ON THE ANALYSIS OF TUBERCULOSIS STUDIES WITH INTERMITTENT MISSING SPUTUM DATA
|
|
|
- Allyson Short
- 5 years ago
- Views:
Transcription
1 Submitted to the Annals of Applied Statistics arxiv: arxiv: ON THE ANALYSIS OF TUBERCULOSIS STUDIES WITH INTERMITTENT MISSING SPUTUM DATA By Daniel Scharfstein, Andrea Rotnitzky, Maria Abraham Aidan McDermott Richard Chaisson and Lawrence Geiter Johns Hopkins University, Universidad Torcuato Di Tella, Statistics Collaborative, Otsuka Novel Products In randomized studies evaluating treatments for tuberculosis (TB), individuals are scheduled to be routinely evaluated for the presence of TB using sputum cultures. One important endpoint in such studies is the time of culture conversion, the first visit at which a patient s sputum culture is negative and remains negative. This paper addresses how to draw inference about treatment effects when sputum cultures are intermittently missing on some patients. We discuss inference under a novel benchmark assumption and under a class of assumptions indexed by a treatment-specific sensitivity parameter that quantify departures from the benchmark assumption. We motivate and illustrate our approach using data from a randomized trial comparing the effectiveness of two treatments for adult TB patients in Brazil. 1. Introduction. In the design of randomized studies evaluating competing treatments for patients with tuberculosis (TB), it is common to culture sputum for the presence of TB at regularly scheduled clinic visits over a specified time horizon. A primary goal in such studies is to estimate the treatment-specific distribution of the visit of first culture conversion [European Medicines Agency, Committee for Medicinal Products for Human Use (2010)], which we refer to as the time of culture conversion. Culture conversion is said to have occurred for a patient at a given visit, if the sputum cultures for that visit and all subsequent visits are negative. A key complication in the analysis is that culture results are missing at some visits, because the culture was contaminated, the patient could not produce sputum, or the patient did not show up. Culture conversion status at a given visit is unknown when from that visit onward at least one culture result is missing and all recorded culture results are negative. For a given patient, the set of visits with unknown culture conversion status is empty, or it consists of either a single visit or a set of consecutive visits. If the set is not empty, the time of culture conversion will be known to lie in an interval; however, Keywords and phrases: Culture Conversion, Curse of Dimensionality, Exponential Tilting, Reverse-Time Hazard, Senstiivity Analysis 1
2 2 D. SCHARFSTEIN ET AL. the time may not be interval censored in the classical sense. This is because certain data configurations may imply that the time of culture conversion cannot occur at certain visit times within the interval. To distinguish this data structure from classic interval censoring, we refer to the set of feasible times that are compatible with an individual s data as the coarsening set. The coarsening set can include a single point or a set of points. The treatment-specific distribution of time of culture conversion is not identified without untestable assumptions about the distribution of culture conversion status within the coarsening sets. There are countless ways of imposing such assumptions. The worst-case and best-case assumptions, leading to bounds on the treatment-specific distribution of time of culture conversion, are that the missing culture associated with the latest visit time at which culture conversion status is unknown is positive and the missing cultures associated with all visit times at which culture conversion status is unknown are negative, respectively. In considering alternative assumptions it is natural to condition on as much of the relevant data as possible. In addition to conditioning on observed culture results, it is natural to condition on auxiliary factors that are associated with the unknown results inside the coarsening set. Most TB studies collect a key auxiliary time-varying variable. Sputum specimens are also evaluated by smear, and the results of the smear may be available when a culture is contaminated. Sputum smear is a less reliable assessment of clinical tuberculosis than sputum culture. It relies on the visualization of the bacteria through a microscope after staining of sputum with dyes that allows the microscopist to see so-called acid-fast TB bacteria. Sensitivity of the sputum smear is about 0.5, but it can vary by staining method used and clinical population. In contrast, sputum culture involves breaking down the sputum (which is very vicous), decontaminating the specimen to kill bacteria other than the mycobacteria, and inoculating it on culture media where the bacilli can grow. Sensitivity of the sputum culture is about (American Thoracic Society, 2000). Roughly 65% of those with a positive culture are expected to have a positive smear, and nearly 100% of those with a positive smear are expected to have a positive culture (American Thoracic Society, 2000). Another important auxiliary variable is baseline cavitation status. Patients with pulmonary tuberculosis who have cavities seen on a chest radiograph (cavitary tuberculosis) are more likely to have positive sputum smears, as they harbor larger numbers of tubercle bacilli than patients without cavities. It generally takes longer for patients with cavitary tuberculosis to convert their smears and cultures to negative during treatment, as they
3 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 3 have a larger bacterial load and the therapy must kill more organisms. This article is motivated by data from a randomized TB study previously analyzed by Conde et al. (2009). 1 This phase II, double-blind, randomized trial compared moxifloxacin vs. ethambutol in adults with smear-positive tuberculosis at baseline in a hospital in Rio de Janeiro, Brazil. All 170 patients randomized (85 to each treatment arm) into the study were treated with a background regimen of isoniazid, rifampicin and pyrazinamide. Patients with negative, contaminated or drug-resistant Mycobacterium tuberculosis at baseline were excluded from the analysis, resulting in an analysis sample of 74 and 72 patients in the moxifloxacin and ethambutol groups, respectively. Treatment was scheduled to be given five days per week and was to be directly observed by study personnel. Smear and culture testing of sputum specimens (spontaneous or induced) was scheduled on specimens collected at baseline and every week for 8 weeks. In this study, 55.4% and 62.5% of patients in the moxifloxacin and ethambutol arms, respectively, had complete culture data through week 8. Time of culture conversion could be determined for 64.9% and 72.2% of patients in these arms; the remaining patients had their time of culture conversion coarsened. In this article, we develop a method that estimates the treatment-specific distribution of time of culture conversion under a class of assumptions on the distribution of the coarsened time of culture conversion that conditions on all of the relevant available data, including sputum cultures, sputum smears and baseline data. Each assumption in the class is indexed by a treatment-specific sensitivity-analysis parameter which quantifies the magnitude of discrepancy from a specific benchmark assumption. In Section 2, we provide a preview of our proposed method. In Section 3, we introduce the data structure and notation. Section 4 discusses our modeling assumptions and the approach to inference. Section 5 presents an analysis of data from the Conde et al. study. The article concludes with a short discussion. 2. Preview. Our approach starts by imposing non-testable assumptions that identify the conditional distribution of time to culture conversion given the data, for every data configuration for which time of culture conversion is unknown. The crucial methodological challenge then is to make sensible identifying assumptions. Because time of culture conversion is determined by the results of sputum cultures, these assumptions ultimately identify the visit-specific probabilities of the last positive sputum culture 1 The dataset provided to us differs slightly from that of Conde et al. (2009). There are small differences in the number of observed cultures and the number of observed negative cultures at each week. All analyses reported in this paper are based on the data provided to us.
4 4 D. SCHARFSTEIN ET AL. Table 1 Example of patient data. - denotes negative culture, + denotes positive culture,? denotes unknown, green? denotes a culture result that is irrelevant for determining culture conversion, red? denotes a culture result that is relevant for determining culture conversion. Visit Line Coarsening Set 1 Culture Results? +? -? Conversion? N N??? Y Y Y {3, 4, 6} 3 Culture Results? +? Conversion? N N N N N Y Y Y {6} 5 Culture Results? +? Conversion? N N? Y Y Y Y Y {3, 4} 7 Culture Results? Conversion? N N N Y Y Y Y Y {4} 9 Culture Results? Conversion? N N Y Y Y Y Y Y {3} given the data. Lack of culture conversion at a given visit can be determined without full knowledge of all subsequent results if the culture at that visit is positive or at least one subsequent culture is observed to be positive. It turns out that with proper book-keeping, we can achieve identifiability by imposing assumptions that suffice to identify the distribution of time of culture conversion but do not fully identify the joint distribution of results across visits. These conditions identify the reverse-time conditional hazards of time of culture conversion. This section illustrates these issues by means of an example. The sputum culture data collected on one patient in the study illustrate the coarsening structure of culture conversion status and the time of culture conversion. The sputum culture data for this patient, we call Mary, are displayed in the first line of Table 1; the associated culture conversion statuses are displayed in line 2. Missing values are indicated by?. Mary has negative cultures at visits 4, 6, 7 and 8 and a positive culture at visit 2. She is a culture converter at visits 6, 7 and 8 (labeled Y in the second line), known not be a culture converter at visits 1 and 2 (labeled N in the second line) and has unknown culture conversion status at visits 3, 4 and 5 (labeled? in the second line). First, even though her culture status at visit 1 (labeled with a green? in the first line) is missing, it is irrelevant for determining whether she is a culture converter at that visit, because she has a positive culture at visit 2 and therefore cannot be a culture converter at visit 1. Second, missingness of cultures at visits 3 and 5 (labeled with a red? in the first line) affects our ability to determine her culture conversion status at visits
5 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 5 3, 4 and 5 because all cultures after visit 5 are negative and the culture at visit 4 is also negative. So even though Mary has a negative culture at visit 4, her culture conversion status is not known at that visit. Third, the visits at which culture conversion status is unknown are consecutive: the earliest and latest visits at are 3 and 5, respectively. Finally, the coarsening set for time of culture conversion comprises visits 3, 4 and 6. This follows because if the culture at visit 5 were positive, then the time of culture conversion would be visit 6, and if the culture at visit 5 were negative, the time of culture conversion would be either visit 3 or visit 4, depending on the result at visit 3. To illustrate our approach for identifying the conditional distribution of time of culture conversion given the data, consider subset A, the subset of patients with the same observed data as Mary. Specifically, we must model the probability that the cultures at visits 3 and 5 are both negative (in which case the time of culture conversion is visit 3), the probability that the culture at visit 3 is positive and the culture at visit 5 is negative (in which case the time of culture conversion is visit 4) and the probability that the culture at visit 5 is positive (in which case the time of culture conversion is visit 6). In modeling these probabilities, it is natural to consider two chronological factorizations. In forward time, we would need to model the probability of a negative culture at visit 3, the conditional probability of a negative culture at visit 5 given negative culture at visit 3, and the conditional probability of a negative culture at visit 5 given a positive culture at visit 3. In reverse time, we would need to model (i) the probability of a positive culture at visit 5 and (ii) the conditional probability of a positive culture at visit 3 given a negative culture at visit 5. We utilize this latter factorization as it requires fewer modeling assumptions. We first turn to the task of imposing assumptions that identify (i). Our identifying assumption specifies that (i) is the same as the probability of a positive culture at visit 5 among patients with the same data as in subset A with the exception that they have an observed culture at visit 5. The observed culture results for these patients are depicted in lines 3 and 5. These patients have observed culture conversion status as depicted in lines 4 and 6. The probability (i) is then assumed to be equal to the ratio of the proportion of patients with observed results as in line 3 to the sum of the proportions of patients with observed results as in lines 3 and 5. Next, we turn to the task of imposing assumptions that identify (ii). Our identifying assumption specifies that (ii) is the same as the probability of a positive result at visit 3 among patients with the same pattern of observed cultures as that in line 5, with the exception that they have an observed
6 6 D. SCHARFSTEIN ET AL. culture at visit 3. The results of these patients are depicted in lines 7 and 9 with associated culture conversion status in lines 8 and 10. The probability (ii) is then assumed to be equal to the ratio of the proportion of patients with observed cultures as in line 7 to the sum of the proportions of patients with observed cultures as lines 7 and 9. In datasets of typical size, we will not be able to obtain estimates of the proportion of patients who have specific patterns of observed data due to the curse of dimensionality. As a result, our inference will require dimension reduction assumptions. We will use fully parametric models for the treatment-specific distributions of the data. These models are described in Section 3.4. In Section 3.3, we evaluate the sensitivity of our results to our identifying assumptions by conducting inference under a class of exponential tilt deviations from the assumed conditional probabilities for the unobserved culture conversion status. 3. Formalization of the problem. Since we focus on inference about the time of culture conversion, separately for each treatment arm, we consider, until Section 4, only data from one arm and suppress notational dependence on treatment assignment Data Structure and Notation. Let X denote baseline cavitation status (1 for cavitation, 0 otherwise), C k denote the indicator that the culture is negative at visit k (1 for negative, 0 for positive) and S k denote the indicator that the smear is negative at visit k (1 for negative, 0 for positive). Let Mk c and M k s be the indicators that C k and S k are missing, respectively (1 for missing, 0 for observed). The data recorded on an individual at visit k are realizations of the random vector O k = (Mk c, Cobs k, M k s, Sobs k ) where Ck obs = C k if C k is observed and Ck obs is empty otherwise and Sk obs is defined likewise. Let K denote the number of scheduled post-baseline visits. For any collection of random vectors {W k } 1 k K, we use the notation W k = (W 1,..., W k ). With this notation, the data recorded on an individual throughout the entire study are realizations of the random vector O = (X, O K ). Define time of culture conversion T to be the earliest visit such that sputum cultures are negative from that visit onward if such a visit exists and T = K + 1 otherwise. As illustrated in Section 2, when the data available on each patient is the vector O defined earlier, T belongs to a set with a single visit time (in which case T is determined from O) or a set with multiple visit times, not necessarily consecutive. We denote the coarsening set where
7 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 7 T is known to lie by T. Given O, T is determined (i.e., the set T has one element), unless either (i) the culture at visit K is missing, i.e. MK c = 1, or (ii) there exists a visit k < K with a missing culture, i.e. Mk c = 1, such that all subsequent visits have sputum cultures that are either negative or missing, i.e. Mj c = 1 or Cobs j = 1 for j > k. When either (i) or (ii) occurs, then T will have multiple visit times. We denote the lowest visit number in the set by L, which is the earliest visit k where the sputum cultures are either missing or negative at and subsequent to visit k. We denote the largest visit number in the set by R + 1, where R = K if the culture at visit K is missing (i.e. MK c = 1) and R = k (k < K) if at visit k the sputum culture is missing and at all subsequent visits the sputum cultures are recorded and negative. The other times in the coarsening set include all visit numbers k such that L < k < R + 1 and Mk 1 c = 1. Formally, our inferential goal is to estimate the distribution of time of culture conversion, i.e., P [T = k] for k = 1,..., K. based on n i.i.d. realizations of the vector O. We use the subscript i to denote data for the ith individual. In the next section we formally describe the identifying assumptions on which our benchmark analysis relies. These assumptions were illustrated in Section 2. The assumptions are identifying in the sense that once they are imposed, we are able to express P [T = k] for k = 1,..., K as a function of the distribution of the observed data O and consequently, we can hope, to estimate P [T = k] consistently. Subsequently, we propose models for departures from the benchmark assumptions that form the basis of our proposed sensitivity analysis. Specifically, our sensitivity analysis consists of repeating estimation of P [T = k] under various plausible departures from the benchmark assumptions. Both our benchmark analysis and the models for our sensitivity analysis rely on assumptions that identify the conditional distribution of T given the observed data O. The marginal distribution of T is then obtained as the mixture of the, now identified, conditional distribution of T given O mixed over the distribution of the observed data O Benchmark Identifying Assumptions. To help guide our choice of benchmark identifying assumptions, we first note that since P [T = k O] = 0 if k / T and P [T = k O] = 1 if k T and T = 1, we only need assumptions that suffice to identify P [T = k O] when k T and T > 1. Thus, we proceed in reverse order through the set T by postulating assumptions that identify, first P [T = R + 1 O], then in reverse sequential order P [T = k T
8 8 D. SCHARFSTEIN ET AL. k, O] where k T, L < k < R + 1. This iterative procedure results in assumptions that identify P [T = k O] for all k T. This follows because P [R + 1 O] is identified and for k T, k < R + 1, P [T = k O] equals P [T R + 1 O] k<s<r+1 k T P [T s T s, O] P [T = k T k, O] We need to focus only on the case where T > 1. To guide our choice of benchmark assumptions for identifying P [T = r + 1 O = o], we first note that given O = o the event T = r + 1 occurs if and only if the culture at visit r is positive, i.e. if C r = 0. Our benchmark assumption equates the unidentified probability P [T = r + 1 O = o] with the identified probability that C r = 0 in the subset of patients for which O = o (r) where o (r) agrees with O in all its components except that the culture at visit r is observed. Now, we write the event O = o as the event (3.1) Mr c = 1, M c r 1 = m c r 1, C obs r 1 = c obs r 1, V = v, Mj c = 0, C j = 1 for r+1 j K we define the event O = o (r) as the event M c r = 0, M c r 1 = m c r 1, C obs r 1 = c obs r 1, V = v,, M c j = 0, C j = 1 for r+1 j K and we postulate that (3.2) P [T = r + 1 O = o] = P [T = r + 1 O = o (r) ] Next, we consider assumptions that identify P [T = k T k, O = o] for k T, l < k < r + 1. Once again, given (T k, O = o), the event T = k occurs if and only if the culture result at visit k 1 is positive, i.e. C k 1 = 0. Because k T, l < k < r + 1, we know that C k 1 is missing, i.e., Mk 1 c = 1. Our benchmark assumption in this case equates the probability P [T = k T k, O = o] with the identified probability that T = k (which equates to the event C k 1 = 0) in the subset of patients for which O = o (k 1), where o (k 1) differs from the subset of patients with (T k, O = o) only in that a sputum culture is observed at visit k 1 and the event T k is observed to occur (i.e., Mj c = 0 and C j = 1 for all k j K). Formally, with the event O = o defined as in (3.1), we define the event O = o (k 1) as the event M c k 1 = 0, M c k 2 = m c k 2, C obs k 2 = c obs k 2,, V = v, M c j = 1, C j = 1 for k j K
9 and assume TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 9 (3.3) P [T = k T k, O = o] = P [T = k O = o (k 1) ] Patients in the subset defined by (T k, O = o) and subjects in the subset O = o (k 1) have the same baseline factors and the same recorded history of the auxiliary smear sputums throughout the study, as well as the same recorded history of sputum cultures up to visit k 2. Finally, for a realization O = o where T > 1, P [T = l T l, O = o] = Sensitivity Analysis. The benchmark assumptions (3.2) and (3.3) are untestable. For realizations O = o with T > 1, the following exponential tilt model [Cox and Barndorff-Nielsen (1994)] expresses departures from our benchmark assumptions: (3.4) P [T = r + 1 O = o] = P [T = r + 1 O = o(r) ] exp(α) h r+1 (o (r) ; α) and for k T, l < k < r + 1, (3.5) P [T = k T k, O = o] = P [T = k O = o(k 1) ] exp(α) h k (o (k 1) ; α) where α is fixed and given and h k (o (k 1) ; α) are normalizing constants equal to E[exp{αI(T = k)} O = o (k 1) ] for k T, l < k r + 1. Under this exponential tilt model, odds(p [T = r + 1 O = o]) odds(p [T = k T k, O = o]) odds(p [T = r + 1 O = o (r) = ]) odds(p [T = k O = o (k 1) = exp(α) ]) Thus, the magnitude of α quantifies the departure from our benchmark assumptions. When α > 0 (< 0), P [T = r + 1 O = o] is greater (less) than P [T = r + 1 O = o (r) ] and P [T = k T k, O = o] is greater (less) than P [T = k O = o (k 1) ]. As α ( ), P [T = r + 1 O = o] and P [T = k T k, O = o] go to one (zero). When α ( ), the worstcase and best-case bounds described in the introduction are attained. α = 0 corresponds to the benchmark assumption. To facilitate sensitivity analysis, our class of models assumes that the departures from the benchmark assumption are not time-specific Modeling. For specified α, estimation of the distribution of time of culture conversion depends on our ability to estimate for each realization
10 10 D. SCHARFSTEIN ET AL. O = o with T > 1, P [T = k O = o (k 1) ] for k T, l < k r + 1. However, in practice these probabilities cannot be estimated non-parametrically. Therefore, we use a parametric model for the law of the observed data O given baseline covariates X. This model induces parametric models for P [T = k O = o (k 1) ], k T, l < k r+1, that ultimately enable estimation of P [T = k O = o] by borrowing information ) across strata O = o (k 1). We first recall that O = (X, O K, where O k = (Mk c, Cobs k, M k s, Sobs k ) are the data available at visit k. We model the law of O by modeling the distribution of O k given O k 1 and X for all k = 1,..., K, where O 0 =. We use separate logistic regression models for 1. the probabililty of M c k = 1 given O k 1 and X, i.e. (3.6) logit{p [M c k = 1 O k 1, X]} = a(k, O k 1, X; γ (a) ) 2. the probability that C obs k = 1 given M c k = 0, O k 1 and X, i.e. (3.7) logit{p [C obs k = 1 M c k = 0, O k 1, X]} = b(k, O k 1, X; γ (b) ) 3. the probability that Mk s = 1 given M k c, Cobs k, O k 1 and X, i.e. (3.8) logit{p [Mk s = 1 Mk, c Ck obs, O k 1, X]} = c(k, Mk, c Ck obs, O k 1, X; γ (c) ) 4. the probability that Sk obs i.e. (3.9) logit{p [Sk obs = 1 given M s k = 0, M c k, Cobs k, O k 1 and X, = 1 Mk s = 0, Mk, c Ck obs, O k 1, X]} = d(k, Mk, c Ck obs, O k 1, X; γ (d) ) where a( ), b( ), c( ) and d( ) are specified functions of their arguments and γ (a), γ (a), γ (a) and γ (a) are unknown parameter vectors. For the distributions in (3.7) and (3.9) we need not consider the cases Mk c = 1 and M k s = 1, respectively, because for such settings the conditional distributions are degenerate (Ck obs is empty when Mk c = 1, and Sobs k is empty when Mk s = 1) Inference. Under models (3.6) - (3.9) we can express for all realizations O = o with T > 1, the conditional probability P [T = k O = o (k 1) ] for k T, l < k r + 1 as a given function of o (k 1) and γ = (γ (a), γ (b), γ (c), γ (d) ) whose expressions, for the special case where the righthand sides of (3.6) - (3.9) only depend on O k 1, are given in the Appendix. We denote this expression as P [T = k O = o (k 1) ; γ]. Consequently, if we additionally assume models (3.4) and (3.5), then we can express P [T = r + 1 O = o] and P [T = k T k, O = o] (k T, l < k r) as
11 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 11 given functions, P [T = r + 1 O = o; γ; α] and P [T = k T k, O = o; γ; α] of o, γ and α. The first step is to estimate γ using maximum likelihood; denote this estimator by γ. This can be done using standard logistic regression software. This step does not rely on specification of the sensitivity analysis parameter α. i=1 P [T i = k; α] For fixed α, we estimate P [T = k] by P [T = k; α] = 1 n where P [T i = k; α] has one of four expressions depending of k and O i. If k / T i then P [T i = k; α] = 0; if T i = 1 and k T i then P [T i = k; α] = 1; if T i > 1 and k = R i + 1 then P [T i = k; α] = P [T = R i + 1 O = O i ; γ; α]; if T i > 1 and k T i, k < R i + 1 then P [T i = k; α] equals P [T R i + 1 O = O i ; γ; α] P [T s T s, O = O i ; γ; α] k T i k<s<r i +1 P [T = k T k, O = O i ; γ; α] To compare the treatment-specific distributions of time to culture conversion, one can estimate a common treatment effect over time. Towards this end, one can utilize the logistic model for discrete survival data proposed by Cox (1972). This model assumes that h z (k) 1 h z (k) = τ k exp(βz) k = 1...., K, z = 0, 1 where z denotes treatment group, h z (k) = P z [T = k T k], and τ 1,..., τ K 0. Here exp(β) is the ratio of the odds of first becoming a culture converter at visit k given culture conversion at or after visit k, comparing moxifloxacin to ethambutol. To estimate the model parameters, one can use equally weighted minimum distance estimation [Newey and McFadden (1994)]. Specifically, for each choice of α z (z = 0, 1), we minimize the following objective function: 1 8 z=0 k=1 { } 2 ĥz (k; α z ) 1 ĥz(k; α z ) τ k exp(βz) with respect τ 1,..., τ 8 0 and β, where ĥz(k; α z ) = P z [T = k T k; α z ]. For each choice of α z (z = 0, 1), this method finds the closest fitting logistic model to the data : {ĥz(k; α z ) : k = 1,..., 8, z = 0, 1}. Even if the model is incorrectly specified, it can still be used to provide a valid test of the null hypothesis of no treatment effect.
12 12 D. SCHARFSTEIN ET AL. To estimate the standard error of our estimator, we propose the use of non-parametric bootstrap. 4. Data Analysis. Figure 1 displays the treatment-specific observed culture results, with rows denoting patients, columns denoting visit number, red indicating a positive culture, green indicating a negative culture and white indicating a missing culture. Figure 2 displays the treatment-specific coarsening sets for time of culture conversion, with red and green indicating the infeasible and feasible points, respectively. The treatment groups were not balanced with respect to cavitation status at baseline; 81.1% and 56.9% have cavitation in the moxifloxacin and ethambutol arms, respectively. It is essential that our analysis adjust for this key confounder. For each treatment group, we estimate the distribution of time of culture conversion by a weighted average of cavitation-specific distribution of time of culture conversion. The weights are taken to be the marginal (i.e., not conditional on treatment arm) proportion of patients with and without cavitation at baseline, respectively. In our data analysis, we considered parsimonious models for the righthand sides of (3.6) - (3.9). Our choice of models was guided by substantive considerations discussed with our scientist collaborators and by data analytic model fitting techniques. Our final model assumed that the right-hand sides of (3.6) - (3.9) depended only on O k 1 = (Mk 1 c, Cobs k 1, M k 1 s, Sobs k 1 ) (i.e., not on the other components of O k 1 ) and the final model for (3.8) further assumed that c(k, Mk c, Cobs k, O k 1, X; γ (c) ) did not depend on Ck obs and Ck 1 obs. The latter assumption was imposed because missingness of a sputum culture at visit k is highly predictive of missingness of smear sputums at visit k and k 1. In the Appendix we show that when the function c(k, Mk c, Cobs k, O k 1, X; γ (c) ) does not depend on Ck obs and Ck 1 obs for all k, then P [T = k O = o (k 1) ] does not depend on c(k, Mk c, Cobs k, O k 1, X; γ (c) ) for any k, thus alleviating the need to further specify the function c(k, Mk c, Cobs k, O k 1, X; γ (c) ). In the remaining models, we borrowed strength
13 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 13 Fig 1. Treatment-specific observed culture results, with rows denoting patients, columns denoting visit number, red indicating a positive culture, green indicating a negative culture and white indicating a missing culture. Ethambutol Moxifloxacin
14 14 D. SCHARFSTEIN ET AL. Fig 2. Treatment-specific coarsening sets, with rows denoting patients, columns denoting visit number, red indicating infeasible points and green indicating feasible points. Ethambutol Moxifloxacin
15 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 15 across treatment groups. Specifically, we assumed a z (k, O k 1, X; γ (a) ) = γ (a) 0,0,k + γ(a) 0,1,k X + γ(a) 1 I(k > 1)M c k 1 + γ (a) 2 I(k > 1)(1 M c k 1)C obs k 1+ γ (a) 3 I(k > 1)M s k 1 + γ (a) 4 I(k > 1)(1 M s k 1)S obs k 1 + γ (a) 5 z b z (k, O k 1, X; γ (b) ) = γ (b) 0,k + γ(b) 1 I(k > 1)M c k 1 + γ (b) 2 I(k > 1)(1 M c k 1)C obs k 1+ γ (b) 3 I(k > 1)M s k 1 + γ (b) 4 I(k > 1)(1 M s k 1)S obs k 1 + γ (b) 5 z + γ(b) 6 X d z (k, M c k, C obs k, O k 1, X; γ (d) ) = γ (d) 0,k + γ(d) 1 M k c + γ (d) 2 (1 M k)c c k obs + γ (d) 3 (1 M k)c c k obs X+ γ (d) 4 I(k > 1)M c k 1 + γ (d) 5 I(k > 1)(1 M c k 1)C obs k 1 + γ (d) 6 I(k > 1)M s k 1+ γ (d) 7 I(k > 1)(1 M s k 1)S obs k 1 + γ (d) 8 z + γ(d) 9 X where the functions are subscripted by treatment z (z = 0 denotes ethambutol, z = 1 denoted moxifloxacin). In Tables 2, 3 and 4, we present estimates of the exponentiated parameters from these models, along with 95% non-parametric bootstrap percentile confidence intervals (based on 1000 resamples within treatment groups). In Table 2, missingness of sputum culture (aor= 4.45; 95% CI: ) and missingness of smear (aor= 3.51; 95% CI: ) at a previous visit are significant predictors of missingness of sputum culture at the next visit. In Table 3, among patients with an observed sputum culture at visit k, missingness of sputum culture (aor = 4.00; 95% CI: ), missingness of a smear culture (aor = 5.16; 95% CI: ), a negative observed culture (aor = 6.37; 95% CI: ) and a negative observed smear (aor = 3.93; 95% CI: ) at visit k 1, as well as assignment to the moxifloxacin arm (aor = 2.06; 95% CI: ), are significant predictors of a negative sputum culture at visit k. In Table 4, among patients with an observed smear at visit k, missingness of smear (aor = 3.66; 95% CI: ) and an observed negative smear (aor = 6.99; 95% CI: ) at visit k 1, as well as an observed negative sputum culture at visit k (aor = 10.73; 95% CI: ), are significant predictors of a negative smear at visit k. Under our benchmark assumption, the estimated hazard ratio is 3.41 (95% CI: [1.33,13.06]), indicating that patients treated with moxifloxacin have a statistically significant shorter time of culture conversion than those treated with ethambutol. Figure 3 displays a contour plot of the estimated odds
16 16 D. SCHARFSTEIN ET AL. Table 2 (Exponentiated) parameter estimates and 95% confidence intervals from the model for missingness of culture results. See a z(k, O k 1, X; γ (a) ) for form of the model. Intercept Odds 95% CI Wk 1 (exp(γ (a) 0,0,1 )) 0.14 [0.04,0.30] Wk 2 (exp(γ (a) 0,0,2 )) 0.11 [0.03,0.24] Wk 3 (exp(γ (a) 0,0,3 )) 0.03 [0.00,0.09] Wk 4 (exp(γ (a) 0,0,4 )) 0.11 [0.03,0.25] Wk 5 (exp(γ (a) 0,0,5 )) 0.07 [0.01,0.19] Wk 6 (exp(γ (a) 0,0,6 )) 0.04 [0.00,0.12] Wk 7 (exp(γ (a) 0,0,7 )) 0.05 [0.00,0.14] Wk 8 (exp(γ (a) 0,0,8 )) 0.07 [0.01,0.18] Predictor Odds Ratio 95% CI Wk 1*Cav (exp(γ (a) 0,1,1 )) 0.19 [0.00,0.90] Wk 2*Cav (exp(γ (a) 0,1,2 )) 0.21 [0.00,0.87] Wk 3*Cav (exp(γ (a) 0,1,3 )) 2.93 [0.81, ] Wk 4*Cav (exp(γ (a) 0,1,4 )) 0.38 [0.09,1.50] Wk 5*Cav (exp(γ (a) 0,1,5 )) 1.63 [0.55,11.28] Wk 6*Cav (exp(γ (a) 0,1,6 )) 1.39 [0.36, ] Wk 7*Cav (exp(γ (a) 0,1,7 )) 1.60 [0.47, ] Wk 8*Cav (exp(γ (a) 0,1,8 )) 1.63 [0.51,8.32] I(k > 1)Mk 1 c (exp(γ (a) 1 )) 4.45 [2.03,9.70] I(k > 1)(1 Mk 1)C c k 1 obs (exp(γ (a) 2 )) 0.89 [0.50,1.68] I(k > 1)Mk 1 s (exp(γ (a) 3 )) 3.51 [1.52,9.66] I(k > 1)(1 Mk 1)S s k 1 obs (exp(γ (a) 4 )) 1.28 [0.72,2.28] Moxifloxacin (exp(γ (a) 5 )) 1.07 [0.67,1.76] here means a big number ratio as a function of α 0 and α 1. The region is white indicates combinations of α 0 and α 1 where the lower bound of the 95% confidence interval is less than one. The green region indicates combinations of α 0 and α 1 where the null of no treatment difference is rejected in favor of moxifloxacin. Inference would change relative to the benchmark assumption (circle in Figure 3), if say α 0 = 5.0, α 1 = 3.0 (triangle in Figure 3) or α 0 = 4.0, α 1 = 10.0 (square in Figure 3). At these combination of treatmentspecific sensitivity analysis parameters, the estimated treatment effects are 2.19 (95% CI: ) and 2.07 (95% CI: ). To understand whether these combinations are far from the benchmark assumption, consider Figure 4. In the first row, we plot for each treatment group, the estimated distribution of time of culture conversion for these sensitivity analysis parameters (dashed and dotted lines) and the estimated distributions under the benchmark assumption (solid lines). In the second row, we plot for each
17 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 17 Fig 3. Contour plot of the estimated ratio of the odds of first becoming a culture converter at visit k given culture conversion at or after visit k, comparing moxifloxacin vs. ethambutol as a function of the sensitivity-analysis parameters α 0 and α 1. The region is white indicates combinations of α 0 and α 1 where the lower bound of the 95% confidence interval is less than one. The green region indicates combinations of α 0 and α 1 where the null of no treatment difference is rejected in favor of moxifloxacin. Circle denotes the benchmark assumption (α 0 = 0.0, α 1 = 0.0). Triangle (α 0 = 5, α 1 = 3) and square (α 0 = 4, α 1 = 10) denotes two combinations discussed in the text. 5 0 α α 0
18 18 D. SCHARFSTEIN ET AL. Table 3 (Exponentiated) parameter estimates and 95% confidence intervals from the model for culture results. See b z(k, O k 1, X; γ (b) ) for form of the model. Intercept Odds 95% CI Wk 1 (exp(γ (b) 0,0,1 )) 0.04 [0.02,0.08] Wk 2 (exp(γ (b) 0,0,1 )) 0.03 [0.02,0.06] Wk 3 (exp(γ (b) 0,0,1 )) 0.08 [0.04,0.14] Wk 4 (exp(γ (b) 0,0,1 )) 0.10 [0.05,0.18] Wk 5 (exp(γ (b) 0,0,1 )) 0.10 [0.05,0.17] Wk 6 (exp(γ (b) 0,0,1 )) 0.24 [ ] Wk 7 (exp(γ (b) 0,0,1 )) 0.22 [0.12,0.40] Wk 8 (exp(γ (b) 0,0,1 )) 0.49 [0.26,0.95] Predictor Odds Ratio 95% CI I(k > 1)Mk 1 c (exp(γ (b) 1 )) 4.00 [2.00,9.45] (k > 1)(1 Mk 1)C c k 1 obs (exp(γ (b) 2 )) 6.37 [4.08,10.39] I(k > 1)Mk 1 s (exp(γ (b) 3 )) 5.16 [1.67,22.47] I(k > 1)(1 Mk 1)S s k 1 obs (exp(γ (b) 4 )) 3.93 [2.51,6.18] Moxifloxacin (exp(γ (b) 5 )) 2.06 [1.46,3.19] Cavitation (exp(γ (b) 6 )) 1.16 [0.81,1.75] treatment group, the signed Kolmogorov distance between the estimated distribution of time of culture conversion for given α and the estimated distribution function of time of culture conversion under the benchmark assumption. The signed Kolmogorov distance for treatment group z with sensitivity analysis parameter α z equals F z (k z; α z ) F z (k z; 0), where k z = argmax k F z (k; α z ) F z (k; 0) and F z (k; α z ) is the estimated cumulative distribution function. When α 0 = 5.0 and α 1 = 3.0, the signed distances for the ethambutol and moxifloxacin arms are and 0.11; the latter being a fairly sizable difference (for the moxifloxacin arm, the estimated probability of culture conversion by Visit 5 is 49.6% under the benchmark assumption and 38.2% when α 1 = 3.0). Further, the distances are of opposite signs (i.e., the bias is differential between arms). When we look at other combinations of sensitivity analysis parameters where the null hypothesis is not rejected, the sensitivity analysis parameter for the moxifloxacin arm is less than or equal to 3.0 and the associated signed distances are at least as extreme as When α 0 = 4.0 and α 1 = 10.0, the signed distances are 0.11 and 0.16 for the ethambutol and moxifloxacin arms, respectively. Here the signs are in the same direction, but the choice of sensitivity analysis parameters yield results that are very close to the worst case bounds that assume that all missing
19 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 19 Table 4 (Exponentiated) parameter estimates and 95% confidence intervals from the model for smear results. See d z(k, Mk, c Ck obs, O k 1, X; γ (d) ) for form of the model. Intercept Odds 95% CI Wk 1 (exp(γ (d) 0,0,1 )) 0.25 [0.13, 0.42] Wk 2 (exp(γ (d) 0,0,1 )) 0.34 [0.20,0.56] Wk 3 (exp(γ (d) 0,0,1 )) 0.35 [0.21,0.58] Wk 4 (exp(γ (d) 0,0,1 )) 0.21 [0.11,0.37] Wk 5 (exp(γ (d) 0,0,1 )) 0.35 [0.19,0.65] Wk 6 (exp(γ (d) 0,0,1 )) 0.20 [0.10,0.39] Wk 7 (exp(γ (d) 0,0,1 )) 0.23 [0.12,0.43] Wk 8 (exp(γ (d) 0,0,1 )) 0.24 [0.11,0.53] Predictor Odds Ratio 95% CI Mk c (exp(γ (d) 1 )) 1.24 [0.61, 2.61] (1 Mk)C c k obs (exp(γ (d) 2 )) [5.58, 31.18] (1 Mk)C c k obs *Cav (exp(γ (d) 3 )) 0.46 [0.16,1.04] I(k > 1)Mk 1 c (exp(γ (d) 4 )) 1.52 [0.66,3.46] I(k > 1)(1 Mk 1)C c k 1 obs (exp(γ (d) 5 )) 1.44 [0.92,2.38] I(k > 1)Mk 1 s 3.66 (exp(γ (d) 6 )) [1.45,14.53] I(k > 1)(1 Mk 1)S s k 1 obs (exp(γ (d) 7 )) 6.99 [4.50,11.81] Moxifloxacin (exp(γ (d) 8 )) 0.97 [0.63,1.48] Cavitation (exp(γ (d) 9 )) 1.18 [0.75,1.85] cultures are positive. From a clinical perspective, inferences relative to the benchmark assumption are fairly robust. 5. Discussion. Conde et al. (2009) did not compare the treatments with respect to time of culture conversion. Rather, they compared the treatmentspecific probabilities of being a culture converter at or prior to week 8, which is equivalent to having a negative culture at week 8. They used two different methods. The primary method assumed that all missing cultures at week 8 were positive (moxifloxacin: 77.0%; ethambutol: 62.5%; difference: 14.5 percentage points, 95% CI [ 0.0 percentage points, 29.0 percentage points]); the secondary method excluded patients who were missing their week 8 culture, assuming that the missing cultures were missing completely at random (moxifloxacin: 90.5%; ethambutol: 73.8%; difference: 16.7 percentage points, 95% CI [3.4 percentage points, 30.4 percentage points]). The former analysis is not statistically significant, while the latter analysis does suggest a statistically significant treatment effect in favor of moxifloxacin. In their analysis, no attempt was made to account for imbalances in baseline cavitation status between treatment groups. It is tempting to think that this problem can be addressed by simply ana-
20 20 D. SCHARFSTEIN ET AL. Fig 4. First Row: Treatment-specific estimated distribution of time of culture conversion for the benchmark (α 0 = 0, α 1 = 0) and alternative (α 0 = 5.0, α 1 = 3) sensitivityanalysis parameters. Second Row: For each treatment group, the signed Kolmogorov distance between the estimated distribution of time of culture conversion for given α and the estimated distribution function of time of culture conversion under the benchmark assumption. Ethambutol Moxifloxacin Probability Probability Visit Visit Ethambutol Moxifloxacin Signed Distance Signed Distance α 0 α 1
21 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 21 lyzing the culture results using standard statistical methods for longitudinal binary data, e.g., marginal models and generalized linear mixed models. A marginal model, fit using generalized estimating equations, identifies, under the assumption that the culture results are missing completely at random, the probability of a negative culture at each visit k; it does not admit identification of the distribution of time of culture conversion. In contrast, a generalized linear mixed model is a fully parametric model for the joint distribution of the culture results and, under the missing at random assumption, admits identification of the distribution of time of culture conversion. However, the modeling assumptions are too strong as they essentially allow the imputation of missing culture results that are not needed to identify the distribution of interest, e.g., the imputation of missing cultures that are proceeded by positive cultures. Further, the model induces testable restrictions and, as discussed by Robins and Gill (1997), the missing at random assumption is often unrealistic in follow-up studies with intermittent missing data. Nonetheless, we fit a logistic-normal generalized linear mixed model to the culture data with fixed effects for time, treatment and cavitation, and a random intercept. Using the model, we estimated, within levels of time, treatment and cavitation, the induced probability of a negative culture among those with missing cultures. Many of the estimated probabilities were either greater than one or less than zero, suggesting inadequate model fit. An alternative way of analyzing the culture data would treat the data for each patient as the set of times of culture conversion that are consistent with their observed culture data (i.e., the coarsening set) and estimate the distribution of time to culture conversion under the coarsening at random (CAR) assumption [Gill, Van Der Laan and Robins (1997); Heitjan and Rubin (1991); Heitjan (1993, 1994)]. This assumption states that the coarsening process provides no information about the time of culture conversion beyond conveying that the true event time is in the observed coarsening set. Under CAR, the coarsening process is ignorable, i.e., it factors out of the likelihood for the observed data. Under this assumption, the estimated value of β is 2.92 (95%: ). The result is statistically significant and favors moxifloxacin. We have assumed, as in most analyses of culture conversion data, that the test results are measured without error. We know that this is not the case. It would be interesting to use known information about the sensitivity and specificity of the culture and smear procedures to learn about the true distribution of time of culture conversion. This will be the subject of future research.
22 22 D. SCHARFSTEIN ET AL. In our analysis, we did not have access to the reasons for missingness. We know them to be a combination of three main sources: culture contamination, inability to produce sputum and skipped clinic visits. In the former two cases, the culture results are more likely to be negative. Contamination of sputum cultures with bacteria from the mouth and airways occurs in 2-10% of specimens and varies by laboratory. Patients producing smaller amounts of sputum that is mixed with saliva are more likely to have contamination; therefore, patients who have responded to therapy (i.e., have negative cultures) and no longer are producing large volumes of sputum may be more likely to have a contaminated specimen. Patients with treated tuberculosis who can no longer produce sputum are also likely to have responded to therapy and have negative cultures. If most of the missing data is due to these two causes, it is not so surprising that the results of the benchmark analysis are so close to the best-case bounds. In summary, we introduced a novel benchmark assumption that allows us to learn about the distribution of just those unknown culture results that are absolutely necessary to identify the distribution of time of culture conversion by borrowing strength from patients as similar as possible (with respect to baseline cavitation status, treatment assignment, and observed culture and sputum results) and on whom the distribution of these culture results is identified. We evaluated the sensitivity of inferences to our benchmark assumption by embedding it within a class of model assumptions indexed by sensitivity-analysis parameters. While the sensitivity-analysis parameters themselves are not scientifically interpretable, the induced distribution of time of culture conversion (and functionals thereof) can be estimated and compared with that under the benchmark assumption. If the differences are judged large by scientific experts, we hope that they will comment on the fragility or robustness of the benchmark inference. Except in rare settings where the treatment effects are so dramatic or missing data are so minor, we see no alternatives to sensitivity analysis aided by scientific judgement. Acknowledgements. The authors would like to thank Jonghyeon Kim, Chad Heilig, Malathi Ram, Pei-Jean Feng and Swarnadip Ghosh for assistance during the conduct of this research.
23 TUBERCULOSIS STUDIES WITH MISSING SPUTUM DATA 23 APPENDIX A straightforward application of the law of total probability entails that under models (3.6) - (3.9), P [T = j O = o (j 1) ; γ] = g j 1 (0, O j, X; γ) 1y=0 g j 1 (y, O j, X; γ) where g j (y, O j+1, X; γ) =(1 expit{a(j + 1, O j (0, y), X; γ (a) )}) expit{b(j, O j 1, X; γ (b) )} y (1 expit{b(j, O j 1, X; γ (b) )}) (1 y) expit{b(j + 1, O j (0, y), X; γ (b) )} (1 expit{c(j, 0, y, O j 1, X; γ (c) )}) (1 M j s) expit{c(j, 0, y, O j 1, X; γ (c) )} M j s (1 expit{c(j + 1, 0, 1, O j (0, y), X; γ (c) )}) (1 M j+1 s ) expit{c(j + 1, 0, 1, O j (0, y), X; γ (c) )} M j+1 s (1 expit{d(j, 0, y, O j 1, X; γ (d) )}) (1 Sj) expit{d(j, 0, y, O j 1, X; γ (d) )} S j (1 expit{d(j + 1, 0, 1, O j (0, y), X; γ (d) )}) (1 Sj+1) expit{d(j + 1, 0, 1, O j (0, y), X; γ (d) )} S j+1 and O j (0, y) represents observed data visit j with Mj c set to 0 and C j set y. Notice that if c(k, Mk c, Cobs k, O k 1, X; γ (c) ) does not depend on Ck obs and Ck 1 obs for all k, then P [T = j O = o(j 1) ; γ] does not depend on γ (c).
24 24 D. SCHARFSTEIN ET AL. REFERENCES American Thoracic Society, (2000). Diagnostic standards and classification of tuberculosis in adults and children. American Journal of Respiratory and Critical Care Medicine Conde, M. B., Efron, A., Loredo, C., De Souza, G. R. M., Graça, N. P., Cezar, M. C., Ram, M., Chaudhary, M. A., Bishai, W. R., Kritski, A. L. and Chaisson, R. E. (2009). Moxifloxacin versus ethambutol in the initial treatment of tuberculosis: a double-blind, randomised, controlled phase II trial. Lancet Cox, D. R. (1972). Regression models and life-tables (with discussion). Journal of the Royal Statistical Society Series B Cox, D. and Barndorff-Nielsen, O. (1994). Inference and asymptotics. Chapman & Hall/CRC. European Medicines Agency, Committee for Medicinal Products for Human Use, (2010). Addendum to the Note for Guidance on Evaluation of Medicinal Products Indicated for Treatment of Bacterial Infections to Specifically Address the Clinical Development of New Agents to Treat Disease due to Mycobacterium Tuberculosis. European Medicines Agency, London. Gill, R. D., Van Der Laan, M. J. and Robins, J. M. (1997). Coarsening at random: characterizations, conjectures and counter-examples. In Proceedings of the First Seattle Symposium in Biostatistics: Survival Analysis (D. Y. Lin and T. R. Fleming, eds.) Springer, Berlin. Heitjan, D. F. (1993). Ignorability and coarse data: Some biomedical examples. Biometrics Heitjan, D. F. (1994). Ignorability in general incomplete-data models. Biometrika Heitjan, D. F. and Rubin, D. B. (1991). Ignorability and coarse data. Annals of Statistics Newey, W. K. and McFadden, D. (1994). Large sample estimation and hypothesis testing. In Handbook of Econometrics, (R. F. Engle and D. L. McFadden, eds.) IV Elsevier, Amsterdam. Robins, J. M. and Gill, R. D. (1997). Non-response models for the analysis of nonmonotone ignorable missing data. Statistics in Medicine Daniel Scharfstein Adian McDermott Department of Biostatistics Johns Hopkins Bloomberg School of Public Health 615 North Wolfe Street Baltimore, Maryland dscharf@jhsph.edu amcdermo@jhsph.edu Maria Abraham Statistics Collaborative 1625 Massachusetts Ave NW Suite 600 Washington, DC maria@statcollab.com Andrea Rotnitzky Department of Economics Universidad Torcuato Di Tella Saenz Valiente Buenos Aires arotnitzky@utdt.edu Richard Chaisson Johns Hopkins Center for Tuberculosis Research 1550 Orleans St., 1M.08 Baltimore, MD rchaiss@jhmi.edu
ON THE ANALYSIS OF TUBERCULOSIS STUDIES WITH INTERMITTENT MISSING SPUTUM DATA
 Submitted to the Annals of Applied Statistics arxiv: arxiv:0000.0000 ON THE ANALYSIS OF TUBERCULOSIS STUDIES WITH INTERMITTENT MISSING SPUTUM DATA By Daniel Scharfstein, Andrea Rotnitzky, Maria Abraham
Submitted to the Annals of Applied Statistics arxiv: arxiv:0000.0000 ON THE ANALYSIS OF TUBERCULOSIS STUDIES WITH INTERMITTENT MISSING SPUTUM DATA By Daniel Scharfstein, Andrea Rotnitzky, Maria Abraham
On the Analysis of Tuberculosis Studies with Intermittent Missing Sputum Data
 On the Analysis of Tuberculosis Studies with Intermittent Missing Sputum Data Daniel Scharfstein Johns Hopkins University dscharf@jhsph.edu July 12, 2013 Happy Birthday! Collaborators Andrea Rotnitzky
On the Analysis of Tuberculosis Studies with Intermittent Missing Sputum Data Daniel Scharfstein Johns Hopkins University dscharf@jhsph.edu July 12, 2013 Happy Birthday! Collaborators Andrea Rotnitzky
Chapter 4. Parametric Approach. 4.1 Introduction
 Chapter 4 Parametric Approach 4.1 Introduction The missing data problem is already a classical problem that has not been yet solved satisfactorily. This problem includes those situations where the dependent
Chapter 4 Parametric Approach 4.1 Introduction The missing data problem is already a classical problem that has not been yet solved satisfactorily. This problem includes those situations where the dependent
A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness
 A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness A. Linero and M. Daniels UF, UT-Austin SRC 2014, Galveston, TX 1 Background 2 Working model
A Bayesian Nonparametric Approach to Monotone Missing Data in Longitudinal Studies with Informative Missingness A. Linero and M. Daniels UF, UT-Austin SRC 2014, Galveston, TX 1 Background 2 Working model
On Testing the Missing at Random Assumption
 On Testing the Missing at Random Assumption Manfred Jaeger Institut for Datalogi, Aalborg Universitet, Fredrik Bajers Vej 7E, DK-9220 Aalborg Ø jaeger@cs.aau.dk Abstract. Most approaches to learning from
On Testing the Missing at Random Assumption Manfred Jaeger Institut for Datalogi, Aalborg Universitet, Fredrik Bajers Vej 7E, DK-9220 Aalborg Ø jaeger@cs.aau.dk Abstract. Most approaches to learning from
Improving Efficiency of Inferences in Randomized Clinical Trials Using Auxiliary Covariates
 Improving Efficiency of Inferences in Randomized Clinical Trials Using Auxiliary Covariates Anastasios (Butch) Tsiatis Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
Improving Efficiency of Inferences in Randomized Clinical Trials Using Auxiliary Covariates Anastasios (Butch) Tsiatis Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs
 Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs Michael J. Daniels and Chenguang Wang Jan. 18, 2009 First, we would like to thank Joe and Geert for a carefully
Discussion of Missing Data Methods in Longitudinal Studies: A Review by Ibrahim and Molenberghs Michael J. Daniels and Chenguang Wang Jan. 18, 2009 First, we would like to thank Joe and Geert for a carefully
PubH 7470: STATISTICS FOR TRANSLATIONAL & CLINICAL RESEARCH
 PubH 7470: STATISTICS FOR TRANSLATIONAL & CLINICAL RESEARCH The First Step: SAMPLE SIZE DETERMINATION THE ULTIMATE GOAL The most important, ultimate step of any of clinical research is to do draw inferences;
PubH 7470: STATISTICS FOR TRANSLATIONAL & CLINICAL RESEARCH The First Step: SAMPLE SIZE DETERMINATION THE ULTIMATE GOAL The most important, ultimate step of any of clinical research is to do draw inferences;
Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials
 Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials Progress, Updates, Problems William Jen Hoe Koh May 9, 2013 Overview Marginal vs Conditional What is TMLE? Key Estimation
Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials Progress, Updates, Problems William Jen Hoe Koh May 9, 2013 Overview Marginal vs Conditional What is TMLE? Key Estimation
7 Sensitivity Analysis
 7 Sensitivity Analysis A recurrent theme underlying methodology for analysis in the presence of missing data is the need to make assumptions that cannot be verified based on the observed data. If the assumption
7 Sensitivity Analysis A recurrent theme underlying methodology for analysis in the presence of missing data is the need to make assumptions that cannot be verified based on the observed data. If the assumption
Analysing longitudinal data when the visit times are informative
 Analysing longitudinal data when the visit times are informative Eleanor Pullenayegum, PhD Scientist, Hospital for Sick Children Associate Professor, University of Toronto eleanor.pullenayegum@sickkids.ca
Analysing longitudinal data when the visit times are informative Eleanor Pullenayegum, PhD Scientist, Hospital for Sick Children Associate Professor, University of Toronto eleanor.pullenayegum@sickkids.ca
SIMPLE EXAMPLES OF ESTIMATING CAUSAL EFFECTS USING TARGETED MAXIMUM LIKELIHOOD ESTIMATION
 Johns Hopkins University, Dept. of Biostatistics Working Papers 3-3-2011 SIMPLE EXAMPLES OF ESTIMATING CAUSAL EFFECTS USING TARGETED MAXIMUM LIKELIHOOD ESTIMATION Michael Rosenblum Johns Hopkins Bloomberg
Johns Hopkins University, Dept. of Biostatistics Working Papers 3-3-2011 SIMPLE EXAMPLES OF ESTIMATING CAUSAL EFFECTS USING TARGETED MAXIMUM LIKELIHOOD ESTIMATION Michael Rosenblum Johns Hopkins Bloomberg
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2004 Paper 155 Estimation of Direct and Indirect Causal Effects in Longitudinal Studies Mark J. van
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2004 Paper 155 Estimation of Direct and Indirect Causal Effects in Longitudinal Studies Mark J. van
Global Sensitivity Analysis for Repeated Measures Studies with. Informative Drop-out: A Fully Parametric Approach
 Global Sensitivity Analysis for Repeated Measures Studies with Informative Drop-out: A Fully Parametric Approach Daniel Scharfstein (dscharf@jhsph.edu) Aidan McDermott (amcdermo@jhsph.edu) Department of
Global Sensitivity Analysis for Repeated Measures Studies with Informative Drop-out: A Fully Parametric Approach Daniel Scharfstein (dscharf@jhsph.edu) Aidan McDermott (amcdermo@jhsph.edu) Department of
Global Sensitivity Analysis for Repeated Measures Studies with Informative Drop-out: A Semi-Parametric Approach
 Global for Repeated Measures Studies with Informative Drop-out: A Semi-Parametric Approach Daniel Aidan McDermott Ivan Diaz Johns Hopkins University Ibrahim Turkoz Janssen Research and Development September
Global for Repeated Measures Studies with Informative Drop-out: A Semi-Parametric Approach Daniel Aidan McDermott Ivan Diaz Johns Hopkins University Ibrahim Turkoz Janssen Research and Development September
FULL LIKELIHOOD INFERENCES IN THE COX MODEL
 October 20, 2007 FULL LIKELIHOOD INFERENCES IN THE COX MODEL BY JIAN-JIAN REN 1 AND MAI ZHOU 2 University of Central Florida and University of Kentucky Abstract We use the empirical likelihood approach
October 20, 2007 FULL LIKELIHOOD INFERENCES IN THE COX MODEL BY JIAN-JIAN REN 1 AND MAI ZHOU 2 University of Central Florida and University of Kentucky Abstract We use the empirical likelihood approach
Robust estimates of state occupancy and transition probabilities for Non-Markov multi-state models
 Robust estimates of state occupancy and transition probabilities for Non-Markov multi-state models 26 March 2014 Overview Continuously observed data Three-state illness-death General robust estimator Interval
Robust estimates of state occupancy and transition probabilities for Non-Markov multi-state models 26 March 2014 Overview Continuously observed data Three-state illness-death General robust estimator Interval
A Sampling of IMPACT Research:
 A Sampling of IMPACT Research: Methods for Analysis with Dropout and Identifying Optimal Treatment Regimes Marie Davidian Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
A Sampling of IMPACT Research: Methods for Analysis with Dropout and Identifying Optimal Treatment Regimes Marie Davidian Department of Statistics North Carolina State University http://www.stat.ncsu.edu/
Sensitivity analysis and distributional assumptions
 Sensitivity analysis and distributional assumptions Tyler J. VanderWeele Department of Health Studies, University of Chicago 5841 South Maryland Avenue, MC 2007, Chicago, IL 60637, USA vanderweele@uchicago.edu
Sensitivity analysis and distributional assumptions Tyler J. VanderWeele Department of Health Studies, University of Chicago 5841 South Maryland Avenue, MC 2007, Chicago, IL 60637, USA vanderweele@uchicago.edu
Targeted Maximum Likelihood Estimation in Safety Analysis
 Targeted Maximum Likelihood Estimation in Safety Analysis Sam Lendle 1 Bruce Fireman 2 Mark van der Laan 1 1 UC Berkeley 2 Kaiser Permanente ISPE Advanced Topics Session, Barcelona, August 2012 1 / 35
Targeted Maximum Likelihood Estimation in Safety Analysis Sam Lendle 1 Bruce Fireman 2 Mark van der Laan 1 1 UC Berkeley 2 Kaiser Permanente ISPE Advanced Topics Session, Barcelona, August 2012 1 / 35
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution
 Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Semiparametric Mixed Effects Models with Flexible Random Effects Distribution Marie Davidian North Carolina State University davidian@stat.ncsu.edu www.stat.ncsu.edu/ davidian Joint work with A. Tsiatis,
Approximation of Survival Function by Taylor Series for General Partly Interval Censored Data
 Malaysian Journal of Mathematical Sciences 11(3): 33 315 (217) MALAYSIAN JOURNAL OF MATHEMATICAL SCIENCES Journal homepage: http://einspem.upm.edu.my/journal Approximation of Survival Function by Taylor
Malaysian Journal of Mathematical Sciences 11(3): 33 315 (217) MALAYSIAN JOURNAL OF MATHEMATICAL SCIENCES Journal homepage: http://einspem.upm.edu.my/journal Approximation of Survival Function by Taylor
Multi-state Models: An Overview
 Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Multi-state Models: An Overview Andrew Titman Lancaster University 14 April 2016 Overview Introduction to multi-state modelling Examples of applications Continuously observed processes Intermittently observed
Bootstrapping Sensitivity Analysis
 Bootstrapping Sensitivity Analysis Qingyuan Zhao Department of Statistics, The Wharton School University of Pennsylvania May 23, 2018 @ ACIC Based on: Qingyuan Zhao, Dylan S. Small, and Bhaswar B. Bhattacharya.
Bootstrapping Sensitivity Analysis Qingyuan Zhao Department of Statistics, The Wharton School University of Pennsylvania May 23, 2018 @ ACIC Based on: Qingyuan Zhao, Dylan S. Small, and Bhaswar B. Bhattacharya.
Biost 518 Applied Biostatistics II. Purpose of Statistics. First Stage of Scientific Investigation. Further Stages of Scientific Investigation
 Biost 58 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture 5: Review Purpose of Statistics Statistics is about science (Science in the broadest
Biost 58 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture 5: Review Purpose of Statistics Statistics is about science (Science in the broadest
Bayesian methods for missing data: part 1. Key Concepts. Nicky Best and Alexina Mason. Imperial College London
 Bayesian methods for missing data: part 1 Key Concepts Nicky Best and Alexina Mason Imperial College London BAYES 2013, May 21-23, Erasmus University Rotterdam Missing Data: Part 1 BAYES2013 1 / 68 Outline
Bayesian methods for missing data: part 1 Key Concepts Nicky Best and Alexina Mason Imperial College London BAYES 2013, May 21-23, Erasmus University Rotterdam Missing Data: Part 1 BAYES2013 1 / 68 Outline
Sample size re-estimation in clinical trials. Dealing with those unknowns. Chris Jennison. University of Kyoto, January 2018
 Sample Size Re-estimation in Clinical Trials: Dealing with those unknowns Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj University of Kyoto,
Sample Size Re-estimation in Clinical Trials: Dealing with those unknowns Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj University of Kyoto,
Minimax-Regret Sample Design in Anticipation of Missing Data, With Application to Panel Data. Jeff Dominitz RAND. and
 Minimax-Regret Sample Design in Anticipation of Missing Data, With Application to Panel Data Jeff Dominitz RAND and Charles F. Manski Department of Economics and Institute for Policy Research, Northwestern
Minimax-Regret Sample Design in Anticipation of Missing Data, With Application to Panel Data Jeff Dominitz RAND and Charles F. Manski Department of Economics and Institute for Policy Research, Northwestern
Selection on Observables: Propensity Score Matching.
 Selection on Observables: Propensity Score Matching. Department of Economics and Management Irene Brunetti ireneb@ec.unipi.it 24/10/2017 I. Brunetti Labour Economics in an European Perspective 24/10/2017
Selection on Observables: Propensity Score Matching. Department of Economics and Management Irene Brunetti ireneb@ec.unipi.it 24/10/2017 I. Brunetti Labour Economics in an European Perspective 24/10/2017
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features. Yangxin Huang
 Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
Bayesian Inference on Joint Mixture Models for Survival-Longitudinal Data with Multiple Features Yangxin Huang Department of Epidemiology and Biostatistics, COPH, USF, Tampa, FL yhuang@health.usf.edu January
UNIVERSITY OF CALIFORNIA, SAN DIEGO
 UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
UNIVERSITY OF CALIFORNIA, SAN DIEGO Estimation of the primary hazard ratio in the presence of a secondary covariate with non-proportional hazards An undergraduate honors thesis submitted to the Department
6 Pattern Mixture Models
 6 Pattern Mixture Models A common theme underlying the methods we have discussed so far is that interest focuses on making inference on parameters in a parametric or semiparametric model for the full data
6 Pattern Mixture Models A common theme underlying the methods we have discussed so far is that interest focuses on making inference on parameters in a parametric or semiparametric model for the full data
Deductive Derivation and Computerization of Semiparametric Efficient Estimation
 Deductive Derivation and Computerization of Semiparametric Efficient Estimation Constantine Frangakis, Tianchen Qian, Zhenke Wu, and Ivan Diaz Department of Biostatistics Johns Hopkins Bloomberg School
Deductive Derivation and Computerization of Semiparametric Efficient Estimation Constantine Frangakis, Tianchen Qian, Zhenke Wu, and Ivan Diaz Department of Biostatistics Johns Hopkins Bloomberg School
36 th Annual Meeting Preconference Workshop P3 Handout
 36 th Annual Meeting Preconference Workshop P3 Handout Global Sensitivity Analysis of Randomized Trials with Missing Data Daniel Scharfstein Johns Hopkins University dscharf@jhu.edu May 17, 2015 1 / 75
36 th Annual Meeting Preconference Workshop P3 Handout Global Sensitivity Analysis of Randomized Trials with Missing Data Daniel Scharfstein Johns Hopkins University dscharf@jhu.edu May 17, 2015 1 / 75
A COMPARISON OF POISSON AND BINOMIAL EMPIRICAL LIKELIHOOD Mai Zhou and Hui Fang University of Kentucky
 A COMPARISON OF POISSON AND BINOMIAL EMPIRICAL LIKELIHOOD Mai Zhou and Hui Fang University of Kentucky Empirical likelihood with right censored data were studied by Thomas and Grunkmier (1975), Li (1995),
A COMPARISON OF POISSON AND BINOMIAL EMPIRICAL LIKELIHOOD Mai Zhou and Hui Fang University of Kentucky Empirical likelihood with right censored data were studied by Thomas and Grunkmier (1975), Li (1995),
Causal inference in epidemiological practice
 Causal inference in epidemiological practice Willem van der Wal Biostatistics, Julius Center UMC Utrecht June 5, 2 Overview Introduction to causal inference Marginal causal effects Estimating marginal
Causal inference in epidemiological practice Willem van der Wal Biostatistics, Julius Center UMC Utrecht June 5, 2 Overview Introduction to causal inference Marginal causal effects Estimating marginal
Personalized Treatment Selection Based on Randomized Clinical Trials. Tianxi Cai Department of Biostatistics Harvard School of Public Health
 Personalized Treatment Selection Based on Randomized Clinical Trials Tianxi Cai Department of Biostatistics Harvard School of Public Health Outline Motivation A systematic approach to separating subpopulations
Personalized Treatment Selection Based on Randomized Clinical Trials Tianxi Cai Department of Biostatistics Harvard School of Public Health Outline Motivation A systematic approach to separating subpopulations
Reconstruction of individual patient data for meta analysis via Bayesian approach
 Reconstruction of individual patient data for meta analysis via Bayesian approach Yusuke Yamaguchi, Wataru Sakamoto and Shingo Shirahata Graduate School of Engineering Science, Osaka University Masashi
Reconstruction of individual patient data for meta analysis via Bayesian approach Yusuke Yamaguchi, Wataru Sakamoto and Shingo Shirahata Graduate School of Engineering Science, Osaka University Masashi
Extending causal inferences from a randomized trial to a target population
 Extending causal inferences from a randomized trial to a target population Issa Dahabreh Center for Evidence Synthesis in Health, Brown University issa dahabreh@brown.edu January 16, 2019 Issa Dahabreh
Extending causal inferences from a randomized trial to a target population Issa Dahabreh Center for Evidence Synthesis in Health, Brown University issa dahabreh@brown.edu January 16, 2019 Issa Dahabreh
Chapter 1 Statistical Inference
 Chapter 1 Statistical Inference causal inference To infer causality, you need a randomized experiment (or a huge observational study and lots of outside information). inference to populations Generalizations
Chapter 1 Statistical Inference causal inference To infer causality, you need a randomized experiment (or a huge observational study and lots of outside information). inference to populations Generalizations
Statistical Methods. Missing Data snijders/sm.htm. Tom A.B. Snijders. November, University of Oxford 1 / 23
 1 / 23 Statistical Methods Missing Data http://www.stats.ox.ac.uk/ snijders/sm.htm Tom A.B. Snijders University of Oxford November, 2011 2 / 23 Literature: Joseph L. Schafer and John W. Graham, Missing
1 / 23 Statistical Methods Missing Data http://www.stats.ox.ac.uk/ snijders/sm.htm Tom A.B. Snijders University of Oxford November, 2011 2 / 23 Literature: Joseph L. Schafer and John W. Graham, Missing
Power and Sample Size Calculations with the Additive Hazards Model
 Journal of Data Science 10(2012), 143-155 Power and Sample Size Calculations with the Additive Hazards Model Ling Chen, Chengjie Xiong, J. Philip Miller and Feng Gao Washington University School of Medicine
Journal of Data Science 10(2012), 143-155 Power and Sample Size Calculations with the Additive Hazards Model Ling Chen, Chengjie Xiong, J. Philip Miller and Feng Gao Washington University School of Medicine
8 Nominal and Ordinal Logistic Regression
 8 Nominal and Ordinal Logistic Regression 8.1 Introduction If the response variable is categorical, with more then two categories, then there are two options for generalized linear models. One relies on
8 Nominal and Ordinal Logistic Regression 8.1 Introduction If the response variable is categorical, with more then two categories, then there are two options for generalized linear models. One relies on
Group Sequential Designs: Theory, Computation and Optimisation
 Group Sequential Designs: Theory, Computation and Optimisation Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj 8th International Conference
Group Sequential Designs: Theory, Computation and Optimisation Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj 8th International Conference
Multivariate Survival Analysis
 Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Multivariate Survival Analysis Previously we have assumed that either (X i, δ i ) or (X i, δ i, Z i ), i = 1,..., n, are i.i.d.. This may not always be the case. Multivariate survival data can arise in
Lecture 7 Time-dependent Covariates in Cox Regression
 Lecture 7 Time-dependent Covariates in Cox Regression So far, we ve been considering the following Cox PH model: λ(t Z) = λ 0 (t) exp(β Z) = λ 0 (t) exp( β j Z j ) where β j is the parameter for the the
Lecture 7 Time-dependent Covariates in Cox Regression So far, we ve been considering the following Cox PH model: λ(t Z) = λ 0 (t) exp(β Z) = λ 0 (t) exp( β j Z j ) where β j is the parameter for the the
Pairwise rank based likelihood for estimating the relationship between two homogeneous populations and their mixture proportion
 Pairwise rank based likelihood for estimating the relationship between two homogeneous populations and their mixture proportion Glenn Heller and Jing Qin Department of Epidemiology and Biostatistics Memorial
Pairwise rank based likelihood for estimating the relationship between two homogeneous populations and their mixture proportion Glenn Heller and Jing Qin Department of Epidemiology and Biostatistics Memorial
Discussion of Identifiability and Estimation of Causal Effects in Randomized. Trials with Noncompliance and Completely Non-ignorable Missing Data
 Biometrics 000, 000 000 DOI: 000 000 0000 Discussion of Identifiability and Estimation of Causal Effects in Randomized Trials with Noncompliance and Completely Non-ignorable Missing Data Dylan S. Small
Biometrics 000, 000 000 DOI: 000 000 0000 Discussion of Identifiability and Estimation of Causal Effects in Randomized Trials with Noncompliance and Completely Non-ignorable Missing Data Dylan S. Small
Research Note: A more powerful test statistic for reasoning about interference between units
 Research Note: A more powerful test statistic for reasoning about interference between units Jake Bowers Mark Fredrickson Peter M. Aronow August 26, 2015 Abstract Bowers, Fredrickson and Panagopoulos (2012)
Research Note: A more powerful test statistic for reasoning about interference between units Jake Bowers Mark Fredrickson Peter M. Aronow August 26, 2015 Abstract Bowers, Fredrickson and Panagopoulos (2012)
Lawrence D. Brown* and Daniel McCarthy*
 Comments on the paper, An adaptive resampling test for detecting the presence of significant predictors by I. W. McKeague and M. Qian Lawrence D. Brown* and Daniel McCarthy* ABSTRACT: This commentary deals
Comments on the paper, An adaptive resampling test for detecting the presence of significant predictors by I. W. McKeague and M. Qian Lawrence D. Brown* and Daniel McCarthy* ABSTRACT: This commentary deals
Introduction to Empirical Processes and Semiparametric Inference Lecture 01: Introduction and Overview
 Introduction to Empirical Processes and Semiparametric Inference Lecture 01: Introduction and Overview Michael R. Kosorok, Ph.D. Professor and Chair of Biostatistics Professor of Statistics and Operations
Introduction to Empirical Processes and Semiparametric Inference Lecture 01: Introduction and Overview Michael R. Kosorok, Ph.D. Professor and Chair of Biostatistics Professor of Statistics and Operations
Alexina Mason. Department of Epidemiology and Biostatistics Imperial College, London. 16 February 2010
 Strategy for modelling non-random missing data mechanisms in longitudinal studies using Bayesian methods: application to income data from the Millennium Cohort Study Alexina Mason Department of Epidemiology
Strategy for modelling non-random missing data mechanisms in longitudinal studies using Bayesian methods: application to income data from the Millennium Cohort Study Alexina Mason Department of Epidemiology
Lecture Outline. Biost 518 Applied Biostatistics II. Choice of Model for Analysis. Choice of Model. Choice of Model. Lecture 10: Multiple Regression:
 Biost 518 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture utline Choice of Model Alternative Models Effect of data driven selection of
Biost 518 Applied Biostatistics II Scott S. Emerson, M.D., Ph.D. Professor of Biostatistics University of Washington Lecture utline Choice of Model Alternative Models Effect of data driven selection of
e author and the promoter give permission to consult this master dissertation and to copy it or parts of it for personal use. Each other use falls
 e author and the promoter give permission to consult this master dissertation and to copy it or parts of it for personal use. Each other use falls under the restrictions of the copyright, in particular
e author and the promoter give permission to consult this master dissertation and to copy it or parts of it for personal use. Each other use falls under the restrictions of the copyright, in particular
A NOTE ON ROBUST ESTIMATION IN LOGISTIC REGRESSION MODEL
 Discussiones Mathematicae Probability and Statistics 36 206 43 5 doi:0.75/dmps.80 A NOTE ON ROBUST ESTIMATION IN LOGISTIC REGRESSION MODEL Tadeusz Bednarski Wroclaw University e-mail: t.bednarski@prawo.uni.wroc.pl
Discussiones Mathematicae Probability and Statistics 36 206 43 5 doi:0.75/dmps.80 A NOTE ON ROBUST ESTIMATION IN LOGISTIC REGRESSION MODEL Tadeusz Bednarski Wroclaw University e-mail: t.bednarski@prawo.uni.wroc.pl
Introduction to Statistical Analysis
 Introduction to Statistical Analysis Changyu Shen Richard A. and Susan F. Smith Center for Outcomes Research in Cardiology Beth Israel Deaconess Medical Center Harvard Medical School Objectives Descriptive
Introduction to Statistical Analysis Changyu Shen Richard A. and Susan F. Smith Center for Outcomes Research in Cardiology Beth Israel Deaconess Medical Center Harvard Medical School Objectives Descriptive
Causal Effect Models for Realistic Individualized Treatment and Intention to Treat Rules
 University of California, Berkeley From the SelectedWorks of Maya Petersen March, 2007 Causal Effect Models for Realistic Individualized Treatment and Intention to Treat Rules Mark J van der Laan, University
University of California, Berkeley From the SelectedWorks of Maya Petersen March, 2007 Causal Effect Models for Realistic Individualized Treatment and Intention to Treat Rules Mark J van der Laan, University
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2015 Paper 334 Targeted Estimation and Inference for the Sample Average Treatment Effect Laura B. Balzer
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2015 Paper 334 Targeted Estimation and Inference for the Sample Average Treatment Effect Laura B. Balzer
Structural Nested Mean Models for Assessing Time-Varying Effect Moderation. Daniel Almirall
 1 Structural Nested Mean Models for Assessing Time-Varying Effect Moderation Daniel Almirall Center for Health Services Research, Durham VAMC & Dept. of Biostatistics, Duke University Medical Joint work
1 Structural Nested Mean Models for Assessing Time-Varying Effect Moderation Daniel Almirall Center for Health Services Research, Durham VAMC & Dept. of Biostatistics, Duke University Medical Joint work
Group Sequential Tests for Delayed Responses. Christopher Jennison. Lisa Hampson. Workshop on Special Topics on Sequential Methodology
 Group Sequential Tests for Delayed Responses Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj Lisa Hampson Department of Mathematics and Statistics,
Group Sequential Tests for Delayed Responses Christopher Jennison Department of Mathematical Sciences, University of Bath, UK http://people.bath.ac.uk/mascj Lisa Hampson Department of Mathematics and Statistics,
Simulation-based robust IV inference for lifetime data
 Simulation-based robust IV inference for lifetime data Anand Acharya 1 Lynda Khalaf 1 Marcel Voia 1 Myra Yazbeck 2 David Wensley 3 1 Department of Economics Carleton University 2 Department of Economics
Simulation-based robust IV inference for lifetime data Anand Acharya 1 Lynda Khalaf 1 Marcel Voia 1 Myra Yazbeck 2 David Wensley 3 1 Department of Economics Carleton University 2 Department of Economics
CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity
 CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity Prof. Kevin E. Thorpe Dept. of Public Health Sciences University of Toronto Objectives 1. Be able to distinguish among the various
CHL 5225H Advanced Statistical Methods for Clinical Trials: Multiplicity Prof. Kevin E. Thorpe Dept. of Public Health Sciences University of Toronto Objectives 1. Be able to distinguish among the various
Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness
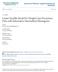 Journal of Modern Applied Statistical Methods Volume 15 Issue 2 Article 36 11-1-2016 Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness Li Qin Yale University,
Journal of Modern Applied Statistical Methods Volume 15 Issue 2 Article 36 11-1-2016 Latent Variable Model for Weight Gain Prevention Data with Informative Intermittent Missingness Li Qin Yale University,
2 Naïve Methods. 2.1 Complete or available case analysis
 2 Naïve Methods Before discussing methods for taking account of missingness when the missingness pattern can be assumed to be MAR in the next three chapters, we review some simple methods for handling
2 Naïve Methods Before discussing methods for taking account of missingness when the missingness pattern can be assumed to be MAR in the next three chapters, we review some simple methods for handling
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2010 Paper 260 Collaborative Targeted Maximum Likelihood For Time To Event Data Ori M. Stitelman Mark
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2010 Paper 260 Collaborative Targeted Maximum Likelihood For Time To Event Data Ori M. Stitelman Mark
Pubh 8482: Sequential Analysis
 Pubh 8482: Sequential Analysis Joseph S. Koopmeiners Division of Biostatistics University of Minnesota Week 8 P-values When reporting results, we usually report p-values in place of reporting whether or
Pubh 8482: Sequential Analysis Joseph S. Koopmeiners Division of Biostatistics University of Minnesota Week 8 P-values When reporting results, we usually report p-values in place of reporting whether or
Dose-response modeling with bivariate binary data under model uncertainty
 Dose-response modeling with bivariate binary data under model uncertainty Bernhard Klingenberg 1 1 Department of Mathematics and Statistics, Williams College, Williamstown, MA, 01267 and Institute of Statistics,
Dose-response modeling with bivariate binary data under model uncertainty Bernhard Klingenberg 1 1 Department of Mathematics and Statistics, Williams College, Williamstown, MA, 01267 and Institute of Statistics,
Statistics in medicine
 Statistics in medicine Lecture 4: and multivariable regression Fatma Shebl, MD, MS, MPH, PhD Assistant Professor Chronic Disease Epidemiology Department Yale School of Public Health Fatma.shebl@yale.edu
Statistics in medicine Lecture 4: and multivariable regression Fatma Shebl, MD, MS, MPH, PhD Assistant Professor Chronic Disease Epidemiology Department Yale School of Public Health Fatma.shebl@yale.edu
Unbiased estimation of exposure odds ratios in complete records logistic regression
 Unbiased estimation of exposure odds ratios in complete records logistic regression Jonathan Bartlett London School of Hygiene and Tropical Medicine www.missingdata.org.uk Centre for Statistical Methodology
Unbiased estimation of exposure odds ratios in complete records logistic regression Jonathan Bartlett London School of Hygiene and Tropical Medicine www.missingdata.org.uk Centre for Statistical Methodology
Variable selection and machine learning methods in causal inference
 Variable selection and machine learning methods in causal inference Debashis Ghosh Department of Biostatistics and Informatics Colorado School of Public Health Joint work with Yeying Zhu, University of
Variable selection and machine learning methods in causal inference Debashis Ghosh Department of Biostatistics and Informatics Colorado School of Public Health Joint work with Yeying Zhu, University of
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials
 Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Two-stage Adaptive Randomization for Delayed Response in Clinical Trials Guosheng Yin Department of Statistics and Actuarial Science The University of Hong Kong Joint work with J. Xu PSI and RSS Journal
Shu Yang and Jae Kwang Kim. Harvard University and Iowa State University
 Statistica Sinica 27 (2017), 000-000 doi:https://doi.org/10.5705/ss.202016.0155 DISCUSSION: DISSECTING MULTIPLE IMPUTATION FROM A MULTI-PHASE INFERENCE PERSPECTIVE: WHAT HAPPENS WHEN GOD S, IMPUTER S AND
Statistica Sinica 27 (2017), 000-000 doi:https://doi.org/10.5705/ss.202016.0155 DISCUSSION: DISSECTING MULTIPLE IMPUTATION FROM A MULTI-PHASE INFERENCE PERSPECTIVE: WHAT HAPPENS WHEN GOD S, IMPUTER S AND
Guideline on adjustment for baseline covariates in clinical trials
 26 February 2015 EMA/CHMP/295050/2013 Committee for Medicinal Products for Human Use (CHMP) Guideline on adjustment for baseline covariates in clinical trials Draft Agreed by Biostatistics Working Party
26 February 2015 EMA/CHMP/295050/2013 Committee for Medicinal Products for Human Use (CHMP) Guideline on adjustment for baseline covariates in clinical trials Draft Agreed by Biostatistics Working Party
Probability and Statistics
 Probability and Statistics Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be CHAPTER 4: IT IS ALL ABOUT DATA 4a - 1 CHAPTER 4: IT
Probability and Statistics Kristel Van Steen, PhD 2 Montefiore Institute - Systems and Modeling GIGA - Bioinformatics ULg kristel.vansteen@ulg.ac.be CHAPTER 4: IT IS ALL ABOUT DATA 4a - 1 CHAPTER 4: IT
Section on Survey Research Methods JSM 2010 STATISTICAL GRAPHICS OF PEARSON RESIDUALS IN SURVEY LOGISTIC REGRESSION DIAGNOSIS
 STATISTICAL GRAPHICS OF PEARSON RESIDUALS IN SURVEY LOGISTIC REGRESSION DIAGNOSIS Stanley Weng, National Agricultural Statistics Service, U.S. Department of Agriculture 3251 Old Lee Hwy, Fairfax, VA 22030,
STATISTICAL GRAPHICS OF PEARSON RESIDUALS IN SURVEY LOGISTIC REGRESSION DIAGNOSIS Stanley Weng, National Agricultural Statistics Service, U.S. Department of Agriculture 3251 Old Lee Hwy, Fairfax, VA 22030,
Fair Inference Through Semiparametric-Efficient Estimation Over Constraint-Specific Paths
 Fair Inference Through Semiparametric-Efficient Estimation Over Constraint-Specific Paths for New Developments in Nonparametric and Semiparametric Statistics, Joint Statistical Meetings; Vancouver, BC,
Fair Inference Through Semiparametric-Efficient Estimation Over Constraint-Specific Paths for New Developments in Nonparametric and Semiparametric Statistics, Joint Statistical Meetings; Vancouver, BC,
A noninformative Bayesian approach to domain estimation
 A noninformative Bayesian approach to domain estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu August 2002 Revised July 2003 To appear in Journal
A noninformative Bayesian approach to domain estimation Glen Meeden School of Statistics University of Minnesota Minneapolis, MN 55455 glen@stat.umn.edu August 2002 Revised July 2003 To appear in Journal
Pubh 8482: Sequential Analysis
 Pubh 8482: Sequential Analysis Joseph S. Koopmeiners Division of Biostatistics University of Minnesota Week 10 Class Summary Last time... We began our discussion of adaptive clinical trials Specifically,
Pubh 8482: Sequential Analysis Joseph S. Koopmeiners Division of Biostatistics University of Minnesota Week 10 Class Summary Last time... We began our discussion of adaptive clinical trials Specifically,
A Simple Way to Calculate Confidence Intervals for Partially Identified Parameters. By Tiemen Woutersen. Draft, September
 A Simple Way to Calculate Confidence Intervals for Partially Identified Parameters By Tiemen Woutersen Draft, September 006 Abstract. This note proposes a new way to calculate confidence intervals of partially
A Simple Way to Calculate Confidence Intervals for Partially Identified Parameters By Tiemen Woutersen Draft, September 006 Abstract. This note proposes a new way to calculate confidence intervals of partially
Published online: 10 Apr 2012.
 This article was downloaded by: Columbia University] On: 23 March 215, At: 12:7 Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number: 172954 Registered office: Mortimer
This article was downloaded by: Columbia University] On: 23 March 215, At: 12:7 Publisher: Taylor & Francis Informa Ltd Registered in England and Wales Registered Number: 172954 Registered office: Mortimer
Marginal Screening and Post-Selection Inference
 Marginal Screening and Post-Selection Inference Ian McKeague August 13, 2017 Ian McKeague (Columbia University) Marginal Screening August 13, 2017 1 / 29 Outline 1 Background on Marginal Screening 2 2
Marginal Screening and Post-Selection Inference Ian McKeague August 13, 2017 Ian McKeague (Columbia University) Marginal Screening August 13, 2017 1 / 29 Outline 1 Background on Marginal Screening 2 2
Prerequisite: STATS 7 or STATS 8 or AP90 or (STATS 120A and STATS 120B and STATS 120C). AP90 with a minimum score of 3
 University of California, Irvine 2017-2018 1 Statistics (STATS) Courses STATS 5. Seminar in Data Science. 1 Unit. An introduction to the field of Data Science; intended for entering freshman and transfers.
University of California, Irvine 2017-2018 1 Statistics (STATS) Courses STATS 5. Seminar in Data Science. 1 Unit. An introduction to the field of Data Science; intended for entering freshman and transfers.
Construction and statistical analysis of adaptive group sequential designs for randomized clinical trials
 Construction and statistical analysis of adaptive group sequential designs for randomized clinical trials Antoine Chambaz (MAP5, Université Paris Descartes) joint work with Mark van der Laan Atelier INSERM
Construction and statistical analysis of adaptive group sequential designs for randomized clinical trials Antoine Chambaz (MAP5, Université Paris Descartes) joint work with Mark van der Laan Atelier INSERM
Dynamic Prediction of Disease Progression Using Longitudinal Biomarker Data
 Dynamic Prediction of Disease Progression Using Longitudinal Biomarker Data Xuelin Huang Department of Biostatistics M. D. Anderson Cancer Center The University of Texas Joint Work with Jing Ning, Sangbum
Dynamic Prediction of Disease Progression Using Longitudinal Biomarker Data Xuelin Huang Department of Biostatistics M. D. Anderson Cancer Center The University of Texas Joint Work with Jing Ning, Sangbum
University of California, Berkeley
 University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2009 Paper 248 Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials Kelly
University of California, Berkeley U.C. Berkeley Division of Biostatistics Working Paper Series Year 2009 Paper 248 Application of Time-to-Event Methods in the Assessment of Safety in Clinical Trials Kelly
BIAS OF MAXIMUM-LIKELIHOOD ESTIMATES IN LOGISTIC AND COX REGRESSION MODELS: A COMPARATIVE SIMULATION STUDY
 BIAS OF MAXIMUM-LIKELIHOOD ESTIMATES IN LOGISTIC AND COX REGRESSION MODELS: A COMPARATIVE SIMULATION STUDY Ingo Langner 1, Ralf Bender 2, Rebecca Lenz-Tönjes 1, Helmut Küchenhoff 2, Maria Blettner 2 1
BIAS OF MAXIMUM-LIKELIHOOD ESTIMATES IN LOGISTIC AND COX REGRESSION MODELS: A COMPARATIVE SIMULATION STUDY Ingo Langner 1, Ralf Bender 2, Rebecca Lenz-Tönjes 1, Helmut Küchenhoff 2, Maria Blettner 2 1
A Flexible Bayesian Approach to Monotone Missing. Data in Longitudinal Studies with Nonignorable. Missingness with Application to an Acute
 A Flexible Bayesian Approach to Monotone Missing Data in Longitudinal Studies with Nonignorable Missingness with Application to an Acute Schizophrenia Clinical Trial Antonio R. Linero, Michael J. Daniels
A Flexible Bayesian Approach to Monotone Missing Data in Longitudinal Studies with Nonignorable Missingness with Application to an Acute Schizophrenia Clinical Trial Antonio R. Linero, Michael J. Daniels
Logistic regression model for survival time analysis using time-varying coefficients
 Logistic regression model for survival time analysis using time-varying coefficients Accepted in American Journal of Mathematical and Management Sciences, 2016 Kenichi SATOH ksatoh@hiroshima-u.ac.jp Research
Logistic regression model for survival time analysis using time-varying coefficients Accepted in American Journal of Mathematical and Management Sciences, 2016 Kenichi SATOH ksatoh@hiroshima-u.ac.jp Research
Applied Multivariate and Longitudinal Data Analysis
 Applied Multivariate and Longitudinal Data Analysis Chapter 2: Inference about the mean vector(s) Ana-Maria Staicu SAS Hall 5220; 919-515-0644; astaicu@ncsu.edu 1 In this chapter we will discuss inference
Applied Multivariate and Longitudinal Data Analysis Chapter 2: Inference about the mean vector(s) Ana-Maria Staicu SAS Hall 5220; 919-515-0644; astaicu@ncsu.edu 1 In this chapter we will discuss inference
Figure 36: Respiratory infection versus time for the first 49 children.
 y BINARY DATA MODELS We devote an entire chapter to binary data since such data are challenging, both in terms of modeling the dependence, and parameter interpretation. We again consider mixed effects
y BINARY DATA MODELS We devote an entire chapter to binary data since such data are challenging, both in terms of modeling the dependence, and parameter interpretation. We again consider mixed effects
Survival Analysis for Case-Cohort Studies
 Survival Analysis for ase-ohort Studies Petr Klášterecký Dept. of Probability and Mathematical Statistics, Faculty of Mathematics and Physics, harles University, Prague, zech Republic e-mail: petr.klasterecky@matfyz.cz
Survival Analysis for ase-ohort Studies Petr Klášterecký Dept. of Probability and Mathematical Statistics, Faculty of Mathematics and Physics, harles University, Prague, zech Republic e-mail: petr.klasterecky@matfyz.cz
High-Throughput Sequencing Course
 High-Throughput Sequencing Course DESeq Model for RNA-Seq Biostatistics and Bioinformatics Summer 2017 Outline Review: Standard linear regression model (e.g., to model gene expression as function of an
High-Throughput Sequencing Course DESeq Model for RNA-Seq Biostatistics and Bioinformatics Summer 2017 Outline Review: Standard linear regression model (e.g., to model gene expression as function of an
Discrete Response Multilevel Models for Repeated Measures: An Application to Voting Intentions Data
 Quality & Quantity 34: 323 330, 2000. 2000 Kluwer Academic Publishers. Printed in the Netherlands. 323 Note Discrete Response Multilevel Models for Repeated Measures: An Application to Voting Intentions
Quality & Quantity 34: 323 330, 2000. 2000 Kluwer Academic Publishers. Printed in the Netherlands. 323 Note Discrete Response Multilevel Models for Repeated Measures: An Application to Voting Intentions
Model Selection in GLMs. (should be able to implement frequentist GLM analyses!) Today: standard frequentist methods for model selection
 Model Selection in GLMs Last class: estimability/identifiability, analysis of deviance, standard errors & confidence intervals (should be able to implement frequentist GLM analyses!) Today: standard frequentist
Model Selection in GLMs Last class: estimability/identifiability, analysis of deviance, standard errors & confidence intervals (should be able to implement frequentist GLM analyses!) Today: standard frequentist
Learning the Semantic Correlation: An Alternative Way to Gain from Unlabeled Text
 Learning the Semantic Correlation: An Alternative Way to Gain from Unlabeled Text Yi Zhang Machine Learning Department Carnegie Mellon University yizhang1@cs.cmu.edu Jeff Schneider The Robotics Institute
Learning the Semantic Correlation: An Alternative Way to Gain from Unlabeled Text Yi Zhang Machine Learning Department Carnegie Mellon University yizhang1@cs.cmu.edu Jeff Schneider The Robotics Institute
Introduction. Chapter 1
 Chapter 1 Introduction In this book we will be concerned with supervised learning, which is the problem of learning input-output mappings from empirical data (the training dataset). Depending on the characteristics
Chapter 1 Introduction In this book we will be concerned with supervised learning, which is the problem of learning input-output mappings from empirical data (the training dataset). Depending on the characteristics
Individualized Treatment Effects with Censored Data via Nonparametric Accelerated Failure Time Models
 Individualized Treatment Effects with Censored Data via Nonparametric Accelerated Failure Time Models Nicholas C. Henderson Thomas A. Louis Gary Rosner Ravi Varadhan Johns Hopkins University July 31, 2018
Individualized Treatment Effects with Censored Data via Nonparametric Accelerated Failure Time Models Nicholas C. Henderson Thomas A. Louis Gary Rosner Ravi Varadhan Johns Hopkins University July 31, 2018
Causality II: How does causal inference fit into public health and what it is the role of statistics?
 Causality II: How does causal inference fit into public health and what it is the role of statistics? Statistics for Psychosocial Research II November 13, 2006 1 Outline Potential Outcomes / Counterfactual
Causality II: How does causal inference fit into public health and what it is the role of statistics? Statistics for Psychosocial Research II November 13, 2006 1 Outline Potential Outcomes / Counterfactual
BIOL 51A - Biostatistics 1 1. Lecture 1: Intro to Biostatistics. Smoking: hazardous? FEV (l) Smoke
 BIOL 51A - Biostatistics 1 1 Lecture 1: Intro to Biostatistics Smoking: hazardous? FEV (l) 1 2 3 4 5 No Yes Smoke BIOL 51A - Biostatistics 1 2 Box Plot a.k.a box-and-whisker diagram or candlestick chart
BIOL 51A - Biostatistics 1 1 Lecture 1: Intro to Biostatistics Smoking: hazardous? FEV (l) 1 2 3 4 5 No Yes Smoke BIOL 51A - Biostatistics 1 2 Box Plot a.k.a box-and-whisker diagram or candlestick chart
AFT Models and Empirical Likelihood
 AFT Models and Empirical Likelihood Mai Zhou Department of Statistics, University of Kentucky Collaborators: Gang Li (UCLA); A. Bathke; M. Kim (Kentucky) Accelerated Failure Time (AFT) models: Y = log(t
AFT Models and Empirical Likelihood Mai Zhou Department of Statistics, University of Kentucky Collaborators: Gang Li (UCLA); A. Bathke; M. Kim (Kentucky) Accelerated Failure Time (AFT) models: Y = log(t
