANOVA) with Equal Variances Across Groups. Acknowledgements:
 Tutorial 5: Power and Sample Size for One-way Analysis of Variance (ANOVA) with Equal Variances Across Groups Anna E. Barón, Keith E. Muller, Sarah M. Kreidler, and Deborah H. Glueck Acknowledgements:
Tutorial 5: Power and Sample Size for One-way Analysis of Variance (ANOVA) with Equal Variances Across Groups Anna E. Barón, Keith E. Muller, Sarah M. Kreidler, and Deborah H. Glueck Acknowledgements:
Summary Report: MAA Program Study Group on Computing and Computational Science
 Summary Report: MAA Program Study Group on Computing and Computational Science Introduction and Themes Henry M. Walker, Grinnell College (Chair) Daniel Kaplan, Macalester College Douglas Baldwin, SUNY
Summary Report: MAA Program Study Group on Computing and Computational Science Introduction and Themes Henry M. Walker, Grinnell College (Chair) Daniel Kaplan, Macalester College Douglas Baldwin, SUNY
Valley Central School District 944 State Route 17K Montgomery, NY Telephone Number: (845) ext Fax Number: (845)
 Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
Valley Central School District 944 State Route 17K Montgomery, NY 12549 Telephone Number: (845)457-2400 ext. 18121 Fax Number: (845)457-4254 Advance Placement Biology Presented to the Board of Education
OCR Biology Checklist
 Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Qualitative Analysis of Tumor-Immune ODE System
 of Tumor-Immune ODE System LG de Pillis and AE Radunskaya August 15, 2002 This work was supported in part by a grant from the WM Keck Foundation 0-0 QUALITATIVE ANALYSIS Overview 1 Simplified System of
of Tumor-Immune ODE System LG de Pillis and AE Radunskaya August 15, 2002 This work was supported in part by a grant from the WM Keck Foundation 0-0 QUALITATIVE ANALYSIS Overview 1 Simplified System of
OCR Biology Checklist
 Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Topic 1. Cell level systems Video: Eukaryotic and prokaryotic cells Compare the structure of animal and plant cells. Label typical and atypical prokaryotic cells. Compare prokaryotic and eukaryotic cells.
Solving the Yang-Baxter Matrix Equation
 The University of Southern Mississippi The Aquila Digital Community Honors Theses Honors College 5-7 Solving the Yang-Baxter Matrix Equation Mallory O Jennings Follow this and additional works at: http://aquilausmedu/honors_theses
The University of Southern Mississippi The Aquila Digital Community Honors Theses Honors College 5-7 Solving the Yang-Baxter Matrix Equation Mallory O Jennings Follow this and additional works at: http://aquilausmedu/honors_theses
Safety Envelope for Load Tolerance and Its Application to Fatigue Reliability Design
 Safety Envelope for Load Tolerance and Its Application to Fatigue Reliability Design Haoyu Wang * and Nam H. Kim University of Florida, Gainesville, FL 32611 Yoon-Jun Kim Caterpillar Inc., Peoria, IL 61656
Safety Envelope for Load Tolerance and Its Application to Fatigue Reliability Design Haoyu Wang * and Nam H. Kim University of Florida, Gainesville, FL 32611 Yoon-Jun Kim Caterpillar Inc., Peoria, IL 61656
Tutorial 1: Power and Sample Size for the One-sample t-test. Acknowledgements:
 Tutorial 1: Power and Sample Size for the One-sample t-test Anna E. Barón, Keith E. Muller, Sarah M. Kreidler, and Deborah H. Glueck Acknowledgements: The project was supported in large part by the National
Tutorial 1: Power and Sample Size for the One-sample t-test Anna E. Barón, Keith E. Muller, Sarah M. Kreidler, and Deborah H. Glueck Acknowledgements: The project was supported in large part by the National
Row and Column Distributions of Letter Matrices
 College of William and Mary W&M ScholarWorks Undergraduate Honors Theses Theses, Dissertations, & Master Projects 5-2016 Row and Column Distributions of Letter Matrices Xiaonan Hu College of William and
College of William and Mary W&M ScholarWorks Undergraduate Honors Theses Theses, Dissertations, & Master Projects 5-2016 Row and Column Distributions of Letter Matrices Xiaonan Hu College of William and
MULTIPLE CHOICE QUESTIONS DECISION SCIENCE
 MULTIPLE CHOICE QUESTIONS DECISION SCIENCE 1. Decision Science approach is a. Multi-disciplinary b. Scientific c. Intuitive 2. For analyzing a problem, decision-makers should study a. Its qualitative aspects
MULTIPLE CHOICE QUESTIONS DECISION SCIENCE 1. Decision Science approach is a. Multi-disciplinary b. Scientific c. Intuitive 2. For analyzing a problem, decision-makers should study a. Its qualitative aspects
Magnetic removal of electron contamination in radiotherapy x-ray beams
 University of Wollongong Research Online University of Wollongong Thesis Collection 1954-2016 University of Wollongong Thesis Collections 2006 Magnetic removal of electron contamination in radiotherapy
University of Wollongong Research Online University of Wollongong Thesis Collection 1954-2016 University of Wollongong Thesis Collections 2006 Magnetic removal of electron contamination in radiotherapy
IDENTIFICATION AND ANALYSIS OF TIME-VARYING MODAL PARAMETERS
 IDENTIFICATION AND ANALYSIS OF TIME-VARYING MODAL PARAMETERS By STEPHEN L. SORLEY A THESIS PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE
IDENTIFICATION AND ANALYSIS OF TIME-VARYING MODAL PARAMETERS By STEPHEN L. SORLEY A THESIS PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE
Qualitative Analysis of Tumor-Immune ODE System
 of Tumor-Immune ODE System L.G. de Pillis and A.E. Radunskaya August 15, 2002 This work was supported in part by a grant from the W.M. Keck Foundation 0-0 QUALITATIVE ANALYSIS Overview 1. Simplified System
of Tumor-Immune ODE System L.G. de Pillis and A.E. Radunskaya August 15, 2002 This work was supported in part by a grant from the W.M. Keck Foundation 0-0 QUALITATIVE ANALYSIS Overview 1. Simplified System
Analysis and Simulation of Biological Systems
 Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
Analysis and Simulation of Biological Systems Dr. Carlo Cosentino School of Computer and Biomedical Engineering Department of Experimental and Clinical Medicine Università degli Studi Magna Graecia Catanzaro,
Follow links Class Use and other Permissions. For more information, send to:
 COPYRIGHT NOTICE: Stephen L. Campbell & Richard Haberman: Introduction to Differential Equations with Dynamical Systems is published by Princeton University Press and copyrighted, 2008, by Princeton University
COPYRIGHT NOTICE: Stephen L. Campbell & Richard Haberman: Introduction to Differential Equations with Dynamical Systems is published by Princeton University Press and copyrighted, 2008, by Princeton University
An Approach to Constructing Good Two-level Orthogonal Factorial Designs with Large Run Sizes
 An Approach to Constructing Good Two-level Orthogonal Factorial Designs with Large Run Sizes by Chenlu Shi B.Sc. (Hons.), St. Francis Xavier University, 013 Project Submitted in Partial Fulfillment of
An Approach to Constructing Good Two-level Orthogonal Factorial Designs with Large Run Sizes by Chenlu Shi B.Sc. (Hons.), St. Francis Xavier University, 013 Project Submitted in Partial Fulfillment of
Lifetime prediction and confidence bounds in accelerated degradation testing for lognormal response distributions with an Arrhenius rate relationship
 Scholars' Mine Doctoral Dissertations Student Research & Creative Works Spring 01 Lifetime prediction and confidence bounds in accelerated degradation testing for lognormal response distributions with
Scholars' Mine Doctoral Dissertations Student Research & Creative Works Spring 01 Lifetime prediction and confidence bounds in accelerated degradation testing for lognormal response distributions with
Basic modeling approaches for biological systems. Mahesh Bule
 Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Basic modeling approaches for biological systems Mahesh Bule The hierarchy of life from atoms to living organisms Modeling biological processes often requires accounting for action and feedback involving
Cell-Cell Communication in Development
 Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Biology 4361 - Developmental Biology Cell-Cell Communication in Development October 2, 2007 Cell-Cell Communication - Topics Induction and competence Paracrine factors inducer molecules Signal transduction
Types of biological networks. I. Intra-cellurar networks
 Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Types of biological networks I. Intra-cellurar networks 1 Some intra-cellular networks: 1. Metabolic networks 2. Transcriptional regulation networks 3. Cell signalling networks 4. Protein-protein interaction
Stoichiometry Of Tumor Dynamics: Models And Analysis
 Stoichiometry Of Tumor Dynamics: Models And Analysis Yang Kuang Department of Mathematics and Statistics Arizona State University Supported by NSF grant DMS-0077790 Y. Kuang, J. Nagy and J. Elser: Disc.
Stoichiometry Of Tumor Dynamics: Models And Analysis Yang Kuang Department of Mathematics and Statistics Arizona State University Supported by NSF grant DMS-0077790 Y. Kuang, J. Nagy and J. Elser: Disc.
Honors Biology Reading Guide Chapter 11
 Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Honors Biology Reading Guide Chapter 11 v Promoter a specific nucleotide sequence in DNA located near the start of a gene that is the binding site for RNA polymerase and the place where transcription begins
Scientific Computing with Case Studies SIAM Press, Lecture Notes for Unit VII Sparse Matrix
 Scientific Computing with Case Studies SIAM Press, 2009 http://www.cs.umd.edu/users/oleary/sccswebpage Lecture Notes for Unit VII Sparse Matrix Computations Part 1: Direct Methods Dianne P. O Leary c 2008
Scientific Computing with Case Studies SIAM Press, 2009 http://www.cs.umd.edu/users/oleary/sccswebpage Lecture Notes for Unit VII Sparse Matrix Computations Part 1: Direct Methods Dianne P. O Leary c 2008
UNIT 4 RANK CORRELATION (Rho AND KENDALL RANK CORRELATION
 UNIT 4 RANK CORRELATION (Rho AND KENDALL RANK CORRELATION Structure 4.0 Introduction 4.1 Objectives 4. Rank-Order s 4..1 Rank-order data 4.. Assumptions Underlying Pearson s r are Not Satisfied 4.3 Spearman
UNIT 4 RANK CORRELATION (Rho AND KENDALL RANK CORRELATION Structure 4.0 Introduction 4.1 Objectives 4. Rank-Order s 4..1 Rank-order data 4.. Assumptions Underlying Pearson s r are Not Satisfied 4.3 Spearman
Final Project Descriptions Introduction to Mathematical Biology Professor: Paul J. Atzberger. Project I: Predator-Prey Equations
 Final Project Descriptions Introduction to Mathematical Biology Professor: Paul J. Atzberger Project I: Predator-Prey Equations The Lotka-Volterra Predator-Prey Model is given by: du dv = αu βuv = ρβuv
Final Project Descriptions Introduction to Mathematical Biology Professor: Paul J. Atzberger Project I: Predator-Prey Equations The Lotka-Volterra Predator-Prey Model is given by: du dv = αu βuv = ρβuv
Multi-state Models: An Overview
 Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
 1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
1 of 13 8/11/2014 10:32 AM Units: Teacher: APBiology, CORE Course: APBiology Year: 2012-13 Chemistry of Life Chapters 1-4 Big Idea 1, 2 & 4 Change in the genetic population over time is feedback mechanisms
ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 7
 ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 7 KINETICS OF ENZYME CATALYSED REACTIONS (CONTD.) So in the last lecture we
ENZYME SCIENCE AND ENGINEERING PROF. SUBHASH CHAND DEPARTMENT OF BIOCHEMICAL ENGINEERING AND BIOTECHNOLOGY IIT DELHI LECTURE 7 KINETICS OF ENZYME CATALYSED REACTIONS (CONTD.) So in the last lecture we
Reactants and Products
 Biological Chemical Reactions Chemical reactions allow living things to grow, develop, reproduce, and adapt. Real-World Reading Link When you lie down for the night, you might think that your body is completely
Biological Chemical Reactions Chemical reactions allow living things to grow, develop, reproduce, and adapt. Real-World Reading Link When you lie down for the night, you might think that your body is completely
Gene Network Science Diagrammatic Cell Language and Visual Cell
 Gene Network Science Diagrammatic Cell Language and Visual Cell Mr. Tan Chee Meng Scientific Programmer, System Biology Group, Bioinformatics Institute Overview Introduction Why? Challenges Diagrammatic
Gene Network Science Diagrammatic Cell Language and Visual Cell Mr. Tan Chee Meng Scientific Programmer, System Biology Group, Bioinformatics Institute Overview Introduction Why? Challenges Diagrammatic
Epidermal Wound Healing: A Mathematical Model
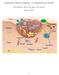 Epidermal Wound Healing: A Mathematical Model Abi Martinez, Sirena Van Epp, Jesse Kreger April 16, 2014 1 Contents 1 Wound Healing History 3 2 Biology of Epidermal Wound Healing 4 3 Mathematical Model
Epidermal Wound Healing: A Mathematical Model Abi Martinez, Sirena Van Epp, Jesse Kreger April 16, 2014 1 Contents 1 Wound Healing History 3 2 Biology of Epidermal Wound Healing 4 3 Mathematical Model
A A A A B B1
 LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LEARNING OBJECTIVES FOR EACH BIG IDEA WITH ASSOCIATED SCIENCE PRACTICES AND ESSENTIAL KNOWLEDGE Learning Objectives will be the target for AP Biology exam questions Learning Objectives Sci Prac Es Knowl
LAKELAND COMMUNITY COLLEGE COURSE OUTLINE FORM
 LAKELAND COMMUNITY COLLEGE COURSE OUTLINE FORM ORIGINATION DATE: 8/2/99 APPROVAL DATE: 3/22/12 LAST MODIFICATION DATE: 3/28/12 EFFECTIVE TERM/YEAR: FALL/ 12 COURSE ID: COURSE TITLE: MATH2800 Linear Algebra
LAKELAND COMMUNITY COLLEGE COURSE OUTLINE FORM ORIGINATION DATE: 8/2/99 APPROVAL DATE: 3/22/12 LAST MODIFICATION DATE: 3/28/12 EFFECTIVE TERM/YEAR: FALL/ 12 COURSE ID: COURSE TITLE: MATH2800 Linear Algebra
8 The SVD Applied to Signal and Image Deblurring
 8 The SVD Applied to Signal and Image Deblurring We will discuss the restoration of one-dimensional signals and two-dimensional gray-scale images that have been contaminated by blur and noise. After an
8 The SVD Applied to Signal and Image Deblurring We will discuss the restoration of one-dimensional signals and two-dimensional gray-scale images that have been contaminated by blur and noise. After an
Structures and Functions of Living Organisms (LS1)
 EALR 4: Big Idea: Core Content: Life Science Structures and Functions of Living Organisms (LS1) Processes Within Cells In prior grades students learned that all living systems are composed of cells which
EALR 4: Big Idea: Core Content: Life Science Structures and Functions of Living Organisms (LS1) Processes Within Cells In prior grades students learned that all living systems are composed of cells which
ESTIMATING STATISTICAL CHARACTERISTICS UNDER INTERVAL UNCERTAINTY AND CONSTRAINTS: MEAN, VARIANCE, COVARIANCE, AND CORRELATION ALI JALAL-KAMALI
 ESTIMATING STATISTICAL CHARACTERISTICS UNDER INTERVAL UNCERTAINTY AND CONSTRAINTS: MEAN, VARIANCE, COVARIANCE, AND CORRELATION ALI JALAL-KAMALI Department of Computer Science APPROVED: Vladik Kreinovich,
ESTIMATING STATISTICAL CHARACTERISTICS UNDER INTERVAL UNCERTAINTY AND CONSTRAINTS: MEAN, VARIANCE, COVARIANCE, AND CORRELATION ALI JALAL-KAMALI Department of Computer Science APPROVED: Vladik Kreinovich,
Using Optimal Control Theory to Optimize the Use of Oxygen Therapy in Chronic Wound Healing
 Western Kentucky University TopSCHOLAR Masters Theses & Specialist Projects Graduate School 5-2013 Using Optimal Control Theory to Optimize the Use of Oxygen Therapy in Chronic Wound Healing Donna Lynn
Western Kentucky University TopSCHOLAR Masters Theses & Specialist Projects Graduate School 5-2013 Using Optimal Control Theory to Optimize the Use of Oxygen Therapy in Chronic Wound Healing Donna Lynn
BMM 305 Biomaterials. Biological Recognition. Dr. Ersin Emre Oren
 BMM 305 Biomaterials Dr. Ersin Emre Oren Department of Biomedical Engineering Department of Materials Science & Nanotechnology Engineering TOBB University of Economics and Technology Ankara - TURKEY Bionanodesign
BMM 305 Biomaterials Dr. Ersin Emre Oren Department of Biomedical Engineering Department of Materials Science & Nanotechnology Engineering TOBB University of Economics and Technology Ankara - TURKEY Bionanodesign
Workbook Exercises for Statistical Problem Solving in Geography
 Workbook Exercises for Statistical Problem Solving in Geography Arthur J. Lembo, Jr. This workbook is for use with the popular textbook Introduction to Statistical Problem Solving in Geography, and includes
Workbook Exercises for Statistical Problem Solving in Geography Arthur J. Lembo, Jr. This workbook is for use with the popular textbook Introduction to Statistical Problem Solving in Geography, and includes
